 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
A Bayesian approach providing design choices and chemical insight for solution-processed thermoelectric polymers
Connor
Ganley
 *a,
Yuqi
Feng
a,
Himadri Shekhar
Karmakar
b,
Howard E.
Katz
*a,
Yuqi
Feng
a,
Himadri Shekhar
Karmakar
b,
Howard E.
Katz
 b and
Paulette
Clancy
b and
Paulette
Clancy
 *a
*a
aDepartment of Chemical and Biomolecular Engineering, Johns Hopkins University, Baltimore, Maryland, USA. E-mail: cganley2@jhu.edu; pclancy3@jhu.edu
bDepartment of Materials Science and Engineering, Johns Hopkins University, Baltimore, Maryland, USA
First published on 20th February 2026
Abstract
The advent of machine learning approaches to materials design is poised to lead experimental validation by selecting a much smaller, more promising, and sometimes unexpected, set of new materials candidates. This work describes one such investigation to characterize novel, hypothetical, and promising polymer candidates based on their calculated molecular-scale electronic and chemical parameters. Subsequently, we optimize them for a chosen chemical property using a chemically informed Bayesian surrogate model. As a test case, over 7300 combinations of novel, hypothetical semiconducting diketopyrrolopyrrole-based (DPP) polymers and commonly used solvents, generated by density functional theory, were screened based on their free energies of solvation, Gsolv, as a guide to selecting optimal processing conditions. From this synthetic data set, we trained a physics-informed Gaussian process model that linked molecular-scale electronic structure properties to Gsolv, and then used Bayesian optimization (BO) to identify key descriptors for predicting optimal enthalpic, single-chain polymer solvation in the infinitely dilute limit. As a result, we predicted an “optimal” solvent dielectric constant value around 10 for the DPP-based polymer class. To validate this result, we showed that the predicted polymer design associated with the minimum ΔGsolv value corresponded to the polymer with the highest experimentally measured conductivity in the literature. The importance of these observations is that our BO approach provided the chemical insight necessary to quickly screen solvents for potential DPP-based polymers based on the polymer repeat unit's quadrupole moments and to identify preferred compatible solvents corresponding to a minimized ΔGsolv. This study also highlights the effectiveness of our BO algorithm: The optimal polymer design was found using just 6% of the parameter space and in a very short execution time (on the order of minutes), a feat which cannot be duplicated experimentally. Finally, to show the extensibility of this machine learning approach, we repeated this exercise with a second class of semiconducting polymers in which the DPP base group is replaced by an indacenodithiophene (IDT) group. We again successfully validated our machine learning predictions of the most promising polymer designs against experimental results.
1 Introduction
Organic semiconducting polymers constitute a large class of materials with a wide range of electronic applications. They can be used in field-effect transistors (FETs), organic solar cells, light-emitting diodes (LEDs) and thermoelectrics, among many others.1–4 These materials have the advantages of being lightweight, solution-processable,5 and exhibiting highly tunable band gaps.6,7 The band gap is a consequence of the theoretically infinite chemical and configurational space within which a polymer can be designed. Rational design within this space is an active area of research.8–10 For example, the highest power conversion efficiency (PCE) of an organic solar cell is now 19.2%, a metric which has seen ∼50% improvement over the past decade.11Polymer design approaches that appear in the literature include functional group engineering,9,10 single-atom substitution,12 solvent engineering,4,13,14 and doping.15–17 More recently, machine learning (ML)-based approaches have accelerated the rate at which we can screen the otherwise daunting numbers of potential organic semiconductor candidates. This can be done in silico with tools like density functional theory (DFT), for example, to subsequently inform experimental design.18–22 Indeed, Gao et al. identified AI-driven approaches as a fourth research paradigm and describe a molecular fingerprint informed by topological descriptors.23 Older large-scale databases like the Harvard Clean Energy Project24 and the Cambridge Structural Database25 serve as useful archives for a portion of the infinite chemical design space spanned by conjugated organic molecules. Despite efforts to tabulate the properties of millions of materials, there are many more candidate materials that remain undiscovered.
Semiconducting polymers can be constructed from numerous building blocks (e.g., functional groups) linked together via a variety of accessible bond-forming reactions so that the building blocks become electronically conjugated, either randomly or in short, repetitive sequences. For example, a repetitive sequence can be an alternation of a donor and an acceptor building block. Yan et al. characterized the efficacy of a diketopyrrolopyrrole (DPP)-derivative building block in an n-doped organic semiconductor, employing a donor–acceptor motif.3 In a similar vein, Ren et al. characterized a DPP-based thin-film transistor (TFT), also employing a donor–acceptor motif.9 The repetitive sequence gives the resulting polymer its tailorable electronic properties: the chosen donor and acceptor units determine the polymer's highest occupied molecular orbital (HOMO) and lowest unoccupied molecular orbital (LUMO).
Mukhopadhyaya et al. also utilized a common DPP building block and varied the nature of the functional groups surrounding it to produce a set of p-doped thermoelectric polymers. They chose this DPP-containing class of polymers to optimize chemical stability, molecular ordering, and electronic energy levels for thermoelectric applications. They identified polymer-dopant combinations with high-performing conductivities (σ) of up to 700 S cm−1.26,27 Inspired by that approach, we take advantage of the known capability to link and conjugate building blocks to propose numerous permutations, with the confidence that synthetic methods to prepare most of the permutations would be available.
There is a well-known relationship between the behavior of a polymer in solution and the thin-film morphology of the resultant device.5,28–30 Thus, it is important to quantify the complex relationship between the structure of a polymer and its solution (and ultimately, morphological) behavior. Identifying an “optimal” solvent for a class of polymers is a useful construct because it creates a high probability of successful processing from the start of work on that class. For the present case of the commonly used donor–acceptor polymer architecture, there is a characteristic subunit that determines a significant portion of the electronic properties of that polymer system (i.e. DPP versus IDT). Therefore, the addition of subunits peripheral to this characteristic subunit would only cause relatively small modulations of the electronic properties, so an “optimal” solvent would correspond to both this characteristic subunit while still maintaining the desired property objectives.
Dhabal et al. made a step toward this goal using a so-called “united-atom” (or coarse-grained) approach to describe polymer conformation upon desolvation.31 But this relies on the fact that this simplified approximation is a good representation of the complex potential energy landscape that arises due to quantum mechanical interactions. Coarse-graining can also suffer from removing the cis-/trans-ordering that defines the morphology, planarity, and hence charge transport that are a key target for this class of semiconducting polymers.32
This work uses a “building block” polymer design approach to computationally screen a combination of 324 hypothetical DPP-based donor–acceptor copolymers in the presence of 26 solvents. We generated density functional theory (DFT) data characterizing the molecular electronic properties of all 8424 combinations of possible copolymer–solvent pairs as our synthetic database. Then we used a BO algorithm, the Physical Analytics pipeLine (PAL) 2.0, that we have developed as a means to identify the optimal design of polymeric building blocks.33 We can vary the combination of polymer building blocks in order to make a set of unique synthetic polymer candidates, each of which might lead to a minimized free energy of solvation, (ΔGsolv). We have found the PAL 2.0 code base to be successful in materials discovery scenarios for a range of applications.34
We discovered that one of the parameters, the repeat unit's quadrupole moment, was highly correlated to ΔGsolv. This parameter, in turn, was retrospectively found to be associated with the highest conductivity composition found in the study by Mukhopadhyaya et al., validating our machine learning predictions.27 We tested the extensibility of our model to polymers with a different basic core group, here, based on the indacenodithiophene (IDT) unit, which is actively being studied for semiconducting applications.35,36 Experimental observations on the solubility of indacenodithiophene-co-benzothiadiazole (IDT-BT) in various solvents with dielectric constants ranging from 2–80 allowed us to validate our findings that a solvation “minimum” exists for a given design at a resulting dielectric constant.
2 Methods
In this study, we defined a set of diketopyrrolopyrrole (DPP)-based p-type semiconducting polymers from commonly used functional group sub-units according to the backbone-ordering motif shown in Fig. 1.27,28 The polymers were segmented into discrete sites, the second of which was always occupied by methyl-terminated DPP in all cases. The remaining four backbone sites (labeled A, B, and C) can be occupied by any of the functional groups shown in the boxes below the polymer motif in Fig. 1. The “A” site is situated on either side of DPP to account for synthesizability considerations, and the “A” functional groups are fewer than “B” or “C” so that the larger functional groups would not sterically interact with the DPP. The IUPAC names of the available functional groups and their associated abbreviations used throughout the text are shown in Fig. S1.Four functional group candidate choices for the A sites and nine functional group choices for both B and C create a combinatorial space of 324 unique polymer candidates. Note that any number of potential motifs can, in principle, be used to construct these hypothetical polymers; we chose to design our polymer using four different motifs arbitrarily. Each of the 324 possibilities we considered has different molecular electronic parameters, which we characterized by performing DFT calculations. For the DFT calculations, we elected to use the B97-D3 functional, as recommended by Goerigk et al. for main-group non-covalent interactions at the GGA level of theory with dispersion corrections. We combined this functional with the accurate def2-TZVP basis set that Oliveira et al. found to effectively balance accuracy and computational cost.37,38
A given polymer repeat unit constructed in this way is theoretically capable of being polymerized, processed, and cast into a thin-film device in conjunction with any number of potential solvents. But some solvents are known to solvate a polymer more effectively than others due to the complex relationship between the chemical properties of solute and solvent. It is this processing aspect of materials design that we wanted to explore. Due to the resources required for modeling the electronic structures of up to hundreds of atoms simultaneously, we represented each polymer as its repeat unit, i.e., n = 1 in Fig. 1. We thus calculated the solvation free energy change (ΔGsolv) for all 324 polymer repeat units immersed in 26 different implicitly modeled solvents using DFT. This yielded a final design space of 8424 possible polymer–solvent interactions. We acknowledge that it is a limitation of our work to have studied single-chain, infinitely-dilute, enthalpic solvation of these polymers. Moreover, due to the structural simplifications necessary for high-fidelity electronic structure modeling within a reasonable time and computational budget, our calculations do not include inter- or intra-chain interaction effects, entanglement molecular weight effects, free-volume contributions, or kinetic constraints to solvation, all of which affect the dissolution process, but at a much larger scale than is feasible to model with DFT. We envision these effects could be the subject of future studies that utilize coarse-grained or classical molecular dynamics (MD) with a suitable interatomic potential. However, the focus of the present study and its general value is on the electrostatic contributions of functional groups within polymer repeat units, and which are the starting point of polymer design. The insight gained herein should therefore be used as a single tool in an arsenal of multi-scale tools for computational polymer design, which can save experimental time and cost with high-throughput screening a priori.
There are many different ways to represent a chemical structure in model space. Commonly used descriptor libraries include Dragon, Mordred, and RDKit.39–41 However, the descriptors from these libraries are primarily topology-based: they only include the connectivity and some basic composition data. Pereira et al. found that including DFT-calculated orbital energy data in addition to connectivity descriptors improved their model's accuracy of predicting HOMO and LUMO energies.42 Understanding the value of electronic structure data, we identified some features characterizing the design space that effectively encode expert chemical knowledge into the training set. A common heuristic for solvation says that “like dissolves like”. Even though this is a simplification of a very complex process, the essence of this dictum holds. The electrostatic properties (governed by the electron density) of the polymer repeat unit and solvent pair must be similar in order to optimize the free energy of solvation which, in turn, will speak to its thermodynamic stability. To this end, we elected to represent our polymers and solvents with the descriptors shown in Table 1.
| Polymer repeat unit | Solvent |
|---|---|
| Isotropic quadrupole | Isotropic quadrupole |
| Isotropic polarizability | Isotropic polarizability |
| ESPmin | ESPmin |
| ESPmax | ESPmax |
| Polarity index | Polarity index |
| % polar surface area | % polar surface area |
| HOMO–LUMO gap | Dielectric constant |
These descriptors relate to the distribution of charge around the respective species and are readily calculable with the ORCA DFT software package,43,44 or using a post-processing software recommended in ORCA's documentation called Multiwfn.45 In Table 1, “ESP” stands for the “electrostatic potential”, and the molecular polarity index (MPI) is a normalization across the area of the autocorrelation of ESP charge over the van der Waals (vdW) surface at a constant potential value, in this case the default value ρ = 0.001 a.u.46 The equation for calculating MPI is:
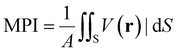 | (1) |
While we did not know, a priori, if the properties listed in Table 1 would correlate with ΔGsolv, it was reasonable to assume that these properties, which relate to electron distribution, might be correlated to solvation, which is governed by polarizable molecular interactions. Test models that included purely topological descriptors, such as those available in the cheminformatics packages mentioned above, did not yield significant correlation to ΔGsolv. In this way, encoding electronic structure data in the descriptors improves model prediction.
The universal solvation model (SMD) is available within ORCA and was employed to calculate the free energy of solvation. According to the SMD model, the solvation free energy can be decomposed into a bulk electrostatic contribution (ENP) and a cavitation dispersion energy (CDS), according to eqn (2).47
| ΔGsolv = ΔGENP + ΔGCDS | (2) |
The SMD model calculates these values by assigning a uniform dielectric constant across the simulation medium that is commensurate with the solvent of interest. The ENP and CDS terms calculated are therefore dependent on the dielectric constant of the solvent and the electron distribution within the polymer.
As a basis of comparison, we also performed implicit solvation calculations of the polymer repeat units according to the conductor-like polarizable continuum model (CPCM), which is available in the ORCA DFT code and which closely follows the methodology described in the seminal CPCM paper by Barone and Cossi.48 In short, the molecular Hamiltonian of the isolated molecular system is perturbed by the solvent according to eqn (3),
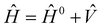 | (3) |
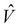 represents the solute–solvent interaction term and is incorporated into the self-consistent field (SCF) procedure to variationally minimize the free energy of the solute.
represents the solute–solvent interaction term and is incorporated into the self-consistent field (SCF) procedure to variationally minimize the free energy of the solute.
3 Results and discussion
Of the 8424 possible combinations of polymer and implicit solvent, 7300 of the 8424 DFT relaxations converged readily and comprised the primary data set. The largest polymer repeat unit contained 136 atoms, so it is possible that the non-converged simulations required more computational resources, especially memory, than were available. Nonetheless, the data we acquired were sufficient to allow us to sample the objective function space and identify relevant variable correlations. The resulting distributions of ΔGsolv for different solvent choices were arranged in ascending values of the dielectric constant for these 7300 data points, as shown in Fig. 2. To properly present this huge amount of data, we chose the box plot. However, remember that, for each solvent, we analyzed the ΔGsolv of hundreds of polymer structures. Thus, the bars on each solvent do not represent error bars and indicate the difference in free energies represented by the polymer choice having the highest and lowest ΔGsolv. For each solvent, the blue box represents the distribution of ΔGsolv of different polymer structures. Consequently, this figure suggests that there exists a minimum ΔGsolv and, therefore, an “optimal” solvent for a given polymer class, in this case DPP-based. | ||
| Fig. 2 Box-plot distributions of DFT-calculated values of the Gibbs free energy of solvation for 324 DPP-based polymers in 26 implicitly modeled solvents. | ||
ΔGsolv is a function of the polymer's composition, and each functional group sub-unit of the polymer contributes differently to its overall electronic density distribution. Accordingly, we sorted the 324 polymer candidates into top, middle, and bottom quantiles (thirds) according to two metrics: polarity index and isotropic quadrupole moment. These were chosen for comparison because they both describe a single value that distills the electronic distribution around the polymer repeat unit (on a per-area or per-direction basis). Then we evaluated their relative correlation to the target value. Fig. 3 compares the average value of ΔGsolv calculated by DFT simulations of the most polar (filled circles) and least polar (empty circles) polymers, as sorted by the two indicated properties. The results show that there is a relatively large, 0.5 eV, difference in the magnitude of ΔGsolv between the solvation of polymers with the highest and lowest quadrupole moments (B). As expected, polymers with higher quadrupole moments are better solvated in general, even by ostensibly “low” dielectric solvents. In contrast, the polarity index (A) does not demonstrate a significant variation in ΔGsolv, suggesting that it may not be an important property, at least in this class of DPP-based polymers. The quadrupole moment is traditionally thought to be structure-directing, as seems to be borne out here. The identification of quadrupole moment as a modulating factor for ΔGsolv is reasonable, given that multipoles serve as an approximation for charge density distribution in density-based solvation models like SMD.47 It is likely that the spatial information, encoded in the quadrupole matrix, leads to its utility over a non-spatial descriptor property like the polarity index, even though it is made to be isotropic to enter the dataset as a single value. The isotropic quadrupole was used instead of the largest eigenvalue of the quadrupole matrix because we assumed that, in a liquid system, the directional dependence of the largest eigenvalue would average out over time such that the isotropic value was more intrinsic to the system at hand. Moreover, this orientation-independent parameter provided a singular basis on which to compare the different polymer repeat units, which varied significantly in their structure, without needing to consider the direction of the largest quadrupole eigenvalue.
A frequency analysis of the sorted quantiles revealed further insight into the structure–property relationships exhibited by the polymer candidates. Fig. 4 shows a comparison of the most frequently encountered functional group subunits for the most (A) and least (B) polar polymers according to their isotropic quadrupole moment. The most frequently appearing functional groups in high-polarity polymers are TT, EDOT, and B3TA, the structures of which are shown in Fig. S1. The presence and configuration of polar atoms, like S and O, in these groups contributes to the overall polarity of the polymer. On the other hand, the most frequently appearing functional groups in the least polar polymers are benzene and thiophene, which are known for their non-polar character.
 | ||
| Fig. 4 Frequency analysis of functional unit subgroups of the highest (A) and lowest (B) quantiles as predicted by PAL. | ||
A component analysis of these data revealed that the isotropic quadrupole of the overall polymer can be described as a linear combination of the isotropic quadrupoles of its individual components to within an error of <7%, under-predicting in all cases. This suggests that there exists a small quadrupole interaction effect as more functional group subunits are added along the conjugated axis. This analysis has implications for polymer design: we found that modeling only the electronic properties of functional group subunits provides a reasonably accurate way to model the composite polymer's electronic properties, at least for the isotropic quadrupole moment. This can cut down immensely on the computational resources required for in silico property prediction.
To validate the results shown in Fig. 2 for implicit solvents that assume a “mean field”-like representation, we used a more accurate explicit representation of the solvent molecule in which every atom in the functional group was modeled. For these explicitly modeled solvents, we chose to study the binding enthalpy of just four of the original 26 solvents to interact with all 324 polymer repeat units to act as representative cases and to keep computational resource-use within reason. We placed a single, atomically explicit solvent molecule 3 Å away from the plane spanned by each of the polymer choices. The system was allowed to relax within the DFT calculation of the polymer–solvent pair. The solvents chosen were n-hexane, ortho-dichlorobenzene, acetonitrile, and water, to cover a broad range of dielectric constants, (ε), with values 1.88, 9.99, 35.69 and 78.36, respectively.47
The results of the relaxed interaction enthalpies are shown in Fig. 5. Each row (A–D) in this Figure shows the distribution of binding enthalpies for all 324 polymers with a single molecule of each respective solvent. The binding enthalpy was calculated with respect to the individual component species, where a negative value for the binding enthalpy indicates greater enthalpic preference for the bound state over the isolated states.
The polymers we tested exhibit a greater “stability” in the presence of solvents with a lower dielectric constant. The average binding enthalpies, in order of increasing solvent dielectric constant, of the four explicitly modeled solvents are: −0.29 eV, −0.29 eV, −0.16 eV, and −0.14 eV. Less polar solvents are predicted to solvate the 324 polymer candidates better than more polar ones. There is a correlation with the results in Fig. 2 in that ortho-dichlorobenzene, with ε ∼ 9.99, exhibits both the highest (negative) magnitude ΔGsolv and the greatest binding enthalpy. A direct comparison of this correlation is displayed in Fig. S9. This suggests that this is an “optimal” solvent range for this set of polymers.
An outcome of this validation is the similarity of results using “implicit solvent” models, like the conductor-like polarizable continuum model (CPCM) or the universal solvation model (SMD), with those of the explicitly modeled solvents. This confirms that accurate and valuable insight into solvation can be obtained using implicit solvent modeling at a fraction of the cost of explicit solvation modeling. It is possible that, with greater sampling of the polymer–explicit solvent interaction space than the four test cases explicitly modeled here, there would be an even stronger correlation. We note that the lower magnitude binding enthalpy between the highly polar solvents (water and acetonitrile) could arise because the most favorable binding sites are highly localized, whereas the polar binding sites available on the polymer are more diffuse. There are other experimental considerations at play, such as dopant miscibility, which affect solvent choice and were not considered in this study.
3.1 Machine-learned feature engineering
A more systematic approach to elucidate the quantitative nature of structure–property relationships in these polymers was conducted on the implicit solvation data using the “feature engineering” capability of PAL 2.0, a physics-informed Bayesian framework for materials discovery developed by the Clancy group.33 Using XGBoost, a gradient-based machine learning algorithm for feature selection,49 and the Least Absolute Shrinkage and Selection Operator (LASSO), a method for estimation of linear models,50 we obtained a ranked list of the most important descriptors (features), as shown in Table 1, for quantifying a target material property, in this case, ΔGsolv.Feature correlation is addressed through the feature selection step, using the well-known XGBoost technique. During tree construction, correlated features compete to reduce the loss, and the algorithm typically selects one representative feature from a correlated set for early ‘splits’. Once such a feature is selected, additional correlated features tend to provide diminished marginal gain and consequently receive lower importance scores. As a result, highly correlated features are often down-weighted or excluded during importance-based ranking. This results in one of the highly correlated features being selected as important and the others being pruned from the model. In this case, this method results in one of the correlated features being arbitrarily selected over the others, but that this does not affect interpretability at all as we know how these features are correlated and can still properly attribute physical meaning to the values selected.
Fig. 6 shows the relative importance of each model descriptor as determined by the labeled algorithms. Both LASSO and XGBoost models identify the polymer's isotropic quadrupole moment as a descriptor with a high relative importance. This supports the observation of a different trend in ΔGsolv for low-quadrupole versus high-quadrupole polymers exhibited in Fig. 3. Both models also identify the relative importance of the explicit polymer–solvent interaction, providing further evidence for the behavior observed in Fig. 5 and the ensuing commentary on the value of implicit versus explicit models of solvent interactions. A key difference between LASSO and XGBoost is that the former employs a linear model while the latter is better suited for non-linear modeling. This could explain the disparity in features identified in Fig. 6. LASSO identifies numerous descriptors with comparable importance, whereas XGBoost clearly identifies the solvent dielectric (ε) as the most important feature. It could be that LASSO simply needs more terms to be able to describe the input–output relationship; it cannot penalize too many terms in order to maintain accurate predictions. XGBoost, on the other hand, may be identifying the same non-linear dependence on ε that is shown in Fig. 3. Within the vast polymer chemical space that exists, feature engineering has identified useful metrics to help uncover the complex relationship between polymer solutes and solvents.
 | ||
| Fig. 6 Relative descriptor importance as quantified by two different machine learning techniques, LASSO (A) and XGBoost (B). | ||
An “expert” surrogate Gaussian process (GP) model using the features identified by XGBoost was created from a 80/20 train/test split within the PAL 2.0 framework. The Matérn 5/2 kernel was used in the Gaussian Process surrogate model. The features are concatenated as an input vector, and there exists a single length scale for the kernel as it operates on the input vectors. With each BO iteration, this length scale hyperparameter is optimized. Fig. 7 shows that BO using the Expected Improvement (EI) acquisition function converged to the surrogate model's optimal value (minimum ΔGsolv) within 25 evaluations, on average, of the objective function for a Gaussian Process (GP) model with a linear mean prior, and within 50 evaluations for the GP with a zero mean prior.51 The difference in the number of function evaluations needed here shows that a GP with a linear mean prior is a better description of the functional behavior than one with a zero mean prior. In the distribution of GP models trained for this task, the linear mean prior GP model with the slowest optimization time (in terms of objective function evaluations) needed to explore only 6% of the compositional space, whereas the slowest zero mean prior GP model required more than 12% (See Fig. S3).
 | ||
| Fig. 7 Function evaluations of GP models with a zero mean prior (blue) and a linear mean prior (red). Shaded regions indicate model uncertainty. | ||
These results further support the hypothesis that the dielectric constant of the solvent is a sufficient description of the variation observed in ΔGsolv. In this way, the Gaussian Process surrogate model acts as reinforcement for the hypothesis that a complex process, like polymer solvation, is, in fact, dependent on very few physically realizable variables.
3.2 Validation from experimental results
To validate the worth of the Bayesian-derived suggestions of potentially high-performing semiconducting doped polymers, we looked to the work of Mukhopadhyaya et al., who studied the properties of four different DPP-based semiconducting polymers. Their DPP-based polymers were doped with one of three dopants: F4TCNQ, a [3]-radialene-based dopant Cp(CN)3-(COOMe)3, and trifluoromethanesulfonic acid CF3SO3H.27 They found that a polymer containing a DPP group, two EDOT groups, and two MEET groups exhibited the highest conductivity of the polymers that they studied when doped with a sulfonic acid dopant. Although it is thought that long-range crystallinity is a property critical to higher electrical conductivity,52 the XRD data showed that this polymer-dopant combination exhibited “virtually no long-range crystallinity”.27 We hypothesize that the high relative conductivity of this polymer-dopant thin film could be due to more favorable solvation leading to a degree of, at the very least, short-range order. Insight into any potential short-range order, in line with our suggestion, would require higher resolution characterization techniques.In Mukhopadhyaya's paper, solution-doping was achieved by combining a polymer, dissolved in chlorobenzene (ε = 5.7), with a dopant, dissolved in a different solvent, acetonitrile (ε = 35.7). During mixing, the dielectric constant of the solution naturally changed, based on the desired dopant concentration, and in the range between 5 and 50 dopant mol%. At 5 mol% dopant concentration, the dielectric constant of the solvent was 6.3; at 50 mol%, it was 8.4. Referring back to the information obtained from Fig. 2 (implicit solvent calculations) and 5 (explicit solvent calculations) that suggested a solvation free energy minimum will occur at a dielectric constant value, ε, around 10, the experiments are clearly operating at a close-to-optimal dielectric constant that would be recommended by theoretical calculations. Post facto, we learned that Imbrogno et al.'s experiments found an ionic conductivity maximum at a polymer dielectric constant ε ∼ 9 for various poly(vinyl ethers) (PVE) and they observed a plateau in ionic conductivity at ε above 9.53 Both of these experimental observations offer very similar outcomes to those we observed via computational studies in this work.
It is possible that the increase in conductivity with dopant mol% observed in experiments could be due to the augmented presence of acetonitrile, which drives the solution's overall dielectric constant closer to the minimum. Moreover, comparison of the relative magnitudes of ΔGsolv among the four polymers studied by Mukhopadhyaya et al. shows that the doped polymer with the highest conductivity also exhibited the highest magnitude of solvation free energy (see Fig. S2). It is possible that these factors contributed to favorable solvation behavior, and therefore some degree of short-range order in the polymer thin film.
3.3 Extensibility to other polymer classes: IDT
To determine the extensibility of our approach, we conducted an identical computational analysis for IDT-based polymers. For this extension, and to keep the system size within the bounds of reasonable resource-use, we deployed a slightly simpler motif due to the relatively large number of atoms present in the IDT unit as compared to DPP. The polymer repeat unit motif and functional group subunits are shown in Fig. S4 and S5. The ΔGsolv distributions of all IDT polymer combinations in 25 implicit solvents (1.88 ≤ ε ≤ 78.36), analogous to Fig. 2, are shown in Fig. 8. We emphasize that these calculations were performed prior to any experimental validation. | ||
| Fig. 8 Box-plot distributions of DFT-calculated values of the Gibbs free energy of solvation for 14 IDT-based polymers in 25 implicitly modeled solvents. | ||
As was the case for the DPP-based systems, we observed a minimum in ΔGsolv at a value around ε = 7, corresponding to solvation in chlorobenzene and ortho-dichblorobenzene. However, we note that the DFT-calculated ΔGsolv in tetrahydrofuran (THF) is consistently higher in magnitude (less solvated) than either of its dielectric neighbors, chlorobenzene and ortho-dichblorobenzene. A follow-up study wherein we conduct an explicit solvation study on these three solvents, analogous to Fig. 5, was conducted for all the A-IDT combinations present in Fig. S4. These results, shown in Fig. 9, demonstrate the same trend as seen in the implicit calculations: THF has a less negative (unfavorable) binding enthalpy with these polymers than its dielectric neighbors. We posit that this may be due to the more localized dipole moment of THF as compared to chlorobenzene and ortho-dichlorobenzene. The larger aromatic rings present in these two dielectric neighbors also allow for greater charge stabilization, which enhances binding enthalpy.
Experimental solvation observations on an indacenodithiophene-benzothiadiazole (IDTBT) polymer revealed insolubility in a dielectric as low as acetone (ε = 20.7) while noting solvation in THF and its neighbors. We observed a maximum ε value at which a polymer is soluble; regardless of this value, solvents with a dielectric lower than this critical point exhibit solubility. An analysis of variance (ANOVA) test was conducted on the solvation data, grouped by solvent, and found a rejection of the null hypothesis: that all mean ΔGsolv values among solvent groups were equal. Consequently, we used Tukey's range test to quantify the confidence intervals of the data within each group. The results of this test are shown in Fig. 10.
There may be minimal overlap in the confidence intervals of the chlorobenzene and ortho-dichlorobenzene ΔGsolv population means but, importantly, they both exhibit a statistical difference (i.e., no overlap) from THF. This conclusion is reinforced with evidence from k-means clustering analysis on the DPP data. Fig. S6 and S7 demonstrate that four clusters is optimal for converging a within-class sum of squared distances to an asymptote, when grouping solvation data by solvent, and that those four clusters occupy discrete ranges along Principal Component (PC) 1. These four clusters have been color-coded and applied as a filter to Fig. 11, whose underlying data is the same as Fig. 2.
THF is clustered separately from its dielectric neighbors, as are cyclohexanone and ethanol, hinting at the highly nonlinear behavior of ΔGsolv as a function of ε. It is possible that the two clusters with the highest magnitude ΔGsolv (green and blue in Fig. 11) define a range of “optimal” solvents for this application. However, further investigation into the entropic effects of solvation is needed in order to assert the existence of a single “optimal” solvent and to better match experimental observations.
3.4 Strengths and limitations of the present approach
Ultimately, the design framework described herein possesses several key strengths which underlie its utility for materials design. The physics-informed Gaussian Process surrogate model, trained on an exhaustive computational survey of the curated design space, demonstrated an accurate learning of the search space. Such an exhaustive survey of a design space is not always necessary, as evident in the rapid convergence of the Bayesian optimization algorithm after examining less than 20% of the search space. PAL 2.0 is agnostic regarding the origin of the data: it can be used with computational or experimental data training sets, or a combination of both. This is consistent with our experimental observations of the solubility of IDT-based polymers. Our framework demonstrated generality across both DPP- and IDT-based polymer systems, suggesting applicability across planar, conjugated systems. From an algorithm performance standpoint, PAL 2.0 required sparse training data and very few surrogate function evaluations when compared to neural network-based methods, which often require thousands of data points to be effective.In addition, experimentalists can find value in the chemical conclusions of the present work due to the physical interpretability of the design framework. Advanced feature selection methods identified an electrostatic descriptor, the quadrupole moment, as a key property that affects the free energy of solvation. When designing polymer repeat units, references on quadrupole moments of functional group subunits can be consulted to gain a deeper understanding of the expected solubility of a given candidate before ever synthesizing it. Moreover, to prepare coating formulations that include two or more conjugated molecules or polymers, such as bulk heterojunction solar cells, it should be possible to identify a priori a common solvent system that optimizes the combined solubility of the multiple components.
The present approach is not without its limits. Although the data and analysis to date suggest generality across planar, conjugated systems, we have only tested the framework's efficacy on two classes of materials systems. Until other, different systems are tried, the limit of PAL's efficacy remains unknown. The ultimate breadth and limitations of its efficacy will be revealed only after we and others apply the method to other systems. With that said, PAL 2.0 has demonstrated its use in multiple application areas from high entropy alloys to Metal Organic Frameworks (MOFs) to solar cell materials.33,34,54 Returning to polymer systems, it could also be that the method might struggle with handling side-chains whose interactions with solvents are more complex than the simplified interactions studied here, e.g., side-chains with larger free volume quantities than the ones described here. Similar approaches to optimize solvents for those side-chains could be taken wherein the side-chain interactions might dominate over the main-chain conjugated subunits of this manuscript. To see how much of an issue this is likely to be, we investigated the free volumes of functional group subunits explored in this work. As provided in Fig. S8 of the SI, our per-subunit free volumes range widely, from 31.1 Å3 to 1739.9 Å3. These quantities were calculated by subtracting the isosurface volume from the DFT calculations described above from the volume of the smallest rectangular prism that enclosed the functional group subunit. Thus, the range of free volumes exhibited by polymer repeat unit candidates as linear combinations of their functional group subunits ranged from 254.9 Å3 to 7090.1 Å3. This goes a long way to mitigating any concerns about the volume issues that might arise.
Future work involving the design approach described here could take two distinct directions. The first is utilizing larger-scale computational methods, like classical MD, to quantify the polymer–solvent interactions of multiple repeat units (n > 1) as described above. The second involves broadening the search space of polymers by removing the “class” restraint imposed in the present work. Such an approach would allow all the discrete sites in a polymer repeat unit to be occupied by any functional group subunit, rather than requiring one site always be occupied by, e.g. DPP or IDT. This would move the approach of this framework away from an “exploitative” approach of searching near known high-performing candidates to a more “exploratory” approach that searches far from currently known structures.
4 Conclusions
In this work, we created a curated database consisting of 324 DPP-based semiconducting polymers with a systematic variation of functional groups commonly found in thermoelectric polymers. Using DFT and the universal solvation model, we calculated the free energies of solvation (ΔGsolv) of all 324 polymers we considered in 26 choices of solvent, a total of over 8000 combinations (of which over 7000 converged energetically), far exceeding experimental consideration of all of these candidates. Validation of our method with a second, distinctly separate class of polymers, those based on IDT, suggests that it is, at the very least, applicable to rigid, planar donor–acceptor conjugated backbones.Using DFT-calculated properties that quantify charge density distribution as descriptors, we trained a GP model and demonstrated the usefulness of the PAL 2.0 Bayesian optimization algorithm to predict ΔGsolv. The PAL 2.0 codebase found the optimum result, in its best replicate performance, within two iterations and identified, using XGBoost and LASSO methods for “feature engineering”, important descriptors needed to predict ΔGsolv, our metric of success for insight into the solution behavior of the system. Explicit all-atom modeling of the polymer–solvent interactions revealed a similar trend to the behavior exhibited by ΔGsolv when an implicit solvent model is used, in which the solvent is represented by a mean field of a given dielectric constant (ε).
We identified a molecular electronic parameter that demonstrates near-linear additivity when constructing an aggregate polymer from functional group sub-units, namely the isotropic quadrupole moment. The importance of this observation is that we have identified a fast route to determining better polymer candidates, without needing to resort to machine learning, as used in this PAL-generated study. It suggests that future studies of different semiconducting polymers, beyond simply DPP-based ones, could conceivably only need to model the isotropic quadrupole moment of the various functional group sub-units in the posited polymer in order to be able to predict the extent of solvation of this novel polymer. Thus, this work lays the groundwork for a unique closed-loop, hybrid computational–experimental study wherein polymers could be designed in silico for a target property and then synthesized and characterized with experimental techniques. Ameliorating computational property predictions with experimentally observable properties will be important for model validation and investigation design.
Author contributions
C. G.: conceptualization, methodology, data curation, visualization, and writing. Y. F.: data curation, visualization, and writing. H. S. K.: experimental data curation and writing. H. E. K.: conceptualization, supervision, and writing. P. C.: conceptualization, funding acquisition, project administration, resources, supervision, and writing.Conflicts of interest
The authors declare no conflict of interest.Data availability
The DPP-based polymer electronic structure calculations dataset generated during this study is available on Github at https://github.com/pclancy-lab/polymer-builder.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5py01172h.
Code availability: the PAL 2.0 software is available on Github at https://github.com/ClancyLab/PAL2.
Acknowledgements
This work has been sponsored by an NSF (CHE) award to Drs Howard E. Katz and Paulette Clancy, award number 2107360 and by the Hopkins Institute for Data-Intensive Engineering and Science (IDIES) through the provision of a seed grant to partially fund this study. The authors gratefully acknowledge the support of both awards for this work. We thank Dr Tushita Mukhopadhyay for sharing her expertise in DPP-based polymer systems, and Dr Maitreyee Sharma Priyadarshini for assistance with the PAL 2.0 methodology. Computing resources were provided by the ARCH high-performance computing (HPC) facilities at Johns Hopkins University, which is supported by National Science Foundation (NSF) grant number OAC 1920103.References
- A. R. Murad, A. Iraqi, S. B. Aziz, S. N. Abdullah and M. A. Brza, Polymers, 2020, 12, 2627 CrossRef CAS PubMed.
- S. A. Lopez, B. Sanchez-Lengeling, J. De Goes Soares and A. Aspuru-Guzik, Joule, 2017, 1, 857–870 CrossRef CAS.
- X. Yan, M. Xiong, J.-T. Li, S. Zhang, Z. Ahmad, Y. Lu, Z.-Y. Wang, Z.-F. Yao, J.-Y. Wang, X. Gu and T. Lei, J. Am. Chem. Soc., 2019, 141, 20215–20221 Search PubMed.
- J. Lee, S. A. Park, S. U. Ryu, D. Chung, T. Park and S. Y. Son, J. Mater. Chem. A, 2020, 8, 21455–21473 Search PubMed.
- T.-Q. Nguyen, R. Y. Yee and B. J. Schwartz, J. Photochem. Photobiol., A, 2001, 144, 21–30 Search PubMed.
- M. C. Scharber and N. S. Sariciftci, Adv. Mater. Technol., 2021, 6, 2000857 Search PubMed.
- I. Kaur, W. Jia, R. P. Kopreski, S. Selvarasah, M. R. Dokmeci, C. Pramanik, N. E. McGruer and G. P. Miller, J. Am. Chem. Soc., 2008, 130, 16274–16286 Search PubMed.
- V. Sharma, C. Wang, R. G. Lorenzini, R. Ma, Q. Zhu, D. W. Sinkovits, G. Pilania, A. R. Oganov, S. Kumar, G. A. Sotzing, S. A. Boggs and R. Ramprasad, Nat. Commun., 2014, 5, 4845 Search PubMed.
- S. Ren, Y. Ding, W. Zhang, Z. Wang, S. Wang and Z. Yi, Polymers, 2023, 15, 3803 Search PubMed.
- K. Lee, M.-K. Jeong, E. H. Suh, W. Jeong, J. G. Oh, J. Jang and I. H. Jung, Adv. Electron. Mater., 2022, 8, 2101105 Search PubMed.
- NREL, Best Research Cell Efficiency Chart, 2024, https://www.nrel.gov/pv/cell-efficiency.html.
- A. T. John, A. Narayanasamy, D. George and M. Hariharan, Cryst. Growth Des., 2022, 22, 1237–1243 CrossRef CAS.
- S. Lee, D. Jeong, C. Kim, C. Lee, H. Kang, H. Y. Woo and B. J. Kim, ACS Nano, 2020, 14, 14493–14527 CrossRef CAS PubMed.
- D. Lungwitz, A. E. Mansour, Y. Zhang, A. Opitz, S. Barlow, S. R. Marder and N. Koch, Chem. Mater., 2023, 35, 672–681 CrossRef CAS PubMed.
- M. Duhandžić, M. Lu-Dìaz, S. Samanta, D. Venkataraman and Z. Akšamija, Phys. Rev. Lett., 2023, 131, 248101 CrossRef PubMed.
- W. J. R. Jayasundara and G. Schreckenbach, J. Phys. Chem. C, 2020, 124, 17528–17537 CrossRef CAS.
- J. Tang, Y. H. Pai and Z. Liang, ACS Energy Lett., 2022, 7, 4299–4324 Search PubMed.
- F. M. A. Alzahrani, M. Sagir, M. Saqib, S. Bashir, T. Sarwar, S. Hussain, S. Murtaza, A. Mushtaq, R. Razzaq, Z. A. Alrowaili and M. S. Al-Buriahi, Comput. Mater. Sci., 2024, 239, 112961 Search PubMed.
- N. Meftahi, M. Klymenko, A. J. Christofferson, U. Bach, D. A. Winkler and S. P. Russo, npj Comput. Mater., 2020, 6, 1–8 Search PubMed.
- S. Wu, Y. Kondo, M. a. Kakimoto, B. Yang, H. Yamada, I. Kuwajima, G. Lambard, K. Hongo, Y. Xu, J. Shiomi, C. Schick, J. Morikawa and R. Yoshida, npj Comput. Mater., 2019, 5, 66 CrossRef.
- P. Xu, T. Lu, L. Ju, L. Tian, M. Li and W. Lu, J. Phys. Chem. B, 2021, 125, 601–611 CrossRef CAS PubMed.
- T. B. Martin and D. J. Audus, ACS Polym. Au, 2023, 3, 239–258 CrossRef CAS PubMed.
- L. Gao, J. Lin, L. Wang and L. Du, Acc. Mater. Res., 2024, 5, 571–584 Search PubMed.
- J. Hachmann, R. Olivares-Amaya, S. Atahan-Evrenk, C. Amador-Bedolla, R. S. Sánchez-Carrera, A. Gold-Parker, L. Vogt, A. M. Brockway and A. Aspuru-Guzik, J. Phys. Chem. Lett., 2011, 2, 2241–2251 CrossRef CAS.
- O. H. Omar, T. Nematiaram, A. Troisi and D. Padula, Sci. Data, 2022, 9, 54 CrossRef CAS PubMed.
- T. Mukhopadhyaya, T. D. Lee, C. Ganley, S. Tanwar, P. Raj, L. Li, Y. Song, P. Clancy, I. Barman, S. Thon and H. E. Katz, Adv. Funct. Mater., 2022, 32, 2208541 CrossRef CAS.
- T. Mukhopadhyaya, J. Wagner, T. D. Lee, C. Ganley, S. Tanwar, P. Raj, L. Li, Y. Song, S. J. Melvin, Y. Ji, P. Clancy, I. Barman, S. Thon, R. S. Klausen and H. E. Katz, Adv. Funct. Mater., 2024, 34, 2309646 CrossRef CAS.
- N. E. Jackson, B. M. Savoie, K. L. Kohlstedt, M. Olvera de la Cruz, G. C. Schatz, L. X. Chen and M. A. Ratner, J. Am. Chem. Soc., 2013, 135, 10475–10483 CrossRef CAS PubMed.
- J. Liu, Y. Shi and Y. Yang, Adv. Funt. Mater., 2001, 11, 420–424 CrossRef CAS.
- S. N. Patel, A. M. Glaudell, K. A. Peterson, E. M. Thomas, K. A. O'Hara, E. Lim and M. L. Chabinyc, Sci. Adv., 2017, 3, e1700434 Search PubMed.
- D. Dhabal, Z. Jiang, A. Pallath and A. J. Patel, J. Phys. Chem. B, 2021, 125, 5434–5442 Search PubMed.
- B. Lukose, S. V. Bobbili and P. Clancy, Mol. Simul., 2017, 43, 743–755 CrossRef CAS.
- M. S. Priyadarshini, O. Romiluyi, Y. Wang, K. Miskin, C. Ganley and P. Clancy, Mater. Horiz., 2024, 11, 781–791 RSC.
- M. S. Priyadarshini, E. Gienger, J. Ren, B. Piloseno, E. A. Pogue, P. K. Lambert, J. S. Ko and P. Clancy, Machine Learning-Driven Closed-Loop Discovery of Hard Multiple Principal Element Alloys, 2025, https://chemrxiv.org/engage/chemrxiv/article-details/68a7f1d1a94eede154053d56.
- X. Zhang, H. Bronstein, A. J. Kronemeijer, J. Smith, Y. Kim, R. J. Kline, L. J. Richter, T. D. Anthopoulos, H. Sirringhaus, K. Song, M. Heeney, W. Zhang, I. McCulloch and D. M. DeLongchamp, Nat. Commun., 2013, 4, 2238 Search PubMed.
- R. Chen, T. Jin, Y. Liu, T. Zhang, X. Liu, L. Zhang, Y. Chen, H. Li, X. Duan and Y. Han, Macromolecules, 2023, 56, 5356–5368 Search PubMed.
- L. Goerigk and S. Grimme, Phys. Chem. Chem. Phys., 2011, 13, 6670–6688 Search PubMed.
- M. T. d. Oliveira, J. M. A. Alves, N. L. Vrech, A. A. C. Braga and C. A. Barboza, Phys. Chem. Chem. Phys., 2023, 25, 1903–1922 Search PubMed.
- I. V. Tetko, J. Gasteiger, R. Todeschini, A. Mauri, D. Livingstone, P. Ertl, V. A. Palyulin, E. V. Radchenko, N. S. Zefirov, A. S. Makarenko, V. Y. Tanchuk and V. V. Prokopenko, J. Comput.-Aided Mol. Des., 2005, 19, 453–463 CrossRef CAS PubMed.
- H. Moriwaki, Y.-S. Tian, N. Kawashita and T. Takagi, J. Cheminf., 2018, 10, 4 Search PubMed.
- G. Landrum, P. Tosco, B. Kelley, R. Rodriguez, D. Cosgrove, R. Vianello, P. sriniker, G. Jones, N. Schneider, E. Kawashima, D. Nealschneider, A. Dalke, M. Swain, B. Cole, S. Turk, A. Savelev, A. Vaucher, M. Wójcikowski, I. Take, tadhurst cdd, V. F. Scalfani, R. Walker, D. Probst, K. Ujihara, J. Lehtivarjo, G. Godin, A. Pahl, F. Bérenger and H. Faara, rdkit/rdkit:2024_09_3 (Q3 2024) Release, 2024, https://zenodo.org/records/14245989 Search PubMed.
- F. Pereira, K. Xiao, D. A. R. S. Latino, C. Wu, Q. Zhang and J. Aires-de-Sousa, J. Chem. Inf. Model., 2017, 57, 11–21 Search PubMed.
- F. Neese, Wiley Interdiscip. Rev.:Comput. Mol. Sci., 2011, 2, 73–78 Search PubMed.
- F. Neese, Wiley Interdiscip. Rev.:Comput. Mol. Sci., 2017, 8, e1327 Search PubMed.
- T. Lu and F. Chen, J. Comput. Chem., 2012, 33, 580–592 CrossRef CAS PubMed.
- Z. Liu, T. Lu and Q. Chen, Carbon, 2021, 171, 514–523 Search PubMed.
- A. V. Marenich, C. J. Cramer and D. G. Truhlar, J. Phys. Chem. B, 2009, 113, 6378–6396 CrossRef CAS PubMed.
- V. Barone and M. Cossi, J. Phys. Chem. A, 1998, 102, 1995–2001 CrossRef CAS.
- T. Chen and C. Guestrin, Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, New York, NY, USA, 2016, pp. 785–794.
- R. Tibshirani, J. R. Stat. Soc. Ser. B: Methodol., 1996, 58, 267–288 CrossRef.
- F. Archetti and A. Candelieri, Bayesian Optimization and Data Science, Springer International Publishing, Cham, 2019, pp. 57–72 Search PubMed.
- Y. Zheng, S. Zhang, J. B.-H. Tok and Z. Bao, J. Am. Chem. Soc., 2022, 144, 4699–4715 Search PubMed.
- J. Imbrogno, K. Maruyama, F. Rivers, J. R. Baltzegar, Z. Zhang, P. W. Meyer, V. Ganesan, S. Aoshima and N. A. Lynd, ACS Macro Lett., 2021, 10, 1002–1007 Search PubMed.
- S. Raguraman, M. S. Priyadarshini, T. Nguyen, R. McGovern, A. Kim, A. J. Griebel, P. Clancy and T. P. Weihs, J. Magnesium Alloys, 2024, 12, 2267–2283 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |






