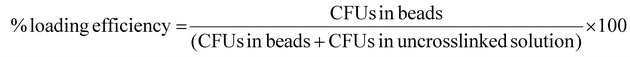Open Access Article
Open Access ArticleEffects of microparticle composition on colony morphology and viability of encapsulated therapeutic yeast for oral delivery
Emma
Etter
ab,
Alita F.
Miller
 c,
Timothy
Little
c,
Sri Sruthi
Potluru
ab,
Srilekha
Venkatraman
ab and
Juliane
Nguyen
c,
Timothy
Little
c,
Sri Sruthi
Potluru
ab,
Srilekha
Venkatraman
ab and
Juliane
Nguyen
 *ac
*ac
aJoint Department of Biomedical Engineering, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA. E-mail: julianen@email.unc.edu
bNorth Carolina State University, Raleigh, NC 27695, USA
cDivision of Pharmacoengineering and Molecular Pharmaceutics, Eshelman School of Pharmacy, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA
First published on 29th December 2025
Abstract
Engineered live biotherapeutic products (LBPs) offer a promising avenue for targeted drug delivery, particularly within the gastrointestinal (GI) tract. Among microbial chassis, Saccharomyces cerevisiae (S. cerevisiae) is recognized as a highly favorable platform due to its safety profile, genetic amenability, and potential for dual functionality as both a therapeutic protein producer and probiotic. However, oral delivery of LBPs remains challenging due to the harsh conditions of the GI tract, which compromise microbial viability and therapeutic efficacy. To address this, we developed alginate-based hydrogel particles designed to encapsulate S. cerevisiae for oral administration and systematically evaluated their performance under simulated physiological conditions. Notably, we demonstrated that colony size can be tuned through specific alginate formulations, and that colony morphology significantly influences cell survival. Our findings establish key design principles for optimizing hydrogel carriers to enhance the viability and therapeutic potential of engineered microbial therapeutics.
1. Introduction
Engineered microorganisms, or microbes, have emerged as powerful tools in biomedical engineering, offering dynamic, living platforms for therapeutic intervention, disease diagnosis and drug delivery.1 These living, non-mammalian organisms are classified as live biotherapeutic products (LBPs) intended for the diagnosis, prevention and treatment of human disease.2 LBPs encompass both intrinsically beneficial strains, commonly known as probiotics, and engineered LBPs which are genetically engineered for enhanced therapeutic function.3 Microbes such as bacteria and fungi have been recombinantly engineered into LBPs to serve as in situ biomanufacturing platforms that produce and secrete therapeutic proteins directly within the body. Unlike free protein therapeutics, which are often unstable especially after oral administration, LBPs can continuously produce therapeutic agents and proteins at the site of disease. This local action enhances bioavailability and has the potential to reduce dosing frequency.4 Additional approaches such as anchoring to physiological targets within the colon lead to prolonged gut retention and improved efficacy of various LBP systems.5 Furthermore, they can be designed for controlled release or to respond to specific physiological cues, allowing for more precise and sustained therapeutic interventions.6Fungi, particularly yeasts, represent a promising class of organisms for use as LBPs. Among them, Saccharomyces cerevisiae (S. cerevisiae), commonly known as baker's yeast, stands out as an especially attractive chassis for therapeutic applications.7 It has a long-standing history of safe use in food and pharmaceutical industries, is generally recognized as safe (GRAS) by the FDA, and is typically non-colonizing and non-pathogenic, making it well-suited for use in humans.8 Moreover, S. cerevisiae is highly amenable to genetic engineering, enabling the integration of complex synthetic circuits and the controlled expression of therapeutic payloads, such as anti-inflammatory cytokines, metabolic enzymes, or immune-modulating proteins.9 Its eukaryotic cellular machinery also supports the correct folding and post-translational modification of human proteins, which is often a limitation in bacterial systems.10 In addition to its engineering flexibility, certain strains of S. cerevisiae, such as S. boulardii, have intrinsic probiotic properties and have been shown to modulate the gut microbiota and improve outcomes in gastrointestinal disorders such as inflammatory bowel disease (IBD), antibiotic-associated diarrhea, and Clostridioides difficile infection.11 This dual functionality, as both a drug-producing chassis and a probiotic, positions S. cerevisiae as a versatile and powerful platform for next-generation microbial therapeutics.
Oral delivery is the preferred method for administering LBPs due to its non-invasive nature, higher patient compliance, and suitability for treating gastrointestinal tract diseases.12 However, delivering LBPs orally can result in decreased viability of the LBPs due to extreme pH changes in the stomach to digestive enzymes and mechanical disruption in the intestines.13 These are known to negatively affect LBP viability which results in lower function as a local drug production factory in the GI tract.14 Previous work in our lab has highlighted this limitation as the LBP was shown to display low viability in acidic pH. To address these limitations in LBP delivery efficiency, yeast and bacteria have been loaded into carriers to protect them during GI transit and deliver them to their target location with improved viability and function.15 Of these delivery vehicles, hydrogels have been employed and are an attractive choice as they can be biocompatible and modifiable to address specific engineering concerns.16 Loading LBPs into hydrogel carriers can protect cells from the harsh environment in the GI tract.17
Alginate is widely utilized in oral drug delivery systems due to its excellent biocompatibility, mucoadhesive properties, and ability to form hydrogels under mild conditions.18 While its use for encapsulating free biologics, such as proteins and peptides, has been extensively studied, its application for delivering LBPs is relatively newer.19 Alginate encapsulation is advantageous for LBP delivery systems because it maintains viability and performance while passaging the harsh conditions of the gastrointestinal tract.20 While these approaches have been developed for general probiotic use, knowledge gaps remain to develop optimal delivery strategies for therapeutic-intended LBPs, particularly for fungal chassis.14 Thus, there is a need to better understand the effect of alginate microencapsulation on therapeutic yeast, such as S. cerevisiae, a relatively unexplored field.21 In particular, there is limited understanding of how the composition and physical properties of alginate hydrogels influence both the structural stability of the particles and the viability of encapsulated yeast during gastrointestinal transit.
Microorganisms, including yeast, are known to form biofilm-like colonies in confined or static environments, which can enhance survival by providing physical protection and facilitating cell-to-cell signaling.22,23 Despite this, few studies have investigated how such colony formation occurs within hydrogel matrices or how it impacts the function of engineered microbes used as LBPs. To our knowledge, no prior work has systematically examined the relationship between hydrogel composition, colony morphology, and gene expression dynamics in therapeutic yeast. Likewise, no studies have investigated how these factors collectively influence LBP viability and therapeutic potential. This represents a critical knowledge gap in the design of optimized delivery systems for engineered microbial therapeutics.
The objective of the study is to develop a biocompatible hydrogel drug delivery system for oral administration of LBPs to the lower GI tract, with potential applications in treating conditions such as inflammatory bowel diseases and colon cancer. To achieve this, the carrier must meet the following key criteria: (1) efficiently encapsulate viable yeast cells with tunable particle size suitable for oral delivery, (2) maintain structural integrity throughout transit in the harsh GI tract environment, and (3) support cell viability and functional activity to ensure ability to exert therapeutic efficacy.
We hypothesize that by systematically optimizing hydrogel particle formulations, we can achieve greater stability and enhanced support for cell viability compared to unmodified alginate particles. Although therapeutic outcomes were not directly assessed in this study, we anticipate that improvements in particle stability and live cell delivery will ultimately translate to more effective drug delivery and clinical benefit. Here, we engineered alginate-based hydrogel particles encapsulating S. cerevisiae and assessed their performance under simulated physiological conditions. Specifically, we evaluated particle stability following exposure to simulated gastric and intestinal fluids and cell viability and functional activity in different nutrient conditions. Notably, we demonstrated for the first time that colony size of S. cerevisiae can be tuned through specific alginate formulations, and that colony size has a significant impact on yeast survival. These findings offer valuable design principles for optimizing hydrogel carriers to maximize viability and therapeutic potential of engineered live biotherapeutics.
2. Materials and methods
2.1. Materials
Sodium alginate, calcium chloride (CaCl2), barium chloride (BaCl2), gelatin, chitosan, poly-L-lysine (0.1% w/v), sodium citrate dihydrate, and phosphate buffered saline (PBS) were purchased from Sigma-Aldrich (St Louis, MO, USA). Yeast peptone dextrose broth (YPD, Difco; 10 g L−1 yeast extract, 20 g L−1 peptone, 20 g L−1 dextrose) was purchased from Becton Dickinson (Franklin Lakes, NJ, USA).2.2. Yeast culturing
S. cerevisiae, strain JK93Dα, was previously engineered to secrete green fluorescent protein (GFP) as a model fluorescent protein for visualization and to model the secretion of engineered biologics from cells.24 Overnight cultures were grown in YPD medium (Becton Dickinson) at 30 °C for 12–16 hours then optical density at 600 nm (OD600) was measured to estimate cell counts then cells were centrifuged at 3000 rpm for 3 minutes and resuspended to obtain the desired concentration for experiments.2.3. Polymer preparation and particle synthesis
Alginate polymer solution was prepared by dissolving sodium alginate (Sigma-Aldrich) in filtered diH2O at a concentration of 2% w/v, unless otherwise stated. Calcium chloride and barium chloride (Sigma-Aldrich) crosslinking solutions were prepared by dissolving each reagent in diH2O to obtain desired concentrations (1% unless otherwise stated). The solutions were then filtered through 0.22 μm PES filters (Genesee Scientific). Yeast culture solutions were added to alginate solutions and mixed well. Then alginate cell solutions were loaded into 5 mL plastic syringes (Becton Dickinson) and added to a syringe pump (Harvard Apparatus). The crosslinking solution was placed below the syringe with a stirring bar keeping solution spinning at 300 rpm. Polymer solutions were added to crosslinking solutions via dropwise extrusion from the syringe pump and were allowed to crosslink for 20 minutes before rinsing with water three times. The resulting alginate particles, with or without encapsulated yeast, were collected with a cell strainer and kept in filtered diH2O solutions until time of use.2.4. Characterization and optimization of methods
Concentrations of alginate and calcium chloride were varied within literature-reported ranges in independent experiments and images of formed particles were taken using an Echo Revolve microscope (Discover Echo Inc., San Diego, CA, USA) to explore particle formation and morphology changes due to changes in these factors.Next, combinations of these solution concentrations were produced to assess the effect on particle formation and morphology. The pH-dependent stability of the particles was determined by incubating particles in pH solutions ranging from 1 to 11 for 4 hours. Particle area was measured using ImageJ image analysis software and plotted as percentage change in diameter over the 0 to 4-hour period.
Evaluation of the pH-dependent stability of the particles in physiological GI conditions was performed by preparing media to mimic the conditions of (1) mouth/esophagus, (2) stomach, (3) small intestine, and (4) large intestine. These media were prepared based on compositions reported in the literature and the pH was adjusted to be physiologically relevant. Media compositions were prepared as follows: saliva fluid25 (14 mg mL−1 NaCl, 0.5 mg mL−1 KCl, 0.1 mg mL−1 CaCl2, 0.15 mg mL−1 NaH2PO4, 0.2 mg mL−1 C6H12O6, 0.7 units per mL lysozyme), simulated gastric fluid (Ricca Chemical Company), and intestinal fluid (Ricca Chemical Company). The base formulations of the commercial simulated fluids used do not include bile salts, a potential limitation of the experimental design. Particles were prepared as described above and sequentially transferred through a series of solutions, each for physiologically relevant durations of human digestion.26 Specifically, particles were first added to Solution 1 (salivary fluid) for 2–8 minutes, then collected and moved to Solution 2 (gastric fluid) for 2 hours, then collected and moved to Solution 3 (intestinal fluid) for the remaining incubation times (4–96 hours). At the beginning and end of each incubation period, optical microscope images were taken to observe particle size and morphology.
2.5. Yeast loading and delivery proof of concept
Particles with and without yeast cell loading were prepared as previously described. Optical images and fluorescent microscopy images were obtained to visualize cells within hydrogel particles. Yeast loaded particles were dried at room temperature overnight then imaged using environmental scanning electron microscopy (ESEM), which does not require sputter coating. Yeast cells within particles were then colorized for better visualization.Yeast cell loading was tested by preparing particles with different concentrations of yeast, achieved by varying the starting concentration of yeast culture added to alginate solutions. Low, medium and high concentrations were prepared by changing yeast concentration 100-fold between groups. To evaluate cells encapsulated in particles, particles were first dissolved by incubating individual particles in 50 mM sodium citrate then homogenizing at the lowest speed (4 m s−1) for 10 seconds. Dissolved particle solutions were then serially diluted and plated on YPD agar plates and left to grow at 30 °C for 48 hours. At this time, colonies were counted and colony-forming units (CFUs) calculated for each group to determine the number of yeast cells loaded into each particle.
Cell growth over time when planktonic versus encapsulated was tested in a similar way. Particles were prepared with either a low or high concentration of yeast. Yeast cell cultures were resuspended in water at the same concentration as cells loaded into alginate solutions. At time points, a sample of either particles or free yeast was collected and particles dissolved then samples were plated and CFUs counted as described above.
Cell viability when exposed to pH stress environments was measured by preparing particles and free yeast solutions with the same starting cell density as described above. Both groups were incubated in either low pH (2) or high pH (6). Particles were dissolved and yeast solutions were collected both at the start of the experiment and after 4 hours incubation. These were serially diluted, plated and CFUs counted and calculated. The percentage change between starting samples and samples after pH incubation was calculated to determine yeast growth.
Cell viability in particles during simulated GI transit was tested by preparing simulated GI fluid solutions as described above. Yeast loaded particles were prepared and incubated in sequential fluids to mimic the GI tract. After incubation in each fluid, a sample of particles were taken, dissolved and plated to analyze CFUs. CFUs from particles incubated in simulated fluids were then compared to a control group of the same particles incubated in PBS to show normal yeast growth in particles when not exposed to simulated fluids.
2.6. Electrospray and diameter control experiments
Particles were prepared as described above but the needle gauge/diameter was varied from 27G (0.21 mm inner diameter) to 15G (1.43 mm inner diameter) needles. Particles were prepared normally and crosslinked in calcium chloride solution then imaged using an optical microscope. Diameters of particles in each group (n = 15) were measured using ImageJ software.Electrosprayed particles were prepared using similar methods, but a voltage was applied to both the extrusion needle and the crosslinking bath using a voltage generator (Gamma High Voltage Research, Inc., Ormond Beach, FL). Particle crosslinking, rinsing, and collection were carried out using the same protocol as that used for particles prepared using the dropwise method. Particles were imaged using an optical microscope and diameters were measured using ImageJ.
A screening was conducted using three needle gauges and three voltage settings, testing all combinations to assess their effects on particle size. Particles were imaged using optical microscope and diameters measured using ImageJ. The relationship between needle gauge and voltage was evaluated using JMP Pro 17.
Particles were prepared as described above and loaded into the electrospray setup. Flow rate from the syringe pump was modified as noted while needle diameter (27G), voltage (9 kV), alginate solution, crosslink bath and drop height were kept constant. Particles were collected and imaged as described previously. Particle diameter from 30 particles was measured using ImageJ.
Similarly, particles were prepared as described above, but yeast concentration loaded into the alginate solution was varied. Overnight yeast cultures were centrifuged and supernatants removed. The yeast pellet was then resuspended in alginate solution to prepare loading concentrations ranging from 0 to 10% w/v. All other electrospray parameters were kept constant as described above. Particles were imaged and diameters measured using ImageJ.
2.7. Stability screening (one factor at a time)
Potential modifications that have been shown to improve the stability of alginate hydrogel particles were identified from the literature. Specifically, higher alginate concentrations,27 alternative crosslinkers,28 and surface coatings29 have been used to strengthen particles. A selection of these modifications was tested for stability in both simulated gastric and simulated intestinal fluid (Ricca Chemical Company) without yeast loaded. In all groups, electrospray setup and parameters were kept constant. Control particles consisted of 2% alginate crosslinked in a calcium chloride solution without any additional surface coating. For modifications to the alginate solutions, higher concentrations of alginate (5%) and gelatin (type A, Sigma-Aldrich) added to alginate were used. A 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixture of alginate and gelatin was sprayed into the crosslinking solution. For modified crosslinking conditions, chitosan (Sigma-Aldrich) was mixed into the calcium chloride solution, while alginate was used as the hydrogel phase.
1 mixture of alginate and gelatin was sprayed into the crosslinking solution. For modified crosslinking conditions, chitosan (Sigma-Aldrich) was mixed into the calcium chloride solution, while alginate was used as the hydrogel phase.
In the other group, barium chloride was prepared and used in place of the calcium solution. For coating modifications, particles were prepared using control parameters. After particles were rinsed and collected, they were moved into either a chitosan or poly-L-lysine (Sigma-Aldrich) solution for 20 minutes. After incubation, particles were rinsed and collected again. Particles were imaged before incubation in simulated fluids then placed into both simulated gastric fluid and simulated intestinal fluid. Images were taken at time points and particle integrity was observed.
2.8. Design of experiments screening for particle stability
To screen a broader range of particle compositions with fewer experimental runs, a custom design of experiments (DoE) was prepared using JMP Pro 17 software. Four factors that improved particle stability were selected from the one-factor-at-a-time screening mentioned above (crosslinker: calcium or barium; crosslink bath additive: none or chitosan; post-crosslinking coating: none, chitosan, or poly-L-lysine (PLL); and alginate concentration: 1%, 2.5%, or 5% w/v). These factors and levels are listed in (Table 1). A total of 12 runs were generated to maximize efficiency. The design did not include factor constraints, and the run order was randomized to minimize systematic bias. All formulations were prepared under identical conditions.| Factor | Levels |
|---|---|
| Alginate concentration | 1%, 5% (2.5% midpoint) |
| Crosslinker | Ca2+, Ba2+ |
| Chitosan added to crosslinking bath | No, Yes |
| Particle coating | None, PLL, chitosan |
Alginate was prepared as previously described in concentrations of 1%, 2.5%, and 5%. A 2% chitosan solution in 10% acetic acid (pH 4.5) (Sigma-Aldrich) was prepared and was added to crosslinking solutions to achieve a final concentration of 0.02% chitosan. Particles were prepared as previously described and according to generated run factors. For coatings, particles were incubated in either chitosan or poly-L-lysine solutions after collection and rinsing.
Equal volumes of particles from each group were placed in simulated intestinal fluid. At time points, images were taken using an optical microscope. The response variable, time to degradation, was calculated as the time point at which no intact particles or particles without significant morphological changes are present. These compositions were also tested in simulated gastric fluid, and all compositions were stable for the 96-hour duration of the experiment.
Descriptive statistics, like mean, standard deviation, median, and interquartile range, were calculated for each run group. A one-way analysis of variance (ANOVA) was used to compare differences in degradation time across runs. Post hoc multiple comparisons were conducted using Tukey's test to identify group-level differences. Survival analysis was conducted, and the event of interest was defined as the time point at which stable particles were no longer observed. Samples that remained stable through the experiment observation period were treated as right censored. Kaplan–Meier survival curves were generated using JMP and statistical differences between levels of factors were assessed using the Log-rank (Mantel–Cox) test in GraphPad Prism.
2.9. Loading efficiency setup, methods, data analysis
Yeast was cultured overnight and was loaded into particles as described above. Particle compositions were prepared using the factors and levels from the stability experiment above but with the inclusion of yeast cells. To determine loading, 1 mL of alginate yeast solution was loaded into electrospray setup. The full volume was extruded into the crosslinking bath and allowed to crosslink. Crosslinking solutions were then collected and a portion was plated on agar plates. CFUs were counted after 48 hours of growth and the total number of uncrosslinked CFUs were calculated. Particles separated from the crosslinking solution were collected, dissolved then serially diluted for plating and CFU counting. Loading efficiency was calculated using the following formula:Descriptive statistics, as mean, standard deviation, median and interquartile range were calculated amongst experimental runs. A one-way ANOVA was performed, and post hoc multiple comparisons were conducted using Tukey's test to identify group-level differences.
2.10. Yeast viability
Yeast cultures grew overnight, and OD600 was measured to estimate yeast growth. Particles were prepared as described above according to generated DoE runs using the same four factors. Yeast peptone dextrose (YPD) media and neutral aqueous media were prepared to provide either a nutrient-rich or nutrient-deficient environment for cell growth. Rinsed and collected particles were diluted then added to 1.5 mL of nutrient-rich or nutrient-deficient medium, supplemented with penicillin–streptomycin in equal volumes. These were incubated at 30 °C with gentle shaking. At each time point (0, 24, 48, 72, and 96 hours), a sample was collected and imaged using an Echo Revolve microscope to visualize cell growth and particle volume changes. Particle diameter (n = 30) of each run was measured using ImageJ and averages were used for further analysis. Samples were then dissolved using sodium citrate as described above. Dissolved particle solutions were serially diluted then plated on agar plates to count CFUs. Once dissolved, the coating materials are able to interact with free yeast, a potential limitation of evaluating inner particle viability. CFU mL−1 at each time point from each run were plotted. The CFU mL−1 for each replicate after 96 hours was plotted as a bar graph to compare final viability for each run composition. Data were analyzed using one-way ANOVA and post hoc multiple comparisons were conducted using Tukey's test to identify group-level differences.2.11. Particle and colony diameter
Changes in particle diameters over time were measured as described above. Mean, standard deviation, median and interquartile range of baseline diameter values were computed for run groups. Particle diameters for each run after fabrication (t = 0) were plotted using a bar graph. The significance of group-level differences was determined by one-way ANOVA followed by post-hoc comparisons using Tukey's HSD test.Colony growth within particles was visualized using optical microscopy and diameters (when applicable = 30 per composition) were quantified using ImageJ for each run at each time point. These were plotted over time and runs were additionally plotted at 96 hours to compare groups at the experimental endpoint. Significance was determined by one-way ANOVA followed by Tukey's multiple comparisons test.
The average colony diameter for each run composition after 24 hours in YPD media was calculated and plotted against the average particle diameter for each run composition after 24 hours to visualize the relationship between colony and particle diameter. The average colony diameter for each run composition was then plotted against the average log(CFU mL−1) for each run composition after 24 hours in YPD media to determine the relationship between colony diameter and cell viability. A simple linear regression was performed on the data in GraphPad Prism to identify a relationship.
2.12. qRT-PCR
Yeast-loaded particles were prepared as described above and incubated in YPD. After 24 hours, particles were imaged then dissolved in sodium citrate. The control group was healthy yeast not encapsulated in particles after incubation in YPD under the same conditions. Total RNA was extracted from yeast using the RiboPure™ RNA Purification Kit (Invitrogen, Waltham, MA) and complementary DNA (cDNA) was synthesized using the First Strand cDNA Synthesis Kit (New England Biolabs, Ipswich, MA), according to the manufacturer's instructions. Gene expression levels were measured by quantitative reverse transcription PCR (qRT-PCR) using the SYBR Green PCR kit (Qiagen). A table of primers used is included in Table 2. Relative gene expression was calculated using the ΔΔCt method. All experiments were performed in triplicate.| Gene of interest | Primer sequence | Source |
|---|---|---|
| ACT1 – FP | 5′ GAATTGAGAGTTGCCCCAGAAGAA 3′ | 30 |
| ACT1 – RP | 5′ AGAAACCAGCGTAAATTGGAACGA 3′ | |
| FLO11 – FP | 5′ CTGGTCCAAAAGATACCGTCCAAC 3′ | 30 |
| FLO11 – RP | 5′ ATGCATATTCAGCGGCACTACCTT 3′ | |
| RIM15 – FP | 5′ CCCAAGGACTAACCACAACATTC 3′ | 31 |
| RIM15 – RP | 5′ ACCTCAACACGGCATCCAC 3′ | |
| STE11 – FP | 5′ TCGTCGAGATCGCCAAGC 3′ | 32 |
| STE11 – RP | 5′ GGCACTTTCATTTCTCCACAGC 3′ | |
| STE12 – FP | 5′ GTATCTCCTAGCGACCCTAC 3′ | 33 |
| STE12 – RP | 5′ AGTTTGCTGGCCAGAGTTGT 3′ | |
| TEC1 – FP | 5′ TCAAGCACAAAACCAACGAG 3′ | 34 |
| TEC1 – RP | 5′ ATGATAGGGTCAGCGAGTCC 3′ | |
| RAS2 – FP | 5′ CGCTCAAAGCGCTAATACGG 3; | 35 |
| RAS2 – RP | 5′ AGGCTTCACTGGTGTTACCG 3′ | |
| TPK2 – FP | 5′ CTGGATCTTTTGGTAGGGTTCA 3′ | 36 |
| TPK2 – RP | 5′ ACGTCGTTCGTCATTGGTATGT 3′ | |
| GLN3 – FP | 5′ TCAACATGACGCCCAGCAACTC 3′ | 37 |
| GLN3 – RP | 5′ CGGGAACAGTGAGGATTTGGAGAC 3′ |
3. Results
3.1. Characterization and optimization of methods
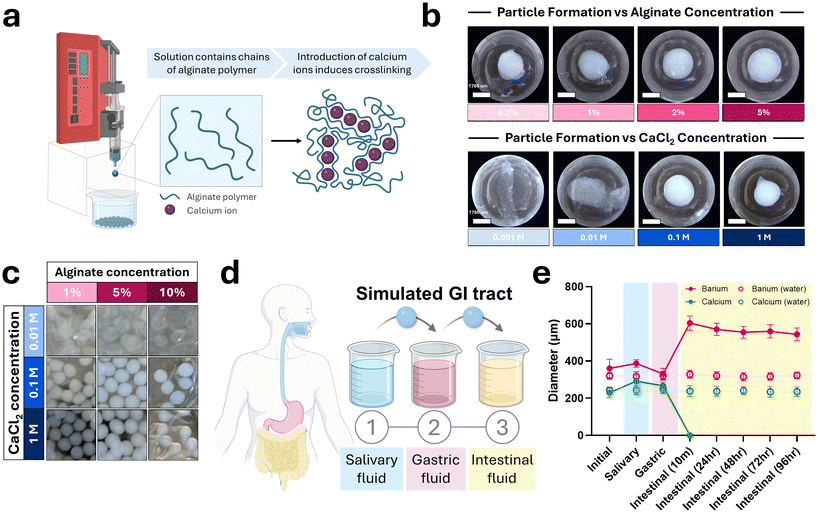
Alginate particles were prepared via dropwise extrusion of alginate solution into a calcium chloride crosslinking bath (Fig. 1a). A range of alginate (0.5%–5%) and calcium chloride (0.001–1 M) concentrations were evaluated for their effects on particle formation and morphology (Fig. 1b). Higher alginate concentrations yielded more spherical and uniform particles but significantly increased solution viscosity, making extrusion and handling more challenging. Based on visual and practical assessment, 2% alginate was selected for subsequent experiments due to its optimal balance between particle uniformity and ease of processing. Similarly, increasing calcium concentrations improved crosslinking density and particle sphericity. Solutions with ≤0.01 M calcium produced poorly crosslinked, irregular particles, while 1 M calcium solutions generated well-formed particles with occasional tailing. A matrix screen of three alginate concentrations and three calcium concentrations confirmed these trends: low calcium resulted in incomplete crosslinking, while high alginate concentrations increased tail formation due to extrusion artifacts (Fig. 1c).Alginate is known to swell as the pH increases, primarily due to deprotonation of carboxyl groups that enhances electrostatic repulsion and water uptake within the polymer matrix.38 To evaluate predicted particle diameter and dissolution during gastrointestinal transit, particles were sequentially incubated with media mimicking each segment of the digestive tract to simulate transit from the oral cavity to the colon (Fig. 1d). Barium and calcium cross-linked particles were fabricated as previously described, then washed and transitioned to the subsequent simulated fluid. Particle diameter was measured at the end of each incubation period, with control particles maintained in water (n = 30 particles per condition). To capture the time to degradation in intestinal fluid following simulated GI transit, particles remained incubated in intestinal fluid and diameters measured from 10 minutes to 96 hours (Fig. 1e). For both formulations, results showed an increase in diameter when incubated in simulated salivary fluid followed by a decrease in diameter when transitioned to simulated gastric fluid. Specifically, the barium cross-linked particles increased from 360 µm to 387 µm then returned to 331 µm while calcium cross-linked particles were initially 230 µm, increased to 291 µm, then returned to 266 µm once incubated in simulated gastric fluid. Upon exposure to intestinal fluid, barium cross-linked particles exhibited significant swelling and had an average diameter of 604 µm. This swelling slightly decreased over time as the barium cross-linked particles remained in the intestinal fluid, with the particles decreasing to 543 µm at 96 hours. The calcium cross-linked particles lost structural integrity within as short as 10 minutes of being incubated in simulated intestinal fluid. Control particles remained stable with no significant changes in size across all time points. These findings confirm the pH-responsive nature of the alginate system and its potential for targeted delivery in the lower GI tract.
3.2. Evaluation of encapsulation on yeast viability
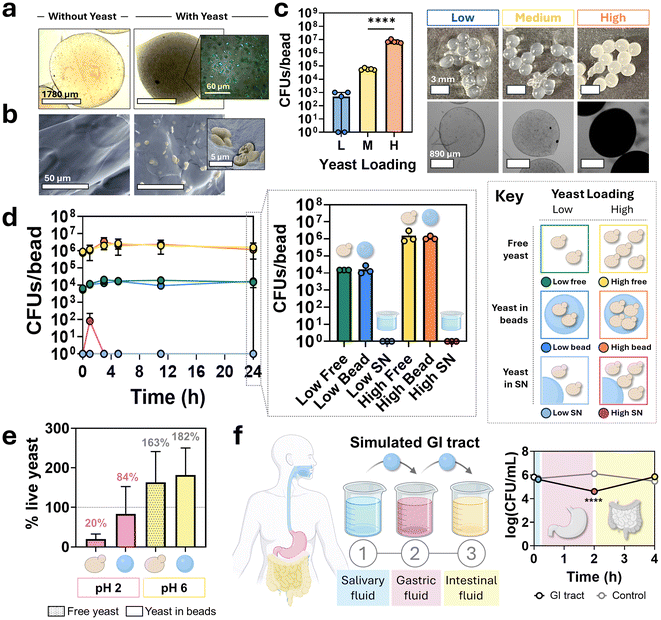
Since alginate is biocompatible and can be crosslinked in mild conditions, cells can be loaded into gels with simple incubation in pre-crosslinked alginate solution.20 Yeast cell suspensions from overnight cultures were mixed into alginate solutions and encapsulated by dropping them into a crosslinking solution. Particles were prepared both with and without yeast cells and imaged using optical and fluorescent microscopy to visualize cells (Fig. 2a). Scanning electron microscopy provided higher magnification images of the embedded S. boulardii, with cells colorized for better visualization (Fig. 2b). These images confirmed that cells were successfully loaded into particles through incubation in the alginate solution.The ability to change loading density was assessed using low, medium and high concentrations of initial yeast cells in alginate pre-cursor solutions (Fig. 2c). Particles were imaged with both a camera and microscope to visualize their density. Particles were dissolved and colony-forming units (CFUs) were counted to determine the number of cells successfully encapsulated particles. The initial cell loading levels were two orders of magnitude apart, and the actual loaded cells followed a similar trend: low concentrations averaged 5 × 102 cells per particle, medium concentrations averaged 6 × 104 cells per particle, and high concentrations averaged 7 × 106 cells per particle, demonstrating the ability to control yeast cells per particle through initial cell density in uncrosslinked solutions.
The growth of yeast within particles was compared to the growth of free yeast to determine if encapsulation negatively impacts growth (Fig. 2d). Both low (104 CFUs per particle) and high (106 CFUs per particle) initial cell densities were tested. CFUs were obtained from free yeast samples, yeast in dissolved particles, and yeast present in the supernatant surrounding the particles. CFUs were plotted over time, with 24-hour CFU values represented in a bar graph for comparison. No statistically significant difference in CFUs between free yeast and encapsulated yeast, regardless of initial cell density (low or high) were found. Additionally, minimal to no yeast cells were detected in the supernatants, indicating strong cell retention within the hydrogel particles throughout the incubation period.
To assess whether encapsulation protects yeast from pH-induced cell death, free yeast and particle-encapsulated yeast (loaded with equivalent cell numbers) were incubated in either low or high pH conditions for 2 hours (Fig. 2e). Following exposure to low pH, the percentage of viable cells dropped to 20% in the free yeast group, whereas encapsulated cells maintained 84% viability. At higher pH conditions, both groups exhibited robust growth, with free yeast showing a 63% increase and particle-encapsulated yeast showing 82% increase in CFUs, indicating comparable proliferation in non-stress environments.
Yeast cell viability within particles was evaluated in a simulated GI solution and compared to control yeast in particles in buffer solution (Fig. 2f). Yeast extracted from particles were plated after each sequential incubation period to determine CFUs. Results showed minimal change in viability following exposure to simulated saliva fluid, but a significant decrease in cell viability after gastric fluid incubation. Notably, yeast viability recovered during incubation in simulated intestinal fluid, with no significant difference observed between this group and the control, indicating the potential for functional recovery after gastric stress.
3.3. Parameters controlling particle diameters
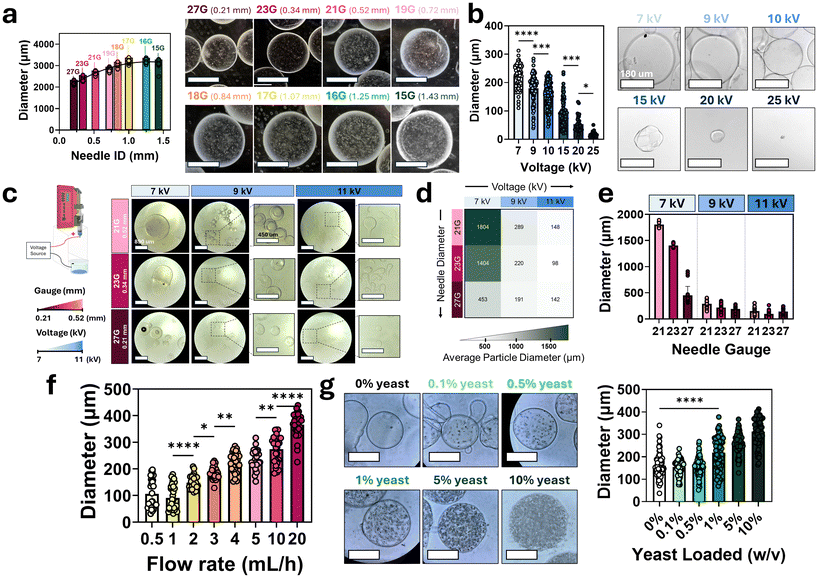
To determine parameters controlling particle diameter, we first evaluated the impact of extrusion needle inner diameter (ID) on particle size (Fig. 3a). The needle gauge was varied from 27G (ID: 0.21 mm) to 15G (ID: 1.43 mm), while keeping all other parameters, formulation, and processing conditions constant. Particle diameters for each group were quantified using ImageJ. A significant increase in particle diameter was observed with each successive increase in needle diameter from 27G to 17G. However, beyond 17G, further increases in needle diameter did not result in significant changes in particle size. Particles were fabricated with average diameters ranging from 2188 μm using a 27G needle to 3194 μm using a 16G needle, an overall average change of around 46%.To further reduce particle size and improve size control, an electrospray system was integrated into the particle fabrication setup. The effect of applied voltage on particle diameter was evaluated by varying voltage while maintaining all other parameters constant (Fig. 3b). Particle diameters were analyzed using ImageJ, revealing a significant inverse relationship between voltage and particle size. This is a well-studied phenomenon in which a stronger electric field increases electrostatic forces on the liquid, breaking it into smaller droplets.39,40 In this study, average diameters ranged from 224 μm at lower voltages to 17.7 μm at higher voltages, demonstrating effective size modulation through voltage control. A screening experiment was then conducted combining needle gauges (27G to 21G) with voltage settings (7 to 11 kV) to assess their individual and combined effects on particle diameter (Fig. 3c). Statistical analysis showed that voltage had a significant effect on particle size, while needle gauge alone did not reach statistical significance but showed a trend toward larger particle sizes with increasing needle diameter (Fig. 3d and e).
The effects of flow rate on particle diameter were also measured by varying flow rates from 0.5 to 20 mL h−1 while keeping all other electrospray parameters constant (Fig. 3f). A significant positive correlation was observed, with particle sizes increasing from 90.9 μm to 366.6 μm across the range. Lastly, the effect of yeast cell loading on particle diameter was evaluated by encapsulating resuspended yeast cell culture pellets in alginate solutions to create concentrations from 0% to 10% w/v (Fig. 3g). Results demonstrated a significant increase in particle diameter with higher yeast loading, indicating that cell density is an important factor in modulating the final particle size, showing an increase from 161 μm average diameter with no yeast loaded to 312 μm on average with higher cell loading.
3.4. Stability
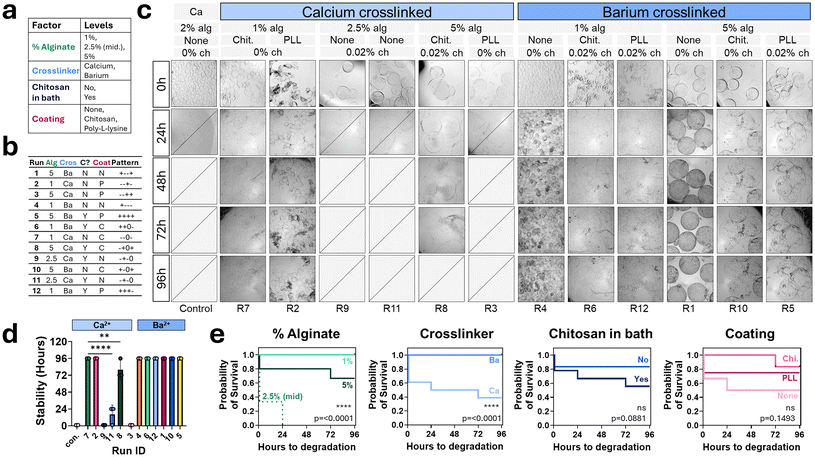
To improve particle stability in simulated gastrointestinal environments, three components of the encapsulation system were identified for potential modification: (1) the alginate solution, (2) the crosslinking solution, and (3) the coating solution (Fig. 4a). Various compositions were screened and a subset demonstrated particle integrity in both simulated intestinal fluid (SIF) and simulated gastric fluid (SGF) (Fig. 4b). Modifications that increased particle stability included: increasing alginate concentration and adding gelatin to the alginate phase; adding chitosan to the crosslinking bath or replacing calcium with barium as the crosslinker; and coating particles with either chitosan or poly-L-lysine (PLL). These factors, excluding gelatin due to resulting particle size and swelling, were used to construct a design of experiments (DoE) framework, with factor levels detailed in Fig. 5a used to systematically generate runs using JMP Pro 17 of different particle compositions designed to evenly distribute combinations of factor levels (Fig. 5b).Representative images from each experimental run are shown in Fig. 5c, highlighting differences in particle degradation rates when incubated in SIF. The primary outcome measured was “time to degradation”, defined as the time point at which no intact particles remained in SIF and these values for each run composition are plotted in the bar graph displayed in Fig. 5d. Statistical analysis revealed significant differences in particle stability across several conditions. There appeared to be two common behaviors amongst particles in SIF, either they degraded immediately within the first 24 hours, or they remained intact through the course of the experiment (96 hours). The majority of run compositions (8 of 13 groups) remained stable in simulated intestinal fluid for over 96 hours. Of these stable runs, six were barium crosslinked, and all compositions crosslinked with barium remained fully stable in simulated intestinal fluid.
The hours to degradation-metric were used to construct a survival curve by separating each run by factor level to visualize stability over time (Fig. 5e). Significance, as assessed using the Log-rank (Mantel–Cox) test in GraphPad Prism, is denoted as p-values on plots. These tests found a significant difference between calcium and barium crosslinkers (p < 0.0001). The other factors did not show significant differences between factor levels. Each run composition was also analyzed for its stability in simulated gastric fluid (SGF) and all compositions were found to retain their integrity in SGF over 96 hours (Fig. S1).
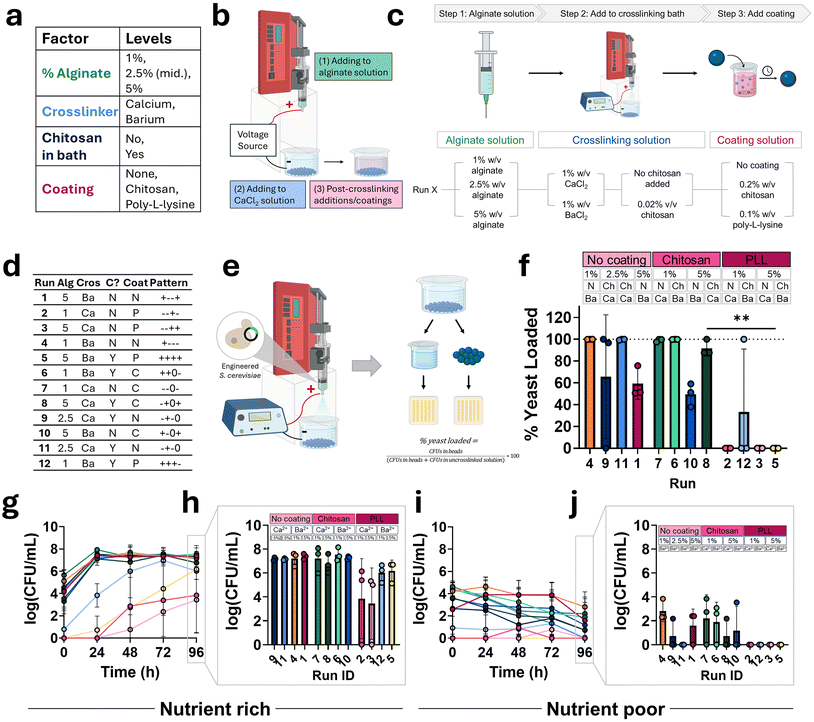
3.5. Cell loading efficiency
The same factors were tested for their cell loading efficiency and viability (Fig. 6a). Modifications to particle compositions were done as previously discussed (Fig. 6b and c) in accordance with determined run factor levels (Fig. 6d). Cell loading efficiency was determined by encapsulating an equal number of cells in each particle composition and quantifying the proportion recovered from dissolved particles relative to the total number of quantified cells (i.e., those encapsulated and those remaining in the uncrosslinked supernatant) (Fig. 6e). Loading efficiencies for each particle composition run are displayed in Fig. 6f. Poly-L-lysine (PLL) coating significantly reduced cell loading efficiency, as demonstrated by near-zero loading across all replicates in the PLL group. While the coating is generally not expected to penetrate deeply into the alginate particles where the yeast reside, partial penetration cannot be ruled out and may influence yeast viability. In addition, the observed differences are likely attributable to the experimental setup, in which particle dissolution enables PLL to exert its antifungal properties on the liberated yeast.41 In contrast, other conditions displayed no statistically significant differences in loading profiles, and no other coating group exhibited an average loading efficiency of 0%, as observed in three of the four PLL-coated compositions.3.6. Encapsulated cell viability in nutrient depleted and nutrient rich conditions
Cell viability was assessed by quantifying viable cells recovered from dissolved particles incubated over time in either nutrient-poor or nutrient-rich (YPD) media. For all compositions in nutrient rich conditions, cell growth occurred since quantified CFU mL−1 for each run exceeded values from the starting point of the experiment (Fig. 6g). For all compositions with no coating or a chitosan coating, a majority of cell growth occurred in the first 24 hours and CFU mL−1 remained stable through 96 hours. For these compositions, there were no significant differences observed between groups after 96 hours of incubation in nutrient-free media (Fig. 6h). A different cell growth pattern is seen in PLL-coated compositions where some growth was observed over time, but none reached the magnitude of non-coated and chitosan-coated compositions. Similar to the observations in yeast loading, these differences may be due to partial penetration of PLL and PLL impacting free yeast after the particles are dissolved. Of the PLL-coated compositions, there was a non-significant trend of higher viability after 96 hours in barium crosslinked compositions compared to calcium crosslinked compositions.In nutrient-poor conditions, cell viability in all compositions declined over time as final CFU mL−1 at 96 hours was lower than starting CFU mL−1 at 0 hours (Fig. 6i). There was an observed difference in endpoint values depending on coating type similar to nutrient rich conditions (Fig. 6j). After 96 hours, all PLL-coated compositions have no viable cells compared to the other coatings where a majority of run compositions have an average non-zero viability.
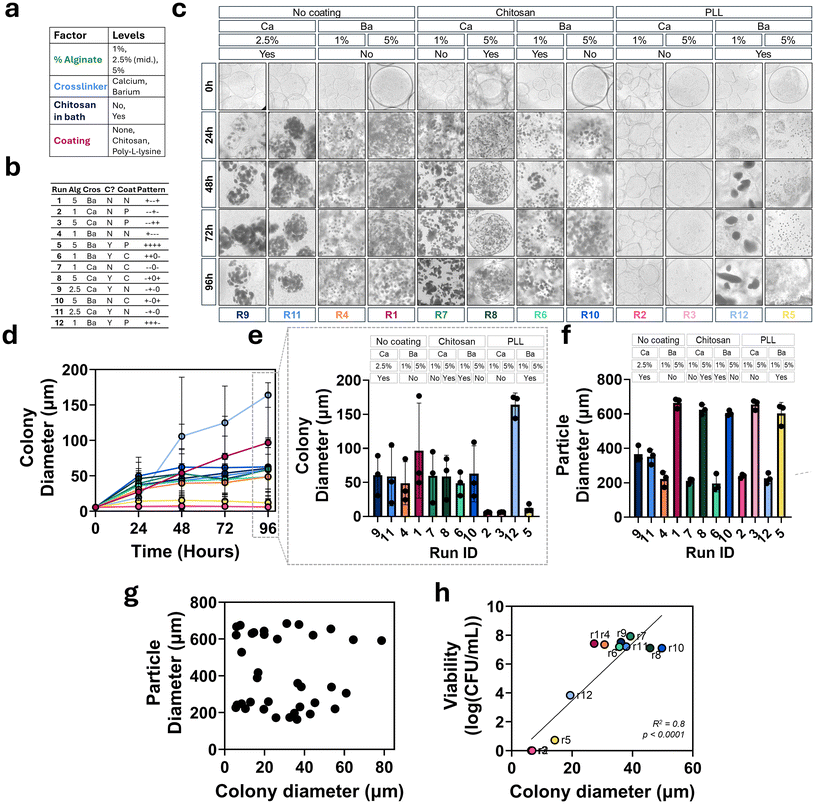
3.7. Effect of particle composition on yeast colony formation
The factor levels (Fig. 7a) and run particle compositions (Fig. 7b) from above were analyzed for their yeast colony formation behaviors. Representative images of each composition every 24 hours are shown in Fig. 7c. Average colony diameter in particles over time was quantified using ImageJ (Fig. 7d) and final average colony diameter after 96 hours is shown in the bar plot in Fig. 7e. The colony growth pattern over time differed depending on composition.For most compositions, yeast went from single cells in particles at 0 hours to clusters or colonies after 24 hours. In most groups, the largest increase in colony diameter occurred over the first 24 hours followed by a gradual increase in diameter through 96 hours. Three compositions, formulation 2 (1% alginate, Ca crosslinked without chitosan, PLL coating), formulation 3 (5% alginate, calcium crosslinked without chitosan, PLL coating), and formulation 5 (5% alginate, barium crosslinked without chitosan, PLL coating), all of which were coated with PLL, remained as single cells or small clusters through the course of the experiment and did not display colony formation behavior. The smallest colonies after 96 hours are seen in these groups, formulation 2 (6 μm), formulation 3 (6.3 μm) and formulation 5 (12 μm), where no little to no colonies formed.
The next largest average colonies, 48.6 μm, 48.8 μm, and 58.7 μm, are seen in formulation 6 (1% alginate, barium crosslinked with chitosan, chitosan coating), formulation 4 (1% alginate, barium crosslinked without chitosan, no coating), formulation 11 (2.5% alginate, calcium crosslinked with chitosan, no coating), respectively. Slightly larger colonies after 96 hours are seen in formulation 8 (5% alginate, calcium crosslinked with chitosan, chitosan coating), formulation 7 (1% alginate, calcium crosslinked without chitosan, chitosan coating), formulation 9 (2.5% alginate, calcium crosslinked with chitosan, no coating), and formulation 10 (5% alginate, barium crosslinked without chitosan, chitosan coating), with average colony diameters of 58.8 μm, 59.6 μm, 60.7 μm, and 62.7 μm, respectively.
The largest colonies are seen in formulation 1 (5% alginate, barium crosslinked without chitosan, no coating) and formulation 12 (1% alginate, barium crosslinked with chitosan, PLL coating). The final colony diameters were 96.7 μm for formulation 1 and 164 μm for formulation 12, which showed statistical significance (p < 0.02). For these two compositions, colony diameter change was not the greatest between 0 and 24 hours like other compositions and instead colonies grew the most between 24 and 96 hours and did not display plateau behavior, resulting in final average colony diameters higher than all other compositions.
The number of colonies per particle was inversely proportional to colony diameter. Formulation 12 with the greatest colony diameter had an average of 2 colonies per particle while those with medium sized colonies had 10–20 colonies per particle. These numbers remained consistent throughout the 96 hours, as expected, since new colonies do not form after encapsulation. Instead, existing colonies grow larger as the original yeast cells continue dividing at their initial location within the solid particle.42 These results also confirm that the particles are robust enough to maintain yeast growth within the particles rather than loss of yeast loading over time.
To test whether colony diameter differences in particle compositions was a result of particle size, we first evaluated particle diameter after fabrication using ImageJ to analyze images of each run composition (Fig. 7f). There was a significant increase in particle diameter when prepared with 5% alginate starting solution versus 1% alginate. Particle diameter differences between other factors were not significant compared to the change in diameter based on percent alginate. Next, particle diameter for each composition was plotted against resulting colony diameter to visualize the relationship, if any, between these variables (Fig. 7g). There was no correlation between particle size and colony size. The data appear grouped based on particle diameter, small particles made with 1% alginate and large particles made with 5% alginate, and each of these groups display a range of colony sizes.
Finally, the relationship between yeast cell viability and colony formation behavior was observed with a plot of colony diameter versus cell count (log(CFU mL−1)) after 24 hours incubation in nutrient rich media (Fig. 7h). A simple linear regression was fit to the data (R2 = 0.8) and found a significant correlation between colony diameter and yeast cell viability (p < 0.0001). In compositions with the smallest (<20 μm) colonies (formulations 2, 3, 5 and 12), the cell viability is the lowest.
3.8. Gene expression in yeast with different colony formation behaviors
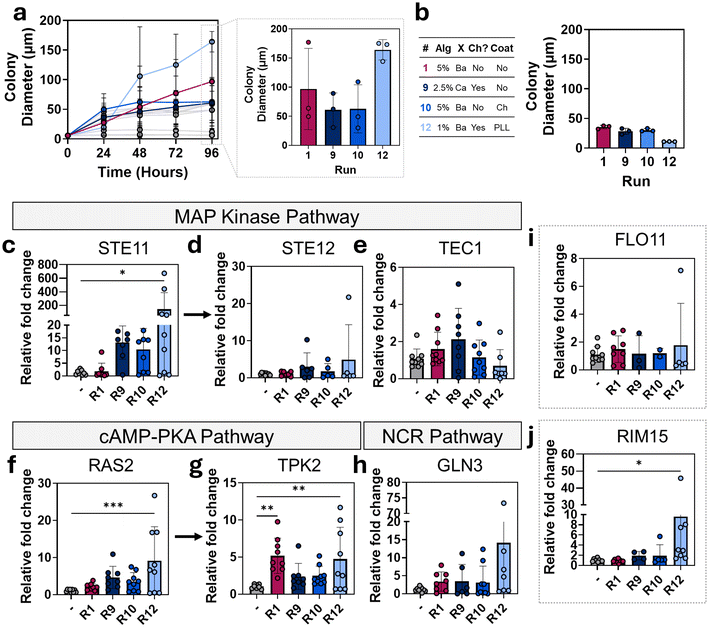
Formulations 1, 9, 10 and 12 were selected for gene expression analysis based on their colony growth behavior (Fig. 8a). Formulations 1 (5% alginate, barium crosslinked without chitosan, no coating) and 12 (1% alginate, barium crosslinked with chitosan, PLL coating) show unique growth behavior where initial colony diameter is low after 24 hours, but increases substantially from 24 to 96 hours, resulting in the largest final colonies of all the compositions. In contrast the other two compositions tested, formulation 9 (2.5% alginate, calcium crosslinked with chitosan, no coating) and 10 (5% alginate, barium crosslinked without chitosan, chitosan coating), display a similar colony growth behavior to the other run compositions where the most change in colony diameter happens in the first 24 hours followed by a gradual increase over the next 24–96 hours. These formulations resulted in the largest final colony diameter after formulations 1 and 12. These compositions and their colony diameters after 24 hours of incubation in YPD are shown in Fig. 8b.Colony formation in S. cerevisiae is a complex behavior, influenced by nutrient access and environmental stresses. To assess the effect of hydrogel composition on yeast colony formation, we focused on three key signaling pathways. Among these, the mitogen-activated protein kinase (MAPK) cascade plays a central role, as it is activated not only by a depletion in carbon nutrient sources but also responds to surface contact, osmotic stress, and cell wall integrity signals. Activation of this pathway induces FLO11 (flocculin 11) expression, which encodes a cell surface glycoprotein essential for cell adhesion and colony formation.30,43
The Ras-cyclic adenosine monophosphate-protein kinase A (Ras-cAMP-PKA) pathway is similarly responsive to carbon source availability but is also influenced by intracellular energy status, pH, and stress signals such as heat shock or oxidative conditions. This pathway regulates both FLO11 expression and RIM15 (regulator of IME3 15), that encodes for a nutrient-sensitive serine/threonine kinase involved in the transition to complex colony architecture under nutrient-poor conditions.43 In addition to carbon limitation, nitrogen availability also influences colony formation and morphology via the nitrogen catabolite repression (NCR) pathway, which leads to the expression of RIM15.43 The NCR pathway regulates the use of nitrogen sources by repressing genes involved in the utilization of poor nitrogen when preferred sources are available.44 The NCR pathway is also sensitive to the TOR (target of rapamycin) signaling axis, linking it to global nutrient sensing, stress response, and autophagy. In addition to these inputs, factors such as oxygen availability, surface stiffness, and quorum-sensing molecules further fine-tune the expression of colony morphogenesis genes. Thus, colony development in S. cerevisiae arises from the integration of diverse environmental and intracellular signals through these core regulatory networks.43
In the MAPK pathway, the gene STE11 (sterile 11) encodes a MAP kinase kinase kinase (MAPKKK) that activates an invasive growth MAPK cascade in S. cerevisiae, which ultimately leads to transcriptional changes that promote biofilm-like colony architectures.43,45 An upregulation in STE11 suggests activation of the MAPK cascade in response to environmental stressors.32 There was a significant upregulation (p = 0.0198) in expression of STE11 with formulation 12 (Fig. 8c). The expression of STE11 in formulation 1 was the lowest of the formulations tested and was near expression levels of the control group. Downstream of the MAPK cascade, the transcription factor STE12 (sterile 12) activates genes involved in cell adhesion, elongation, and morphological transitions.33TEC1 (transcription enhancer complex 1) encodes a transcription factor (Tec1p) that forms a complex with Ste12p to regulate a subset of filamentation-specific genes, including FLO11.33,34 An upregulation in these genes suggests activation of the filamentation pathway in the MAPK cascade and represents a shift towards complex colony formation and cell adhesion in yeast.33 Expression levels of STE12 from yeast in chosen particle compositions was not significantly different, although average expression was highest in formulation 12, followed by formulation 9, then formulation 10, and formulation 1 (Fig. 8d). For TEC1 expression, levels were highest in formulation 9, then formulation 1, which were higher than controls followed by formulation 10 and lowest in formulation 12, which were lower than control levels (Fig. 8e).
In the Ras-cAMP-PKA signaling pathway, RAS2 (rat sarcoma 2) encodes a small GTPase that activates adenylate cyclase to increase intracellular cAMP levels, leading to activation of protein kinase A (PKA).35,46 An upregulation in RAS2 expression suggests activation of the Ras-cAMP-PKA cascade in response to nutrient limitation.35 There was a significant (p = 0.006) increase in expression of RAS2 in formulation 12 compared to controls (Fig. 8f). The next highest expression of RAS2 was seen in formulation 9, followed by formulation 10 and finally formulation 1 with the lowest expression levels. Three prime protein kinase 2 (TPK2) is one of three catalytic subunits of PKA and is specifically associated with regulation of filamentous growth and cell differentiation following Ras-cAMP-PKA activation.36 An upregulation in TPK2 suggests activation of enhanced filamentous growth pathways in the Ras-cAMP-PKA cascade that may contribute to biofilm colony-like structures.43 There was significant upregulation in expression of TPK2 in both formulation 1 (p = 0.0032) and formulation 12 (p = 0.0097) compared to control (Fig. 8g). Expression levels of TPK2 in formulation 9 and 10 were similar and less than formulation 1 and 12.
In the NCR pathway, glutamine metabolism 3 (GLN3) encodes a transcription factor that activates the expression of genes required for alternative nitrogen source utilization under nitrogen-limiting conditions.37 Upregulation of GLN3 suggests the activation of nitrogen-scavenging programs which may contribute to adaptive colony formation under nitrogen limited conditions.43 The highest upregulation of GLN3 was observed in formulation 12, followed by formulation 1, then formulation 9 and finally formulation 10 (Fig. 8h).
Finally, expression of FLO11 and RIM15, key downstream indicators of colony formation behavior regulated by multiple pathways was measured. FLO11, an essential genetic marker of colony formation, encodes cell surface glycoprotein necessary for both cell–cell and cell–substrate adhesion during colony development.47 Upregulation of FLO11 indicates activation of adhesion and colony formation gene programs and is essential for colony growth in response to environmental stressors.43 All formulations exhibited comparable FLO11 expression levels, with formulation 12 showing a trend toward elevated expression (Fig. 8i). RIM15 encodes a nutrient-responsive kinase that integrates signals from multiple pathways.48 While RIM15 is not essential for colony growth like FLO11, it plays an important role in regulating FLO11 expression.31,43 Upregulation of RIM15 suggests a shift toward a nutrient-limited or stress-adaptive transcriptional state and modulation of FLO11 expression. There was a significant (p = 0.0303) increase in expression of RIM15 in formulation 12 compared to controls (Fig. 8j). The next highest expression levels of RIM15 were seen in formulation 10, followed by formulation 9, and formulation 1.
4. Discussion
The goal of this study was to develop an alginate formulation with improved stability under intestinal conditions that maintains encapsulated yeast viability. To our knowledge, this is one of the first studies to demonstrate that controlling yeast colony size within alginate particles can be used to enhance cell survival. This strategy may be broadly applicable to live microbial delivery systems.Our study systematically optimized alginate hydrogel particles for encapsulating live yeast cells for oral delivery applications. We first confirmed that alginate particles exhibit well-characterized, cation-dependent crosslinking behavior and pH-responsiveness suitable for gastrointestinal transit.15,18 Optimal particle formation required a minimum calcium concentration of 0.1 M; however, at higher calcium concentrations and alginate concentrations >2%, viscosity and tail formation limited handling and extrusion. We therefore selected 2% alginate and 0.1 M calcium as a baseline for further experiments.
Consistent with previous reports, alginate particles demonstrated pH-responsive swelling and shrinkage, protecting encapsulated cells under gastric conditions and enabling cargo release in the intestines.15 Yeast encapsulation was successful using mild fabrication conditions. We observed that cell loading scaled with initial yeast input, supporting a tunable loading strategy. Importantly, encapsulation did not significantly impair yeast viability, and particles provided substantial protection against low-pH stress compared to free cells.
To achieve particle sizes suitable for oral delivery, we integrated electrospray with fine needle extrusion. Increasing voltage substantially reduced particle diameter, enabling production of particles <300 μm, appropriate for both murine and human studies.49 Needle gauge and flow rate further influenced particle size, with a 27G needle at 9 kV and a 4 mL h−1 flow rate yielding uniform particles (∼191 μm average diameter) with a high throughput. Higher yeast loading slightly increased particle size but remained within acceptable limits.
A major limitation of alginate particles is their susceptibility to disintegration in the presence of competing cations in the gastrointestinal tract.19,38 To address this, we screened multiple particle modifications for enhanced stability. Barium crosslinking markedly improved stability, consistent with its higher binding affinity to alginate compared to calcium. A range of divalent cations can be used to crosslink alginate and we initially chose a calcium crosslinker, the most widely adopted crosslinker for its biocompatibility.50 While crosslinkers like lead and copper have the highest affinity for alginate, their toxicity limits their use. Barium provides a compromise, offering higher affinity for alginate than calcium and lower toxicity than lead and copper;50 however, its potential toxicity to encapsulated yeast warranted further investigation to confirm its feasibility. Although barium crosslinking appeared to be the main driver of stability, in calcium-crosslinked compositions, chitosan and poly-L-lysine coatings also trended toward improved stability, though not statistically significant across all conditions. Chitosan and poly-L-lysine are positively charged polycations that form polyelectrolyte complexes with alginate, enhancing GI stability and reducing degradative diffusion.50 This behavior likely explains the improved stability observed with these coatings in our results.
We further evaluated how fabrication parameters affected cell loading and viability. Coating with poly-L-lysine significantly reduced both cell loading efficiency and viability, likely due to its known potential cytotoxicity.51 In contrast, chitosan coating preserved viability and enhanced loading. Barium crosslinking did not adversely affect yeast viability, supporting its suitability for stable, cell-supportive particles. Under nutrient-rich conditions, all formulations except poly-L-lysine-coated particles supported robust yeast growth over 96 hours. Under nutrient-poor conditions, cell viability declined over time, but chitosan-coated compositions showed a trend of increased resistance to cell death compared to uncoated and PLL-coated particles, although not statistically significant.
Interestingly, we observed the formation of yeast colonies within particles, a phenomenon linked to enhanced cell survival.43 Colony growth behavior and morphology differed amongst particle compositions. For most compositions with no coating and chitosan coatings, a majority of colony expansion happened in the first 24 hours followed by gradual expansion over 96 hours. In other compositions, particularly those coated with PLL, no visible colonies were observed over 96 hours. Partial penetration of PLL into the alginate matrix prior to complete particle dissolution cannot be ruled out and may have influenced yeast viability. Furthermore, during particle dissolution to assess yeast viability, PLL likely exhibited its antifungal mechanisms on the liberated yeast and induced membrane and cell wall damage in the yeast, triggering a stress response.41 These mechanisms likely reduced cell viability that was not recovered in most cases. However, formulation 12 appears to have caused moderate, rather than lethal, damage. Sublethal stress of this kind is known to activate the yeast cell wall integrity MAP kinase pathway, which initiates a compensatory response to reinforce the cell wall, restore membrane function, and promote cell survival under stress.45 This activation could explain why colonies were not observed at first but rapidly grew after 24 hours into the largest colonies seen in any composition. Future studies should monitor cell behavior without the need for particle dissolution (i.e., use live/dead stains) to assess differences in the internal colony behavior across various formulations.
Particle diameter was significantly larger in particles with a higher concentration of alginate, a known phenomenon due to solution viscosity and greater polymer content per droplet.52 We found no significant relationship between alginate particle size and colony diameter, indicating colony growth was likely independent of spatial constraints within our particles. Colony formation correlated with higher overall viability, suggesting a potential stress-adaptive behavior akin to biofilm formation observed in other microbial systems.53,54
To investigate the genetic basis of colony formation in our particle systems, we examined key genes associated with stress-induced cellular responses that typically drive biofilm-like colony development. Formulation 12 exhibited significant upregulation of genes associated with the MAP kinase pathway, STE11, and the Ras-cAMP-PKA pathway, RAS2 and TPK2, as well as increased expression of RIM15, a downstream regulator involved in the activation of cell adhesion genes like FLO11. This formulation also had the highest upregulation of STE12, associated with the MAP kinase pathway, GLN3, associated with the NCR pathway, and FLO11, an essential marker of colony formation although these levels are non-significant. This formulation showed slow colony growth and reduced viability initially, an indicator of increased exposure to environmental stressors. As other compositions coated with PLL showed a similar slow growth, exposure to the PLL coating is likely a contributing factor. Upregulation of these genes at 24 hours suggests activation of the MAP kinase, Ras-cAMP-PKA, and NCR pathways in response to these stressors, marking a transition from a stress-tolerant, low growth state to a stress-adaptive colony-forming program. Notably, this particle formulation exhibited the highest expression of these pathway genes and formed the largest colonies of any composition, suggesting a strong link between pathway activation and its unique robust colony-forming behavior.
Additionally, formulation 1 showed significant upregulation in TPK2, a component of the Ras-cAMP-PKA pathway, along with high expression of FLO11 and GLN3, second only to formulation 12. This composition also exhibited the second highest colony size by the end of the experiment, suggesting a possible link between gene expression and colony growth.
In summary, we developed a robust, tunable alginate particle system optimized for oral delivery of live yeast. Key factors influencing particle performance included crosslinker type, coating, and fabrication parameters. Barium-crosslinked, chitosan-coated particles emerged as the most promising formulation, balancing particle stability, cell viability, and oral delivery suitability. Future studies will further characterize the biological mechanisms underlying yeast colony formation in hydrogels and evaluate in vivo performance of optimized particles.
5. Conclusions
The results of these studies offer a method for optimizing hydrogel compositions for oral delivery of LBPs and more specifically a study of conditions that can improve polymer stability in the GI tract and that result in high cell viability and function. This could be used to deliver biologics or probiotics to the GI tract as a treatment of diseases like IBD, colon cancer and many other conditions that are starting to be shown to be implicated by activities in the gut microbiome which is an exciting new field of study that could be very influential in how we treat inflammatory conditions in the future.Conflicts of interest
There are no conflicts of interest to declare.Data availability
All raw data and images for this article are available at https://doi.org/10.15139/S3/T79UDV.Supplementary information (SI): images of particle compositions in simulated gastric fluid over time (PDF). See DOI: https://doi.org/10.1039/d5pm00288e.
Acknowledgements
This work was supported by the NIH 1R01DK140954. AM acknowledges financial support through the UNC Royster Society of Fellows Joseph E. Pogue Fellowship. SEM images were acquired at the Chapel Hill Analytical and Nanofabrication Laboratory (CHANL) core. Illustrations were created with BioRender.com. Microsoft Copilot was used to improve clarity and readability in certain sections of the manuscript. All AI-assisted content was verified and edited by the authors to ensure accuracy and integrity.References
- X. Yan, X. Liu, C. Zhao and G. Q. Chen, Applications of synthetic biology in medical and pharmaceutical fields, Signal Transduction Targeted Ther., 2023, 8(1), 199, DOI:10.1038/s41392-023-01440-5.
- M. Cordaillat-Simmons, A. Rouanet and B. Pot, Live biotherapeutic products: the importance of a defined regulatory framework, Exp. Mol. Med., 2020, 52(9), 1397–1406, DOI:10.1038/s12276-020-0437-6.
- M. R. Charbonneau, V. M. Isabella, N. Li and C. B. Kurtz, Developing a new class of engineered live bacterial therapeutics to treat human diseases, Nat. Commun., 2020, 11(1), 1738, DOI:10.1038/s41467-020-15508-1.
- Y. Lee, H. G. Koh, K. H. Kim, Y. S. Jin, B. H. Sung and J. Kim, Enhancing the persistence of engineered biotherapeutics in the gut: Adhesion, glycan metabolism, and environmental resistance, Adv. Drug Delivery Rev., 2025, 221, 115591, DOI:10.1016/j.addr.2025.115591.
- A. F. Miller, S. S. Potluru, S. M. Thormann, Y. Wang and J. Nguyen, Advanced adhesion and targeting strategies to prolong gut residence time and improve eLBP efficacy in colonic diseases, Adv. Drug Delivery Rev., 2025, 227, 115722, DOI:10.1016/j.addr.2025.115722.
- J. W. Rutter, L. Dekker, K. A. Owen and C. P. Barnes, Microbiome engineering: engineered live biotherapeutic products for treating human disease, Front. Bioeng. Biotechnol., 2022, 10, 1000873, DOI:10.3389/fbioe.2022.1000873.
- M. L. Palma, D. Zamith-Miranda, F. S. Martins, F. A. Bozza, L. Nimrichter, M. Montero-Lomeli, E. T. A. Marques and B. Douradinha, Probiotic Saccharomyces cerevisiae strains as biotherapeutic tools: is there room for improvement?, Appl. Microbiol. Biotechnol., 2015, 99(16), 6563–6570, DOI:10.1007/s00253-015-6776-x.
- M. Zhao, J. Ma, L. Zhang and H. Qi, Engineering strategies for enhanced heterologous protein production by Saccharomyces cerevisiae, Microb. Cell Fact., 2024, 23(1), 32, DOI:10.1186/s12934-024-02299-z.
- M. Duman-Scheel, Saccharomyces cerevisiae (Baker’s Yeast) as an Interfering RNA Expression and Delivery System, Curr. Drug Targets, 2019, 20(9), 942–952, DOI:10.2174/1389450120666181126123538.
- M. K. Heavey, A. Hazelton, Y. Wang, M. Garner, A. C. Anselmo, J. C. Arthur and J. Nguyen, Targeted delivery of the probiotic Saccharomyces boulardii to the extracellular matrix enhances gut residence time and recovery in murine colitis, Nat. Commun., 2024, 15(1), 3784, DOI:10.1038/s41467-024-48128-0.
- T. Culpepper, K. Senthil, J. Vlcek, A. Hazelton, M. K. Heavey, R. S. Sellers, J. Nguyen and J. C. Arthur, Engineered Probiotic Saccharomyces boulardii Reduces Colitis-Associated Colorectal Cancer Burden in Mice, Dig. Dis Sci., 2025, 70(7), 2348–2367, DOI:10.1007/s10620-025-09008-9.
- M. N. Hasan, S. M. S. Shahriar, J. Mondal, M. Nurunnabi and Y.-k. Lee, Bioinspired and biomimetic materials for oral drug delivery, in Bioinspired and Biomimetic Materials for Drug Delivery, ed. M. Nurunnabi, Woodhead Publishing, 2021, pp. 89–104 Search PubMed.
- S. Han, Y. Lu, J. Xie, Y. Fei, G. Zheng, Z. Wang, J. Liu, L. Lv, Z. Ling and B. Berglund, et al., Probiotic Gastrointestinal Transit and Colonization After Oral Administration: A Long Journey, Front. Cell. Infect. Microbiol., 2021, 11, 609722, DOI:10.3389/fcimb.2021.609722.
- M. K. Heavey, D. Durmusoglu, N. Crook and A. C. Anselmo, Discovery and delivery strategies for engineered live biotherapeutic products, Trends Biotechnol., 2022, 40(3), 354–369, DOI:10.1016/j.tibtech.2021.08.002.
- M. T. Cook, G. Tzortzis, D. Charalampopoulos and V. V. Khutoryanskiy, Microencapsulation of probiotics for gastrointestinal delivery, J. Controlled Release, 2012, 162(1), 56–67, DOI:10.1016/j.jconrel.2012.06.003.
- A. Rodrigo-Navarro, S. Sankaran, M. J. Dalby, A. Del Campo and M. Salmeron-Sanchez, Engineered living biomaterials, Nat. Rev. Mater., 2021, 6(12), 1175–1190, DOI:10.1038/s41578-021-00350-8.
- X. Liu, M. E. Inda, Y. Lai, T. K. Lu and X. Zhao, Engineered Living Hydrogels, Adv. Mater., 2022, 34(26), 2201326, DOI:10.1002/adma.202201326.
- P. Severino, C. F. Da Silva, L. N. Andrade, D. De Lima Oliveira, J. Campos and E. B. Souto, Alginate Nanoparticles for Drug Delivery and Targeting, Curr. Pharm. Des., 2019, 25(11), 1312–1334, DOI:10.2174/1381612825666190425163424.
- K. Y. Lee and D. J. Mooney, Alginate: Properties and biomedical applications, Prog. Polym. Sci., 2012, 37(1), 106–126, DOI:10.1016/j.progpolymsci.2011.06.003.
- X. Wang, S. Gao, S. Yun, M. Zhang, L. Peng, Y. Li and Y. Zhou, Microencapsulating Alginate-Based Polymers for Probiotics Delivery Systems and Their Application, Pharmaceuticals, 2022, 15(5), 644, DOI:10.3390/ph15050644.
- A. Bevilacqua, D. Campaniello, B. Speranza, A. Racioppo, C. Altieri, M. Sinigaglia and M. R. Corbo, Microencapsulation of Saccharomyces cerevisiae into Alginate Beads: A Focus on Functional Properties of Released Cells, Foods, 2020, 9(8), 1051 CrossRef CAS PubMed.
- B. Speranza, M. R. Corbo, D. Campaniello, C. Altieri, M. Sinigaglia and A. Bevilacqua, Biofilm formation by potentially probiotic Saccharomyces cerevisiae strains, Food Microbiol., 2020, 87, 103393, DOI:10.1016/j.fm.2019.103393.
- M. W. Harding, L. L. Marques, R. J. Howard and M. E. Olson, Can filamentous fungi form biofilms?, Trends Microbiol., 2009, 17(11), 475–480, DOI:10.1016/j.tim.2009.08.007.
- M. K. Heavey and A. C. Anselmo, Modulating Oral Delivery and Gastrointestinal Kinetics of Recombinant Proteins via Engineered Fungi, AAPS J., 2021, 23(4), 76, DOI:10.1208/s12248-021-00606-9.
- J. L. Patarroyo, J. S. Florez-Rojas, D. Pradilla, J. D. Valderrama-Rincón, J. C. Cruz and L. H. Reyes, Formulation and characterization of gelatin-based hydrogels for the encapsulation of kluyveromyces lactis-Applications in packed-bed reactors and probiotics delivery in humans, Polymers, 2020, 12(6), 1287, DOI:10.3390/POLYM12061287.
- V. Calero, P. M. Rodrigues, T. Dias, A. Ainla, A. Vilaça, L. Pastrana, M. Xavier and C. Gonçalves, A miniaturised semi-dynamic in vitro model of human digestion, Sci. Rep., 2024, 14(1), 11923, DOI:10.1038/s41598-024-54612-w.
- H. Choukaife, A. A. Doolaanea and M. Alfatama, Alginate Nanoformulation: Influence of Process and Selected Variables, Pharmaceuticals, 2020, 13(11), 335, DOI:10.3390/ph13110335.
- S.-L. Huang and Y.-S. Lin, The Size Stability of Alginate Beads by Different Ionic Crosslinkers, Adv. Mater. Sci. Eng., 2017, 2017(1), 9304592, DOI:10.1155/2017/9304592.
- M. I. García-Briega, J. Ródenas-Rochina, L. A. Martins, S. Lanceros-Méndez, G. Gallego Ferrer, A. Sempere and J. L. Gómez Ribelles, Stability of Biomimetically Functionalised Alginate Microspheres as 3D Support in Cell Cultures, Polymers, 2022, 14(20), 4282, DOI:10.3390/polym14204282.
- C. Fischer, O. Valerius, H. Rupprecht, M. Dumkow, S. Krappmann and G. H. Braus, Posttranscriptional regulation of FLO11 upon amino acid starvation in Saccharomyces cerevisiae, FEMS Yeast Res., 2008, 8(2), 225–236, DOI:10.1111/j.1567-1364.2007.00331.x.
- D. Watanabe, A. Kaneko, Y. Sugimoto, S. Ohnuki, H. Takagi and Y. Ohya, Promoter engineering of the Saccharomyces cerevisiae RIM15 gene for improvement of alcoholic fermentation rates under stress conditions, J. Biosci. Bioeng., 2017, 123(2), 183–189, DOI:10.1016/j.jbiosc.2016.08.004.
- S. Fauquenoy, V. Migeot, O. Finet, C. Yague-Sanz, O. Khorosjutina, K. Ekwall and D. Hermand, Repression of Cell Differentiation by a cis-Acting lincRNA in Fission Yeast, Curr. Biol., 2018, 28(3), 383–391, DOI:10.1016/j.cub.2017.12.048.
- D. Jung, J. S. Seo, J. Nam and J. Kim, Functional association of Loc1 and Puf6 with RNA helicase Dhh1 in translational regulation of Saccharomyces cerevisiae Ste12, PLoS One, 2019, 14(7), e0220137, DOI:10.1371/journal.pone.0220137.
- C. Louw, P. R. Young, P. van Rensburg and B. Divol, Regulation of endo-polygalacturonase activity in Saccharomyces cerevisiae, FEMS Yeast Res., 2010, 10(1), 44–57, DOI:10.1111/j.1567-1364.2009.00584.x.
- L. M. Sun and K. Liao, Saccharomyces cerevisiae Hog1 MAP kinase pathway is activated in response to honokiol exposure, J. Appl. Microbiol., 2018, 124(3), 754–763, DOI:10.1111/jam.13649.
- J. Dong, X. M. Fu, P. F. Wang, S. S. Dong, X. Li, D. G. Xiao and C. Y. Zhang, Construction of industrial baker’s yeast with high level of cAMP, J Food Biochem., 2019, 43(7), e12846, DOI:10.1111/jfbc.12846.
- Y. Chen, W. Zeng, W. Ma, W. Ma and J. Zhou, Chromatin Regulators Ahc1p and Eaf3p Positively Influence Nitrogen Metabolism in Saccharomyces cerevisiae, Front. Microbiol., 2022, 13, 883934, DOI:10.3389/fmicb.2022.883934.
- A. Hurtado, A. A. A. Aljabali, V. Mishra, M. M. Tambuwala and A. Serrano-Aroca, Alginate: Enhancement Strategies for Advanced Applications, Int. J. Mol. Sci., 2022, 23(9), 4486, DOI:10.3390/ijms23094486.
- B. Alallam, S. Altahhan, M. Taher, M. H. Mohd Nasir and A. A. Doolaanea, Electrosprayed Alginate Nanoparticles as CRISPR Plasmid DNA Delivery Carrier: Preparation, Optimization, and Characterization, Pharmaceuticals, 2020, 13, 158 Search PubMed.
- S. K. Boda, X. Li and J. Xie, Electrospraying an enabling technology for pharmaceutical and biomedical applications: A review, J. Aerosol Sci., 2018, 125, 164–181, DOI:10.1016/j.jaerosci.2018.04.002.
- F. Padilla-Garfias, L. Rios-Cifuentes, N. S. Sanchez, M. Calahorra and A. Pena, Study of the mechanism of epsilon-poly-L-lysine as an antifungal on Candida albicans and Saccharomyces cerevisiae, Biochim. Biophys. Acta, Gen. Subj., 2022, 1866(10), 130197, DOI:10.1016/j.bbagen.2022.130197.
- T. Gaizer, J. Juhász, B. Pillér, H. Szakadáti, C. I. Pongor and A. Csikász-Nagy, Integrative analysis of yeast colony growth, Commun. Biol., 2024, 7(1), 511, DOI:10.1038/s42003-024-06218-1.
- J. A. Granek and P. M. Magwene, Environmental and genetic determinants of colony morphology in yeast, PLoS Genet., 2010, 6(1), e1000823, DOI:10.1371/journal.pgen.1000823.
- A. E. Beeser and T. G. Cooper, Control of nitrogen catabolite repression is not affected by the tRNAGln-CUU mutation, which results in constitutive pseudohyphal growth of Saccharomyces cerevisiae, J. Bacteriol., 1999, 181(8), 2472–2476, DOI:10.1128/JB.181.8.2472-2476.1999.
- C. Jin, S. Kim, S. Willis and K. Cooper, The MAPKKKs Ste11 and Bck1 jointly transduce the high oxidative stress signal through the cell wall integrity MAP kinase pathway, Microb. Cell, 2015, 2(9), 329–342, DOI:10.15698/mic2015.09.226.
- H. U. Mosch, E. Kubler, S. Krappmann, G. R. Fink and G. H. Braus, Crosstalk between the Ras2p-controlled mitogen-activated protein kinase and cAMP pathways during invasive growth of Saccharomyces cerevisiae, Mol. Biol. Cell, 1999, 10(5), 1325–1335, DOI:10.1091/mbc.10.5.1325.
- W. S. Lo and A. M. Dranginis, The cell surface flocculin Flo11 is required for pseudohyphae formation and invasion by Saccharomyces cerevisiae, Mol. Biol. Cell, 1998, 9(1), 161–171, DOI:10.1091/mbc.9.1.161.
- S. Nagarajan, A. L. Kruckeberg, K. H. Schmidt, E. Kroll, M. Hamilton, K. McInnerney, R. Summers, T. Taylor and F. Rosenzweig, Uncoupling reproduction from metabolism extends chronological lifespan in yeast, Proc. Natl. Acad. Sci. U. S. A., 2014, 111(15), E1538–E1547, DOI:10.1073/pnas.1323918111.
- S. F. Jang, B. A. Goins, W. T. Phillips, C. Santoyo, A. Rice-Ficht and J. T. McConville, Size discrimination in rat and mouse gastric emptying, Biopharm. Drug Dispos., 2013, 34(2), 107–124, DOI:10.1002/bdd.1828 , (acccessed 2024/01/12/17:53:56).
- S. H. Ching, N. Bansal and B. Bhandari, Alginate gel particles–A review of production techniques and physical properties, Crit. Rev. Food Sci. Nutr., 2017, 57(6), 1133–1152, DOI:10.1080/10408398.2014.965773.
- G. Orive, S. K. Tam, J. L. Pedraz and J.-P. Hallé, Biocompatibility of alginate–poly-L-lysine microcapsules for cell therapy, Biomaterials, 2006, 27(20), 3691–3700, DOI:10.1016/j.biomaterials.2006.02.048.
- A. Letocha, A. Michalczyk, M. Miastkowska and E. Sikora, Effect of Encapsulation of Lactobacillus casei in Alginate-Tapioca Flour Microspheres Coated with Different Biopolymers on the Viability of Probiotic Bacteria, ACS Appl. Mater. Interfaces, 2024, 16(39), 52878–52893, DOI:10.1021/acsami.4c10187.
- V. S. Tadimarri, T. A. Tyagi, C. N. Duong, S. Rasheed, R. Muller and S. Sankaran, Adaptations of Gram-negative and Gram-positive Probiotic Bacteria in Engineered Living Materials, ACS Biomater. Sci. Eng., 2025, 11(6), 3773–3784, DOI:10.1021/acsbiomaterials.5c00325.
- V. Plocek, L. Vachova, V. Stovicek and Z. Palkova, Cell Distribution within Yeast Colonies and Colony Biofilms: How Structure Develops, Int. J. Mol. Sci., 2020, 21(11), 3873, DOI:10.3390/ijms21113873.
| This journal is © The Royal Society of Chemistry 2026 |