Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Rapid plasma membrane reorganisation and endocytosis in HER2 breast cancer cells incubated with trastuzumab decorated polymer nanoparticles
Carwyn S.
Hughes
a,
Saeed
Tayeb
ab,
Duncan
Muir
c,
Anthony J.
Hayes
d,
Peter
Watson
d and
Arwyn T.
Jones
 *a
*a
aSchool of Pharmacy and Pharmaceutical Sciences, Cardiff University, Cardiff, UK. E-mail: jonesat@cardiff.ac.uk
bDepartment of Pharmaceutical Sciences, College of Pharmacy, Umm Al-Qura University, Makkah 24381, Saudi Arabia
cSchool of Earth Sciences, Cardiff University, Cardiff, UK
dSchool of Biosciences, Cardiff University, Cardiff, UK
First published on 10th February 2026
Abstract
Knowledge on HER2+ breast cancer biology has informed drug design leading to targeted therapies giving improved clinical outcomes. Drug resistance and disease relapse, however still drive a continuous need for more efficacious and reliable therapeutics. Nanoparticles (NPs) as HER2 targeting nanomedicines offer new hope for selective targeting of HER2 within and beyond solid tumours, together with concomitant delivery of therapeutic cargo. For initial preclinical characterisation, studies on NP endocytosis and drug delivery are often performed in HER2+ breast cancer cell models, but information on initial NP-HER2 dynamics at the plasma membrane and how this impacts endocytic uptake and delivery efficiency is largely missing. Here using polymer poly(lactic-co-glycolide) acid NPs decorated with different valencies of the HER2 targeting monoclonal antibody trastuzumab, we have designed approaches to immediately study the impact of NP-HER2 targeting on high and low HER2 expressing breast cancer cell models. Using resonant scanning confocal imaging of live cell plasma membrane dynamics, we show in very high detail and within 10 minutes of cell exposure of the receptor to the NPs, extensive blebbing and ruffling of the plasma membrane, manifesting before much longer uptake of the NPs into the cell interior. Plasma membrane reorganisation was rapidly reversible, with cells reaching baseline morphology in 30 minutes. Our findings were confirmed at the ultrastructural level by scanning electron microscopy in cells fixed within 10 and 30 minutes of exposure to the NPs. Endocytic traffic of the NPs was in part directed to lysosomes and we discover a relationship between antibody valency and the ability of the NPs to deliver the chemotherapeutic agent doxorubicin to mediate cell death. Knowledge gained from these studies offers new approaches to study NP-cell dynamics in different NP-receptor settings and how receptor targeting influences plasma membrane organisation, endocytosis and delivery.
Introduction
Breast cancer remains a global health concern with 2.3 million new cases and 670![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 deaths reported in 2022. Of these cases, 15% were classified as HER2+, representing cells that overexpress the HER2 growth factor receptor.1 In 1998 the HER2 targeting monoclonal antibody trastuzumab was FDA approved for treating HER2+ breast cancer and despite improving therapeutic outcomes, resistance to trastuzumab has emerged as a major limitation.2,3 Efforts to address this challenge have included using trastuzumab as an antibody drug conjugate with Kadcyla representing an example, with the antibody attached to the microtubule disrupting drug emtansine.4 The success of treating patients with trastuzumab alone or as an ADC largely relies on inhibiting the ability of HER2 to signal growth and cell division, to mediate antibody mediated cell cytotoxicity and also endocytosis of the trastuzumab-HER2 complex to endolysosomes to enable payload release and also proteolytic degradation, and thus downregulation, of the receptor. This for HER2 is a particular challenge as the receptor has inherent resistance to endocytosis mediated by a number of factors.5–7
000 deaths reported in 2022. Of these cases, 15% were classified as HER2+, representing cells that overexpress the HER2 growth factor receptor.1 In 1998 the HER2 targeting monoclonal antibody trastuzumab was FDA approved for treating HER2+ breast cancer and despite improving therapeutic outcomes, resistance to trastuzumab has emerged as a major limitation.2,3 Efforts to address this challenge have included using trastuzumab as an antibody drug conjugate with Kadcyla representing an example, with the antibody attached to the microtubule disrupting drug emtansine.4 The success of treating patients with trastuzumab alone or as an ADC largely relies on inhibiting the ability of HER2 to signal growth and cell division, to mediate antibody mediated cell cytotoxicity and also endocytosis of the trastuzumab-HER2 complex to endolysosomes to enable payload release and also proteolytic degradation, and thus downregulation, of the receptor. This for HER2 is a particular challenge as the receptor has inherent resistance to endocytosis mediated by a number of factors.5–7
We previously identified that strategic crosslinking of HER2 on the plasma membrane of HER2+ breast cancer cells incubated with trastuzumab resulted in a dramatic increase in endocytosis of the antibody and receptor leading to lysosomal degradation of the latter.8,9 This opens up opportunities for further studies, such as those showing that engineered biparatropic antibodies can mediate the same effects,10 or the possibility of combining multiple antibodies on a single platform. For this, nanoparticles offer an approach that can harness the benefits of selective cell targeting via, for example ligand conjugation, with encapsulation of a drug for its uptake and intracellular release from endolysosomes. NPs decorated with trastuzumab have been developed for carrying a number of drugs, based on polymer, liposome and inorganic frameworks.11–13 A N-(2-hydroxypropyl)methacrylamide (HPMA) polymer formulation decorated with a HER2 targeting polypeptide affibody was designed based on the hypothesis that it could mediate HER2 crosslinking.14 Developed formulations showed very high capacity for internalisation to HER2+ ovarian cancer cells with concomitant paclitaxel delivery to cause cell toxicity.
The described in vitro characterisation of nanotherapeutics targeting breast and other cancers is most often based on initial pre-clinical analysis in suitable cell lines and progression to in vivo studies in xenograft models. Very few of these or other NP-cell studies have attempted the challenge of immediately (minutes rather than hours/days) analysing their effects on cells and how this could influence uptake and cargo delivery. Analysis of these early timepoints is important to understand how cells immediately respond to the drug delivery challenge. It, for example, informs on how activation of the plasma membrane could dictate the mechanism of uptake that will ultimately influence endocytic traffic to a specific subcellular destination – this will affect its performance as a therapeutic entity. In this study we prepared a range of PLGA based nanoparticles containing different trastuzumab valencies and developed a Resonant Scanning Confocal Microscopy technique to demonstrate in living cells how binding to HER2 caused, within 10 minutes, extensive and reversible plasma membrane remodelling. The extent of the impact on the plasma membrane also influenced the ability to which they were able to deliver doxorubicin to cause cell toxicity.
Experimental
Materials
Unless otherwise described all materials were from Fisher Scientific.Cell culture
All cell lines were obtained directly from LCG Standards UK, the UK supplier of authenticated American Type Culture Collection (ATCC) cells. SKBR3 and BT474 breast cancer cells were used as the high expressing models (HER2++) positive models whilst MCF7 cells were used as a low-HER2 expression model.15 All cells were cultured at 37 °C in a 5% CO2 humidified atmosphere (referred to as tissue culture conditions) in high-glucose Dulbecco's Modified Eagle Medium (DMEM) containing phenol red and supplemented with 10% v/v foetal bovine serum (FBS)-designated complete medium. Cells were routinely passaged upon reaching 70% confluency. For live cell imaging the media was replaced with phenol red free RPMI supplemented with 10% v/v FBS; here referred to as imaging medium.NP manufacture & characterisation
For PLGA NP manufacture 50 mg of both poly(lactic-co-glycolic) acid (PLGA, lactide![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) glycolide 50
glycolide 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50, Mw 30
50, Mw 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–60
000–60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, P2191 Sigma-Aldrich) and pegylated PLGA-COOH (PEG average Mn 5000, PLGA average Mn 15
000, P2191 Sigma-Aldrich) and pegylated PLGA-COOH (PEG average Mn 5000, PLGA average Mn 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, lactide
000, lactide![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) glycolide 50
glycolide 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50, 902071 Sigma-Aldrich) were dissolved in 8 ml of acetone which was then added dropwise to a 14 ml 2% PVA w/v (Mw 89
50, 902071 Sigma-Aldrich) were dissolved in 8 ml of acetone which was then added dropwise to a 14 ml 2% PVA w/v (Mw 89![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000–98
000–98![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000, 98+% hydrolyzed, 341584 Sigma Aldrich) solution in water and left to stir overnight – depending on NP cargo, 10 mg of Doxorubicin (Dox) (Stratech Scientific) or 1 mg of rhodamineB (RhB) (Sigma-Aldrich) was added to the PVA solution to prepare respectively PLGA-Dox or PLGA-RhB. Following acetone evaporation, the NPs were centrifuged at 14
000, 98+% hydrolyzed, 341584 Sigma Aldrich) solution in water and left to stir overnight – depending on NP cargo, 10 mg of Doxorubicin (Dox) (Stratech Scientific) or 1 mg of rhodamineB (RhB) (Sigma-Aldrich) was added to the PVA solution to prepare respectively PLGA-Dox or PLGA-RhB. Following acetone evaporation, the NPs were centrifuged at 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 15 min at 4 °C and washed repeatedly by aspirating the supernatant then resuspending the NPs in distilled water then sonicated at 300 W on ice for 60 seconds (two 30 s intervals with 10 s pause in-between) using a Soniprep 150 probe sonicator. The NPs were then again centrifuged and washed as previously described. This was repeated for a minimum of 5 cycles. NP concentration was determined using a Malvern Nanosight LM10 nanoparticle tracking analysis. NPs and Trastuzumab (Tz, a kind gift from the Velindre Cancer centre, Cardiff UK) were added together in a 10
000g for 15 min at 4 °C and washed repeatedly by aspirating the supernatant then resuspending the NPs in distilled water then sonicated at 300 W on ice for 60 seconds (two 30 s intervals with 10 s pause in-between) using a Soniprep 150 probe sonicator. The NPs were then again centrifuged and washed as previously described. This was repeated for a minimum of 5 cycles. NP concentration was determined using a Malvern Nanosight LM10 nanoparticle tracking analysis. NPs and Trastuzumab (Tz, a kind gift from the Velindre Cancer centre, Cardiff UK) were added together in a 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 or 30
1 or 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 molar ratio (Tz
1 molar ratio (Tz![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NP) on ice under anaerobic conditions prior to adding a N-hydroxysuccinimide (NHS) and 1-ethyl-3-(3-dimethylamionopropyl)carbodiimide (EDC) solution giving final concentrations of 50 mM and 100 mM respectively prior to a further 3 h incubation under these conditions, providing us with Tz decorated NPs. The NPs were centrifuged, washed, sonicated and their concentration remeasured. Size and zeta potential measurements were performed using a Malvern Zetasizer Nano ZS. For morphology analysis NPs were air dried overnight onto glass coverslips that were then gold–palladium sputter coated (BioRad SC-500 for sputter coating with argon gas for plasma) and imaged using a Zeiss Sigma HD Field Emission Gun Analytical SEM. The generated NP formulations are listed in Table 1. Particle valency was determined by comparing the fluorescence of a known concentration of unloaded nanoparticles against a standard curve generated with Tz488. We assume the degree of valency to be the same between unloaded and Dox-loaded NPs.
NP) on ice under anaerobic conditions prior to adding a N-hydroxysuccinimide (NHS) and 1-ethyl-3-(3-dimethylamionopropyl)carbodiimide (EDC) solution giving final concentrations of 50 mM and 100 mM respectively prior to a further 3 h incubation under these conditions, providing us with Tz decorated NPs. The NPs were centrifuged, washed, sonicated and their concentration remeasured. Size and zeta potential measurements were performed using a Malvern Zetasizer Nano ZS. For morphology analysis NPs were air dried overnight onto glass coverslips that were then gold–palladium sputter coated (BioRad SC-500 for sputter coating with argon gas for plasma) and imaged using a Zeiss Sigma HD Field Emission Gun Analytical SEM. The generated NP formulations are listed in Table 1. Particle valency was determined by comparing the fluorescence of a known concentration of unloaded nanoparticles against a standard curve generated with Tz488. We assume the degree of valency to be the same between unloaded and Dox-loaded NPs.
| Nanoparticle | Size (nm) | PDI | Zeta potential (mV) |
|---|---|---|---|
| PLGA | 182.18 ± 35.68 | 0.14 ± 0.09 | −19.84 ± 9.09 |
| PLGA-TZ | 205.80 ± 12.85 | 0.11 ± 0.06 | −11.87 ± 11.92 |
| PLGA-RhB | 216.96 ± 21.26 | 0.17 ± 0.06 | −3.50 ± 4.01 |
| PLGA-RhB-Tz | 207.40 ± 8.03 | 0.14 ± 0.03 | 1.87 ± 0.52 |
| PLGA-Dox | 197.70 ± 27.99 | 0.16 ± 0.09 | −27.65 ± 9.26 |
| PLGA-Dox-9Tz | 193.03 ± 4.38 | 0.06 ± 0.01 | −7.13 ± 5.69 |
| PLGA-Dox-21Tz | 209.95 ± 13.30 | 0.13 ± 0.08 | −15.23 ± 2.80 |
Encapsulation efficiency
To determine the efficiency of Dox encapsulation, a portion of the NPs were dissolved in DMSO and the concentration of the Dox within was compared against a standard curve of known concentrations. Each datapoint was prepared in triplicate in a clear-bottomed 96 well plate and the absorbance at 490 nm was measured using a Tecan infinite M-plex plate reader.Binding and endocytosis of NPs
Cells were grown in MatTek dishes (MatTek Corporation) until 70% confluent, then following removal of the media they were incubated with 500 μg ml−1 PLGA-RhB-Tz in complete media under tissue culture conditions for 5 hours. Following 2× washes in imaging media the cells were imaged by confocal microscopy on a Leica SP5 confocal laser scanning microscope equipped with a 37 °C heated stage, CO2 perfused humidified chamber (ibidi), 488 nm Argon, 543/633 nm HeNe lasers and a 100× oil immersion objective. Images were then processed using FIJI. All fluorescence microscopy studies were performed on this instrument.Analysis of NP endocytosis to lysosomes
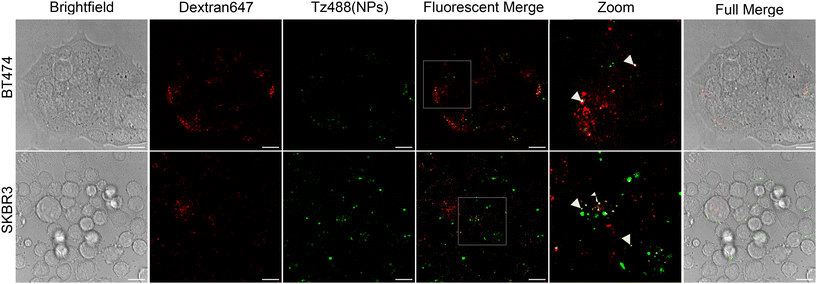
Cells were treated with 75 μg ml−1 Dextran-647 in complete medium for 2 hours, washed 3 times prior to incubation with 500 μg ml−1 PLGA-Tz488 for 24 hours before 3× washing and then immediately analysed in imaging medium by confocal microscopy as previously described.16,17 To allow for comparative analysis of endocytic traffic, cells were treated with dextran-647 and washed as previously described however rather than adding NPs they were treated with dextran-488 for 24 hours before being washed and imaged. Pearsons and Manders colocalization coefficients were obtained for each individual slices of the z-stacks taken. To calculate the coefficients the Otsu thresholding algorithm was used to threshold each individual slice and then the BIOP JACoP plugin was run in imageJ.18,19 The data shown represents coefficient values from one representative slice per individual z-stack.Analysing plasma membrane organisation
Cells were grown in complete medium in MatTek dishes until 70% confluent. Prior to staining, the cells were washed and equilibrated in imaging medium for 10 min under tissue culture conditions then stained for 10 min with 1× CellMask Deep Red (ThermoFisher). Cells were then washed with imaging medium and then treated with 500 μg ml−1 of PLGA-21Tz with imaging commencing immediately following treatment and continuing for 10 min. To minimise the possible impact of temperature changes the various treatments, washes and stains were kept at 37 °C throughout. For the monitoring of cell ruffling across a single focal plane each frame was acquired every 4.9 seconds. Quantification of these cell ruffles was done by using a plugin that measures the total cell-free area over time.20 For 3D visualization, imaging was performed utilising the resonant scanner of the microscope allowing acquisition of a complete z-stack in <30 seconds, with the data then displayed as maximum intensity projections. For temporal colour coding and mapping these projections we used the temporal colour code in FiJi.21 BitPlane Imaris software was used to generate cell surface representations of live 3D data stacks.SEM imaging of membrane ruffling
Analysis was based on our published method with some minor modifications.9 SKBR3 cells were grown on coverslips within a 12 well plate and treated with 500 μg ml−1 PLGA-21Tz in complete media for 10 min under tissue culture conditions prior to 3× washing in PBS. The cells were then immediately fixed with 2% glutaraldehyde in PBS (Sigma-Aldrich) for 30 min, washed 3× with PBS prior to post fixing in 1% v/v osmium tetroxide in PBS for a further 30 min. Cells were then washed 3× with PBS then dehydrated using sequential 10 min incubations with increasing concentrations of ethanol (50%, 60%, 70%, 80%, 90%, 95%, 100% × 3) diluted with water. The dehydrated cells were then subject to further 10 min sequential incubations using increasing concentrations of Hexamethyldisilazane (HMDS) diluted in ethanol (50%, 60%, 70%, 80%, 90%, 95%, 100% × 3) before being air dried overnight in a fume cupboard. Coverslips were then mounted onto stage inserts and gold–palladium sputter coated (BioRad SC-500 for sputter coating with argon gas for plasma) before imaging by SEM (Zeiss Sigma HD Field Emission Gun Analytical SEM) under an accelerating voltage (EHT) of 5.00 kV with probe current of <100 pA and a working distance of 5 mm.Cell viability analysis
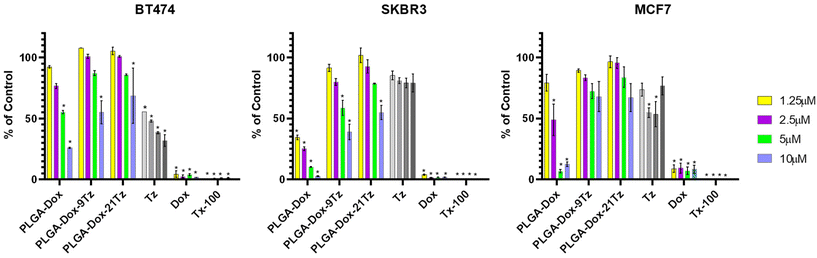
The CellTiter Blue (Promega) metabolic assay was used to determine cell viability. Cells (100 µl) were seeded in a black 96-well plate at a density of 4000 cells per well and left to grow for 48 h prior to addition of PLGA-Dox-Tz (100 μl) of increasing Tz valences (0–21 per NP). The number of particles added was determined based on Dox concentration giving final well concentrations of 0–10 μM. Each treatment group was done in duplicate and Triton X-100 (Tx-100) was used as positive cell death control. After 72 hours the well media was aspirated and replaced with 110 μl of cell titre blue solution (10 μl CTB in 100 μl complete medium). Cells were then incubated for 4 hours under tissue culture conditions and the fluorescence recorded (560Ex/590Em) on the plate reader (Tecan Infinite M Plex). Cell viability was expressed as a percentage of untreated diluent controls.Statistical analysis
A two-way ANOVA was used to assess cell viability analysis for statistical significance (p < 0.05) using Graphpad Prism. For all experiments details on the numbers of repeat experiments are provided in the figure legends.Results and discussion
PLGA, as an FDA approved polymer has been extensively studied as a biodegradable nanoparticle scaffold affording easy functionalisation that is capable of encapsulating a broad range of cargos with therapeutic interest.22,23 This interest has already seen the development of PLGA based NPs encapsulating anti-cancer agents such as doxorubicin, including those decorated with Tz for HER2 targeting.11,24–27 None of these have reached the clinic and there is a general paucity of information of cell and NP analysis immediately following their initial interaction that is likely to influence how they enter cells, by which mechanism, and how this relates to therapeutic efficacy.28,29NP characterisation
PLGA NPs variants including Tz decorated variants were generated using nanoprecipitation based on established methods and characterized as having an average size of approximately 200 nm (Table 1 and SI Fig. S1) with PDI of ≤0.2. SEM analysis supports our size measurements highlighting their spheroidal nature (Fig. 1). All NP formulations were negatively charged with the exception of those containing RhB that were closer to neutral (Table 1 and SI Fig. S1). For Dox encapsulation we observed a low encapsulation efficiency of 55% (SI Fig. S2) due to the passive encapsulation methodology utilised here as compared to higher efficiencies observed using techniques such as microfluidics or active drug loading.21,28 Overall, the data on the physicochemical nature of our NPs agrees with that of other PLGA NPs found within the literature, including those decorated with Tz.24,25,30,31 Furthermore, using PLGA NPs of this size, Shipunova et al. demonstrated that HER2 crosslinking was a feasible method to ensure the efficient delivery of doxorubicin in vivo.25PLGA-Tz NPs binding and uptake in HER2++ cells
We initially confirmed that the NPs bound to the plasma membrane in a Tz and HER2 dependant manner (Fig. 2). Confocal microscopy analysis after 5 hours of PLGA-RhB-Tz488 incubation highlighted that they have a strong affinity for HER2++ cells (SKBR3 and BT474) with very little, if any fluorescence observed in the low HER2 expressing MCF7 model. The labelling of the NPs with fluorescent Tz and RhB allowed us to identify the fluorophores as colocalising and highly enriched on the plasma membrane, often as aggregates with evidence of fluorophore separation inside the cells in smaller structures that are likely to be endolysosomes that we then characterise.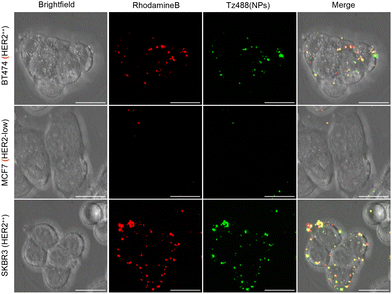 | ||
| Fig. 2 After 5 hours of treatment PLGA-RhB-Tz NPs show preferential binding to HER2++ breast cancer cells (BT474 and SKBR3). Scale bar = 20 µm. | ||
PLGA-Tz endocytosis to lysosomes
To further investigate the endocytosis of the particles and their intracellular fate we used the fluid phase endocytosis and macropinocytosis probe dextran that as a 10 kDa conjugate will enter cells and traffic to lysosomes.16,17 We initially incubated cells with this molecule as an Alexa 647 conjugate to label the entire endolysosomal pathway and then incubated the cells with the PLGA-Tz488 to identify whether they were able to reach dextran labelled lysosomes; noting that during the NP incubation phase the previously internalised dextran would have trafficked to and accumulated in this organelle. We also prepared experiments where the NPs were substituted with dextran Alexa 488 that should be trafficked to the same organelle as the Alexa 647 variant thus giving a representation of high colocalization. Data in Fig. 3 highlights in both cell lines, emissions from single fluorophores showing lysosomes labelled with dextran alone, endosome containing NPs or lysosomes containing both. Given the lack of internalisation seen in the MCF7 cells (SI Fig. S3) we consider that the uptake is driven by Tz-HER2 binding rather than via any other non-specific interactions between antibody and cell.To quantify the degree of colocalization in lysosomes we compared the coefficient value of the NPs/Dex647 against the values for cells treated with Dex488/Dex647. Both Pearson and Manders coefficients can be used to quantify colocalization however each approach has limitations.18 Both methods highlighted a higher degree of colocalization between the two dextran probes but this was not 100% most possibly due to the fact that the Pearson coefficient considers the difference between fluorophore intensity within individual endocytic structures whilst the Manders coefficient reflects the fraction of the second probe that is still residing in early and late endosomal compartments. Because of this we believe a combination of both coefficients provides a better understanding about the nature of the colocalization. Table 2 highlights the highest degree of NP-dextran647 colocalization can be seen in the SKBR3 cells (0.396) using the Manders coefficient. The higher equivalent Pearson coefficient in the BT474 cells (0.367) combined with the lower Manders coefficient (0.252) suggests that the NPs are more concentrated within the lysosomes resulting in a more intense overall signal intensity but from fewer sources. Across both cell lines whilst the estimated overlap between the two dextrans is higher and could in theory colocalize perfectly, in practice this is not the case and based on this, we have also reported the coefficients as a % of what should be the maximum colocalization achievable (SI Fig. S4). The exact mechanism by which our Tz decorated NPs enter cells is unknown. Based on their dimensions, and knowledge on endocytic portals for macromolecules it would be assumed that they would be too large for entry via clathrin-mediated or clathrin-independent endocytosis (CME or CIE). An exception is macropinocytosis and also phagocytosis, both representing processes involving outward, though distinct, rearrangement of the plasma membrane.29,32 It is also unknown as to whether the trafficking of the HER2-NP complexes follows the same route as HER2 when it forms homodimers, or when it forms heterodimers with other family members such as EGFR.29,33,34 Analysing HER2 endocytosis and recycling is challenging, with evidence that in nature it can enter via CME and CIE pathways and that it is recycled back to the plasma membrane.7 There is also little consensus as to the exact uptake mechanism of HER2 when bound to Tz or any other soluble ligand or when these are attached to drug delivery vectors. Given the size of our nanoparticles however, it is unlikely that they are internalised by either CME or CIE with most of the uptake being attributed to macropinocytosis.35–37 Whilst we considered the possibility of studying this further using endocytic inhibitors, we ultimately decided against this due to concerns about lack of specificity and the potential effects on untargeted pathways.38,39
| Coefficient | BT474 | SKBR3 | ||
|---|---|---|---|---|
| Pearsons | Manders | Pearsons | Manders | |
| NPs/Dex | 0.367 | 0.252 | 0.258 | 0.396 |
| Dex488/Dex647 | 0.636 | 0.732 | 0.628 | 0.571 |
| NPs/Dex % | 57.724 | 34.449 | 41.128 | 69.292 |
PLGA-Tz induced plasma membrane reorganisation
Based on our previous observations on the ability of Tz to cluster HER2 and induce membrane ruffling we investigated events at this location immediately following incubation of cells with the NPs labelled with and without Tz.9 Having established the capacity of our NPs to selectively bind HER2 we decided to control the degree of Tz488 binding to our particles and measure this. Particle valency was determined by comparing the fluorescence of a known concentration of labelled NPs against a standard curve generated with Tz488. When generating NPs using a Tz![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NP ratio of 10
NP ratio of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 or 30
1 or 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, valencies were calculated to be 8.8 and 21.1 respectively. However, the calculated valency from this, or other similar methods does not consider the orientation of the antibodies and their accessibility to the antigen. We could not reliably determine the valency of the original PLGA-Tz NPs used in our study, created with a much higher Tz
1, valencies were calculated to be 8.8 and 21.1 respectively. However, the calculated valency from this, or other similar methods does not consider the orientation of the antibodies and their accessibility to the antigen. We could not reliably determine the valency of the original PLGA-Tz NPs used in our study, created with a much higher Tz![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NP ratio. This is most likely due to excess antibody causing stearic hindrance to binding, and further analyses were conducted using only the two conjugates termed PLGA-9Tz and PLGA-21Tz from the 10
NP ratio. This is most likely due to excess antibody causing stearic hindrance to binding, and further analyses were conducted using only the two conjugates termed PLGA-9Tz and PLGA-21Tz from the 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 and 30
1 and 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
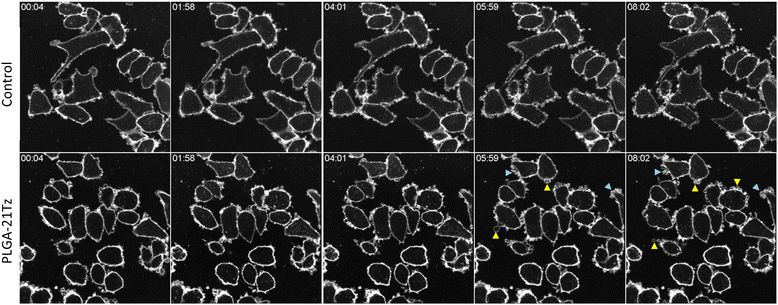
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratios respectively. For phenotypic characterisation we separate membrane reorganisation as blebbing and ruffling defined respectively as actin-free membrane protrusions or actin-rich fold-like membrane protrusions.40,41 To capture plasma membrane activity in real time over the first 10 min of conjugate addition to the cells we recorded the cells across a single focal plane (Fig. 4 and SI Fig. S5). Time lapse imaging analysis shows the dynamic fluidic nature of the plasma membrane in untreated cells (Fig. 4, SI Movies S1 and S2) but also demonstrate phenotypic variation within a field of view and thus the population. Prominent plasma membrane movements were visualised as regions having different levels of blebbing and ruffling. To indirectly quantify surface area changes we have utilised an imageJ plugin that measures the amount of “cell free space” available within a field of view. This showed an increase in total cell area over time within both BT474 cells and SKBR3 (SI Fig. S6). In the SKBR3 cells the untreated control show some variation however it does not substantially deviate from the baseline, in comparison the PLGA-21Tz treated cells show 10% reduction in cell-free area. As for the BT474 cells there is far less deviation in the untreated group representing their clumped nature and a smaller 5% reduction in cell-free area.
1 ratios respectively. For phenotypic characterisation we separate membrane reorganisation as blebbing and ruffling defined respectively as actin-free membrane protrusions or actin-rich fold-like membrane protrusions.40,41 To capture plasma membrane activity in real time over the first 10 min of conjugate addition to the cells we recorded the cells across a single focal plane (Fig. 4 and SI Fig. S5). Time lapse imaging analysis shows the dynamic fluidic nature of the plasma membrane in untreated cells (Fig. 4, SI Movies S1 and S2) but also demonstrate phenotypic variation within a field of view and thus the population. Prominent plasma membrane movements were visualised as regions having different levels of blebbing and ruffling. To indirectly quantify surface area changes we have utilised an imageJ plugin that measures the amount of “cell free space” available within a field of view. This showed an increase in total cell area over time within both BT474 cells and SKBR3 (SI Fig. S6). In the SKBR3 cells the untreated control show some variation however it does not substantially deviate from the baseline, in comparison the PLGA-21Tz treated cells show 10% reduction in cell-free area. As for the BT474 cells there is far less deviation in the untreated group representing their clumped nature and a smaller 5% reduction in cell-free area.
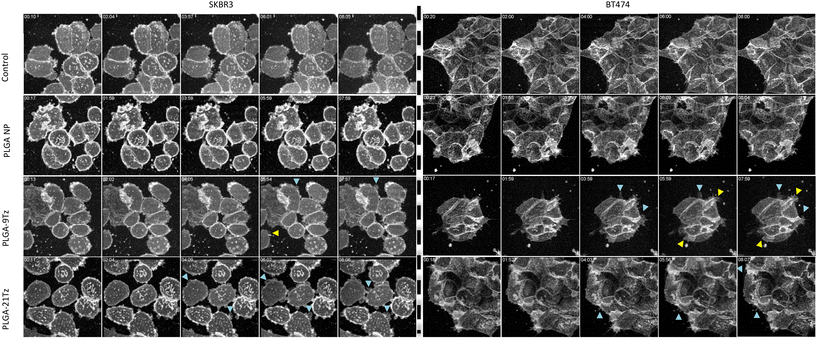
While this approach provided us with rapid imaging acquisition it became apparent that membrane reorganisation was occurring outside of our field of view. We therefore developed a resonant scanning confocal microscopy method to allow more rapid data acquisition that also provided the cell profile in 3D. Resonant scanners facilitate rapid acquisition of confocal 3D data stacks by driving a mirror at a high resonant frequency, creating a sinusoidal laser scan that enables video-rate frame acquisition. This minimizes phototoxicity and photobleaching, which is essential for gentle, high-speed volumetric imaging of living specimens. This allowed us to capture the full depth of the cells at a timescale suitable to observe changes in the whole plasma membrane and not of a single section. This data can then be observed and examined as a maximum intensity projection for each timepoint (Fig. 5) or as a volumetric dataset (Fig. 6).
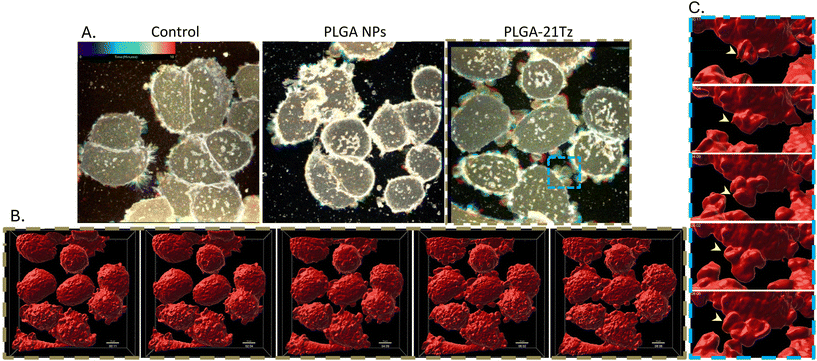
Using this method, we observe that cells incubated with undecorated NPs showed similar phenotypes to untreated cells (Fig. 5 and SI Movies S3–S10). However, the Tz-NPs caused, within 5 min, extensive membrane reorganisation manifesting as large membrane protrusions and ruffling that rapidly appear and disappear; this is most apparent in the cells treated with PLGA-21Tz. A greater extent of blebbing was noted in the PLGA-9Tz conjugates, with the reason behind this being hypothesised that it is due to the extent of HER2 binding influencing downstream effects on plasma membrane organisation. We hypothesise that this might be caused by an increasing degree of stearic hinderance as the valency increases. Work exploring HER2 affibody valencies on a HPMA polymer backbone have reported that HER2 binding influenced the internalisation of the conjugates with the possibility of “overcrowding” reducing the degree of receptor crosslinking.14 Our previous observations highlighted that HER2 crosslinking causes Erk phosphorylation that is linked to actin polymerisation and very likely to be involved in our observed effects with the Tz-NPs.9,42 By individually colour coding each frame of our maximum intensity projections we are able to provide a temporal map of cell surface events (Fig. 6A) with stationary area of the image represented as white/grey, and moving areas colour coded. This allows us to easily identify areas where membrane restructuring occurs, and the lifespan of the various protrusions.
Remodelling of the plasma membrane can lead to internalisation of extracellular fluid and local membranes through the formation of macropinosomes. In order to visualise the cell surface, volume data taken by the resonant scanner were processed using Bitplane's Imaris software to create a surface rendered visualisation of the membrane. Overall 3D topography of the plasma membrane across all cells can be followed over time (Fig. 6B and SI Movie S11), with individual protrusions forming cup-like structures that fold over and fuse with the plasma membrane (example given in Fig. 6C and SI Movie S12) when treated with PLGA-21Tz.
There is other evidence that HER2 crosslinking stimulates rapid plasma membrane rearrangement. Zanidatamab, an FDA approved antibody (Ziihera®) biparatopic antibody binding two different extracellular regions of HER2 simultaneously, has been shown to crosslink HER2.43,44 In SKBR3 breast cancer cells it formed HER2 “caps” or “microstructures” across the plasma membrane during the first 15 min of treatment that we believe to be the same structures/ruffling observed in Fig. 4–6.
This aspect of the work demonstrates the importance of obtaining a detailed analysis of membrane dynamics at early timepoints in live cells so that any rapid reorganisation of the plasma membrane caused by the formulation can be assessed. Resonant scanning confocal microscopy allowed for clear identification of 3-dimensional structures forming at the plasma membrane. For high spatial and temporal resolution studies, lattice light sheet time lapse microscopy shows great promise in providing higher resolution and a more complete picture of surface dynamics over short periods of time.45
SEM imaging of plasma membrane reorganization
The 2D and 3D live cell imaging data gave us valuable insights to rapid plasma membrane restructuring after NP-Tz addition. We next investigated whether this could be captured in SKBR3 cells at ultrastructural level using SEM to explore similarities in phenotype compared to previous work exploring the addition of soluble Tz to specifically induce HER2 crosslinking within 10 min of antibody addition.9Fig. 7 shows the extensive plasma membrane reorganisation observed in PLGA-21Tz treated cells mirrored our previous observations9 highlighting several relatively large ‘petal-like’ ruffles across the surface of the cell. Interestingly, we now show that these effects are reversible as 30 min later the cells return to baseline ruffling. It is well known that clustering receptors on the plasma membrane affects plasma membrane dynamics especially if these are, like HER2, signalling molecules with links to actin dynamics.46,47 This can result in membrane ruffling manifesting as macropinocytosis and/or membrane blebbing as a result of its detachment from the tethered cortical actin giving it freedom to blister in the way that we show, with increased hydrostatic pressure also contributing to the contracted cells.40,48 A separate study could investigate how our NPs could induce signalling pathways that cause ruffling and blebbing, focusing for example on roles of RAC1 and other GTPases, Arp and cofilin, and how signalling is switched off to regain homeostasis.41,49,50 Interestingly reversibility on the same timescale to shown here was observed with zanidatamab following its impact on membrane reorganisation.44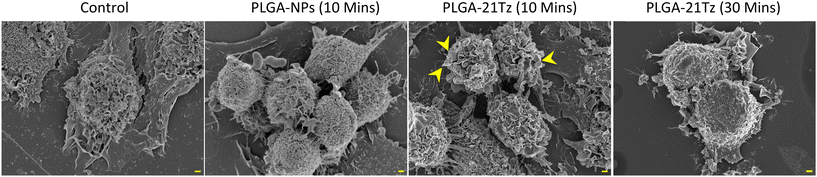 | ||
| Fig. 7 SEM analysis of untreated and NP incubated SKBR3 cells for 10 min and for PLGA-21Tz for 30 min. Yellow arrowheads highlight prominent ruffling. Scale bar = 1 µm. | ||
The observed effects on the plasma membrane are likely to involve actin and our previous observations highlighted that incubating cells with the actin disrupting agent cytochalasin D inhibited the stimulatory effects of Tz mediated HER2 crosslinking and endocytosis.9 A detailed examination of the role of actin in this process is now needed, but care must be taken due to the global effects when using actin inhibitors on cell morphology thus making data interpretation difficult.51
Cell viability
Numerous studies have investigated encapsulating anti-cancer agents within Tz-targeting nanoparticles (reviewed in Selepe, C. T. et al.).52 These include doxorubicin, utilised here to identify whether our particles showed any selective toxicity to HER2 expression and the impact of Tz valency. Unconjugated PLGA-Dox NPs showed toxicity against all cell lines, irrespective of HER2 status (Fig. 8). Conversely, Tz decorated NPs showed no significant toxicity against the MCF7 cells at equivalent dox concentrations. In the HER2++ cell lines the Tz decorated NPs showed very different profiles highlighting higher relative toxicity that was significantly different to untreated cells. Across both HER2++ cell lines increasing the valency resulted in reduced toxicity. For this trend it is possible that the additional antibodies were, by stearic hindrance, resulting in less efficient NP binding. Indeed, as we have no control over the orientation of Tz on the NP surface it is possible that this could also contribute to our findings and that it could further compound potential stearic hinderance encountered when increasing the valency. Though we demonstrate that our PLGA-Dox-Tz NPs are capable of selectively targeting HER2++ to deliver Dox and offering greater selectivity than free drug, the data highlights that increasing the concentration of the targeting ligand across the surface of a nanoparticle does not guarantee an improvement in drug delivery.Conclusions
Here we highlight how detailed real time analysis of nanoparticle–ligand–plasma membrane interactions reveals immediate effects on the cell surface dynamics. It is likely that simultaneously targeting two HER2 epitopes or using a combination of two antibodies causes similar effects. It is also likely that crosslinking results in cell signalling to induce blebbing/ruffling manifesting as macropinocytosis. As HER2 is well characterised as an endocytosis-resistant receptor, our studies along with others, highlight that this crosslinking approach could translate to more effective treatment options for HER2+ breast and other cancers.5,33,53,54 Our work also opens opportunities for gaining much needed knowledge on ligand binding of decorated nanoparticles targeting other receptors that are of therapeutic interest. However, further work will be required to study if these changes in membrane dynamics occur within an in vivo model.Author contributions
CSH: data acquisition, data processing, methodology, validation and writing. ST: methodology. DM: methodology and SEM imaging. AJH: data processing. PW: conceptualization, supervision, data processing and writing. ATJ: conceptualization, supervision, funding acquisition, data processing and writing.Conflicts of interest
There are no conflicts to declare.Data availability
Data is available upon reasonable request to the corresponding author.Supplementary information (SI) is available. Supplementary Information contains all video recordings and additional data. See DOI: https://doi.org/10.1039/d5pm00168d.
Acknowledgements
This work was funded by the Coleg Cymraeg Cenedlaethol as part of a PhD award granted to CSH and ATJ. The authors would like to thank Nourhan Adel Abdulla, Cardiff University for critically reading this manuscript. The authors gratefully acknowledge Cardiff University Bioimaging Hub Core Facility, RRID:SCR_022556, for their support and assistance in this work. Bitplane Imaris software and PC workstation was funded by a Cardiff University Research Infrastructure Fund award.References
- WHO Cancer Factsheet, https://www.who.int/news-room/fact-sheets/detail/cancer, (accessed 22 January 2025).
- P. R. Pohlmann, I. A. Mayer and R. Mernaugh, Clin. Cancer Res., 2009, 15, 7479–7491 Search PubMed.
- S. Ahmad, S. Gupta, R. Kumar, G. C. Varshney and G. P. S. Raghava, Sci. Rep., 2014, 4, 4483 Search PubMed.
- S. Verma, D. Miles, L. Gianni, I. E. Krop, M. Welslau, J. Baselga, M. Pegram, D.-Y. Oh, V. Diéras, E. Guardino, L. Fang, M. W. Lu, S. Olsen, K. Blackwell and EMILIA Study Group, N. Engl. J. Med., 2012, 367, 1783–1791 Search PubMed.
- F. Shen, Q. Lin, C. Childress and W. Yang, Cell. Signalling, 2008, 20, 779–786 CrossRef PubMed.
- A. M. Hommelgaard, M. Lerdrup and B. van Deurs, Mol. Biol. Cell, 2004, 15, 1557–1567 CrossRef PubMed.
- V. Bertelsen and E. Stang, Membranes, 2014, 4, 424–446 CrossRef PubMed.
- P. R. Moody, E. J. Sayers, J. P. Magnusson, C. Alexander, P. Borri, P. Watson and A. T. Jones, Mol. Ther., 2015, 23, 1888 CrossRef PubMed.
- J. M. Wymant, E. J. Sayers, D. Muir and A. T. Jones, J. Cancer, 2020, 11, 3288–3302 CrossRef.
- J. Y. Li, S. R. Perry, V. Muniz-Medina, X. Wang, L. K. Wetzel, M. C. Rebelatto, M. J. M. Hinrichs, B. Z. Bezabeh, R. L. Fleming, N. Dimasi, H. Feng, D. Toader, A. Q. Yuan, L. Xu, J. Lin, C. Gao, H. Wu, R. Dixit, J. K. Osbourn and S. R. Coats, Cancer Cell, 2016, 29, 117–129 CrossRef PubMed.
- R. Domínguez-Ríos, D. R. Sánchez-Ramírez, K. Ruiz-Saray, P. E. Oceguera-Basurto, M. Almada, J. Juárez, A. Zepeda-Moreno, A. del Toro-Arreola, A. Topete and A. Daneri-Navarro, Colloids Surf., B, 2019, 178, 199–207 CrossRef PubMed.
- T. Kubota, S. Kuroda, N. Kanaya, T. Morihiro, K. Aoyama, Y. Kakiuchi, S. Kikuchi, M. Nishizaki, S. Kagawa, H. Tazawa and T. Fujiwara, Nanomedicine, 2018, 14, 1919–1929 CrossRef PubMed.
- C. A. Juul, T. B. Engel, F. P. Fliedner, L. Ringgaard, R. Eliasen, F. Melander, M. Bak, A. Kjær, J. R. Henriksen, D. R. Elema, A. E. Hansen and T. L. Andresen, J. Controlled Release, 2024, 371, 288–297 CrossRef PubMed.
- D. C. Radford, J. Yang, M. C. Doan, L. Li, A. S. Dixon, S. C. Owen and J. Kopeček, J. Controlled Release, 2020, 319, 285–299 CrossRef PubMed.
- A. Tsherniak, F. Vazquez, P. G. Montgomery, B. A. Weir, G. Kryukov, G. S. Cowley, S. Gill, W. F. Harrington, S. Pantel, J. M. Krill-Burger, R. M. Meyers, L. Ali, A. Goodale, Y. Lee, G. Jiang, J. Hsiao, W. F. J. Gerath, S. Howell, E. Merkel, M. Ghandi, L. A. Garraway, D. E. Root, T. R. Golub, J. S. Boehm and W. C. Hahn, Cell, 2017, 170, 564–576.e16 Search PubMed.
- E. J. Sayers, S. E. Peel, A. Schantz, R. M. England, M. Beano, S. M. Bates, A. S. Desai, S. Puri, M. B. Ashford and A. T. Jones, Mol. Ther., 2019, 27, 1950–1962 Search PubMed.
- W. H. Humphries, C. J. Szymanski and C. K. Payne, PLoS One, 2011, 6, e26626 CrossRef PubMed.
- K. W. Dunn, M. M. Kamocka and J. H. McDonald, Am. J. Physiol.: Cell Physiol., 2011, 300, C723–C742 CrossRef PubMed.
- N. Otsu, IEEE Trans. Syst. Man. Cybern., 1979, 9, 62–66 Search PubMed.
- A. Suarez-Arnedo, F. Torres Figueroa, C. Clavijo, P. Arbeláez, J. C. Cruz and C. Muñoz-Camargo, PLoS One, 2020, 15, e0232565 CrossRef PubMed.
- J. Schindelin, I. Arganda-Carreras, E. Frise, V. Kaynig, M. Longair, T. Pietzsch, S. Preibisch, C. Rueden, S. Saalfeld, B. Schmid, J.-Y. Tinevez, D. J. White, V. Hartenstein, K. Eliceiri, P. Tomancak and A. Cardona, Nat. Methods, 2012, 9, 676–682 CrossRef PubMed.
- H. O. Alsaab, F. D. Alharbi, A. S. Alhibs, N. B. Alanazi, B. Y. Alshehri, M. A. Saleh, F. S. Alshehri, M. A. Algarni, T. Almugaiteeb, M. N. Uddin and R. M. Alzhrani, Pharmaceutics, 2022, 14, 2728 CrossRef PubMed.
- E. K. Marecki, K. W. Oh, P. R. Knight and B. A. Davidson, Biomicrofluidics, 2024, 18, 051503 Search PubMed.
- E. N. Komedchikova, O. A. Kolesnikova, E. D. Tereshina, P. A. Kotelnikova, A. S. Sogomonyan, A. V. Stepanov, S. M. Deyev, M. P. Nikitin and V. O. Shipunova, Pharmaceutics, 2022, 15, 52 Search PubMed.
- V. O. Shipunova, E. N. Komedchikova, P. A. Kotelnikova, M. P. Nikitin and S. M. Deyev, Pharmaceutics, 2023, 15, 833 CrossRef PubMed.
- J. Park, P. M. Fong, J. Lu, K. S. Russell, C. J. Booth, W. M. Saltzman and T. M. Fahmy, Nanomedicine, 2009, 5, 410–418 CrossRef PubMed.
- J. Sun, L. Zhang, J. Wang, Q. Feng, D. Liu, Q. Yin, D. Xu, Y. Wei, B. Ding, X. Shi and X. Jiang, Adv. Mater., 2015, 27, 1402–1407 Search PubMed.
- S. Varma, S. Dey and D. Palanisamy, Curr. Pharm. Biotechnol., 2022, 23, 679–706 Search PubMed.
- G. Griffiths, J. Gruenberg, M. Marsh, J. Wohlmann, A. T. Jones and R. G. Parton, Adv. Drug Delivery Rev., 2022, 188, 114403 CrossRef PubMed.
- N. Eivazi, R. Rahmani and M. Paknejad, Life Sci., 2020, 261, 118361 CrossRef PubMed.
- B. Colzani, L. Pandolfi, A. Hoti, P. A. Iovene, A. Natalello, S. Avvakumova, M. Colombo and D. Prosperi, Int. J. Nanomed., 2018, 13, 957–973 CrossRef.
- S. Mylvaganam, S. A. Freeman and S. Grinstein, Curr. Biol., 2021, 31, R619–R632 CrossRef.
- C. D. Austin, A. M. De Mazière, P. I. Pisacane, S. M. van Dijk, C. Eigenbrot, M. X. Sliwkowski, J. Klumperman and R. H. Scheller, Mol. Biol. Cell, 2004, 15, 5268–5282 CrossRef.
- Y. Yarden and M. X. Sliwkowski, Nat. Rev. Mol. Cell Biol., 2001, 2, 127–137 CrossRef.
- V. T. Cong, R. D. Tilley, G. Sharbeen, P. A. Phillips, K. Gaus and J. J. Gooding, Chem. Sci., 2021, 12, 15407–15417 RSC.
- A. Akinc and G. Battaglia, Cold Spring Harbor Perspect. Biol., 2013, 5, a016980 CrossRef.
- Y.-X. Li and H.-B. Pang, J. Controlled Release, 2021, 329, 1222–1230 CrossRef CAS.
- D. Vercauteren, R. E. Vandenbroucke, A. T. Jones, J. Rejman, J. Demeester, S. C. De Smedt, N. N. Sanders and K. Braeckmans, Mol. Ther., 2010, 18, 561–569 CrossRef CAS PubMed.
- M. Itagaki, Y. Nasu, C. Sugiyama, I. Nakase and N. Kamei, FASEB J., 2023, 37, e22764 CrossRef CAS PubMed.
- G. T. Charras, J. Microsc., 2008, 231, 466–478 CrossRef CAS PubMed.
- G. Yan, J. Zhou, J. Yin, D. Gao, X. Zhong, X. Deng, H. Kang and A. Sun, Int. J. Mol. Sci., 2024, 25, 10971 CrossRef CAS PubMed.
- N. Martinez-Quiles, H.-Y. H. Ho, M. W. Kirschner, N. Ramesh and R. S. Geha, Mol. Cell. Biol., 2004, 24, 5269–5280 CrossRef CAS PubMed.
- J. J. Harding, J. Fan, D.-Y. Oh, H. J. Choi, J. W. Kim, H.-M. Chang, L. Bao, H.-C. Sun, T. Macarulla, F. Xie, J.-P. Metges, J. Ying, J. Bridgewater, M.-A. Lee, M. A. Tejani, E. Y. Chen, D. U. Kim, H. Wasan, M. Ducreux, Y. Bao, L. Boyken, J. Ma, P. Garfin, S. Pant, G. Abou-Alfa, J. A. Alfonso, M. Aglietta, A. Baron, M. Beg, P. C. Aguirre, E. Chen, Y. Cheng, A. C. Gracián, L. Dahan, F. D. Braud, E. Fenocchio, O. Gbolohon, R. Gillmore, M. Jary, M. Javle, Y. Jiang, J. H. Kang, G. G. King, M. Kundranda, L. Layos, D. Li, T. Liang, S. Lonardi, O. Marathe, S. Mondaca, A. J. M. Martin, J. O. Park, R. P. Cid, P. R. Fernandez, L. Rimassa, R. R. Alonso, S. Sadeghi, A. Scott, B. Tan, D. Tougeron, Q. Yan, X. Yin and H. Zhao, Lancet Oncol., 2023, 24, 772–782 CrossRef CAS.
- N. E. Weisser, M. Sanches, E. Escobar-Cabrera, J. O'Toole, E. Whalen, P. W. Y. Chan, G. Wickman, L. Abraham, K. Choi, B. Harbourne, A. Samiotakis, A. H. Rojas, G. Volkers, J. Wong, C. E. Atkinson, J. Baardsnes, L. J. Worrall, D. Browman, E. E. Smith, P. Baichoo, C. W. Cheng, J. Guedia, S. Kang, A. Mukhopadhyay, L. Newhook, A. Ohrn, P. Raghunatha, M. Zago-Schmitt, J. D. Schrag, J. Smith, P. Zwierzchowski, J. M. Scurll, V. Fung, S. Black, N. C. J. Strynadka, M. R. Gold, L. G. Presta, G. Ng and S. Dixit, Nat. Commun., 2023, 14, 1394 CrossRef CAS.
- S. E. Quinn, L. Huang, J. G. Kerkvliet, J. A. Swanson, S. Smith, A. D. Hoppe, R. B. Anderson, N. W. Thiex and B. L. Scott, Nat. Commun., 2021, 12, 4838 CrossRef CAS.
- D. B. Peckys, U. Korf and N. de Jonge, Sci. Adv., 2015, 1, e1500165 CrossRef.
- R. Malla, A. Tumbali, P. Chode, K. Manda, A. S. Samudrala, Y. Nuthalapati, C. Mangam, P. Bhamidipati, M. Srilatha and G. P. Nagaraju, J. Natl. Cancer Cent., 2025, 5, 263–275 Search PubMed.
- W. Strychalski and R. D. Guy, Biophys. J., 2016, 110, 1168–1179 CrossRef CAS PubMed.
- F. Leyden, S. Uthishtran, U. K. Moorthi, H. M. York, A. Patil, H. Gandhi, E. P. Petrov, T. Bornschlögl and S. Arumugam, BMC Biol., 2021, 19, 72 CrossRef CAS.
- A. C. Bendell, E. K. Williamson, C. S. Chen, J. K. Burkhardt and D. A. Hammer, Integr. Biol. Quant. Biosci. Nano Macro, 2017, 9, 695–708 CAS.
- L. He, E. J. Sayers, P. Watson and A. T. Jones, Sci. Rep., 2018, 8, 7318 CrossRef CAS PubMed.
- C. T. Selepe, K. S. Dhlamini, L. Tshweu, M. Moralo, L. Kwezi, S. S. Ray and B. Ramalapa, Nano Sel., 2024, 5, 2300191 CrossRef CAS.
- K. Cortese, M. T. Howes, R. Lundmark, E. Tagliatti, P. Bagnato, A. Petrelli, M. Bono, H. T. McMahon, R. G. Parton and C. Tacchetti, Mol. Biol. Cell, 2013, 24, 129–144 CrossRef CAS.
- J. Cheng, M. Liang, M. F. Carvalho, N. Tigue, R. Faggioni, L. K. Roskos and I. Vainshtein, Antibodies, 2020, 9, 49 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |