Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Backbone extension by methylene insertion as a strategy to restore duplex stability of sulfonamide-linked oligodeoxynucleotides while preserving RNase H activity
Kohji
Seio
 *,
Hiroki
Mikagi
,
Haruka
Tanabe
,
Shigetoshi
Tachibana
,
Yoshiaki
Masaki
and
Eitaro
Murakami
*,
Hiroki
Mikagi
,
Haruka
Tanabe
,
Shigetoshi
Tachibana
,
Yoshiaki
Masaki
and
Eitaro
Murakami

Department of Life Science and Technology, Institute of Science Tokyo, 4259 Nagatsuta-cho, Midori-ku, Yokohama 226-8501, Japan. E-mail: seio.k.2552@m.isct.ac.jp
First published on 24th April 2026
Abstract
A backbone-extended sulfonamide nucleic acid was developed for use in gapmer antisense oligonucleotides. Backbone extension restores duplex stability while preserving the ability of the oligonucleotides to induce RNase H-mediated RNA cleavage. Structural analysis suggests that the extended linkage adopts unusual backbone conformations yet remains compatible with canonical A-type duplex formation, highlighting backbone extension as a new design parameter for antisense oligonucleotide therapeutics.
Antisense oligonucleotides (ASOs) are clinically useful synthetic nucleic acids that exert their biological activity through sequence-specific binding to target RNAs.1 Among these, gapmer-type ASOs consist of a central DNA (“gap”) region flanked by segments of 2′-O-modified nucleotides (“wings”) and induce RNase H-mediated cleavage of the RNA strand.2 Although phosphorothioate backbones3 are widely used to enhance the resistance of ASOs to nucleases, their inherent chirality3,4 and associated off-target interactions with various proteins5–7 have motivated the development of alternative internucleotidic linkages.8–10 Although unmodified phosphodiester linkages can be inserted at specific positions, such as the wing regions of gapmers, their application in the gap region is hampered by instability toward nucleases.11
A sulfonamide linkage (Fig. 1 left panel) represents an achiral backbone modification and has been investigated as a candidate internucleotidic linkage lacking a phosphate group because of its charge-neutral character and resistance to nuclease digestion.12,13 In this linkage, the 3′-oxygen, the phosphorus atom, and the 5′-oxygen of ![[t with combining low line]](https://www.rsc.org/images/entities/char_0074_0332.gif) hymidyl(3,5′)
hymidyl(3,5′)![[t with combining low line]](https://www.rsc.org/images/entities/char_0074_0332.gif) hymidine were replaced with
hymidine were replaced with ![[n with combining low line]](https://www.rsc.org/images/entities/char_006e_0332.gif) itrogen,
itrogen, ![[s with combining low line]](https://www.rsc.org/images/entities/char_0073_0332.gif) ulfur, and
ulfur, and ![[c with combining low line]](https://www.rsc.org/images/entities/char_0063_0332.gif) arbon, respectively; we therefore named this residue TnscT. We have previously reported the incorporation of TnscT into gapmer-type oligonucleotides and shown that the modified gapmers support RNase H-dependent RNA cleavage.14 However, TnscT exhibited reduced affinity for complementary RNA, probably because of the limited rotational flexibility of the S–N bond and steric hindrance of the 6′-CH2 group adjacent to the SO2 group (Fig. 1). Herein, we report a new sulfonamide nucleic acid (Fig. 1, TnsccT) in which backbone extension is achieved by inserting a 7′-CH2 group.
arbon, respectively; we therefore named this residue TnscT. We have previously reported the incorporation of TnscT into gapmer-type oligonucleotides and shown that the modified gapmers support RNase H-dependent RNA cleavage.14 However, TnscT exhibited reduced affinity for complementary RNA, probably because of the limited rotational flexibility of the S–N bond and steric hindrance of the 6′-CH2 group adjacent to the SO2 group (Fig. 1). Herein, we report a new sulfonamide nucleic acid (Fig. 1, TnsccT) in which backbone extension is achieved by inserting a 7′-CH2 group.
 | ||
| Fig. 1 Structures of the sulfonamide-modified thymidine dimer TnscT and the backbone-extended TnsccT. | ||
We became interested in backbone extension based on our previous studies15 showing that contrary to the general expectation that backbone extension destabilizes duplex structures, extension of the carbon chain in phosphodiester-linked oligodeoxynucleotides can allow the formation of stable canonical duplexes with complementary RNA. In some cases, duplex stability was slightly increased, suggesting that the extended backbone could be accommodated within the DNA–RNA duplex. Based on this, we hypothesized that introducing a similar backbone extension into sulfonamide-linked nucleic acids might restore duplex stability by alleviating the conformational restrictions imposed by the sulfonamide linkage. In this paper, we report a backbone-extended sulfonamide nucleic acid (TnsccT) that restores the duplex stability of sulfonamide-linked oligonucleotides while maintaining their ability to induce RNase H-mediated RNA cleavage.
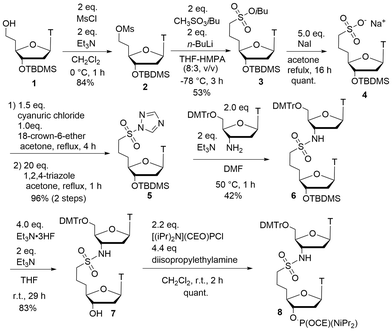
The phosphoramidite dimer of TnsccT (8) was synthesized as shown in Scheme 1. Starting from protected 5′-homothymidine 1,15,16 selective mesylation of the 5′-hydroxy group followed by substitution with isobutyl methanesulfonate afforded isobutyl sulfonate 3. Subsequent deprotection by treatment with sodium iodide generated the sodium salt of sulfonate 4, which was converted into the corresponding sulfonamide precursor by reacting with cyanuric chloride14,17 and 1,2,4-triazole to give intermediate 5. Coupling of 5 with a 3′-aminothymidine18 in the presence of triethylamine afforded the sulfonamide-linked dinucleoside 6. Removal of the tert-butyldimethylsilyl group, followed by phosphitylation of the free hydroxy group using standard chlorophosphoramidite chemistry, provided the target phosphoramidite 8 suitable for automated solid-phase oligonucleotide synthesis.
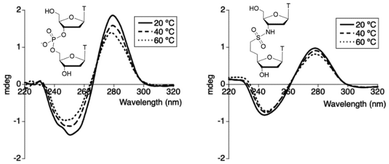
First, we measured the circular dichroism (CD) spectra of the TnsccT dimer obtained by deprotection of 7 (Fig. 2). Temperature-dependent CD spectra provide insight into the conformational behavior of the natural thymidyl(3′,5′)thymidine and TnsccT. As shown in the upper panel of Fig. 2, the CD spectra recorded at 20, 40, and 60 °C exhibited a characteristic negative Cotton effect at 250–260 nm and a positive band at 280–290 nm, consistent with π–π* transitions associated with base stacking. Upon increasing the temperature, a gradual attenuation of the CD intensity was observed, indicating a transition from stacked to unstacked conformations. In contrast, the CD spectra of TnsccT were largely independent of temperature, and the intensity was comparable to that of TpT at 60 °C. These results suggest that TnsccT does not adopt a stacked conformation and that the unit likely undergoes a conformational adjustment upon binding to a complementary strand.
The oligonucleotides synthesized in this study are listed in Table 1. ODN1 is an oligodeoxynucleotide with the sequence 5′-CGCACTTTTTTTTCGACC-3′. The UV-melting temperature (Tm) of the duplex with complementary RNA was 58.4 °C. When the 9th and 10th thymidine residues were replaced with a TnscT unit (ODN2), the Tm decreased by 3.3 °C to 55.1 °C. In contrast, replacement of the same residues with TnsccT (ODN3) resulted in a Tm of 57.7 °C, only 0.7 °C lower than that of ODN1. These results indicate that the destabilizing effect is significantly reduced in backbone-extended ODN3. This mitigating effect was more pronounced when multiple TnsccT units were incorporated. For example, in ODN4, in which two TnscT units replaced the 7th to 10th thymidine residues, the Tm decreased by 6.2 °C to 52.2 °C. In contrast, ODN7, containing TnsccT at the same positions, showed only a 1.0 °C decrease, giving a Tm of 57.4 °C. A similar trend was observed for ODN5 and ODN8, in which residues at position 9–12 were replaced. Moreover, in ODN6 and ODN9, where six thymidine residues at positions 7 to 12 were replaced with three TnscT or TnsccT units, respectively, each Tm was 49.3 °C and 55.2 °C, giving a difference of 5.9 °C. These results suggest that the extended backbone of TnsccT improves the accommodation of the sulfonamide linkage within the A-type duplex.
| Name | Sequence (5′ to 3′) | T m [°C] | ΔTm [°C] |
|---|---|---|---|
| Values in parentheses are standard deviations of three independent measurements. ΔTm is the difference between Tm and that of ODN1 or ASO1. For ASO1–3, the underline and lower case ‘c’ indicates LNA and 5-methylcytosine, respectively. The internucleotide linkages of ASOs are phosphorothioates. | |||
| ODN1 | CGCACTTTTTTTTCGACC | 58.4 (0.3) | — |
| ODN2 | CGCACTTTxxTTTCGACC | 55.1 (0.1) | −3.3 |
| ODN3 | CGCACTTTyyTTTCGACC | 57.7 (0.2) | −0.7 |
| ODN4 | CGCACT[xx][xx]TTTCGACC | 52.2 (0.2) | −6.2 |
| ODN5 | CGCACTTT[xx][xx]TCGACC | 52.7 (0.3) | −5.7 |
| ODN6 | CGCACT[xx][xx][xx]TCGACC | 49.3 (0.7) | −9.1 |
| ODN7 | CGCACT[yy][yy]TTTCGACC | 57.4 (0.4) | −1.0 |
| ODN8 | CGCACTTT[yy][yy]TCGACC | 57.6 (0.4) | −0.8 |
| ODN9 | CGCACT[yy][yy][yy]TCGACC | 55.2 (0.2) | −3.2 |
| ASO1 | ![[A with combining low line]](https://www.rsc.org/images/entities/char_0041_0332.gif) ![[c with combining low line]](https://www.rsc.org/images/entities/char_0063_0332.gif) ![[c with combining low line]](https://www.rsc.org/images/entities/char_0063_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) AGGcTTGcAT AGGcTTGcAT![[A with combining low line]](https://www.rsc.org/images/entities/char_0041_0332.gif) ![[c with combining low line]](https://www.rsc.org/images/entities/char_0063_0332.gif) |
70.1 (0.3) | — |
| ASO2 | ![[A with combining low line]](https://www.rsc.org/images/entities/char_0041_0332.gif) ![[c with combining low line]](https://www.rsc.org/images/entities/char_0063_0332.gif) ![[c with combining low line]](https://www.rsc.org/images/entities/char_0063_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) AGGcxxGcAT AGGcxxGcAT![[A with combining low line]](https://www.rsc.org/images/entities/char_0041_0332.gif) ![[c with combining low line]](https://www.rsc.org/images/entities/char_0063_0332.gif) |
68.0 (0.9) | −2.1 |
| ASO3 | ![[A with combining low line]](https://www.rsc.org/images/entities/char_0041_0332.gif) ![[c with combining low line]](https://www.rsc.org/images/entities/char_0063_0332.gif) ![[c with combining low line]](https://www.rsc.org/images/entities/char_0063_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) AGGcyyGCAT AGGcyyGCAT![[A with combining low line]](https://www.rsc.org/images/entities/char_0041_0332.gif) ![[c with combining low line]](https://www.rsc.org/images/entities/char_0063_0332.gif) |
69.3 (0.5) | −0.8 |
Next, we studied the properties of TnsccT in the gapmer antisense oligonucleotides. ASO1–3 contained a TT dimer at positions 9 and 10. The sequences were originally designed in our laboratory. As discussed in Fig. 4 and 5, RNase H recognizes the phosphate group corresponding to the sulfonamide linkage during cleavage of the complementary RNA; therefore, this sequence was chosen for the present study to see if the phosphate binding pocket of RNase H recognizes sulfonamide linkage. In this sequence, cytosine residues were replaced with 5-methylcytosine in accordance with the general design principles of antisense oligonucleotides, where such modifications are commonly employed to reduce activation of the innate immune system. The reference sequence, ASO1, incorporating natural TT, exhibited Tm of 70.1 °C. Introduction of the TnscT modification (ASO2) resulted in a noticeable decrease in duplex stability, with Tm of 68.0 °C, corresponding to a decrease of 2.1 °C relative to the unmodified control. In contrast, the incorporation of the TnsccT modification (ASO3) led to only minor changes in Tm. ASO3 showed a Tm of 69.3 °C, representing a modest decrease of 0.8 °C compared with ASO1. These results indicate that TnsccT modification largely preserves duplex stability, whereas TnscT modification induces a more pronounced destabilizing effect.
To understand the structural basis of the improved duplex stability of TnsccT-containing oligonucleotides despite the extended backbone, we modeled the possible conformations of the TnsccT unit. The CD spectrum of the ODN3/RNA duplex was measured and compared with those of ODN1/RNA and ODN2/RNA (Fig. S4). Although the Cotton effects at ∼210 nm and ∼260 nm are reduced in magnitude, the spectrum of ODN3/RNA still exhibits a characteristic positive band at ∼260 nm together with a negative band at ∼210 nm, which are typical features of A-form duplexes. These results indicate that the ODN3/RNA duplex retains an overall A-type helical conformation, and that the decreased intensities reflect subtle conformational modulation probably due to the below mentioned backbone conformation rather than a structural transition away from the A-form. As shown in Fig. 3, the C3′–C4′–C5′–C6′ (γ) dihedral angle can adopt either trans (left panel) or the gauche- (right panel) conformations. These conformations were not found in the natural A-type and B-type nucleic acid duplexes but were found in distorted duplexes such as protein–DNA complexes or mismatched duplexes.19 This suggested that the oligonucleotide containing a backbone-extended sulfonamide, such as TnsccT, formed a canonical A-type duplex while adopting an unusual backbone conformation.
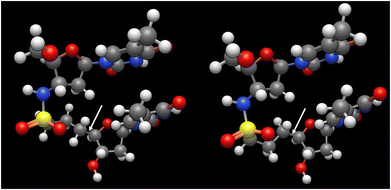 | ||
| Fig. 3 The TnsccT unit adopted to the A-type duplex structure. γ = trans (left panel), and gauche- (right panel). The C4′–C5′ bond is indicated by arrows. | ||
To examine how the introduction of the sulfonamide linkage affected RNase H recognition, we analyzed the initial RNase H cleavage pattern before extensive secondary cleavage. ASO1–3 were hybridized with a perfectly matched 16-mer RNA labeled with 5′-6FAM, and the cleavage reactions in the presence of Escherichia coli RNase H were analyzed using polyacrylamide gel electrophoresis. As shown in Fig. 4, in the presence of ASO1, 5% of the 16-mer RNA remained, and a 7-mer RNA was generated as the major product. Additionally, 8-, 9-, 10-, and 12-mer RNA were identified as minor products. When TnscT was incorporated (ASO2), 10% of the 16-mer RNA remained, and a 7-mer RNA fragment was generated as the major product. Importantly, cleavage at position 7 remained the major pathway, indicating that the introduction of the sulfonamide linkage did not interfere with RNase H cleavage of the phosphate located opposite to the modification (Fig. 5). Among the minor products, the amounts of 9-mer and 10-mer fragments decreased. When TnsccT was incorporated (ASO3), 28% of the 16-mer RNA remained, and a 7-mer RNA fragment was generated as the major product. In the case of ASO3, not only the 9- and 10- but also the 8-mer products disappeared, whereas the 12-mer product increased. These results indicate that sulfonamide-modified gapmers, such as ASO2 and ASO3, retain the ability to induce RNase H-mediated RNA cleavage while altering the cleavage pattern.
 | ||
| Fig. 4 The PAGE analyses of the cleavage of the 5′-6FAM-labeled complementary 16-mer RNA in the presence of E. coli RNase H and ASOs. | ||
To understand the cleavage pattern shown in Fig. 4, we mapped the positions of the phosphates as shown in Fig. 5. Arabic numerals on the RNA indicate the phosphates whose cleavage generates RNA fragments of the indicated length. The Roman numerals on the ASOs represent the phosphates recognized by the phosphate-binding pocket of RNase H.19 For example, cleavage at position 7 requires the recognition of phosphate vii. The cleavage pattern of ASO1 suggests that RNase H primarily recognizes phosphate vii to generate the major product, whereas phosphates viii, ix, x, and xii are recognized to produce minor products. In the case of ASO2, where phosphate ix is replaced by a sulfonamide linkage, recognition at position ix and additionally at x was diminished. These results suggest that the sulfonamide linkage and adjacent upstream phosphate are not efficiently recognized by the phosphate-binding pocket of RNase H. In the case of ASO3, the recognition of phosphate viii, which leads to cleavage at position 8, is also diminished. Because cleavage at position 8 requires the recognition of phosphate viii across the modification at position ix, the unusual backbone conformation shown in Fig. 3 may alter the static or dynamic structure of the duplex and weaken the RNase H-substrate complex responsible for cleavage at position 8.
Conclusions
In conclusion, we demonstrated that backbone extension restores the duplex stability of sulfonamide-linked oligonucleotides while preserving their ability to induce RNase H-mediated RNA cleavage. Analysis of RNase H cleavage patterns revealed that the sulfonamide linkage alters the recognition of the backbone by the enzyme, while the major cleavage pathway remains preserved. This indicates that sulfonamide linkages can be incorporated without abolishing RNase H activity when placed at positions that avoid critical phosphate recognition sites. Additionally, because sulfonamide linkages are resistant to nuclease digestion, backbone-extended sulfonamides may serve as nuclease-resistant backbone modifications that do not rely on phosphorothioate modifications and can therefore be incorporated into the gap region.Molecular modeling suggests the possibility that oligonucleotides containing TnsccT form canonical A-type duplexes with complementary RNA despite adopting unusual backbone conformations, reflecting the increased conformational flexibility of the extended backbone. Such structural adaptability may influence protein–nucleic acid interactions, suggesting that backbone extension could serve as a new design parameter for modulating undesired protein–nucleic acid interactions in antisense oligonucleotide therapeutics. Recent studies have highlighted the importance of replacing phosphorothioate linkages9,20 or controlling nucleic acid–protein interactions through chemical modification of the gap region.21,22 In this context, the incorporation of backbone-extended sulfonamide linkages may provide a useful strategy for developing antisense oligonucleotides with reduced phosphorothioate content and potentially improved safety profiles.
The synthesis of additional backbone-extended sulfonamide dimers beyond TnsccT, in vitro and in vivo applications of the ASOs containing nscc backbone are currently underway to expand the general applicability of this modification and will be reported elsewhere.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article are included in the supplementary information (SI). Supplementary information: NMR spectra and further experimental details. See DOI: https://doi.org/10.1039/d6ob00490c.Acknowledgements
K. S thanks the Japan Agency for Medical Research and Development (AMED) under grant number JP24fk0210147. This study was supported by the Japan Science and Technology Agency (grant number GTIE2024_EX07). The authors thank the Materials Analysis Division, Core Facility Center, Research Infrastructure Management Center, Institute of Science, Tokyo, for their support with the MALDI-TOF MS analyses.References
- S. T. Crooke, J. L. Witztum, C. F. Bennett and B. F. Baker, Cell Metab., 2018, 27, 714–739 CrossRef CAS PubMed.
- B. P. Monia, E. A. Lesnik, C. Gonzalez, W. F. Lima, D. McGee, C. J. Guinosso, A. M. Kawasaki, P. D. Cook and S. M. Freier, J. Biol. Chem., 1993, 268, 14514–14522 CrossRef CAS PubMed.
- F. Eckstein, Nucleic Acid Ther., 2014, 24, 374–387 CrossRef CAS PubMed.
- T. Poredoš, M. Trampuž, T. Gornik, K. Naveršnik, M. Sinreih Tisnikar, S. Pirc and Z. Časar, Org. Process Res. Dev., 2024, 28, 4194–4214 CrossRef.
- S. T. Crooke, T. A. Vickers and X. H. Liang, Nucleic Acids Res., 2020, 48, 5235–5253 CrossRef PubMed.
- S. T. Crooke, S. A. Baker, T. Pham, A. Hughes, H. S. Kwoh, A. Vickers, X. H. Liang, X. H. Sun and E. E. Swayze, Nucleic Acids Res., 2021, 49, 733–755 Search PubMed.
- M. Hyjek-Składanowska, B. A. Anderson, V. Mykhaylyk, C. Orr, A. Wagner, J. T. Poznański, K. Skowronek, P. Seth and M. Nowotny, Nucleic Acids Res., 2023, 51, 1409–1423 CrossRef PubMed.
- Y. Takahashi, I. Kato, K. Sato and T. Wada, J. Org. Chem., 2025, 90, 8966–8985 CrossRef CAS PubMed.
- M. T. Migawa, W. Shen, W. B. Wan, G. Vasquez, M. E. Øestergaard, A. Low, C. L. De Hoyos, R. Gupta, S. Murray, M. Tanowitz, M. Bell, J. G. Nichols, H. Gaus, X. H. Liang, E. E. Swayze, S. T. Crooke and P. P. Seth, Nucleic Acids Res., 2019, 47, 5465–5479 CrossRef CAS PubMed.
- J. Lovrić, J. Yan, X. Li, T. Karlsborn, M. Bood, A. Dahlén, C. Hilgendorf and D. K. Bhatt, Pharmacol. Res. Perspect., 2025, 13, e70096 CrossRef PubMed.
- E. B. McElroy, R. Bandaru, J. Huang and T. S. Widlanski, Bioorg. Med. Chem. Lett., 1994, 4, 1071–1076 CrossRef CAS.
- V. Korotkovs, L. F. Reichenbach, C. Pescheteau, G. A. Burley and R. M. J. Liskamp, J. Org. Chem., 2019, 84, 10635–10648 CrossRef CAS PubMed.
- K. Seio, R. Ohnishi, S. Tachibana, H. Mikagi and Y. Masaki, Org. Biomol. Chem., 2025, 23, 400–409 RSC.
- Y. Masaki, A. Tabira, S. Hattori, S. Wakatsuki and K. Seio, Org. Biomol. Chem., 2022, 20, 8917–8924 RSC.
- T. Kofoed, P. B. Rasmussen, P. Valentin-Hansen, E. B. Pedersen, K. Rissanen, W. Shi, S. Styring, C. Tommos, K. Warncke and B. R. Wood, Acta Chem. Scand., 1997, 51, 318–324 CrossRef CAS.
- T. Tomori, K. Uekusa, A. Koyama, T. Kanagawa, Y. Masaki and K. Seio, Bioorg. Med. Chem., 2022, 73, 117002 CrossRef CAS PubMed.
- R. Eisenhuth and C. Richert, J. Org. Chem., 2009, 74, 26–37 CrossRef CAS PubMed.
- D. Svozil, J. Kalina, M. Omelka and B. Schneider, Nucleic Acids Res., 2008, 36, 3690–3706 CrossRef CAS PubMed.
- M. Nowotny, S. A. Gaidamakov, R. Ghirlando, S. M. Cerritelli, R. J. Crouch and W. Yang, Mol. Cell, 2007, 28, 264–276 CrossRef CAS PubMed.
- B. A. Anderson, G. C. Freestone, A. Low, C. L. De-Hoyos, W. J. D. Iii, M. E. Østergaard, M. T. Migawa, M. Fazio, W. B. Wan, A. Berdeja, E. Scandalis, S. A. Burel, T. A. Vickers, S. T. Crooke, E. E. Swayze, X.-H. Liang and P. P. Seth, Nucleic Acids Res., 2021, 49, 9026–9041 CrossRef CAS PubMed.
- W. Shen, C. L. De Hoyos, M. T. Migawa, T. A. Vickers, H. Sun, A. Low, T. A. Bell III, M. Rahdar, S. Mukhopadhyay, C. E. Hart, M. Bell, S. Riney, S. F. Murray, S. Greenlee, R. M. Crooke, X.-H. Liang, P. P. Seth and S. T. Crooke, Nat. Biotechnol., 2019, 37, 640–650 CrossRef CAS PubMed.
- T. Kuroda, K. Yoshioka, S. S. Lei Mon, M. Katsuyama, K. Sato, E. Isogai, K. Yoshida-Tanaka, R. Iwata-Hara, T. Yamaguchi, S. Obika and T. Yokota, Mol. Ther.–Nucleic Acids, 2025, 36, 102692 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |