 Open Access Article
Open Access ArticleProtease-triggered self-immolative acyl phosphates for controlled phosphate release
Hunter J. Clark,
Yoojeong Chun and
Mark Nitz
and
Mark Nitz *
*
Department of Chemistry, University of Toronto, Toronto, Ontario, Canada. E-mail: mark.nitz@utoronto.ca
First published on 28th April 2026
Abstract
The intracellular delivery of phosphate esters remains a challenge, which often requires the use of a prodrug strategy. We report a protease-responsive scaffold that links an enzyme-catalyzed peptide cleavage to a spontaneous intramolecular lactamization onto a sterically encumbered acyl phosphate, resulting in phosphate release. This design represents a distinct mechanistic class of phosphate unmasking, differing fundamentally from phosphoramidate-based ProTides and other esterase-triggered systems. Using a phenylalanine based monoalkyl acyl phosphate as a model substrate, we demonstrate that the scaffold undergoes, chymotrypsin-dependent decomposition to liberate phosphate bearing substrates under mild conditions. The modular synthesis via a modified Staudinger ligation suggests that other protease-recognition sequences could be generated. This proof-of-concept establishes a general platform for enzyme-triggered phosphate release via acyl phosphates with potential applications in prodrug development and intracellular probe delivery.
Introduction
The intracellular delivery of phosphate-bearing small molecules remains a challenge. The negatively charged and highly polar nature of phosphate esters severely restricts passive diffusion across lipid membranes, while transporter-mediated uptake is often substrate-specific and tightly regulated.1–3 Consequently, phosphate-containing drugs and probes typically suffer from poor bioavailability and limited intracellular accumulation, motivating the development of diverse phosphate-masking strategies.1–4Among these strategies (Fig. 1), ProTide has been the most widely adopted. In this design, the phosphate is masked as a phosphoramidate, thereby enhancing lipophilicity and circumventing the need for transporter-mediated uptake.2,3 Once internalized, the prodrug is sequentially activated by esterases such as Cathepsin A or CES1 and the phosphoramidase HinT1, ultimately liberating the phosphorylated cargo.2,3,5–7 Although highly effective in mammalian systems for the delivery of nucleoside phosphates, this multi-enzyme activation pathway limits broader applicability, as homologous hydrolases, and their substrate specificities, differ substantially across species and cell types.8,9
 | ||
| Fig. 1 Established strategies for enzyme-triggered phosphate release (examples 1–3) and a proposed peptidase-activated cyclization mechanism (example 4). | ||
Several alternative phosphate-masking strategies (Fig. 1) have been explored, including non-enzymatic systems such as cycloSaligenyl (cycloSal)10,11 and esterase-dependent acyloxymethyl or S-acyl-2-thioethyl methods.12–15 These approaches highlight the chemical diversity available for phosphate protection but depend strongly on the intracellular environment for activation. The cycloSal design, for example, releases phosphate through a purely chemical mechanism without enzymatic assistance.10,11 Although conceptually general and applicable to diverse cargos, cycloSal prodrugs often require high concentrations to elicit biological activity and typically display lower potency than established analogues.11 This behaviour is attributed to limited passive diffusion and suboptimal stability, whereby either excessive persistence hinders intracellular release or premature degradation occurs extracellularly.11 By contrast, acyloxymethyl and S-acyl-2-thioethyl esters rely on intracellular esterases for unmasking,12–15 making their performance heavily dependent on host enzyme expression and thus less suitable in systems with low esterase activity. Collectively, these observations underscore the continuing need for robust and versatile phosphate-releasing strategies capable of functioning across diverse biological contexts.
Here, we present a protease-responsive phosphate-releasing scaffold that couple's enzyme-catalyzed peptide cleavage to a spontaneous intramolecular lactamization. Enzyme-triggered self-immolative release is well established in prodrug design, where esterase- or protease-mediated cleavage initiates decomposition of a para-aminobenzyl carbamate (PABC) spacer to liberate the active cargo.16–18 Phosphate-containing self-immolative linkers have also been described that release two cargos simultaneously; in those systems, however, phosphate functions only as a transient trigger rather than as the delivered species.19–21 Garbaccio and co-workers likewise demonstrated phosphate-bridged PABC linkers “CatPhos” that undergo Cathepsin B-mediated activation in antibody–drug conjugates releasing the desired cargo and a reactive aza-quinone methide.18 In contrast, our approach mediates phosphate release from an acyl phosphate intermediate via lactam formation, generating an inert lactam as the sole byproduct.
Acylphosphates are naturally occurring, high-energy intermediates that play central roles in metabolism and exhibit a distinctive reactivity profile arising from their dual capacity to function as both acylating and phosphorylating agents.22,23 Studies by Kluger, showed the moderate stability of acylphosphates to hydrolysis in aqueous systems facilitating their use in amide bond formation in protein conjugation and peptide synthesis.24–28 The useful reactivity profile of acyl phosphates has more recently been leveraged in self-assembling systems and as reactive intermediates.29–32 Additionally, work from Bearne, has built upon previous studies highlighting acylphosphate based inhibitors,33–35 and has highlighted their potential to act as highly specific probes for reactivity profiling.36
This study establishes a proof-of-concept of a stable acyl phosphate-based scaffold that releases phosphate through amide cleavage. Because proteases are ubiquitous across biological systems, this platform offers a foundation for future development of enzyme-triggered phosphate prodrugs and chemical biology probes. A monoalkylphosphate was employed here as a model leaving group to validate the mechanistic design.
Results and discussion
At the outset of this study, we sought to evaluate whether acyl phosphates could serve as viable structural elements within a prodrug scaffold. Acyl phosphates are well established as naturally occurring, high-energy intermediates in central metabolism, notable for their intrinsic electrophilicity and dual capacity to participate in either acyl transfer or phosphoryl transfer reactions.31,37–39 While these properties underpin their biological utility, they also raise concerns regarding chemical stability in aqueous and nucleophile-rich environments. Previous work on methyl acetyl phosphate has shown the hydrolysis half-life to be on the order of days near neutral pH but related aminoacyl phosphates hydrolyze in minutes.25,27,28 Accordingly, it was important to determine if the reactivity of the acyl phosphates could be reduced to provide a molecule sufficiently stable prior to a self immolative fragmentation. To this end, we synthesized a series of acyl phosphate derivatives featuring varied substitution at the α-position of the carbonyl to study how both steric (1c–e) and electronic effects (1f) contribute towards overall stability.Compounds 1c–f (Fig. 2) were selected for initial stability evaluation based on their ease of preparation in a single step from the corresponding acid anhydrides. The compounds were evaluated under pseudo-first-order conditions (pH 7.0, 1 M MOPS) using excess lysine (100 mM) as a representative nucleophile. The high concentration of lysine was selected to mimic the large excess of nucleophile that would be present in biological contexts.40 Acylphosphates undergo aminolysis more rapidly than hydrolysis, and the experimental design was intended to preferentially interrogate aminolysis reactivity.23,28,41 Accordingly, these conditions enabled us to probe the limits of aminolysis reactivity relevant to future prodrug applications. Rates of aminolysis were determined by 31P NMR spectroscopy, in which formation of the aminolysis product (δ ≈ −0.6 ppm) resulted in a pronounced downfield shift relative to the parent acylphosphate signal (δ ≈ −12 ppm), allowing reaction progress to be monitored over time. As shown in Fig. 2, increasing the degree of methyl substitution at the α-carbon resulted in a marked increase in acylphosphate stability. The measured half-lives for compounds 1c–f were approximately 5 min, 27 min, 636 min, and 66 min respectively. Compounds 1d and 1f exhibited similar half-lives, with the enhanced stability of 1f possibly arising from electronic effects associated with the aryl substituent, which may reduce the electrophilicity of the carbonyl center. Compound 1f can be compared to benzoyl methyl phosphate for which the reported aminolysis rates were comparable within a factor of two.28 Comparing aminolysis rates of the ethyl 1d and pivaloyl 1f derivatives, the two order of magnitude reduction in rate observed is consistent with the differences observed for aminolysis of p-nitrophenyl esters with the same acyl substituents.42 Collectively, these results indicate that degradation under biologically relevant conditions proceeds predominantly via C–O bond cleavage through nucleophilic attack at the carbonyl center and that this pathway can be substantially attenuated through steric modification at the α-position.
Guided by the results of the aminolysis studies, the target model compound contains a phosphate ester cargo, a self-immolative linker, and a protease recognition sequence. Phenyl phosphate (1a) and methyl phosphate (2a) were selected as the model phosphate ester cargos in our system, as they are synthetically accessible and representative of biologically relevant organophosphates.43–49 The linker design was guided by the need for it to remain chemically stable prior to enzymatic activation while reacting readily following proteolytic cleavage. Introduction of an α,α-dimethyl substitution at the carbonyl-phosphate stabilizes the unactivated prodrug, while simultaneously accelerating intramolecular cyclization after amide hydrolysis, consistent with the Thorpe–Ingold effect. It is well established that both 5 and 6-membered lactams form spontaneously via intramolecular cyclization under physiological conditions with appropriate esters.50,51 We opted for a six-atom linker over a five-atom analogue due to synthetic accessibility. Phenylalanine was incorporated into our model as the protease recognition sequence to serve as a chymotrypsin (CT) substrate; this enzyme is extensively characterized, readily available, and displays a strong preference for cleavage at the carboxy terminus of aromatic residues. Moreover, chymotrypsin is known to accommodate a broad range of substrates, further supporting its practicality in initial evaluations.52–54
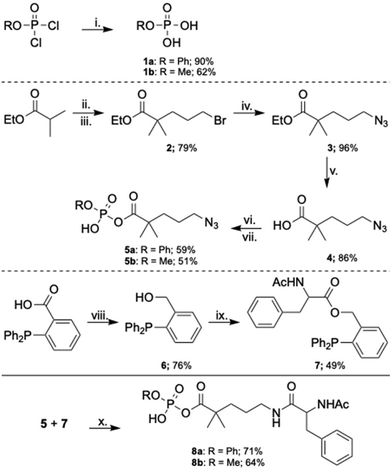
A modular synthesis was envisioned, where an azido acid could be first coupled to the desired phosphate ester cargo. The azide would then serve as the protected amine for subsequent Staudinger ligation with the preferred peptide (Scheme 1). 1a and 1b were readily prepared by basic hydrolysis of their respective commercially available dichlorophosphates. Synthesis of the self-immolative linker began by alkylation of ethyl isobutyrate with 1,3-dibromopropane to afford 2.55 Subsequent nucleophilic displacement of the primary bromide with sodium azide, followed by saponification of the ester, furnished 4 in 86% yield. Intermediate 4 was converted into the corresponding acyl chloride and subsequently coupled with the desired alkyl phosphate to give 5a and 5b in an analogous fashion to methods reported by Joeseph et al.56 Although acylphosphates can be prepared efficiently via coupling with anhydrides, typically affording higher yields and cleaner product profiles,39 this approach proved suboptimal due to its reduced atom economy and a low yield likely arising from the steric hindrance imposed by the α,α-dimethyl substitution. Formation of the required phenylalanine amide bond is complicated by the propensity a free amine to cyclize to the lactam in the presence of the electrophilic acyl phosphate. To circumvent this issue, a traceless Staudinger ligation strategy was adopted, enabling direct amide bond formation from the stable azide precursor.57 N-Acetyl-L-phenylalanine was first coupled to 6, prepared by reduction of commercially available 3-(diphenylphosphanyl)benzoic acid.57 This phosphine scaffold was selected over alternatives such as (diphenylphosphaneyl)methan(thi)ol due to its improved stability under ambient conditions,58 and was further favored over 3-(diphenylphosphanyl)phenol based on its increased reaction efficiency.57 Upon treatment of phosphine amino acid 7 with acylphosphate 5a or 5b under mild conditions,57 the traceless Staudinger ligation proceeded cleanly to afford the final compounds 8a and 8b in 71% and 64% respectively.
The performance of the self-immolative phosphate-releasing scaffold was evaluated by monitoring the hydrolysis by chymotrypsin (CT) (37 °C, 100 mM HEPES pH 8, 10 mM CaCl2). The addition of Ca2+ to the reaction mixture was necessary to ensure both the optimal stability and activity of CT.59–61 Divalent metal ions are reported to accelerate acyl phosphate hydrolysis but require high millimolar concentrations of Ca2+ are required for significant effects.62 Reaction progress was assessed by both 1H and 31P NMR spectroscopy (Fig. 3a and b). Successful amide bond cleavage and intramolecular cyclization of 8a was expected to liberate phenylphosphate, which appears as a characteristic downfield absorbance relative to the acylphosphate starting material in the 31P NMR spectrum. Under our assay conditions however, the phenylphosphate signal was attenuated by its precipitation with the Ca2+ ions associated with the chymotrypsin. To confirm the release of phenylphosphate the reaction mixture was analyzed HPLC and a signal consistent with phenylphosphate release was observed (Fig. S1). The 1H NMR spectrum provided a complementary measurement of reaction progress as the signal corresponding to the diastereotopic gem-dimethyl protons of the substrate collapse to a singlet, reflecting loss of the adjacent stereocenter.
The NMR time-course (Fig. 3a and b) revealed a progressive decrease in the intensity of the starting gem-dimethyl doublet (1.42 ppm), accompanied by the emergence and growth of a new downfield singlet (1.47 ppm), consistent with conversion to the cyclized product 10. The lactam 10 (3,3-dimethylpiperidin-2-one) was confirmed by the addition of authentic product to the crude reaction mixture which led to an increased signal intensity of the newly formed singlet (Fig. 3c), validating our proposed mechanism for the protease-triggered phosphate release. To benchmark the enzymatic reaction, we compared our experimental initial rate (vo) with rates for the established chromogenic substrates N-succinyl-L-phenylalanine-p-nitroanilide (Suc-Phe-pNA).63 This analysis indicated that chymotrypsin activity on 8a was noticeably slower, proceeding at approximately 10% the rate of the benchmark substrate, consistent with Phe-pNA being an activated substrate. Interestingly, complete conversion of 8a into its corresponding products was not observed. By extending the reaction times and adding additional enzyme it was only possible to achieve 50% conversion (Fig. S2). Interested by these results, we next evaluated if 8a was also an inhibitor of chymotrypsin. The activity of Chymotrypsin (10 μM) was monitored with the chromogenic substrate Suc-Phe-pNA (2 mM, 50 mM HEPES pH 8, 5 mM CaCl2) with increasing concentrations of 8a (0.75–3.0 mM). No significant reduction in reaction velocity was observed in the presence of the inhibitor (Fig. S3). This suggest that the limited turnover does not arise from loss of enzymatic activity due to inhibition, but rather from heterogeneity of the substrate. These findings are consistent with racemization of the N-acetyl phenylalanine during formation of the ester 7. To verify this, a sample of 7 was hydrolyzed and the liberated amino acid analyzed for its optical rotation relative to authentic N-acetyl-L-phenylalanine. The hydrolyzed material exhibited no optical activity. Although acylated amino acids are known to racemize at accelerated rates during coupling when compared to amino acid carbamates, racemization was not previously reported for the preparation of these structures and further work is required to optimize the synthesis of these esters.57,64
To explore whether the nature of the phosphate substituent influences scaffold stability or performance, we prepared an alternative derivative in which the phenyl group was replaced with a smaller methyl substituent. Compound 8b was synthesized analogously to 8a, differing only in the identity of the phosphate substituent. The performance of 8b was evaluated under identical conditions (37 °C, pH 8 HEPES buffer) using 1H NMR spectroscopy. In addition to the expected signals described above, phosphate release could be observed by following the shift in signal corresponding to the methyl protons of the alkyl phosphate (3.95 to 3.77 ppm for starting material to product respectively). However, time-course NMR analysis (Fig. 4) revealed no evidence of lactam formation, expected at 1.47 ppm. Instead, a doublet at 1.34 ppm was observed, consistent with the phenylalanyl amide remaining intact indicating slow acyl phosphate bond cleavage in leu of amide hydrolysis. Concomitantly, a doublet at 3.77 ppm corresponding to methyl phosphate increased over time. The parallel rates of appearance of the amino-acid-linker and appearance of the methyl phosphate signal suggested that 8b predominantly undergoes slow non-enzymatic hydrolysis (Fig. 4 and Fig. S4).
The contrasting outcomes between 8a and 8b underscore the influence of the phosphate substituent on enzymatic activation. Given that acyl phosphates are known inhibitors of related β-lactamases,33 we next sought to confirm that the divergent outcomes observed were not attributed to inhibition of chymotrypsin. Control activity assays were performed as described above. Compound 8b exhibited no evidence of enzyme inhibition (Fig. S3) at the highest concentration evaluated (3 mM).
Having established that 8b shows no detected inhibition, the divergent reactivity of 8a and 8b can be rationalized in terms of substrate recognition. Chymotrypsin's catalytic efficiency is not dictated solely by the P1 residue, but rather by contributions from residues spanning the P4–P3′ positions that occupy complementary subsites within the enzyme.53,65,66 In the Schellenberger model, substrate affinity and turnover are enhanced when each fragment is matched to a site with corresponding preference.53,65,66 Both the S1 and S2′ pockets exhibit a substantial preference for hydrophobic substituents.65,66 Within this framework, our scaffold positions the phenylalanine residue in S1, the immolative linker and the phosphate substituents in P′ positions. Placement of a phenyl group in a position for hydrophobic contacts may improve recognition and cleavage of this substrate, in contrast the predominantly polar methyl phosphate would be poorly recognized by the P′ subsites leading to poor cleavage.
Although chymotrypsin was used as a model protease in this study, the modular nature of the scaffold enables substitution of the peptide recognition sequence to target alternative proteolytic environments. For instance, in plant systems, papain-like cysteine proteases represent attractive candidates, while in microbial or in vitro contexts, broadly active enzymes such as proteinase K or subtilisin-like proteases may be employed.67–71 More generally, tailoring the terminal peptide sequence to the specificity of a target protease should enable this strategy to be adapted across diverse biological settings for controlled phosphate release.
Conclusion
In this work, we described the design and evaluation of a modular, protease-activated scaffold for controlled phosphate release. Deconstruction of the scaffold into three functionally independent components enabled a flexible architecture that supports modifications while limiting synthetic complexity. The incorporation of an α,α-dimethyl-substituted acyl phosphate linker was central to the design, providing both enhanced chemical stability prior to enzymatic activation and promoting rapid intramolecular cyclization following proteolytic cleavage. Using chymotrypsin as a model system, we showed that the scaffold 8a undergoes efficient enzymatic cleavage and lactam formation, resulting in phosphate release without detectable accumulation of hydrolytic byproducts. In contrast, analogue 8b does not undergo productive enzymatic activation and instead undergoes slow, non-enzymatic hydrolysis, underscoring the importance of the phosphate substituent in determining protease recognition. Control experiments confirmed that these divergent outcomes are not attributable to enzyme inhibition, but rather reflect differences in substrate engagement with chymotrypsin. Collectively, these results establish a proof-of-concept for a peptidase-triggered, self-immolative strategy for phosphate release and define key structure–reactivity relationships governing scaffold performance. Ongoing efforts will focus on extending this platform to more complex phosphate cargos and peptide recognition sequences, with the goal of enabling selective, enzyme-guided delivery of phosphorylated metabolites in biologically relevant settings.Conflicts of interest
There are no conflicts to declare.Data availability
The NMR spectra of all compounds, supplemental figures and the synthetic protocols have been included in the supplementary information (SI). See DOI: https://doi.org/10.1039/d6ob00405a.Acknowledgements
The authors gratefully acknowledge Dr Darcy Burns of the Centre for Spectroscopic Investigation of Complex Organic Molecules and Polymers (CSICOMP) NMR Facility for assistance with NMR experimental programming, and Dr Matt Forbes of the Advanced Instrumentation for Molecular Structure (AIMS) Mass Spectrometry Facility for their support with sample preparation and mass spectrometric analysis. We gratefully acknowledge financial support from the Natural Sciences and Engineering Research Council of Canada.References
- A. J. Wiemer and D. F. Wiemer, Prodrugs of Phosphonates and Phosphates: Crossing the Membrane Barrier, in Phosphorus Chemistry I: Asymmetric Synthesis and Bioactive Compounds, ed. J.-L. Montchamp, Springer International Publishing, Cham, 2015, pp. 115–160. DOI:10.1007/128_2014_561.
- M. Serpi and F. Pertusati, An Overview of ProTide Technology and Its Implications to Drug Discovery, Expert Opin. Drug Discov., 2021, 16(10), 1149–1161, DOI:10.1080/17460441.2021.1922385.
- M. Slusarczyk, M. Serpi and F. Pertusati, Phosphoramidates and Phosphonamidates (ProTides) with Antiviral Activity, Antivir. Chem. Chemother., 2018, 26, 2040206618775243, DOI:10.1177/2040206618775243.
- A. Usvalampi, H. Li and A. D. Frey, Production of Glucose 6-Phosphate From a Cellulosic Feedstock in a One Pot Multi-Enzyme Synthesis, Front. Bioeng. Biotechnol., 2021, 9 DOI:10.3389/fbioe.2021.678038.
- J. Cheng, X. Zhou, T.-F. Chou, B. Ghosh, B. Liu and C. R. Wagner, Identification of the Amino Acid-AZT-Phosphoramidase by Affinity T7 Phage Display Selection, Bioorg. Med. Chem. Lett., 2009, 19(22), 6379–6381, DOI:10.1016/j.bmcl.2009.09.067.
- E. Murakami, T. Tolstykh, H. Bao, C. Niu, H. M. M. Steuer, D. Bao, W. Chang, C. Espiritu, S. Bansal, A. M. Lam, M. J. Otto, M. J. Sofia and P. A. Furman, Mechanism of Activation of PSI-7851 and Its Diastereoisomer PSI-7977, J. Biol. Chem., 2010, 285(45), 34337–34347, DOI:10.1074/jbc.M110.161802.
- T. Yadati, T. Houben, A. Bitorina and R. Shiri-Sverdlov, The Ins and Outs of Cathepsins: Physiological Function and Role in Disease Management, Cells, 2020, 9(7), 1679, DOI:10.3390/cells9071679.
- S. Bardaweel, B. Ghosh, T.-F. Chou, M. J. Sadowsky and C. R. E. Wagner, Coli Histidine Triad Nucleotide Binding Protein 1 (ecHinT) Is a Catalytic Regulator of D-Alanine Dehydrogenase (DadA) Activity In Vivo, PLoS One, 2011, 6(7), e20897, DOI:10.1371/journal.pone.0020897.
- T.-F. Chou, P. Bieganowski, K. Shilinski, J. Cheng, C. Brenner and C. R. Wagner, 31P NMR and Genetic Analysis Establish hinT as the Only Escherchia Coli Purine Nucleoside Phosphoramidase and as Essential for Growth under High Salt Conditions *, J. Biol. Chem., 2005, 280(15), 15356–15361, DOI:10.1074/jbc.M500434200.
- C. Meier and J. Balzarini, Application of the cycloSal-Prodrug Approach for Improving the Biological Potential of Phosphorylated Biomolecules, Antiviral Res., 2006, 71(2–3), 282–292, DOI:10.1016/j.antiviral.2006.04.011.
- M. Munier, D. Tritsch, D. Lièvremont, M. Rohmer and C. Grosdemange-Billiard, New Application of cycloSaligenyl Prodrugs Approach for the Delivery of Fosfoxacin Derivatives in Mycobacteria, Molecules, 2023, 28(23), 7713, DOI:10.3390/molecules28237713.
- K. Schlüter, R. D. Walter, B. Bergmann and T. Kurz, Arylmethyl Substituted Derivatives of Fosmidomycin: Synthesis and Antimalarial Activity, Eur. J. Med. Chem., 2006, 41(12), 1385–1397, DOI:10.1016/j.ejmech.2006.06.015.
- T. Kurz, K. Schlüter, U. Kaula, B. Bergmann, R. D. Walter and D. Geffken, Synthesis and Antimalarial Activity of Chain Substituted Pivaloyloxymethyl Ester Analogues of Fosmidomycin and FR900098, Bioorg. Med. Chem., 2006, 14(15), 5121–5135, DOI:10.1016/j.bmc.2006.04.018.
- J. Wiesner, R. Ortmann, H. Jomaa and M. Schlitzer, Double Ester Prodrugs of FR900098 Display Enhanced In-Vitro Antimalarial Activity, Arch. Pharm., 2007, 340(12), 667–669, DOI:10.1002/ardp.200700069.
- C. Courtens, F. Van Charante, T. Quennesson, M. Risseeuw, P. Cos, G. Caljon, T. Coenye and S. Van Calenbergh, Acyloxymethyl and Alkoxycarbonyloxymethyl Prodrugs of a Fosmidomycin Surrogate as Antimalarial and Antibacterial Agents, Eur. J. Med. Chem., 2023, 245, 114924, DOI:10.1016/j.ejmech.2022.114924.
- J. R. McCombs and S. C. Owen, Antibody Drug Conjugates: Design and Selection of Linker, Payload and Conjugation Chemistry, AAPS J., 2015, 17(2), 339–351, DOI:10.1208/s12248-014-9710-8.
- C. C. Marvin, A. D. Hobson, M. McPherson, T. A. Dunstan, T. R. Vargo, M. E. Hayes, M. M. Fettis, A. Bischoff, L. Wang, L. Wang, A. Hernandez, Y. Jia, J. Z. Oh and Y. Tian, Self-Immolative Carbamate Linkers for CD19-Budesonide Antibody–Drug Conjugates, Bioconjugate Chem., 2023, 34(10), 1835–1850, DOI:10.1021/acs.bioconjchem.3c00354.
- J. C. Kern, D. Dooney, R. Zhang, L. Liang, P. E. Brandish, M. Cheng, G. Feng, A. Beck, D. Bresson, J. Firdos, D. Gately, N. Knudsen, A. Manibusan, Y. Sun and R. M. Garbaccio, Novel Phosphate Modified Cathepsin B Linkers: Improving Aqueous Solubility and Enhancing Payload Scope of ADCs, Bioconjugate Chem., 2016, 27(9), 2081–2088, DOI:10.1021/acs.bioconjchem.6b00337.
- P. Šimon, M. Tichotová, M. García Gallardo, E. Procházková and O. Baszczyňski, Phosphate–Based Self–Immolative Linkers for Tuneable Double Cargo Release, Chem. – Eur. J., 2021, 27(50), 12763–12775, DOI:10.1002/chem.202101805.
- O. Baszczyňski, Phosphorus-based Self-immolative Linkers, Habilitation thesis, Charles University, 2025 Search PubMed.
- E. Procházková, P. Šimon, M. Straka, J. Filo, M. Májek, M. Cigáň and O. Baszczyňski, Phosphate Linkers with Traceable Cyclic Intermediates for Self-Immolation Detection and Monitoring, Chem. Commun., 2021, 57(2), 211–214, 10.1039/D0CC06928K.
- F. Ramirez, S. Glaser, P. Stern, P. D. Gillespie and I. Ugi, Five–Membered Cyclic Acyl Phosphates, a New Class of Extremely Reactive Phosphorylating Agents, Angew. Chem., Int. Ed. Engl., 1973, 12(1), 66–67, DOI:10.1002/anie.197300661.
- G. Di Sabato and W. P. Jencks, Mechanism and Catalysis of Reactions of Acyl Phosphates. II. Hydrolysis, J. Am. Chem. Soc., 1961, 83(21), 4400–4405, DOI:10.1021/ja01482a025.
- R. S. Dhiman, L. G. Opinska and R. Kluger, Biomimetic Peptide Bond Formation in Water with Aminoacyl Phosphate Esters, Org. Biomol. Chem., 2011, 9(16), 5645–5647, 10.1039/C1OB05660C.
- R. Kluger, R. W. Loo and V. Mazza, Biomimetically Activated Amino Acids. Catalysis in the Hydrolysis of Alanyl Ethyl Phosphate, J. Am. Chem. Soc., 1997, 119(50), 12089–12094, DOI:10.1021/ja972406q.
- R. Kluger, A. S. Grant, S. L. Bearne and M. R. Trachsel, Dicarboxylic Acid Bis(Methyl Phosphates): Anionic Biomimetic Crosslinking Reagents, J. Org. Chem., 1990, 55(9), 2864–2868, DOI:10.1021/jo00296a056.
- R. Kluger and W.-C. Tsui, Methyl Acetyl Phosphate. A Small Anionic Acetylating Agent, J. Org. Chem., 1980, 45(13), 2723–2724, DOI:10.1021/jo01301a039.
- J. Wodzinska and R. Kluger, pKa -Dependent Formation of Amides in Water from an Acyl Phosphate Monoester and Amines, J. Org. Chem., 2008, 73(12), 4753–4754, DOI:10.1021/jo800667b.
- K. Dai, M. D. Pol, L. Saile, A. Sharma, B. Liu, R. Thomann, J. L. Trefs, D. Qiu, S. Moser, S. Wiesler, B. N. Balzer, T. Hugel, H. J. Jessen and C. G. Pappas, Spontaneous and Selective Peptide Elongation in Water Driven by Aminoacyl Phosphate Esters and Phase Changes, J. Am. Chem. Soc., 2023, 145(48), 26086–26094, DOI:10.1021/jacs.3c07918.
- G. L. Thomas and R. J. Payne, Phosphate-Assisted Peptide Ligation, Chem. Commun., 2009,(28), 4260, 10.1039/b906492c.
- A. Whicher, E. Camprubi, S. Pinna, B. Herschy and N. Lane, Acetyl Phosphate as a Primordial Energy Currency at the Origin of Life, Orig. Life Evol. Biosph., 2018, 48(2), 159–179, DOI:10.1007/s11084-018-9555-8.
- A. Sharma, K. Dai, M. D. Pol, R. Thomann, Y. Thomann, S. K. Roy and C. G. Pappas, Selective Peptide Bond Formation via Side Chain Reactivity and Self-Assembly of Abiotic Phosphates, Nat. Commun., 2025, 16(1), 1306, DOI:10.1038/s41467-025-56432-6.
- S. A. Adediran, M. Nukaga, S. Baurin, J.-M. Frère and R. F. Pratt, Inhibition of Class D β-Lactamases by Acyl Phosphates and Phosphonates, Antimicrob. Agents Chemother., 2005, 49(10), 4410, DOI:10.1128/AAC.49.10.4410-4412.2005.
- M. J. Morrison, N. Li and R. F. Pratt, Inverse Acyl Phosph(on)Ates: Substrates or Inhibitors of β-Lactam-Recognizing Enzymes?, Bioorg. Chem., 2001, 29(5), 271–281, DOI:10.1006/bioo.2001.1218.
- R. Kluger and W.-C. Tsui, Reaction of the Anionic Acetylation Agent Methyl Acetyl Phosphate with D-3-Hydroxybutyrate Dehydrogenase, Biochem. Cell Biol., 1986, 64(5), 434–440, DOI:10.1139/o86-061.
- L. C. McGary, G. L. Regan and S. L. Bearne, Reactive Architecture Profiling with a Methyl Acyl Phosphate Electrophile, Biochim. Biophys. Acta, Proteins Proteomics, 2023, 1871(6), 140945, DOI:10.1016/j.bbapap.2023.140945.
- C. S. Francklyn and P. Mullen, Progress and Challenges in Aminoacyl-tRNA Synthetase-Based Therapeutics, J. Biol. Chem., 2019, 294(14), 5365–5385, DOI:10.1074/jbc.REV118.002956.
- T. L. Mayover, C. J. Halkides and R. C. Stewart, Kinetic Characterization of CheY Phosphorylation Reactions: Comparison of P-CheA and Small-Molecule Phosphodonors, Biochemistry, 1999, 38(8), 2259–2271, DOI:10.1021/bi981707p.
- A. Englert, F. Majer, J. L. Schiessl, A. J. C. Kuehne and M. Von Delius, Acylphosphates, as Versatile Transient Species in Reaction Networks and Optical Catalyst Screenings, Chem, 2024, 10(3), 910–923, DOI:10.1016/j.chempr.2023.11.015.
- G. Gauthier-Coles, J. Vennitti, Z. Zhang, W. C. Comb, S. Xing, K. Javed, A. Bröer and S. Bröer, Quantitative Modelling of Amino Acid Transport and Homeostasis in Mammalian Cells, Nat. Commun., 2021, 12(1), 5282, DOI:10.1038/s41467-021-25563-x.
- G. Di Sabato and W. P. Jencks, Mechanism and Catalysis of Reactions of Acyl Phosphates. I. Nucleophilic Reactions, J. Am. Chem. Soc., 1961, 83(21), 4393–4400, DOI:10.1021/ja01482a024.
- R. C. Knowlton and L. D. Byers, Acyl Substituent Effects on Ester Aminolysis, J. Org. Chem., 1988, 53(16), 3862–3865, DOI:10.1021/jo00251a041.
- A. Narmandakh, N. Gad'on, F. Drepper, B. Knapp, W. Haehnel and G. Fuchs, Phosphorylation of Phenol by Phenylphosphate Synthase: Role of Histidine Phosphate in Catalysis, J. Bacteriol., 2006, 188(22), 7815–7822, DOI:10.1128/jb.00785-06.
- A. Lack and G. Fuchs, Evidence That Phenol Phosphorylation to Phenylphosphate Is the First Step in Anaerobic Phenol Metabolism in a Denitrifying Pseudomonas Sp, Arch. Microbiol., 1994, 161(2), 132–139, DOI:10.1007/BF00276473.
- N. H. Williams, Models for Biological Phosphoryl Transfer, Biochim. Biophys. Acta, Proteins Proteomics, 2004, 1697(1), 279–287, DOI:10.1016/j.bbapap.2003.11.031.
- X. Xie and N. Müller, Enzymes Involved in the Anaerobic Degradation of Phenol by the Sulfate-Reducing Bacterium Desulfatiglans Anilini, BMC Microbiol., 2018, 18(1), 93, DOI:10.1186/s12866-018-1238-0.
- K. Schühle and G. Fuchs, Phenylphosphate Carboxylase: A New C-C Lyase Involved in Anaerobic Phenol Metabolism in Thauera Aromatica, J. Bacteriol., 2004, 186(14), 4556–4567, DOI:10.1128/jb.186.14.4556-4567.2004.
- E. A. Salter, A. Wierzbicki and R. E. Honkanen, Quantum-Based Modeling of Dephosphorylation in the Catalytic Site of Serine/Threonine Protein Phosphatase-5 (PPP5C), Catalysts, 2020, 10(6), 674, DOI:10.3390/catal10060674.
- J. G. Zalatan, T. D. Fenn and D. Herschlag, Comparative Enzymology in the Alkaline Phosphatase Superfamily to Determine the Catalytic Role of an Active Site Metal Ion, J. Mol. Biol., 2008, 384(5), 1174–1189, DOI:10.1016/j.jmb.2008.09.059.
- M. A. DeWit and E. R. Gillies, Design, Synthesis, and Cyclization of 4-Aminobutyric Acid Derivatives: Potential Candidates as Self-Immolative Spacers, Org. Biomol. Chem., 2011, 9(6), 1846–1854, 10.1039/C0OB00890G.
- Z. Qin, X. Zhang, X. Sang, W. Zhang, G. Qu and Z. Sun, Carboxylic Acid Reductases Enable Intramolecular Lactamization Reactions, Green Synth. Catal., 2022, 3(3), 294–297, DOI:10.1016/j.gresc.2022.05.009.
- W. Ma, C. Tang and L. Lai, Specificity of Trypsin and Chymotrypsin: Loop-Motion-Controlled Dynamic Correlation as a Determinant, Biophys. J., 2005, 89(2), 1183–1193, DOI:10.1529/biophysj.104.057158.
- G. J. C. Vreeke, J.-P. Vincken and P. A. Wierenga, The Path of Proteolysis by Bovine Chymotrypsin, Food Res. Int., 2023, 165, 112485, DOI:10.1016/j.foodres.2023.112485.
- L. Hedstrom, Serine Protease Mechanism and Specificity, Chem. Rev., 2002, 102(12), 4501–4524, DOI:10.1021/cr000033x.
- M. Kuwahara, Y. Kawano, M. Kajino, Y. Ashida and A. Miyake, Synthetic Studies on Condensed-Azole Derivatives. V. Synthesis and Anti-Asthmatic Activities of ω-Sulfamoylalkyloxy[1, 2, 4]Triazolo[1, 5-b]-Pyridazines, Chem. Pharm. Bull., 1997, 45(9), 1447–1457, DOI:10.1248/cpb.45.1447.
- A. A. Joseph, V. M. Dhurandhare, C.-W. Chang, V. P. Verma, G. P. Mishra, C.-C. Ku, C.-C. Lin and C.-C. Wang, Chemoselective Per-O-Trimethylsilylation and Homogeneous N-Functionalisation of Amino Sugars, Chem. Commun., 2014, 51(1), 104–106, 10.1039/C4CC06645F.
- K. Bajaj, G. G. Pillai, R. Sakhuja and D. Kumar, Expansion of Phosphane Treasure Box for Staudinger Peptide Ligation, J. Org. Chem., 2020, 85(19), 12147–12159, DOI:10.1021/acs.joc.0c01319.
- C. Grandjean, A. Boutonnier, C. Guerreiro, J.-M. Fournier and L. A. Mulard, On the Preparation of Carbohydrate−Protein Conjugates Using the Traceless Staudinger Ligation, J. Org. Chem., 2005, 70(18), 7123–7132, DOI:10.1021/jo0505472.
- M. Kotormán, I. Laczkó, A. Szabó and L. M. Simon, Effects of Ca2+ on Catalytic Activity and Conformation of Trypsin and α-Chymotrypsin in Aqueous Ethanol, Biochem. Biophys. Res. Commun., 2003, 304(1), 18–21, DOI:10.1016/S0006-291X(03)00534-5.
- M. M. Green, J. A. Gladner, L. W. Cunningham and H. Neurath, The Effects of Divalent Cations on the Enzymatic Activities of Trypsin and of A-Chymotrypsin, J. Am. Chem. Soc., 1952, 74(8), 2122–2123, DOI:10.1021/ja01128a523.
- F. C. Wu and M. Laskowski, The Effect of Calcium on Chymotrypsins α and B, Biochim. Biophys. Acta, 1956, 19, 110–115, DOI:10.1016/0006-3002(56)90391-2.
- R. Kluger, R. W. Loo and V. Mazza, Biomimetically Activated Amino Acids. Catalysis in the Hydrolysis of Alanyl Ethyl Phosphate, J. Am. Chem. Soc., 1997, 119(50), 12089–12094, DOI:10.1021/ja972406q.
- Q. Mao and P. Walde, Substrate effects on the enzymatic activity of α-chymotrypsin in reverse micelles, Biochem. Biophys. Res. Commun., 1991, 178(3), 1105–1112, DOI:10.1016/0006-291X(91)91006-X.
- E. Sturabotti, F. Vetica, G. Toscano, A. Calcaterra, A. Martinelli, L. M. Migneco and F. Leonelli, N-Acetyl-l-Phenylalanine, Racemization during TBTU Amidation: An In-Depth Study for the Synthesis of Anti-Inflammatory 2-(N-Acetyl)-l-Phenylalanylamido-2-Deoxy-d-Glucose (NAPA), Molecules, 2023, 28(2), 581, DOI:10.3390/molecules28020581.
- L. Kiczak, M. Kasztura, K. Koscielska-Kasprzak, M. Dadlez and J. Otlewski, Selection of Potent Chymotrypsin and Elastase Inhibitors from M13 Phage Library of Basic Pancreatic Trypsin Inhibitor (BPTI), Biochim. Biophys. Acta, Protein Struct. Mol. Enzymol., 2001, 1550(2), 153–163, DOI:10.1016/S0167-4838(01)00282-5.
- V. Schellenberger, K. Braune, H.-J. Hofmann and H.-D. Jakubke, The Specificity of Chymotrypsin, Eur. J. Biochem., 1991, 199(3), 623–636, DOI:10.1111/j.1432-1033.1991.tb16163.x.
- A. Ghaffar, G. J. J. Draaisma, G. Mihov, A. A. Dias, P. J. Schoenmakers and S. J. Van Der Wal, Monitoring the in Vitro Enzyme-Mediated Degradation of Degradable Poly(Ester Amide) for Controlled Drug Delivery by LC-ToF-MS, Biomacromolecules, 2011, 12(9), 3243–3251, DOI:10.1021/bm200709r.
- P. J. Sweeney and J. M. Walker, Proteinase K (EC 3.4.21.14), in Enzymes of Molecular Biology, Humana Press, New Jersey, 1993, Vol. 16, pp. 305–312. DOI:10.1385/0-89603-234-5:305.
- K. H. Richau, F. Kaschani, M. Verdoes, T. C. Pansuriya, S. Niessen, K. Stüber, T. Colby, H. S. Overkleeft, M. Bogyo and R. A. L. Van der Hoorn, Subclassification and Biochemical Analysis of Plant Papain-Like Cysteine Proteases Displays Subfamily-Specific Characteristics1[C][W], Plant Physiol., 2012, 158(4), 1583–1599, DOI:10.1104/pp.112.194001.
- T. A. Semashko, E. A. Vorotnikova, V. F. Sharikova, K. S. Vinokurov, Y. A. Smirnova, Y. E. Dunaevsky, M. A. Belozersky, B. Oppert, E. N. Elpidina and I. Y. Filippova, Selective Chromogenic and Fluorogenic Peptide Substrates for the Assay of Cysteine Peptidases in Complex Mixtures, Anal. Biochem., 2014, 449, 179–187, DOI:10.1016/j.ab.2013.12.032.
- C. J. Gray, J. Boukouvalas, R. J. Szawelski and C. W. Wharton, Benzyloxycarbonylphenylalanylcitrulline P-Nitroanilide as a Substrate for Papain and Other Plant Cysteine Proteinases, Biochem. J., 1984, 219(1), 325–328, DOI:10.1042/bj2190325.
| This journal is © The Royal Society of Chemistry 2026 |




![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:
