 Open Access Article
Open Access ArticleExpeditious synthesis of multiglycopeptides with heterogeneous glycan cores derived from an α-dystroglycan mucin-like domain
Dror
Ben Abba Amiel
 a and
Mattan
Hurevich
a and
Mattan
Hurevich
 *ab
*ab
aInstitute of Chemistry, Hebrew University of Jerusalem, Edmond J. Safra Campus, Givat Ram, Jerusalem 9190401, Israel. E-mail: mattan.hurevich@mail.huji.ac.il
bCenter of Nanoscience and Nanotechnology, Hebrew University of Jerusalem, Edmond J. Safra Campus, Givat Ram, Jerusalem 9190401, Israel
First published on 10th February 2026
Abstract
Glycosylation is a post-translational modification prevalent in the majority of proteins. Many glycoproteins contain several glycosylation sites, often bearing different glycan moieties. The inherent difficulties of glycopeptide synthesis worsen for heterogeneously glycosylated peptides, as each glycan introduces unique synthetic hurdles. Stirring-assisted solid-phase synthesis proved extremely valuable in accessing post-translationally modified peptides. We present the stirring-assisted synthesis of a heterogeneous glycopeptide library, derived from α-dystroglycan, bearing a variety of glycosylation patterns combining both mannose and GalNAc cores. The developed strategy streamlined the expeditious assembly with post-assembly manipulation, enabling the procurement of heterogeneously glycosylated peptides with high purity.
Introduction
Glycosylation is the most complex post-translational modification (PTM) of proteins.1 Glycoproteins are crucial in every aspect of biology, with glycosylation playing an imperative role in many functions, from binding to signaling.2–6 Most glycoproteins contain more than one glycosylation site (glycosite), and many of those incorporate more than one kind of glycan core.1 The complexity of glycoproteins goes even further, as the same glycosites might alternate between different glycan cores.7α-Dystroglycan (α-DG) is an extracellular membrane glycoprotein that provides structural integrity to muscle tissues.8,9 α-DG has a dumbbell-like shape with a heavily glycosylated “mucin-like” domain.10,11 This domain presents two types of α-linked O-glycosylation cores, mannose and GalNAc.12 Several sites within this region can be glycosylated either by mannose or by α-GalNAc (Tn antigen).13 Abnormal mannosylation of the mucin-like domain is linked to several types of muscular dystrophies, predominantly Duchenne muscular dystrophy.8 It is crucial to evaluate the effect of glycosylation on α-DG-related diseases, considering that the synthetic construct must mimic both the α-mannose 1,2-trans glycan linkage and the 1,2-cis linked α-GalNAc core.14–16
Due to the complexity and variety of glycoproteins, glycopeptides (GPs) are often used as a simpler synthetic alternative.17–20 GPs are difficult to obtain efficiently by solid-phase peptide synthesis (SPPS).21–25 This difficulty is exacerbated as the complexity of the desired GP increases. Singly-glycosylated GPs already pose a greater synthetic challenge compared to standard peptides due to the steric hindrance of the bulky glycan moiety and the tendency of glycosylated amino acids (GAAs) to undergo racemization or elimination.23,26,27 The addition of multiple glycosylation sites, let alone different ones, increases the synthetic difficulty to a level at which standard SPPS protocols and technologies fail to access them efficiently.28
There are several common methods to obtain synthetic GPs, most of which hinge on the use of a large excess of difficult-to-obtain GAAs or on long coupling reactions.29–32 The use of equimolar quantities of GAAs and the conduction of short coupling reactions are mutually exclusive, requiring researchers to pick one and give up on the other. These impediments are multiplied in the case of multiply glycosylated peptides (MGPs), thus bringing about the great difficulty previously discussed.
The different steps in the solid-phase synthesis process can be classified into diffusion-dependent (DD) and diffusion-independent (DI) reactions. DD reactions, such as AA coupling or Fmoc deprotection, are characterized by kinetics that are faster than the diffusion of solvents and reagents into the resin.33–35 This makes diffusion the rate-limiting process in these reactions. In DD processes, increasing the mixing efficiency facilitates mass transfer via diffusion and hence accelerates reactions. Conversely, DI reactions, such as elimination or epimerization, are slow enough not to be limited by the rate of diffusion and thus do not benefit from improved mixing strategies.36
The solid phase assembly of peptides and GPs is a constant competition between DD reactions that provide the desired outcome and destructive DI reactions that lead to accumulation of side-products.36–39 Inefficient mixing with slow diffusion does not promote DD over DI reactions and therefore drives toward obtaining more side products. As effective diffusion benefits mostly DD reactions, it can assist in obtaining the desired products before DI-promoted side-products accumulate.
Our group has reported the utilization of fast overhead stirring at a high temperature to enable a fast and equimolar assembly of peptides.28,40,41 The enhanced diffusion achieved through stirring promotes fast coupling and deprotection reactions and surpasses common solid-phase techniques.42–45 The strategy became especially beneficial for the synthesis of peptides with complex PTMs.28,41,46 A stirring-assisted assembly of O-xylosylated GPs and their subsequent streamlined deacetylation proved extremely efficient in terms of time and molarity (expeditious glycopeptide synthesis, EGPS, Fig. 1, red).46 The process was further generalized to enable the synthesis of multi-O-mannosylated GPs (expeditious multiply-glycosylated peptide synthesis, EMGPS, Fig. 1, blue).47 EMGPS employs fast stirring at a high temperature (1200 rpm at 90 °C) to accelerate DD processes, such as AA coupling and Fmoc deprotection, and applies a significant excess of hexafluorophosphate azabenzotriazole tetramethyl uronium (HATU) to in situ reactivate the hydrolyzed GAA following activator decomposition. The combination of these techniques allowed the synthesis of triply mannosylated GPs with high crude purity and in large quantity, while using only 1.2 equiv. of GAA and conducting GAA coupling reactions in 1 min. Additionally, glycan deacetylation in EGPS and EMGPS is done swiftly on the solid support, thus making lengthy and wasteful solution deacetylation reactions redundant. Both EGPS and EMGPS have shown no indication of racemization or elimination of GAAs.
The progressive character of EGPS and EMGPS showed that each increase in complexity required reevaluation of the process and optimization of protocols. While working towards this goal, we have encountered obstacles that were insignificant for singly-glycosylated peptides, but became substantial once MGPs were synthesized. This forced us to develop a strategy for adaptation of EMGPS to accommodate multiple and hindered GAAs and obtain complex MGPs.
The heterogeneity of the α-DG “mucin-like” domain presents a complexity that challenges the limits of EMGPS. The methodological evolution of stirring-assisted GP synthesis guided us toward the adaptation of EMGPS to access α-DG-derived heterogeneous multiply glycosylated peptides (HGPs) bearing several types of glycan cores on the same peptide (Fig. 1, green).
Besides mannose, which has already been a focus of our previous project, α-DG contains α-O-GalNAc glycosylation sites, also known as the Tn antigen.48 This glycosylation can be found on membrane-bound and secreted proteins, clustered or in single sites, and as a monosaccharide or in extended glycans.49 The combination of O-Man and the Tn antigen in α-DG, possibly competing on the same glycosites, is intriguing. Exploring the biological significance of these unique combinations requires accessibility to GPs that can mirror such complexity. Several reports have been published on the preparation of an α-GalNAc GAA and the synthesis of Tn-bearing GPs, mostly hinging on a single type of core connectivity.32,50–52
Apart from the obvious novelty of a hitherto-unreported accelerated synthesis of HGPs, the inclusion of α-GalNAc as a target glycosylation in addition to O-Man raises several challenges. An optimization of the assembly process for a new glycan core was required. The adaptation of EMGPS, developed for MGPs with a homogeneous core type, for the synthesis of HGPs was equally important. Unlike α-Man, α-GalNAc has a 1,2-cis AA–glycan linkage, which increases the difficulty in accessing these GAAs in sufficient quantities required for the preparation of HGP libraries. An HGP synthetic protocol must also comply with both 1,2-cis and 1,2-trans GAAs. α-GalNAc also contains an amide C-2 moiety, which requires additional synthetic consideration and is known to promote racemization and elimination.26
The preparation of the Tn antigen almost invariably goes through an azidogalactose.51,53 This GAA then needs to be reduced and acetylated to provide an amidated glycan moiety. Azide reduction has been reported both on the solid support and in solution.54 While the solution reaction allows the reduction of larger quantities, the solid-phase reaction enables easy purification of the product. The diffusion dependence of azide reduction on the solid support was crucial to evaluate in the context of the fast-stirring accelerated GP synthesis process. Understanding the diffusion dependence of solid phase processes is a vital tool for their optimization.
We realized that adaptation of EMGPS to α-DG-derived HGPs will require methodological development and reevaluation of steps that were optimized for other homogeneous GPs. We set out to synthesize a library of α-DG-derived GPs combining O-Man and O-α-GalNAc glycosites.
We opted for a robust synthetic procedure for the efficient preparation and purification of GAAs bearing a 2-azidogalactose moiety. This was followed by an optimization of protocols developed for homogeneous MGP assembly via EMGPS to provide HGPs with both O-Man and O-α-GalNAc cores. We evaluated the ability to perform on-resin azide reduction and to simultaneously deacetylate the different glycan cores while retaining stereochemistry and avoiding other side-reactions. After the method was adapted successfully, a library of α-DG fragments, bearing all possible glycosylation combinations, was synthesized. Finally, we reorganized the HGP synthesis based on diffusion dependency and streamlined the process.
Results and discussion
We aimed to synthesize a library of MGPs derived from the 471–478 “mucin-like” domain of α-DG. This short sequence contains two glycosites that can bear either O-Man or O-GalNAc, one on a Ser and one on a Thr.7 Nine peptides with the α-DG 471–478 sequence LETASPPT were envisioned, wherein αDG-1 is the non-glycosylated peptide, αDG-2–αDG-5 are monoglycopeptides, αDG-6 and αDG-7 are homogeneous MGPs, and αDG-8 and αDG-9 are the target HGPs (Fig. 2, table). This set of peptides covers all possible core glycosylation patterns of this sequence.The assembly of this library would require the use of four different GAAs: mannosylated Ser and Thr (1s and 1t, respectively, Fig. 2), which were prepared in our previous work, and α-galactosylated Ser and Thr (2s and 2t, respectively).47
The synthesis of GAAs containing α-GalNAc is more challenging than the mannosylated ones because of the inherent difficulty in installing a cis linkage of a 2-deoxy monosaccharide that can serve as a precursor for the native 2-acetamide Tn antigen.55 We aimed to prepare 2s and 2t, both bearing 2-azidogalactose triacetate, which could later be reduced to the amide. The reduction step could be done on the solid support in order to test its diffusion dependence and our ability to avoid its unpleasant purification process. We looked for a robust and scalable route to provide enough GAA to use for the synthesis of the entire α-DG 471–478 GP library. We aimed first to evaluate a route for synthesizing 2s. To achieve that, 1-selenophenyl-2-azidogalactose triacetate 3 was hydrolyzed in the anomeric position to give 4 (Scheme 1).53 In parallel, Fmoc-L-serine tert-butyl ester 8 was prepared.56 From this point, we compared two synthetic pathways. In route I (Scheme 1, green), 4 was introduced with a trifluoro-N-phenyl imidate moiety to produce the highly reactive glycosyl donor 6.
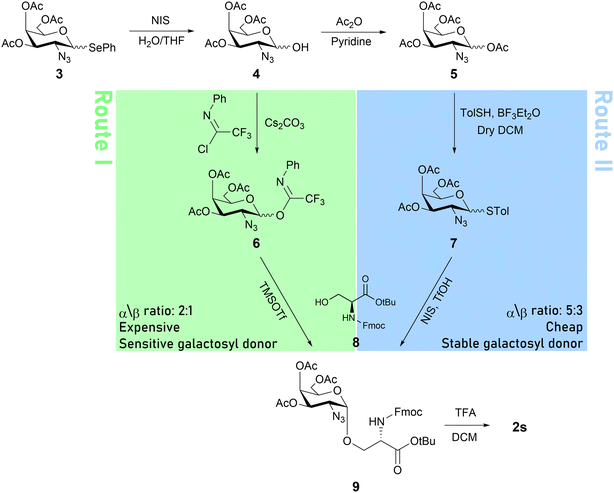 | ||
| Scheme 1 The synthetic routes to obtain 2s. Green: route I, relying on trifluoro-N-phenyl imidate as the donor LG. Blue: route II, relying on p-tolylthiol as the donor LG. | ||
Serine acceptor 8 was glycosylated with the azide galactosyl donor 6 using a catalytic amount of TMSOTf activator to provide GAA 9 in a yield of about 25% with 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 α/β selectivity in a mixture with unreacted 8. In route II (Scheme 1, blue), the anomeric position of 4 was acetylated to give 5, which was thioglycosylated with p-thiocresol to yield donor 7 at a 1
1 α/β selectivity in a mixture with unreacted 8. In route II (Scheme 1, blue), the anomeric position of 4 was acetylated to give 5, which was thioglycosylated with p-thiocresol to yield donor 7 at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 α/β ratio. Protected Ser 8 was glycosylated with 7 to give 9 in a yield of about 25% with 5
1 α/β ratio. Protected Ser 8 was glycosylated with 7 to give 9 in a yield of about 25% with 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 α/β selectivity, also in a mixture with unreacted 8.
3 α/β selectivity, also in a mixture with unreacted 8.
At this point, we had to choose between the two methods to upscale the synthesis of 9. Route I employs an expensive reagent (2,2,2-trifluoro-N-phenylacetimidoyl chloride) to obtain the unstable donor 6, which is sensitive to temperature, acidic conditions and moisture. Route II, on the other hand, utilizes much cheaper reagents to obtain 7, which is benchtop-stable. While route I provided 9 at a slightly higher α/β ratio, in both cases purification was required. An optimized chromatography facilitated the isolation of 9 and its separation from the β anomer.57 Given this separation and the marginal α-selectivity advantage of route I over route II, we decided to use the latter for the synthesis of 9, as it relies on stable intermediates, which allows upscaling and reproducibility. To complete the preparation of 2s, pure 9, obtained via route II, was reacted with diluted trifluoroacetic acid to remove the tBu. Route II was also used for the synthesis of 2t without any setbacks (see the SI for more details).
After GAAs 2s and 2t were obtained in sufficient quantities, EMGPS was optimized for the use of these azidogalactosylated AAs. We conducted our optimization on a relatively hydrophobic sequence, derived from the huDKK1 protein, that contains a GalNAc-serine (gS), as it enabled following the assembly, deprotection, and reduction steps via HPLC analysis (D1, SAVgSAAPGIL, Fig. 3).58 The GP D1 was assembled via the EMGPS protocol using 2s as a GAA. The last AA was left protected by Fmoc to avoid acetylation of the terminal amine during azide reduction (see the SI for more details).59 A small sample of the resin was cleaved and analyzed (Fig. 3, violet), revealing that the target peptide was obtained with a crude purity of 79%, with the other 21% being a gS deletion sequence (des, Fig. 3). This showed that although 2s coupling does not proceed to completion, the protocol developed for mannosylated GAAs is applicable for GPs containing azidogalactosylated ones.
With the assembled GP in hand, we set to reduce the azidogalactose to GalNAc while the GP is still attached to the solid support. On-resin azide reduction usually proceeds in a lengthy overnight reaction, and we decided to check its applicability to EMGPS.53,54 The reaction was performed first by shaking resin with D1-A with 3 mL of a thioacetic acid/pyridine mixture (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 v/v) overnight to give a 95% conversion of the azide to NAc (D1-N, Fig. 3, green, des remained unchanged).26 In parallel, 1200 rpm stirring of a resin with D1-A for 30 min at rt in the above reducing mixture resulted in minor conversion to D1-N (Fig. 3, dashed blue). This showed that stirring does not expedite the azide reduction under these conditions, suggesting that diffusion is not a rate-limiting factor in this reaction.
1 v/v) overnight to give a 95% conversion of the azide to NAc (D1-N, Fig. 3, green, des remained unchanged).26 In parallel, 1200 rpm stirring of a resin with D1-A for 30 min at rt in the above reducing mixture resulted in minor conversion to D1-N (Fig. 3, dashed blue). This showed that stirring does not expedite the azide reduction under these conditions, suggesting that diffusion is not a rate-limiting factor in this reaction.
To complete the evaluation of EMGPS protocols, on-resin deacetylation of D1-N was performed in three 5 minute intervals of fast stirring with a 160 mM solution of NaOMe in MeOH at rt.46,47 HPLC analysis of the product has shown a complete deacetylation to D1 (Fig. 3, red). The reaction also removes the Fmoc group, which permitted a parallel unmasking of the peptide terminal amine and the glycan hydroxyls. This places Fmoc as a good protecting group for this strategy, as it protects the amine from acetylation during manipulation of the azide and could then be removed without adding more steps to the existing protocol. The above evaluations showed that by adding a single synthetic step, the EMGPS protocol could be used to synthesize GPs with an α-GalNAc moiety using an equimolar amount of GAA. This ability to use equimolar amounts of GAAs is an enabling capacity crucial for the synthesis of mucin-derived GPs. While the GAAs 1s and 1t were accessed from the readily available Fmoc-Ser/Thr and the peracetylated monosaccharides in a single synthetic step and minimal workup, obtaining 2s and 2t from similar starting materials was possible only via a tedious multistep process with nontrivial purification steps. Consequently, it is imperative to use them as efficiently as possible for the process to be viable.
We named this EMGPS-adapted method Protocol A, describing a process in which the expeditious assembly of the peptides is followed by reduction, deacetylation, and cleavage, in this order (Fig. 4, top). We utilized Protocol A and EMGPS to synthesize a library of mono-, multi-, and heterogeneously glycosylated α-DG fragments. The HGP αDG-9, which bears both gS and Thr-O-Man, was synthesized following Protocol A as a first example for heterogeneous MGP (Fig. 4, pink). The post-cleavage analysis shows that αDG-9 was obtained with 79% crude purity. αDG-8, with the Tn antigen on Thr and O-Man on Ser, was similarly synthesized using Protocol A. All GPs containing α-GalNAc, i.e., αDG-4, αDG-5, and αDG-7, were prepared via Protocol A, while the rest of the peptides were synthesized via EMGPS. The assembly of each of the sequences produced via Protocol A was done in about 30 min, and their subsequent deacetylation was completed in 20 min. All the GPs were obtained with moderate to good crude purity (Fig. 4), isolated and characterized with HPLC and HRMS. The library represents all possible combinations of glycosylations and non-glycosylations on these two sites.
 | ||
| Fig. 4 Top: synthetic procedure of Protocol A used for synthesis of the αDG-n library. Bottom left: preparative HPLC chromatograms of αDG-n products and their calculated crude purities according to peak integration. The legend of αDG-n glycosylations from Fig. 2 is provided again in this figure to assist the reader in comprehension. Bottom right: HSQC NMR spectra of the anomeric regions of αDG-8 and αDG-9. It can be seen that the Thr-bonded anomeric signals are shifted slightly downfield, both in 1H and in 13C, compared to their Ser-bonded counterparts in the other HGP. | ||
α-DG-8 and αDG-9 were characterized by NMR to validate their structure and the integrity of the glycan cores (Fig. 4, right). The characteristic aliphatic signals (Leu, Ala, Thr, and Pro) and the acetyl group of GalNAc confirmed that the common peptide features of the HGPs are present. HSQC-NMR in both GPs showed two anomeric-indicative carbon signals at around 100 ppm.60 These carbons correlated with proton signals at 4.90 and 4.94 ppm in αDG-9 and at 4.90 and 4.97 ppm in αDG-8. These signals prove that the GP has two different types of glycosidic linkages, and the distinctive 13C shifts are indicative of a difference between the glycan moieties. The signal at 101.1 ppm in the spectrum of αDG-9, correlating with the 1H signal at 4.90 ppm, is typical of O-mannosylated GPs (100.3–101.3 ppm) and in agreement with our previous studies.47 Similarly, the azidogalactosylated AAs 2s and 2t presented anomeric signals at 97.9 and 98.9 ppm, respectively, which match the signal at 97.7 ppm, correlating with the 1H signal at 4.94. The 1H–13C correlations found for the anomeric carbon and protons in αDG-8 follow the same trend. When comparing the two spectra, both anomeric protons bonded to the threonine are shifted slightly downfield compared to the same peak in the other GP, which is to be expected due to the rigidity of the glycan–Thr bond.61–63 Both HGPs were also analyzed by coupled HSQC, with all signals showing characteristic coupling constants of 170–174 Hz associated with the α configuration, proving that no epimerization took place.64
In our previous work, we aimed for a streamlined approach where all synthetic stages are done continuously. This is done in the interest of efficiency, as every vessel change is time-consuming and may reduce the overall yield. As Protocol A alternates between the use of expeditious fast-stirring DD steps and slower DI ones, we wanted to streamline the process by eliminating much of this oscillation. To do that, we evaluated if all DD steps could be grouped, followed by the DI ones (Fig. 5, blue and orange, respectively). We realized that the coupling and deprotection steps of the peptide assembly, as well as the acetyl deprotection step, benefit from fast stirring and can be done in succession using the same fast-stirring reactor. The reduction and cleavage can be done at a post-assembly synthesis stage as both are DI processes that do not benefit much from fast stirring. To that end, we devised the sequence αDG-9-Ac (Fig. 5), which is similar to αDG-9 except that we used N-acetylleucine instead of Fmoc-leucine as the last AA to be coupled, rendering the terminal amine permanently protected. By blocking the terminal amine with a base-stable group, the deprotection of the glycan acetyls can be done directly after the assembly of the HGP and before the reduction of the azide. This will allow grouping of DD and DI processes. αDG-9-Ac was assembled and deacetylated in succession according to the EMGPS procedure within less than 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 h and then left overnight in AcSH/pyridine and cleaved. This protocol, named Protocol B (Fig. 5, bottom), provided αDG-9-AC with a crude purity of 51%, indicating that the process can be streamlined by using a base-stable terminal amine PG. αDG-9-Ac was purified by HPLC and analyzed by NMR and HRMS, confirming its identity and similarity to αDG-9. Given that Protocol B allowed the efficient and streamlined synthesis of HGPs, we have termed it expeditious heterogeneous glycosylated peptide synthesis (EHGPS). Interestingly, while in the synthesis of αDG-9 deacetylation takes place in the presence of an azidogalactose moiety, in the synthesis of αDG-9-Ac deacetylation is done in the presence of the reduced GalNAc moiety. In both cases, no elimination was observed, proving that the fast deacetylation can be used for GPs containing both forms of Tn precursors. The HGPs αDG-8 and αDG-9 were obtained via Protocol A at quantities of 4.7 mg and 7 mg, respectively, representing overall yields of 22% and 33%. The HGP αDG-9-Ac was obtained via EHGPS in 6 mg, representing a yield of 27%. This indicates that both protocols provide similar overall product yields.
30 h and then left overnight in AcSH/pyridine and cleaved. This protocol, named Protocol B (Fig. 5, bottom), provided αDG-9-AC with a crude purity of 51%, indicating that the process can be streamlined by using a base-stable terminal amine PG. αDG-9-Ac was purified by HPLC and analyzed by NMR and HRMS, confirming its identity and similarity to αDG-9. Given that Protocol B allowed the efficient and streamlined synthesis of HGPs, we have termed it expeditious heterogeneous glycosylated peptide synthesis (EHGPS). Interestingly, while in the synthesis of αDG-9 deacetylation takes place in the presence of an azidogalactose moiety, in the synthesis of αDG-9-Ac deacetylation is done in the presence of the reduced GalNAc moiety. In both cases, no elimination was observed, proving that the fast deacetylation can be used for GPs containing both forms of Tn precursors. The HGPs αDG-8 and αDG-9 were obtained via Protocol A at quantities of 4.7 mg and 7 mg, respectively, representing overall yields of 22% and 33%. The HGP αDG-9-Ac was obtained via EHGPS in 6 mg, representing a yield of 27%. This indicates that both protocols provide similar overall product yields.
The adaptability of the EMGPS process is one of its great features. As the EGPS protocol was adapted for the coupling of several mannosylated AAs to provide EMGPS, we aimed to broaden the scope of usable GAAs and the variety of synthetic manipulations. GAAs, and especially cis-linked ones, are very precious and difficult to obtain. After we have successfully prepared and purified 2s and 2t, it is imperative to use the minimal necessary quantity for every synthesis. EMGPS allowed conducting an optimization experiment within an hour, which could then be analyzed the same day and enable the optimization of the next synthetic step. The fast and GAA-economical optimization was detrimental for the development of a robust and reproducible strategy to access MGPs and HGPs derived from the α-DG 471–478 region.
The optimization had additional merit, as it allowed categorizing the steps based on their diffusion dependency. The reduction and acetylation of the azide were not expedited by fast stirring. This indicated, given the high concentration of reagent, that diffusion is not the rate-limiting factor in these reactions. However, there is still a practical advantage to this, as it allows facilitating the synthesis of the library. In practice, several GPs were assembled in succession quickly and efficiently via EMGPS in a single day and then left for reduction in parallel. Additionally, being able to conduct the reduction on the solid support meant that this could be done before deacetylation, allowing the use of Fmoc as a terminal amine protecting group.
Deacetylation is a problematic step with a risk of elimination, especially for α-GalNAc.27,62 Although the fast on-resin deacetylation used here significantly accelerates the diffusion of the methoxide, it did not result in significant unwanted products in any of the GPs. The efficiency of deacetylation demonstrated for αDG-2–9 shows that the conditions can be applied for the α-azidogalactose core, and the deacetylation of αDG-9-Ac shows that they can be equally applicable for the GalNAc core. This adds to the successful deacetylations of a hexose (O-Man) and a pentose (O-Xyl), showing the versatility and generality of the protocol.46,47 The results further prove that simultaneous deacetylation of different glycan cores can be done with no change in the protocol. The ability to perform a streamlined, on-resin deacetylation process surpasses other state-of-the-art SPPS acceleration methods.
Another important facet of the library synthesis is the additional insights obtained regarding the azide reduction process. D1 provided the initial confirmation that the reaction is DI and that the overnight on-resin reduction was effective. αDG-4 and αDG-5 have shown that reduction could be done equally on Ser and Thr glycosites. αDG-7 demonstrated that the reaction was compatible with multiple azide moieties on the same GP, and αDG-8 and9 established its effectiveness for GPs that contain other glycan cores. In all GalNAc-bearing GPs, a near-perfect conversion rate without significant side-products was observed. This indicates that the reduction does not affect other functionalities of the protected GPs and hence could be integrated with other GAAs. In contrast to bulk reduction of the GAA in solution, the reduction on the solid support is performed separately for each GP. As the reduction step in EHGPS takes place after the accelerated assembly and deacetylation of the HGPs, it can be done simultaneously for several GPs without significantly increasing the time consumption. Additionally, avoiding the extra reaction and the resulting purification step required for converting the azide to the GalNAc-bearing GAA in the already lengthy multistep process in solution is another advantage, which rationalizes the decision to perform the reduction on the solid support.
The development of EHGPS provided us with an evaluation of the design and organization of synthetic processes based on their diffusion dependency. There are five processes involved in the synthesis of these GPs – three DD reactions (coupling, Fmoc deprotection, and deacetylation) and two DI ones (azide reduction and cleavage). In EMGPS, all DD processes are done in succession in the same vessel, leaving only the DI cleavage to be done later. The adaptation of the process to azidogalactose has upended this order, since the DI reduction in Protocol A takes place between the DD coupling–deprotection cycles and the DD deacetylation step. This resulted in three unfavored transitions between the DD and DI steps. Moving the reduction to follow the deacetylation in EHGPS results in an advantageous arrangement where all DD reactions are grouped. When streamlining EHGPS, the terminal amine of the peptide must be protected during thioacetic acid-assisted reduction. In this study, Ac blocking of the terminal AA was used. In cases where a terminal free amine is important for the activity, an acid-labile PG (e.g., Boc) can be used and removed in situ with the cleavage.
The streamlined EHGPS strategy can also be advantageous for the synthesis of larger quantities of HGPs. The reproducibility of the process, demonstrated here and in previous reports, suggests that when done in the same reaction vessel and scale, repeating the process multiple times in the accelerated process can provide large quantities of GPs in a short time.47 As the DI azide reduction can be performed in bulk for the combined synthesized HGP batches, the overall time for scaling up will not be affected. This will enable the procurement of significant amounts of HGPs within a few days of work.
The α-DG fragment library testifies to the evolution of EGPS, providing access to simple peptides, through monoglycosylated GPs with either Man or GalNAc and homogeneous MGPs, all the way to HGPs. The assembly of a library of eight GPs is a daunting task that would take a long time and an excess of GAAs in the state-of-the-art methods, even without the additional challenges of multiple glycosylations and heterogeneous glycan cores. The simple requirements of EHGPS and its efficient optimization enabled the synthesis of an α-DG library at a remarkable speed using standardized steps.
We have previously stressed that the major highlights of the fast-stirring approach to peptide synthesis are its speed and equimolarity. As we broaden the scope of attainable products, we show that adaptability is another great advantage. We demonstrate that the stirring-assisted protocols could be adapted to complex glycan moieties and enable multiple types of manipulations. Adaptation and generalization allow access to GPs with pentoses, hexoses, azidohexoses and hexoseamides. This shows that countless possible GPs and applications can be accessed by EGPS and its advanced protocols, providing us with a glimpse of what further optimization and adaptation can achieve.
Conclusions
We introduced an adapted variant of the EGPS protocol, developed for the synthesis of GalNAc-bearing GPs. A procedure for the synthesis and purification of α-azidogalactosylated GAAs was found, and an optimization of the coupling, reduction, and deacetylation of these GAAs was carried out. We found that these GAAs can be used in conjunction with mannosylated ones to perform an accelerated synthesis of HGPs. The fast deacetylation protocol is equally effective for multiple and different types of glycosyl cores, cementing its robustness. The adaptability of the process was utilized to streamline it, culminating in EHGPS. The EHGPS protocol can provide access to a great variety of HGP libraries, similar to the αDG-n library. As HGP libraries are a vital need in the characterization of protein glycosylation, their fast and efficient synthesis will propel applications to understand, detect, and tackle diseases.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: Table S1, Scheme S1 and Fig. S1–S63 provide HPLC chromatograms, NMR spectra and further experimental details. See DOI: https://doi.org/10.1039/d6ob00013d.Acknowledgements
The authors would like to thank Dr Deepak Ganesh for his assistance with flash chromatography. M. H. thanks the Israel Science Foundation grant no. 1805/22 for the support.References
- R. Apweiler, H. Hermjakob and N. Sharon, Biochim. Biophys. Acta, Gen. Subj., 1999, 1473, 4–8 CrossRef CAS PubMed.
- Z. Chen, H. Yu, X. Chen, W. Chen, W. Song and Z. Li, Int. J. Biol. Macromol., 2023, 236, 123818 CrossRef CAS PubMed.
- M. He, X. Zhou and X. Wang, Signal Transduction Targeted Ther., 2024, 9, 1–33 Search PubMed.
- H. Läubli and L. Borsig, Front. Immunol., 2019, 10, 484841 Search PubMed.
- F. Ye, C. Li, F. L. Liu, X. Liu, P. Xu, R. H. Luo, W. Song, Y. T. Zheng, T. Ying, B. Yu and P. Wang, Natl. Sci. Rev., 2024, 11, nwae030 CrossRef CAS PubMed.
- A. C. Conibear, E. E. Watson, R. J. Payne and C. F. W. Becker, Chem. Soc. Rev., 2018, 47, 9046–9068 RSC.
- S. H. Stalnaker, S. Hashmi, J. M. Lim, K. Aoki, M. Porterfield, G. Gutierrez-Sanchez, J. Wheeler, J. M. Ervasti, C. Bergmann, M. Tiemeyer and L. Wells, J. Biol. Chem., 2010, 285, 24882–24891 CrossRef CAS PubMed.
- T. Endo, J. Biochem., 2015, 157, 1–12 Search PubMed.
- T. Endo, Acta Myol., 2007, 26, 165–170 CAS.
- L. Wan, X. Ge, Q. Xu, G. Huang, T. Yang, K. P. Campbell, Z. Yan and J. Wu, Nature, 2024, 637, 1252–1260 Search PubMed.
- P. T. Martin, Glycobiology, 2003, 13, 55R–66R Search PubMed.
- D. T. Tran, J. M. Lim, M. Liu, S. H. Stalnaker, L. Wells, K. G. Ten Hagen and D. Live, J. Biol. Chem., 2012, 287, 20967–20974 Search PubMed.
- S. H. Stalnaker, S. Hashmi, J. M. Lim, K. Aoki, M. Porterfield, G. Gutierrez-Sanchez, J. Wheeler, J. M. Ervasti, C. Bergmann, M. Tiemeyer and L. Wells, J. Biol. Chem., 2010, 285, 24882–24891 CrossRef CAS PubMed.
- C. Quereda, À. Pastor and J. Martín-Nieto, Cancer Cell Int., 2022, 22, 1–30 Search PubMed.
- H. Manya and M. Kanagawa, J. Hum. Genet., 2025, 2025, 1–5 Search PubMed.
- M. Hopkinson and A. A. Pitsillides, Int. J. Exp. Pathol., 2025, 106, e12525 CrossRef CAS PubMed.
- C. M. Potel, M. L. Burtscher, M. Garrido-Rodriguez, A. Brauer-Nikonow, I. Becher, C. Le Sueur, A. Typas, M. Zimmermann and M. M. Savitski, Nat. Struct. Mol. Biol., 2025, 2025, 1–16 Search PubMed.
- J. Hang, J. Wang, M. Lu, Y. Xue, J. Qiao and L. Tao, Biomed. Pharmacother., 2022, 148, 112685 CrossRef CAS PubMed.
- Z. Roth, G. Yehezkel and I. Khalaila, Int. J. Carbohydr. Chem., 2012, 2012, 640923 Search PubMed.
- B. Wei, P. Huang, X. Wang, Z. Liu, F. Tang, W. Huang, B. Liu, F. Ye and P. Wang, Angew. Chem., Int. Ed., 2025, 64, e202415565 CrossRef CAS PubMed.
- L. Wang, N. Wang, W. Zhang, X. Cheng, Z. Yan, G. Shao, X. Wang, R. Wang and C. Fu, Signal Transduction Targeted Ther., 2022, 7, 1–27 Search PubMed.
- A. J. Pereira, L. J. de Campos, H. Xing and M. Conda-Sheridan, Med. Chem. Res., 2024, 33, 1275–1280 CrossRef CAS.
- O. Seitz, ChemBioChem, 2000, 1, 214–246 CrossRef CAS PubMed.
- C. P. R. Hackenberger, M. K. O'Reilly and B. Imperiali, J. Org. Chem., 2005, 70, 3574–3578 CrossRef CAS PubMed.
- D. M. M. Jaradat, H. Hamouda and C. P. R. Hackenberger, Eur. J. Org. Chem., 2010, 5004–5009 CrossRef CAS.
- Y. Zhang, S. M. Muthana, D. Farnsworth, O. Ludek, K. Adams, J. J. Barchi and J. C. Gildersleeve, J. Am. Chem. Soc., 2012, 134, 6316–6325 CrossRef CAS PubMed.
- P. Sjölin and J. Kihlberg, J. Org. Chem., 2001, 66, 2957–2965 CrossRef PubMed.
- P. Strauss, F. Nuti, M. Quagliata, A. M. Papini and M. Hurevich, Org. Biomol. Chem., 2023, 21, 1674–1679 RSC.
- P. Dong, S. Cheng, Y. Wang, H. Gao, Y. Zhang, T. Zhu, P. Yu and X. Meng, Chem. Commun., 2022, 58, 8642–8645 RSC.
- Y. Maki, S. K. Mong, C. Chandrashekar, B. E. Forbes, M. A. Hossain, S. Yamaguchi, C. M. Fadzen, Y. Kajihara and B. L. Pentelute, Chem. Sci., 2025, 16, 7929–7935 RSC.
- J. Zhao, F. Ye, P. Huang and P. Wang, Curr. Opin. Chem. Biol., 2023, 77, 102405 CrossRef CAS PubMed.
- A. Galashov, E. di Gregorio, P. Ponomareva, M. Safferthal, E. Kazakova, L. Bechtella, K. Pagel, M. Pigaleva, B. Joseph and O. Seitz, Angew. Chem., 2025, 137, e202508278 CrossRef.
- C. Viklund, F. Svec, J. M. J. Fréchet and K. Irgum, Chem. Mater., 1996, 8, 744–750 Search PubMed.
- J. Wood and L. F. Gladden, Chem. Eng. Sci., 2002, 57, 3033–3045 CrossRef CAS.
- W. Rudzinski and W. Plazinski, J. Phys. Chem. C, 2007, 111, 15100–15110 Search PubMed.
- J. N. Naoum, I. Alshanski, A. Gitlin-Domagalska, M. Bentolila, C. Gilon and M. Hurevich, Org. Process Res. Dev., 2019, 23, 2733–2739 Search PubMed.
- F. Ye, H. Bai, X. Liu, P. Xu, G. Ding, P. Huang, X. Zhang, B. Yu and P. Wang, Angew. Chem., Int. Ed., 2025, 64, e202518002 CrossRef CAS PubMed.
- X. Sun, F. Ye, D. Hu and P. Wang, J. Am. Chem. Soc., 2025, 147, 48244–48253 Search PubMed.
- H. Bai, F. Ye, D. Purnachandar, X. Liu, P. Huang and P. Wang, J. Am. Chem. Soc., 2025, 147, 34011–34018 Search PubMed.
- J. N. Naoum, I. Alshanski, G. Mayer, P. Strauss and M. Hurevich, Org. Process Res. Dev., 2022, 26, 129–136 Search PubMed.
- D. Grunhaus, E. R. Molina, R. Cohen, T. Stein, A. Friedler and M. Hurevich, Org. Process Res. Dev., 2022, 26, 2492–2497 CrossRef CAS PubMed.
- S. Pinzón-López, M. Kraume, J. Danglad-Flores and P. H. Seeberger, React. Chem. Eng., 2023, 8, 2951–2962 RSC.
- S. Pinzón-López, D. Ebert, E. E. Reuber, M. Kraume, P. H. Seeberger and J. Danglad-Flores, Org. Process Res. Dev., 2025, 29, 2275–2286 CrossRef.
- J. N. Naoum, I. Alshanski, A. Gitlin-Domagalska, M. Bentolila, C. Gilon and M. Hurevich, Org. Process Res. Dev., 2019, 23, 2733–2739 CrossRef CAS.
- I. Alshanski, M. Bentolila, A. Gitlin-Domagalska, D. Zamir, S. Zorsky, S. Joubran, M. Hurevich and C. Gilon, Org. Process Res. Dev., 2018, 22, 1318–1322 CrossRef CAS.
- D. Ben Abba Amiel and M. Hurevich, Eur. J. Org. Chem., 2022, 2022(38), e202200623 CrossRef.
- D. Ben Abba Amiel, H. Okshtein, I. Alshanski, Z. Hayouka, S. Yitzchaik and M. Hurevich, J. Med. Chem., 2025, 68, 26513–26524 Search PubMed.
- I. Brockhausen, H. H. Wandall, K. G. Ten Hagen and P. Stanley, In Essentials of Glycobiology [Internet], ed. A. Varki, R. D. Cummings and J. D. Esko, Cold Spring Harbor Laboratory Press, New York, 4th edn, Chapter 10, 2022 Search PubMed.
- C. Bello, E. Pranzini, E. Piemontese, M. Schrems, M. L. Taddei, L. Giovannelli, M. Schubert, C. F. W. Becker, P. Rovero and A. M. Papini, ChemBioChem, 2023, 24, e202200741 CrossRef CAS PubMed.
- W. Ma, J. Luo, H. Liu, Q. Du, T. Hao, Y. Jiang, Z. Huang, L. Lan, Z. Li and T. Li, Angew. Chem., Int. Ed., 2025, 64, e202424312 Search PubMed.
- W. Liu, P. He, S. Shang and Z. Tan, J. Carbohydr. Chem., 2023, 42, 223–251 CrossRef CAS.
- A. K. Mishra, E. E. Reuber, D. Rai, L. E. Sander, J. Duerr, S. Y. Graeber, M. A. Mall, B. C. Fries, P. H. Seeberger and S. S. Kulkarni, Angew. Chem., 2025, 2025, e24231 Search PubMed.
- M. Hurevich and P. H. Seeberger, Chem. Commun., 2014, 50, 1851–1853 RSC.
- H. Paulsen, T. Bielfeldt, S. Peters, M. Meldal and K. Bock, Liebigs Ann. Chem., 1994, 1994, 369–379 CrossRef.
- A. Galashov, E. Kazakova, C. E. Stieger, C. P. R. Hackenberger and O. Seitz, Chem. Sci., 2024, 15, 1297–1305 Search PubMed.
- S.-X. Liu, Y.-T. Tsai, Y.-T. Lin, J.-Y. Li and C.-C. Chang, Tetrahedron, 2019, 75, 130776 Search PubMed.
- Y. Nakahara, Y. Nakahara and T. Ogawa, Carbohydr. Res., 1996, 292, 71–81 Search PubMed.
- M. Haniu, T. Horan, C. Spahr, J. Hui, W. Fan, C. Chen, W. G. Richards and H. S. Lu, Protein Sci., 2011, 20, 1802–1813 CrossRef CAS PubMed.
- T. Rosen, I. M. Lico and D. T. W. Chu, J. Org. Chem., 1988, 53, 1580–1582 CrossRef CAS.
- G. D. Brown, J. Bauer, H. M. I. Osborn and R. Kuemmerle, ACS Omega, 2018, 3, 17957–17975 CrossRef CAS PubMed.
- F. Corzana, J. H. Busto, G. Jiménez-Osés, M. G. De Luis, J. L. Asensio, J. Jiménez-Barbero, J. M. Peregrina and A. Avenoza, J. Am. Chem. Soc., 2007, 129, 9458–9467 CrossRef CAS PubMed.
- J. J. Barchi, Biopolymers, 2013, 99, 713–723 CrossRef CAS PubMed.
- J. J. Barchi and C. N. Strain, Front. Mol. Biosci., 2023, 10, 1117850 CrossRef CAS PubMed.
- C. Fontana and G. Widmalm, Chem. Rev., 2023, 123, 1040–1102 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |




