Open Access Article
Open Access ArticleAn integrative chemical and genomic similarity approach linking fungal secondary metabolites and biosynthetic gene clusters
Karin
Steffen
a,
Manuel
Rangel-Grimaldo
bc,
Thomas J. C.
Sauters
a,
David C.
Rinker
a,
Huzefa A.
Raja
 b,
Tyler N.
Graf
b,
Adiyantara
Gumilang
a,
Olivia L.
Riedling
a,
Gustavo H.
Goldman
d,
Nicholas H.
Oberlies
b and
Antonis
Rokas
b,
Tyler N.
Graf
b,
Adiyantara
Gumilang
a,
Olivia L.
Riedling
a,
Gustavo H.
Goldman
d,
Nicholas H.
Oberlies
b and
Antonis
Rokas
 *a
*a
aDepartment of Biological Sciences and Evolutionary Studies Initiative, Vanderbilt University, Nashville, TN 37235, USA. E-mail: antonis.rokas@vanderbilt.edu
bDepartment of Chemistry and Biochemistry, University of North Carolina at Greensboro, Greensboro, NC 27402, USA
cDepartment of Natural Products, Institute of Chemistry, Universidad Nacional Autónoma de México, Mexico City, 04510, Mexico
dFaculdade de Ciencias Farmacêuticas de Ribeirão Preto, Universidade de São Paulo, Ribeirão Preto 14040-903, Brazil
First published on 14th January 2026
Abstract
Fungi are well known to biosynthesize structurally complex secondary metabolites (SMs) with diverse bioactivities. These fungal SMs are frequently produced by biosynthetic gene clusters (BGCs). Linking SMs to their BGCs is key to understanding their chemical and biological functions. Reasoning that structural similarity of SMs arises from similarities in the genes involved in their biosynthesis, we developed an integrative approach that leverages known SM–BGC pairs to infer links between detected SMs and genome-predicted BGC regions in fungi. As proof of concept, we structurally characterized 60 metabolites from metabolomic data of 16 strains of the filamentous fungus Aspergillus fischeri. Our approach assigned 22 to known SM–BGC pairs and proposed specific links to BGCs and genetic pathways for the remaining 38 metabolites. These results suggest that coupling chemical structure similarity and genomic sequence similarity is a straightforward and high-throughput approach for linking fungal SMs to their BGCs.
Introduction
Secondary or specialized metabolites (SMs), together with allelochemicals, effectors, and extrolites, are all molecules isolated from Nature that are instrumental to fungal ecology.1 Fungal SMs contribute to various functions, including micronutrient acquisition (e.g., siderophores such as ferrichrome2), defense (e.g., antibacterials such as penicillin3), and pathogenicity (e.g., virulence factors such as gliotoxin4 and ToxA5,6). By virtue of their potent bioactivities, SMs are essential to drug discovery pipelines7,8 and for medical and agricultural research more broadly.9In fungal genomes, the pathways involved in SM biosynthesis typically contain a set of neighboring, co-regulated genes, collectively referred to as biosynthetic gene clusters or BGCs.1 A typical BGC contains genes coding for ‘core’ or ‘backbone-forming’ enzymes responsible for the biosynthesis of the scaffold of the SM, tailoring enzymes that modify the scaffold, and cluster-specific transcription factors and transporters.1 The clustering and content of genes in fungal secondary metabolite pathways led to the development of many different methods to predict BGCs.10 These include CASSIS, a tool for predicting BGCs around a given anchor (or backbone) gene;11 CLOCI, which predicts BGCs based on co-occurring loci and orthologous clusters;12 DeepBGC, a machine learning-based tool trained on distinguishing BGC genomic regions from non-BGC regions in prokaryotic genomes;13 the fai and zol set of tools that employ sequence orthology information for targeted detection of BGCs across genomes,14 and protein domain-based tools like the popular antiSMASH15 that predict BGC presence using profile hidden Markov models targeting required biosynthetic domains, along with BGC class-specific rules.15,16
Widespread access of column chromatography coupled with mass spectrometry (i.e., LC-MS and LC-MS/MS or LC-MSn) has driven the annotation of metabolites from extracts of fungal cultures and even in situ from the cultures themselves.17–19 Yet, due to technical limitations, the degree of certainty of an observation of a SM can vary based on the approach used.20 Assigning the chemical identity, and hence structure, of compounds within an extract of an organism can be categorized into four levels of certainty:21 (1) identified compounds for which there are orthogonal supporting structural data, (2) putatively annotated compounds for which there are matches to spectral libraries, (3) putatively characterized compound classes for which there are matches to the class of compounds, if not the specific compound, and (4) unknown compounds. For the purposes of this report, we focused on the identified compounds (#1 in the list above), where the compounds were isolated and characterized by mass spectrometry and NMR spectroscopy or there were matches to a dereplication database that was built upon fully characterized compounds.22,23
To date, more than 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 fungal metabolites have been characterized,24 and genomic examinations suggest that there are likely millions of predicted BGCs in fungal genomes.25–29 In contrast, there are only about 608 experimentally verified SM–BGC pairs in fungi.27,30–32 This 50-fold discrepancy between identified metabolites and linked BGCs arises largely because SM–BGC pairings are typically established on a case-by-case basis, since confirmation of their pairing requires experimental validation.16,33,34 Thus, the SMs biosynthesized by predicted BGCs in fungal genomes have not yet been discovered, and as such, most of these BGCs are considered “orphans”. Similarly, the biosynthetic pathways responsible for the vast majority of characterized fungal metabolites also remain uncharacterized, hindering efforts to study their biosynthesis.
000 fungal metabolites have been characterized,24 and genomic examinations suggest that there are likely millions of predicted BGCs in fungal genomes.25–29 In contrast, there are only about 608 experimentally verified SM–BGC pairs in fungi.27,30–32 This 50-fold discrepancy between identified metabolites and linked BGCs arises largely because SM–BGC pairings are typically established on a case-by-case basis, since confirmation of their pairing requires experimental validation.16,33,34 Thus, the SMs biosynthesized by predicted BGCs in fungal genomes have not yet been discovered, and as such, most of these BGCs are considered “orphans”. Similarly, the biosynthetic pathways responsible for the vast majority of characterized fungal metabolites also remain uncharacterized, hindering efforts to study their biosynthesis.
The very small number of SM–BGC pairs identified to date, coupled with the much larger numbers of fungal metabolites and predicted orphan BGCs in fungal genomes, underscores the need for methods and strategies to predict SM–BGC pairs. To bridge this gap between chemotype and genotype, several general and specific methodologies have been developed to associate SMs and their cognate BGCs.16,34–37 At the heart of these general approaches lies the independent identification of BGCs via predictions from the genome, and structural identification of SMs via metabolomics, followed by an algorithm predicting connections. Importantly, many of these algorithms take advantage of the MIBiG database,30,32 a community effort cataloguing BGCs and their SMs, which includes information on the gene/protein sequences of the BGC with their known or putative functions, the organism the SM–BGC pair was identified, and the resulting SM structures and bioactivities.
Strategies have sought to enhance SM–BGC prediction by integrating large metabolomics data. For example, correlation-based approaches statistically associate BGC or gene cluster family (GCF)–SM pairs based on co-occurrence patterns,36 while feature-based approaches rely on specific, searchable attributes (e.g., core enzymes, transcription factors or metabolomic spectral features like fragments and isotopes) to generate “forward” (BGC to SM) or “reverse” (SM to BGC) associations. These approaches have recently uncovered a novel class of BGCs, the isocyanide synthases,37 and linked peptide natural products (e.g., ribosomally synthesized and post-translationally modified peptides (RiPPs) or non-ribosomal peptide synthetases (NRPSs)) to their core genes.35,38,39 Stable isotope labelling has also been used to connect mass spectrometric features (i.e., mass to charge values coupled with chromatographic retention times for metabolites/SMs) to BGCs by tracing the biosynthesis from known BGC substrates.40
Here, we introduce a new strategy to link the chemical structures of experimentally identified SMs to their cognate BGCs via structural similarity to known SM–BGC pairs. We then applied this strategy to the metabolomes and genomes of 16 strains of the filamentous fungus Aspergillus fischeri and the known SM–BGC pairs in the MIBiG database. This enabled us to confidently assign more than one third of detected metabolites to known BGCs that are present in A. fischeri genomes, and generate testable SM–BGC hypotheses in a straightforward, fast and ab initio manner for all the remaining SMs. Our results suggest that coupling chemical structure-based similarity with genomic similarity is a powerful approach for linking detected SMs to their BGCs in fungal genomes.
Results
Leveraging chemical and genomic similarity to infer SM–BGC pairs
We developed an integrative approach based on chemical structural similarity to link SMs to BGCs (Fig. 1A and B). This approach evaluates structural similarity by matching machine-readable molecular fingerprints from candidate compounds to those stored in the MIBiG database, allowing for the inference of putative SM–BGC relationships. Leveraging the MIBiG database, which contains 3158 structures from 1896 bacterial and eukaryotic BGCs, including 692 SMs from 377 fungal BGCs, allows us to connect listed SMs and their BGC genes via their BGC accession IDs.30 Our study demonstrates that metabolomics data from fungal culture extracts can be used to improve the quality and accuracy of genome-based BGC predictions.Establishing chemical structure similarity
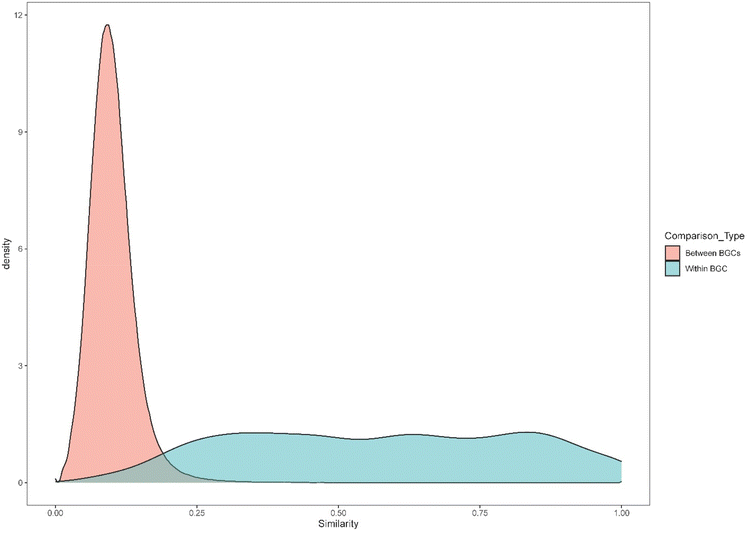
Structural similarity of small molecules can be assessed via digital fingerprints, i.e., a bit vector of each structure generated from SMILES (simplified molecular-input line-entry system, i.e. text abstractions of 2D or 3D structures of molecules).28,41–43 The similarity between a pair of fingerprints is then expressed using the Tanimoto (Jaccard) index, which is the ratio of the number of shared fingerprint bits (i.e., substructures) to the union of bits in a pairwise comparison. As proposed here, Tanimoto similarity is a heuristic for generating SM–BGC links. Similarity can be computed between any two given structures and we opted to provide users with the result(s) and leave it up to them to evaluate the quality of the match(es).The structure similarity linking approach that we employ assumes that SMs from the same BGC are much more similar (as expressed by Tanimoto pairwise similarity) than SMs from different BGCs. To validate this assumption, we calculated the pairwise structural similarity among all SMs in MIBiG (Fig. 2). We found that SMs from the same BGC are, on average, significantly more similar than SMs from different BGCs (average Tanimoto pairwise similarity for SMs from the same BGC = 0.568; for SMs from different BGCs = 0.101; permutation test with 1000 permutations gave no permuted statistic as extreme as the observed and a p value ≤0.001).
Experimental data: sixty structurally characterized metabolites from A. fischeri
We next applied our approach to a data set containing the metabolomes and genomes of 16 strains of Aspergillus fischeri, a filamentous fungus that is gaining attention as a close, non-pathogenic relative of the major human pathogen Aspergillus fumigatus.44–46 Using aspects of the “one strain many compounds” approach, the production of SM was evaluated at two temperatures (30 °C and 37 °C) using UPLC-MS/MS, recently.46,47 In doing so, the number of compounds detected per strain increased,46 as would have growing them on e.g., different media.44,48 Metabolites were identified based on either a direct match in LC-MS/MS to reference standards, all of which had been fully characterized by NMR, or to a class of fungal metabolites (i.e., via mass defect filtering22,23). A total of 60 metabolites were identified at two levels of confidence (Table 1, ‘A’ and ‘B’ respectively), subsequently referred to as ‘identified SMs’. Three biological replicates provided insight into the consistency of SM production by the various BGCs and strains (Fig. S1). Overall, we found the most biosynthetically rich strains across all replicates and temperatures yielded up to three times more SMs than the least-producing strains (e.g., CBS 150748: N = 45 vs. CBS 54465: N = 15). Interestingly, strains with the greatest consistency of SM production across all biological replicates produced fewer metabolites (e.g., 18/20 SMs were detected in all replicates of strain CBS 150750 at 30 °C (90%)) (Fig. S2).46| SM group # | SM | BGC link | BGC present | Confidence | Ref. |
|---|---|---|---|---|---|
| a MS/MS and NMR or MS/MS and dereplication with in-house database/standard. b MS/MS only. For each SM, the BGC(s) linked by structural similarity clustering are indicated, with the underlined BGCs yielding the highest Tanimoto similarity match. Hypothetical links that were confirmed post-hoc based on experimental data (e.g., identical SM structures, evidence from the literature) are denoted as ‘reported’, and all newly generated hypotheses without additional evidence are denoted as ‘predicted’. For SMs of known BGCs, all generated hypotheses were accurate. For an overview of all structurally similar metabolites from A. fischeri together with their top SM hits in MIBiG database, where available, see Fig. S5. | |||||
| 1 | Ilicicolin Eb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[9 with combining low line]](https://www.rsc.org/images/entities/char_0039_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[3 with combining low line]](https://www.rsc.org/images/entities/char_0033_0332.gif) , BGC0001924 (new BGC 1) , BGC0001924 (new BGC 1) |
No | Predicted | 51 |
| 2 | (3β,22E)-Ergosta-4,6,8(14),22-tetraene-3-ola | Primary metabolism | — | — | 49 |
| 3 | Fumagillolb | BGC0001067 | Yes | Reported | 52 |
| 4 | Brevianamide A/Bb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[8 with combining low line]](https://www.rsc.org/images/entities/char_0038_0332.gif) ![[4 with combining low line]](https://www.rsc.org/images/entities/char_0034_0332.gif) , BGC0000816 (new BGC 2) , BGC0000816 (new BGC 2) |
No | Predicted | |
| 4 | Brevianamide C/Db | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[8 with combining low line]](https://www.rsc.org/images/entities/char_0038_0332.gif) ![[4 with combining low line]](https://www.rsc.org/images/entities/char_0034_0332.gif) , BGC0000816 (new BGC 2) , BGC0000816 (new BGC 2) |
No | Predicted | |
| 5 | Brevianamide Qb | BGC0000442 (new BGC 3) | No | Predicted | |
| 5 | Brevianamide Rb | BGC0000442 (new BGC 3) | No | Predicted | |
| 5 | Brevianamide Tb | BGC0000442 (new BGC 3) | No | Predicted | |
| 5 | Brevianamide Ub | BGC0000442 (new BGC 3) | No | Predicted | |
| 5 | Brevianamide V/Wb | BGC0000356 (new BGC 3) | No | Predicted | |
| 5 | Brevianamide Kb | BGC0000442 (new BGC 3) | No | Predicted | |
| 6 | Cottoquinazoline Ea | BGC0000355 | Putative | Predicted | 53 and 54 |
| 6 | Cottoquinazoline Fa | BGC0000355 | Putative | Predicted | 53 and 54 |
| 6 | Cottoquinazoline Ga | BGC0000355 | Putative | Predicted | 53 and 54 |
| 7 | Fumitremorgin Fb | BGC0001142, ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[3 with combining low line]](https://www.rsc.org/images/entities/char_0033_0332.gif) ![[5 with combining low line]](https://www.rsc.org/images/entities/char_0035_0332.gif) ![[5 with combining low line]](https://www.rsc.org/images/entities/char_0035_0332.gif) (new BGC 4) (new BGC 4) |
No | Predicted | 55 |
| 7 | Fumitremorgin G/Lb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[4 with combining low line]](https://www.rsc.org/images/entities/char_0034_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) , BGC0000355 (new BGC 4) , BGC0000355 (new BGC 4) |
No | Predicted | 55 |
| 8 | 4-Hydroxyaszonaleninb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[9 with combining low line]](https://www.rsc.org/images/entities/char_0039_0332.gif) ![[3 with combining low line]](https://www.rsc.org/images/entities/char_0033_0332.gif) , (BGC0002272) , (BGC0002272) |
Yes | Predicted | 56 |
| 8 | Acetylaszonalenina | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[9 with combining low line]](https://www.rsc.org/images/entities/char_0039_0332.gif) ![[3 with combining low line]](https://www.rsc.org/images/entities/char_0033_0332.gif) , (BGC0002272) , (BGC0002272) |
Yes | Reported | 56 |
| 8 | Aszonalenina | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[9 with combining low line]](https://www.rsc.org/images/entities/char_0039_0332.gif) ![[3 with combining low line]](https://www.rsc.org/images/entities/char_0033_0332.gif) , (BGC0002272) , (BGC0002272) |
Yes | Reported | 56 and 57 |
| 9 | Isoroquefortine Cb | BGC0000420 | Yes | Reported | 58 and 59 |
| 9 | Roquefortine Cb | BGC0000420 | Yes | Reported | 58 and 59 |
| 10 | Brevianamide Eb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[7 with combining low line]](https://www.rsc.org/images/entities/char_0037_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) , BGC0002617 , BGC0002617 |
No | Predicted | |
| 11 | 13-O-Prenylfumitremorgin Ba | BGC0000356 | Yes | Predicted | 60 and 61 |
| 11 | Brevianamide Fb | BGC0000356 | Yes | Reported | 60 and 61 |
| 11 | Deoxybrevianamide Eb | BGC0000356 | Yes | Predicted | 60 and 61 |
| 11 | Fumitremorgin Aa | BGC0000356 | Yes | Reported | 62 |
| 11 | Fumitremorgin Ba | BGC0000356 | Yes | Reported | 60 and 61 |
| 11 | Fumitremorgin Ca | BGC0000356 | Yes | Reported | 60 and 61 |
| 11 | Spiro[5H,10H-dipyrrolo-[1,2-a:1′,2′-d]pyrazine-2-(3H),2′-[2H]-indole]-3′,5,10(1′H)trionea | BGC0000356 | Yes | Predicted | 63 |
| 11 | Tryprostatin Bb | BGC0000356 | Yes | Reported | 60 and 61 |
| 11 | Tryprostatin C/Db | BGC0000356 | Yes | Predicted | 60 and 61 |
| 11 | Verruculogenb | BGC0000356 | Yes | Reported | 60 and 61 |
| 12 | Hexadehydroastechrome (monomer)b | BGC0000372 | Yes | Reported | 64 |
| 12 | Tryhistatina | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[4 with combining low line]](https://www.rsc.org/images/entities/char_0034_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) , BGC0000372 , BGC0000372 |
Yes | Predicted | 64 |
| 13 | 16-O-Deacetyl helvolic acid 21,16-lactoneb | BGC0000686 | Yes | Predicted | 65 |
| 13 | Helvolic acida | BGC0000686 | Yes | Reported | 65 |
| 14 | Pyripyropene Fb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[9 with combining low line]](https://www.rsc.org/images/entities/char_0039_0332.gif) , BGC0001068 , BGC0001068 |
Yes | Predicted | 66 |
| 14 | Pyripyropene Hb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[9 with combining low line]](https://www.rsc.org/images/entities/char_0039_0332.gif) , BGC0001068 , BGC0001068 |
Yes | Predicted | 66 |
| 14 | Pyripyropene Ib | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[9 with combining low line]](https://www.rsc.org/images/entities/char_0039_0332.gif) , BGC0001068 , BGC0001068 |
Yes | Predicted | 66 |
| 14 | Pyripyropene Ob | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[9 with combining low line]](https://www.rsc.org/images/entities/char_0039_0332.gif) , BGC0001068 , BGC0001068 |
Yes | Predicted | 66 |
| 15 | Azonapyrone Aa | BGC0002604 | Yes | Predicted | 67 |
| 15 | Sartorypyrone Aa | BGC0002604 | Yes | Reported | 67 |
| 16 | Circumdatin Ca | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[3 with combining low line]](https://www.rsc.org/images/entities/char_0033_0332.gif) ![[5 with combining low line]](https://www.rsc.org/images/entities/char_0035_0332.gif) ![[5 with combining low line]](https://www.rsc.org/images/entities/char_0035_0332.gif) , BGC0001652, BGC0000448, BGC0000409, BGC0000303 (new BGC 5) , BGC0001652, BGC0000448, BGC0000409, BGC0000303 (new BGC 5) |
No | Predicted | |
| 16 | Dimetoxycircumdatin Ca | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[3 with combining low line]](https://www.rsc.org/images/entities/char_0033_0332.gif) ![[5 with combining low line]](https://www.rsc.org/images/entities/char_0035_0332.gif) ![[5 with combining low line]](https://www.rsc.org/images/entities/char_0035_0332.gif) , BGC0001652, BGC0000448, BGC0000409, BGC0000303 (new BGC 5) , BGC0001652, BGC0000448, BGC0000409, BGC0000303 (new BGC 5) |
No | Predicted | |
| 17 | Betaenone Eb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[6 with combining low line]](https://www.rsc.org/images/entities/char_0036_0332.gif) ![[5 with combining low line]](https://www.rsc.org/images/entities/char_0035_0332.gif) , BGC0001264 (new BGC 6) , BGC0001264 (new BGC 6) |
No | Predicted | 68 |
| 17 | Betaenone G/I/Jb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[6 with combining low line]](https://www.rsc.org/images/entities/char_0036_0332.gif) ![[5 with combining low line]](https://www.rsc.org/images/entities/char_0035_0332.gif) , BGC0001264 (new BGC 6) , BGC0001264 (new BGC 6) |
No | Predicted | 68 |
| 17 | Betaenone Hb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[6 with combining low line]](https://www.rsc.org/images/entities/char_0036_0332.gif) ![[5 with combining low line]](https://www.rsc.org/images/entities/char_0035_0332.gif) , BGC0001264 (new BGC 6) , BGC0001264 (new BGC 6) |
No | Predicted | 68 |
| 18 | Clavaric acidb | BGC0001248 | Yes | Reported | 69 and 70 |
| 19 | Chaetoglobosin 542b | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[5 with combining low line]](https://www.rsc.org/images/entities/char_0035_0332.gif) ![[3 with combining low line]](https://www.rsc.org/images/entities/char_0033_0332.gif) ![[9 with combining low line]](https://www.rsc.org/images/entities/char_0039_0332.gif) , BGC0000968, BGC0001182 , BGC0000968, BGC0001182 |
Yes | Predicted | 71 |
| 20 | Neosartoricinb | BGC0001144 | Yes | Reported | 72 and 73 |
| 20 | Neosartoricin Cb | BGC0001144 | Yes | Reported | 72 and 73 |
| 20 | Neosartoricin Db | BGC0001144 | Yes | Reported | 72 and 73 |
| 21 | Brevianamide Lb | ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[8 with combining low line]](https://www.rsc.org/images/entities/char_0038_0332.gif) , BGC0002242 (new BGC 7) , BGC0002242 (new BGC 7) |
No | Predicted | 74 |
| 21 | Brevianamide Ob | BGC0002208, ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[4 with combining low line]](https://www.rsc.org/images/entities/char_0034_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) (new BGC 7) (new BGC 7) |
No | Predicted | 74 |
| 21 | Brevianamide Pb | BGC0002208, ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[4 with combining low line]](https://www.rsc.org/images/entities/char_0034_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) (new BGC 7) (new BGC 7) |
No | Predicted | 74 |
| 22 | Secalonic acids (A/B/C/D/E/F/F1/G; 4,4′-secalonic acid E)b | BGC0002063, BGC0001886, BGC0001988 | Yes | Reported | 75 |
| 23 | Nidiascin Ca | BGC0002275, ![[B with combining low line]](https://www.rsc.org/images/entities/char_0042_0332.gif) ![[G with combining low line]](https://www.rsc.org/images/entities/char_0047_0332.gif) ![[C with combining low line]](https://www.rsc.org/images/entities/char_0043_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[0 with combining low line]](https://www.rsc.org/images/entities/char_0030_0332.gif) ![[2 with combining low line]](https://www.rsc.org/images/entities/char_0032_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) ![[7 with combining low line]](https://www.rsc.org/images/entities/char_0037_0332.gif) ![[1 with combining low line]](https://www.rsc.org/images/entities/char_0031_0332.gif) (new BGC 8) (new BGC 8) |
No | Predicted | |
| 24 | Neosartorina | BGC0001988 | Yes | Reported | 76 |
| 25 | Bisdethiobis(methylthio)-gliotoxina | BGC0000361 | Yes | Reported | 4 |
| 25 | Gliotoxina | BGC0000361 | Yes | Reported | 4 |
Predicting the BGCs linked to experimentally identified SMs
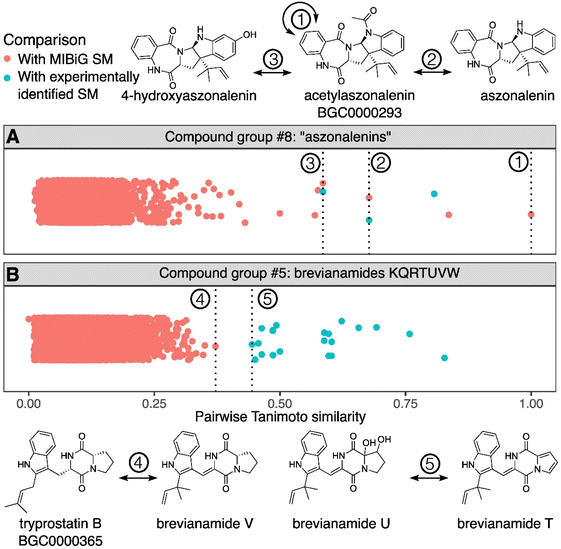
To generate hypotheses about the biosynthetic origin of SMs from A. fischeri, we calculated pairwise Tanimoto similarities for all 60 experimentally identified chemical structures from A. fischeri and all known SMs from the MIBiG database. We next used the all-versus-all structural similarity matrix to perform hierarchical clustering and generate groups of highly similar SMs (Table 1, Fig. 3; Fig. S3). The highest match between an identified SM and an SM (or a set of SMs) from MIBiG, which is already linked to a BGC, enabled us to assign the identified SM to that corresponding BGC; we refer to these assignments as hypothetical SM–BGC links (Table 1).Identified SMs were thus linked to putative BGCs via their highest structural similarity to SMs from MIBiG. In doing so, we generated BGC hypotheses for all 60 identified metabolites from A. fischeri. Of these, 22 A. fischeri metabolites were identical to SMs in the MIBiG database, i.e., representing known SM–BGC links, and 37 metabolites were structurally similar, but not identical, to SMs in MIBiG (Table S1 ‘confidence’ column: ‘reported’ and ‘predicted’, respectively). The sole remaining metabolite is a sterol, which was not linked to a BGC, as sterol biosynthesis is part of primary metabolism.49,50 Structural similarity between identified SMs and SMs in MIBiG varied substantially for the 37 metabolites examined (Fig. 3). For example, the experimentally identified SM acetylaszonalenin produces an exact match with the acetylaszonalenin SM present in the MIBiG database (Fig. 3A). Two additional experimentally identified SMs have a high similarity with acetylaszonalenin: aszonalenin and 4-hyrdroxyaszonalenin. Upon further investigation, the link of aszonalenin with the acetylaszonalenin BGC is confirmed by literature (but not recorded in MIBiG), while the link between 4-hyrdroxyaszonalenin and the acetylaszonalenin BGC remains a hypothetical connection not yet experimentally confirmed. In other cases, such the breviamides (Fig. 3B), the similarity score between experimentally identified SMs and MIBiG SMs is lower, which suggests that these metabolites may be biosynthesized by a BGC not currently present in the MIBiG database. All SM groups and hypotheses are described in Table 1, and are subsequently evaluated in depth.
Assigning BGCs to identical pairs of structures
There were 13 A. fischeri SMs that had an identical SM structure included in the MIBiG database (Table 1). While unsurprising and seemingly trivial, the ability of our approach to quickly assign BGCs for experimentally identified SMs also present in the MIBiG database offers considerable practical utility, since the natural products literature does not dictate a consistent nomenclature process for SMs, which makes lookups by name futile. Lack of well-catalogued data further complicates fast identification.43The 13 SMs identified from A. fischeri with an identical SM match in the MIBiG database are: acetylaszonalenin, brevianamide F, clavaric acid, fumagilol, fumitremorgin B and C, helvolic acid, hexadehydroastechrom, neosartorin, roquefortine C, sartorypyrone A, tryprostatin B, and verruculogen. Notably, three known SM–BGC pairs were missed by our structure similarity approach due to database limitations. These were bisdethiobis(methylthio)gliotoxin and gliotoxin, both produced by gliotoxin BGC0000361, which was retired in MIBiG v3.1, and secalonic acid(s) for which the SM(s) were not structurally identified in our study nor when describing the BGC (and hence neither in the corresponding MIBiG BGC0001886 entry).
Uncovering BGCs for SMs not present in the MIBiG database
Not all known biosynthetic intermediates, shunt products or possible SMs are deposited in the MIBiG database. Thus, six additional SM–BGC links were confirmed based on primary literature. These are aszonalenin in BGC0000293,56 fumitremorgin A in BGC0000356,62 isoroquefortine C in BGC0000420,58 and neosartoricin, neosartoricin C, and neosartoricin D in BGC0001144.72 Notably, isoroquefortine C is an artifact produced by the isomerization of roquefortine C caused by pH or light.58 Similarly, neosartoricin C and D might be artifacts related to the production of neosartoricin B.72 Indeed, artifacts – compounds that were isolated but whose structure slightly differs from the true SM, possibly due to extraction solvents or sample handling – are a well-known challenge in the natural products literature.77 Finally, fumitremorgin A is technically not considered a product of the verruculogen BGC (BGC0000356), as the gene encoding the FtmPT3 protein responsible for converting verruculogen to fumitremorgin A is not part of the BGC.62 However, this variation in the degree of biosynthetic gene clustering is not unusual.78 Given that fumitremorgin A is produced from verruculogen, an SM of this BGC, it is reasonable to include it in the set of SMs attributed to BGC0000356. In summary, our approach directly assigned BGCs for 22 of the 60 experimentally identified SMs (36%).The remaining 38 A. fischeri identified metabolites did not have identical matches to SM structures included in the MIBiG database or biosynthetic information in the literature. Thus, we augmented the SM–BGC hypotheses for each of these metabolites based on structural similarity by examining whether A. fischeri genomes contained the SM-linked BGCs (for details on BGC prediction/detection, see Methods and the next section). Our predictions can be broadly grouped into three level-of-confidence categories: (i) attributing the metabolite to a known BGC that is present in the respective A. fischeri strain genome(s) (e.g., 4-hydroxyaszonalenin – BGC0000293, Fig. 3A), (ii) linking the SM to a BGC not present in the respective A. fischeri genome(s), and (iii) ascribing the SM (or SM group) as a novel metabolite(s) likely encoded by an unknown BGC (i.e., no similar SMs are present in the MIBiG database, Fig. 3B). Given the dearth of fungal BGCs in MIBiG (i.e., only 377), we were pleased that our approach predicted 13 SMs in category (i), 11 SMs in category (ii), and 13 SMs in category (iii) (see extended Table S1 for more details, and Table S2 for all Tanimoto similarities). Notably, we found that the BGCs associated with 38 of the 59 predicted SM–BGC links (64%) are in the curated list of A. fischeri BGCs (Table S1). For the remaining “undetected” BGCs, hypotheses to explain this pattern are consistent with the presence of a homologous but divergent/convergent BGC or with genome incompleteness. The latter possibility is less likely because the estimated genome completeness is very high.46
Genomic characterization of A. fischeri BGCs
To evaluate the hypotheses generated for the 38 remaining metabolites without known BGCs, we next examined the BGC content of the A. fischeri genomes. We first analyzed the 16 genomes using antiSMASH v7, which predicts ‘BGC regions’ – i.e., continuous stretches in the genome containing BGC(s) and other genes (Fig. 4). For traceability, we also identified and grouped homologous BGCs across the individual genomes. Across all 16 genomes, antiSMASH predicted 44 BGC regions that corresponded to 42 unique BGCs (BGC0001248 and BGC0002710 were each detected in two regions of the genomes), as well as 20 candidate BGC regions (‘unnamed’ or ‘orphan’ putative BGCs). The mean number of BGCs per strain was 53.3 (range 51–56), a number consistent with previous reports.79 Note that we refer to these predicted BGCs by the accession numbers of their reference BGCs in the MIBiG database.antiSMASH-predicted BGCs were classified as ‘present’ in A. fischeri when they contained all the genes present in the reference MIBiG entry, or when they were incomplete but supported by evidence from structurally identified SMs. Additionally, BGCs were classified as ‘putative’ in A. fischeri when they were incomplete with at least half of the genes detected but without evidence from the metabolomics study. Otherwise, BGCs were classified as ‘absent’ (i.e., fewer than half of the genes were found and no evidence from metabolomics was present). Examining each of the 44 antiSMASH-predicted BGC regions across A. fischeri genomes, we classified 20 BGCs as ‘present’, 9 as ‘putative’, and 15 as ‘absent’ (Table S3). We also specifically searched for the protein sequences of each MIBiG BGC in the RNAseq-based gene annotations,46 allowing us to manually curate and revise the antiSMASH predictions. These additional analyses enabled us to classify 7 additional BGCs as ‘present’ and 4 BGCs as ‘putative’. A full list of BGCs is given in Table S3, with information on sequence identity with known BGC genes and genomic location in Table S4. Subsequently, we detail issues and difficulties in faithfully assessing the number and identity of BCGs present in genomes.
Among all 40 BGCs classified as ‘present’ or ‘putative’, five artifacts reduce the total BGC count. These primarily stem from the current cataloging approach for BGCs and the scientific community's limited understanding of them. MIBiG defines each BGC in the genome it is reported in, sometimes listing the ‘same’ (i.e., homologous) BGC multiple times from different organisms. Similarly, one BGC that biosynthesizes one SM can be nested within another, larger BGC that biosynthesizes a different SM. These situations can lead to ‘collisions’, i.e. the assignment of the same set of genes to multiple BGCs.
There were collisions in two pairs of BGCs where the same set of proteins in A. fischeri is classified as two different BGCs due to similarity of the MIBiG reference sequences (BGC0000361 gliotoxin/BGC0001609 gliovirin, and BGC0001144 neosartoricin B/BGC0002646 hancockinone A), reducing the number of unique BGCs by two. The BGC for biotin is listed twice in the MIBiG database (BGC0001238 and BGC0001239) but was counted only once, as it matches the same set of genes. Similarly, there are two slightly different BGCs matching a congruent set of genes for the metabolite ilicicolin H (BGC0002035 and BGC0002093), which were counted as one BGC, further decreasing the total count by two. Additionally, the BGC for clavaric acid (BGC0001248), which is composed of a single gene, was found twice (Table S4). However, only one of the two homologs identified (homolog ID 221721_1) contains the sequence motif VSDCISE, which was previously found in Fusarium graminearum to be involved in clavaric acid production.70
In total, we infer that A. fischeri contains 35 ‘present’ and ‘putative’ BGCs (Fig. 4 and Table S3). Overall, BGC content was largely conserved and consistent across the 16 strains, with most BGCs (82%; 29/35 total ‘present’ and ‘putative’ BGCs) detected in all strains.
Caveats for using antiSMASH as tool for accurate BGC surveys
We chose the comparison and validation of antiSMASH, since it is a widely used (and very useful) tool for BGC prediction in fungal genomes.34 antiSMASH is designed to discover regions containing known or novel BGCs.15 In practice, the tool is frequently also used to discover BGCs (rather than regions containing BGCs) in fungal genomes, with the results being taken at face value without further scrutiny. While examining the correspondence between A. fischeri BGCs identified by antiSMASH and their inferred references in the MIBiG database, we noted five sources of error associated with the common practice of conflating the BGC regions identified by antiSMASH with individual BGCs.First, in most known BGCs, the locus predicted by antiSMASH to contain a BGC was much larger (up to approximately three times the number of genes) than the actual BGC. This is by design, as BGC boundaries are difficult to define (formerly possible with CASSIS11) and hence the more relaxed/inclusive ‘region’ concept in antiSMASH (Fig. 4 and 5A). Second, as BGCs are known to co-localize, particularly in telomeric or low complexity regions of genomes,80,81 their physical proximity on chromosomes, in combination with this ‘regions’ concept, can lead to BGCs masking each other (Fig. 5B). This masking occurred in the proximal BGCs BGC0001403 for trypacidin and BGC0001988 for neosartorin, and with BGC0000356 for verruculogen and BGC0001067 for fumagillin. Examination of 16 strains of A. fischeri revealed some instances where the same homologous genes were predicted as part of different BGCs in different strains (Fig. 5C). Third, at low identities, the BGC predictions from the module ‘–cb-knownclusters’ may be misleading/arbitrary as we found several instances of the same orthologous region being labeled as different BGCs. A fourth source of inaccuracies stems from version differences of the MIBiG database used for the BGC prediction. Curation processes continually expand the knowledge base,32 but sometimes, valid BGCs are removed or lacking, thus leading to missed predictions (e.g., the extensively studied gliotoxin BGC, which was retired, i.e. removed in MIBiG v3.1/v4.0) (Fig. 4). Finally, we noted instances of BGCs missed by antiSMASH (but detected by protein sequence searches) for reasons that are not apparent (Fig. 4).
Discussion
There are at least 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 reported fungal metabolites24,82–84 and millions of BGCs predicted in fungal genomes26–28 but only a few hundred SM–BGC pairs,32 suggesting that linking SMs and BGCs remains challenging. To address this challenge, we developed an SM–BGC linking approach based on chemical similarity, that requires a minimum of input data (e.g., a single SM) and can be performed using experimental data or data retrieved from natural product databases.82–85 Across 16 strains of a single fungal species, our approach recovered 22 known SM–BGC pairs and generated hypotheses for 37 more, including 11 that could be SMs attributed to BGCs present in MIBiG. Thus, our approach efficiently automated SM-driven linking of SM and BGCs, and faithfully recovered known connections, additional links not included in MIBiG, and new hypotheses. This approach offers two advantages: (1) it can provide orthogonal BGC validation (in case of known links i.e., Tanimoto similarity = 1), and (2) it can generate hypotheses for SMs whose biosynthetic pathways are not known (i.e., Tanimoto similarity <1).
000 reported fungal metabolites24,82–84 and millions of BGCs predicted in fungal genomes26–28 but only a few hundred SM–BGC pairs,32 suggesting that linking SMs and BGCs remains challenging. To address this challenge, we developed an SM–BGC linking approach based on chemical similarity, that requires a minimum of input data (e.g., a single SM) and can be performed using experimental data or data retrieved from natural product databases.82–85 Across 16 strains of a single fungal species, our approach recovered 22 known SM–BGC pairs and generated hypotheses for 37 more, including 11 that could be SMs attributed to BGCs present in MIBiG. Thus, our approach efficiently automated SM-driven linking of SM and BGCs, and faithfully recovered known connections, additional links not included in MIBiG, and new hypotheses. This approach offers two advantages: (1) it can provide orthogonal BGC validation (in case of known links i.e., Tanimoto similarity = 1), and (2) it can generate hypotheses for SMs whose biosynthetic pathways are not known (i.e., Tanimoto similarity <1).
Method development in BGC detection from genomic data has produced many tools (e.g., SMURF,86 antiSMASH,15 BiG-SCAPE,87 cblaster,88 DeepBGC,13 BGCFlow,89 CLOCI,12 zol and fai14). Additionally, SM–BGC links can be established via correlation analyses, an approach termed ‘metabologenomics’, or specific experiments such as ‘IsoAnalyst’.40 Metabologenomics can yield de novo SM–BGC links, but requires large datasets (>100 species) from extensive experimental data as well as sophisticated fine-tuning of scoring functions and parameters, dependent on BGC class (NP Linker).36,90 These genome- or BGC-driven innovations stand in contrast to the number of integrative tools for linking SMs (SANDPUMA,91 GNP,92 PARAS93), which typically are limited to specific taxa or classes of SMs. Furthermore, only two tools are available that can link SMs to BGCs: RIPPminer94 and Prism,95,96 which again are limited to specific classes of SMs or taxa. The method outlined here fills a gap, where the strategy of connecting SMs to BGC is agnostic to chemical structural class, organism, data size or specificity.
As a consequence of the aforementioned challenges in SM–BGC linking, existing strategies for straightforward orthogonal validation of BGCs are lacking. SM–BGC links are typically validated via gene knock-out studies (e.g., ref. 65, 97 and 98). In contrast, in silico tools linking SMs to BGCs deliver unvalidated predictions or connections. The wealth of BGC prediction tools with various strategies, focused on specific BGC classes or more general tools10 poses a challenge because presence/absence or identity of a predicted BGC are frequently a function of arbitrary cutoffs (Fig. 5). BGCs can be interpreted with some fluidity, e.g., many genes in described BGCs are of unknown function and may not be essential to the BGC, synteny conservation is sometimes low, and with increasing phylogenetic/evolutionary distance, gene and protein sequences naturally diverge. Using metabolomics as orthogonal validation can be a means to avoid arbitrary thresholds confirming the presence of a BGC with the unambiguous presence of its SM product(s).
In our structural similarity analyses, we refrained from setting a similarity threshold, due to the known patchiness of SMs present in the MIBiG database (i.e., only 692 fungal SMs out of >30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 reported in the literature), general limitations in SM–BGC pairing knowledge, and previously documented challenges with threshold-based approaches.36 Moreover, some SMs (particularly biosynthetic intermediates) are not necessarily unique to any single BGC. For example, the diketopiperazine brevianamide F (cyclo-L-Trp-L-Pro) is the first product of the biosynthesis of verruculogen by BGC000356 in A. fumigatus60 as well as of the biosynthesis of notoamide A by BGC0000818 in Aspergillus versicolor99 and brevianamide A by the bvn gene cluster (currently not listed in MIBiG) in Penicillium brevicompactum NRRL 864.100
000 reported in the literature), general limitations in SM–BGC pairing knowledge, and previously documented challenges with threshold-based approaches.36 Moreover, some SMs (particularly biosynthetic intermediates) are not necessarily unique to any single BGC. For example, the diketopiperazine brevianamide F (cyclo-L-Trp-L-Pro) is the first product of the biosynthesis of verruculogen by BGC000356 in A. fumigatus60 as well as of the biosynthesis of notoamide A by BGC0000818 in Aspergillus versicolor99 and brevianamide A by the bvn gene cluster (currently not listed in MIBiG) in Penicillium brevicompactum NRRL 864.100
Our chemical structure similarity approach also has caveats. Our results are based on the examination of strains of an Aspergillus species, one of the most well studied fungal genera in terms of prior knowledge of SM–BGC pairs. Studies of less-studied organisms may be more challenging, especially if their chemodiversity differs from the SMs currently represented in the MIBiG database, resulting in hypotheses (SM–BGC links and SM groups) that may be a poor fit. As we have shown, some published SM–BGC pairs are not currently included in the MIBiG database. Yet, as databases grow, so does the utility of this approach. This methodology could further be expanded to work on partial structures or m/z fingerprints in the same manner as using SMILES as input. Additionally, chemical conversions that alter the backbone or skeleton of a SM sufficiently could mask a better clustering fit. Furthermore, when SMs are produced by multiple non-homologous BGCs (e.g., brevianamide F), genomic evidence is necessary to determine which BGC it is produced by. Such instances of convergently evolved SMs would only be detected in this strategy when finding the SM and not the BGC (but this inference would be based on the absence of evidence). Putative examples of this in our data are chaetoglobosin 542 and ilicicolin E. Chaetoglobosin 542 is structurally very similar to chaetoglobosin A produced by BGC0000968,101 which is similar to two different A. fischeri BGCs. Interestingly, the presence-absence patterns of the BGC and the SM match only for one of the BGCs. In the second case, ilicicolin E differs from ascochlorin of BGC000192351 only by the presence of an α,β-unsaturated ketone instead of the aliphatic ketone, respectively, in the 6-membered ring. However, A. fischeri genomes do not contain any related BGCs, suggesting that the observed structural similarity of the two SMs may result from convergent evolution. Of course, another possibility is that the BGC is present in the genome but not part of the genome assembly (e.g., because it resides in an otherwise highly repetitive region).
These caveats notwithstanding, our approach successfully inferred SM–BGC pairs for nearly one third of the fungal metabolites identified and predicted SM–BGC pair hypotheses for nearly all the rest. Ultimately, our approach is a hypothesis-generating strategy and must be validated experimentally (e.g., by modifying putative BGCs in the native host or through heterologous expression of the putative BGC).33,102 The approach applied in this work leveraged similarity among known SM structures and BGCs to bidirectionally link SMs and BGCs via the MIBiG database and thereby successfully generated testable biosynthetic hypotheses in a high-throughput fashion and validated the presence of predicted BGCs. This increases the fidelity of the biological conclusions drawn based on the BGCs and their implications for the chemotype (i.e., SM profile), lifestyle, and niche of an organism. While our approach is hypothesis-generating and requires further validation, it can augment the fidelity of stand-alone tools that operate solely either on metabolomic or genomic data.
Methods
All genomic and metabolomic data were taken from Rinker et al.46 and are available via FigShare (https://doi.org/10.6084/m9.figshare.25316452).Chemical fingerprinting and clustering
Structures (SMILES, simplified molecular-input line-entry system; a text string representing the molecule) for all SMs identified from untargeted metabolomics were collected via ChemDraw v23.1.1 (Revvity) and combined with structures from known BGCs deposited in the MIBiG database.32 Chemoinformatic analyses were carried out in Jupyter notebook103 using RDKit and PubChemPy.104 To facilitate the subsequent search for the detected metabolites, we prefixed the names of structures from MIBiG with the BGC accession ID, and those of metabolites found in extracts with ‘chem_’. For comparing structural similarity and clustering the metabolites, we calculated the Morgan fingerprint for each metabolite with GetMorganFingerprintAsBitVect() using chirality with a radius of 2, and 2048 bits, and converted the fingerprints to binary strings using ToBitString(). We calculated Tanimoto similarity (Jaccard index, the intersection of set bitflags divided by the union) between all pairwise comparisons using calculate_tanimoto() resulting in a symmetric similarity matrix of all-vs.-all comparisons. With linkage(method = ‘average’, metric = ‘euclidean’) and dendrogram() from scipy, we performed hierarchical clustering of the metabolites based on the distance matrix and used matplotlib to plot and save the resulting figure (Fig. S3). SM groups were initially delineated by searching the dendrogram for the tag “chem_” and grouping similar structures. Subsequently, for every identified SM, the highest pairwise similarity scores with a SM (or a set of SMs) in the MIBiG database was extracted from the similarity matrix, thereby generating biosynthetic hypotheses for each SM. To test validity, we performed bootstrap resampling (1000 replicates) yielding a 95% confidence interval of [0.475, 0.498] for the median difference, confirming the robustness of the result.BGC predictions
BGCs were predicted using antiSMASH v7.1.015 and DIAMOND v2.1.6.160 blastp searches105 of the MIBiG database v3.1.30 All subsequent analyses were performed in R v4.4.0.106 Conventionally, BGCs are defined in a specific genome. However, in this manuscript, we refer to the predicted candidate BGCs by their MIBiG accession number for convenience.After the antiSMASH prediction (–fullhmmer –rre –cc-MIBiG –cb-knownclusters –cb-subclusters –cb-general, using the corresponding gff3 annotation file), we aggregated results from individual runs into a single file. Across the different strains, known and unknown BGCs were aggregated by comparing gene content. This approach yielded meaningful clusters as evidenced by the correct grouping of known BGCs (with their MIBiG BGC accession ID). This clustering revealed instances in which the same genes were attributed to different BGCs (both known and “anonymous” candidate clusters) by antiSMASH.
For the amino acid sequence search, the 16 genomes were queried with all sequences in MIBiG v3.1 (diamond blastp -f6 qseqid sseqid pident length mismatch gapopen qstart qend sstart send evalue bitscore qcovhsp qlen slen full_sseq) and the results concatenated.
To validate the antiSMASH BGC predictions, the genes in each predicted region were searched using DIAMOND blastp. Additionally, the hits were filtered for high identity (pident >80%, minimum 50% query coverage), as well as for runs of hits against the same BGC in proximity (low identity clustering of putative, diverged BGCs). By using DIAMOND blastp to confirm A. fischeri BGC genes based on known BGCs, we tagged every BGC gene with a BGC ID from MIBiG hence allowing for interoperability of biological and chemical data.
BGCs were classified as present if all genes were found in proximity, regardless of whether a corresponding SM was detected, or if they were recovered partially, i.e. incomplete but with evidence from SMs. BGCs were classified as putative if more than half of the genes were present but there was no evidence for their presence based on metabolomics. BGCs were classified as absent if fewer than half of the genes were found and no evidence from metabolomics was present.
Additional data (https://doi.org/10.6084/m9.figshare.28380443) and analysis code (https://doi.org/10.6084/m9.figshare.28380431) for this study can be found on FigShare.
Conflicts of interest
AR is a scientific consultant for LifeMine Therapeutics, Inc. NHO has ownership interests in Ionic Pharmaceuticals, LLC and is a member of the Scientific Advisory Board of Mycosynthetix, Inc. HAR, TNG, and N.H.O. are members of the Scientific Advisory Board of Clue Genetics, Inc. KS is a data scientist at Olink part of Thermo Fisher Scientific.Data availability
Analysis code is deposited together with supplementary information (SI) in Figshare (https://doi.org/10.6084/m9.figshare.28380431). The genomic and metabolomic data for the 16 A. fischeri strains used in this study were published by Rinker et al.43 and can be accessed in GenBank via BioProject accession number PRJNA1129834, and in the corresponding Figshare repository (https://doi.org/10.6084/m9.figshare.25316452).Supplementary information is available. See DOI: https://doi.org/10.1039/d5ob01965f.
Acknowledgements
KS was supported by a PostDoc stipend of the Swedish Pharmaceutical Society. Research in the AR lab is supported by the National Science Foundation (DEB-2110404) and the National Institutes of Health/National Institute of Allergy and Infectious Diseases (R01 AI153356).OLR was supported by the National Science Foundation Graduate Research Fellowship Program under Grant No. 2444112. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation. GHG thanks the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and Fundação Coordenação de Aperfeiçoamento do Pessoal do Ensino Superior (CAPES) grant number 405934/2022-0 (The National Institute of Science and Technology INCT Funvir), and CNPq 301058/2019-9 from Brazil.
References
- N. P. Keller, Fungal secondary metabolism: regulation, function and drug discovery, Nat. Rev. Microbiol., 2019, 17(3), 167–180 CrossRef CAS.
- P. Wiemann, B. E. Lechner, J. A. Baccile, T. A. Velk, W. B. Yin and J. W. Bok, et al., Perturbations in small molecule synthesis uncovers an iron-responsive secondary metabolite network in Aspergillus fumigatus, Front. Microbiol., 2014, 5 Search PubMed . Available from: https://journal.frontiersin.org/article/10.3389/fmicb.2014.00530/abstract.
- A. Fleming, On the Antibacterial Action of Cultures of a Penicillium, with Special Reference to their Use in the Isolation of B. influenzæ, Br. J. Exp. Pathol., 1929, 10(3), 226–236 CAS.
- S. K. Dolan, G. O'Keeffe, G. W. Jones and S. Doyle, Resistance is not futile: gliotoxin biosynthesis, functionality and utility, Trends Microbiol., 2015, 23(7), 419–428 CrossRef CAS PubMed.
- A. Tomas, Purification of a Cultivar-Specific Toxin from Pyrenophora tritici-repentis, Causal Agent of Tan Spot of Wheat, Mol. Plant-Microbe Interact., 1990, 3(4), 221 CrossRef CAS.
- T. L. Friesen, E. H. Stukenbrock, Z. Liu, S. Meinhardt, H. Ling and J. D. Faris, et al., Emergence of a new disease as a result of interspecific virulence gene transfer, Nat. Genet., 2006, 38(8), 953–956 CrossRef CAS PubMed.
- D. J. Newman and G. M. Cragg, Natural Products as Sources of New Drugs over the Nearly Four Decades from 01/1981 to 09/2019, J. Nat. Prod., 2020, 83(3), 770–803 CrossRef CAS PubMed.
- the International Natural Product Sciences Taskforce, A. G. Atanasov, S. B. Zotchev, V. M. Dirsch and C. T. Supuran, Natural products in drug discovery: advances and opportunities, Nat. Rev. Drug Discovery, 2021, 20(3), 200–216 CrossRef PubMed.
- A. G. T. Niego, C. Lambert, P. Mortimer, N. Thongklang, S. Rapior and M. Grosse, et al., The contribution of fungi to the global economy, Fungal Diversity, 2023, 121(1), 95–137 CrossRef CAS.
- T. Weber and H. U. Kim, The secondary metabolite bioinformatics portal: Computational tools to facilitate synthetic biology of secondary metabolite production, Synth. Syst. Biotechnol., 2016, 1(2), 69–79 CrossRef PubMed.
- T. Wolf, V. Shelest, N. Nath and E. Shelest, CASSIS and SMIPS: promoter-based prediction of secondary metabolite gene clusters in eukaryotic genomes, Bioinformatics, 2016, 32(8), 1138–1143 CrossRef CAS PubMed.
- Z. Konkel, L. Kubatko and J. C. Slot, CLOCI: unveiling cryptic fungal gene clusters with generalized detection, Nucleic Acids Res., 2024, gkae625 Search PubMed.
- G. D. Hannigan, D. Prihoda, A. Palicka, J. Soukup, O. Klempir and L. Rampula, et al., A deep learning genome-mining strategy for biosynthetic gene cluster prediction, Nucleic Acids Res., 2019, 47(18), e110–e110 CrossRef CAS PubMed.
- R. Salamzade, P. Q. Tran, C. Martin, A. L. Manson, M. S. Gilmore and A. M. Earl, et al., zol and fai: large-scale targeted detection and evolutionary investigation of gene clusters, Nucleic Acids Res., 2025, 53(3), gkaf045 CrossRef PubMed.
- K. Blin, S. Shaw, H. E. Augustijn, Z. L. Reitz, F. Biermann and M. Alanjary, et al., antiSMASH 7.0: new and improved predictions for detection, regulation, chemical structures and visualisation, Nucleic Acids Res., 2023, 51(W1), W46–W50 CrossRef CAS PubMed.
- J. J. J. Van Der Hooft, H. Mohimani, A. Bauermeister, P. C. Dorrestein, K. R. Duncan and M. H. Medema, Linking genomics and metabolomics to chart specialized metabolic diversity, Chem. Soc. Rev., 2020, 49(11), 3297–3314 Search PubMed.
- N. H. Oberlies, S. L. Knowles, C. S. M. Amrine, D. Kao, V. Kertesz and H. A. Raja, Droplet probe: coupling chromatography to the in situ evaluation of the chemistry of nature, Nat. Prod. Rep., 2019, 36(7), 944–959 RSC.
- S. A. Jarmusch, J. J. J. Van Der Hooft, P. C. Dorrestein and A. K. Jarmusch, Advancements in capturing and mining mass spectrometry data are transforming natural products research, Nat. Prod. Rep., 2021, 38(11), 2066–2082 Search PubMed.
- Y. Dong and A. Aharoni, Image to insight: exploring natural products through mass spectrometry imaging, Nat. Prod. Rep., 2022, 39(7), 1510–1530 Search PubMed.
- S. Alseekh, A. Aharoni, Y. Brotman, K. Contrepois, J. D'Auria and J. Ewald, et al., Mass spectrometry-based metabolomics: a guide for annotation, quantification and best reporting practices, Nat. Methods, 2021, 18(7), 747–756 CrossRef CAS PubMed.
- L. W. Sumner, A. Amberg, D. Barrett, M. H. Beale, R. Beger and C. A. Daykin, et al., Proposed minimum reporting standards for chemical analysis: Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI), Metabolomics, 2007, 3(3), 211–221 CrossRef CAS PubMed.
- T. El-Elimat, M. Figueroa, B. M. Ehrmann, N. B. Cech, C. J. Pearce and N. H. Oberlies, High-Resolution MS, MS/MS, and UV Database of Fungal Secondary Metabolites as a Dereplication Protocol for Bioactive Natural Products, J. Nat. Prod., 2013, 76(9), 1709–1716 CrossRef CAS.
- N. D. Paguigan, T. El-Elimat, D. Kao, H. A. Raja, C. J. Pearce and N. H. Oberlies, Enhanced dereplication of fungal cultures via use of mass defect filtering, J. Antibiot., 2017, 70(5), 553–561 CrossRef CAS PubMed.
- J. Bérdy, Thoughts and facts about antibiotics: Where we are now and where we are heading, J. Antibiot., 2012, 65(8), 385–395 CrossRef PubMed.
- G. C. A. Amos, T. Awakawa, R. N. Tuttle, A. C. Letzel, M. C. Kim and Y. Kudo, et al., Comparative transcriptomics as a guide to natural product discovery and biosynthetic gene cluster functionality, Proc. Natl. Acad. Sci. U. S. A., 2017, 114(52), E11121–E11130 CrossRef CAS PubMed.
- S. A. Kautsar, K. Blin, S. Shaw, T. Weber and M. H. Medema, BiG-FAM: the biosynthetic gene cluster families database, Nucleic Acids Res., 2021, 49(D1), D490–D497 CrossRef CAS PubMed.
- K. Palaniappan, I. M. A. Chen, K. Chu, A. Ratner, R. Seshadri and N. C. Kyrpides, et al., IMG-ABC v.5.0: an update to the IMG/Atlas of Biosynthetic Gene Clusters Knowledgebase, Nucleic Acids Res., 2019, gkz932 CrossRef PubMed.
- S. Zhang, G. Shi, X. Xu, X. Guo, S. Li and Z. Li, et al., Global Analysis of Natural Products Biosynthetic Diversity Encoded in Fungal Genomes, J. Fungi, 2024, 10(9), 653 CrossRef CAS PubMed.
- O. L. Riedling and A. Rokas, mGem: How many fungal secondary metabolites are produced by filamentous fungi? Conservatively, at least 1.4 million. Rodrigues M, editor, mBio, 2025, 16(10), e01381–e01325 Search PubMed.
- B. R. Terlouw, K. Blin, J. C. Navarro-Muñoz, N. E. Avalon, M. G. Chevrette and S. Egbert, et al., MIBiG 3.0: a community-driven effort to annotate experimentally validated biosynthetic gene clusters, Nucleic Acids Res., 2023, 51(D1), D603–D610 CrossRef CAS.
- O. Riedling, A. S. Walker and A. Rokas, Predicting fungal secondary metabolite activity from biosynthetic gene cluster data using machine learning, Microbiol. Spectrum, 2024, 12(2), e03400–e03423 Search PubMed.
- M. M. Zdouc, K. Blin, N. L. L. Louwen, J. Navarro, C. Loureiro and C. D. Bader, et al., MIBiG 4.0: advancing biosynthetic gene cluster curation through global collaboration, Nucleic Acids Res., 2024, gkae1115 Search PubMed.
- I. Kjærbølling, U. H. Mortensen, T. Vesth and M. R. Andersen, Strategies to establish the link between biosynthetic gene clusters and secondary metabolites, Fungal Genet. Biol., 2019, 130, 107–121 CrossRef.
- H. W. Lv, J. G. Tang, B. Wei, M. D. Zhu, H. W. Zhang and Z. B. Zhou, et al., Bioinformatics assisted construction of the link between biosynthetic gene clusters and secondary metabolites in fungi, Biotechnol. Adv., 2025, 81, 108547 CrossRef CAS PubMed.
- C. A. Dejong, G. M. Chen, H. Li, C. W. Johnston, M. R. Edwards and P. N. Rees, et al., Polyketide and nonribosomal peptide retro-biosynthesis and global gene cluster matching, Nat. Chem. Biol., 2016, 12(12), 1007–1014 CrossRef CAS PubMed.
- L. K. Caesar, F. A. Butun, M. T. Robey, N. J. Ayon, R. Gupta and D. Dainko, et al., Correlative metabologenomics of 110 fungi reveals metabolite–gene cluster pairs, Nat. Chem. Biol., 2023, 19(7), 846–854 CrossRef CAS PubMed.
- G. R. Nickles, B. Oestereicher, N. P. Keller and M. T. Drott, Mining for a new class of fungal natural products: the evolution, diversity, and distribution of isocyanide synthase biosynthetic gene clusters, Nucleic Acids Res., 2023, 51(14), 7220–7235 CrossRef CAS PubMed.
- R. D. Kersten, Y. L. Yang, Y. Xu, P. Cimermancic, S. J. Nam and W. Fenical, et al., A mass spectrometry–guided genome mining approach for natural product peptidogenomics, Nat. Chem. Biol., 2011, 7(11), 794–802 CrossRef CAS.
- B. Behsaz, E. Bode, A. Gurevich, Y. N. Shi, F. Grundmann and D. Acharya, et al., Integrating genomics and metabolomics for scalable non-ribosomal peptide discovery, Nat. Commun., 2021, 12(1), 3225 Search PubMed.
- C. S. McCaughey, J. A. Van Santen, J. J. J. Van Der Hooft, M. H. Medema and R. G. Linington, An isotopic labeling approach linking natural products with biosynthetic gene clusters, Nat. Chem. Biol., 2022, 18(3), 295–304 Search PubMed.
- T. M. Voser, M. D. Campbell and A. R. Carroll, How different are marine microbial natural products compared to their terrestrial counterparts?, Nat. Prod. Rep., 2022, 39(1), 7–19 RSC.
- H. L. Morgan, The Generation of a Unique Machine Description for Chemical Structures-A Technique Developed at Chemical Abstracts Service, J. Chem. Doc., 1965, 5(2), 107–113 CrossRef CAS.
- K. Steffen, N. H. Oberlies and A. Rokas, Machine-Readable Structural Information Is Essential for Natural Products Research, J. Nat. Prod., 2025, 88(11), 2815–2821 CrossRef CAS PubMed.
- M. E. Mead, S. L. Knowles, H. A. Raja, S. R. Beattie, C. H. Kowalski and J. L. Steenwyk, et al., Characterizing the Pathogenic, Genomic, and Chemical Traits of Aspergillus fischeri, a Close Relative of the Major Human Fungal Pathogen Aspergillus fumigatus, mSphere, 2019, 4(1), e00018–e00019 CrossRef CAS PubMed.
- A. Rokas, Evolution of the human pathogenic lifestyle in fungi, Nat. Microbiol., 2022, 7(5), 607–619 CrossRef CAS PubMed.
- D. C. Rinker, T. J. C. Sauters, K. Steffen, A. Gumilang, H. A. Raja and M. Rangel-Grimaldo, et al., Strain heterogeneity in a non-pathogenic Aspergillus fungus highlights factors associated with virulence, Commun. Biol., 2024, 7(1), 1082 CrossRef CAS PubMed.
- H. B. Bode, B. Bethe, R. Höfs and A. Zeeck, Big Effects from Small Changes: Possible Ways to Explore Nature's Chemical Diversity, ChemBioChem, 2002, 3(7), 619 Search PubMed.
- K. M. VanderMolen, H. A. Raja, T. El-Elimat and N. H. Oberlies, Evaluation of culture media for the production of secondary metabolites in a natural products screening program, AMB Express, 2013, 3(1), 71 CrossRef PubMed.
- E. Desmond and S. Gribaldo, Phylogenomics of Sterol Synthesis: Insights into the Origin, Evolution, and Diversity of a Key Eukaryotic Feature, Genome Biol. Evol., 2009, 1, 364–381 Search PubMed.
- S. Dhingra and R. A. Cramer, Regulation of Sterol Biosynthesis in the Human Fungal Pathogen Aspergillus fumigatus: Opportunities for Therapeutic Development, Front. Microbiol., 2017, 8 Search PubMed . Available from: https://journal.frontiersin.org/article/10.3389/fmicb.2017.00092/full.
- Y. Araki, T. Awakawa, M. Matsuzaki, R. Cho, Y. Matsuda and S. Hoshino, et al., Complete biosynthetic pathways of ascofuranone and ascochlorin in Acremonium egyptiacum, Proc. Natl. Acad. Sci. U. S. A., 2019, 116(17), 8269–8274 Search PubMed.
- H. C. Lin, Y. H. Chooi, S. Dhingra, W. Xu, A. M. Calvo and Y. Tang, The Fumagillin Biosynthetic Gene Cluster in Aspergillus fumigatus Encodes a Cryptic Terpene Cyclase Involved in the Formation of β-trans-Bergamotene, J. Am. Chem. Soc., 2013, 135(12), 4616–4619 CrossRef CAS PubMed.
- K. A. O'Hanlon, L. Gallagher, M. Schrettl, C. Jöchl, K. Kavanagh and T. O. Larsen, et al., Nonribosomal Peptide Synthetase Genes pesL and pes1 Are Essential for Fumigaclavine C Production in Aspergillus fumigatus, Appl. Environ. Microbiol., 2012, 78(9), 3166–3176 CrossRef.
- B. D. Ames, S. W. Haynes, X. Gao, B. S. Evans, N. L. Kelleher and Y. Tang, et al., Complexity Generation in Fungal Peptidyl Alkaloid Biosynthesis: Oxidation of Fumiquinazoline A to the Heptacyclic Hemiaminal Fumiquinazoline C by the Flavoenzyme Af12070 from Aspergillus fumigatus, Biochemistry, 2011, 50(40), 8756–8769 CrossRef CAS PubMed.
- X. Gao, Y. H. Chooi, B. D. Ames, P. Wang, C. T. Walsh and Y. Tang, Fungal Indole Alkaloid Biosynthesis: Genetic and Biochemical Investigation of the Tryptoquialanine Pathway in Penicillium aethiopicum, J. Am. Chem. Soc., 2011, 133(8), 2729–2741 CrossRef CAS PubMed.
- W. B. Yin, A. Grundmann, J. Cheng and S. M. Li, Acetylaszonalenin Biosynthesis in Neosartorya fischeri, J. Biol. Chem., 2009, 284(1), 100–109 CrossRef CAS.
- D. Wakana, T. Hosoe, T. Itabashi, K. Nozawa, K. i. Kawai and K. Okada, et al., Isolation of Isoterrein from Neosartorya fischeri, Mycotoxins, 2006, 56(1), 3–6 CrossRef CAS.
- N. Shangguan, W. J. Hehre, W. S. Ohlinger, M. P. Beavers and M. M. Joullié, The Total Synthesis of Roquefortine C and a Rationale for the Thermodynamic Stability of Isoroquefortine C over Roquefortine C, J. Am. Chem. Soc., 2008, 130(19), 6281–6287 Search PubMed.
- C. García-Estrada, R. V. Ullán, S. M. Albillos, M.Á Fernández-Bodega, P. Durek and H. von Döhren, et al., A Single Cluster of Coregulated Genes Encodes the Biosynthesis of the Mycotoxins Roquefortine C and Meleagrin in Penicillium chrysogenum, Chem. Biol., 2011, 18(11), 1499–1512 Search PubMed.
- S. Maiya, A. Grundmann, S. Li and G. Turner, The Fumitremorgin Gene Cluster of Aspergillus fumigatus: Identification of a Gene Encoding Brevianamide F Synthetase, ChemBioChem, 2006, 7(7), 1062–1069 CrossRef CAS PubMed.
- A. Grundmann, T. Kuznetsova, S. Afiyatullov and S. Li, FtmPT2, an N –Prenyltransferase from Aspergillus fumigatus, Catalyses the Last Step in the Biosynthesis of Fumitremorgin B, ChemBioChem, 2008, 9(13), 2059–2063 CrossRef CAS PubMed.
- K. Mundt, B. Wollinsky, H. Ruan, T. Zhu and S. Li, Identification of the Verruculogen Prenyltransferase FtmPT3 by a Combination of Chemical, Bioinformatic and Biochemical Approaches, ChemBioChem, 2012, 13(17), 2583–2592 CrossRef CAS PubMed.
- Y. Tsunematsu, N. Ishikawa, D. Wakana, Y. Goda, H. Noguchi and H. Moriya, et al., Distinct mechanisms for spiro-carbon formation reveal biosynthetic pathway crosstalk, Nat. Chem. Biol., 2013, 9(12), 818–825 CrossRef CAS PubMed.
- W. B. Yin, J. A. Baccile, J. W. Bok, Y. Chen, N. P. Keller and F. C. Schroeder, A Nonribosomal Peptide Synthetase-Derived Iron(III) Complex from the Pathogenic Fungus Aspergillus fumigatus, J. Am. Chem. Soc., 2013, 135(6), 2064–2067 CrossRef CAS PubMed.
- J. M. Lv, D. Hu, H. Gao, T. Kushiro, T. Awakawa and G. D. Chen, et al., Biosynthesis of helvolic acid and identification of an unusual C-4-demethylation process distinct from sterol biosynthesis, Nat. Commun., 2017, 8(1), 1644 CrossRef.
- T. Itoh, K. Tokunaga, Y. Matsuda, I. Fujii, I. Abe and Y. Ebizuka, et al., Reconstitution of a fungal meroterpenoid biosynthesis reveals the involvement of a novel family of terpene cyclases, Nat. Chem., 2010, 2(10), 858–864 CrossRef CAS PubMed.
- W. G. Wang, L. Q. Du, S. L. Sheng, A. Li, Y. P. Li and G. G. Cheng, et al., Genome mining for fungal polyketide-diterpenoid hybrids: discovery of key terpene cyclases and multifunctional P450s for structural diversification, Org. Chem. Front., 2019, 6(5), 571–578 RSC.
- H. Li, J. Hu, H. Wei, P. S. Solomon, K. A. Stubbs and Y. Chooi, Biosynthesis of a Tricyclo[6.2.2.02,7]dodecane System by a Berberine Bridge Enzyme–Like Aldolase, Chem. – Eur. J., 2019, 25(66), 15062–15066 CrossRef CAS PubMed.
- R. P. Godio, R. Fouces and J. F. Martín, A Squalene Epoxidase Is Involved in Biosynthesis of Both the Antitumor Compound Clavaric Acid and Sterols in the Basidiomycete H. sublateritium, Chem. Biol., 2007, 14(12), 1334–1346 Search PubMed.
- R. P. Godio and J. F. Martín, Modified oxidosqualene cyclases in the formation of bioactive secondary metabolites: Biosynthesis of the antitumor clavaric acid, Fungal Genet. Biol., 2009, 46(3), 232–242 Search PubMed.
- B. Perlatti, C. B. Nichols, N. Lan, P. Wiemann, C. J. B. Harvey and J. A. Alspaugh, et al., Identification of the Antifungal Metabolite Chaetoglobosin P From Discosia rubi Using a Cryptococcus neoformans Inhibition Assay: Insights Into Mode of Action and Biosynthesis, Front. Microbiol., 2020, 11, 1766 Search PubMed.
- W. B. Yin, Y. H. Chooi, A. R. Smith, R. A. Cacho, Y. Hu and T. C. White, et al., Discovery of Cryptic Polyketide Metabolites from Dermatophytes Using Heterologous Expression in Aspergillus nidulans, ACS Synth. Biol., 2013, 2(11), 629–634 Search PubMed.
- Y. H. Chooi, J. Fang, H. Liu, S. G. Filler, P. Wang and Y. Tang, Genome Mining of a Prenylated and Immunosuppressive Polyketide from Pathogenic Fungi, Org. Lett., 2013, 15(4), 780–783 CrossRef CAS PubMed.
- L. Zheng, H. Wang, L. Ludwig-Radtke and S. M. Li, Oxepin Formation in Fungi Implies Specific and Stereoselective Ring Expansion, Org. Lett., 2021, 23(6), 2024–2028 Search PubMed.
- L. Neubauer, J. Dopstadt, H. U. Humpf and P. Tudzynski, Identification and characterization of the ergochrome gene cluster in the plant pathogenic fungus Claviceps purpurea, Fungal Biol. Biotechnol., 2016, 3(1), 2 Search PubMed.
- Y. Matsuda, C. H. Gotfredsen and T. O. Larsen, Genetic Characterization of Neosartorin Biosynthesis Provides Insight into Heterodimeric Natural Product Generation, Org. Lett., 2018, 20(22), 7197–7200 Search PubMed.
- R. J. Capon, Extracting value: mechanistic insights into the formation of natural product artifacts – case studies in marine natural products, Nat. Prod. Rep., 2020, 37(1), 55–79 Search PubMed.
- A. Rokas, J. H. Wisecaver and A. L. Lind, The birth, evolution and death of metabolic gene clusters in fungi, Nat. Rev. Microbiol., 2018, 16(12), 731–744 Search PubMed.
- J. L. Steenwyk, M. E. Mead, S. L. Knowles, H. A. Raja, C. D. Roberts and O. Bader, et al., Variation Among Biosynthetic Gene Clusters, Secondary Metabolite Profiles, and Cards of Virulence Across Aspergillus Species, Genetics, 2020, 216(2), 481–497 Search PubMed.
- N. P. Keller, G. Turner and J. W. Bennett, Fungal secondary metabolism—from biochemistry to genomics, Nat. Rev. Microbiol., 2005, 3(12), 937–947 CrossRef CAS PubMed.
- X. Zhang, I. Leahy, J. Collemare and M. F. Seidl, Secondary metabolite biosynthetic gene clusters and their genomic localization in the fungal genus Aspergillus, 2024. Available from: https://biorxiv.org/lookup/doi/10.1101/2024.02.20.581327 Search PubMed.
- J. A. Van Santen, G. Jacob, A. L. Singh, V. Aniebok, M. J. Balunas and D. Bunsko, et al., The Natural Products Atlas: An Open Access Knowledge Base for Microbial Natural Products Discovery, ACS Cent. Sci., 2019, 5(11), 1824–1833 Search PubMed.
- A. Rutz, M. Sorokina, J. Galgonek, D. Mietchen, E. Willighagen and A. Gaudry, et al., The LOTUS initiative for open knowledge management in natural products research, eLife, 2022, 11, e70780 Search PubMed.
- Chemnetbase Dictionary of Natural Products 33.2. 2024. Available from: https://dnp.chemnetbase.com/chemical/ChemicalSearch.xhtml?dswid=918.
- V. Chandrasekhar, K. Rajan, S. R. S. Kanakam, N. Sharma, V. Weißenborn and J. Schaub, et al., COCONUT 2.0: a comprehensive overhaul and curation of the collection of open natural products database, Nucleic Acids Res., 2025, 53(D1), D634–D643 Search PubMed.
- N. Khaldi, F. T. Seifuddin, G. Turner, D. Haft, W. C. Nierman and K. H. Wolfe, et al., SMURF: Genomic mapping of fungal secondary metabolite clusters, Fungal Genet. Biol., 2010, 47(9), 736–741 Search PubMed.
- J. C. Navarro-Muñoz, N. Selem-Mojica, M. W. Mullowney, S. A. Kautsar, J. H. Tryon and E. I. Parkinson, et al., A computational framework to explore large-scale biosynthetic diversity, Nat. Chem. Biol., 2020, 16(1), 60–68 Search PubMed.
- C. L. M. Gilchrist, T. J. Booth, B. Van Wersch, L. Van Grieken, M. H. Medema and Y. H. Chooi, cblaster: a remote search tool for rapid identification and visualization of homologous gene clusters, Bioinform. Adv., 2021, 1(1), vbab016 Search PubMed.
- M. Nuhamunada, O. S. Mohite, P. V. Phaneuf, B. O. Palsson and T. Weber, BGCFlow: systematic pangenome workflow for the analysis of biosynthetic gene clusters across large genomic datasets, Nucleic Acids Res., 2024, 52(10), 5478–5495 Search PubMed.
- G. Hjörleifsson Eldjárn, A. Ramsay, J. J. J. Van Der Hooft, K. R. Duncan, S. Soldatou and J. Rousu, et al., Ranking microbial metabolomic and genomic links in the NPLinker framework using complementary scoring functions, PLoS Comput. Biol., 2021, 17(5), e1008920 Search PubMed.
- M. G. Chevrette, F. Aicheler, O. Kohlbacher, C. R. Currie and M. H. Medema, SANDPUMA: ensemble predictions of nonribosomal peptide chemistry reveal biosynthetic diversity across Actinobacteria, Bioinformatics, 2017, 33(20), 3202–3210 Search PubMed.
- C. W. Johnston, M. A. Skinnider, M. A. Wyatt, X. Li, M. R. M. Ranieri and L. Yang, et al., An automated Genomes-to-Natural Products platform (GNP) for the discovery of modular natural products, Nat. Commun., 2015, 6(1), 8421 Search PubMed.
- B. R. Terlouw, C. Huang, D. Meijer, J. D. D. Cediel-Becerra, R. He and M. L. Rothe, et al., PARAS: high-accuracy machine-learning of substrate specificities in nonribosomal peptide synthetases, Bioinformatics, 2025 Search PubMed . Available from: https://biorxiv.org/lookup/doi/10.1101/2025.01.08.631717.
- P. Agrawal, S. Khater, M. Gupta, N. Sain and D. Mohanty, RiPPMiner: a bioinformatics resource for deciphering chemical structures of RiPPs based on prediction of cleavage and cross-links, Nucleic Acids Res., 2017, 45(W1), W80–W88 Search PubMed.
- M. A. Skinnider, N. J. Merwin, C. W. Johnston and N. A. Magarvey, PRISM 3: expanded prediction of natural product chemical structures from microbial genomes, Nucleic Acids Res., 2017, 45(W1), W49–W54 Search PubMed.
- N. R. Spencer, M. Gunabalasingam, K. Dial, X. Di, T. Malcolm and N. A. Magarvey, An integrated AI knowledge graph framework of bacterial enzymology and metabolism, Proc. Natl. Acad. Sci. U. S. A., 2025, 122(15), e2425048122 Search PubMed.
- K. Ma, P. Zhang, Q. Tao, N. P. Keller, Y. Yang and W. B. Yin, et al., Characterization and Biosynthesis of a Rare Fungal Hopane-Type Triterpenoid Glycoside Involved in the Antistress Property of Aspergillus fumigatus, Org. Lett., 2019, 21(9), 3252–3256 Search PubMed.
- S. C. Heard, G. Wu and J. M. Winter, Discovery and characterization of a cytochalasan biosynthetic cluster from the marine-derived fungus Aspergillus flavipes CNL-338, J. Antibiot., 2020, 73(11), 803–807 CrossRef CAS PubMed.
- S. Li, K. Srinivasan, H. Tran, F. Yu, J. M. Finefield and J. D. Sunderhaus, et al., Comparative analysis of the biosynthetic systems for fungal bicyclo[2.2.2]diazaoctane indole alkaloids: the (+)/(−)-notoamide, paraherquamide and malbrancheamide pathways, MedChemComm, 2012, 3(8), 987 RSC.
- Y. Ye, L. Du, X. Zhang, S. A. Newmister, M. McCauley and J. V. Alegre-Requena, et al., Fungal-derived brevianamide assembly by a stereoselective semipinacolase, Nat. Catal., 2020, 3(6), 497–506 CrossRef CAS PubMed.
- J. Schümann and C. Hertweck, Molecular basis of cytochalasan biosynthesis in fungi: gene cluster analysis and evidence for the involvement of a PKS-NRPS hybrid synthase by RNA silencing, J. Am. Chem. Soc., 2007, 129(31), 9564–9565 CrossRef PubMed.
- L. K. Caesar, M. T. Robey, M. Swyers, M. N. Islam, R. Ye and P. P. Vagadia, et al., Heterologous Expression of the Unusual Terreazepine Biosynthetic Gene Cluster Reveals a Promising Approach for Identifying New Chemical Scaffolds, mBio, 2020, 11(4), e01691–e01620 CrossRef CAS PubMed.
- T. Kluyver, B. Ragan-Kelley, F. Pérez, B. Granger, M. Bussonnier and J. Frederic, et al., Jupyter Notebooks – a publishing format for reproducible computational workflows, in Positioning and Power in Academic Publishing: Players, Agents and Agendas, 2016, pp. 87–90 Search PubMed.
- G. Landrum, P. Tosco, B. Kelley, R. Rodriguez, D. Cosgrove and R. Vianello, et al., rdkit/rdkit: 2024_09_3 (Q3 2024) Release, Zenodo, 2024. Available from: https://zenodo.org/doi/10.5281/zenodo.591637 Search PubMed.
- B. Buchfink, C. Xie and D. H. Huson, Fast and sensitive protein alignment using DIAMOND, Nat. Methods, 2015, 12(1), 59–60 CrossRef CAS PubMed.
- R Core Team, R: A Language and Environment for Statistical Computing, R Foundation for Statistical Computing, 2024. Available from: https://www.R-project.org/ Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |