Open Access Article
Open Access ArticleEnantioselective synthesis of antibiotic inducing gamma-butenolides from Streptomyces rochei and derivatives
Haylie E. Hennigana,
Manuela Frias-Gomeza,
Kylie G. Castatora,
Lauren E. Wilbanksa,
Caroline Zua,
Gracie Sandersa and
Elizabeth I. Parkinson *ab
*ab
aJames Tarpo Jr. and Margaret Tarpo Department of Chemistry, Purdue University, West Lafayette, IN, USA. E-mail: eparkins@purdue.edu
bBorch Department of Medicinal Chemistry and Molecular Pharmacology, Purdue University, West Lafayette, IN, USA
First published on 11th February 2026
Abstract
Given the challenges of discovering new natural products, further investigations into the regulatory systems that control their production are needed. Herein, we describe an enantioselective, divergent synthesis of two known autoregulators from Streptmyces rochei, the Streptomyces rochei butenolides (SRBs), which regulate production of the antibiotics lankacidin and lankamycin. The synthesis hinges on two key reactions: enantioselective alkynylation and regioselective photo-oxidation. The synthetic route also enabled access to nine derivatives, allowing for structure activity relationship studies. Specifically, we developed a quantitative green-fluorescence protein (GFP) based reporter assay and used this assay to assess key structural features for natural product induction in Streptomyces rochei.
Introduction
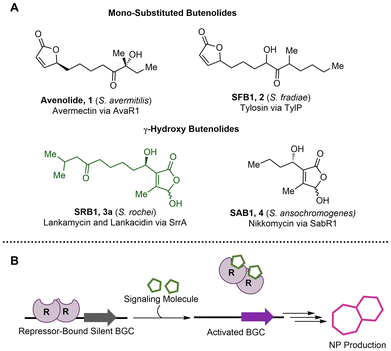
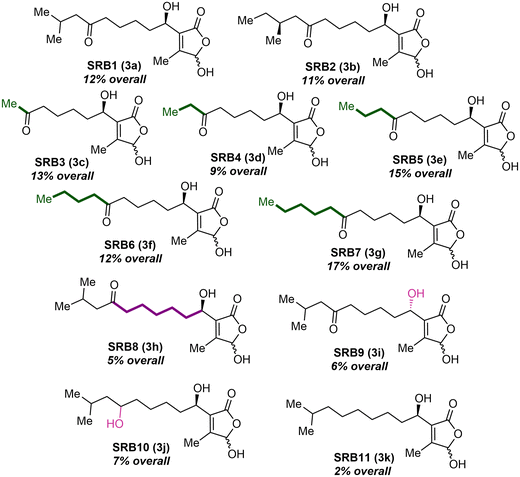
γ-Butenolides are commonly used by Streptomyces as signaling molecules in natural product regulation (Fig. 1A).1 These molecules induce natural product production via binding to cluster situated regulators, such as TetR-type repressors.2 This binding results in conformational changes to the repressors that inhibit DNA binding and ultimately enable generation of increased amounts of natural products (Fig. 1B). Avenolide (1) was the first γ-butenolide signaling molecule discovered, regulating production of the anthelmintic avermectin via derepression of AvaR1.3,4 Since then, other mono-substituted butenolides, such as the avenolide derivatives from Streptomyces albus and the Streptomyces fradiae butenolide (SFB1, 2), have been discovered to regulate avermectin and the macrolide antibiotic, tylosin, production, respectively.5,6 Tri-substituted γ-hydroxy butenolides have also been found to be signaling molecules but are generally less well studied compared to avenolide. The Streptomyces rochei butenolides (SRB1,2, 3) were isolated as a mixture with titers of 1.6 μg L−1 by the Arakawa group in 2012.7 These ligands regulate production of the antibiotics lankacidin and lankamycin via binding and subsequent release of the repressor SrrA.8 The only other structurally similar ligands that have been isolated are the Streptomyces ansochromogenes butenolides (SAB1–3, 4), which regulate production of the antifungal nikkomycins through binding to the repressor SabR1.9 Because of the low in vivo production of the tri-substituted γ-hydroxy butenolides, it is challenging to isolate sufficient quantities of these molecules to study their effects on repressor homologs. Access to a library of structurally diverse SRB type ligands through organic synthesis would provide enhanced insight into their role as signaling molecules for natural product regulation. Described herein is an enantioselective, divergent synthesis of two natural SRBs and derivatives, as well as development of a green fluorescent protein (GFP) reporter assay, which allowed for determination of the structure–activity relationship between the SrrA repressor and SRB derivatives.Divergent and enantioselective route to access the natural SRB1 and SRB2
To confirm the structure and determine the configuration of SRB1 and SRB2 the Arakawa group synthesized these compounds through a vinyl lithium addition between known butenolide 7 and an aldehyde sidechain corresponding to SRB1 or SRB2 (Scheme 1A).10,11 Global deprotection with boron tribromide afforded the two C1′ epimers of SRB1 and SRB2 in 5% and 8% overall yield, respectively. Comparison of the optical rotation data with the isolated mixture confirmed the configuration at C1′ to be R. While this synthesis was essential to confirming the structure of the SRB ligands, the low diastereoselectivity has a significant effect on overall yield of these compounds. Additionally, the acidity at C4 of butenolide 7 typically results in vinylogous addition when the menthyl ether is absent. We envisioned these compounds could be accessed more efficiently via a regioselective photo-oxidation of a chiral disubstituted furyl alcohol (8) which could be obtained through an enantioselective alkynylation with known 4-methylfuran-3-carbaldehyde (9, Scheme 1B).12,12 This route would allow for easy diversification to access a library of unique SRB type molecules.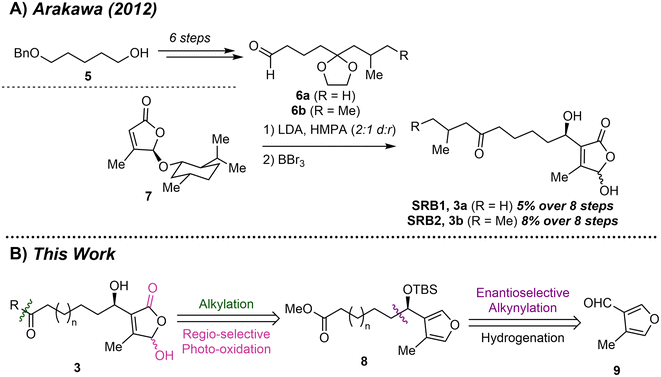 | ||
| Scheme 1 (A) Synthetic route to access SRB1 and SRB2 by the Arakawa group. (B) Retrosynthesis of the SRBs. | ||
Results and discussion
Model system of alkynylation
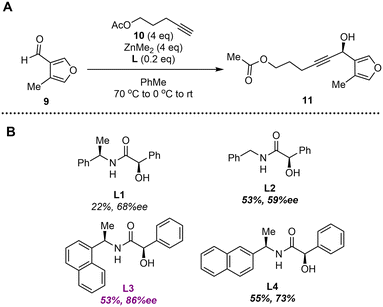
The alkynylation was initially tested on the more encumbered 4-methylfuran-3-carbaldehyde (9) which is easily accessed in 6 steps from methacrolein, and an excess of acetyl protected pent-4-yn-1-ol (10).11 Using the previously reported mandelamide ligand, L1, the reaction proceeded in 22% yield and 68% ee (Fig. 2A and B).12 Removing the methyl group alpha to the nitrogen of the amide (L2) decreased the enantioselectivity. Increasing the bulk of the amino side of the ligand through substituting the phenyl ring with a naphthyl group (L3, L4) increased the enantioselectivities with the best selectivity being observed with a ligand bearing substitution at the 1 position of the naphthyl rings, L3, affording 86% ee. Due to the increased enantioselectivity, this ligand was implemented in the synthesis of the SRB compounds.Synthesis of SRB1 and SRB2
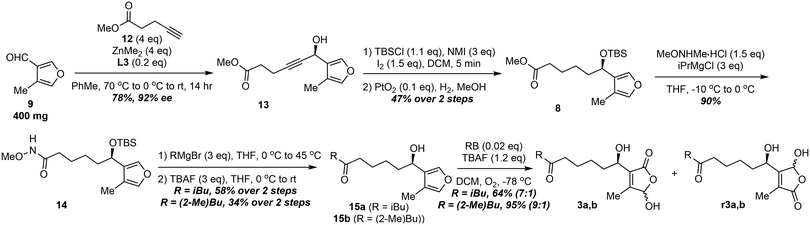
Initially, a convergent approach to the SRBs was employed via synthesizing the complete alkyne side chain and then subjecting to an enantioselective alkynylation (Fig. S4). Unfortunately, the overall yields were low (2% over 11 steps) due to the difficulty in accessing the alkyne partner; therefore, a divergent approach to these molecules was employed. The synthesis of SRB1 and SRB2 commenced with an enantioselective alkynylation between 4-methylfuran-3-carbaldehyde (9) and 5-pentynoic acid methyl ester (12) using the optimized mandelamide ligand, L3 to afford the propargly alcohol (13) in 78% yield and 92% ee (Scheme 2). The resulting alcohol was subjected to silyl protection with catalytic iodine and N-methylimidazole, followed by alkyne hydrogenation with platinum dioxide. The resulting methyl ester (8) was converted to the Weinreb amide (14) using isopropylmagnesium chloride. Alkylation of the amide with either isobutylmagnesium bromide or (S)-2-methylbutylmagnesium bromide allowed for diversification to the precursors for SRB1 (15a) and SRB2 (15b). Deprotection of the silyl ether afforded the photo-oxidation precursors, which were subjected to furan photo-oxidation conditions reported by Liu and coworkers.13 Photo-oxidation using singlet oxygen generated from Rose Bengal in the presence of the base, tetrabutylammonium fluoride, afforded primarily the desired regioisomers of the SRBs (7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 and 9
1 and 9![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixtures of SRB1 (3a)/SRB2 (3b): the regioisomer of SRB1 (r3A)/SRB2 (r3b), respectively) in 12% (SRB1) and 7% (SRB2) over 7 steps and allows for facile diversification. Unfortunately, the regioisomers were inseparable using standard silica gel chromatography. SRB1 (3a) and rSRB1 (regioisomer of SRB1, r3a) were separated using preparative scale high-performance liquid chromatography (HPLC, Fig. S17–19).
1 mixtures of SRB1 (3a)/SRB2 (3b): the regioisomer of SRB1 (r3A)/SRB2 (r3b), respectively) in 12% (SRB1) and 7% (SRB2) over 7 steps and allows for facile diversification. Unfortunately, the regioisomers were inseparable using standard silica gel chromatography. SRB1 (3a) and rSRB1 (regioisomer of SRB1, r3a) were separated using preparative scale high-performance liquid chromatography (HPLC, Fig. S17–19).
Proposed ligands for SAR with the receptor SrrA
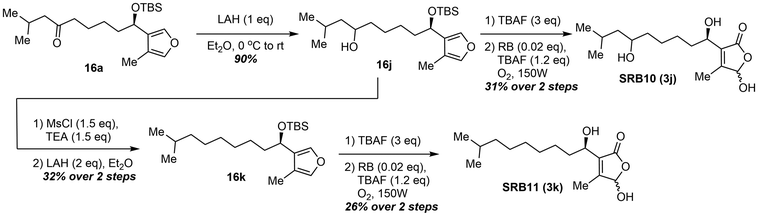
With a diversifiable and enantioselective route developed, SRB derivatives were synthesized to assess structural attributes necessary for derepressing SrrA, with a particular focus on chain length and branching (Fig. 2). Other known Streptomyces signaling molecules have similar scaffolds but vary in chain length and branching at C2.3,14,15 Because the two natural SRBs are branched at the tail of the C6′ ketone, a series of linear derivatives ranging from methyl to pentyl (SRB3–7) were explored. These compounds were synthesized by alkylating the Weinreb amide intermediate (14) with the corresponding Grignard reagent. Additionally, the distance between the hydroxyl group and the ketone was assessed (SRB8). Previously studied avenolide-type derivatives contain different oxidation patterns on the sidechain, resulting in different activities.5 The one carbon elongated SRB8 (3h), which was synthesized using 5-hexyne methyl ester, was synthesized to determine the importance in the location of the ketone. Three additional ligands were designed to probe the importance of the oxidation state on the sidechain, SRB9–11. SRB9 is the opposite epimer at C1′ of SRB1 and was synthesized using the opposite enantiomer of the ligand. Previous studies have shown that C1′ epimeric γ-butyrolactone signaling molecules can have different activities.16 The last two derivatives were proposed to determine the importance of the C6′ ketone. Specifically, SRB10 replaces the ketone at C6′ with a hydroxyl while SRB11 is completely deoxygenated at C6′. While deoxygenation at C6′ has previously been shown to greatly reduce activity,11 substitution of a hydroxyl for the ketone is currently unexplored. These compounds were synthesized by the reduction of intermediate 16a in route to SRB1 (Scheme 3). With a library of substrates in hand, their activity against the cognate receptor, SrrA, was tested.Development of a GFP based reporter assay to assess ligand binding
Previously, the Arakawa group tested a subset of these compounds, specifically the two natural SRB1,2, the enantiomer SRB9, the C6′ deoxo SRB11, and the C1′ ketone for their biological activities. Initial reports tested the induction of lankacidin and lankamycin using SRB1,2,9, and the C1′ ketone via induction assays.10 While the two natural compounds showed induction, SRB9 showed a 10-fold decrease in induction, whereas the C1′ ketone showed no induction. In a study of the SRB biosynthesis, they obtained SRB11 and tested its ability to bind SrrA using a gel-shift assay and observed a 100-fold decrease in activity.11 While both studies provide insight into the activity of these compounds, they lack a quantitative measurement of molecule derepression. Because of this, we chose to assess the activity of the ligand library using a green fluorescent protein (GFP) based reporter assay using the repressor, SrrA, similar to assays we have previously developed for other TetR type repressors (Fig. 3).17–19 First, biological evaluation of the purified SRB1 and rSRB1 was performed and compared to the mix (7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, desired
1, desired![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) undesired) (Fig. S7). As expected, the pure undesired regioisomer rSRB1 had little-to-no activity at up to the highest concentration texted (125 µM). The pure desired regioisomer (pSRB1) was the most active with a dissociation constant (KD) of 5.6 µM. The mixed SRB1 regioisomers showed very similar activity (KD = 5.7 µM) to that of pSRB1. Given the difficulty of the HPLC purification and the very similar activities of the mix to the pure regioisomer, all other SRBs and derivatives were tested as the mixtures. We note that all of the mixtures were at least 5
undesired) (Fig. S7). As expected, the pure undesired regioisomer rSRB1 had little-to-no activity at up to the highest concentration texted (125 µM). The pure desired regioisomer (pSRB1) was the most active with a dissociation constant (KD) of 5.6 µM. The mixed SRB1 regioisomers showed very similar activity (KD = 5.7 µM) to that of pSRB1. Given the difficulty of the HPLC purification and the very similar activities of the mix to the pure regioisomer, all other SRBs and derivatives were tested as the mixtures. We note that all of the mixtures were at least 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 desired
1 desired![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) undesired regioisomer with an average ratio of 7
undesired regioisomer with an average ratio of 7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.
1.
As expected, both the natural branched ligands (SRB1 and SRB2) had relatively low KDs. However, they also have low Emax values, indicating that their structure confers a high affinity for SrrA, but not necessarily maximum GFP expression. This differs from the linear derivatives of SRB1–2, SRB5 and SRB6, which exhibit comparable KDs but significantly higher Emax values. All linear ligands (SRB4–8), apart from SRB3, have Emax values higher than SRB1–2, indicating that branching plays an important role in natural product regulation. Previously, we have reported that shorter acyl chain lengths can have a dramatic effect on repressor binding, with the shortest derivatives exhibiting low repressor affinity and low GFP maximum (Fig. 4).17,19
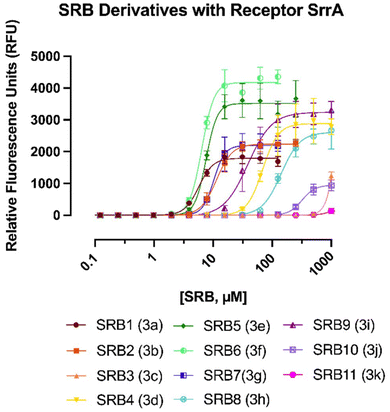 | ||
| Fig. 4 Results of the GFP reporter assay with SRB1-11 and the repressor SrrA. Each point is an average of three biological replicates analyzed on different days. | ||
Unsurprisingly, the shorter ligands SRB3 and SRB4 have higher KDs. This is likely because shorter chains cannot make important contacts with amino acids in the ligand binding domain. The C6′ ketone was also found to be very important for binding SrrA and GFP expression, as the ligands that either modify the ketone's position (SRB8) or reduce it (SRB10 or SRB11) have much higher KDs and generally lower GFP expression. The ligands lacking the C6′ ketone (SRB10 and SRB11) are two of the worst performing ligands showcasing the importance of the oxidation state on the sidechain. This is in agreement with the previous reports by Arakawa and co-workers.11 Interestingly, the Emax of the branched SRBs (SRB1, 2, 8 and 10) are all relatively low, indicating that branching plays an important role in derepression of SrrA. Although the Emax for SRB9, the enantiomer of the natural SRB1, is one of the highest, its KD is ∼7× greater than that of SRB1 which is in accordance with what was reported by the Arakawa group.10 Overall, SrrA has the highest affinity for ligands between 9–12 carbons that contain the C6′ ketone (SRB1,2,5,6,7) with the unnatural linear ligands generally having the highest GFP expression. The cause for the lower Emax observed in the natural substrates (SRB1,2) is still being investigated. It may be that the branching does not cause as great of a conformational change in SrrA compared to the linear compounds. S. rochei may have purposely adapted this medium level of induction as a protective measure to avoid self-toxicity during the production of lankamycin and lankacidin.
Conclusions
The ligands SRB1 and SRB2 bind the repressor SrrA to upregulate production of the antibiotics lankamycin and lankacidin. A diversifiable route featuring enantioselective alkynylation and regioselective photo-oxidation was employed to provide access to 11 total SRBs. This ligand library was tested in a GFP reporter assay with the cognate receptor SrrA. Linear ligands with chain lengths between 9–12 carbons performed best with the ligand SRB6 having a very low KDs and high Emax. Synthetic access to this class of signaling molecules in combination with a quantifiable reporter assay will help better study the roles of this class of ligands in other Streptomyces species.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. The SI contains experimental details along with supplementary figures, tables, and spectra. See DOI: https://doi.org/10.1039/d5ob01862e.Acknowledgements
This work was funded by an NSF CAREER Award to E. I. P. (CHE 223689). L. E. W. was supported by a Purdue Research Foundation Ross-Lynn Grant. This work was supported in part by the Research Instrumentation Center in the Department of Chemistry at Purdue University. The authors acknowledge the support from the Purdue Center for Cancer Research, NIH grant P30 CA023168.References
- G. Niu, K. F. Chater, Y. Tian, J. Zhang and H. Tan, Specialised Metabolites Regulating Antibiotic Biosynthesis in Streptomyces Spp, FEMS Microbiol. Rev., 2016, 40(4), 554–573, DOI:10.1093/femsre/fuw012.
- J. L. Ramos, M. Martínez-Bueno, A. J. Molina-Henares, W. Terán, K. Watanabe, X. Zhang, M. T. Gallegos, R. Brennan and R. Tobes, The TetR Family of Transcriptional Repressors, Microbiol. Mol. Biol. Rev., 2005, 69(2), 326–356, DOI:10.1128/MMBR.69.2.326-356.2005.
- S. Kitani, K. T. Miyamoto and S. Takamatsu, Avenolide, a Streptomyces Hormone Controlling Antibiotic Production in Streptomyces Avermitilis, Proc. Natl. Acad. Sci. U. S. A., 2011, 108(39), 16410–16415, DOI:10.1073/pnas.1113908108.
- J. Zhu, Z. Chen, J. Li and Y. Wen, AvaR1, a Butenolide-Type Autoregulator Receptor in Streptomyces Avermitilis, Directly Represses Avenolide and Avermectin Biosynthesis and Multiple Physiological Responses, Front. Microbiol., 2017, 8, 2577, DOI:10.3389/fmicb.2017.02577.
- T. B. Nguyen, S. Kitani, S. Shimma and T. Nihira, Butenolides from Streptomyces Albus J1074 Act as External Signals To Stimulate Avermectin Production in Streptomyces Avermitilis, Appl. Environ. Microbiol., 2018, 84(9), e02791–e02717, DOI:10.1128/AEM.02791-17.
- H. Chen, X. Liu, B. Wei, Y. Tian, Y. Li, J. Zhang, H. Tan and J. Li, Identification of a Novel Butenolide Signal System to Regulate High Production of Tylosin in Streptomyces Fradiae, Appl. Microbiol. Biotechnol., 2025, 109(1), 18, DOI:10.1007/s00253-024-13396-9.
- K. Arakawa, N. Tsuda, A. Taniguchi and H. Kinashi, The Butenolide Signaling Molecules SRB1 and SRB2 Induce Lankacidin and Lankamycin Production in Streptomyces Rochei, ChemBioChem, 2012, 13(10), 1447–1457, DOI:10.1002/cbic.201200149.
- K. Arakawa, S. Mochizuki, K. Yamada, T. Noma and H. Kinashi, γ-Butyrolactone Autoregulator-Receptor System Involved in Lankacidin and Lankamycin Production and Morphological Differentiation in Streptomyces Rochei, Microbiology, 2007, 153(6), 1817–1827, DOI:10.1099/mic.0.2006/002170-0.
- W. Wang, J. Zhang, X. Liu, D. Li, Y. Li, Y. Tian and H. Tan, Identification of a Butenolide Signaling System That Regulates Nikkomycin Biosynthesis in Streptomyces, J. Biol. Chem., 2018, 293(52), 20029–20040, DOI:10.1074/jbc.RA118.005667.
- The Butenolide Signaling Molecules SRB1 and SRB2 Induce Lankacidin and Lankamycin Production in Streptomyces rochei – Arakawa – 2012 – ChemBioChem – Wiley Online Library. https://chemistry-europe.onlinelibrary.wiley.com/doi/full/10.1002/cbic.201200149 (accessed 2024-07-24).
- A. Teshima, N. Hadae, N. Tsuda and K. Arakawa, Functional Analysis of P450 Monooxygenase SrrO in the Biosynthesis of Butenolide-Type Signaling Molecules in Streptomyces Rochei, Biomolecules, 2020, 10(9), 1237, DOI:10.3390/biom10091237.
- A. L. Silva, R. A. Toscano and L. A. Maldonado, An Enantioselective Approach to Furanoeremophilanes: (+)-9-Oxoeuryopsin, J. Org. Chem., 2013, 78(11), 5282–5292, DOI:10.1021/jo400399q.
- S. N. Patil and F. Liu, Fluoride-Assisted Regioselective Conversion of Functionalized Furans to α-Substituted γ-Hydroxybutenolides Using Singlet Oxygen, J. Org. Chem., 2007, 72(16), 6305–6308, DOI:10.1021/jo070666c.
- J. D. Sidda, V. Poon, L. Song, W. Wang, K. Yang and C. Corre, Overproduction and Identification of Butyrolactones SCB1–8 in the Antibiotic Production Superhost Streptomyces M1152, Org. Biomol. Chem., 2016, 14(27), 6390–6393, 10.1039/C6OB00840B.
- Y. Yamada, K. Sugamura, K. Kondo, M. Yanagimoto and H. Okada, The Structure of Inducing Factors for Virginiamycin Production in Streptomyces Virginiae, J. Antibiot., 1987, 40(4), 496–504, DOI:10.7164/antibiotics.40.496.
- G.-Y. Tan, Y. Peng, C. Lu, L. Bai and J.-J. Zhong, Engineering Validamycin Production by Tandem Deletion of γ-Butyrolactone Receptor Genes in Streptomyces Hygroscopicus, 5008, Metab. Eng., 2015, 28, 74–81, DOI:10.1016/j.ymben.2014.12.003.
- L. E. Wilbanks, H. E. Hennigan, C. D. Martinez-Brokaw, H. Lakkis, S. Thormann, A. S. Eggly, G. Buechel and E. I. Parkinson, Synthesis of Gamma-Butyrolactone Hormones Enables Understanding of Natural Product Induction, ACS Chem. Biol., 2023, 18(7), 1624–1631, DOI:10.1021/acschembio.3c00241.
- K. G. Castator, M. Frias-Gomez, L. E. Wilbanks and E. I. Parkinson, Diastereoselective, Diversifiable Synthesis and Biological Evaluation of the Virginiamycin Inducers, the Virginiae Butanolides, ChemBioChem, 2025, 26(17), e202500386, DOI:10.1002/cbic.202500386.
- L. E. Wilbanks, C. B. Roberts, M. Frias-Gomez, H. E. Hennigan, K. G. Castator, Z. L. Budimir, C. Zu and E. I. Parkinson, Streptomyces Autoregulator Biosensors from Natural Product Cluster-Situated Regulators, bioRxiv, 2025, preprint, p. 2025.09.05.673737. DOI:10.1101/2025.09.05.673737.
| This journal is © The Royal Society of Chemistry 2026 |