Open Access Article
Open Access ArticleConnecting the mechanistic steps of cyclic dipeptide formation by a proton-transfer network: pH, temperature, pressure, and nuclear quantum effects
Pimjai Pimbaothama,
John K. Villanueva b,
Siriporn Jungsuttiwong
b,
Siriporn Jungsuttiwong a,
Masanori Tachikawa
a,
Masanori Tachikawa c and
Robert K. Szilagyi
c and
Robert K. Szilagyi *b
*b
aDepartment of Chemistry, Faculty of Science, Ubon Ratchathani University, Ubon Ratchathani 34190, Thailand
bDepartment of Chemistry, I.K. Barber Faculty of Science, University of British Columbia Okanagan, 1177 Research Rd., Kelowna, BC V1 V 1 V7, Canada. E-mail: Robert.Szilagyi@UBC.CA
cQuantum Chemistry Division, Faculty of Nanobioscience, Yokohama City University, Yokohama 236-0027, Japan
First published on 21st January 2026
Abstract
We expanded our previous mapping of the peptide condensation reaction mechanism from the linear dipeptide formation to the cyclization reaction that results in diketopiperazines. The overarching theme of our computational investigations is a reaction network that connects all intermediates via proton-transfer pathways. We conducted the simulations designed to be predictive in a range of environments, such as the gas phase, hydrothermal aqueous conditions, deliquescent salts, and bulk water. While the free-energy profiles are similar to the linear peptide, the presence of the cis amide bond leading to a pre-arranged vicinity of the two reacting groups and the role of explicit solvent molecules revealed new mechanistic insights that differentiate the linear versus cyclic peptide formation/hydrolysis reactions. The rate-determining step corresponds to the final water-elimination reaction using the most realistic computational models with both implicit and explicit water solvation models at neutral pH. At high pH, the highest barrier corresponds to the C–N bond formation at a significantly lower free energy, while at low pH, the water elimination step's barrier increases by close to 30%; thus, effectively shutting down the reaction in agreement with experiments. Due to the central role of proton transfer, we studied the impact of nuclear wave functions on all active H-centers. By utilizing two quantum protons, we document up to 0.1 Å impact on H positions, ca. 20 kJ mol−1 tunneling effects, and a significant change in the shape of the potential energy surface in comparison with the classical DFT calculations. The calculated reaction rates well reproduce the experimentally determined values under hydrothermal conditions.
Introduction
Peptide bond formation and its hydrolysis are fundamental chemical reactions with significant roles and scopes in organic1–4 and biochemical5–8 transformations. Condensation of amino acids is also a long-standing discussion topic in pre-biotic chemistry9–12 as a key process in the chemical evolution of proto-enzymes. The generalized mechanism of the forward reaction of peptide formation10,13 is commonly described by a nucleophilic attack of the amine lone pair on the carboxyl carbon, which is followed by proton transfer from the amine to the carboxylic or to the carbonyl oxygen of the acid. These steps can result in direct water elimination or geminal-diol formation, respectively. Hydrolysis is described as a nucleophilic attack of water on the amide carbonyl carbon to form either the gem-diol intermediate or, in a concerted step, cleave the C–N bond with the dissociation of the amine and carboxyl groups. In the two-step mechanism with the presence of a transient gem-diol intermediate, a proton is transferred between the two hydroxyl groups in order to form the amide carbonyl group and eliminate water for peptide bond formation or vice versa for hydrolysis. However, depending on the chemical environment, the dielectric constant of the solvent and the presence of Lewis acid/base salts, these oversimplified mechanistic steps branch out to new intermediates that are one or two proton-transfer steps away from the aforementioned central intermediates. The energetic significance of the alternative pathways remains obscured, when only considering gas-phase models or even water solvated amino acids with neutral functional groups.Our earlier computational investigation14 systematically built on the gas phase reaction toward implicitly solvated neutral and zwitterionic reactants and products in a microsolvation environment. We documented a complex proton-transfer network with up to 18 different transition states connecting 11 intermediates for the linear diglycine (GlyGly) formation/hydrolysis. Among them, we found alternative, low-energy pathways to the abovementioned generic one or two-step mechanisms. The remarkable richness of the free-energy landscape was particularly evident, when considering zwitterionic forms of the reacting functional groups and the presence of a bulk water solvent environment or microsolvation. Starting with the compositionally well-defined, bulk water reaction at ambient pH, we uncovered a ‘ping-pong mechanism’ of shuttling protons between the newly formed amide bond and the C- and N-termini of GlyGly. The lowest free energy path is marked by the shift in the acidity/alkalinity of the secondary amine/amide groups in the middle and amine/carboxylic groups at the periphery of the molecule. Specifically, upon formation of the C–N bond through a transition state (TS) of Δ‡G° = 92 kJ mol−1 relative to the free energy dissociation limit of two glycine molecules, the negatively charged O atom of the half-formed amide group gets protonated by the N-terminal group (Δ‡G° = 107 kJ mol−1). This triggers the shift of another proton from the positively charged secondary amine N to the C-terminal group (Δ‡G° = 98 kJ mol−1). The consecutive proton-transfer steps give the most stable intermediate along a plateau of the reaction coordinate at 74 kJ mol−1 with a gem-diol moiety and neutral end-groups. The rate-determining water-elimination step is achieved by the now neutral N-terminal group deprotonating the gem-diol (Δ‡G° = 110 kJ mol−1), followed by proton transfer from the neutral C-terminal group to the central hydroxyl group (Δ‡G° = 118 kJ mol−1). Only in the rate-determining last step is the amide π-bond formed, which gives rise to cis/trans stereochemistry. The trans amide bond formation barrier (Δ‡G°) is 6 kJ mol−1 lower and 9 kJ mol−1 more spontaneous, which is relevant to the present study, since the formation of the cis amide conformer can shortcut to the path to the cyclization of a dipeptide without peptide bond isomerisation. The overall reaction driving force for the linear peptide formation was found to be exergonic (ΔG° = −14 kJ mol−1) and significantly exothermic (ΔH° = −22 kJ mol−1) for the trans stereoisomer.
It is important to highlight that this multi-step proton ‘ping-pong mechanism’ can only exist for dipeptides, given the critical vicinity of the acid/base groups for effective intramolecular proton-transfer within ∼3 Å. For oligopeptides with the C- and N-termini being several amino acid groups away, only the higher energy pathway is available with 189 kJ mol−1 TS for the gem-diol formation and 176 kJ mol−1 for water elimination. The significant increase in the barrier height explains why, in a closely related experimental work,15 only dipeptide molecules were detected. When we take into account explicit solvent molecules in assisting the proton transfer through the rearrangements of H-bonding, we documented the inhibitory effect on the last water-elimination step, since the path of the proton movement increased from 2.46 Å to 3.95 Å that corresponds to a 36 kJ mol−1 increase in the barrier height. At the same time, up to four explicit solvent molecules microsolvating the reactive functional groups in the rate-determining step increased the barrier by 11 kJ mol−1. One of the main experimentally relevant conclusions of our previous study on linear dipeptides was the favorable energetics of the C- and N-termini-assisted water-elimination/condensation reaction that explains the absence of oligomeric peptides in bulk water under hydrothermal conditions.
The concept of a proton-transfer network in peptide bond condensation and hydrolysis reactions was a guiding principle to organize the large number of transition states and intermediates along the reaction mechanism. However, it is also informative for considering peptide condensation/hydrolysis reactions at mineral/nanoparticle surfaces16–19 and liquid/gas interfaces.20–23 The arrangement and spacing of acidic/alkaline groups can catalyze or inhibit proton transfer from the secondary amine and the gem-diol groups of the newly formed amide bond. Hereby, we describe a follow-up reaction pathway to dipeptide formation with the cyclization steps that lead to the smallest representative of diketopiperazines (DKPs).24 The kinetics of cyclic GlyGly formation and decomposition have been investigated experimentally25 under systematically controlled hydrothermal conditions. Assuming Arrhenius behavior at pH 9.8, the activation energies were determined to be 93 and 134 kJ mol−1 for DKP formation and the reverse process of ring opening, respectively. This specific pH value was chosen to maximize the linear GlyGly formation, which corresponds to a dominantly anionic Gly presence at 140 °C with a neutral amino group as a requirement for the nucleophilic attack on the C center of the negatively charged carboxylate group. Therefore, we also considered the effect of elevated temperature on barrier heights. Prior computational modelling of cyclization reactions26–28 provided incomplete geometric and energetic descriptions of the reaction mechanism due to either being oversimplified with intermediates and transition states missing or presented in the absence of the cyclic peptide formation. Thus, comparative analyses between linear and cyclic peptide formation remained challenging due to the differences in the employed level of theory and composition of the computational models. Furthermore, the biologically and environmentally relevant solvation environment was not always considered or simplified to a polarizable continuum solvation model without taking into account the effects of covalent bonding from explicit solvent molecules.
In order to create a unified free energy surface for both the linear and cyclic peptide condensation reactions, we applied the proton-transfer network concept for mapping the reaction mechanism. The computational method and the level of theory were maintained as in our previous publication,14 which was selected from a detailed comparison of the accuracy of various density functionals and ab initio correlated MO methods. For being able to provide experimentally relevant rate constants, we reevaluated the barrier heights at the non-standard state that corresponds to the hydrothermal conditions. The pH effects (above 8 and below 3) were taken into account by explicitly considering the presence of hydronium and hydroxide ions for low and high pH conditions, respectively. An enhancement of our quantum chemical description of the peptide formation and hydrolysis reactions was the incorporation of nuclear quantum effects (NQE) using multi-component DFT (MC-DFT) calculations.29–32 Each active proton involved in a proton-transfer network along the reaction coordinate was treated with a nuclear wave function, which allowed for calculating explicitly the proton/proton and proton/electron quantum interactions within polarizable continuum solvation models. The NQE has already been shown to be impactful for low-barrier H-bonded systems, describing kinetic isotope effects, and impacting the geometric structures, barrier heights, and relative energies of intermediates.33–37
Computational models and methods
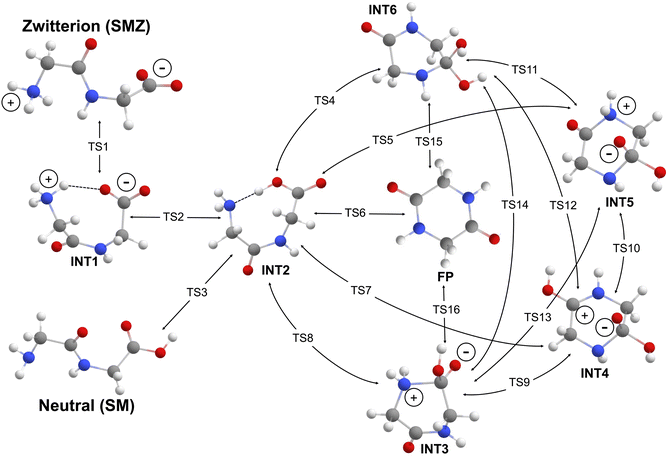
The map of all the intermediates and their connecting transition states are presented in Scheme 1, starting from the linear, zwitterionic dipeptide structure (SMZ) on the top left-hand side. Unique to the dipeptide cyclization are TS1 for the zwitterionic dipeptide and TS3 for the neutral linear dipeptide which are required to ensure that the C- and N-termini of the linear dipeptide will be in proximity despite that the first peptide bond is in the energetically less stable cis stereoisomeric form (INT1). However, INT1 can be directly formed during the linear peptide condensation reaction as a function of which the hydroxyl group of the gem-diol intermediate is the source of the eliminated water molecule. The corresponding transition state is only 10 kJ mol−1 higher than that for the trans isomer. When considering the zwitterionic form, another mandatory process is neutralization via TS2 to form INT2, which poises the N lone pair for nucleophilic attack on the carbonyl carbon. Concomitantly, the C–N bond formation can take place via several pathways (TS4–TS8). The final DKP product can be achieved directly from INT2 via TS6 in a single-step process. Alternatively, the reaction can proceed through neutral INT6 and zwitterionic INT3 intermediates. Notably, in this representation, the proton can be substituted with another Lewis acid, which can be another analyte from the solution (transition metal ions, ammonium cations) and part of a coordinatively unsaturated metal site on a nanoparticle or mineral surface. TS5 and TS7 open up the reaction toward the right-hand side of Scheme 1, where the amide bond of the dipeptide can act as a proton acceptor from the secondary amine group. Notably, INT4 and INT5 are also likely intermediates for the cleavage of the first peptide bond, which would result in a non-productive cyclization reaction.All potential energy surface scans and localization of equilibrium structures without imaginary normal modes were carried out using the MN15/def2TZVPP level of theory.38,39 This level of theory has been systematically validated in our previous work14 on the condensation and hydrolysis reactions of linear dipeptides to produce CCSD(T)-F12/haTZ intermediates and transition states for amide models.40 Transition-state structures characterized by a single imaginary normal mode were confirmed by intrinsic reaction coordinate (IRC) calculations41 to connect their respective intermediates. The intermediate structures presented in the free-energy plots shown here correspond to the lowest-energy conformations between the two transition states without explicitly showing the exit and entry structures of the IRC calculations for the sake of clarity. For implicit solvation, the SMD continuum model42 was utilized with default water parameters. Similarly to the linear peptide condensation reaction, we identified post-TS structures along the cyclic peptide pathways that were higher in enthalpy and free energy than the corresponding TSs. However, analysis of the QM potential energy surface employing IRC analysis confirmed that all structures used in this study correspond to valid equilibrium structures. Upon evaluation of the quantum-chemical potential energies (ΔEQM), we documented excessive energetic stabilization of the transition states that can result in negative barrier heights for the reverse reaction (see discussion below, TS2 in Fig. 2).
The impact of pH was considered by changing the protonation states of the amino acids and bringing into the reaction mechanism the presence of hydronium and hydroxide ions for low and high pH environments, respectively. The temperature effects on relative energies were estimated by recalculating thermal energy corrections to enthalpy and entropy for adjusting the free-energy values. We also considered the effects of the condensed phase by considering a significantly higher pressure (1748 atm),43 which corrects the translational entropy by taking into account the density of the solution versus gas phase free mean path. Nuclear quantum effects on the protonic wave function were estimated by using the multi-component DFT (MC-DFT) formalism with a Gaussian-type protonic wave function.32 Given that the MC-MO or MC-DFT methods are relatively recent developments,30,31,44,45 we provide here a brief introduction to their unique features. MC-MO simultaneously treats both electrons and nuclei within a quantum framework using a multi-component scheme.46,47 The total Hamiltonian for the combined electronic and protonic wave functions is expressed as
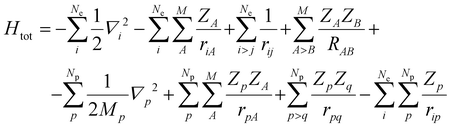 | (1) |
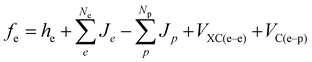 | (2) |
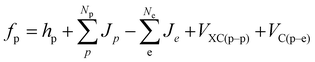 | (3) |
Results and discussion
Similarly to the description of the linear peptide condensation reaction,14 we defined four scenarios: (I) gas-phase models, (II) neutral dipeptide reactions in a continuum, (III) zwitterionic dipeptides with implicit solvation and (IV) zwitterionic dipeptides with explicit water solvation in order to describe environmental models for atmospheric or vacuum conditions, a simplified hydrothermal bulk water environment, and wet/dry conditions of deliquescent salts, respectively.Scenarios I and II: neutral species
Starting from the extended linear peptide with a trans amide bond, the cyclization requires access to the cis isomer through TS3, which has a barrier of Δ‡G° = 96 kJ mol−1. Notably, the arrangement of the amide carbonyl and N–H bonds in TS3 can be characterized with a dihedral angle of 57°. The deviation from the ideal perpendicular arrangement arises from intramolecular H-bonding between the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and N–H moieties. When considering an aqueous continuum solvent, the free energy of solvation for the linear trans SM is −75 kJ mol−1, while it is −82 kJ mol−1 for INT2; hence, the 7 kJ mol−1 stabilization for INT2, as shown in Fig. 1. The cis isomer was calculated to be +34 and +27 kJ mol−1 less stable than the trans isomer in the gas and condensed phases, respectively. Importantly, the rotational barrier in the gas-phase and implicit-solvation models is approximately half of that associated with water hydrolysis steps in linear peptide formation (179 kJ mol−1);14 thus, during isomerization, the first peptide bond is expected to remain intact.
O and N–H moieties. When considering an aqueous continuum solvent, the free energy of solvation for the linear trans SM is −75 kJ mol−1, while it is −82 kJ mol−1 for INT2; hence, the 7 kJ mol−1 stabilization for INT2, as shown in Fig. 1. The cis isomer was calculated to be +34 and +27 kJ mol−1 less stable than the trans isomer in the gas and condensed phases, respectively. Importantly, the rotational barrier in the gas-phase and implicit-solvation models is approximately half of that associated with water hydrolysis steps in linear peptide formation (179 kJ mol−1);14 thus, during isomerization, the first peptide bond is expected to remain intact.
The specific orientations of INT2 and then TS4 and TS6 in Fig. 1 illustrate well how the reaction coordinate splits into one-step and two-step mechanisms. Upon the nucleophilic attack of the N-terminal lone pair on the carbonyl carbon through TS4, one of the H atoms of the reacting amine group can transfer either to the carbonyl or the carboxylic group. The former results in the formation of a gem-diol intermediate (INT6, black and blue traces; hence, a two-step process), while the second leads to an instant water elimination step (FP, in a one-step process). The cyclization transition states for the gas phase (205 kJ mol−1) and condensed phase (189 kJ mol−1) parallel the analogous structures for linear peptide formation (216 and 203 kJ mol−1). The approximately 11 and 14 kJ mol−1 lower barrier heights in free energy can be attributed to the presence of the cis amide bond that brings the two reacting ends in proximity and, thus, lowers the reorganization energy contribution to the transition state. The impact of the structural rigidity and compactness is also evident in the formation of the gem-diol intermediate INT6 via TS4 at 176 and 156 kJ mol−1 as compared to 199 and 189 kJ mol−1 for the linear peptide pathways in the gas phase and condensed phase, respectively. The lowering of the transition state energy and the consequent increase in the reaction rate for the gem-diol conversion to the final product DKP suggest that cyclization can proceed more favorably than hydrolysis of the first amide bond in linear peptides under gas-phase and low-dielectric/hydrophobic conditions. The resulting accumulation of DKP as one of the condensation products was also observed experimentally.25 The overall thermodynamic driving force for the second amide bond formation in the cyclic peptide is of a similar magnitude (−18 kJ mol−1) to that of the first in the linear peptide (−23 kJ mol−1). These indicate that cyclization is a spontaneous reaction when a cis-amide bond is present with the rate-determining step in the gas phase being C–N bond formation, while water is eliminated in model aqueous environments, although the preference is only by a small extent.
Scenario III
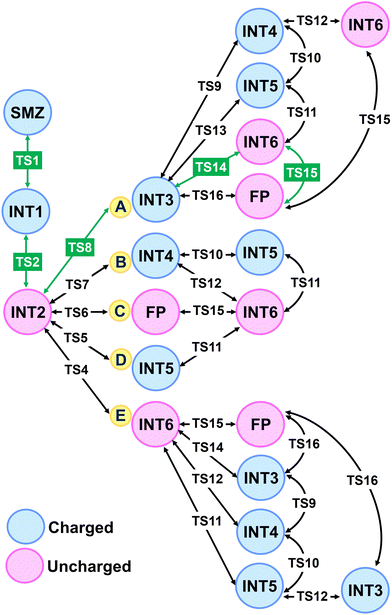
For the bulk water scenario using ionized functional groups in a continuum model, we conducted the mapping of all possible proton-transfer pathways described in Scheme 1. Already for a modest system of 9 intermediates and 16 transition states, the combinatorial possibilities are remarkable as summarized in Scheme 2. Notably, all reactions need to proceed through a bottleneck intermediate, INT2 with neutral N- and C-termini groups; TS4–TS8 connect the linear peptide with INT3–INT6, FP, and the entire right-hand side of Scheme 1. It is also notable that the shortest pathway with the least number of steps via TS6 does not necessarily correspond to the lowest free-energy pathway as discussed below.Among all reactivity pathways shown in Fig. S1 of the SI, pathways A and E are the most energetically competitive via TS14 (Δ‡G° = 130 kJ mol−1 in path A) and TS4 (Δ‡G° = 135 kJ mol−1 in path E) gem-diol formation barriers from the secondary ammonium and deprotonated diol INT3 and the proton-transferred INT2, respectively. Eventually, both processes will need to pass through TS15 (133 kJ mol−1) to reach the DKP final product by water elimination from the gem-diol intermediate (INT6). The calculated barriers slightly favor path A; thus, INT3 with a charged secondary ammonium group and the adjacent deprotonated gem-diol moiety will be stabilized by the solvation model in comparison with the gem-diol intermediate INT3 in path B. TS9–TS13 exhibit significantly higher activation barriers leading to INT4 and INT5 zwitterionic intermediates than anything up to the formation of INT3 and INT6; therefore, these pathways will not be expected to be operative in bulk condensed-phase water environments, although they may gain importance under high ionic strength conditions and potentially in deliquescent salt solutions.
Following the path highlighted in green in Scheme 2, the free-energy landscape shifts significantly to lower energies, and the overall activation barrier heights are reduced due to the presence of zwitterionic species in a water-solvated environment in contrast to Scenarios I and II. The rotational barrier through TS1 is reduced by 20 kJ mol−1 in comparison with the neutral solvated model (Fig. 1 vs. 2). Proton transfer in TS2 exhibits a negligible barrier due to the rigidity of the cis amide bond that brings together the C- and N-termini within 1.35 Å distance. When considering the potential-energy plot (pink trace, ΔEQM in Fig. S2), the apparent anomaly between TS2 and INT2 can be rationalized by the significant enthalpic stabilization and increased entropic contribution of TS2 relative to the pre- and post-TS intermediates. Importantly, the identity of TS2 as a genuine transition state connecting INT2 to the subsequent intermediate has been explicitly confirmed by intrinsic reaction coordinate (IRC) calculations (Fig. S4). Furthermore, the carboxylic H–O group position is in a rotated anti-conformation in INT2, which destabilizes the structure by 5 kJ mol−1 compared to the more common conformer with syn-arrangement of the H–O and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O groups. Following a conformational change of +8 kJ mol−1, the lone pair of the N-terminus is aligned for nucleophilic attack on a 24 kJ mol−1 more stable structure due to the presence of the continuum (INT2 in Fig. 1 vs. 2). Importantly, the implicit solvation stabilizes the protonated secondary amine and the ionized gem-diol groups of INT3 through a late transition state of the C–N bond formation step (TS8). The potential-energy plot (ΔEQM) in Fig. 2 for TS8 shows the slightly lower energy of INT3 than that of TS8. In support of the importance of considering the proton-transfer network as shown in Scheme 1, INT3 does not convert directly to the FP through TS16 at 163 kJ mol−1, but it passes along a lower energy path through TS14, connecting to INT6 at 130 kJ mol−1. This is followed by TS15 to afford the FP after passing through the rate-limiting step of water elimination at a free energy of 133 kJ mol−1. Upon dissociation of the eliminated water molecule, the DKP exhibits a second amide bond that is energetically less stable than the first in the linear peptide due to ring strain arising from keeping the methylene groups in plane and the formation of a near-C2h-symmetric dipeptide. The solvation of the zwitterionic form of the FP enhances the thermodynamic driving force by about 19 kJ mol−1 for the cyclization in relation to the neutral species at the endpoints in Scenarios I and II in Fig. 1.
O groups. Following a conformational change of +8 kJ mol−1, the lone pair of the N-terminus is aligned for nucleophilic attack on a 24 kJ mol−1 more stable structure due to the presence of the continuum (INT2 in Fig. 1 vs. 2). Importantly, the implicit solvation stabilizes the protonated secondary amine and the ionized gem-diol groups of INT3 through a late transition state of the C–N bond formation step (TS8). The potential-energy plot (ΔEQM) in Fig. 2 for TS8 shows the slightly lower energy of INT3 than that of TS8. In support of the importance of considering the proton-transfer network as shown in Scheme 1, INT3 does not convert directly to the FP through TS16 at 163 kJ mol−1, but it passes along a lower energy path through TS14, connecting to INT6 at 130 kJ mol−1. This is followed by TS15 to afford the FP after passing through the rate-limiting step of water elimination at a free energy of 133 kJ mol−1. Upon dissociation of the eliminated water molecule, the DKP exhibits a second amide bond that is energetically less stable than the first in the linear peptide due to ring strain arising from keeping the methylene groups in plane and the formation of a near-C2h-symmetric dipeptide. The solvation of the zwitterionic form of the FP enhances the thermodynamic driving force by about 19 kJ mol−1 for the cyclization in relation to the neutral species at the endpoints in Scenarios I and II in Fig. 1.
Scenario IV
The free-energy landscape of the cyclization reaction in the presence of both explicit and implicit solvation is shown in Fig. 3. Various microsolvated models were constructed in order to capture most of the critical covalent solvent/solute interactions that can impact the reaction coordinate by stabilizing ionized molecules in addition to the electrostatic interactions from the implicit polarizable continuum. The presence of explicit water solvent molecules also allows spontaneous proton transfer to take place, which is important when pH effects are considered. Notably, for the linear peptide condensation reaction,14 we found that the presence of explicit solvent molecules actually increases the proton-transfer activation barriers. This can also be seen for the neutralizing proton-transfer TS2 from the zwitterionic cis-isomer, INT1, where the barrier height (Δ‡G°) relative to the zwitterionic trans-dipeptide reference point increased from 4 to 36 kJ mol−1. However, this destabilization effect of transition states becomes computationally advantageous in our case, since the microsolvation environment around the reactive part of the peptide eliminates anomalies in the relative energies of TS and adjacent INT structures as shown in Fig. 3. The TS1 amide bond isomerization energy remains the same, and all other transition states are lowered by 16–18 kJ mol−1. A notable exception to this trend is the significant lowering of TS14 (gem-diol forming proton-transfer path) which decreases to 91 kJ mol−1 in Fig. 3 from 130 kJ mol−1 in Fig. 2. While the proton-transfer process is highly similar to TS2, the key difference from TS14 is the adjacent location of formal cationic (–NH2+–) and anionic (–C(O−)(OH)–) groups. The charge separation is attenuated by the presence of a polarized water solvent molecule, which contributes to the lowered barrier. The impact of these interactions is further evaluated in the nuclear quantum effect section (see below). Importantly, for the outcome of the reaction, the free energy of the rate-determining step in TS15 is only about 3 kJ mol−1 higher than the relevant saddle points in Scenario III, which shows that explicit water has a diminishing impact on the overall reaction energetics than is generally concluded from the peptide condensation literature.4,20,58–61Impact of nuclear quantum effects (NQE)
A potential critique of the abovementioned free-energy profiles for all scenarios is the Coulombic point-charge treatment of the protons in the nuclear/nuclear repulsion and nuclear/electron attraction terms of the Hamiltonian. This theoretical level can be enhanced by describing the active protons as nuclear wave functions between donor ammonium and acceptor carboxylate groups in the neutralization transition state TS2, assisted by explicit water molecules in Scenario IV. Hydrogens along the most dominant normal-mode displacements shown by arrows in Fig. 3 for TS2 were treated as quantum protons in SMZ, pre- and post-TS intermediates, INT1 and INT2, respectively. Furthermore, in Fig. 3, two additional steps can be highlighted that involve proton transfers: the gem-diol formation (TS14) and the water elimination step (TS15), which are prime candidates for assessing the impact of the presence of the nuclear wave function.Table 1 summarizes the nuclear quantum effects (rows marked with Δ) on the potential energies of the TS2, TS14, and TS15 transition states along the lowest-energy pathway as a function of the composition of the density functional and basis set. The segments of the potential-energy plots at each level of theory are also summarized in Fig. S3. Fig. S4 shows the differences among IRC path calculations with or without nuclear quantum effects. An extended discussion of the impact of the protonic wave function is also provided in the SI.
| Level of theory | Δ‡EQM, kJ mol−1 | ||
|---|---|---|---|
| Functional/basis set | TS2 | TS14 | TS15 |
| The TS energies are given relative to the zwitterionic linear dipeptide (SMZ). The rows containing multicomponent DFT calculations (“Δ(MC−)”) correspond to the Δ(Δ‡EQM) values as the differences between MC-DFT and classical DFT calculations at the same level of theory. | |||
| MN15/def2TZVPP | 44 | 81 | 139 |
| MN15/cc-pVTZ | 43 | 105 | 135 |
| Δ(MC-MN15/cc-pVTZ) | −18 | −20 | −19 |
| B3LYP/cc-pVTZ | 47 | 105 | 170 |
| Δ(MC-B3LYP/cc-pVTZ) | −20 | −14 | −26 |
| B3LYP/6-31G(d,p) | 30 | 87 | 148 |
| Δ(MC-B3LYP/6-31G(d,p)) | −21 | −15 | −24 |
As anticipated, incorporation of the protonic wave function reduces the relative potential energy differences for the proton-transfer transition states by adding more stabilizing quantum interactions. The magnitude of the energy decrease or the tunnelling effect can be up to 20 kJ mol−1. As we move away from the MN15 functional (with 44% HF exchange, hybrid meta-GGA functional) to B3LYP (with 20% HF exchange, hybrid GGA functional, as well as lower chemical accuracy for the peptide condensation reactions14), the tunnelling effect gradually reduces for TS14, while it increases for both TS2 and TS15 up to −26 kJ mol−1. A modest basis set effect (at most 2 kJ mol−1 difference) can be detected for using a double-ζ quality basis set with the density functional that exhibits extreme barrier heights without considering NQE. Importantly, a change in the amount of HF exchange has a significantly less influence on the barrier heights than the presence of protonic wave functions at centers involved in proton transfer. In terms of geometrical differences, the most notable change is the dramatic shortening of the secondary ammonium–cation distance from 2.65 Å to 1.94 Å in INT3 before TS14, as the quantum proton becomes shared between the N and O centers of the amide. The quantum mechanical treatment of the nuclear interactions reshapes the potential-energy surface as seen from the comparisons of the IRC scans (shown in Fig. S5) and contributes to the increase in the barrier height for TS14 in comparison with TS2 and TS15 due to the over-stabilization of INT3.
Impact of hydrothermal conditions
Continuing with Scenario IV, we assessed the temperature and pressure influence on the potential-energy surface. Experiments on bulk amino acid condensation15 were carried out at an elevated temperature of 140 °C. Even without molecular dynamics simulations, we can correct for the occupations of translational, rotational, and vibrational quantum levels as a function of temperature increase by recalculating the thermal corrections to both enthalpy and free energy using statistical mechanics (see Table S1 for the numerical results). Another consideration is the non-standard state due to the condensed phase, which is simulated by 1748 atm pressure that corresponds to the approximately 1 kg dm−3 density for the bulk water system. The non-standard pressure had a negligible (<1 kJ mol−1) impact on the free energy (Table S1), mainly due to the dominantly equimolar reaction steps. However, the temperature (Fig. S5) introduced some notable changes, since the reaction barrier (Δ‡G) for the rate-limiting water-elimination step increased by about 8 kJ mol−1 in comparison with TS15 in Scenario IV from Fig. 3. Other notable differences are the smaller destabilization of TS8 (C–N bond formation step) but a significantly more altered barrier for TS14 (+14 kJ mol−1) for the neutralization reaction by forming the gem-diol intermediate with the assistance of an explicit water molecule. The overall thermodynamic driving force also increased by elevating the temperature by 20 kJ mol−1, due to the entropically favorable water elimination.Impact of pH
In an attempt to bring the simulation results even closer to the experimental observations, we considered the pH-induced changes in protonation states at the reactive ends of the amino acids. The experimental activation barriers for DKP formation and hydrolysis were specifically determined for the highest rates at pH 9.8.25 In contrast, there was barely observable condensation reaction products at low pH; hence, no rate constants were determined. The key structural differences in the GlyGly dipeptide are due to the pKa values of 3.14 and 8.25, which are less spread than those of the pKa values of the Gly amino acid (2.3 and 9.6).62 Therefore, high pH conditions were modelling with the presence of anionic GlyGly with neutral N-terminal and anionic C-terminal groups; while at low pH, both termini are protonated to give cationic GlyGly structure.Fig. 4 summarizes the simplified cyclization pathways at low pH (below pH = 3), where INT2-a (‘a’ refers to protonated, acidic conditions) forms the C–N bond along with the simultaneous transfer of the ammonium proton to the carbonyl group to form the gem-diol intermediate INT5-a. We exhausted our toolchest to locate a transition state for INT2-a → INT5-a. The lack of a well-defined barrier can be chemically rationalized, since as the proton transfers from the cationic N-terminus to the C-terminal carbonyl, the newly formed N lone pair is already in range for the instantaneous nucleophilic attack on the carboxyl carbon with a significant carbocation character in the –C+(OH)2 moiety. If a water molecule is involved in the proton transfer, the deprotonated gem-diol upon the formation of the C–N bond would immediately pick up a proton from the adjacent hydronium and end up in the INT5-a state. The proton transfer from the secondary ammonium group to the gem-diol hydroxyl group needs to overcome a much higher barrier (Δ‡G′ = 192 kJ mol−1) than estimated for neutral pH conditions (136 kJ mol−1, Fig. 3). The deprotonation of the carbonyl group in INT7-a is another barrierless process as the elimination product of the hydronium cation is formed in FP-a. The overall thermodynamic driving force is significant (−10 kJ mol−1); however, the low pH cyclization process is kinetically controlled.
The high pH (pH > 8) reaction path starting from the anionic GlyGly dipeptide changes the kinetic control of the cyclization reactions, as shown in Fig. 5. The rate-determining step now shifts to the C–N bond formation (TS1-b, ‘b’ refers to deprotonated, alkaline conditions) and the overall barrier height (Δ‡G′) is reduced by close to 15 kJ mol−1 relative to the analogous step for zwitterionic GlyGly (139 kJ mol−1). This is mainly due to the favorable pre-TS complex with the N-terminal lone pair pre-arranged to be aiming at the carboxylate carbon already in INT2-b. The proton from the amine end-group transfers to form the singly deprotonated gem-diol intermediate INT5-b in order to avoid the accumulation of −2 charges on the adjacent two hydroxyl groups. Due to a lack of a secondary ammonium cation and need for additional proton-transfer steps, the leaving group is hydroxide through TS2-b with a significantly lower barrier than that observed for the neutral or acidic pH elimination step (85 kJ mol−1). Due to the formation of an identical product molecule, DKP, the pH dependence of the thermodynamic driving force can be determined by the species eliminated: H2O at neutral pH and H3O+ or HO− at low or high pH, respectively. Given that DKP is an electron-rich molecule, the −42 kJ mol−1 more stable cation interaction with H3O+ than HO− is justified.
In order to assess the experimentally relevant impact of various barrier heights at different temperatures, we estimated reaction rate constants using the Eyring equation, which relates hypothetical equilibrium conditions (K‡) and the vibrational motion coupling the reactants and their transition states. With Δ‡G°(25 °C) = 136 kJ mol−1 and Δ‡G(140 °C) = 144 kJ mol−1, the rate constants can be estimated to be 9.2 × 10−12 and 5.4 × 10−6 s−1, respectively. At the alkaline pH range and elevated temperature, the rate drastically increases to 1.3 × 10−4 s−1 corresponding to Δ‡G′ = 124 kJ mol−1. The experimental rates at a non-standard temperature of 140 °C were determined to be 2.5 × 10−5 and 4.0 × 10−5 s−1 for neutral and high pH, respectively.25 Some low amounts of products were detected experimentally at low pH; however, no rate constant was reported. While our computational models cannot reproduce quantitatively the experimental numbers due to the inherent compositional limitations of the model and likely the incomplete explicit solvation shell for taking into account solvent reorganization or solvation/desolvation processes, the relative differences among the estimated rate constants highlight the adequate chemical accuracy that can be used to predict and rationalize reaction mechanisms.
Conclusions
The mechanistic insights obtained from the systematic mapping of the proton-transfer network for cyclic diglycine (diketopiperazine, DKP) formation complete our previous work on glycine/glycine condensation reaction in bulk water at neutral pH. Importantly, the activation energies to obtain both linear or cyclic GlyGly peptides are low to moderate. The reverse process of hydrolysis has consistently larger barriers in agreement with the experiment, which allows for the accumulation of both linear and cyclic products. This is in part due to the thermodynamic stability of the peptide bond. For dipeptides, the cyclic product is only accessible if the cis-peptide isomer is present. The peptide bond rotation energy is about half that of the rate-determining step, H2O elimination's activation barrier; thus, this likely takes place in bulk water to a limited extent. An alternative pathway that can directly lead to the cis isomer and then to the cyclic peptide product is from the water elimination step during the linear peptide oligomerization. All transition states are in agreement with a significant thermodynamic preference toward the linear oligomerization; however, once the cyclic dipeptide is formed, its kinetic stability prevents the decomposition. Common to both processes is the unique role of the adjacent functional groups that can accept or donate excess protons, as dictated by the relative stability of acidic or basic groups. The ping-pong mechanism defined by the lowest free-energy pathway can also be considered as a reference for proton-transfer reactions in our ongoing automated reaction route mapping studies. The central focus on proton-transfer processes invited the employment of nuclear wave functions in order to improve the quantum description of protons. Using the MC-DFT method, we found up to 0.1 Å elongation and up to 20 kJ mol−1 tunnelling effect in interatomic distances and thermochemistry, respectively. Dominant changes were manifested in the restructuring of the potential energy curvature in the presence of quantum protons that in part rationalizes some of the challenges in localizing transition state structures and using specific basis sets for MC-DFT implementation. In order to understand the pH-dependent condensation reactions, we defined the free energy profiles for acidic (pH < 3) and alkaline (pH > 8) scenarios. We found mechanistic evidence for the kinetic control under acidic conditions that shuts down the condensation reactions, while alkaline conditions reorganize the potential energy surface and shift the rate-determining step toward the C–N bond formation with a significantly lower activation energy. The estimated rate constants show the correct qualitative differences and afford experimentally meaningful values to use the computational mechanistic investigation as a tool for amide bond condensation and hydrolysis design.Author contributions
Computations, data curation, visualization, & preparation of the initial draft: P. Pimbaotham. Implementation and benchmarking of NQE calculations: J. Villanueva. Method and software development, and supervision of nuclear quantum calculations: M. Tachikawa. Synchronization of experimental conditions and computational modelling: S. Jungsuttiwong. Funding acquisition, project administration, resources, and research supervision: R. K. Szilagyi. Conceptualization, investigation, methodology, software, and writing – review and editing were contributed by all authors.Conflicts of interest
There are conflicts to declare.Data availability
Data supporting this article have been included as part of the supplementary information (SI). Supplementary information: extended discussions, poster presentations, and supporting figures and tables for enriching the presentation and discussion of the cyclization mechanism as well as to provide an explanation for unexpected trends and anomalies discovered during the research. See DOI: https://doi.org/10.1039/d5ob01815c.The molecular structures, formatted checkpoint files with electronic structure information, and electronic spreadsheets with all data are deposited. We created a new version of the dataset that combines the results of both linear and cyclic dipeptide formation mechanisms (https://doi.org/10.5281/zenodo.18264551).
Acknowledgements
P. P. is thankful for receiving financial support from the Canada-ASEAN Scholarships and Educational Exchanges for Development Program for a visiting Ph.D. fellowships at UBC Okanagan. This project was also supported in part by the National Research Council of Thailand, Thailand (NRCT): N41A650123. We acknowledge the financial support from the UBC Aspire Program (AWD-019715 UBCOVPR 2021) for R. K. Sz. Computational modelling was enabled in part by support provided by the Advanced Research Computing at the University of British Columbia (arc.ubc.ca) and the Digital Research Alliance of Canada (alliancecan.ca).References
- J. S. Carey, D. Laffan, C. Thomson and M. T. Williams, Org. Biomol. Chem., 2006, 4, 2337–2347 RSC.
- R. M. de Figueiredo, J. S. Suppo and J. M. Campagne, Chem. Rev., 2016, 116, 12029–12122 CrossRef CAS.
- X. Wang, Nat. Catal., 2019, 2, 98–102 CrossRef.
- F. S. Brigiano, M. Gierada, F. Tielens and F. Pietrucci, ACS Catal., 2022, 12, 2821–2830 CrossRef CAS.
- T. M. Schmeing, K. S. Huang, D. E. Kitchen, S. A. Strobel and T. A. Steitz, Mol. Cell, 2005, 20, 437–448 CrossRef CAS.
- K. Swiderek, S. Marti, I. Tuñón, V. Moliner and J. Bertran, J. Am. Chem. Soc., 2015, 137, 12024–12034 CrossRef CAS.
- M. Kazemi, J. Socan, F. Himo and J. Åqvist, Nucleic Acids Res., 2018, 46, 5345–5354 CrossRef CAS PubMed.
- R. David, I. Tuñón and D. Laage, J. Am. Chem. Soc., 2024, 146, 14213–14224 CrossRef CAS PubMed.
- K. H. Lemke, R. J. Rosenbauer and D. K. Bird, Astrobiology, 2009, 9, 141–146 CrossRef CAS.
- E. C. Griffith and V. Vaida, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 15697–15701 CrossRef CAS PubMed.
- T. Stolar, S. Grubesic, N. Cindro, E. Mestrovic, K. Uzarevic and J. G. Hernández, Angew. Chem., Int. Ed., 2021, 60, 12727–12731 CrossRef CAS PubMed.
- Y. L. Yang, Z. B. Wang, J. Bai and H. Qiao, J. Mol. Evol., 2025, 93, 193–211 CrossRef CAS PubMed.
- J. H. Jones, in Major Methods of Peptide Bond Formation, ed. E. Gross and J. Meienhofer, Academic Press, New York, U.S.A., 2014, vol. 1, ch. 3, pp. 65–104 Search PubMed.
- T. Herriman and R. K. Szilagyi, Org. Biomol. Chem., 2023, 21, 5953–5963 RSC.
- W. Takahagi, K. Seo, T. Shibuya, Y. Takano, K. Fujishima, M. Saitoh, S. Shimamura, Y. Matsui, M. Tomita and K. Takai, ACS Earth Space Chem., 2019, 3, 2559–2568 CrossRef CAS.
- K. Marshall-Bowman, S. Ohara, D. A. Sverjensky, R. M. Hazen and H. J. Cleaves, Geochim. Cosmochim. Acta, 2010, 74, 5852–5861 CrossRef CAS.
- E. Schreiner, N. N. Nair, C. Wittekindt and D. Marx, J. Am. Chem. Soc., 2011, 133, 8216–8226 CrossRef CAS PubMed.
- J. F. Lambert, M. Jaber, T. Georgelin and L. Stievano, Phys. Chem. Chem. Phys., 2013, 15, 13371–13380 RSC.
- V. Erastova, M. T. Degiacomi, D. G. Fraser and H. C. Greenwell, Nat. Commun., 2017, 8, 9 CrossRef.
- A. G. Gale, T. T. Odbadrakh, B. T. Ball and G. C. Shields, J. Phys. Chem. A, 2020, 124, 4150–4159 CrossRef CAS PubMed.
- A. M. Deal, R. J. Rapf and V. Vaida, J. Phys. Chem. A, 2021, 125, 4929–4942 CrossRef CAS.
- W. X. Wang, L. N. Qiao, J. He, Y. Ju, K. Yu, G. F. Kan, C. L. Guo, H. Zhang and J. Jiang, J. Phys. Chem. Lett., 2021, 12, 5774–5780 CrossRef CAS.
- W. Wei, F. J. Chu, G. R. Chen, S. W. Zhou, C. R. Sun, H. R. Feng and Y. J. Pan, Chem. – Eur. J., 2024, 30, 8 Search PubMed.
- A. D. Borthwick, Chem. Rev., 2012, 112, 3641–3716 CrossRef CAS PubMed.
- K. Sakata, N. Kitadai and T. Yokoyama, Geochim. Cosmochim. Acta, 2010, 74, 6841–6851 CrossRef CAS.
- Y. Li, F. F. Li, Y. Y. Zhu, X. Li, Z. Y. Zhou, C. M. Liu, W. J. Zhang and M. S. Tang, Struct. Chem., 2016, 27, 1165–1173 CrossRef CAS.
- S. Freza, J. Mol. Model., 2019, 25, 7 CrossRef PubMed.
- S. Achour, Z. Hosni, S. Darghouthi and C. Syme, Heliyon, 2021, 7, e07276 CrossRef PubMed.
- T. Ishimoto, M. Tachikawa and U. Nagashima, J. Chem. Phys., 2006, 125, 8 Search PubMed.
- T. Udagawa and M. Tachikawa, J. Chem. Phys., 2006, 125, 9 CrossRef PubMed.
- T. Udagawa, T. Tsuneda and M. Tachikawa, Phys. Rev. A, 2014, 89, 6 CrossRef.
- M. Hashimoto, T. Ishimoto, M. Tachikawa and T. Udagawa, Int. J. Quantum Chem., 2016, 116, 961–970 CrossRef.
- T. Ishimoto, M. Tachikawa and U. Nagashima, Int. J. Quantum Chem., 2009, 109, 2677–2694 CrossRef.
- T. Udagawa, K. Suzuki and M. Tachikawa, ChemPhysChem, 2015, 16, 3156–3160 CrossRef PubMed.
- H. Sugimoto, M. Tachikawa and T. Udagawa, Int. J. Quantum Chem., 2019, 119, 10 CrossRef.
- N. Yodsin, H. Sakagami, T. Udagawa, T. Ishimoto, S. Jungsuttiwong and M. Tachikawa, Mol. Catal., 2021, 504, 10 Search PubMed.
- Y. Shimohata, H. Sakagami, Y. Kanematsu, D. S. R. Rocabado, M. Tachikawa and T. Ishimoto, J. Phys. Chem. C, 2023, 127, 24316–24323 CrossRef.
- F. Weigend and R. Ahlrichs, Phys. Chem. Chem. Phys., 2005, 7, 3297–3305 RSC.
- H. S. Yu, X. He, S. L. Li and D. G. Truhlar, Chem. Sci., 2016, 7, 5032–5051 RSC.
- E. Van Dornshuld, R. A. Vergenz and G. S. Tschumper, J. Phys. Chem. B, 2014, 118, 8583–8590 Search PubMed.
- K. Fukui, Acc. Chem. Res., 1981, 14, 363–368 CrossRef.
- A. V. Marenich, C. J. Cramer and D. G. Truhlar, J. Phys. Chem. B, 2009, 113, 6378–6396 Search PubMed.
- R. L. Martin, P. J. Hay and L. R. Pratt, J. Phys. Chem. A, 1998, 102, 3565–3573 CrossRef.
- T. Kreibich and E. K. U. Gross, Phys. Rev. Lett., 2001, 86, 2984–2987 CrossRef PubMed.
- Y. Kita, T. Udagawa and M. Tachikawa, Chem. Lett., 2009, 38, 1156–1157 Search PubMed.
- M. Tachikawa, K. Mori, H. Nakai and K. Iguchi, Chem. Phys. Lett., 1998, 290, 437–442 Search PubMed.
- T. Udagawa, T. Ishimoto and M. Tachikawa, Molecules, 2013, 18, 5209–5220 CrossRef PubMed.
- K. Suzuki, M. Tachikawa and M. Shiga, J. Chem. Phys., 2010, 132, 7 Search PubMed.
- T. Yoshikawa, S. Sugawara, T. Takayanagi, M. Shiga and M. Tachikawa, Chem. Phys. Lett., 2010, 496, 14–19 Search PubMed.
- A. A. Auer, J. Gauss and J. F. Stanton, J. Chem. Phys., 2003, 118, 10407–10417 CrossRef.
- N. S. Golubev, S. M. Melikova, D. N. Shchepkin, I. G. Shenderovich, P. M. Tolstoy and G. S. Denisov, Z. Phys. Chem., 2003, 217, 1549–1563 Search PubMed.
- K. Sugimori and H. Kawabe, Int. J. Quantum Chem., 2010, 110, 2989–2995 Search PubMed.
- L. J. Yu, E. Golden, N. N. Chen, Y. Zhao, A. Vrielink and A. Karton, Sci. Rep., 2017, 7, 13 Search PubMed.
- L. L. Lin, W. Zhou, R. Gao, S. Y. Yao, X. Zhang, W. Q. Xu, S. J. Zheng, Z. Jiang, Q. L. Yu, Y. W. Li, C. Shi, X. D. Wen and D. Ma, Nature, 2017, 544, 80–81 Search PubMed.
- C. Rivera-Cárcamo, I. C. Gerber, R. I. Del, B. Guicheret, R. C. Contreras, L. Vanoye, A. Favre-Réguillon, B. F. Machado, J. Audevard, C. de Bellefon, R. Philippe and P. Serp, Catal. Sci. Technol., 2021, 11, 984–999 Search PubMed.
- L. Rochette, M. Zeller, Y. Cottin and C. Vergely, Cancers, 2021, 13, 10 CrossRef.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery Jr., J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Revision C01, Gaussian Inc, 2016 Search PubMed.
- R. K. Lindsey, M. P. Kroonblawd, L. E. Fried and N. Goldman, in Computational Approaches for Chemistry under Extreme Conditions, ed. N. Goldman, Springer International Publishing Ag, Cham, 2019, vol. 28, pp. 71–93 Search PubMed.
- A. G. Gale, T. T. Odbadrakh and G. C. Shields, Int. J. Quantum Chem., 2020, 120, e26469 Search PubMed.
- S. E. Harold, S. L. Warf and G. C. Shields, Phys. Chem. Chem. Phys., 2023, 25, 28517–28532 Search PubMed.
- M. Bin Jassar, Q. W. Yao, F. S. Brigiano, W. L. Chen and S. Pezzotti, J. Phys. Chem. Lett., 2024, 15, 11961–11968 CrossRef PubMed.
- J. W. Zheng and O. Lafontant-Joseph, IUPAC Digitized pKa Dataset, v2.3, 2025, DOI:10.5281/zenodo.15375522.
| This journal is © The Royal Society of Chemistry 2026 |