 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Nanoscale protonation limits and charge density in polymer films govern the activity of immobilized LacZ under acid stress
Huida Duan†
a,
Junxing Chen†b,
Felicia Fianua,
Wei Sun
 *b and
Yifan Cheng
*b and
Yifan Cheng
 *a
*a
aVirginia Polytechnic Institute and State University, Department of Food Science and Technology, Blacksburg, VA 24061, USA. E-mail: yifancheng@vt.edu
bVirginia Polytechnic Institute and State University, Department of Biochemistry, Blacksburg, VA 24061, USA. E-mail: sunwei@vt.edu
First published on 24th April 2026
Abstract
Under acidic conditions, polycationic polymer coatings can serve as protective immobilization matrices that buffer local acidity and help preserve enzyme function. However, it remains unclear how polymer support design parameters, particularly film thickness and effective cationic charge density, govern the vital protonation process. Leveraging the nanometer-scale control of film thickness and copolymer composition enabled by initiated chemical vapor deposition (iCVD), we systematically investigated how these parameters govern the protonation behavior of poly[glycidyl methacrylate-co-2-(dimethylamino)ethyl methacrylate] (pGD) thin films and, in turn, the normalized initial ONPG hydrolysis rate of immobilized β-galactosidase protein (LacZ protein). Infrared spectroscopy suggests that proton penetration was capped at a depth of ∼250 nm in pGD with 65 mol% DMAEMA, limiting the polycationic thickness in pGD films thicker than this value. Consistent with this limit, immobilized LacZ activity under acidic stress (pH 4) increased with protonated thickness up to ∼250 nm and then plateaued. Raising the polycationic monomer content from 25 to 65 mol% increased LacZ activity at pH 4 by up to 83%, consistent with a higher positive charge density providing stronger local pH buffering. To test whether this behavior depends on immobilization methods, we evaluated two approaches: random immobilization (via amine-epoxy ring-opening reactions) and site-directed immobilization (via SpyCatcher/SpyTag binding). Directed immobilization preserved higher free LacZ activity than random immobilization, but the protonation-dependent protection trend remained consistent for both strategies. Together, these results identify protonation depth and charge density as orthogonal, tunable design parameters and establish a thickness regime that maximizes protection without unnecessary film growth.
1. Introduction
Immobilized enzymes are foundational tools in biotechnology and industrial biocatalysis, offering advantages such as enhanced stability, recoverability, and recyclability.1–3 Despite these advantages, maintaining the catalytic efficiency of immobilized enzymes under suboptimal environmental conditions, particularly at low pH, remains a critical challenge.4,5 The support material's microenvironment, especially its surface chemistry and morphology, plays a critical role in preserving enzyme activity by regulating local pH, hydrophilicity and conformational stability.6,7 To this end, functional polymer coatings have emerged as versatile platforms that allow functional groups to be tailored for robust enzyme attachment, including covalent anchoring, and microenvironment modulation.8,9 Among covalent strategies, epoxide-functionalized polymers are widely adopted for enzyme immobilization because they form stable covalent linkages with nucleophiles, such as the amines in amino acid side chains on enzyme surfaces.10Meanwhile, pH-responsive polymers, particularly those bearing tertiary amine groups such as poly[2-(dimethylamino)ethyl methacrylate] (pDMAEMA) have shown promise in buffering local microenvironments by changing their ionization state in response to bulk pH shifts.11–13 For instance, pDMAEMA brushes grafted onto surfaces underwent reversible transition between collapsed (neutral) and swollen (protonated) conformations as pH changed, thereby modulating enzyme accessibility and stability.13,14 These pH-responsive polymers are also advantageous due to their tunable charge density, which can be adjusted by controlling the molar fraction of the ionizable groups during polymerization.15–17 Together, these advancements suggest a path toward designing enzyme immobilization platforms that not only anchor enzymes but also tailor their microenvironment, particularly under pH extremes. These pH-modulating enzyme supports may benefit a variety of applications. For example, orally administered lactase must retain sufficient activity after exposure to highly acidic gastric fluids in order to remain effective during gastrointestinal transit, which has motivated the development of protected lactase delivery systems.18 Enzyme-based biosensors used for environmental and water-quality monitoring likewise operate in sample matrices where pH can vary and may compromise sensor performance.19,20 In addition, immobilized-enzyme reactors and porous biocatalyst particles frequently develop local pH gradients that differ from the bulk phase, which can reduce catalytic rates and long-term stability unless the support is engineered to dynamically regulate the enzyme microenvironment.21 In the dairy sector, lactose valorization in acid whey provides another relevant setting for robust biocatalysis, because acid whey is a lactose-rich byproduct and its enzymatic upcycling into lactose-hydrolyzed products, galacto-oligosaccharides, and rare sugars depends strongly on maintaining sufficient β-galactosidase activity under process-relevant conditions.22 These examples underscore the potential benefits of developing immobilization supports that actively preserve a favorable local chemical environment.23
Despite the promise of pH-responsive polymers for enzyme immobilization, a fundamental understanding of how polymer thickness and charge density modulate protonation behavior—and how this, in turn, influences biocatalytic performance—remains lacking.24,25 In addition, while both random immobilization (e.g., amine-epoxy ring opening) and (site-)directed immobilization (e.g., SpyTag/SpyCatcher binding) strategies are established,26 studies integrating site-specific orientation with charge-tunable polymer and evaluating performance under acidic conditions remain limited. Clarifying these relationships would enable more effective local pH modulation and improved activity retention of immobilized enzymes under acidic conditions. Moreover, identifying the minimum polycationic-support thickness required for enzyme protection would guide cost-effective support design.19,20
In this study, we derived design rules for pH-responsive polymer supports by independently tuning film thickness and positive charge density. We copolymerized glycidyl methacrylate (GMA, “G”) and 2-(dimethylamino)ethyl methacrylate (DMAEMA, “D”) to form pGMA-co-DMAEMA (pGD) thin films, where G supplies epoxide groups for covalent protein conjugation27,28 and D supplies tertiary amines that protonate under pH < pKa (∼8.4) to generate positive charge.7 To precisely control thickness and copolymer composition, we synthesized these supports using initiated chemical vapor deposition (iCVD), an all-dry, solvent-free vapor-deposition polymerization method.29,30 This method enables the formation of highly conformal polymer coatings on complex and high–aspect ratio substrates due to its vapor-phase, surface-limited polymerization mechanism.31 During the iCVD process, monomer and initiator vapors (typically tert-butyl peroxide, TBPO, as the initiator) are metered into a reactor held at moderate vacuum (commonly ∼0.1–1 Torr).32 Monomers physisorb on a cooled substrate (often ∼20–40 °C), while TBPO thermally decomposes to generate radicals that initiate surface-confined chain-growth polymerization.33 As a result, iCVD enables nanometer-scale thickness control and high retention of monomer functional groups (e.g., epoxides and tertiary amines) under mild-temperature conditions.
In addition to iCVD-synthesized polycationic supports, we incorporated self-assembled monolayers (SAMs) bearing epoxysilane and tertiaryaminosilane, which are chemically analogous to the iCVD functional groups, in order to access the ultrathin limit of a single molecular layer (typically ∼0.5–3 nm, depending on chain length and substrate).34,35 SAMs are a well-established, solution-phase surface modification approach for presenting reactive groups and mediating covalent enzyme immobilization,34,36 and here they provide an extreme-thickness benchmark for comparison with tens to hundreds of nm iCVD films.
Combining iCVD films and silane-based SAMs, we fabricated chemically parallel polycationic immobilization supports with well-defined thicknesses spanning ∼1 to 800 nm to systematically examine protonation behavior and enzyme activity at pH 4. To probe the interplay between film thickness and charge density, we varied the DMAEMA content in p(GMA-co-DMAEMA) films (25 mol% and 65 mol%). We hypothesized that proton penetration becomes self-limiting above a critical thickness and this effect, in turn, may modulate local pH buffering and the protection of immobilized enzymes under acidic stress. Overall, the novelty of this work lies in decoupling two immobilization-support-level design variables—effective protonated thickness and fixed positive charge density—and linking them directly to the catalytic performance of immobilized LacZ under acidic stress. Moreover, unlike prior studies that have typically examined pH-responsiveness or enzyme orientation separately, this study probes their interplay. Leveraging the precise compositional and thickness control afforded by iCVD, with SAMs as low-thickness controls, this work defines a protonation limit in chemically matched polymer supports, establishes the impact of this limit on enzyme protection under acid stress, and compares LacZ specific activity under random and directed immobilization on the same acid-responsive platform.
2. Materials and methods
2.1 Nomenclature
In this study, we employed four monomers: glycidyl methacrylate (GMA) and N,N-dimethylaminoethyl methacrylate (DMAEMA) for iCVD, and 3-glycidylpropyl trimethoxysilane (GPTMS) and 3-(dimethylamino)propyltrimethoxysilane (DMAPTMS) for SAM coatings. We adopted a concise naming convention in which “G” denotes an epoxide-bearing monomer (GMA or GPTMS) and “D” a tertiary amine monomer (DMAEMA or DMAPTMS), while a leading lowercase “p” signifies an iCVD synthesized thin film and a leading uppercase “S” signifies a SAM monolayer; thus, for example, pGD refers to an iCVD film copolymerized from GMA and DMAEMA and SGD to a SAM composed of GPTMS and DMAPTMS.2.2 Chemical reagents
All reagents were used as received: the iCVD monomers glycidyl methacrylate (GMA, 97.0% purity with ∼0.01% hydroquinone monomethyl ether stabilizer), N,N-dimethylaminoethyl methacrylate (DMAEMA, 98% purity with 700–1000 ppm monomethyl ether hydroquinone inhibitor) and the initiator tert-butyl peroxide (TBPO, 98% purity) were obtained from Sigma-Aldrich Inc. (St Louis, MO, USA), while the SAM reagents 3-glycidylpropyl trimethoxysilane (GPTMS, 98%) and 3-(dimethylamino)propyl trimethoxysilane (DMAPTMS, 96%) were procured from TCI America via Fisher Scientific Company LLC (Suwanee, GA, USA).2.3 Preparation of iCVD-derived polycationic nanolayer supports
Polymers were deposited via iCVD to serve as LacZ immobilization supports. Glycidyl methacrylate (GMA) introduced epoxy groups for covalent enzyme attachment and N,N-dimethylaminoethyl methacrylate (DMAEMA) that impart pH-responsive buffering capacity to the coating. tert-Butyl peroxide (TBPO) was used as the radical initiator, in line with established iCVD polymerization processes.37 Liquid monomers and initiator were loaded in excess (>10 mL) to ensure a continuous, steady vapor supply throughout the deposition duration. The deposition process was controlled by regulating vapor flow rates using needle valves for the monomers and a mass flow controller (MKS Instruments, Inc.) for the initiator. All precursors were vaporized and metered into the reactor at a combined flow of 3–5 sccm, with TBPO held constant at 0.6 sccm. The substrate stage was maintained at 35 °C under a total chamber pressure of 0.6 Torr. A resistively heated filament array at 220 °C generated the free radicals necessary to initiate polymerization. Corning™ polystyrene 96-well plates were mounted on a custom aluminum platen for efficient thermal coupling to the reactor's cooled stage, and a silicon slide atop the wells allowed in situ thickness monitoring via a He–Ne interferometer (350–700 nm; Thorlabs Inc., Newton, NJ, USA). Deposition parameters yielded 12–200 nm coatings on the Si slide (Pm/Psat ≈ 0.01). Considering the quadratic dependence of deposition rate on monomer partial pressure at low Pm/Psat (<0.04 for acrylates), corresponding films deposited in 96-well plates where monomer accumulation increases local Pm, are estimated to be 50–800 nm. Detailed gas flow, Pm/Psat ratios, deposition rates and copolymer composition are compiled in Tables S1, S2 and S3.2.4 Fabrication of self-assembled monolayers (SAMs)
To simulate critical functional chemistries of our iCVD films, we formed mixed SAMs from 3-glycidyloxypropyltrimethoxysilane (GPTMS) and 3-(dimethylamino)propyltrimethoxysilane (DMAPTMS) on both 96-well plates and silicon wafers. Substrates were first activated by oxygen plasma (45 W, 500–900 mTorr, 3 min; Harrick Plasma Inc., NY, USA) to generate surface hydroxyl groups for silane coupling. Separately, 1% v/v ethanolic solutions of GPTMS and DMAPTMS were stirred at 650 rpm and 20 °C for 1 h, then combined to yield a mixture containing 35 mol% GPTMS and 65 mol% DMAPTMS—matching the DMAEMA fraction in the pGD(65) iCVD film. Plasma-treated substrates were submerged in the GPTMS/DMAPTMS blend for 18 h at room temperature to allow silanization. Finally, SAM-coated surfaces were rinsed sequentially with ethanol and deionized water and dried under ambient conditions, following established processes.372.5 Ellipsometry
To confirm the iCVD film thickness prior to LacZ immobilization, ex situ measurements were performed on silicon wafer samples using a J.A. Woollam alpha-SE spectroscopic ellipsometer (Lincoln, NE, USA). Data were collected at three incident angles (65°, 70° and 75°) and fit to a Cauchy–Urbach model to extract thickness values. Each sample was measured at three separate locations, and the average thickness was reported to ensure consistency before proceeding with enzyme immobilization and downstream characterizations.2.6 Fourier transform infrared spectroscopy
Fourier-transform infrared spectroscopy (FTIR) analysis was conducted to verify the presence and preservation of characteristic functional groups in the polymer films synthesized by iCVD. The FTIR spectroscopy of all immobilization supports was performed utilizing a Thermo Scientific Nicolet iS50 Model (Austin, TX, USA). To characterize the composition of copolymers, FTIR analysis was conducted in transmission mode. The experiment employed attenuated total reflectance Fourier transform infrared spectroscopy (ATR-FTIR) using the Thermo Scientific Nicolet iS50 Model, which was equipped with a VariGATR grazing angle ATR accessory (GAT-V-N18, Harrick Scientific Products, NY). The incident angle of the ATR accessory, 60°, was utilized in this study. A MCT detector cooled with liquid nitrogen was utilized over the spectral range of 600–4000 cm−1, with a resolution of 4 cm−1. Measurements were averaged over 128 scans to improve the signal-to-noise ratio. Baseline correction was applied by subtracting a background spectrum obtained from an uncoated Si wafer substrate.2.7 Characterization of polymer protonation
The protonation behavior of synthesized polymer films (pGD and SGD) was first examined by ATR-FTIR spectroscopy. Specifically, polymer-coated substrates were immersed in acidic solution (pH 4, 0.1 mM HCl) or basic solution (pH 10, 0.1 mM NaOH) for 5 min, followed by air-drying for 1 min. To quantify the protonation extent across polymer films of varying thicknesses, their FTIR spectra were additionally collected in transmission mode after immersion in acidic solution (pH 4, 0.1 mM HCl) for 5 min followed by air-drying for 1 min.2.8 Bioengineering of enzymes
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g, 4 °C, 30 min), and the supernatants incubated with 5 mL pre-equilibrated HisPur™ Ni-NTA resin (Thermo Scientific) for 1 h at 4 °C with continuous rotation. The resin slurry was transferred onto a Poly-Prep® chromatography column, washed three times with 20 mL wash buffer (20 mM Tris-HCl, pH 8.8, 400 mM NaCl, 20 mM imidazole, 2 mM DTT), and proteins eluted in five fractions of 3 mL each using elution buffer (20 mM Tris-HCl, pH 8.8, 400 mM NaCl, 500 mM imidazole, 2 mM DTT). Eluates were concentrated via Amicon Ultra-15 centrifugal filters (30 kDa cutoff for LacZ variants; 3 kDa for SC), buffer-exchanged into storage buffer (20 mM Tris-HCl, pH 8.8, 400 mM NaCl, 2 mM DTT, 10% glycerol) using Cytiva PD-10 Sephadex G-25M desalting columns, flash-frozen in liquid nitrogen, and stored at −80 °C.
000g, 4 °C, 30 min), and the supernatants incubated with 5 mL pre-equilibrated HisPur™ Ni-NTA resin (Thermo Scientific) for 1 h at 4 °C with continuous rotation. The resin slurry was transferred onto a Poly-Prep® chromatography column, washed three times with 20 mL wash buffer (20 mM Tris-HCl, pH 8.8, 400 mM NaCl, 20 mM imidazole, 2 mM DTT), and proteins eluted in five fractions of 3 mL each using elution buffer (20 mM Tris-HCl, pH 8.8, 400 mM NaCl, 500 mM imidazole, 2 mM DTT). Eluates were concentrated via Amicon Ultra-15 centrifugal filters (30 kDa cutoff for LacZ variants; 3 kDa for SC), buffer-exchanged into storage buffer (20 mM Tris-HCl, pH 8.8, 400 mM NaCl, 2 mM DTT, 10% glycerol) using Cytiva PD-10 Sephadex G-25M desalting columns, flash-frozen in liquid nitrogen, and stored at −80 °C.2.9 Enzyme immobilization
For RI, LacZ-WT solution (200 µL, 0.06 mg mL−1) was added to a Corning 96-well plate previously coated with a polymer layer. The plate was incubated statically at 37 °C for 24 h to achieve covalent attachment via amine-epoxide coupling. DI proceeded in three sequential steps: (i) random immobilization of SC, (ii) blocking of unreacted epoxides with glycine, and (iii) site-specific capture of LacZ-ST via SC/ST interaction. Briefly, SC (200 µL, 0.05 mg mL−1, 2.8 × 10−4 mM) was randomly immobilized under conditions identical to LacZ-WT. Remaining epoxide groups were blocked using 1.0 M glycine solution following Gao et al. (https://doi.org/10.1016/j.foodchem.2022.134199). Subsequently, LacZ-SpyTag solution (0.06 mg mL−1, 1.0 × 10−4 mM) was added, and the plate was incubated at 25 °C with shaking (30 rpm) for 2 h as previously described. After immobilization, residual enzyme solutions were removed, wells were rinsed twice with 1× PBS to remove unbound proteins, and immobilized LacZ was immediately subjected to activity assays and protein quantification.2.10 Protein quantification and enzyme activity assays
Protein quantification of immobilized LacZ variants was performed using a Pierce bicinchoninic acid (BCA) assay kit (Thermo Fisher Scientific, Waltham, MA, USA). Briefly, 25 μL of 0.1 M phosphate buffer (pH 7.0) was added to each enzyme-immobilized well, followed by 200 μL of BCA working reagent. Plates were mixed thoroughly for 30 s on a plate shaker and incubated at 37 °C for 30 min. Protein quantification was performed for all immobilized proteins, including randomly immobilized LacZ-WT, and sequentially for SC and LacZ-ST in directed immobilization. Immobilization yield (IY) was calculated using the formula:| IY = 100 × Cf/Ci% |
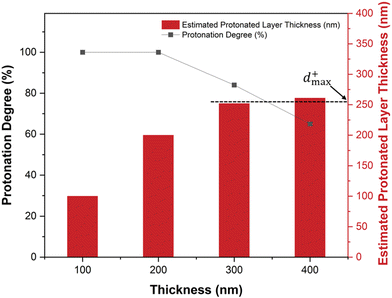
Enzyme activity assays of immobilized and free LacZ were carried out at 37 °C using o-nitrophenyl-β-D-galactopyranoside (ONPG) as substrate. Activity was measured across a pH range using 0.1 M lactate buffers (pH 4) and 0.1 M phosphate buffers (pH 7). Because the absorptivity of ONP/ONP− at 420 nm is pH dependent, activity comparisons in this study are restricted to samples measured within the same pH condition. Prior to the assays, polymer-coated plates with immobilized LacZ were equilibrated at 37 °C for 10–20 min. Reactions were initiated by adding ONPG solutions to a final concentration of 16.6 mM; parallel reaction solutions lacking ONPG were included for background subtraction. The generation of o-nitrophenol was monitored at 420 nm for 5 min using a BioTek Synergy H1 Multi-Mode microplate reader. The normalized initial rate 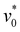 defined as the initial reaction rate measured at 16.6 mM ONPG normalized by the concentration of LacZ, was employed to compare enzyme activity among samples.
defined as the initial reaction rate measured at 16.6 mM ONPG normalized by the concentration of LacZ, was employed to compare enzyme activity among samples.
2.11 Enzyme activity under different protonation depths
To evaluate the influence of polymer protonation depth on the catalytic performance of immobilized enzymes, LacZ activity was measured for both RI and DI configurations on iCVD-synthesized pGD(65) films of different thicknesses (50, 100, 200, 400, 800 nm) and SGD(65) following the procedures of the abovementioned enzyme activity assay.2.12 Statistical analysis
All experiments were performed in triplicate (n = 3). Data analysis and visualization were carried out using RStudio and OriginLab, respectively. Significant differences among treatments were assessed by analysis of variance (ANOVA), followed by Tukey's Honest Significant Difference (HSD) post-hoc test to pinpoint specific differences between groups. Results were reported at a significance level of 0.05 and visualized using compact letter displays to summarize Tukey's test outcomes.3. Results and discussion
3.1 Characterization of immobilization supports and protonation
To illustrate the acid-induced protonation of tertiary amine groups in both pGD and SAM systems (Fig. 1a), we characterized the chemical structures and molar compositions of the thin films synthesized by iCVD, along with the corresponding SAMs, through FTIR spectroscopy (Fig. 1b and c). First, we focused on several characteristic absorption peaks associated with key functional groups relevant to enzyme immobilization and microenvironment modulation. The absorption peak at approximately 900 cm−1 corresponds to epoxide ring-deformation vibrations from the monomer GMA in pGD(65) and from the GPTMS in its SAM counterpart SGD(65).38–40 The presence of this peak confirms the retention of epoxide groups, which serve as critical bioconjugation sites for enzyme attachment. This mechanism is widely reported to facilitate multipoint attachment and enhanced enzyme stability and activity in immobilized systems.41Additionally, a weak absorption around 2770 cm−1 arises from symmetric C–H stretching vibrations of methyl groups linked to nitrogen atoms in tertiary amines. The consistent detection of this peak in both pGD(65) and SGD(65) indicates successful incorporation of tertiary amine groups derived from DMAEMA (iCVD) and DMAPTMS (SAMs).7 Furthermore, a strong carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) stretching vibration at 1730 cm−1 was clearly observed in all iCVD-based polymers but absent in SAMs, confirming successful polymerization and correct composition. The disappearance of absorption peaks around the vinyl C
O) stretching vibration at 1730 cm−1 was clearly observed in all iCVD-based polymers but absent in SAMs, confirming successful polymerization and correct composition. The disappearance of absorption peaks around the vinyl C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C stretch (1650 cm−1) further supports complete monomer conversion during iCVD synthesis. For completeness, the FTIR spectra of the individual monomers (DMAEMA and GMA) together with the resulting pGD(65) film are provided in Fig. S1, where the characteristic vinyl stretching band (1650 cm−1) of the monomers disappears after iCVD deposition, further confirming successful polymerization.
C stretch (1650 cm−1) further supports complete monomer conversion during iCVD synthesis. For completeness, the FTIR spectra of the individual monomers (DMAEMA and GMA) together with the resulting pGD(65) film are provided in Fig. S1, where the characteristic vinyl stretching band (1650 cm−1) of the monomers disappears after iCVD deposition, further confirming successful polymerization.
To confirm protonation of the tertiary amine groups, spectra were compared before and after exposure to acidic (pH 4) and basic (pH 10) solutions. Acid treatment produced two new features: (i) a broad N–H stretch at 3000–3200 cm−1 and (ii) a strong N–H bending band near ∼1600 cm−1, both characteristic of protonated tertiary amines. These features were absent under basic conditions.42 Notably, these functional groups are derived from DMAEMA, whose tertiary amine has a known pKa of ∼8.4.7 In polymerized form (pDMAEMA), the effective pKa is somewhat lower (7.0–7.5) due to local electrostatic and steric effects.43,44 Thus, at pH 4, essentially full protonation is anticipated, in alignment with the observed FTIR features.45 In contrast, the absence of these bands in basic conditions indicates deprotonation and neutrality of tertiary amine groups. In addition, film thickness remained unchanged within experimental error after soaking at pH 4 for 5 min (Table S5), supporting the integrity of the pGD coatings under the acid-exposure conditions used for protonation and activity assays.
In summary, FTIR analyses (Fig. 1b and c) clearly verify the successful incorporation and preservation of epoxide functionalities essential for bioconjugation and ionizable tertiary amine groups crucial for pH-responsive modulation. The acid-induced protonation verified by FTIR further substantiates the pH-responsive capability of our designed polymeric supports. Collectively, these structural and functional confirmations underscore the suitability of our iCVD-synthesized polymer films and SAMs as advanced immobilization platforms.
3.2 Self-limiting proton penetration
To further investigate how protonation varies with polycationic polymer thickness, pGD(65) films with varying thicknesses were immersed in an acidic solution (pH 4, 0.1 mM HCl) for 5 minutes, then air-dried for 1 minute prior to analysis. The resulting FTIR spectra were analyzed (Fig. 2). Specifically, protonation was tracked by the area ratio of the N–H bending vibration peak (∼1600 cm−1) relative to the carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) peak (∼1730 cm−1). As shown in Fig. 2, the protonation ratio increased with thickness up to ∼200 nm, after which it declined for thicker films. This observation suggests that protonation became transport-limited in thicker films within the experimental timeframe, such that the outer film region was protonated more extensively than the interior.
O) peak (∼1730 cm−1). As shown in Fig. 2, the protonation ratio increased with thickness up to ∼200 nm, after which it declined for thicker films. This observation suggests that protonation became transport-limited in thicker films within the experimental timeframe, such that the outer film region was protonated more extensively than the interior.
 | ||
| Fig. 2 FTIR spectra of pGD(65) polymer films with different thicknesses after soaking in acidic solution (pH 4) for 5 min. | ||
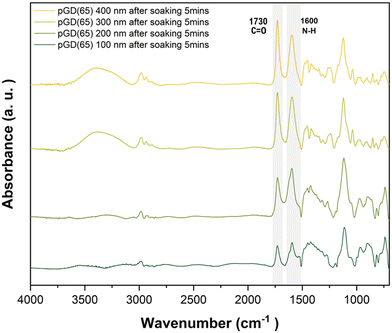
For quantitative comparison, the N–H/C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O area ratio was calculated for each thickness using FTIR spectra collected after immersion in acidic solution (pH 4, 0.1 mM HCl) for 5 min followed by air-drying for 1 min. Each ratio was then normalized to that of the 100 nm pGD(65) film--treated as the “fully protonated” reference--to derive the protonation degree (%). The integrated peak areas and normalized N–H/C=O ratios used for this analysis are summarized in Table S6 and the corresponding normalized protonation percentages for films of varying thicknesses are presented in Fig. 3. The invariant N–H/C=O area ratio between the 100 nm and 200 nm films indicates that both thicknesses reached near-complete protonation, suggesting that protonation capacity was saturated at 100 nm, whereas thicker films (300 nm and 400 nm) showed partial protonation, with the protonation degree decreasing with thickness. Together, these results suggest that the maximum protonation depth likely lies between 200 and 300 nm. To confirm this, the estimated protonated layer thickness was calculated by multiplying the as-synthesized iCVD film thickness, determined by ellipsometry in ambient conditions, by the protonation degree derived from FTIR spectra. The resultant estimated protonated thickness plateaued ∼250 nm, which we define here as the maximum proton penetration depth (dmax+) under the present exposure conditions, beyond which additional thickness no longer contributes to the protonated volume. In this estimation, swelling of the film under aqueous conditions was not explicitly taken into account.
O area ratio was calculated for each thickness using FTIR spectra collected after immersion in acidic solution (pH 4, 0.1 mM HCl) for 5 min followed by air-drying for 1 min. Each ratio was then normalized to that of the 100 nm pGD(65) film--treated as the “fully protonated” reference--to derive the protonation degree (%). The integrated peak areas and normalized N–H/C=O ratios used for this analysis are summarized in Table S6 and the corresponding normalized protonation percentages for films of varying thicknesses are presented in Fig. 3. The invariant N–H/C=O area ratio between the 100 nm and 200 nm films indicates that both thicknesses reached near-complete protonation, suggesting that protonation capacity was saturated at 100 nm, whereas thicker films (300 nm and 400 nm) showed partial protonation, with the protonation degree decreasing with thickness. Together, these results suggest that the maximum protonation depth likely lies between 200 and 300 nm. To confirm this, the estimated protonated layer thickness was calculated by multiplying the as-synthesized iCVD film thickness, determined by ellipsometry in ambient conditions, by the protonation degree derived from FTIR spectra. The resultant estimated protonated thickness plateaued ∼250 nm, which we define here as the maximum proton penetration depth (dmax+) under the present exposure conditions, beyond which additional thickness no longer contributes to the protonated volume. In this estimation, swelling of the film under aqueous conditions was not explicitly taken into account.
 | ||
| Fig. 3 Normalized protonation degree (%) and estimated protonated layer thickness of pGD(65) films with varying thickness after soaking in pH 4 solutions, calculated from FTIR peak area ratios. | ||
In the present pGD system, this thickness-dependent behavior seems consistent with the proton-penetration-limited regime under the present experimental conditions. In hydrated weak-polybase films, proton uptake is expected to arise from a coupled diffusion-reaction process, in which protons migrate through water-accessible regions of the polymer while tertiary amines are protonated along the transport pathway. In pGD, film thickness changed little upon protonation (Table S5), indicating that swelling was not a major feature of the present system. Accordingly, the apparent proton penetration depth in pGD is more likely governed by coupled positive charge buildup and the film's transport landscape. That transport landscape is defined by water-accessible free volume, local chain packing, segmental mobility, and, where relevant, the degree of structural order as well as the connectivity and tortuosity of transport pathways.46,47 These structural features can strongly influence local proton diffusivity and determine whether the outer protonated region becomes kinetically resistive to further inward transport. In weak polyelectrolyte layers, protonation behavior is also known to depend on ionic strength, ionizable-group fraction, and polymer hydrophobicity,48–50 while neutron-reflectivity studies further show that pH-responsive polymer brushes can exhibit nonuniform internal density profiles near interfaces.51 We therefore interpret the ∼250 nm protonation plateau observed here as most likely arising from the combined effects of coupled diffusion, protonation, progressive positive charge buildup, and film structure under the present experimental conditions, rather than from electrostatic repulsion alone.52 Because free volume, porosity, and structural order were not independently measured in this study, these features are treated as plausible contributors to transport limitation rather than directly established properties of pGD.
3.3 Effect of protonated layer thickness on immobilized enzyme activity
Building upon the FTIR and quantitative analyses described above, which revealed a protonation depth of ∼250 nm under acidic conditions, we next investigated how this protonation depth influences the catalytic performance of immobilized LacZ. Fig. 4a–d show the normalized initial rates (v0*, at 16.6 mM ONPG) of LacZ immobilized via random immobilization (RI) and directed immobilization (DI), along with that of soluble, surface-free LacZ as control, under both neutral (pH 7) and acidic (pH 4) conditions. Initial reaction rates were normalized to the LacZ molar concentration in the reaction volume, as determined by BCA assay. This normalization step isolates the intrinsic response of individual enzymes to different microenvironments, effectively decoupling it from potential extensive effects caused by difference in enzyme loading.Under neutral conditions (pH 7), randomly immobilized LacZ retained only ∼25% of the catalytic activity of the free enzyme, suggesting that random covalent attachment to surface epoxides imposed conformational constraints and steric hindrance, thereby limiting substrate accessibility.53 However, when LacZ was immobilized site-specifically through the SpyTag/SpyCatcher conjugation of DI, the activity was significantly preserved, closely matching that of free enzyme controls. This result underscores the advantage of DI over RI, highlighting that spatial orientation control substantially enhances enzyme function post-immobilization by reducing structural perturbation. Notably, across a support-thickness range of ∼1 to 800 nm, immobilized LacZ activity did not differ significantly (p > 0.05) for either DI or RI. Because protonation is negligible at pH 7, these neutral-pH data serve as a thickness-matched control free of electrostatic and local pH-buffering effects, demonstrating that thickness-dependent properties (e.g., mechanical compliance) exert minimal influence on immobilized LacZ activity regardless of immobilization strategy.
Under acidic conditions (pH 4), the free LacZ showed no detectable catalytic activity, consistent with prior reports that E. coli LacZ exhibits maximal activity near neutral pH and strongly reduced activity and stability under acidic conditions.54 Remarkably, LacZ immobilized even on the SGD(65) monolayer exhibited measurable catalytic activity, indicating that a single molecular layer of tertiary amines can confer electrostatic proton shielding and/or local pH buffering around the enzyme. Enzyme activity increased monotonically with support thickness from ∼1 nm (SGD(65)) to 200 nm (pGD(65)), with a more pronounced trend for DI than for RI, reflecting greater activity retention in DI. Neither immobilization yield nor surface enzyme density increased monotonically with support thickness (Tables S7 and S8), indicating that the thickness-dependent activity trend under acidic conditions was not associated with enzyme loading. Beyond a polymer thickness of ∼200 nm (up to 800 nm), no further increase in activity was observed at pH 4 for either immobilization strategy. The slightly different mean value observed for the 800 nm film under RI or DI (Fig. 4b,d) does not indicate a renewed thickness dependence; rather, it remains within the same plateau regime as the 200 and 400 nm samples, consistent with the conclusion that only the outer ∼250 nm of the support contributes to protonation-mediated protection, while the additional inner thickness is effectively inactive with respect to acid buffering. This plateau aligns with the protonation-depth limit determined earlier (section 3.2), where polymer layers thicker than ∼250 nm could not undergo additional protonation, indicating a finite proton-penetration depth. Consequently, the activity plateau arises from the saturation of polymer protonation rather than polymer thickness itself—once this limit is reached, additional film thickness offers negligible improvement in local buffering or enzyme protection.
Fig. 4e summarizes a two-regime conceptual model defined by the relationship between the total thickness of the polycationic support, h, and the maximum protonation depth, dmax+. When the film is thinner than the protonation depth (h < dmax+), protons can fully penetrate the support and convert tertiary amines into protonated tertiary ammonium groups, placing the system in a total-thickness-limited regime (top row). In this regime, very thin supports (e.g., SGD) provide weak proton shielding, whereas increasing h increases the amount of protonated polycationic material and correspondingly enhances LacZ activity under acidic stress.
As the film thickness approaches the protonation limit (h ≈ dmax+; middle row), the entire support becomes protonated. Further increases in h no longer increase the protonated thickness, defining an optimal or “sweet-spot” thickness that maximizes enzyme protection while minimizing coating material usage and deposition time.
When the support thickness exceeds the protonation depth (h > dmax+; bottom row), the system enters a proton-penetration-limited regime, in which the protonated layer thickness remains constant despite additional film growth. Consequently, further increases in total thickness do not improve the acid resistance of immobilized LacZ.
Collectively, these results explicitly demonstrate that the protonation depth, rather than the absolute polymer thickness, determines the extent to which local buffering and proton shielding can preserve enzyme activity under acidic conditions. Controlling polymer thickness within the protonation saturation limit (around 200–300 nm) emerges as a useful strategy for optimizing proton shielding for relieving acidic stress.
3.4 Influence of polycationic monomer density on immobilized enzyme activity
Beyond polymer thickness, the molar ratio of protonatable monomers within polymer formulations could influence the catalytic activity of immobilized enzymes. To systematically investigate this effect, we synthesized copolymers with different compositions of DMAEMA monomer, specifically pGD(25) and pGD(65), representing lower and higher polycationic monomer ratio. Enzyme activities were measured across polymer films of varying thicknesses (200 nm, 400 nm, 800 nm), and results are summarized in Fig. 5. | ||
| Fig. 5 Normalized activities of immobilized LacZ on polymer films with varying thicknesses and compositions, evaluated at pH 4. | ||
Fig. 5 shows that for a given polymer composition (either pGD(25) or pGD(65)), no statistically significant difference in enzyme activity was observed among films of different thicknesses. This outcome is consistent with previous observations in Fig. 4 that, beyond a certain thickness (where protonation saturation occurs), further increases in polymer thickness alone do not influence enzyme activity. However, when we compare enzyme activities immobilized on polymers with different polycationic compositions, a pronounced difference emerged. Specifically, enzyme activity on pGD(65) was significantly higher, 83% greater—than on pGD(25) at all tested thicknesses. This notable increase can be attributed to the substantially higher density of protonatable tertiary amine groups present in pGD(65): the increased DMAEMA content in pGD(65) generates a higher local positive charge density, offering more effective electrostatic repulsion and stronger local buffering capability. Consequently, enzyme immobilized on pGD(65) experienced substantially less acidic stress compared to those on lower-DMAEMA-content polymers such as pGD(25).
From an enzyme catalytic mechanism perspective, this protection likely arises because the polycationic support helps preserve a more favorable local ionization environment around the LacZ active site under acidic bulk conditions. E. coli LacZ protein follows a double-displacement mechanism involving two essential glutamate residues, Glu461 and Glu537, which function as the general acid/base catalyst and nucleophile, respectively.55–58 A schematic summary of the LacZ catalytic residues and their expected acid sensitivity is provided in Fig. S2. Under strongly acidic conditions, excessive proton activity is expected to suppress the catalytic roles of these carboxylate residues, thereby impairing ONPG hydrolysis.55–58 In addition to this active-site effect, our Circular Dichroism (CD) measurements showed a marked loss of ellipticity at 208 and 222 nm at pH 4, consistent with reduced α-helical content and partial structural destabilization (SI Methods). The corresponding CD spectra are shown in Fig. S3. These observations suggest that acid-induced deactivation of LacZ arises from both catalytic-residue protonation and global conformational disruption. In this context, the protonated DMAEMA-containing support does not directly activate the enzyme; rather, it likely attenuates proton influx and buffers the local environment surrounding the immobilized enzyme, helping maintain both catalytic competence and structural integrity.21 Greater protonated thickness and charge density each contribute to this protective effect.
The practical implication of this result is that, once the film is sufficiently thick to establish a protonated interfacial buffering layer, increasing charge density becomes a more effective design parameter than further increasing total thickness. In this context, the 83% enhancement in LacZ activity observed for pGD(65) relative to pGD(25) is not merely incremental but functionally significant, as it reflects an orhtogonal design strategy for recovering catalytic performance under acidic conditions (pH 4), where unprotected enzymes typically exhibit severely diminished activity. Importantly, this improvement is achieved without increasing film thickness, which would otherwise prolong iCVD deposition time and raise material costs—both are detrimental to economic viability at scale. Because only two DMAEMA levels were tested, the present data establish the trend but do not define its upper limit. Increasing %D further may improve acid protection, but likely with a tradeoff in enzyme loading as %G decreases. Building on the mechanistic insights from this work, future work may optimize composition to maximize total catalytic activity per support area or mass, providing a more pratically relevant performance metric. Taken together, these results show that enriching protonatable amine content provides an complementary acid-protection route to increasing thickness, providing another degree of freedom to balance catalytic performance with practical constraints such as fabrication throughput, material usage, and substrate diffusion.
More broadly, these findings suggest a generalizable support-design principle for acid-stressed biocatalysis: a minimum thickness is first required to establish a protonated interfacial region, beyond which further performance gains are more effectively achieved by tuning local charge density rather than adding inactive material. This strategy is directly relevant to systems where enzymes must retain activity in acidic or pH-variable environments, including oral lactase delivery through gastric conditions, biosensing platforms operating in complex or unbuffered samples, immobilized-enzyme reactors where local pH gradients can arise during catalysis, and LacZ-driven lactose valorization in acid whey and related biomanufacturing processes, where a mismatch often exists between the bulk feed conditions and the optimal pH window of the enzyme.
4. Conclusion
This study provides a framework for understanding the effect of protonation depth and charge density on polycationic polymer support, and in turn, on enzyme activity under acidic conditions. By combining iCVD and SAM immobilization supports with precisely controlled thickness and tunable copolymer composition, we systematically investigated how film thickness and the molar ratio of protonatable monomers (DMAEMA) influence the spatial distribution and extent of protonation.We demonstrated that at pH 4 protonation in pGMA-co-DMAEMA (DMAEMA ∼ 65 mol%) films is confined to a finite nanoscale depth of ∼250 nm, which we term dmax+, a material parameter that dictates the upper limit of enzymatic protection under the tested conditions. A two-regime model can be defined based on dmax+: when protonatable film thickness is smaller than dmax+, the system is in a total-thickness-limited regime; when protonatable film thickness is greater than dmax+, the system is placed in a proton-penetration-limited regime. By correlating this depth with the activity of immobilized LacZ, we showed that matching film thickness to this saturation limit is essential for efficient design: dmax+ defines an optimal or “sweet-spot” thickness that maximizes enzyme protection while minimizing coating material usage and deposition time. Furthermore, while site-directed immobilization offers better absolute performance by preserving enzyme orientation, the protective buffering mechanism is eventually governed by the polymer's charge density and protonation depth. These findings shed light on protonation depth and charge density as physicochemical design parameters for developing functional polycationic immobilization supports for enzyme-based biocatalysis, biosensors, and biomedical systems.
Conflicts of interest
There are no conflicts to declare.Data availability
All data supporting the results of this study are provided in the supplementary information (SI) accompanying this article. Supplementary information: iCVD process parameters, primer information, additional experimental measurements, e.g., circular dichroism spectra, ONP standard curves at pH 4 and 7, enzyme immobilization yields, polymer characterization data, and any other datasets analyzed during this work. See DOI: https://doi.org/10.1039/d5nr05409e.Acknowledgements
Y. C. acknowledges the support of the Foundation for Food and Agriculture Research (FFAR) New Innovator Award (23-000576) and award No. USDA NIFA 2021-67034-35040 (2021-67034-39810) and start-up funds provided by Virginia Tech for this work. W. S. acknowledges support from start-up funds provided by Virginia Tech. J. C. acknowledges support from the Virginia Tech Biochemistry Graduate Program. The authors thank members of the Cheng and Sun laboratories for technical assistance and constructive feedback.References
- S. M. Mirsalami, M. Mirsalami and A. Ghodousian, Results Chem., 2024, 7, 101486 Search PubMed.
- F. Asaduzzaman and S. Salmon, Mol. Syst. Des. Eng., 2022, 7, 1385–1414 Search PubMed.
- J. M. Guisan, G. Fernandez-Lorente, J. Rocha-Martin and D. Moreno-Gamero, Curr. Opin. Green Sustainable Chem., 2022, 35, 100593 CrossRef CAS.
- Y. R. Maghraby, R. M. El-Shabasy, A. H. Ibrahim and H. M. E. S. Azzazy, ACS Omega, 2023, 8, 5184–5196 CrossRef CAS PubMed.
- M. S. Robescu and T. Bavaro, Molecules, 2025, 30(4), 939 Search PubMed.
- M. G. Holyavka and V. G. Artyukhov, Biophys. Rev., 2025, 1–39 Search PubMed.
- P. Cotanda, D. B. Wright, M. Tyler and R. K. O'Reilly, J. Polym. Sci., Part A:Polym. Chem., 2013, 51, 3333–3338 Search PubMed.
- W. A. A. Wahab, Microb. Cell Fact., 2025, 24, 167 Search PubMed.
- X. Lyu, R. Gonzalez, A. Horton and T. Li, Catalysts, 2021, 11, 1211 CrossRef CAS.
- T. Prabhakar, J. Giaretta, R. Zulli, R. J. Rath, S. Farajikhah, S. Talebian and F. Dehghani, Chem. Eng. J., 2025, 503, 158054 CrossRef CAS.
- G. Kocak, C. Tuncer and V. Bütün, Polym. Chem., 2017, 8, 144–176 Search PubMed.
- J. Zdarta, A. S. Meyer, T. Jesionowski and M. Pinelo, Catalysts, 2018, 8(2), 92 CrossRef.
- H. Zhou, X. Wang, J. Tang and Y. W. Yang, Polymers, 2016, 8, 277 Search PubMed.
- S. N. H. Ishak, A. H. M. Saad, W. Latip, R. N. Z. R. A. Rahman, A. B. Salleh, N. H. A. Kamarudin, A. T. C. Leow and M. S. M. Ali, Int. J. Biol. Macromol., 2025, 316, 144278 CrossRef CAS PubMed.
- A. Vardaxi and S. Pispas, Polymers, 2023, 15, 1519 CrossRef CAS.
- X. Fu, L. Hosta-Rigau, R. Chandrawati and J. Cui, Chem, 2018, 4, 2084–2107 CAS.
- J. Niskanen, C. Wu, M. Ostrowski, G. G. Fuller, S. Hietala and H. Tenhu, Macromolecules, 2013, 46, 2331–2340 CrossRef CAS.
- K. C. Kwon, D. Verma, N. D. Singh, R. Herzog and H. Daniell, Adv. Drug Delivery Rev., 2013, 65, 782–799 Search PubMed.
- A. P. F. Turner, Chem. Soc. Rev., 2013, 42, 3184 Search PubMed.
- B. L. Vallee and D. D. Ulmer, Annu. Rev. Biochem., 1972, 41, 91–128 Search PubMed.
- R. A. Sheldon and S. van Pelt, Chem. Soc. Rev., 2013, 42, 6223–6235 RSC.
- P. Kolev, D. Rocha-Mendoza, S. Ruiz-Ramírez, J. Ortega-Anaya, R. Jiménez-Flores and I. García-Cano, JDS Commun., 2022, 3, 1–6 CrossRef.
- C. Mateo, J. M. Palomo, G. Fernandez-Lorente, J. M. Guisan and R. Fernandez-Lafuente, Enzyme Microb. Technol., 2007, 40, 1451–1463 CrossRef CAS.
- S. Tippner, D. Hernández-Castillo, F. H. Schacher and L. González, J. Phys. Chem. B, 2025, 129, 2105–2114 CrossRef CAS PubMed.
- X. Laloyaux, B. Mathy, B. Nysten and A. M. Jonas, Macromolecules, 2010, 43, 7744–7751 CrossRef CAS.
- X. Pei, Z. Luo, L. Qiao, Q. Xiao, P. Zhang, A. Wang and R. A. Sheldon, Chem. Soc. Rev., 2022, 51, 7281–7304 RSC.
- M. Gupta and K. K. Gleason, Thin Solid Films, 2006, 515, 1579–1584 CrossRef CAS.
- Y. H. Youn, S. J. Lee, G. R. Choi, H. R. Lee, D. Lee, D. N. Heo, B. S. Kim, J. B. Bang, Y. S. Hwang, V. M. Correlo, R. L. Reis, S. G. Im and I. K. Kwon, Mater. Sci. Eng., C, 2019, 100, 949–958 CrossRef CAS PubMed.
- K. K. Gleason, Nat. Rev. Phys., 2020, 2, 347–364 CrossRef CAS.
- K. K. Gleason, Adv. Mater., 2024, 36(8), 2306665 CrossRef CAS PubMed.
- K. Park, K. K. Gleason and R. Yang, Adv. Funct. Mater., 2025, 35(24), 2417620 CrossRef CAS.
- Y. Cheng, A. Khlyustova, P. Chen and R. Yang, Macromolecules, 2020, 53, 10699–10710 CrossRef CAS.
- K. K. S. Lau and K. K. Gleason, Macromolecules, 2006, 39, 3688–3694 CrossRef CAS.
- I. Luzinov, D. Julthongpiput, A. Liebmann-Vinson, T. Cregger, M. D. Foster and V. V. Tsukruk, Langmuir, 2000, 16, 504–516 CrossRef CAS.
- J. C. Love, L. A. Estroff, J. K. Kriebel, R. G. Nuzzo and G. M. Whitesides, Chem. Rev., 2005, 105, 1103–1170 CrossRef CAS.
- S. Libertino, F. Giannazzo, V. Aiello, A. Scandurra, F. Sinatra, M. Renis and M. Fichera, Langmuir, 2008, 24, 1965–1972 CrossRef CAS PubMed.
- J. Pinson and D. Thiry, Surface modification of polymers: methods and applications, Wiley-VCH, 2019 Search PubMed.
- I. D. Vukoje, E. S. Džunuzović, V. V. Vodnik, S. Dimitrijević, S. P. Ahrenkiel and J. M. Nedeljković, J. Mater. Sci., 2014, 49, 6838–6844 CrossRef CAS.
- M. Sobiesiak, Adsorption, 2019, 25, 257–266 CrossRef CAS.
- S. Ü. Çelik and A. Bozkurt, Solid State Ion., 2011, 199, 1–5 CrossRef.
- C. Mateo, V. Grazú, G. Grazú, B. C. C. Pessela, T. Montes, J. M. Palomo, R. Torres, F. L. Ópez-Gallego, R. Fernández, F. Fernández-Lafuente, J. M. Guisán and G. Guisán, Biochem. Soc. Trans., 2007, 35, 1593–1601 CrossRef CAS PubMed.
- B. Smith, Spectroscopy, 2019, 34, 30–37 Search PubMed.
- E. W. Kent, E. M. Lewoczko and B. Zhao, Polym. Chem., 2021, 12, 265–276 RSC.
- J. C. Y. Wu, C. H. Hutchings, M. J. Lindsay, C. J. Werner and B. C. Bundy, J. Biotechnol., 2015, 193, 83–90 CrossRef CAS PubMed.
- R. A. Heacock and L. Marion, Can. J. Chem., 1956, 34, 782–1795 CrossRef.
- J. Sharma, K. Tewari and R. K. Arya, Prog. Org. Coat., 2017, 111, 83–92 CrossRef CAS.
- A. Mansuri, M. Völkel, T. Feuerbach, J. Winck, A. W. P. Vermeer, W. Hoheisel and M. Thommes, Macromolecules, 2023, 56, 3224–3237 CrossRef CAS.
- J. D. Willott, T. J. Murdoch, B. A. Humphreys, S. Edmondson, G. B. Webber and E. J. Wanless, Langmuir, 2014, 30, 1827–1836 CrossRef CAS PubMed.
- S. Ramezani Bajgiran, F. Safi Samghabadi, S. Li, J. C. Conrad and A. B. Marciel, Macromolecules, 2023, 56, 9218–9228 CrossRef CAS.
- J. D. Willott, B. A. Humphreys, T. J. Murdoch, S. Edmondson, G. B. Webber and E. J. Wanless, Phys. Chem. Chem. Phys., 2015, 17, 3880–3890 RSC.
- M. Moglianetti, J. R. P. Webster, S. Edmondson, S. P. Armes and S. Titmuss, Langmuir, 2010, 26, 12684–12689 CrossRef CAS PubMed.
- G. Ferrand-Drake del Castillo, R. L. N. Hailes and A. Dahlin, J. Phys. Chem. Lett., 2020, 11, 5212–5218 CrossRef CAS PubMed.
- M. J. Paiva, F. C. Paula-Elias, L. A. Pereira, S. C. Carreiro, E. C. Vieira-Almeida, E. M. Silva, G. S. Dias, M. A. C. Xavier, S. A. Morales, R. F. Perna and A. F. Almeida, J. Braz. Chem. Soc., 2023, 34, 1743–1752 CAS.
- A. A. Hamed, M. Khedr and M. Abdelraof, J. Genet. Eng. Biotechnol., 2020, 18, 80 CrossRef PubMed.
- J. Yuan, M. Martinez-Bilbao and R. E. Huber, Biochem. J., 1994, 299, 527–531 CrossRef CAS PubMed.
- J. C. Gebler, R. Aebersold and S. G. Withers, J. Biol. Chem., 1992, 267, 11126–11130 CrossRef CAS.
- J. P. Richard, R. E. Huber, S. Lin, C. Heo and T. L. Amyes, Biochemistry, 1996, 35, 12377–12386 CrossRef CAS PubMed.
- C. G. Cupples, J. H. Miller and R. E. Huber, J. Biol. Chem., 1990, 265, 5512–5518 CrossRef CAS PubMed.
Footnote |
| † Equal contributions. |
| This journal is © The Royal Society of Chemistry 2026 |




