Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Nanoscale topology of γH2AX and 53BP1 foci in U87 cancer cells and normal NHDF after high-LET radiation-induced DSB repair
Kim Annabel
Küntzelmann
a,
Laura Rozo
Pardo
 a,
Myriam
Schäfer
a,
Jonas
Weidner
a,
Iva
Falkova
b,
Jiri
Toufar
b,
Lucie
Toufarova
b,
Felix
Bestvater
c,
Michael
Hausmann†
a,
Myriam
Schäfer
a,
Jonas
Weidner
a,
Iva
Falkova
b,
Jiri
Toufar
b,
Lucie
Toufarova
b,
Felix
Bestvater
c,
Michael
Hausmann†
 *a and
Martin
Falk†
*a and
Martin
Falk†
 *b
*b
aKirchhoff Institute for Physics, Heidelberg University, 69120 Heidelberg, Germany. E-mail: hausmann@kip.uni-heidelberg.de
bInstitute of Biophysics, Czech Academy of Sciences, 612 00 Brno, Czech Republic. E-mail: falk@ibp.cz
cGerman Cancer Research Center (DKFZ), Heidelberg, Germany
First published on 9th January 2026
Abstract
DNA damage repair is essential for maintaining genomic integrity, thereby preventing diseases like cancer. Traditionally, radioresistance has been linked to the ability of cells to repair DNA double-strand breaks (DSBs) accurately. Recent research emphasizes the critical role of spatial chromatin organization and its dynamic reorganization in regulating repair and gene expression. In this study, we have employed single-molecule localization microscopy (SMLM) and Python-based mathematical methods of statistics and topology to locally analyze the spatial organization of γH2AX and 53BP1 foci in 15N-ion irradiated normal human dermal fibroblasts (NHDF) and highly radioresistant U87 glioblastoma cells over extended post-irradiation (PI) periods with nanoscale resolution. Our findings reveal that U87 cancer cells fail to regulate chromatin changes at DSB sites during and after repair. Specifically, Ripley's statistics and cluster analysis showed that both NHDF and U87 cells exhibit smaller, denser, and better separated γH2AX nano-foci surrounded by 53BP1 nano-foci. Mathematical topology approaches, including persistent homology, revealed that γH2AX nano-foci (clusters) have lower topological similarity compared to the more conserved 53BP1 nano-foci during the 24-hour repair period. These findings support the non-random, functional spatial organization of DSB repair (nano)foci and demonstrate its preservation in cancer cells. However, principal component analysis of persistent images showed that γH2AX- and 53BP1 nano-foci in normal fibroblasts exhibit stable, closed cycles, while U87 cells display chaotic, open shifts in topology in the 2D latent space. Combined with DSB repair kinetics measurements, this observation indicates that although U87 cells rejoin DSBs similar to normal cells, they experience more pronounced, dysregulated chromatin alterations during repair, ultimately failing to restore it to its pre-irradiation state. These alterations correlate with topologically more variable DSB sites, slower (more challenging) repair focus formation but faster repair once foci are established, compared to NHDF cells. More disorganized repair and persistent topological alterations likely contribute to genetic instability of cancer cells after irradiation and the development of radioresistant clones, posing challenges for effective radiotherapy.
Introduction
Chromatin in cell nuclei is constantly subjected to damage from environmental factors, as well as from metabolic processes essential for cell function. DNA/chromatin damage repair is thus crucial for maintaining the genomic integrity of the cell and preventing the development of cancer and other genetic diseases. Repair of chromatin double-strand breaks (DSBs), the most harmful DNA damage ionizing radiation can induce, is particularly challenging for the cell and the cellular repair mechanisms. In particular, for complex and clustered DSBs caused, for instance, by high-LET (linear energy transfer) radiation,1–4 cell nuclei are challenged to repair and finally reconstitute their genome.5 High-LET radiation thus poses the number one health risk for astronauts during interplanetary space missions.6 On the other hand, high-LET radiation treatment,7–9 especially by proton10 and C-ions,11 is a very promising and highly efficient anti-cancer therapy, especially for slowly growing brain tumors where other radiation therapies fail.These examples illustrate that understanding the mechanisms underlying cellular responses to ion radiation exposure is a key focus of contemporary radiobiological research. Densely ionizing particles cause damage responses and outcomes that are fundamentally different from those induced by sparsely ionizing radiation.12–15 Moreover, chromatin damage is complex since it forms clusters, which occurs within the limited volume of channel-like trajectories and cannot be separated into distinct formations.
DSB repair has been studied in terms of sequential protein interactions and deemed successful if free DNA ends are accurately rejoined. Radiobiology gives relatively detailed knowledge of the DSB repair proteins, their biochemical modifications during repair and the entire protein sequences involved in the different DSB repair pathways.16,17 However, this perspective may oversimplify the complex repair processes taking place within the chromatin environment. Since the cell nucleus is known to be a self-organizing system,18,19 spatial chromatin organization has to be considered as an additional component in controlling repair processes.1
The most common repair mechanisms for DSBs are homologous recombination repair (HR) and non-homologous end joining (NHEJ). The main difference is that HR20 requires a DNA sequence template, whereas NHEJ21 is independent of sequence homology. In NHEJ, damage is repaired by end ligation of the DNA strands within about 1 h after induction. NHEJ is considered as a fast but less accurate repair pathway compared to HR. Besides these two most prominent pathways, so called backup pathways, are also known as, for instance, alternative end joining (a-EJ)22etc.
Importantly, it appears that spatial chromatin organization and its changes during DNA damage repair are directly involved in the spatio-temporal regulation of repair processes23 and possibly also in the selection of repair mechanisms at given sites of individual DSBs.1 To support this hypothesis, Single Molecule Localization Microscopy (SMLM)11,12,24 was found to be a useful method since specimen preparation is compatible to that known for standard fluorescence microscopy.25 By means of SMLM, it was shown that after photon irradiation with a dose of 2 Gy, γH2AX and Mre11 showed cell type specific, nonrandom similarities in their topology at the damaged sites.26 Also changes in nano-scaled γH2AX organization were observed after heat treatment of the cells.27 Although early investigations on 53BP1 repair clusters induced by high-LET ion irradiation showed a considerable, cell type dependent difference in the development of the spatial organization of 53BP1 during repair,28 further systematic studies of topological changes of 53BP1 in relation to γH2AX cluster organization are still missing.
Here, we are going to present a comprehensive confocal microscopy and SMLM study on γH2AX and 53BP1 cluster organization and topology changes during repair after 15N-ion irradiation of U87 cells in comparison with those in NHDF cells. U87 cells are regarded as being highly radio-resistant cells in comparison with NHDF cells which are supposed to show a normal radio-sensitivity.28,29 For data evaluation the established in-house made software package30 including DBScan,31 Ripley statistics,32 persistent homology,33,34 persistent imaging35,36 and principal component analysis37 was applied.
Results
γH2AX and 53BP1 show a different visual appearance in normal human skin fibroblasts (NHDF) and human glioblastoma U87 cells
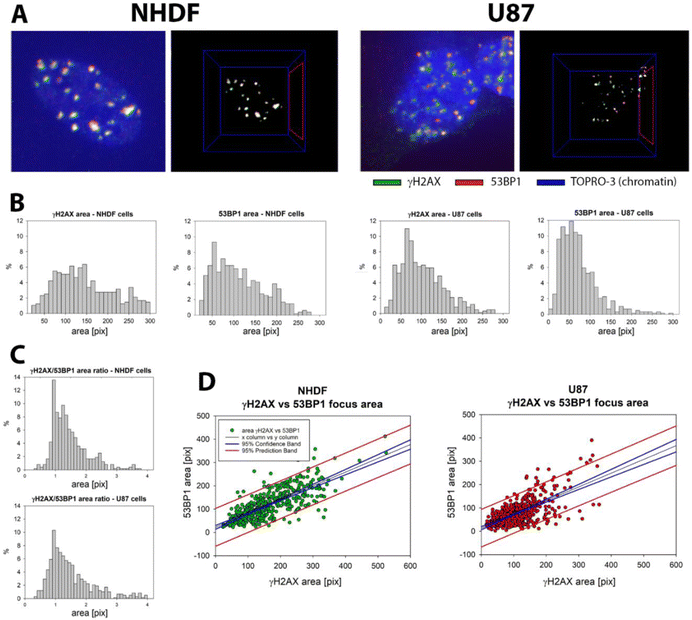
Using a combined approach of standard immunofluorescence confocal (3D) microscopy and SMLM, we examined γH2AX and 53BP1 focus formation, persistence, and organizational changes during DNA repair following exposure to high-LET 15N-ions in two differently radio-resistant cell types – normal human skin fibroblasts (NHDF) and U87 glioblastoma cells. This strategy enabled us to analyze chromatin reorganization and the temporal development of repair complexes (foci) at individual DNA double-strand break (DSB) sites, with mutual correlation within the same samples at both micro- and nanoscale resolutions. The selection of NHDF and U87 cell types was motivated by the following reasons: (a) their clinical relevance – fibroblasts are a typical cell type exposed during external radiation, such as in radiotherapy or space missions, whereas glioblastomas are tumors extremely resistant to standard photon therapy and chemotherapy; (b) the aim to explore whether the high radio-resistance of U87 cancer cells is associated with differences in chromatin responses to radiation-induced damage; and (c) the expectation that even subtle differences in DNA damage responses would be detectable in our study.First, post-irradiation changes of chromatin organization were examined, evidenced by the formation of γH2AX foci at DNA DSB sites, and the concomitant formation of 53BP1 foci at the microscale was analyzed. Fig. 1 and 2 compare γH2AX and 53BP1 focus formation in NHDF and U87 cells 0.5 h after irradiation with high-LET 15N-ions. Fig. 1 presents representative images of view-fields along with selected nuclei, which are magnified and displayed separately for γH2AX (green), 53BP1 (red), and overlaid channels (γH2AX + 53BP1 + chromatin, shown in R–G–B false colors) to enable comparison of focus type formation and micro-morphology. As in previous experiments,28,38 cells were exposed to 4 Gy of 15N-ions (E = 13.0 MeV per n, LET = 182.9 keV μm−1) in a 90° geometry, perpendicular to the cell monolayer and aligned parallel to the microscopy z-stack axis. When confocal 3D images (z-stacks) are superimposed into maximum intensity projections (as shown in Fig. 1 and 2), this irradiation setup facilitates easy qualitative (Fig. 1) and quantitative (Fig. 2) analyses of γH2AX and 53BP1 focus tracks as well as out-of-track foci.
It is evident that radiation induces microscopically visible γH2AX and 53BP1 foci in both NHDF and U87 cells (Fig. 1) at 0.5 h PI. However, while γH2AX foci in NHDF cells are typically compact and sharply bordered, and exhibit intense fluorescence, indicating well-developed foci, the corresponding foci in U87 glioblastoma cells appear more diffuse, dispersed, and less mature at this time point. Similar, but even more pronounced, differences are observed for 53BP1 foci. Although 53BP1 foci appear to be similarly compact in both cell types, in contrast to the pattern observed for γH2AX foci, they are significantly less developed and smaller in U87 cells compared to that in NHDF.
The observed differences between γH2AX and 53BP1 foci, as well as their focus track projections, visible even qualitatively, prompted us to compare the micro-characteristics of these foci in NHDF and U87 cells quantitatively. While automated software analysis of γH2AX and 53BP1 foci at the microscale is highly accurate in high-resolution immunofluorescence confocal microscopy images of nuclei after low-LET photon irradiation, the same approach remains challenging for cells exposed to high-LET ions even when using deep machine learning algorithms.39 To ensure the highest fidelity, we automatically analyzed the micro-morphological parameters of γH2AX and 53BP1 foci in Adobe Photoshop after manual demarcation of individual foci (see the Methods section).
Table 1 summarizes the mean areas of γH2AX and 53BP1 foci, as well as the ratios of γH2AX to 53BP1 focus areas and fluorescence intensities, in NHDF and U87 cells (n = 500 γH2AX and 53BP1 foci in NHDF cells and n = 602 γH2AX and 53BP1 foci in U87 cells). Notably, both γH2AX and 53BP1 focus areas were significantly smaller in U87 cells as compared to those in NHDF (P < 0.001). Due to difficulties in comparing absolute fluorescence intensities across samples with standard confocal microscopy, we instead analyzed the ratios of co-localized γH2AX and 53BP1 fluorescence intensities between the two cell types. Similarly, we examined the ratios of γH2AX to 53BP1 focus areas. While the ratio of focus areas showed no significant difference (P = 0.669), the ratio of fluorescence intensities was significantly different (P < 0.001). These findings suggest that the micro-morphology of 53BP1 foci and the extent of 53BP1 recruitment to DSB sites are largely dictated by the size-characteristics of γH2AX foci, and this relationship is maintained in U87 cells. However, the kinetics or efficiency of 53BP1 recruitment appears to be reduced in U87 cells, likely due to defects in γH2AX focus formation, possibly reflecting suboptimal chromatin organization.
| γH2AX area [pix] ± SE | Δ | Δ [%] | P | |
|---|---|---|---|---|
| NHDF | 159.14 ± 3.83 | 51.42 | 32.31 | <0.001 |
| U87 | 107.73 ± 2.59 |
| 53BP1 area [pix] ± SE | Δ | Δ [%] | P | |
|---|---|---|---|---|
| NHDF | 115.92 ± 3.00 | 39.34 | 33.94 | <0.001 |
| U87 | 76.58 ± 2.29 |
| γH2AX/53BP1 area ratio | P | Average fluorescence intensity γH2AX/53BP1 ratio | P | |
|---|---|---|---|---|
| SE, standard error; Δ, difference of values; and P, statistical significance according to Mann–Whitney rank sum test. | ||||
| NHDF | 1.51 | 0.669 | 1.71 | <0.001 |
| U87 | 1.83 | 2.04 | ||
Fig. 2 presents more detailed results from the same dataset. Panel 2A shows qualitative micro-morphological differences in γH2AX and 53BP1 foci within 2D projections of confocal 3D images, along with co-localized γH2AX and 53BP1 signals in 3D space. The histograms in Fig. 2B indicate that U87 glioblastoma cells have difficulties in forming larger γH2AX and 53BP1 foci, while the formation of smaller foci appears unaffected. It remains to be determined whether U87 cells are entirely incapable of efficiently forming larger foci or if this process is simply slower. Furthermore, the histograms and scatter plots in Fig. 2C and D demonstrate that the micro-parameters of 53BP1 foci strongly depend on those of γH2AX foci, and this relationship is preserved in U87 cells, albeit with a delay or defect in γH2AX focus formation.
In parallel to IRIF sizes, we also quantified the relative fluorescence intensities (RFU, 0–255) of γH2AX and 53BP1 IRIFs in NHDF and U87 cells. Cells were irradiated with 15N ions or γ-rays under the described conditions and analyzed 0.5- or 1 h PI (Fig. 3).
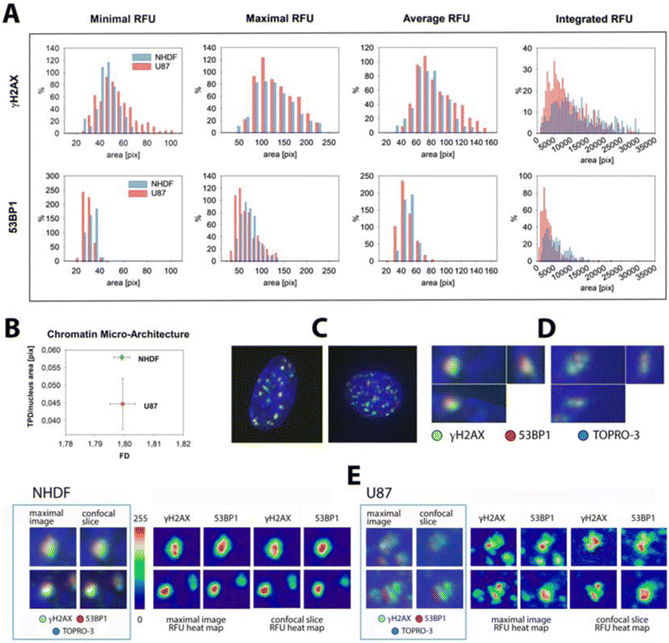 | ||
| Fig. 3 γH2AX and 53BP1 focus intensities 0.5 and 1 h PI in NHDF and U87 cells exposed to 15N-ions (4 Gy; E = 13.1 MeV per n; LET = 181.4 keV μm−1) or 60Co γ-rays (2 Gy; 1 Gy min−1). Irradiation with 15N-ions was performed as in Fig. 2 and Materials and methods section. (A) Relative fluorescence intensity (RFU) histograms for γH2AX foci at 0.5 h PI (NHDF vs. U87), showing minimum, maximum, mean, and integrated focus intensities (left to right). (B) Chromatin architecture parameters—fractal dimension (FD) and total chromatin domain perimeter (TPD)—compared between NHDF and U87; decreased TPD in U87 is consistent with reduced chromatin condensation. (C) Representative nuclei of NHDF (left) and U87 (right) irradiated with 2 Gy γ-rays and stained for γH2AX (green), 53BP1 (red), and total chromatin (TO-PRO-3, blue) at 1 h PI. (D) Colocalized γH2AX (green) and 53BP1 (red) IRIF at 1 h PI (corresponding to IRIFs in the top row of panel E) shown in orthogonal (x–y, x–z, y–z) views; NHDF (left) vs. U87 (right) after 2 Gy γ-rays; raw fluorescence (no normalization), linear LUTs adjusted for print contrast. (E) Representative colocalized γH2AX + 53BP1 IRIF in NHDF (left) and U87 (right) shown as maximum-intensity projections, a single confocal slice through the focus plane, and RFU (0–255) heatmaps. | ||
RFU histograms (Fig. 3A; 15N ions, 0.5 h PI) revealed significantly higher γH2AX RFU values compared to 53BP1 in both NHDF and U87 cells. In addition, a striking difference emerged between cell types: the integrated RFU values for both γH2AX and 53BP1 were significantly lower in U87 cells compared to those in NHDF, with an even more prominent decrease observed for 53BP1 (Fig. 3A).
In contrast to the integrated RFU, which is the sum of the RFU values of all individual pixels present within a defined focus, the histograms of average, maximal, and minimal γH2AX RFU showed shifts to higher values in U87 cells compared to those in NHDF cells (Fig. 3A). This shift was most pronounced for the minimal RFU, moderate for the average RFU, and only slight for the maximal RFU. Conversely, RFU histograms for 53BP1 consistently demonstrated significantly lower values across all measured parameters (average, maximal, minimal, and integrated) in cancer U87 cells compared to those of normal NHDF (Fig. 3A).
These quantitative results, complemented by the smaller IRIF areas for both markers in U87 cells, initially suggested a potentially denser local concentration of γH2AX phosphorylation and reduced 53BP1 recruitment to γH2AX IRIFs in U87 cells compared to those of NHDF. Alternatively, these observations are also highly compatible with more intensive H2AX-ser139 phosphorylation and/or more efficient immunofluorescence detection of γH2AX molecules in U87 cells. This could be attributed to a more open chromatin architecture, which is also indicated by the diffuse and irregular micro-morphology observed for γH2AX foci.
To investigate this, we evaluated the global chromatin architecture in NHDF and U87 cells by analyzing Fractal Dimension (FD) and Total Perimeter of Chromatin Domain (TPD) (Fig. 3B). While no significant difference was observed in FD, TPD was significantly lower in U87 cells compared to that in NHDF, strongly suggesting that chromatin in U87 cells is, in general, more decondensed or ‘relaxed’.
Considering this evidence for a more open chromatin state, the histograms showing higher minimal and average γH2AX RFU in U87 cells compared to those of NHDFs most probably reflect more intensive H2AX phosphorylation and/or enhanced detection of γH2AX within IRIFs, attributable to the increased chromatin accessibility in U87 cells.
The trend described above for γH2AX is, however, not recapitulated by 53BP1. Despite the increased γH2AX intensities in U87 cells compared to those in NHDF, the decreased intensities observed for 53BP1 likely reflect that decondensed chromatin and/or other alterations in U87 cells make 53BP1 binding to γH2AX IRIFs significantly less efficient than in normal NHDF.
Only a slight increase in the maximum RFU in U87 cells compared to that in NHDF cells could be explained by a saturation of γH2AX phosphorylation and/or antibody accumulation at the sites of RFU maxima. This only slightly higher maximal fluorescence, together with significantly higher minimal fluorescence, may point to an altered or alternative internal organization of γH2AX (and consequently 53BP1) IRIFs in U87 cells.
We therefore analyzed the spatial distribution of relative fluorescence intensities within IRIFs. For this purpose, we irradiated cells with γ-rays, which predominantly result in single DSBs. This approach ensures that the observed internal IRIF architecture cannot be misinterpreted as clustering of multiple IRIFs (DSBs), a potential artifact when using high-LET radiation. Examples of NHDF (left) and U87 (right) cells irradiated with 2 Gy γ-rays and analyzed at 1 h PI are shown in Fig. 3C. The 1 h PI time (Fig. 3C–E), rather than the 0.5 h PI time (Fig. 3A), was chosen to analyze more mature and morphologically developed IRIFs, though still within their assembly and active repair phase.
While γH2AX IRIFs in NHDF cells were typically compact, generally rounded, and sharply delineated, their counterparts in U87 cells were more irregular, diffuse, or even punctuated, particularly for larger IRIFs (smaller foci, however, roughly resembled those in NHDF cells; Fig. 3C). Therefore, before analyzing the internal architecture of these IRIFs, we confirmed that the larger, potentially punctuated foci represent real repair sites and were not merely artifacts of 2D projection from numerous IRIFs separated in 3D space (Fig. 3D). The internal architecture of γH2AX and 53BP1 IRIFs, including representative examples for both NHDF and U87 cells shown in Fig. 3D, was analyzed by means of relative fluorescence intensity heatmaps shown in Fig. 3E.
In agreement with fluorescence distribution histograms from Fig. 3A, it is evident from Fig. 3E that the spatial fluorescence distribution within IRIFs in U87 cells, especially for γH2AX IRIFs, is more dispersed, less organized, and sometimes forms multiple maximum intensity hotspots, in stark contrast to the regular, symmetric core of γH2AX/53BP1 IRIFs observed in NHDF. Importantly, γH2AX maxima do not frequently overlay with 53BP1 maxima in U87 cells, even when accounting for potential color shifts due to optical aberrations. These results thus reveal both quantitative and qualitative differences between γH2AX and 53BP1 repair foci in normal NHDF and cancer U87 cells, which might be related to functional differences/defects in DSB repair in U87 cancer cells.
γH2AX and 53BP1 are each arranged differently during the time course of repair in NHDF and U87
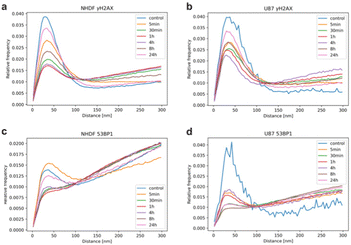
Ripley statistics of the normalized relative frequency of pairwise distances between the coordinates of the detected labelling points was performed.30,32Fig. 4 shows the results as an envelope of the distance histograms. A distinct peak at short distances indicates clustering as expected for nano-foci. The linear trend at higher distances indicates a random distribution. γH2AX was present in more distinct clusters in both cell types, especially for early repair times (Fig. 4a and b). This indicates that clusters were formed quickly after irradiation. At late time points, the clusters were smaller and the tendency towards random distribution increased, indicating a decay of γH2AX. In U87 cells the sizes were more irregular than in NHDF cells. Note that also the control specimen and the specimen analyzed at 24 h PI showed some clusters due to damage occurring during DNA replication. In contrast to γH2AX clusters 53BP1 showed less compactness and a stronger tendency towards random distribution (Fig. 4c and d). The size of clusters was, on average, slightly larger or equal for 53BP1 compared to that for γH2AX.53BP1 accumulates into more and larger clusters colocalizing and surrounding γH2AX clusters in both NHDF and U87
The spatial and temporal distribution of the accumulations of the different markers was quantified using the DBScan algorithm.31 In Fig. 5a, an exemplary nucleus is shown with γH2AX clusters as well as regions with dense and sparse label accumulations, presumably identifying heterochromatin and euchromatin, respectively. 53BP1 was found to be present throughout the nucleus over time and even in the non-irradiated NHDF samples (control). This supports the assumption that 53BP1 is permanently available in the cell nucleus although not in clusters but mostly dispersed (see also ref. 28). In Fig. 5a and b, the cluster arrangement along ionizing radiation induced tracks is visible as is typically observed from 5 min to 1 h PI. Note that in contrast to the confocal images, irradiation was performed by an ion beam passing the specimen under an angle of 10°, i.e., parallel to the microscope plane (E = 13.1 MeV per n, LET = 181.4 keV μm−1). Due to repair, these traces dissolved into smaller, separated, and randomly distributed clusters during the time course of repair (Fig. 4b). While largely co-localizing (Fig. 4b, bottom row), γH2AX formed fewer and smaller clusters than 53BP1 in both cell lines. This supports the assumption that 53BP1 surrounds γH2AX in larger, less organized structures at the nanoscale. Cluster formation differed between NHDF and U87. While the number of clusters immediately increased after irradiation in NHDF cells, the cluster formation was delayed in U87 (Fig. 5c and d). From these results, summarizing the outcome of all cells analyzed, we hypothesized that damage response is reduced in a substantial subset of cells of U87 for up to 8 h PI.The average number of clusters (Fig. 5c and d) showed a similar trend to the total number of detected events and the percentage of signals per cluster for each cell type (data not shown). Altogether, NHDF nuclei exhibited a stronger damage response in the initial phase after irradiation compared to U87 nuclei, not only in the number and size of repair protein clusters but also in the (faster) kinetics of their formation. These findings suggest that repair in NHDF nuclei is largely completed, though not necessarily fully successful, within the analyzed time course, whereas in U87 nuclei repair persists beyond 24 h.
γH2AX and 53BP1 clusters show different topological similarities in NHDF and U87 cells
To compare the topological and geometric properties among clusters, persistent homology analyses were performed.33,34 Here, we focused on holes (dimensions in persistent homology: 1) instead of components (dimensions in persistent homology: 0). In algebraic topology, these properties are called the Betti numbers40 for zero-dimensional (components) and one-dimensional (holes) simplicial complexes. They turned out to be very important topological invariants which help to distinguish between different topological spaces and to identify characteristic topological patterns. Based on the results for holes, the clusters were compared over the whole cell samples. As a measure of comparison, the Jaccard index was calculated between 0 (no similarity) and 1 (identity).34,41 In Fig. 6A, examples are shown for 53BP1 clusters in NHDF cells. Compared to the control (very low similarity), the similarity increases during repair until 4 h PI; for later time points, it decreases again. By comparing single clusters with each other, the so-called first-generation heatmaps26 were obtained. The diagonals are colored in the darkest blue, since they represent the identity. | ||
| Fig. 6 (A) Example for a series of first-generation heatmaps for 53BP1 in NHDF cells. (B) Second generation heatmaps for γH2AX and 53BP1 in NHDF and U87 cells (for details see the text). | ||
By averaging the first-generation heatmaps for each comparison, γH2AX and 53BP1 at all different repair times in both cell types, the so-called second-generation heatmaps26 were prepared in which one pixel represents the mean value of a complete first-generation heatmap (Fig. 6B). The second-generation heatmaps present an overall trend of each time cohort. The highest degree of similarity occurred at 1 h PI for γH2AX clusters in NHDF cell nuclei (upper left section in Fig. 6B) while it occurred with a delay at 4 h PI in U87 cell nuclei. Although the cluster number for γH2AX was upregulated at 24 h PI, the topological analysis showed a comparably lower similarity score as for the control (top left section in Fig. 6B).
This would agree with the hypothesis that ionizing radiation-induced clusters are different from the ones occurring naturally. Thus, after 24 hours, the regulation of γH2AX accumulation could be similar to that of the control since ionizing radiation induced DSB repair mechanisms were finished. Cluster similarity of the control and the 24-hours sample was comparatively low. So the question remains whether the regulation of γH2AX returned into its initial state.
For 53BP1 in NHDF (bottom right section in Fig. 6B), the lowest similarity degree was observed for the control and 5 min PI samples. Compared to γH2AX, the similarity, however, is in general much higher, indicating repair structure persistence up to 4–8 hours. A clear temporal development was not observed for U87 cells. Altogether, 53BP1 clusters showed a higher level of topological similarity than γH2AX for both NHDF and U87.
γH2AX is more evenly dispersed throughout clusters than 53BP1 in NHDF and U87
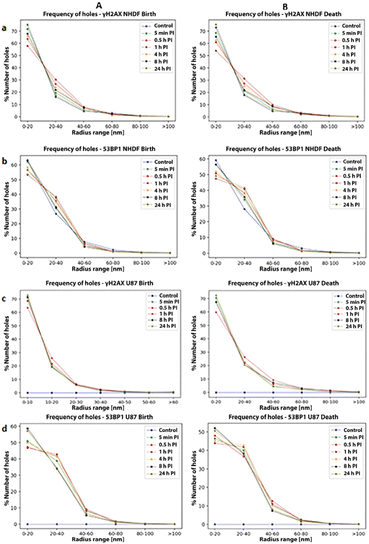
As part of the persistent homology analysis, identified clusters and their development and distribution over time were investigated. The frequency of births and deaths of holes was calculated for different times after irradiation. This represents an overview of the total percentage of holes that were formed within a certain radius range of the artificial circles of persistent homology.34For γH2AX, overall, more (>60%) small holes (0–10 nm) than big holes (>20 nm) were observed. Only 2% of the holes were further apart than 60 nm, indicating the absence of large separations. Comparing U87 with NHDF for γH2AX clusters, a lower number of big holes was found for U87, indicating a higher dispersion within γH2AX clusters. More topological changes in hole size were observed for NHDF than for U87 over time. More precisely, NHDF showed an increase of approximately 12% in the number of large holes between 0.5 and 4 h PI. This correlates with an increased repair-protein activity, which reached its maximum at 1 h PI.
Although the point density within clusters was much higher for 53BP1, the high number of large holes indicated a widely dispersed arrangement. Furthermore, the fewer small holes in 53BP1 clusters compared to those in γH2AX reassured the more even point dispersion throughout the clusters.
For 53BP1 cluster in NHDF, the number of small holes was reduced and big holes increased from 0.5 h onwards, reaching a maximum number of large holes (55%) at 1 h PI. This was followed by a decrease in large holes from 4 h PI onwards. The arrangement of 53BP1 into nano-foci with halo substructures indicated its location around γH2AX.
In U87, the number of small holes for 53BP1 clusters was 3–5% less and the number of big holes was 10% higher than that in NHDF. Furthermore, 53BP1 clusters in U87 contained 4% holes with radii larger than 40 nm. The highest number of big holes occurred at about 0.5 to 1 h PI (Fig. 7).
Principal component analysis reveals the major organization development of γH2AX and 53BP1 foci during repair
By means of principal component analysis (PCA)37 the topological data of γH2AX and 53BP1 were reduced to those main features that showed the largest variations. These uncovered biological features changed significantly during chromatin repair development.30 The plots shown in Fig. 8 revealed the first two components with the largest effects on the chromatin organization. The latent spaces for these two components (called components 0 and 1) based on the normalized data of each labeled protein and cell line are shown in Fig. 8a–d. Because the number of recorded events also contains information about DNA repair, the results were normalized to the number of events in order to investigate solely the underlying structural properties.γH2AX and 53BP1 were located at distinct regions of the latent PCA space both in NHDF and U87 cells (Fig. 8a and b). As shown recently,30 a temporal development during the repair process was observed as indicated by the arrows for both proteins and cell types. In NHDF cell nuclei, the 24-hour postirradiation sample showed a topology very similar to the control for γH2AX, while the 53BP1 data tended towards the 5-minute postirradiation sample. These findings may support the hypothesis that, after successful DSB repair, repair proteins re-establish their initial spatial organization.
U87 cells behaved differently, with the final 53BP1 topology significantly diverging from its original state. Here, the 24 h γH2AX position (i.e., topology) shifted back towards the 5-minute position, whereas 53BP1 adopted a completely different topology relative to the control. Direct comparison of γH2AX (damaged chromatin) topology cycles between NHDF and U87 cell lines (Fig. 8c) provides further support for the above hypothesis that damaged chromatin topology and organization return to their original, pre-irradiation states—particularly in normal NHDF cells, but not in U87 cells.
In contrast, for 53BP1, the 24-hour outcome differed significantly from the control in both normal NHDF and cancer U87 cells (Fig. 8d). This suggests that, in control specimens, 53BP1 is more evenly dispersed throughout the nucleus, whereas at 24 h PI it still remains partly clustered.
In Fig. 8e–l, the persistent images of the holes (component 0 only) were compared for the controls and the 24 h PI sample. For γH2AX in NHDF cell nuclei (Fig. 8e and f), the distributions were very similar, but for 24 h PI larger holes disappeared while smaller ones appeared (Fig. 8f). This is consistent with the observed movements in latent space shown in Fig. 8a and c. In contrast to NHDF, the 24 hours value in U87 cell nuclei showed disappearing small holes (Fig. 8h) compared to the control (Fig. 8g).
For 53BP1, the situation was completely different (Fig. 8i–l). In NHDF cell nuclei the small holes disappeared in the controls (Fig. 8i) while the 24 hours data revealed the opposite (Fig. 8j). The latter was also found for U87 at 24 h PI (Fig. 8l). In U87 controls, all sizes indicated a growing appearance (Fig. 8k).
Discussion
DNA double-strand breaks (DSB) are the most deleterious lesions induced by IR. The radioresistance of cells is thus closely linked to the efficiency of DSB repair, particularly in certain cell types. While the biochemical aspects of DSB repair pathways have been studied in detail, over the past decades it has become evident that chromatin organization plays a pivotal role in both DSB repair1 and carcinogenesis. This raises the intriguing question of how this phenomenon influences the radioresistance of highly radioresistant cancer cells, such as U87 glioblastoma cells42,43 studied in the present work.It has only recently been hypothesized that the local topology of chromatin actively shapes the assembly or IRIFs and DDR at individual damage sites.23,28,44–46 Furthermore, the architecture of the entire chromatin network—acting as a system14,47—also appears to participate in the DDR and genome stability determination.14,18,47,48 Therefore, we examined the interplay between γH2AX-marked chromatin organization at DSB sites, 53BP1 repair foci (IRIF) architecture, global chromatin network, and the radiosensitivity of NHDF (normal control) and U87 cells, achieving a comprehensive understanding of the critical roles of chromatin organization in radiation-induced DNA damage response.
γH2AX specifically marks DSBs and, as an incorporated histone, inherently reflects local chromatin organization at these sites.14,16,26,28,30,46,49–52 In contrast, 53BP1 is an early recruited repair protein that preferentially promotes NHEJ over HR.53 By binding directly to γH2AX-marked chromatin, 53BP1 provides a scaffold for the recruitment of additional repair factors15 and is thus critical for assembling pathway-specific repair complexes. Parallel labeling of γH2AX and 53BP1 thus enabled the investigation of how dynamic chromatin alterations at DSBs influence the timing and organization of repair complex assembly.
The unique aspects of this study are: (a) multi-scale (micro- to nanoscale) analyses of chromatin and IRIFs, ranging from repair foci to single-molecule distributions, and (b) the examination of cells irradiated with 15N ions. Accelerated high-LET heavy ions generate more profound yet less understood effects on chromatin8,9 than photon radiation,11,12,46 as they are rarely available for experimental study. This knowledge gap limits hadron therapy development and hinders manned space mission to Mars.6
Microscale differences in γH2AX and 53BP1 IRIFs between normal NHDF and radioresistant U87 cancer cells
We first analyzed γH2AX and 53BP1 IRIF features in 3D-fixed cells using immunofluorescence confocal microscopy. While 15N ions induced visible γH2AX and 53BP1 IRIFs already at 0.5 h PI, marked differences were evident in their constitution and size between normal NHDF and U87 glioblastoma cells. IRIFs in U87 cells appeared smaller, more irregular, diffuse, and immature compared to NHDF cells, with even more pronounced differences for 53BP1. IRIF size distributions revealed that, while U87 cells retained the ability to form small γH2AX and 53BP1 IRIFs, they failed to generate larger ones.The reduced capacity of U87 cells to form larger γH2AX and 53BP1 IRIFs early (0.5 h PI) after irradiation correlated with decreased 53BP1 fluorescence intensity. In contrast, γH2AX exhibited significantly higher minimal and average intensities, with only a slight increase in maximal intensities compared to those of NHDF cells. This disproportionate increase in minimal versus maximal γH2AX fluorescence, along with altered 53BP1 binding, suggests an altered internal IRIF architecture in U87 cells.
The observed changes in the IRIF architecture were accompanied by pan-nuclear chromatin decondensation in U87 cells, indicated by a similar fractal dimension (FD) but a decreased total chromatin domain perimeter (TPD) compared to NHDF cells.
Collectively, our results highlight local chromatin and IRIF architecture as significant regulators of DSB repair, which is distinctly altered in radioresistant U87 cells, although further time-course analyses are needed to distinguish between a permanent structural defect and delayed IRIF formation. Also, it should be noted that fluorescence intensity studies, along with FD and TPD measurements obtained through confocal microscopy, should be interpreted with caution due to their threshold sensitivity.
Nanoscale topology of γH2AX and 53BP1 IRIFs in normal NHDF and radioresistant U87 cancer cells: changes during the repair period
We analyzed the topology of γH2AX and 53BP1 foci in NHDF and U87 cells at single-molecule resolution (10–20 nm) and over a prolonged post-irradiation period (24 h PI) by SMLM microscopy. In combination with Ripley distance analysis and topological investigations of network formation, this approach offered a unique opportunity to overcome the Abbe limit and achieve the desired resolution, utilizing the same cell preparations previously employed for confocal microscopy. To maximize the precision of SMLM experiments, irradiation was performed parallel (10°-angle) to the cell monolayer28 and the dose was reduced to 1.25 Gy, ensuring only a few ion tracks per nucleus.SMLM analysis of γH2AX and 53BP1 cluster sizes (the term “cluster” is used instead of “IRIF” due to the unclear relationship between them) revealed similar behavior for γH2AX in both NHDF and U87 cells, with a rapid increase after irradiation reaching a peak at 1 h PI, followed by a return to the baseline at 24 h PI. In contrast, 53BP1 cluster areas in U87 cells showed significant fluctuations around the baseline up to 4 h PI, only starting to increase thereafter, peaking at 8 h PI. This indicates a delayed formation of 53BP1 IRIFs and potential repair initiation in U87 glioblastoma cells compared to those in NHDF cells, consistent with the slower increase in both γH2AX and 53BP1 cluster numbers per nucleus after irradiation, present confocal microscopy results, and our previous study results.28 While the above-introduced results cannot be fully interpreted without additional knowledge, collectively they suggest a delayed and perhaps diminished repair process in radioresistant U87 cells, as discussed later.
Persistent homology was employed to further investigate the nanostructures of identified clusters and their regulation over time. This analysis quantified topological features by measuring similarities among the clusters’ components (denser protein accumulations) and holes (the empty spaces between them). Consequently, persistent homology enabled the comparison of scale- and rotation-invariant features, highlighting rearrangements in chromatin and protein organization over time.
Comparing the averaged similarities of both cell lines using second-generation heatmaps, 53BP1 clusters were generally found to be more similar to one another during the repair process than γH2AX clusters. Since DSBs induced by high-LET ion irradiation create chromatin damage of varying complexity and/or multiplicity, γH2AX is expected to form more diverse and less similar arrangements. In contrast, 53BP1 accumulations may be more controlled and organized around DSB lesions. This higher mutual similarity of 53BP1 foci to γH2AX foci, regardless of cell type, may also indicate that while the chromatin architecture must be preserved, 53BP1 adopts a liquid droplet-like structure.54 Notably, a substantial similarity between 53BP1 and γH2AX clusters was observed in U87 cells, suggesting an adaptation of both arrangements.
However, by comparing the cell types, NHDF cells showed a conclusive temporal development for both repair factors, whereas this trend was not observed in U87 cancer cells. U87 displayed small and apparently random changes of averaged cluster similarities. For γH2AX, a more distinct peak in similarity was only found around 4 h PI, while for 53BP1, a broader peak occurred from 0.5 to 24 h PI.
It was hypothesized that the DDR was initially upregulated in a more random manner due to severe and complex damage.28 A so-called “state of alarm” might pre-dominate the cell. After the initial repair activities were completed and acute damage was resolved, the remaining potentially more complex lesions were subsequently addressed. Thus, the peaks in cluster similarity for both markers could represent stricter regulated repair after overcoming the acute phase. This hypothesis was further supported by the result that 53BP1 persisted in the most similar structures for almost 24 h, whereas γH2AX showed the highest similarity only in the early hours PI.
As another possibility, but not necessarily mutually exclusive, functionally and structurally distinct chromatin domains may be affected by DSBs, leading to substantial variability in γH2AX foci. With increasing PI time, chromatin tends to adopt a more uniform structure that is better suited for DSB processing. After free DNA ends are rejoined, the chromatin architecture attempts to revert to its original state, which differs across individual domains. This reversion process can also proceed with distinct kinetics, for example between heterochromatin and euchromatin.
By juxtaposing individual clusters within each cohort using the first generation heatmaps,26 another temporal trend for 53BP1 clusters was observed in NHDF. Clusters in the non-irradiated control and at early PI times appeared to be evenly non-similar. Afterwards the similarity increased up to 4 h PI. Finally, at 24 h PI, some 53BP1 clusters became more similar, even though the overall similarity decreased. Therefore, 53BP1 clusters seemed to increase in heterogeneity over time. This trend was not observed for γH2AX.
Despite an unexpected increase in the number of γH2AX clusters at 24 h PI (perhaps due to the recurrence of S/G2-cells after cell cycle arrest release), the topological analysis30 revealed a similarity score nearly comparable to that of the control, but slightly shifted in both components (0 and 1). This supports the hypothesis that clusters with radiation-induced damage differ from naturally occurring ones in the control. After 24 h, the regulation of γH2AX accumulation may resemble that of the control, indicating the completion of radiation-induced DSB repair.
The principal component analysis identified repair steps that were accompanied by changes in repair focus features, i.e., shifts in the average topology values within the latent space. Therefore, this analysis method was used to further investigate whether different DDR factors (γH2AX and 53BP1) and cell types (NHDF and U87) show different shifts during the PI-period (tracks) in PCA imaginary space.
When comparing γH2AX and 53BP1 proteins within a given cell line, γH2AX and 53BP1 tracks were consistently separated in the latent spaces for both cell types. This indicates that the differences in γH2AX and 53BP1 focus structures, as well as in the kinetics of their alterations, are more pronounced and may represent a conserved feature across both cell types.
Comparing the cell lines for a given protein, a highly similar temporal development in the latent space was found for both, γH2AX and 53BP1, respectively. This supports the trend of adaptation as described above.
For γH2AX, the controls and the 24-hour cohorts were located in the negative space compartments, whereas the postirradiation cohorts corresponding to the active repair period were positioned in the positive ones. This observed separation is consistent with the proposed hypothesis that the regulation and topology of γH2AX domains return to their initial state. For 53BP1, such behavior was neither expected nor observed. Interestingly, the active repair cohorts up to 24 h PI overlapped in the latent space, whereas the non-irradiated controls were the most distant with respect to component 1.
Using persistence images, the development of hole sizes for the individual time cohorts was quantified. Here, both NHDF and U87 cell lines and γH2AX and 53BP1 proteins differed fundamentally. For NHDF, no small holes (50–150 nm) were observed in 53BP1 clusters in the control samples, whereas they were consistently present in all irradiated samples. The constant presence of equal sized holes indicates a controlled arrangement of 53BP1 around the damage.
In contrast, in U87 cells, small holes were also observed but accompanied by larger ones during the period from 0.5 to 4 h PI. These differences in hole sizes may reflect a less consistent response or indicate chromatin decondensation, in agreement with more heterogeneous arrangement of IRIFs in U87 cells according to the confocal microscopy heatmaps.
For 53BP1, the situation was just complementary in NHDF cell nuclei. While larger holes dominated in the control, the small holes considerably increased in 24 h PI. This was also found in U87 cells, indicating that this is a principal feature of 53BP1 cluster relaxation. This, however, differed considerably from the 53BP1 control in U87 nuclei. Altogether, the PCA revealed fundamental differences between the cell types, the repair factors, and the repair development.
Certain limitations of the present SMLM study should be acknowledged. Cells were visually preselected for SMLM data acquisition based on specific criteria (Methods section), as both the acquisition and data evaluation processes are extremely time-consuming. This preselection may introduce bias; however, the same criteria were applied consistently across all datasets, reflecting common practices in microscopy to minimize outcomes affected by sample preparation quality.
Additionally, slides were not always prepared in triplicate due to limited beam time at high-energy particle accelerators and logistical challenges related to transporting samples between CR, Germany, and Russia for irradiation and analysis at micro- and nanoscale.
Finally, biological variability must be considered. Alekseenko et al. highlighted the challenges in achieving reproducibility of quantitative results across individual cancer cells. Slight variations in the microenvironment for each cell may lead to functional differences, even among cells from the same tumor or cell type. While such variability might be negligible in bulk experiments, it can become significant in single-cell analyses. In other words, the greater the precision of single-cell analyses, the more apparent individual variations may be recorded, even without the use of independent experimental replicates.
Implications for DSB repair and radioresistance of cancer cells
Our results have significant implications for DSB repair and cancer cell radioresistance, although many questions remain unanswered. The more intense yet less/differently organized γH2AX IRIFs observed in U87 cells suggest that decondensed chromatin enhances repair protein trafficking within the nucleus and their access to DSB sites.55 However, the associated chromatin disorganization within IRIFs seems to hinder 53BP1 binding and may also mobilize free DNA ends. These observations suggest a deregulation or adaptation of DSB repair, potentially shifting towards alternative, resection-based mechanisms56,57 that are more robust yet highly mutagenic. This is further supported by the anticipated shift of U87 cells to the G2 phase (unpublished results), in contrast to NHDF, which, under the experimental conditions of this study (80% confluence, see “Materials and Methods section”), were predominantly irradiated in the G1 phase and repaired via NHEJ or alternative pathways.53Collectively, these observations may explain the greater capacity of U87 cells to remove complex or clustered DSBs,28 though potentially at the expense of repair fidelity.28 Some authors consider (altered) DSB repair mechanisms to be a primary driver of radioresistance in U87 cells.58 U87 cells may also adapt to survive despite having unrepaired DNA. In fact, cancer cells, in particular, can often recover from cell cycle arrest even with residual DSBs present.59
Furthermore, our results imply that the relationship between the chromatin architecture and IRIF formation is complex and not always straightforward. The observation that precise tuning of the chromatin architecture is crucial for balancing DSB repair efficiency and precision aligns with our previous studies, which have shown that both excessive chromatin condensation (induced by hyperosmotic media) and decondensation (caused by hypoosmotic media or Trichostatin A) negatively impact DSB repair, leading to delays or even complete failure.60 Notably, while repairing DSBs resumed after returning cells to isotonic conditions, chromatin decondensation led to long-term or even permanent repair impairments. This evidence and similar observations during natural differentiatian55,61 suggest that the altered chromatin architecture can prevent IRIF formation and hinder the appropriate restoration of the original chromatin structure during and after the repair process.
Our hypothesis that the irregular and often punctuated γH2AX IRIFs in U87 cells, characterized by multiple, dispersed 53BP1 fluorescence intensity cores on spatial fluorescence intensity heatmaps, arise from chromatin decondensation is strongly supported by Granzotto et al.62 They observed concentration-dependent fragmentation of γH2AX IRIFs into numerous subfoci (up to about 13) in ‘mutation-free’ normal fibroblasts treated with sodium butyrate, which induces chromatin condensation without causing additional DSBs.
Additionally, using SIM2 microscopy combined with super-resolution FRAP (FRAP-SR), Wu et al.63 identified two types of 53BP1 IRIFs: compact, stationary IRIFs and amorphous IRIFs with dynamic shape changes. In FRAP-SR, amorphous IRIFs exhibited rapid recovery of 53BP1-EGFP within distinct subcompartments, indicating varied protein mobilities and functions within a single IRIF. In contrast, compact IRIFs recovered 53BP1-EGFP uniformly as a single compartment but displayed heterogeneous recovery rates. These two types of 53BP1 IRIFs parallel the differences we observed between NHDF and U87 cells; however, the protein dynamics within these IRIFs still require further investigation.
The inability of U87 cells to form larger γH2AX IRIFs and their preference for resection-based repair support the hypothesis that IRIFs associated with NHEJ, HR, and alternative repair mechanisms exhibit distinct morphological features. Remarkable differences in the γH2AX/53BP1 IRIF morphology between G1 vs. S/G2 nuclei have been observed also in ref. 64. In addition, the PCA of γH2AX/53BP1 IRIF features in individual head-and-neck tumors revealed inter-patient variability,39 which in at least some cases appeared to correlate with DSB repair kinetics.65
In contrast, the highly similar size- and intensity-ratios of γH2AX and 53BP1 foci in NHDF and U87 cells demonstrate that the general rules of repair complex formation are preserved even in cancer cells. This suggests an architectural relationship between γH2AX and 53BP1 foci, indicating that γH2AX-marked chromatin at DSB sites dictates the organization of repair complexes and is therefore functionally relevant.
Causes of IRIF defects in U87 cells: which came first, the chicken or the egg?
A fundamental yet complex question remains: what underlies the observed DSB domain and IRIF defects or adaptations in U87 cells, and how do these changes affect radiosensitivity? One possibility is that alterations in the global architecture of the chromatin network influence the local chromatin environment at DSB sites. This, in turn, impacts the access and binding of DSB repair proteins, including various chromatin remodelers, to the damaged areas, thereby affecting subsequent steps in IRIF formation and DSB repair, with implications for the cell's radioresistance. Moreover, disrupted or un-restored chromatin organization following DNA repair may contribute to increased genomic instability in cancer cells after irradiation. This can occur, for instance, through the activation of specific epigenetic pathways, driving these cells toward distinct fates compared to their normal counterparts,49 including the generation of radioresistant tumor cell clones.49U87 cells exhibit a broad spectrum of mutations, with 512 genes homozygously mutated, which may directly or indirectly influence the chromatin architecture. PTEN mutation promotes the dissociation of histone H1 and increases histone H4 acetylation at lysine 16.66,67 Another significant genetic defect contributing to global chromatin decondensation in glioblastoma cells is the overexpression of class I HDACs.68
In high-grade glioma, mutations in histone H3, such as the H3K27M ‘oncohistone’, reduce the repressive mark H3K27me3 and increase activating marks H3K27ac and H3K36me2/3.69 Sharda et al. then illustrated that dynamic changes in histone H3 phospho-acetylation correlate with the radiosensitivity of mitotic cells.70 Furthermore, 3D genome studies revealed hypermethylation of a CCCTC-binding factor (CTCF) motif in certain IDH1 mutant gliomas, leading to the removal of boundaries between chromatin domains.71 Collins et al. demonstrated that the deletion of a CTCF motif reduces γH2AX spread within a TAD,72 which agrees well with the smaller sizes of γH2AX observed in U87 cells in the present study.
These examples, while not always directly relevant to U87 cells specifically, highlight the complex effects of mutations on the epigenetic chromatin landscape in gliomas. Consequently, the combined effects of mutations and other factors shaping the chromatin architecture likely create a specific environment that regulates IRIF formation at both global and DSB site-specific levels.1 Notably, about 50% of cancer driver genes encode chromatin-bound proteins.73
Moreover, cancer-associated mutations can indirectly alter the chromatin architecture by affecting cell signaling and metabolic pathways.74 For instance, Sulkowski et al. demonstrated that oncometabolites produced by IDH1 or IDH2 mutations inhibit the lysine demethylase KDM4B, leading to aberrant hypermethylation of H3K9 at DSB sites. This masks local H3K9 trimethylation signals and disrupts the binding of TIP60, ATM, and downstream proteins to IRIFs.75 Although U87 cells possess wild-type IDH1 and IDH2 genes, similar metabolic mechanisms may still occur.
Conversely, IRIF defects and adaptations may result from mutations and altered functions of IRIF-binding proteins, but the literature does not provide a conclusive perspective on this matter.
In p53-mutated glioblastoma cell lines, PTEN deficiency enhanced RAD51 levels throughout the cell-cycle, leading to the formation of RAD51 IRIFs, potentially with an altered architecture, even outside the G2/S phase.76 However, a more recent study did not confirm PTEN-induced RAD51 overexpression in U87 cells.77 RAD18—significantly overexpressed in U87 cells78—stabilized 53BP1 IRIFs, thereby supporting NHEJ and positively correlating with U87 radioresistance.79 In contrast, the same genetic defect has also been linked to stimulated HR,80 which is better compatible with the compromised/delayed 53BP1 IRIF formation observed in U87 cells in the present study. In glioblastoma cells—though not specifically in U87 cells, where GOS2 is not overexpressed—GOS2 overexpression via the mTOR/S6K/RNF168/53BP1 signaling pathway inhibited γ-H2AX IRIF assembly. This corresponds to our observations of inefficient/delayed γ-H2AX IRIF formation; however, overexpressed GOS2 also stabilized 53BP1 and 53BP1 IRIFs.81
The stability of 53BP1 appears unaffected in U87 cells, as we observed a significant pool of this protein in U87 nuclei.28 Its inability to form IRIFs thus stems from the altered chromatin architecture in IRIFs or unknown mutations of 53BP1 and/or its interacting partners. The former possibility is supported by a recent study by Chen et al.,82 which demonstrated that epigenetics likely drives the cellular heterogeneity of glioblastoma cells, while mutation load plays a modifying role. The significant impact of the chromatin architecture on IRIF formation was further demonstrated by Granzotto et al., who found that γH2AX IRIFs fragmented into subclusters due to chromatin decondensation, even in ‘mutation-free’ normal fibroblasts.62 However, with approximately 35 interchromosomal translocations, 1315 structural variations, 191![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 743 small insertions and deletions, and 2
743 small insertions and deletions, and 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 384
384![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 470 single nucleotide variations in U87 cells,66,83 along with additional alterations from alternative splicing84,85 or aberrant non-coding RNA expression,86 it is challenging to rule out that IRIF defects in U87 cells result partly from genetic defects in repair and related proteins.87
470 single nucleotide variations in U87 cells,66,83 along with additional alterations from alternative splicing84,85 or aberrant non-coding RNA expression,86 it is challenging to rule out that IRIF defects in U87 cells result partly from genetic defects in repair and related proteins.87
The chromatin environment, gene mutations (in repair proteins), and other factors likely operate in a highly interconnected manner. For example, overexpressed HDACs globally open the chromatin architecture while also interacting directly or through partners, such as RAD18, with HR88 and NHEJ members.76 Notably, RAD18 directly interacts with 53BP1, ensuring its retention at DSB sites via monoubiquitination at lysine 1268. Reduced RAD18 expression in glioma cells89 can thus contribute to decreased 53BP1 accumulation at DSBs. This complexity makes it challenging to clearly define the effects of individual factors on the chromatin and IRIF architecture, and U87 cell radioresistance, complicating the establishment of causality.
High-LET versus low-LET radiation
The spatial heatmaps analyzing the internal γH2AX and 53BP1 IRIF architecture, shown in Fig. 3, were generated using γ-rays to avoid artifacts caused by multiple DSB clustering after high-LET IR exposure. These heatmaps showed differences in γH2AX and 53BP1 IRIF architectures in U87 cells, consistent with observations from other experiments using 15N ions. Thus, high-LET and low-LET radiation appears to be associated with similar defects and adaptations of IRIFs and DSB repair at the microscale. Furthermore, a comparison of γH2AX and 53BP1 nano-topologies during the repair period in the latent PCA space for normal NHDF fibroblasts exposed to γ-rays and 15N ions revealed similar trends (Discussion section and Fig. 9). However, while this movement was present for both low- and high-LET radiation, the extent and specific characteristics varied significantly across different cancer types. These analyses will be the focus of a comprehensive separate study. | ||
| Fig. 9 Principal component analysis of persistent imaging for (a) γH2AX and (b) 53BP1 in NHDF fibroblasts after irradiation with 2 Gy g-ray photons (compare with Fig. 8a–d for 15N ions). | ||
An additional factor requiring detailed investigation in the context of radiation LET, cell type, IRIF architecture, repair pathway choice, and cellular radioresistance is the contribution of the cell cycle. The distribution of normal and cancer cells across cell-cycle phases can differ substantially, as can the duration of post-irradiation cell-cycle arrest following exposure to γ-rays and high-LET ions.90–96
Conclusions and future perspectives
In conclusion, we found that the morphological and topological parameters of γH2AX and 53BP1 IRIFs are mutually correlated—both at the micro- and nanoscale—and that this relationship is preserved even in cancer cells, at least to some extent. This observation supports the hypothesis that the formation of IRIFs is controlled, at least in part, by the local chromatin architecture at individual DSB sites and by its remodeling during the initial phase of repair.1 However, U87 glioblastoma cells display specific morphological and topological alterations in γH2AX and 53BP1 micromorphology and nanotopology compared to normal NHDF fibroblasts. These alterations may indicate an initial delay in DSB repair, altered repair pathway choice, more efficient removal of complex and/or multiple DSB lesions, increased post-irradiation genomic instability, and tolerance to such instability. The conclusions on increased post-irradiation genomic instability in U87 cells are strongly supported by SMLM combined with PCA analysis, which revealed significant differences in the movement of γH2AX and 53BP1 focus topology in the latent PCA space during the repair process. While in NHDF fibroblasts the topology of repair foci returned to values close to those of non-irradiated controls after 24 h—forming closed loops in the PCA space—U87 cells exhibited open trajectories, indicative of incomplete restoration of the pre-irradiation chromatin architecture at DSB sites. As similar conclusions were previously reported for the global chromatin architecture,14,47,48,97 our findings suggest that the observed defects in chromatin restoration may promote epimutations98 and increased genetic instability, for example through less efficient silencing of transposons in heterochromatin or deregulating transcription networks. Importantly, genomic instability can be further augmented by the fact that especially cancer cells can recover from the cell cycle arrest even in the presence of residual DSBs, once their number per nucleus decreases to approximately 10–20.59Comparing our results on the DSB markers γH2AX and 53BP1—the latter being predominantly associated with NHEJ—with proteins such as MRE11 or RAD51, which serve as mediators of HR, could provide valuable insights into the differences between repair pathways and the mechanisms underlying pathway choice in future. The spatial organization of chromatin at individual DSB sites seems to play a crucial role in determining the most appropriate repair mechanism for each specific DSB site, with crucial implications for cancer cell radioresistance and genomic (in)stability.1,14,48
Methodologically, super-resolution SMLM provided detailed insights into nuclear and cluster-level structures, underscoring the value of this method for studying DNA repair.99–102 Accordingly, this approach should be upgraded, e.g., to 3D-SMLM, and integrated into future investigations.
Materials and methods
Sample preparation
For staining, the cells were incubated overnight at 4 °C with primary antibodies: mouse anti-γH2AX (ab22551, Abcam, Cambridge, UK) and rabbit anti-53BP1 (ab21083, Abcam). After rinsing with 0.2% Triton X-100, the cells were washed three times in PBS (5 min each). Secondary antibodies were AlexaFluor 647-conjugated goat anti-mouse for γH2AX (Johnson Laboratories, New Brunswick, NJ, USA) and Alexa 568-conjugated goat anti-rabbit for 53BP1 (Johnson Laboratories). They were applied for 30 min at room temperature in the dark. Nuclei were counterstained with DAPI to visualize total nuclear chromatin.
Finally, air-dried samples were embedded in ProLong Gold (Thermo Fisher Scientific, Waltham, MA, USA) and allowed to polymerize for 24 hours in the dark at room temperature before sealing.
Image acquisition
Data analysis
For this study, nuclei containing a minimum of 2000 and a maximum of 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 detected fluorophores (excluding controls) were averaged on a sample-by-sample basis. A minimum of 20 data stacks were collected for each sample. Manual masks were applied to reduce the background signal. The nucleus was delineated in the DAPI channel and subsequently applied to other channels to ensure that only proteins located within the nuclei were considered in the analysis.
000 detected fluorophores (excluding controls) were averaged on a sample-by-sample basis. A minimum of 20 data stacks were collected for each sample. Manual masks were applied to reduce the background signal. The nucleus was delineated in the DAPI channel and subsequently applied to other channels to ensure that only proteins located within the nuclei were considered in the analysis.
Only cells of sufficient preparation quality were included in the analysis. Specifically, inclusion criteria required the absence of bright background signals, a sufficiently high number of blinking events to allow reliable computing, and comparable numbers of blinking events across samples, particularly for persistent imaging followed by PCA. Thus, from the beginning, only cells meeting defined acquisition standards were subjected to SMLM measurements.
 | ||
| Fig. 10 Schematic representation of the different evaluation steps of SMLM datasets. After acquisition of a time series of image frames, the coordinates and other values of all the blinking events of a cell nucleus were integrated into a matrix, the so-called “orte-matrix” (top-left). In the one-channel analysis (top right), the coordinate values of the points were used for the production of pointillist images, or the data were subjected to cluster analysis using Ripley statistics. For the representative graph (top right), the point-to-point distances were calculated and represented in a normalized distance frequency histogram. In the lower part, the processes of persistent homology, persistent imaging, and principal component analysis are shown. The point pattern is transferred into a bar code description of components and holes. The lengths of the bars (difference of the α values of the end and the beginning of a particular bar) provide the lifetime; the beginning provides the birth in the one-point cloud. The one-point clouds for all cell nuclei that are considered for evaluation are transferred to pixel images (persistent imaging). Each pixel is compared for all images. The results of these comparisons span an n-dimensional orthogonal vector space. The variations of the pixel values determine the components of the principal component analysis. The scree plot shows the two components with the largest variances (= components 0 and 1). Finally, the outcome for the components (orthogonal vector values) with the largest variation and the second largest variation determines the latent space (graph bottom right). Note: this figure was originally published under CC-BY license in ref. 97. | ||
The persistent homology algorithm was employed to investigate the topological similarity of clusters. Barcodes were generated to represent the pointillist structure of these clusters by drawing circles with increasing radii (α) around each point within a cluster. When circles intersected, lines connecting the points were drawn, thus forming components. As intersection continued, closed areas—referred to as holes—emerged. All components begin with α = 0 and “die” upon intersection with another component. The evolution of all components and holes was plotted as barcodes, illustrating their lifespan from birth to death as a function of the radius (α).
In the experiments on chromatin analyses by SMLM, the attention will be focused on two quantifiable parameters (Betti numbers):40 the number of components (= number of points with which the analysis starts) and holes (= closed configurations like meshes of a net that occur and disappear during the mathematical evaluation process). These elements are independent of each other. In algebraic topology, these elements are also called zero-dimensional and one-dimensional simplicial complexes. By persistent homology, significant structures of a point pattern are, therefore, obtained by the following process (Fig. 8, bottom left): each point (= component = simplicial complex of dimension 0) registered in the orte-matrix is surrounded by a virtual circle with an increasing radius. Each component is represented by a bar that starts at 0 (= the given point) and ends at the radius value where two circles attach. In this way, two components merge into one. During this process, enlarged components can form a virtual closed loop (in its simplest form a triangle) with free space inside not covered by the increasing virtual circles. The area of this free space is considered as a hole (simplicial complex of dimension 1). At this moment, another bar starts. When the hole is completely covered by the increasing virtual circles, this bar also ends. With connecting components and, thus, reducing their number, the number of holes first increases and then decreases because more and more holes are closed. By this process, the point pattern of fluorescent molecules obtained by SMLM is transferred into a bar code pattern of two different types of bars: one for the components and one for the holes.
For each labelling cluster, the results of the barcodes are transferred into a diagram of “bar lifetime” versus “bar birth”, which is then transferred into a pixel scan (persistent image) in which the pixel intensity correlates with the number of points in a particular pixel of this diagram. These persistent images are obtained for each cell of an experiment. Afterwards, the persistent images are subjected to principal component analysis (PCA).37 This means that the values of the 1st, the 2nd, the 3rd, the 4th, the nth pixel, etc., of all individual persistent images are separately compared and represented in an n-dimensional orthogonal vector space. The dimension (i.e., the respective vector of this n-dimensional vector space) with the largest variances then forms the component 0 in the final diagram, the latent space (see Fig. 8, right bottom). Component 1 is a dimension (vector) perpendicular to component 0 and shows the second large variance of values. In this way, the results of a complex point pattern can be reduced to one value (±standard deviation) of a diagram (latent space) of two dimensions. This means that only two principal components describe the main features that represent the main variances, while small variations (“biological noise”; other orthogonal vectors with fewer variations of values) are neglected. In this way, changes of chromatin organization can be described by shifts of the outcome in the latent space.
The similarity values are visually represented in heatmaps, where dark blue signifies higher similarity and red indicates less similarity. The first-generation heatmap26 illustrates comparisons of each cluster with itself and the remaining clusters within the sample nuclei.
Due to the extensive volume of data analyzed, numerous individual heatmaps were generated, allowing for a comprehensive examination of topological similarities. To streamline this analysis, second-generation heatmaps26 were created to represent the average similarities of components and holes across all clusters within each nucleus. This approach enabled the identification of overall trends in the topological features of the clusters for different samples.
Author contributions
Conceptualization, M. F. and M. H.; methodology, K. A. K., L. R. P., M. F., and M. S.; software, J. W. and M. H.; validation, K. A. K., L. R. P., M. S. and M. H.; formal analysis, K. A. K. and L. R. P.; investigation, I. F., J. T., L. T., M. F. and M. H.; resources, I. F., F. B., M. F. and M. H.; data curation, M. S. and M. F.; writing – original draft preparation, K. A. K., M. F. and M. H.; writing – review and editing, K. A. K., L. R. P., M. S., J. W., I. F., J. T., L. T., F. B., M. F. and M. H.; visualization, K. A. K., L. R. P. and M. F.; supervision, M. F. and M. H.; project administration, M. F. and M. H.; funding acquisition, M. F. and M. H.; all authors have read and agreed to the published version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The raw data supporting the conclusions of this article will be made available by the authors on request.The data are preserved in the Kirchhoff-Institute data archive. Due to the rules of the funding agency “Deutsche Forschungsgemeinschaft”, this archive has to be preserved for minimum 10 years after publication of the data. Reusing existing data whenever possible is explicitly recommended to avoid time-consuming additional measurements. Reusing of the data can be made in collaboration with the authors who have the expertise in data handling.
Acknowledgements
The authors gratefully acknowledge the financial support provided by the DFG (H1601/16-1) and GACR (project GACR 20-04109J) to M. H. and M. F., as well as the projects EU COST MultiChem, Action CA20129, to M. F. and BMBF (FKZ 02NUK058A) for supporting M. H. This research collaboration between Heidelberg University and Institute of Biophysics, Brno, was partly funded by the DAAD (Deutscher Akademischer Austausch Dienst = German Academic Exchange Service) PPP Program for Project-related Person exchange (to M. H. and M. F. Brno), grant project-ID: 57702816. The authors thank Fabio Oscar Marcela for providing some parts of Fig. 8. The authors also acknowledge the possibilities and support of the ion irradiation facilities at the Joint Institute for Nuclear Research (JINR) in Dubna, Russia. Furthermore, the authors thank Dr Corinna-Cornelia Seegler-Sandbanck, European Institute for Feasibility Studies, Strasburg, MVP, Germany, for her continuous support to make researchers’ lives easier. In addition the authors also acknowledge the support of MS-RADAM. The MS-RADAM project has received funding from the Horizon Europe Programme (HORIZON-MSCA-2024-DN-01 call) under Grant Agreement no. 101225527.References
- M. Falk and M. Hausmann, Cancers, 2021, 13, 18 CrossRef CAS PubMed.
- L. Jezkova, M. Zadneprianetc, E. Kulikova, E. Smirnova, T. Bulanova, D. Depes, I. Falkova, A. Boreyko, E. Krasavin and M. Davidkova, et al. , Nanoscale, 2018, 10, 1162–1179 RSC.
- J. A. Nickoloff, N. Sharma and L. Taylor, Genes, 2020, 11, 99 Search PubMed.
- V. Mladenova, E. Mladenov, S. Chaudhary, M. Stuschke and G. Iliakis, Front. Cell Dev. Biol., 2022, 10 Search PubMed.
- B. Jakob, M. Dubiak-Szepietowska, E. Janiel, A. Schmidt, M. Durante and G. Taucher-Scholz, Sci. Rep., 2020, 10, 1443 Search PubMed.
- A. Rizzo, E. M. Borra, L. Ciciani, L. Di Fino, G. Romoli, G. S. Amantini, L. Sperandio, I. Vilardi and L. Narici, Eur. Phys. J. Plus, 2023, 138, 1001 CrossRef.
- A. Helm and C. Fournier, Strahlenther. Onkol., 2023, 199, 1225–1241 CrossRef PubMed.
- M. Durante, J. Debus and J. S. Loeffler, Nat. Rev. Phys., 2021, 3, 777–790 Search PubMed.
- H. Paganetti, Int. J. Radiat. Oncol., Biol., Phys., 2022, 112, 222–236 CrossRef PubMed.
- W. Tinganelli and M. Durante, Cancers, 2020, 12, 1–43 Search PubMed.
- J.-H. Lee and M. Hausmann, in DNA-Repair, ed. P. Behzadi, Intech-Open, Rijeka, 2021, ISBN 978-1-83881-094-8 Search PubMed.
- A. V. Solov'yov, A. V. Verkhovtsev, N. J. Mason, R. A. Amos, I. Bald, G. Baldacchino, B. Dromey, M. Falk, J. Fedor, L. Gerhards and M. Hausmann, et al. , Chem. Rev., 2024, 124, 8014–8129 Search PubMed.
- J. Reindl, A. M. Abrantes, V. Ahire, O. Azimzadeh, S. Baatout, A. Baeyens, B. Baselet, V. Chauhan, F. Da Pieve and W. Delbart, et al., in Radiobiology Textbook, ed. S. Bartout, 2023, pp. 83–189 Search PubMed.
- M. Falk, M. Hausmann, M. Schäfer, I. Falková, J. Toufar and L. Toufarová, in Chromosomal Abnormalities - From DNA Damage to Chromosome Aberrations, ed. S. Soloneski, Intech-Open, Rijeka, 2025, DOI:10.5772/intechopen.1010246.
- R. Scully, A. Panday, R. Elango and N. A. Willis, Nat. Rev. Mol. Cell Biol., 2019, 20, 698–714 Search PubMed.
- O. Jung-Min and M. Kyungjae, Mutat. Res., Genet. Toxicol. Environ. Mutagen., 2022, 873, 503438 Search PubMed.
- E. Mladenov, V. Mladenova, M. Stuschke and G. Iliakis, Int. J. Mol. Sci., 2023, 24, 14956 Search PubMed.
- J. Erenpreisa, K. Salmina, N. M. Vainshelbaum, I. Inashkina and T. Freivalds, Postepy Biochem., 2024, 70, 33–38 CrossRef PubMed.
- J. Erenpreisa, A. Giuliani, K. Yoshikawa, M. Falk, G. Hildenbrand, K. Salmina, T. Freivalds, N. Vainshelbaum, J. Weidner and A. Sievers, et al. , Int. J. Mol. Sci., 2023, 24, 2658 CrossRef CAS PubMed.
- W. D. Wright, S. S. Shah and W.-D. Heyer, J. Biol. Chem., 2018, 293, 10524–10535 Search PubMed.
- A. Shibata and P. A. Jeggo, Br. J. Radiol., 2020, 93, 20190966 Search PubMed.
- G. Iliakis, Radiother. Oncol. J. Eur. Soc. Ther. Radiol. Oncol., 2009, 92, 310–315 CrossRef CAS PubMed.
- M. Falk, M. Hausmann, E. Lukášová, A. Biswas, G. Hildenbrand, M. Davídková, E. Krasavin, Z. Kleibl, I. Falková and L. Ježková, et al. , Crit. Rev. Eukaryotic Gene Expression, 2014, 24, 225–247 Search PubMed.
- M. Lelec, M. T. Gyparaki, G. Beliu, F. Schueder, J. Griffie, S. Manley, R. Jungmann, M. Sauer, M. Lakadamyali and C. Zimmer, Nat. Rev. Methods Primers, 2021, 1, 39 CrossRef PubMed , Erratum in: M. Lelec, M. T. Gyparaki, G. Beliu, F. Schueder, J. Griffie, S. Manley, R. Jungmann, M. Sauer, M. Lakadamyali and C. Zimmer, Nat. Rev. Methods Primers, 2022, 2, 70.
- P. Lemmer, M. Gunkel, Y. Weiland, P. Muller, D. Baddeley, R. Kaufmann, A. Urich, H. Eipel, R. Amberger, M. Hausmann and C. Cremer, J. Microsc., 2009, 235, 163–171 CrossRef CAS PubMed.
- H. Hahn, C. Neitzel, O. Kopecná, D. W. Heermann, M. Falk and M. Hausmann, Cancers, 2021, 13, 5561 CrossRef CAS PubMed.
- S. Chakraborty, M. Singh, R. Pandita, V. Singh, C. Lo, F. Leonard, N. Horikoshi, E. Moros, D. Guha and C. Hunt, et al. , iScience, 2022, 25, 104142 Search PubMed.
- E. Bobkova, D. Depes, J.-H. Lee, L. Jezkova, I. Falkova, E. Pagacova, O. Kopecna, M. Zadneprianetc, A. Bacikova and E. Kulikova, et al. , Int. J. Mol. Sci., 2018, 19, 3713 CrossRef PubMed.
- J. R. Williams, D. S. Gridley and J. M. Slater, in Advances in the Biology, Imaging and Therapies for Glioblast Radiobiology of Radioresistant Glioblastoma, ed. C. Cheng, IntechOPEN, London, UK, 2011, pp. 3–22 Search PubMed.
- J. Weidner, C. Neitzel, M. Gote, J. Deck, K. Küntzelmann, G. Pilarczyk, M. Falk and M. Hausmann, Comput. Struct. Biotechnol. J., 2023, 21, 2018–2034 CrossRef CAS PubMed.
- M. Ester, H.-P. Kriegel, J. Sander and X. Xu, Proc.Sec. Intern. Conf. Knowledge Disc. Data Mining (KDD-96), 1996, 226–231.
- B. D. Ripley, J. R. Stat. Soc., Ser. B, 1977, 39, 172–192 Search PubMed.
- G. Máté and D. W. Heermann, Front. Phys., 2014, 2, 20 Search PubMed.
- A. Hofmann, M. Krufczik, D. W. Heermann and M. Hausmann, Int. J. Mol. Sci., 2018, 19, 2263 Search PubMed.
- R. Ghrist, Bull. Am. Math. Soc., 2008, 45, 61–75 Search PubMed.
- H. Adams, S. Chepushtanova, T. Emerson, E. Hanson, M. Kirby, F. Motta and L. Ziegelmeier, J. Mach. Learn. Res., 2017, 18, 1–35 Search PubMed.
- K. Pearson, London, Edinburgh Dublin Philos. Mag. J. Sci., 1901, 2, 559–557 CrossRef.
- D. Depes, J.-H. Lee, E. Bobkova, L. Jezkova, I. Falkova, F. Bestvater, E. Pagacova, O. Kopecna, M. Zadneprianetc and A. Bacikova, et al. , Eur. Phys. J. D, 2018, 72, 158 Search PubMed.
- T. Vicar, J. Gumulec, R. Kolar, O. Kopecna, E. Pagacova, I. Falkova and M. Falk, Comput. Struct. Biotechnol. J., 2021, 19, 6465–6480 Search PubMed.
- J. R. Munkres, S. G. Krantz and H. R. Parks, Elements of Algebraic Topology in Textbooks in Mathematics, 2nd edn, 2025, ISBN 978-1032765549 Search PubMed.
- P. Jaccard, Bull. Soc. Vaudoise Sci. Nat., 1901, 37, 547–579 Search PubMed.
- C. Adamson, O. O. Kanu, A. I. Mehta, C. Di, N. Lin, A. K. Mattox and D. D. Bigner, Expert Opin. Invest. Drugs, 2009, 18, 1061–1083 CrossRef CAS PubMed.
- Q. T. Ostrom, H. Gittleman, G. Truitt, A. Boscia, C. Kruchko and J. S. Barn-Holtz-Sloan, Neuro-Oncology, 2018, 20, iv1–iv86 CrossRef PubMed.
- K. Rothkamm, I. Krüger, L. H. Thompson and M. Löbrich, Mol. Cell Biol., 2003, 23, 5706–5715 Search PubMed.
- Y. Xu and D. Xu, Essays Biochem., 2020, 64, 765–777 Search PubMed.
- M. Hausmann, G. Hildenbrand and G. Pilarczyk, Results Probl. Cell Differ., 2022, 70, 3–34 Search PubMed.
- G. Zimatore, M. Tsuchiya, M. Hashimoto, A. Kasperski and A. Giuliani, Biophys. Rev., 2021, 2(3), 031303 CrossRef CAS PubMed.
- J. Toufar, L. Toufarová and I. Falková, et al. , Eur. Biophys. J., 2025, 54, 547–572 CrossRef PubMed.
- M. Bayer, J. Zajakina, M. Schäfer, K. Salmina, F. Rumnieks, J. Jansons, F. Bestvater, R. Kurg, J. Erenpreisa and M. Hausmann, Cancers, 2025, 17, 2480 Search PubMed.
- T. T. Paull, E. P. Rogakou, V. Yamazaki, C. U. Kirchgessner, M. Gellert and W. M. Bonner, Curr. Biol., 2000, 10, 886–895 Search PubMed.
- R. Scully and A. Xie, Mutat. Res., 2013, 750, 5–14 Search PubMed.
- M. Hausmann, E. Wagner, J.-H. Lee, G. Schrock, W. Schaufler, M. Krufczik, F. Papenfuß, M. Port, F. Bestvater and H. Scherthan, Nanoscale, 2018, 10, 4320–4331 RSC.
- F. Ochs, K. Somyajit, M. Altmeyer, M. B. Rask, J. Lukas and C. Lukas, Nat. Struct. Mol. Biol., 2016, 23, 714–721 CrossRef CAS PubMed.
- L. Zhang, X. Geng and F. Wang, et al. , Nat. Commun., 2022, 13, 360 Search PubMed.
- M. Falk, E. Lukášová, L. Štefančíková, E. Baranová, I. Falková, L. Ježková, M. Davídková, A. Bačíková, J. Vachelová, A. Michaelidesová and S. Kozubek, Appl. Radiat. Isot., 2014, 83(PtB), 177–185 Search PubMed.
- R. Huang and P. K. Zhou, Signal Transduction Targeted Ther., 2021, 6, 254, DOI:10.1038/s41392-021-00648-7.
- Z. Hou, T. Yu and Q. Yi, et al. , Commun. Biol., 2024, 7, 936, DOI:10.1038/s42003-024-06640-5.
- S. Bao, Q. Wu and R. McLendon, et al. , Nature, 2006, 444, 756–760, DOI:10.1038/nature05236.
- D. Deckbar, P. A. Jeggo and M. Löbrich, Crit. Rev. Biochem. Mol. Biol., 2011, 46, 271–283 Search PubMed.
- M. Falk, E. Lukásová and S. Kozubek, Biochim. Biophys. Acta, Mol. Cell Res., 2008, 1783(12), 2398–2414, DOI:10.1016/j.bbamcr.2008.07.010.
- E. Lukášová, Z. Kořistek, M. Klabusay, V. Ondřej, S. Grigoryev, A. Bačíková, M. Řezáčová, M. Falk, J. Vávrová, V. Kohútová and S. Kozubek, Biochim. Biophys. Acta, Mol. Cell Res., 2013, 1833(3), 767–779 CrossRef PubMed.
- A. Granzotto, L. El Nachef, J. Restier-Verlet, L. Sonzogni, J. Al-Choboq, M. Bourguignon and N. Foray, Biomolecules, 2024, 14, 703 CrossRef CAS PubMed.
- C. Wu, J. C. Manjarrez-González, M. Choudhury, N. Shamkhi, S. Ding, V. M. Nair and V. M. Draviam, Cells Rep. Methods, 2025, 5(8), 101118 CrossRef CAS PubMed.
- M. Falk, I. Falková and O. Kopečná, et al. , Sci. Rep., 2018, 8, 14694 CrossRef PubMed.
- M. Falk, Z. Horakova and M. Svobodova, et al. , Eur. Phys. J. D, 2017, 71, 241 CrossRef.
- M. J. Clark, N. Homer, B. D. O'Connor, Z. Chen, A. Eskin, H. Lee, B. Merriman and S. F. Nelson, PLoS Genet., 2010, 6(1), e1000832 CrossRef PubMed.
- Z. H. Chen, M. Zhu, J. Yang, H. Liang, J. He, S. He, P. Wang, X. Kang, M. A. McNutt, Y. Yin and W. H. Shen, Cell Rep., 2014, 8(6), 2003–2014 CrossRef CAS PubMed.
- D. Hanisch, A. Krumm and T. Diehl, et al. , Cell Death Dis., 2022, 13, 293 Search PubMed.
- J. Wang, T. Y. Huang, Y. Hou, E. Bartom, X. Lu, A. Shilatifard, F. Yue and A. Saratsis, Sci. Adv., 2021, 7, eabg4126 CrossRef CAS PubMed.
- A. Sharda, T. Verma, N. Gadewal and S. Gupta, Dynamic Alterations of Histone H3 Phospho- acetylation Correlate With Radio Sensitivity of Mitotic Cells During DNA Damage, 2020, DOI:10.21203/rs.3.rs-80009/v1 (preprint).
- V. S. Tanwar, C. C. Jose and S. Cuddapah, Mutat. Res., Rev. Mutat. Res., 2019, 780, 61–68 CrossRef CAS PubMed.
- P. L. Collins, C. Purman and S. I. Porter, et al. , Nat. Commun., 2020, 11, 3158 CrossRef CAS PubMed.
- M. H. Bailey, C. Tokheim, E. Porta-Pardo, S. Sengupta and D. Bertrand, et al. , Cell, 2018, 173(2), 371–385, DOI:10.1016/j.cell.2018.02.060.
- T. Ersahin, N. Tuncbag and R. Cetin-Atalay, Mol. Biosyst., 2015, 11(7), 1946–1954, 10.1039/c5mb00101c.
- P. L. Sulkowski, S. Oeck and J. Dow, et al. , Nature, 2020, 582, 586–591 CrossRef CAS PubMed.
- S. C. Short, C. Martindale, S. Bourne, G. Brand, M. Woodcock and P. Johnston, Neuro-Oncology, 2007, 9(4), 404–411 CrossRef CAS PubMed.
- A. Sinha, A. Saleh, R. Endersby, S. H. Yuan and C. R. Chokshi, et al. , Cancers, 2020, 12(11), 3178 CrossRef CAS PubMed.
- C. Xie, H. Wang, H. Cheng, J. Li, Z. Wang and Y. Wu, Biochem. Biophys. Res. Commun., 2014, 445(1), 263–268 CrossRef CAS PubMed.
- K. Watanabe, K. Iwabuchi, J. Sun, Y. Tsuji and T. Tani, et al. , Nucleic Acids Res., 2009, 37(7), 2176–2193 Search PubMed.
- M. Palek, N. Palkova, consortium CZECANCA, P. Kleiblova, Z. Kleibl and L. Macurek, Nucleic Acids Res., 2024, 52(13), 7687–7703 CrossRef CAS PubMed.
- Y. Wang, Y. Hou and W. Zhang, et al. , J. Exp. Clin. Cancer Res., 2019, 38, 147 Search PubMed.
- H. M. Chen, A. Nikolic, D. Singhal and M. Gallo, Cancers, 2022, 14(19), 4942 CrossRef CAS PubMed.
- J. Han, Y. Jun, S. H. Kim, H. Hoang and Y. Jung, et al. , Proc. Natl. Acad. Sci. U. S. A., 2016, 113(50), 14283–14288 CrossRef CAS PubMed.
- J. Sevcik, M. Falk, P. Kleiblova, F. Lhota and L. Stefancikova, et al. , Cell. Signalling, 2012, 24(5), 1023–1030 CrossRef CAS PubMed.
- J. Sevcik, M. Falk, L. Macurek, P. Kleiblova and F. Lhota, et al. , Cell. Signalling, 2013, 25(5), 1186–1193 Search PubMed.
- B. Han, X. Meng, H. Chen, L. Chen, X. Liu, H. Wang, D. Liu, F. Gao, L. Lin, J. Ming, B. Sun, S. Yin and R. Wang, et al. , Aging, 2017, 9, 1885–1897 CrossRef CAS PubMed.
- L. Xia, K. Qin, X. R. Wang, X. L. Wang, A. W. Zhou, G. Q. Chen and Y. Lu, Oncotarget, 2017, 8(65), 109120–109134 CrossRef PubMed.
- D. Hanisch, A. Krumm and T. Diehl, et al. , Cell Death Dis., 2022, 13, 293 CrossRef CAS PubMed.
- J. B. Vilar, M. Christmann and M. T. Tomicic, Cancers, 2022, 14(10), 2416 Search PubMed.
- M. Scholz, W. Kraft-Weyrather, S. Ritter and G. Kraft, Int. J. Radiat. Biol., 1994, 66, 59–75 Search PubMed.
- S. Sora, N. Hamada, T. Hara, T. Funayama, T. Sakashita, Y. Yokota, T. Nakano and Y. Kobayashi, Biol. Sci. Space, 2008, 22, 54–58 CrossRef.
- K. Tsuboi, T. Moritake, Y. Tsuchida, K. Tokuuye, A. Matsumura and K. Ando, J. Radiat. Res., 2007, 48, 317–325 CrossRef CAS PubMed.
- M. Bach, C. Savini, M. Krufczik, C. Cremer, F. Rösl and M. Hausmann, Int. J. Mol. Sci., 2017, 18, 1726 CrossRef PubMed.
- J. Schwarz-Finsterle, H. Scherthan, A. Huna, P. González, P. Müller, E. Schmitt, J. Erenpreisa and M. Hausmann, Mutat. Res., 2013, 756, 56–65 CAS.
- H. Scherthan, B. Geiger, D. Ridinger, J. Müller, D. Riccobono, F. Bestvater, M. Port and M. Hausmann, Biomolecules, 2023, 13, 1518 CrossRef CAS PubMed.
- D. Tsao, P. Kalogerinis, I. Tabrizi, M. Dingfelder, R. D. Stewart and A. G. Georgakilas, Radiat. Res., 2007, 168, 87–97 CrossRef CAS PubMed.
- M. Schäfer, G. Hildenbrand and M. Hausmann, Int. J. Mol. Sci., 2024, 25, 12843 CrossRef PubMed.
- L. Ježková, M. Falk, I. Falková, M. Davídková, A. Bačíková and L. Štefančíková, et al. , Appl. Radiat. Isot., 2014, 83, 128–136 CrossRef PubMed.
- D. Depes, J. Lee and E. Bobkova, et al. , Eur. Phys. J. D, 2018, 72, 158 CrossRef.
- M. Hausmann, C. Neitzel, E. Bobkova, D. Nagel and A. Hofmann, et al. , Front. Phys., 2020, 8, 578662 Search PubMed.
- M. Hausmann, M. Falk, C. Neitzel, A. Hofmann, A. Biswas, T. Gier, I. Falkova, D. W. Heermann and G. Hildenbrand, Int. J. Mol. Sci., 2021, 22, 3636 Search PubMed.
- M. Falk and M. Hausmann, Cas. Lek. Cesk., 2020, 159(7–8), 286–297 Search PubMed.
- P. Matula, M. Maška, O. Danek, P. Matula and M. Kozubek, Proc – IEEE Int. Symp. Biomed. Imag. Nano Macro ISBI, 2009, 1138–1141.
- H. Scherthan, J.-H. Lee, E. Maus, S. Schumann, R. Muhtadi, R. Chojowski, M. Port, M. Lassmann, F. Bestvater and M. Hausmann, Cancers, 2019, 11, 1877 CrossRef CAS PubMed.
- M. Roushenas, M. Salerno and V. Bazzurro, et al. , Eur. Biophys. J., 2025 DOI:10.1007/s00249-025-01770-y.
- F. Grüll, M. Kirchgessner, R. Kaufmann, M. Hausmann and U. Kebschull, Proc. – 21st Int. Conf. Field Program. Logic Appl, FPL, 2011, pp. 1–5.
Footnote |
| † These authors share last authorship. |
| This journal is © The Royal Society of Chemistry 2026 |