 Open Access Article
Open Access ArticleMechanism of polypeptide translocation through gold nanopores in view of sequencing applications
Mirko
Vanzan
 ab,
Agostino
Migliore
ab,
Agostino
Migliore
 *a,
Maria
Blanco-Formoso
*a,
Maria
Blanco-Formoso
 *c,
Francesco
De Angelis
*c,
Francesco
De Angelis
 c and
Stefano
Corni
c and
Stefano
Corni
 *ad
*ad
aDepartment of Chemical Sciences, University of Padova, Via Marzolo 1, 35131 Padova, Italy. E-mail: agostino.migliore@unipd.it; stefano.corni@unipd.it
bDepartment of Physics, University of Milan, Via Celoria 16, 20133 Milano, Italy
cIstituto Italiano di Tecnologia, Via Morego 30, 16163, Genova, Italy. E-mail: Maria.BlancoFormoso@iit.it
dCNR Institute of Nanoscience, 41125 Modena, Italy
First published on 10th February 2026
Abstract
A crucial aspect of the label-free sequencing of peptides and single proteins in solid-state nanopores via optical methods is the ability to control the translocation dynamics of the biomolecule, especially its speed. Very often, this dynamics is studied in terms of its effect on ionic currents through the nanopore. Herein, with attention to label-free optical sequencing methods, we directly study the translocation motion of (poly)peptides by molecular dynamics. By analysis of a vast set of simulations of polyglutamic acids, we show that the peptide elongation is determined by the electrostatic repulsion between the side chains, with less dependence on ionic strength and a more prominent effect of ion type, which emerges as a factor to control the peptide elongation for sequential amino acid detection. Instead, the ionic strength influences the speed of translocation under driving electrostatic fields. Through comparative analysis of simulations with and without confinement of the peptide in a gold nanopore, we quantify the influence of the nanopore on the sequential transit of amino acids and clarify the role of the peptide–pore interaction in promoting the peptide elongation and slowing down their translocation. We identify a stop-and-go translocation mechanism that can be controlled by lateral electric fields, such as in experimental “hot spots”, to achieve translocation velocities adequate for single-amino acid detection, while the use of appropriate ions also favors elongated peptide poses suitable for single-amino acid detection. We also present experiments on polyglutamic acid translocation which, compared with the theoretical results, turn out to be compatible with the stop-and-go translocation mechanism. The translocation mechanism, which is characterized by the proximity of the peptide to the nanopore surface, raises expectations for the promising use of plasmonic hot spots in single-amino acid detection.
1. Introduction
The sequencing of the primary structure of peptides and proteins is of fundamental importance for proteomics1–3 and its medical applications, as even subtle alterations to the primary structure of a protein can lead to serious pathological conditions.4–7 The development of effective, commercially feasible methods for sequencing the genome (DNAs), the transcriptome (RNAs), and the proteome (the entire set of proteins) is critically important to advance precision and personalized medicine.8,9 The importance of these methods has been further highlighted by the recent research on COVID-19,10 as well as the associated outstanding prospects for the development of mRNA-based vaccines and immunotherapies for cancer and infectious diseases.11Remarkable advances in molecular biology and nanobiotechnology over the past two decades have drawn growing attention to nanopore sensing techniques for highly sensitive and versatile investigation of biomolecules. The great progress in nanopore-based techniques for detecting and sequencing nucleic acids12–30 has boosted the development of different nanopore-based techniques for the conformational analysis and sequencing of peptides and proteins,1–3,29–67 with the potential to perform label-free sequencing or at least to overcome the known limitations of traditional methods for protein analysis.68–73 In particular, surface-enhanced Raman scattering (SERS) is a powerful spectroscopic tool for label-free and sensitive analysis at the single-molecule level, especially in conjunction with nanoscale particles and nanopores where plasmonic effects can be effectively enhanced (also inside the nanopore, thanks to the electronic properties at its surfaces, whether external or internal).3,8,74–80 SERS-based sequencing can support the use of solid-state nanopores, overcoming the limitations of biological nanopores in terms of chemical and mechanical stability,67 while compensating for the loss in chemical specificity of the nanopore through the possibility of achieving higher sensitivity with SERS detection compared to other approaches to detection based on current measurements (yet, notable progress of the latter approaches is to be mentioned3,47).
Although very promising, nanopore-based protein sequencing techniques with spectroscopic (e.g., SERS) reading still face key challenges. One is the proper unfolding of the proteins to enable the sequential passage of amino acids through a nanopore for their clear reading,3,32,39,73,81–84 and another is the related difficulties in controlling the protein motion through nanopores (beginning with the translocation speed67), which are exacerbated by the number and different charges of the amino acids.3,40 To address these issues and thus gain control of the translocation dynamics, as is especially required in sequencing applications, it is necessary to gain a deeper understanding of the protein translocation mechanisms and their dependence on pore fabrication, application of external fields,67,85–87 and environmental conditions. This motivates the growing experimental and theoretical investigations of peptide and protein motion through nanopores,56,67,81,83,87–97 and an increasing role for molecular dynamics (MD) simulations to inquire the determinants of molecular translocation (useful discussions and a rich bibliography can be found in ref. 67, 87 and 98). These studies benefit from earlier investigations of ion channels99 and polymers (in particular, DNA) translocation.100–110 Despite tremendous advances in this research area, a unified theory of polymer translocation is not yet available. This is also due to the dependence of the translocation on structural factors that vary between different biomolecular chains and include the interaction of the biomolecule with the pore, the interaction with the solvent, and therefore the characteristics of the latter. In fact, the translocation of biomolecules does not follow any universal scaling law.67 In this regard, the specific investigation of protein translocation is essential, despite many previous studies on nucleic acids. Furthermore, studies on translocation through metallic nanopores have been very scarce to date, unlike those on biological and other (insulating) solid-state nanopores.
To enable proper sequencing of proteins through nanopores, one should be able to control the protein translocation speed and then tailor it to the sequencing data acquisition time. The other key challenge is understanding how much the protein backbone needs to be extended in order to allow for the sequential identification of different amino acids. The analysis of these two points is further complicated by the fact that the driven translocation of polymers (and even more so that of biological heteropolymers) is not an equilibrium process because of the long relaxation times that characterize the polymer dynamics.85,111 The study of the translocation of homopeptides emerges from this state of affairs as a first main step to identify the basic aspects of protein dynamics through nanopores.
The translocation of biomolecules through pores is commonly studied in terms of variation or blockage of the ionic current caused by the passage of the macromolecule, as is relevant to the sensing of the molecular structure and properties by current measurement. Where the label-free detection of a protein sequence is used, it is advantageous to directly study the dynamics of a (poly)peptide chain as it interacts with the nanopore, the solution, and external fields. This is our approach, which uses MD simulations to study protein dynamics in bulk solution and inside a gold nanopore.
As a probe molecule to validate the in silico results, we chose polyglutamic acid because of its chemical and physical stability, ease of use, and commercial availability. In common saline buffers, polyglutamic acid is negatively charged, thus limiting the formation of random coils. We performed the experiments on a 5000-unit-long molecule to facilitate monitoring of the events via electrical measurements.
The use of gold reflects the common experimental choice of noble metals such as gold and silver to build nanopores and nanoparticles for use in visible-range plasmonics, and their potential for use in label-free optical sensing.3,30,77,112–120 Moreover, their greater structural stability and durability compared to that of biological nanopores can be complemented with enhanced chemical properties conferred by suitable surface functionalization, which can be performed in a site-selective way by exploiting plasmonic effects.121–125 Difficulties are still faced in the proper shaping of solid-state pores at the nanoscale, but the past decade has witnessed great advances in nanopore fabrication.67,121 In fact, the nanopore size simulated in our study is within the range of feasibility (vide infra). Moreover, the extensive use of Au nanoparticles and electrodes in experimental molecular electronics has progressed in parallel with advances in their theoretical study,126 including the development of force fields for their MD simulation.127–131 In particular, Au(111) has been the most studied gold surface for peptide and protein binding, both theoretically and experimentally, and is known for its chemical stability.3,130,132–134 In fact, Au(111) is the most used gold surface in nanobioelectronic studies, and particularly in systems for optical detection that use surface plasmon resonance.135–139
Here, we first study the peptide motion in aqueous solution head-on (sections 3.1 and 3.2), analyzing the dependence of its structural and dynamical properties on the type of counterion and ionic strength, the presence of other molecular species in solution (in particular, urea), and the application of electrostatic fields of different intensities, with consequent electrophoretic effects (section 3.3). Two different polymer lengths are also considered to ascertain the validity and scaling of our mechanistic conclusions with the system size, and thus their applicability to the longer polypeptides that make up proteins. The study is then conducted in the presence of a gold nanopore (section 3.4), in order to model the translocation of the real system and gain insight into the effects of the confinement and double layer formation on the biomolecular motion by comparison with the dynamics in the absence of the nanopore. To this aim, our analysis includes MD simulations with different charge densities on the nanopore surface. Our MD analysis explains the non-equilibrium translocation dynamics that, apart from the slow relaxation processes typical of polymers,67,85,111 are determined by the interplay between driving electrostatic field and interaction with the surface of the nanopore, so as to determine the transition between different regimes of translocation motion depending on the surface polarization charge. The study of this interplay in different conditions of solution and ionic force, as well as a convenient description of the spatial distinguishability of the different glutamic acid residues also helps us understand the balance of electrophoretic and driving forces that can be used to control the translocation speed, as is necessary for the detection of single amino acid residues using SERS nanopores. Finally, the feasibility of the translocation mechanism emerging from the theory is tested with experiments on polyglutamic acid motion through a gold nanopore (section 3.5), and the distinguishability of the residues during the peptide translocation required for sensing is analyzed (section 3.6). It is worth noting that, although our analysis is focused on peptides and can be used on elongated proteins, some mechanistic conclusions may be valid for other translocating polymers, including DNA.
2. Methods
The methods described below were primarily employed to study a peptide with 10 glutamic acid residues (10GLU) interacting with different counterions in an aqueous solvent at different ionic strengths, in the presence and absence of a gold nanopore and under different electrostatic fields (two electrostatic fields in the longitudinal direction of the nanopore, with intensities of 0.5 V nm−1 and 1.0 V nm−1, and transversal potential differences through the solvent-nanopore interface from 0 to 260 mV). Some MD simulations were also repeated with other field values (see SI) and with a longer polypeptide (20GLU) to examine possible effects of the peptide length on its structural and dynamical properties. Table S1 in the SI summarizes all simulations performed and analyzed below.2.1. Design and construction of model systems
Our analysis is based on classical MD simulations over a timescale ranging from 0.5 μs to a few μs. Unless otherwise specified, the simulated molecule is a glutamic acid peptide consisting of 10 residues, denoted here as 10GLU (see Fig. 1a), in aqueous solution (either free or confined in a gold nanopore). The use of a homopeptide deliberately sets the worst-case scenario for exploring the possibility of physically distinguishing the passage of successive amino acid residues at a hotspot in the metal nanopore, since such residues are of the same type. This peptide was generated using standard structural parameters for amino acid alpha helices, implemented in the PeptideBuilder software.140 The secondary structure was lost in a few simulation steps, and therefore far before the MD production runs. The chosen peptide length is a compromise between accuracy and feasibility, allowing us to perform a vast set of MD simulations while maintaining chain properties, such as the propensity to randomly coil (also observed in the simulations), that characterize the polypeptide that would result from the unfolding and elongation of a protein to be sequenced. | ||
| Fig. 1 Structures of the peptide and Au nanopore studied. (a) 10GLU peptide capped with methyl groups (20GLU is similarly capped). (b) Side view of the icosagonal-base pore with inner Au(111) surface. The peptide structure in an MD snapshot is drawn (an electrostatic field of 0.5 V nm−1 was applied along the axis of the nanopore during the MD). The Au-like atoms yielding the Au polarization properties and the solution surrounding the peptide are omitted. (c) Top view of the system. All figures with molecular representations were rendered with VMD.141 | ||
Capping of the N- and C-termini of the peptide with methyl groups prevents their spurious direct interaction, which would be very unlikely in long polypeptides. The results obtained using the 10GLU system are further supported by simulations of a system with 20 glutamic acid residues, 20GLU (vide infra), again constructed using PeptideBuilder.
The gold nanopore used in the simulations has the shape of the lateral surface of an icosagonal prism. In fact, it was built from 20 slabs of Au atoms, each consisting of 5 atomic layers and having two Au(111) surfaces (Au(111) is the most stable among gold surfaces), with a lattice constant of 2.93 Å, a width of 8.80 Å, and a length of 5.075 Å. Each slab contains extra atomic species that account for Au polarization, electrostatic interactions, and chemisorption effects.130
The gold slabs expose one Au(111) surface to the interior of the nanopore. The large number of slabs in the icosagonal nanopore surface makes adjacent slabs roughly parallel to each other, so as to approximately retain a local Au(111) structure at the joints between the slabs (see Fig. 1b), consistent with the Au force field130 used for MD simulation. For the same purpose, the nanopore was assembled so that neighboring gold atoms from different slabs are at the same distance as the analogous atomic pairs from the same slab. The deviation between the slabs is more appreciable on the outer surface of the nanopore, but the simulations were performed with fixed Au atoms and all systems moving in the MD simulation were inside the pore.
The resulting nanopore diameter is 5.6 nm, which is within the range of current feasibility.3,30,47,142 Although pore diameters no greater than 3–4 nm would be ideal for protein sequencing, diameters of 10–20 nm are more routinely achieved and their appropriate shaping can enable the creation of plasmonic hot spots with much smaller linear sizes (e.g., ∼3 nm in ref. 143). It is worth noting that pore diameters of 3–4 nm, important for achieving strong Raman enhancements, still allow folding of the peptide chain, but unfolding can be maintained by chemical means. Since we use short polypeptides in our analysis, another issue might arise if they do not line up with the nanopore surface. However, as is shown below, despite the short length of 10GLU, its propensity to stay near the surface of the Au nanopore makes our analysis meaningful, and this consideration is further supported by the additional study of 20GLU's behavior.
The length of the nanopore used in the MD simulation is infinite, as it goes from one side to the other of the simulation cell and periodic boundary conditions are applied. This is not an issue in our analysis, as it is not focused on edge SERS and plasmonic enhancement effects themselves.
MD simulations were performed using solutions with different ionic strengths. Systems with 0 M solutions were obtained by adding 10 and 20 counterions to neutralize the carboxylates of 10GLU and 20GLU, respectively. Considering the size of the MD simulation cell (see next subsection), as well as the fact that GLU has a pKa of 4.1 ± 0.8 (ref. 144) (and therefore its carboxyl group is negatively charged at neutral pH), the 0.5 M solutions of 10GLU in the absence of the nanopore were obtained by adding MCl salts (that is, practically, Cl− and M+ = Na+, Li+, K+), so as to have a total of 29 counterions. The same number of ions was used for the higher-molarity (namely, 0.3 M) solutions of 10GLU in the nanopore. Considering the cell size on the z direction in the simulations with the nanopore (this size coincides with the length of the pore) and reducing the geometric radius of the pore by the van der Waals radius of Au atoms (166 pm) to calculate the effective radius of the Au nanopore, the use of 29 counterions led to a molarity of 0.3 M for the solvated system simulated inside the nanopore.
2.2. MD simulations
Classical MD simulations were performed using the GROMACS package, version 2018.4.145–153 As usual, the intramolecular bonds and dihedral angles were parametrized as harmonic oscillators, and intermolecular forces were modeled using the Lennard-Jones potential. In particular, the OPLS-all atom force field154–156 was used to describe all interactions, while water molecules were parameterized according to a refined single-point-charge (SPC) model.157To model the interaction of the gold nanopore with the solvated molecular system, we used the GolP force field of Iori et al.,130 which requires the use of two additional (fictitious) atomic species to describe the electrostatic response of metal atoms through a dipole rod-model approach (“AUC” atomic species) and to account for surface chemisorption effects (“AUI”).
The pair list was generated by a Verlet algorithm. The electrostatic potentials were calculated by using a fast smooth Particle Mesh Ewald method, and the distance for cutoff of the direct-space Coulomb interactions was set to 1 nm. The same cutoff was used for the Lennard-Jones potential. Energy and pressure long-range dispersion corrections were applied.
The initial conformations of the 10GLU and 20GLU polypeptides for their simulations in the presence of the pore were drawn from the last MD snapshots in the absence of the nanopore. Each polypeptide was initially positioned along the nanopore axis, around its center.
The geometries of all systems were initially optimized with 5 × 104 minimization steps through the steepest descent algorithm. Equilibration in temperature followed, using a leap-frog algorithm to integrate Newton's equations of motion. The hydrogen bonds were constrained with the LINCS algorithm.158 The temperature coupling was performed via the Bussi–Donadio–Parrinello velocity rescaling with a stochastic term,159 with a time constant of 0.5 ps. The initial Maxwell velocity distribution was set to that for 10 K, and the systems were allowed to equilibrate at a temperature of 298.15 K. The equilibrations with respect to temperature lasted 250 ps in the absence of the Au nanopore and 500 ps for the systems in the nanopore. These were reasonable choices considering that the gold atoms were fixed and the molecular systems were relatively small. In both cases, the integration timestep was set to 0.5 fs, and the neighbor list was updated every 20 timesteps.
The pressure was also equilibrated in the next 250 ps of simulation, with a 1 fs timestep. Velocity generation was not used at this stage. The system was coupled to a Parrinello–Rahman barostat160 with a time constant of 10 ps. The pressure was set to 1 bar and the compressibility to the value of 4.5 × 10−5 bar−1 (which is typical of water at a temperature of 300 K and atmospheric pressure). The following MD production runs (at constant temperature and pressure) used a timestep of 2 fs and lasted 500 ns unless otherwise specified.
The pressure control was not used for the simulations in the presence of the nanopore. In fact, as the pore atoms were fixed, these simulations were performed using the NVT ensemble and the pressure inside the Au nanopore was controlled through the density of the water molecules, which were added and monitored until their density was similar to that in bulk water (more precisely, the average water density for the same water force field in the absence of the nanopore was used as the reference value).
The cell size depended on the molecular system. The cell vectors (in nm) were (4, 4, 4) for the simulation of the solvated 10GLU system and (6, 6, 6) for 20GLU, which allowed for a separation between the peptide and its replicas by at least 1.5 nm. In the NPT simulations that were performed in the absence of the pore, these clearly were the initial cell sizes. In the presence of the Au nanopore, the unit cell vectors were (9.2, 9.2, 5.075) nm to substantially prevent replica interactions on the xy plane. The match between the cell size in the z direction and the nanopore length mimicked an infinitely long nanopore, i.e., in practice, a nanopore much longer than the peptide. This choice enabled the study of the peptide translocation motion inside the pore without the complicacies of the dynamics at the entrance and exit of the pore.
2.3. MD analysis
GROMACS tools were used to obtain time-dependent descriptors of the evolving peptides, that is, the root mean square deviation (RMSD), the radius of gyration (Rg), and the solvent accessible surface area (SASA). GROMACS tools were also used to obtain radial distribution functions, the profile densities, and the peptide center-of-mass (CM) coordinates. The latter were used to evaluate the global translocation velocity of the peptide. The CM coordinates were stored every τ = 10 ps. Hence, the coarse-grained instantaneous speed of the CM was obtained as δzCM/τ, where δzCM is the variation of zCM over τ. To remove the correlations between data points and to associate an error with the mean velocity of the CM, we used the simplest blocking approach.161 The data points were grouped in blocks of increasing size. Then, for each block size and corresponding number of blocks, the mean velocities in the blocks and the associated standard deviations were calculated and used as independent data points. The standard error associated with the global average velocity was then calculated. The standard error is expected to fluctuate around a plateau as the block size increases sufficiently. Since the selected number of MD snapshots allowed a rather small number of divisions in blocks of the same size (and therefore the calculation of a correspondingly small number of standard errors), we used the largest standard error in the approximate plateau region as a reasonably safe estimate of the error on the mean speed (see examples in SI Fig. S8).To investigate the elongation of the peptide along the nanopore axis, and hence the possibility of individually detecting amino acid residues to sequence the peptide, we analyzed the overlap of amino acid residues along the z axis, as observed perpendicularly to this axis. The overlap was quantified using the heavy atoms of the side chains, which are important for the discrimination of the different amino acids in a protein. After dividing the simulation cell into 100 slices along z, we removed the translation of the first amino acid between subsequent selected MD snapshots and calculated the average portion of its side chain contained in each slice, thus defining the set of slices with nonzero occupation by the side chain of the first amino acid residue. Then, we computed the average portion (that is, a generally fractional number of atoms) of each of the other side chains contained in this set of slices and thus overlapped with the first side chain. We proceeded similarly by taking as a reference each of the next amino acid side chains in the peptide sequence. After that, we averaged the numbers of side chain atoms overlapping with next neighbor side chains, second-next neighbors, etc.
2.4. Surface potential and diffuse layer
We also studied the dependence of the peptide translocation speed on the potential difference Vt between the Au surface and the bulk solution.We used Gouy–Chapman's theory to calculate the extra charge on each Au atom necessary to obtain the desired transversal electrostatic potential difference, that is, the potential drop across the diffusion layer. Since the peptide was solvated by aqueous solutions at 25 °C, we used the relation
 | (1) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) z electrolyte (it is z = 1 here) in mol L−1 (eqn (1) is equation 13.3.20b on p. 550 of ref. 162, which is valid for dilute aqueous solutions at 25 °C).163
z electrolyte (it is z = 1 here) in mol L−1 (eqn (1) is equation 13.3.20b on p. 550 of ref. 162, which is valid for dilute aqueous solutions at 25 °C).163
2.5. In vitro experimental section
The gold nanopores were realized following the procedure described in ref. 143 upon minor modifications, as noted here. The chip (Si3N4 window: 0.5 × 0.5 mm2 and SiO2 frame: 1 × 1 cm2, thickness: 100 nm) was covered with a 5 nm-thick layer of Ti and a 20 nm layer of Au (Kenosistec KE500ET conjugated e-beam/thermal evaporator), and patterned with 6 holes with a diameter of 20 nm using a focusing ion beam scanning electron microscope (FEI Helios NanoLab 650, voltage: 30 kV, current: 0.24 nA). The diameter of the pore was reduced to 10 nm by sputtering another 10 nm-thick gold layer.3. Results and discussion
3.1. Structural dynamics of free solvated peptides
MD simulations performed in the absence of the nanopore and of an applied electrostatic field aimed to understand the structural and dynamical properties of solvated peptides per se and as compared to those inside the gold nanopore (vide infra), so as to gain a deeper understanding of the direct and indirect effects of the interaction with the pore on such properties.Fig. 2 describes the dynamical evolution of 10GLU in water, where sodium ions neutralize the overall system charge. As usual, the root mean square deviation (RMSD) is used as the main quantity to assess the achievement of an equilibrated structural dynamics. Fig. 2a shows that the time evolution of the RMSD closely correlates with the other quantities studied. In fact, after about 50 ns, the RMSD fluctuates around a plateau, while the radius of gyration (Rg), the maximum atomic distance in the peptide (dmax) and the solvent accessible surface area (SASA) approximately stabilize at values significantly larger than the initial ones, thus implying an elongation of 10GLU and its consequent wider exposure to the solvent. The variations of these geometric parameters correspond to thermal fluctuations of a random coil structure typical of polyglutamic acids in neutral-pH solutions.164,165 For example, the negative spike that is clearly visible in the SASA around 230 ns of MD simulation corresponds to a transient coiling of the peptide to a rather compact structure of linear dimensions similar to the initial ones (see Fig. 2b). However, the mean values and standard deviations of all structural parameters after 50 ns show that 10GLU generally takes conformations that are much more elongated than a globular or alpha helix structure. This is particularly clear from the fact that dmax is, on average, significantly larger than 2Rg; see also the approximately linear correlation between dmax and Rg in SI Fig. S1. The random-coil nature of the polypeptide fluctuations is consistent with the findings of the study in ref. 166, despite the much smaller amount of water and much shorter MD simulations (i.e., 1 ns of production run) in ref. 166.
We find a similar dynamics for the 20GLU polypeptide, although, as expected, the larger size of this system compared to 10GLU makes the alpha helix-to-random coil transition slower. The compact poses rarely observed for 10GLU (which correspond to drops in the SASA) are even more disfavored in the case of 20GLU (Fig. S2). It is not surprising that the alpha helix-to-random coil transition is not observed at room temperature in ref. 166, where the MD sampling of the system, and thus the pertinent energy analysis, is limited to 1.5 ns, against the timescale for conformational change of about 150 ns shown in Fig. S2. On the whole, the comparison between the dynamics of the two polyglutamic acids supports our use of 10GLU as the benchmark amino acid chain.
3.2. Free solvated peptide: peptide-ion interaction
Given the negative charge on the carboxylates of the amino acids, the electric interactions of the (poly)peptides with the solvent and the ionic species dissolved in it are expected to influence their static and dynamic structural properties. In turn, understanding and controlling these properties is critical to protein sequencing.Through radial distribution function analysis (see SI section S4), we find that, in the MD simulation with a solution neutralized by Na+ counterions (the same as in Fig. 2), each of the ten carboxylic groups is on average coordinated by 0.16 Na+ ions, namely, 1 or 2 COO− groups engage in significant interactions with Na+ ions during the MD. This relatively small coordination number suggests that the elongation of the peptide is primarily driven by the decrease in Coulomb repulsion between the negatively charged carboxylates because of their distancing and their screening by the water solvent. A similar conclusion is reached for the 20GLU system, as the average coordination number of the COO− groups with respect to sodium ions is 0.17. This result further confirms the adequacy of the 10GLU model used throughout the following analysis.
To study the effects of the nature and concentration of the dissolved ions on the dynamics of 10GLU, we performed MD simulations with Li+ and K+ counterions, as well as simulations using the three ionic species with an ionic strength of 0.5 M, as obtained by adding suitable amounts of the corresponding chloride salts (see Methods) to the solutions already containing the counterions necessary to neutralize the molecular charge. Fig. 3 shows the time evolutions of the structural descriptors for the different systems, while their statistical analysis is reported in Table 1.
 | ||
| Fig. 3 Evolution of RMSD, Rg, and SASA along the MD simulation of 10GLU in solutions with the indicated counterions and NS (black) or 0.5 M in added salt (blue). The black traces in the central panels are the same as the respective ones in Fig. 2. | ||
| Ion | Li+ | Na+ | K+ | |||
|---|---|---|---|---|---|---|
| I (M) | NS | 0.5 | NS | 0.5 | NS | 0.5 |
a For the system with Li+ ions, the mean values in parentheses refer to the MD time ranges 50–200 ns (left) and 200–500 ns (right), as identified by the RMSD evolution, although Rg and SASA are not at equilibrium in the second range. I refers to the ionic strength of MCl salt added to the solution already containing the M (![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Li, Na, and K) ions that balance the peptide charge. The mean values and standard deviations for the other quantities are obtained excluding the first 50 ns of MD simulation.
b This value larger than unity means that, on average, each COO− group is coordinated to more than one Li+ (see SI Fig. S4). Li, Na, and K) ions that balance the peptide charge. The mean values and standard deviations for the other quantities are obtained excluding the first 50 ns of MD simulation.
b This value larger than unity means that, on average, each COO− group is coordinated to more than one Li+ (see SI Fig. S4).
|
||||||
| RMSD (nm) | 0.40 ± 0.13 (0.22/0.49) | 0.27 ± 0.07 | 0.65 ± 0.03 | 0.65 ± 0.04 | 0.61 ± 0.04 | 0.65 ± 0.04 |
| R g (nm) | 0.66 ± 0.04 (0.62/0.68) | 0.63 ± 0.02 | 0.91 ± 0.08 | 0.89 ± 0.08 | 0.93 ± 0.07 | 0.92 ± 0.07 |
| SASA (nm2) | 14.6 ± 0.6 (14.3/14.7) | 14.2 ± 0.5 | 18.0 ± 0.6 | 17.8 ± 0.7 | 18.1 ± 0.5 | 18.1 ± 0.5 |
| n ion–COO− | 1.0 | 1.6b | 0.16 | 0.25 | 0.06 | 0.11 |
The differences in the behaviors of the systems with Na+ and K+ counterions are secondary with respect to what emerges from the comparison with the Li+-containing systems. Nevertheless, we can appreciate that the relaxation to an equilibrium conformation (as is identified by fluctuations of the RMSD around an approximately constant value) is faster in the K+ system than in the Na+ one, in agreement with a previous study of 12GLU.167 This fact may argue in favor of using Na+ rather than K+ ions to have a slower dynamics in sequencing experiments, although its specific relevance and helpfulness to sequencing deserves a future dedicated investigation. At any rate, in our subsequent analysis of 10GLU in the presence of the nanopore, Na+ ions are used.
Using Rg as a measure of peptide extension, we see that the Li+ solutions lead to significantly more compact peptide structures, and accordingly reduced SASAs, compared to the solutions with the Na+ and K+ species. Furthermore, in the left panels of Fig. 3, the abrupt changes in the three descriptors around 200 ns exemplify the multiple timescale of peptide dynamics (which, in general, is also responsible for complications in the translocation dynamics of polymers driven by external fields67), with the possible occurrence of conformational transitions on a hundred nanosecond time scale. In particular, the top-left panel shows an abrupt increase in the RMSD of the Li+-containing system, which suggests an appreciably different conformation of the peptide. The comparison with the corresponding change in the Rg value indicates a temporary and relatively small elongation of the peptide, which however remains shorter, on average, than the peptide in the other two solutions. The subsequent decrease in Rg despite the persistent change in RMSD indicates that the peptide rearranges in the new conformation, which, once stabilized, is no more extended than the previous one. Overall, the comparison of the peptide Rg and SASA evolutions in the Li+ solution with those in the Na+ and K+ solutions shows that, following the transition, the peptide structure remains more compact in the presence of Li+ counterions (a test simulation with the peptide neutralized by Li+ ions but no additional LiCl salt, starting from the last snapshot of the MD simulation shown, fully confirms the compact peptide structures emerging from Fig. 3 compared to those obtained in the presence of the two other ion species). The use of a 0.5 M ionic strength leads to results similar to those in solutions with no added salt (henceforth denoted NS), except for the absence of the event of conformation change in the Li+ system (the blue curves on the left column of Fig. 3 show neither jump in the RMSD nor transient change in the radius of gyration). In particular, the ionic strength does not significantly affect the peptide elongation, although a slight decrease in size is obtained for the polypeptide in the presence of Li+ ions, as is shown by the comparisons of the Rg and SASA values for 10GLU in the NS and 0.5 M solutions. A previous study167 found a decrease in the end-to-end distance of the polypeptide with increasing concentrations of NaCl and KCl salts, most noticeable in the Na+ system. However, (i) the maximum effect was seen at a much higher salt concentration (∼2 M) than that used here, and the chain size showed some increase by further raising the concentration of the solution to ∼4 M; (ii) the precise value of the chain size depends on the force fields used (as was appreciated in ref. 167 even just by changing the force field for NaCl); (iii) most importantly, the relative changes in size reported in ref. 167 fell within the range of size fluctuations emerging from Fig. 3.
To gain insight into the dependence of peptide dynamics on the counterion species used, we calculated (with the same type of procedure described above) the average counterion–coordination number per carboxyl group along the MD simulations with the different ions. Table 1 shows that, irrespective of the ionic strength, this coordination number is about one order of magnitude greater for Li+ than for Na+ and K+ (incidentally, we note that the coordination numbers obtained for the Na+ and K+ counterions correlate with those in ref. 167, despite quantitative differences that are reasonably expected due to the use of different force fields). Moreover, we find that the first peak of the radial distribution function occurs at a shorter distance for the Li+ species than for the other two ions (see SI section S4), thus indicating a closer proximity of the Li+ ions to the COO− groups.
The following picture emerges from the above analysis. The strong COO−–Li+ interaction, as is manifested by the proximity of the two ionic species, shields the electrostatic repulsion between the carboxylates, which is the major driving force for the greater peptide elongation in the Na+ and K+ solutions, thus favoring more compact conformations of the peptide in the presence of Li+ ions. This conclusion is consistent with the increasing formation of alpha-helix motifs from random coil polyglutamic acids with decreasing pH found in earlier studies.164,168,169 In fact, Li+ is the smallest ion used in our simulations, which can closely approach the carboxylates and therefore play a role quite similar to that of H+ in the mentioned studies.
The simulation with Li+ in a 0.5 M solution does not show the peptide conformation transition seen without salt (compare the red and black traces in the left panels of Fig. 3). This result agrees with the fact that, according to the Rg and SASA descriptors, the transition causes a relatively less folded peptide structure, which is less favorable as the Li+ coordination to COO− increases due to the salt. Therefore, we expect that the timescale for the occurrence of such a transition is expanded in the presence of the salt. Another possible explanation of the different results in the two solutions containing Li+ is that the transition observed in the NS case is an accidental event that, on average, occurs over a much longer timescale and therefore it is expectedly not seen in the 0.5 M solution. In fact: (i) the top-left panel of Fig. 3 describes a persistence of the conformational change, whose duration is, therefore, at least on the timescale of the simulations; (ii) performing an MD simulation of the system with Li+ counterions from different initial conditions (i.e., starting from the final snapshot of the simulation with Na+ counterions), after about 200 ns of re-equilibration we observed again the same compact structure of the peptide as on the left panels of Fig. 3, but without the occurrence of the conformational change, which suggests the occurrence of the conformational change on a longer timescale. However, the significant relative increase of nLi+–COO− from 1 to 1.6 with the addition of the salt still supports an effect of the increased ionic strength on the occurrence of conformational changes of the peptide, as discussed above. A notable relative increase in nLi+–COO− with the ionic strength of the solution is also found for the systems containing Na+ and K+ ions. However, the resulting nLi+–COO− values are small enough not to cause significant changes in the peptide structural dynamics (compare red and black traces in the middle and right panels of Fig. 3 and their statistical analysis in Table 1).
3.3. Driven peptide dynamics in the absence of the nanopore
The translocation of the peptide requires the application of an external (electric) field, which is also generally used in experimental investigations of peptide and protein translocation. In this study, we use uniform electrostatic fields along the z axis, both in the absence (this subsection) and in the presence (next subsection) of the nanopore (whose principal axis of symmetry is along z), which allows us to highlight the effect of the nanopore on the motion of the peptide.The field intensities tested are 0.25, 0.5, 0.75 and 1 V nm−1. Although similar driving field intensities (a few hundred mV per nm) have been employed to investigate the passage of DNA through nanopores,170 the label-free optical sequencing of DNA, peptides, and proteins may require that the molecular translocation be slow enough for sufficient optical signal acquisition. For example, driving fields about two orders of magnitude weaker than those used here were applied to DNA translocating through a plasmonic nanopore in ref. 171. On the other hand, from a theoretical point of view, much more intense driving fields are necessary to observe appreciable translocation dynamics over reasonable simulation time scales. Therefore, the use of different field intensities helps us extrapolate the MD results and validate them at smaller field magnitudes.
Using a field with an intensity of 0.5 V nm−1 and focusing on the case without the nanopore, we do not see any significant change in the evolution of the RMSD, Rg and SASA descriptors (Fig. S5). This is not trivial, given the differences in the local structure, charge, and polarization properties of the components of the solution. In fact, we obtain nNa+–COO−= 0.11, a value with an appreciable relative difference from that obtained without a field (nNa+–COO− = 0.16). This difference may be ascribed to the fact that the cations and the peptide are dragged in opposite directions by the field.
Similar results are obtained with the other driving fields tested, except for a linear variation in the translocation speed of the peptide, which is globally estimated as the velocity of the center of mass (CM) along the direction of the nanopore axis (see Table 2, Table S2 and SI Fig. S6). This result provides an indication that a linear drag regime56 still holds, with good approximation, at such driving field strengths, thus supporting the extension of the present results to systems subjected to weaker external fields by reducing the magnitudes of the CM speeds proportionally to the electric field intensity. This consideration can be extended to the motion of a peptide all inside the pore, that is, without taking into account the dynamics of the biomolecule at the entrance and exit of a finite-length nanopore. Clearly, the observed significant effect of the electric field in slowing down the translocation motion of the polypeptide suggests that this consideration requires further investigation, offering a starting point for future studies on such systems.
| Nanopore | Counterion | I (M) | Δ c (nm) | N w | t 0(ns)a | v(m s−1) |
|---|---|---|---|---|---|---|
| a Time after which the dynamics of 10GLU reaches a plateau according to the structural descriptors, and the statistical analysis is thus performed. b The solution contains urea with a 6 M concentration. c A field strength of 1.0 V nm−1 is used, while a strength of 0.5 V nm−1 is used for all other data points. Further information on the relationship between the field intensity and peptide velocity is provided in SI Table S2 and Fig. S6. | ||||||
| No | Na+ | NS | 3.997 | 2.093 | 50 | 25.2 ± 0.2 |
| NSb | 3.859 | 1107 (Nurea = 308) | 50 | 12.1 ± 0.1 | ||
| 0.5 | 3.997 | 2055 | 50 | 16.6 ± 0.1 | ||
| NSc | 3.981 | 2093 | 50 | 51.5 ± 0.3 | ||
| No | Li+ | NS | 4.005 | 2093 | 200 | 9.3 ± 0.5 |
| 0.5 | 3.991 | 2055 | 100 | 4.8 ± 0.4 | ||
| No | K+ | NS | 4.004 | 2093 | 150 | 30.5 ± 0.2 |
| 0.5 | 4.017 | 2055 | 50 | 23.7 ± 0.1 | ||
| Yes | Na+ | NS | 4016 | 150 | 4.5 ± 0.2 | |
| 0.3 | 3978 | 300 | 3.2 ± 0.2 | |||
| Yes | Li+ | NS | 4016 | 100 | 1.6 ± 0.2 | |
| 0.3 | 3978 | 50 | 0.5 ± 0.1 | |||
Table 2 shows that the peptide velocity significantly depends on the environmental conditions, but decreases with ionic strength regardless of the counterion species. This fact can be rationalized in terms of the Coulomb interaction between the carboxylic groups and the surrounding alkaline ions. Due to the attractive interaction, nM+–COO− (M = K, Li, Na) increases with ionic strength. Thus, it causes more screening of the peptide charge, namely, less effective charge on the peptide subjected to the electrostatic field. Since the M+ and COO− ions move in opposite directions in the presence of the field, the increased number of positive ions surrounding the negatively charged peptide also oppose a more effective drag to the peptide motion. The considerably larger value of nM+–COO− for M![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) Li than for M
Li than for M![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) K, Na accordingly explains the significantly smaller translocation velocity of the peptide in the Li+ solution and its more pronounced decrease with ionic strength. Therefore, the ionic species and strength need to be considered jointly with the intensity of the driving field in order to optimally control the translocation speed for peptide or protein sequencing.
K, Na accordingly explains the significantly smaller translocation velocity of the peptide in the Li+ solution and its more pronounced decrease with ionic strength. Therefore, the ionic species and strength need to be considered jointly with the intensity of the driving field in order to optimally control the translocation speed for peptide or protein sequencing.
The velocity of the peptide CM was also calculated in a solution with a concentration 6 M of urea, since urea is an agent commonly used for protein denaturation,172–174 and the latter is required to unfold the protein and thus allow for sequencing. Table 2 shows the peptide slowdown due to hydrodynamic friction with urea at the concentration used in the simulation (which is on the lower side of the range in which complete denaturation of proteins is expected to occur; e.g., see ref. 173).
Table 2 also reports the number Nw of water molecules present in the MD simulation box in the different computations (clearly, this number is greatly decreased in the presence of Nurea = 308 molecules of urea) and the MD cell size Δc in the final snapshot of each MD simulation. This Δc value is reported as an example of our observation that the relative changes in cell size were very small during all NPT simulations. For the simulation with Na+ counterions and no added salt (first entry in the table) mostly used for the comparison with the NVT simulations in the presence of the nanopore, the relative difference between Δc and the nominal cell size (in each direction) of 4 nm is, e.g., as small as 7.5 × 10−4. On doubling the field, the relative difference is still 4.8 × 10−3. This observation encouraged us both to compare the NPT simulations in the absence of the nanopore with the NVT simulations in the presence of the nanopore, and to use an isotropic barostat despite the presence of a directional electric field. The latter choice can be rationalized by considering the isotropic nature of the aqueous solvent (and the translational symmetry of the Au nanopore in the direction of the electrostatic field applied), which is almost entirely responsible for the pressure in our simulations. In fact, our computational choice is in line with NPT simulations in the presence of a directional electric field in the literature,175–178 where the use of an anisotropic barostat is usually limited to systems with strong intrinsic anisotropies,179,180 and also in the case of ‘side effects’ of using anisotropic pressure coupling.181
3.4. Field-driven peptide dynamics inside the nanopore
We finally investigate the technologically most relevant scenario, in which the 10GLU peptide moves in the Au nanopore under an external electrostatic field with the amplitude Ez = 0.5 V nm−1. We emphasize that, since the peptide system is relatively small and the theory refers to sequencing not based on measurements of ion current, we are not interested in the blockage or reduction of ionic currents through the pore due to the peptide. Rather, we focus on the motion of the peptide interacting with the nanopore surface.Fig. 4 compares the time evolutions of the structural descriptors during the dynamics of the peptide inside the Au nanopore in the presence and absence of the driving electrostatic field. The peptide leans on the nanopore surface in both cases. However, the peptide can more easily take horizontal poses in the absence of the field, while it is generally elongated along the z axis in the presence of the field (see Fig. S7), thus clearly favoring its sequencing. The interaction with the pore surface makes the peptide structure appreciably less flexible and more persistently elongated compared to the structure free in the solution. This is shown by the fact that the RMSD is, on average, appreciably smaller than that in Fig. 2 (compare the black RMSD traces in Fig. 2 and Fig. 4, which correspond to mean RMSD values of 0.65 Å and 0.54 Å, respectively), while Rg is consistently larger (namely, its mean value passes from 0.91 nm in Fig. 2 to 1.07 nm in Fig. 4). Furthermore, we find that the pore-peptide distance lies in the range of 0.3–0.6 nm. With an eye to optical sequencing applications, this range of relatively short distances fuels the expectation of effective interaction with plasmonic fields generated within the nanostructure in the experiment. Consistent with such results, the SASA is significantly reduced by the interaction of the peptide with the pore. At the beginning of the MD simulation, the peptide is located at the center of the Au nanopore. Therefore, the SASA values are similar to those for the free peptide in Fig. 2a. Then, the peptide approaches the surface of the pore and remains attached to it for the remainder of the simulation. Therefore, the exposure to solvent is considerably reduced and the SASA accordingly decreases.
As Fig. 4 shows, the external field enhances the adhesion of the peptide to the Au surface, which is completed between 100 and 150 ns if the driving field with intensity Ez = 0.5 V nm−1 is applied. In contrast, it takes about 400 ns without a driving field (compare black and cyan lines in Fig. 4). We thus conclude that the joint action of the Au surface and the dragging external field promotes the peptide steering along the nanotube, as is desirable for its optical sequencing (see below).
Table 2 shows that the translocation speed of the peptide decreases with ionic strength also inside the nanopore and is lower in the solution with Li+ counterions than in the one with Na+ ions. The increase in ionic strength can be generally expected to weaken the interaction of the peptide with the Au pore surface (as has been observed for very different molecule-pore systems182,183). This weakening should favor translocation, but the effect of shielding the peptide charge on the ability to drive it by an electric field is predominant.
Importantly, in all cases, the interaction of the peptide with the Au pore acts as a frictional force slowing down its translocation motion. This result agrees with the literature on very different systems, such as neutral and charged polyethylene glycols moving inside organic nanopores coated with oxides to favor their adhesion to the polymer.81 Our analysis not only extends this conclusion to the case of peptides in Au nanopores, but also shows that the interaction with the nanopore is similarly effective in slowing down the dynamics of the peptide in solutions with different ions at different ionic strengths. The elongation of the peptide and its smaller translocation velocity in the presence of the nanopore concur in facilitating its sequencing by optical means.
The translocation speeds obtained are too high for adequate optical sequencing even after scaling them down, e.g., by two orders of magnitude to represent the situation with an accordingly weaker translocation field (see above). However, on the one hand, our analysis shows that the interaction with the metal surface slows down the peptide translocation. On the other hand, in experimental contexts, one expects the realization of hot spots for the detection of the molecule3 with an accumulation of excess charge on the metal surface and hence a larger interaction with the molecule.
To perform a more realistic simulation of the peptide motion and to describe the effect of the excess charge on the Au surface and the consequent diffuse layer on such motion, we investigated the translocation of the peptide for different potentials of the nanopore surface, that is, for different excess charges on the gold surface, which were balanced by Cl− counterions added to the solution. To this end, the charges on the Au atoms were varied so as to reach the desired electrostatic potentials at the nanopore surface with respect to the bulk solution (that is, with respect to the solution outside the diffuse layer), according to the Poisson–Boltzmann approach of Gouy and Chapman162 (see details in Methods). In our work, in the absence of data on the effective potential of the gold surface immersed in the electrolytic solution (also due to the experimental intricacies arising from the coexistence of surface and electrophoretic potentials), we explored the range of interface potentials (or lateral potentials, in contrast to the driving potential difference along the nanotube) from 0 to 260 mV (Table 3). Negative values of the lateral potential were not considered, since they would only determine a repulsion between the Au nanopore surface and the negatively charged polypeptide, thus not allowing the translocation velocity to be controlled via the surface–molecule interaction and reduced appropriately for amino acid detection purposes.
| V t (mV) |
σ (μC cm−2)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) a a |
l
GC (Å)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) b b |
t
0 (ns)![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) c c |
v (m s−1) |
|---|---|---|---|---|
| a Net charge density on the internal surface of the nanopore. b Gouy–Chapman length. c Time after which the RMSD reaches a plateau and the peptide speed is calculated. d At these Vt values, the velocity of the CM along z fluctuates around zero, with a modulus of the order of 10−4 m s−1. | ||||
| 160 | 12.7 | 2.8 | 50 | 17.0 |
| 180 | 19.5 | 1.8 | 150 | 10.4 |
| 200 | 29.0 | 1.2 | 50 | 2.2 |
| 210 | 35.1 | 1.0 | 50 | 8.0 × 10−1 |
| 215 | 38.7 | 0.9 | 50 | 1.3 × 10−1 |
| 220 | 43.3 | 0.8 | 150 | 5.9 × 10−3 |
| 230 | 51.9 | 0.7 | 310 | }<10−4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) d d |
| 240 | 62.5 | 0.6 | 150 | |
| 250 | 77.0 | 0.5 | 100 | |
| 260 | 91.4 | 0.4 | 100 | |
The dependence of the peptide translocation speed on the interface potential drop Vt is shown in Table 3 and Fig. 5a. For zero Vt, the translocation speed is smaller than that for moderate values of Vt (up to Vt = 180 mV in Table 3). This can be interpreted as resulting from the accumulation of Cl− ions at the surface (as we observed by inspection of the MD trajectories, and in general agreement with the reducing binding to different surfaces of charged peptides with increasing salt concentration182,183), thus preventing its direct contact with the polypeptide and promoting the sliding of the latter through the gold nanopore. In fact, the comparison between Tables 2 and 3 shows that the translocation speed value for Vt = 160 mV is within the range between the speed without a pore and that in the pore without a lateral potential. As the Vt value increases further, the peptide experiences a prevailing and increasing effective friction due to a direct interaction with the Au surface (as we have also seen by examining MD simulations like the one used to produce the video in the SI) and starts slowing down significantly. For large enough Vt, the speed of the CM fluctuates around zero over the time interval explored by the MD simulations (Table 3). In a range of intermediate values of Vt, we find a discontinuous motion of the peptide characterized by periods of translocation interspersed with periods in which the peptide is stuck on the nanopore surface. For Vt = 220 mV, we inspected this kind of dynamics by means of an MD simulation of 2 μs (Fig. 5b). We expect that the same type of motion can be observed for larger values of Vt in sufficiently long MD simulations.
Table 3 also reports Gouy–Chapman's length, lGC,184 at which the average Coulomb attraction between a unit charge and the charged surface equals the thermal energy. As expected for the rather high charge density on the Au surface, lGC is on the order of 1 Å and decreases with the surface charge. This also means that the electroosmotic flow related to the presence of counterions decreases with the magnitude of Vt (incidentally, we notice that this effect, as well as the viscous drag effect on the water farther from the nanopore surface, is intrinsically included at the atomistic level of our simulations). Furthermore, the ion concentration is maintained at the same value in all MD simulations at different Vt values reported in Table 3 and Fig. 5, and lGC decreases with the surface charge. Therefore, the electroosmotic effect, which we assessed to be negligible for Vt = 220 mV (at this Vt value we find that the average velocity of the water molecules is about 2.2 × 10−6 m s−1, which is three orders of magnitude less than the peptide mean velocity and may be ascribed to the ion distribution and the width of the nanopore), is not responsible for the observed behavior of the translocation speed with increasing transversal potential difference.
The intermittent translocation of the peptide, and other essential features of its motion in response to Vt, can be captured through the following formally simple physical model.
By inspection of the polypeptide trajectory at Vt = 220 mV, we see that one to a few COO− groups strongly interact with the gold surface polarization charge and transiently pin the polypeptide to the surface (which cannot be considered a uniformly charged continuous surface at the MD atomistic level). Therefore, the polypeptide jumps between the local minima of the electrostatic interaction energy, in the direction determined by the driving electrostatic field.
For a given value of Vt, the peptide has at each time a probability P to be detached from the surface, and therefore a probability 1 − P to be stuck on it. Assuming that the difference in peptide–nanopore interaction energy between situations in which the peptide is pinned or not to the surface is proportional to Vt through a coefficient a (due to the very small Gouy–Chapman length, the two situations practically correspond to the displacement of the partly screened peptide charge through a region in which there is a potential drop of Vt), we consider that P decreases with Vt proportionally to the Boltzmann factor 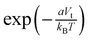 . The average translocation speed while the peptide is detached from the nanopore surface, vd, also depends on Vt. On average, over a time interval Δt, the CM of the peptide travels a distance Δz = vdPΔt, and therefore:
. The average translocation speed while the peptide is detached from the nanopore surface, vd, also depends on Vt. On average, over a time interval Δt, the CM of the peptide travels a distance Δz = vdPΔt, and therefore:
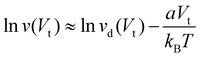 | (2) |
Accurate applications of eqn (2) require knowledge of vd(Vt), and vd can be a complicated function of Vt for a flexible solvated peptide near a gold surface. However, we expect vd to vary slowly with Vt as long as the transversal potential difference is not large enough to make the peptide lean closely on the pore surface, thus leading to the very slow initial variation of the peptide speed with Vt in Fig. 5a. As Vt is sufficiently large, the exponential dependence of P on Vt is expected to dominate the behavior of v(Vt), so as to determine a linear dependence of ln v on Vt. The data points in Fig. 5a are compatible with—but not sufficient to determine—such a linear dependence, whose investigation is computationally very expensive (due to the very slow translocation dynamics in this regime) but worthy of future analysis. At any rate, the drop in translocation speed with increasing Vt shown in Fig. 5a prevents observing an appreciable motion of the peptide over the MD simulation time for Vt ≥ 230 mV (Table 3).
The stop-and-go translocation mechanism underlying eqn (2) and shown in Fig. 5b (the stop-and-go mechanism is clearly visible in a video of the polypeptide translocation through the gold nanopore with Vt = 220 mV, which is provided in the supplementary Material; for this Vt value, SI section S10 also reports a comparison of the peptide average velocities for replicas of the first 100 ns of the MD run) is reminiscent of the slowdown of cisplatin diffusion inside a nanopore due to molecular adsorption onto the pore surface,92,93 although the two types of systems and their motion are very different. The flat regions of the plot correspond to the peptide being stuck on the surface, while the peaks represent the time intervals in which the peptide detaches and proceeds in the main direction of the pore under the action of the electric field. Clearly, the variability of the local environment and of the extent of peptide detachment from the surface determines peaks of variable duration and height (i.e., of different maximum speed reached during the forward motion). The slow modes of the peptide structural dynamics,85,111 which involve large portions of the polymer, are expected to be primarily involved in dragging the peptide away from the anchoring poses. While 10GLU is anchored to the pore, such modes determine a velocity of the CM on the order of magnitude of 10−4 m s−1 (see Fig. S9, for example). Therefore, we expect that for Vt > 220 mV, the translocation speed of the detached peptide, vd, is between these values and those in Fig. 5b.
Given the smooth change in vd with Vt, and the approximately linear dependence of ln v on Vt for sufficiently strong lateral fields, we obtain a/e ≈ 23.87 (e is the elementary charge) from the last two data points in Fig. 5a. With this value of paramter a, a lateral potential as close to 220 mV as 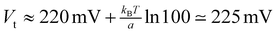 is already sufficient to reduce the translocation velocity to the value v ≈ 6 × 10−5 m s−1, which is two orders of magnitude smaller than the velocity obtained for Vt = 220 mV. This explains why we cannot see jumps in the translocation motion during the MD simulations with Vt > 220 mV. It is worth noting that a similar translocation speed on the order of 10−5 m s−1 can also be obtained for Vt = 220 mV by reducing the driving electrostatic field, Ez, by two orders of magnitude, as vd is expected to scale in an approximately linear way with Ez.
is already sufficient to reduce the translocation velocity to the value v ≈ 6 × 10−5 m s−1, which is two orders of magnitude smaller than the velocity obtained for Vt = 220 mV. This explains why we cannot see jumps in the translocation motion during the MD simulations with Vt > 220 mV. It is worth noting that a similar translocation speed on the order of 10−5 m s−1 can also be obtained for Vt = 220 mV by reducing the driving electrostatic field, Ez, by two orders of magnitude, as vd is expected to scale in an approximately linear way with Ez.
3.5. Measurement of polyglutamic acid translocation through a gold nanopore
To compare these theoretical results with those in practical experimental setups, we performed electrophoretic translocation experiments in solid-state gold nanopores. The nanopores were fabricated by ion beam milling and have a minimum inner diameter of 10 nm.143 Further details are provided in the Materials and Methods section. Translocation dynamics were monitored by electrical readout, following standard electrical measurement. The protocol for the Au nanopores was adapted from ref. 84. As anticipated above, the probe molecule used is a 5000-unit-long polyglutamic acid. The concentration was set to 10 nM. Potassium chloride (KCl) was used as the electrolyte, with a concentration of 100 mM. The electrophoretic bias was 100 mV. Typical current spikes due to the translocation of molecules into pores are shown in Fig. 6a. By analyzing the duration of the spike, it is possible to trace the duration of the translocation, usually called dwell time. The dwell time distribution was extracted from the ion current files using Clampfit software. The half-amplitude threshold method was applied to discriminate events from noise where the signal exceeds a set amplitude threshold. More specifically, events smaller than 0.5 ms were discarded since they could not be differentiated from noise. The log–normal distribution of the dwell time is shown in Fig. 6b. The analysis shows an average duration of 1.2 ms per translocation event, which, considering the length of the polymer (1.5 μm), means a translocation velocity of 1.25 × 10−3 m s−1.Since the bias potential drop occurred across a nanopore length of about 100 nm, the driving field was about 1 mV nm−1. Assuming a linear dependence of the polypeptide translocation velocity on the electrophoretic bias, based on the results shown above, the speed expected in the nanopore under our driving field of 0.5 V nm−1 is about 0.6 m s−1 (or less, since the dependence of the velocity on the driving field may be less than linear at the experimental bias). This translocation velocity is lower than that predicted for Vt = 0, but within an order of magnitude of the latter. This is a reasonable agreement, considering the ideal character of the model (defect-free) nanopore surface compared to what can be expected in the experiments. Also, our simulation was performed using Na+ counterions, while the experiment was carried out in the presence of the KCl salt. Comparing with the velocities obtained under bias, the experimental translocation velocity is in between those obtained with lateral potentials of 210 (last data point in black color in Fig. 5a) and 215 mV (first data point in red in Fig. 5b), in a critical range of lateral potential values where a significant stop-and-go translocation mechanism is triggered. A closer comparison between the theoretical and experimental results is hampered by the differences between the model and real systems, including the much longer polyglutamic acid used in the experiment, the different nanopore sizes and ionic environments (despite the similarities between the theoretical results using the Na+ and K+ ions, the ionic strengths also differ in the simulation and experiment), and other aspects. Although the comparison between theory and experiment must be treated with due caution, it still suggests the stop-and-go motion as a plausible translocation mechanism in the real system, where it might also be triggered by lateral potential differences smaller than 220 mV due to the additional friction of the longer polymer as it enters and exists the nanopore.
The above analysis clearly indicates opportunities for using electrochemical control of the pore potential to slow down the translocation of peptides and stretched proteins through nanopores in SERS-based sequencing. Another critical point for peptide or protein sequencing is the ability to distinguish different amino acids depending on the degree of molecular stretching. This point is examined in the next section.
3.6. Amino acid distinguishability
An ideal sequencing procedure would require that amino acids sequentially pass through a suitably focused hot spot in the pore, with a translocation velocity that allows for the collection of sufficient spectroscopic information. Assuming that the molecular detection can be focused on a spot of the size of one or very few amino acids, their distinction still requires that the stretching of the peptide in the presence of the Au nanopore ensures a sufficiently small spatial overlap between the amino acids in the direction of the Au nanopore axis.To investigate the above point, including the effect of the peptide-nanopore interaction on the distinguishability of amino acids, we determined the amino acid overlaps shown in Fig. 7, after deriving the relative average density profiles of the amino acid residues in the z direction from the MD simulations of the 10GLU peptide free in solution and confined in the nanopore, as described in Methods. We calculated the overlaps considering only the heavy atoms of the side chains, as we obtained similar results in tests performed using the full amino acids.
Fig. 7 shows that, in the presence of the nanopore, the next-neighbor side chains overlap by about a third, the second next-neighbors overlap by significantly less than 10%, and the farther side chains show negligible overlap. In the absence of the nanopore, each amino acid side chain appreciably overlaps, on average, with several others. In terms of overlapping atoms, in the absence of the nanopore, each GLU side chain overlaps (along the z direction), on average, with almost two atoms in the next-neighbor side chains, almost one atom in the second next-neighbors, and an appreciable fraction of atom in several farther GLU side chains. Instead, in the presence of the nanopore, ∼1.6 atoms of a side chain overlap with atoms in the next-neighbor side chains, while there is a small to negligible overlap with the farther side chains. Therefore, the pore–peptide interaction significantly increases the distinguishability of amino acids, as is desirable for protein sequencing. It is worth noting that our conclusion is even more true since the degree of distinguishability we showed refers to the most difficult scenario in which the amino acids are all of the same type.
4. Conclusions
This study presents a mechanistic view (at an atomistic level) of peptide translocation through gold nanopores, which is comprehensive and simple at the same time, with the aim to provide guidance for the experimental implementation of metal nanopores for label-free protein sequencing through optical techniques. To this end, we take on a fresh perspective, in which we focus directly on the motion of the peptide, rather than evaluating the effects of the peptide on ionic currents. Through a vast analysis of MD simulations, using several structural descriptors, we dissect the behavior of the peptide in terms of its dependence on different environmental conditions, including the composition and ionic strength of the solution and the polarization charge on the internal surface of the solid-state nanopore, which can be externally varied by electrochemical control. Over a range of reasonable ionic strengths, we find that the type of ion in the solution is more important than the ionic strength itself to determine the degree of peptide elongation and hence the opportunities for sequential translocation of the amino acids and their suitable detection. For example, Li+ more closely interacts with the carboxylates of the peptide compared to Na+ and K+, thus more effectively screening their electric interaction and allowing more compact conformations of the peptide. This fact brings about the advantage of a slower peptide translocation under a driving electrostatic field, but also the significant disadvantage of an appreciably less sequential transit of the amino acid residues, thus disfavoring their individual detection. In addition, the more moderate peptide charge screening by Na+ and K+ ions enables a more effective control of the dynamics by lateral fields.The use of larger ions such as Na+ and K+, together with the application of lateral electric fields of appropriate intensities, emerges from our analysis (which includes the quantification of nanopore effects on the sequential passage of the amino acids) as an effective strategy to control the rapidity of translocation of sufficiently elongated peptides, and thus enable their sequencing. Our study also shows (Fig. 5) a crucial range of intensities of the surface potential over which the peptide translocation gradually acquires a stop-and-go mechanism, which can be used to fine-tune the translocation velocity of the peptide. Depending on the value of the surface potential and the driving field, the stop-and-go translocation can also occur on timescales that cannot be grasped by the MD simulations, but may be relevant to suitable sequencing of the amino acid chain.
It is worth noting that the presence of a negative charge on the GLU residues does not limit the significance of this mechanism for applications to polypeptide and protein sensing and sequencing. In fact, a growing number of studies demonstrate the use of sodium dodecyl sulfate (SDS) to unfold proteins, while simultaneously charging them, for translocation through solid-state nanopores. Combined theoretical-experimental investigations have indeed shown the threading of individual SDS-denatured and negatively charged polypeptides through sub-5 nm pores.94,185 In the present study, our experiments on a 5000-unit-long polyglutamic acid moving through a solid-state gold nanopore yielded a translocation velocity in the range where theory predicts the stop-and-go translocation mechanism to be at stake and critically controlled by the intensity of the interface potential field.
Author contributions
Mirko Vanzan: conceptualization, investigation (lead), formal analysis, software, data curation, validation, visualization, writing – review & editing; Agostino Migliore: supervision, conceptualization, investigation (supporting), methodology, formal analysis, software, data curation, validation, visualization, writing – original draft, writing – review & editing; Maria Blanco-Formoso: conceptualization, investigation (lead), methodology, formal analysis, data curation, validation, visualization, writing – review & editing; Francesco De Angelis: supervision, conceptualization, methodology, writing – review & editing, resources, funding acquisition, project administration; and Stefano Corni: supervision, conceptualization, methodology, writing – review & editing, resources, funding acquisition, project administration.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: Tables S1–S2, Figs. S1–S10, a brief discussion of the length dependence of the peptide dynamics in solution, further analysis of the peptide-ion interactions, the statistical analysis of the relationship between peptide velocity and diving field strength, and an MD replica analysis. The coordinates of the pore are available in the file Au_nanopore_coordinates.xyz. A video of the polypeptide stop-and-go translocation through the pore with a lateral potential Vt = 220 mV (from 1359 to 1364 ns of the MD trajectory) is contained in the file translocation_for_Vt_220mV.mp4. See DOI: https://doi.org/10.1039/d5nr03385c.Acknowledgements
All authors acknowledge funding of this research by the European Union's Horizon 2020 research and innovation programme – Grant Agreement no. 964363 “ProID” - H2020-FETOPEN-2018-2020. M. V. acknowledges the University of Milan for funding his postdoctoral fellowship “La bellezza degli aggregati: da nano a astro particelle”.References
- W. Q. Shi, A. K. Friedman and L. A. Baker, Anal. Chem., 2017, 89, 157–188 CrossRef CAS PubMed.
- A. Asandei, G. Di Muccio, I. Schiopu, L. Mereuta, I. S. Dragomir, M. Chinappi and T. Luchian, Small Methods, 2020, 4, 1900595 Search PubMed.
- Y. Q. Zhao, M. Iarossi, A. F. De Fazio, J. A. Huang and F. De Angelis, ACS Photonics, 2022, 9, 730–742 Search PubMed.
- R. McDaniell, D. M. Warthen, P. A. Sanchez-Lara, A. Pai, I. D. Krantz, D. A. Piccoli and N. B. Spinner, Am. J. Hum. Genet., 2006, 79, 169–173 CrossRef CAS PubMed.
- J. S. Valastyan and S. Lindquist, Dis. Models Mech., 2014, 7, 9–14 CrossRef CAS PubMed.
- E. M. Shaikho, A. H. Habara, A. Alsultan, A. M. Al-Rubaish, F. Al-Muhanna, Z. Naserullah, A. Alsuliman, H. O. Qutub, P. K. Patra, P. Sebastiani, K. Baltrusaitis, J. J. Farrell, Z. Jiang, H. Y. Luo, D. H. Chui, A. K. Al-Ali and M. H. Steinberg, Blood Cells, Mol., Dis., 2016, 59, 49–51 CrossRef CAS PubMed.
- I. Rehman, M. Farooq and S. Botelho, Biochemistry, Secondary Protein Structure. in: StatPearls, StatPearls Publishing, Treasure Island (FL), 2025 Search PubMed.
- T. J. Moore, A. S. Moody, T. D. Payne, G. M. Sarabia, A. R. Daniel and B. Sharma, Biosensors, 2018, 8, 46 CrossRef PubMed.
- S. Y. Wang, Z. Y. Zhao, F. Haque and P. X. Guo, Curr. Opin. Biotechnol., 2018, 51, 80–89 CrossRef CAS PubMed.
- D. K. Kim, B. Weller, C. W. Lin, D. Sheykhkarimli, J. J. Knapp, G. Dugied, A. Zanzoni, C. Pons, M. J. Tofaute, S. B. Maseko, K. Spirohn, F. Laval, L. Lambourne, N. Kishore, A. Rayhan, M. Sauer, V. Young, H. Halder, N. Marín-de la Rosa, O. Pogoutse, A. Strobel, P. Schwehn, R.-J. Li, S. T. Rothballer, M. Altmann, P. Cassonnet, A. G. Cote, L. E. Vergara, I. Hazelwood, B. B. Liu, M. Nguyen, R. Pandiarajan, B. Dohai, P. A. R. Coloma, J. Poirson, P. Giuliana, L. Willems, M. Taipale, Y. Jacob, T. Hao, D. E. Hill, C. Brun, J. C. Twizere, D. Krappmann, M. Heinig, C. Falter, P. Aloy, C. Demeret, M. Vidal, M. A. Calderwood, F. P. Roth and P. Falter-Braun, Nat. Biotechnol., 2023, 41, 140–149 CrossRef CAS PubMed.
- A. J. Barbier, A. Y. J. Jiang, P. Zhang, R. Wooster and D. G. Anderson, Nat. Biotechnol., 2022, 40, 840–854 CrossRef CAS PubMed.
- K. K. Chen, I. Jou, N. Ermann, M. Muthukumar, U. F. Keyser and N. A. W. Bell, Nat. Phys., 2021, 17, 1043–1049 Search PubMed.
- J. J. Kasianowicz, E. Brandin, D. Branton and D. W. Deamer, Proc. Natl. Acad. Sci. U. S. A., 1996, 93, 13770–13773 CrossRef CAS PubMed.
- M. Akeson, D. Branton, J. J. Kasianowicz, E. Brandin and D. W. Deamer, Biophys. J., 1999, 77, 3227–3233 CrossRef CAS PubMed.
- S. Howorka, S. Cheley and H. Bayley, Nat. Biotechnol., 2001, 19, 636–639 CrossRef CAS PubMed.
- D. W. Deamer and D. Branton, Acc. Chem. Res., 2002, 35, 817–825 CrossRef CAS PubMed.
- A. J. Storm, J. H. Chen, H. W. Zandbergen and C. Dekker, Phys. Rev. E: Stat., Nonlinear, Soft Matter Phys., 2005, 71, 051903 Search PubMed.
- D. Branton, D. W. Deamer, A. Marziali, H. Bayley, S. A. Benner, T. Butler, M. Di Ventra, S. Garaj, A. Hibbs, X. H. Huang, S. B. Jovanovich, P. S. Krstic, S. Lindsay, X. S. S. Ling, C. H. Mastrangelo, A. Meller, J. S. Oliver, Y. V. Pershin, J. M. Ramsey, R. Riehn, G. V. Soni, V. Tabard-Cossa, M. Wanunu, M. Wiggin and J. A. Schloss, Nat. Biotechnol., 2008, 26, 1146–1153 CrossRef CAS PubMed.
- T. Z. Butler, M. Pavlenok, I. M. Derrington, M. Niederweis and J. H. Gundlach, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 20647–20652 CrossRef CAS PubMed.
- D. Stoddart, A. J. Heron, E. Mikhailova, G. Maglia and H. Bayley, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 7702–7707 CrossRef CAS PubMed.
- N. An, A. M. Fleming, H. S. White and C. J. Burrows, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 11504–11509 CrossRef CAS PubMed.
- M. Ayub, D. Stoddart and H. Bayley, ACS Nano, 2015, 9, 7895–7903 CrossRef CAS PubMed.
- C. W. Fuller, S. Kumar, M. Porel, M. Chien, A. Bibillo, P. B. Stranges, M. Dorwart, C. Tao, Z. Li, W. Guo, S. Shi, D. Korenblum, A. Trans, A. Aguirre, E. Liu, E. T. Harada, J. Pollard, A. Bhat, C. Cech, A. Yang, C. Arnold, M. Palla, J. Hovis, R. Chen, I. Morozova, S. Kalachikov, J. J. Russo, J. J. Kasianowicz, R. Davis, S. Roever, G. M. Church and J. Ju, Proc. Natl. Acad. Sci. U. S. A., 2016, 113, 5233–5238 CrossRef CAS PubMed.
- O. K. Zahid, F. Wang, J. A. Ruzicka, E. W. Taylor and A. R. Hall, Nano Lett., 2016, 16, 2033–2039 CrossRef CAS PubMed.
- C. Cao, Y. L. Ying, Z. L. Hu, D. F. Liao, H. Tian and Y. T. Long, Nat. Nanotechnol., 2016, 11, 713–718 CrossRef CAS PubMed.
- J. A. Huang, M. Z. Mousavi, Y. Q. Zhao, A. Hubarevich, F. Omeis, G. Giovannini, M. Schutte, D. Garoli and F. De Angelis, Nat. Commun., 2019, 10, 5321 CrossRef PubMed.
- M. Jain, R. Abu-Shumays, H. E. Olsen and M. Akeson, Nat. Methods, 2022, 19, 1160–1164 CrossRef CAS PubMed.
- P. Chen, Z. P. Sun, J. W. Wang, X. L. Liu, Y. Bai, J. Chen, A. N. Liu, F. Qiao, Y. Chen, C. Y. Yuan, J. J. Sha, J. H. Zhang, L. Q. Xu and J. Li, Front. Microbiol., 2023, 14, 1043967 CrossRef PubMed.
- M. Di Ventra and M. Taniguchi, Nat. Nanotechnol., 2016, 11, 117–126 CrossRef CAS PubMed.
- X. Shi, D. V. Verschueren and C. Dekker, Nano Lett., 2018, 18, 8003–8010 CrossRef CAS PubMed.
- L. Movileanu, J. P. Schmittschmitt, J. M. Scholtz and H. Bayley, Biophys. J., 2005, 89, 1030–1045 CrossRef CAS PubMed.
- D. S. Talaga and J. L. Li, J. Am. Chem. Soc., 2009, 131, 9287–9297 CrossRef CAS PubMed.
- R. S. Wei, V. Gatterdam, R. Wieneke, R. Tampé and U. Rant, Nat. Nanotechnol., 2012, 7, 257–263 CrossRef CAS PubMed.
- L. Mereuta, I. Schiopu, A. Asandei, Y. Park, K.-S. Hahm and T. Luchian, Langmuir, 2012, 28, 17079–17091 CrossRef CAS PubMed.
- S. Y. Wang, F. Haque, P. G. Rychahou, B. M. Evers and P. X. Guo, ACS Nano, 2013, 7, 9814–9822 Search PubMed.
- C. Plesa, S. W. Kowalczyk, R. Zinsmeester, A. Y. Grosberg, Y. Rabin and C. Dekker, Nano Lett., 2013, 13, 658–663 CrossRef CAS PubMed.
- C. B. Rosen, D. Rodriguez-Larrea and H. Bayley, Nat. Biotechnol., 2014, 32, 179–181 CrossRef CAS PubMed.
- J. Nivala, L. Mulroney, G. Li, J. Schreiber and M. Akeson, ACS Nano, 2014, 8, 12365–12375 CrossRef CAS PubMed.
- M. Pastoriza-Gallego, M.-F. Breton, F. Discala, L. Auvray, J.-M. Betton and J. Pelta, ACS Nano, 2014, 8, 11350–11360 CrossRef CAS PubMed.
- A. Asandei, M. Chinappi, H.-K. Kang, C. H. Seo, L. Mereuta, Y. Park and T. Luchian, ACS Appl. Mater. Interfaces, 2015, 7, 16706–16714 CrossRef CAS PubMed.
- Y. A. Zhao, B. Ashcroft, P. M. Zhang, H. Liu, S. M. Sen, W. Song, J. Im, B. Gyarfas, S. Manna, S. Biswas, C. Borges and S. Lindsay, Nat. Nanotechnol., 2014, 9, 466–473 CrossRef CAS PubMed.
- L. J. Steinbock, S. Krishnan, R. D. Bulushev, S. Borgeaud, M. Blokesch, L. Feletti and A. Radenovic, Nanoscale, 2014, 6, 14380–14387 RSC.
- Z. L. Hu, M. Z. Huo, Y. L. Ying and Y. T. Long, Angew. Chem., Int. Ed., 2021, 60, 14738–14749 CrossRef CAS PubMed.
- A. Biesemans, M. Soskine and G. Maglia, Nano Lett., 2015, 15, 6076–6081 CrossRef CAS PubMed.
- M. Fahie, C. Chisholm and M. Chen, ACS Nano, 2015, 9, 1089–1098 Search PubMed.
- Y. Wang, L. Q. Gu and K. Tian, Nanoscale, 2018, 10, 13857–13866 RSC.
- E. Kennedy, Z. Dong, C. Tennant and G. Timp, Nat. Nanotechnol., 2016, 11, 968–976 CrossRef CAS PubMed.
- V. Van Meervelt, M. Soskine, S. Singh, G. K. Schuurman-Wolters, H. J. Wijma, B. Poolman and G. Maglia, J. Am. Chem. Soc., 2017, 139, 18640–18646 CrossRef CAS PubMed.
- R. Hu, J. V. Rodrigues, P. Waduge, H. Yamazaki, B. Cressiot, Y. Chishti, L. Makowski, D. P. Yu, E. Shakhnovich, Q. Zhao and M. Wanunu, ACS Nano, 2018, 12, 4494–4502 CrossRef CAS PubMed.
- A. E. Rossini, F. Gala, M. Chinappi and G. Zollo, Nanoscale, 2018, 10, 5928–5937 RSC.
- A. Asandei, I. S. Dragomir, G. Di Muccio, M. Chinappi, Y. Park and T. Luchian, Polymers, 2018, 10, 885 Search PubMed.
- F. Piguet, H. Ouldali, M. Pastoriza-Gallego, P. Manivet, J. Pelta and A. Oukhaled, Nat. Commun., 2018, 9, 966 CrossRef PubMed.
- N. Varongchayakul, J. X. Song, A. Meller and M. W. Grinstaff, Chem. Soc. Rev., 2018, 47, 8512–8524 RSC.
- C. V. de Lannoy, M. Filius, R. van Wee, C. Joo and D. de Ridder, iScience, 2021, 24, 103239 CrossRef CAS PubMed.
- J. W. F. Robertson and J. E. Reiner, Proteomics, 2018, 18, 1800026 CrossRef PubMed.
- M. Chinappi and F. Cecconi, J. Phys.: Condens. Matter, 2018, 30, 204002 CrossRef PubMed.
- K. Kuehnel, Nat. Methods, 2019, 16, 145 Search PubMed.
- A. K. Thakur and L. Movileanu, Nat. Biotechnol., 2019, 37, 96–101 CrossRef CAS PubMed.
- J. S. Yu, S. C. Hong, S. Wu, H. M. Kim, C. Lee, J. S. Lee, J. E. Lee and K. B. Kim, Nanoscale, 2019, 11, 2510–2520 Search PubMed.
- J. A. Alfaro, P. Bohlander, M. J. Dai, M. Filius, C. J. Howard, X. F. van Kooten, S. Ohayon, A. Pomorski, S. Schmid, A. Aksimentiev, E. V. Anslyn, G. Bedran, C. Cao, M. Chinappi, E. Coyaud, C. Dekker, G. Dittmar, N. Drachman, R. Eelkema, D. Goodlett, S. Hentz, U. Kalathiya, N. L. Kelleher, R. T. Kelly, Z. Kelman, S. H. Kim, B. Kuster, D. Rodriguez-Larrea, S. Lindsay, G. Maglia, E. M. Marcotte, J. P. Marino, C. Masselon, M. Mayer, P. Samaras, K. Sarthak, L. Sepiashvili, D. Stein, M. Wanunu, M. Wilhelm, P. Yin, A. Meller and C. Joo, Nat. Methods, 2021, 18, 604–617 Search PubMed.
- M. A. Bakshloo, J. J. Kasianowicz, M. Pastoriza-Gallego, J. Mathe, R. Daniel, F. Piguet and A. Oukhaled, J. Am. Chem. Soc., 2022, 144, 2716–2725 Search PubMed.
- Y. L. Ying, Z. L. Hu, S. L. Zhang, Y. J. Qing, A. Fragasso, G. Maglia, A. Meller, H. Bayley, C. Dekker and Y. T. Long, Nat. Nanotechnol., 2022, 17, 1136–1146 CrossRef CAS PubMed.
- A. Meller, Mol. Cell, 2022, 82, 237–238 CrossRef CAS PubMed.
- K. Motone and J. Nivala, Nat. Methods, 2023, 20, 336–338 Search PubMed.
- H. Brinkerhoff, A. S. W. Kang, J. Q. Liu, A. Aksimentiev and C. Dekker, Science, 2021, 374, 1509–1513 Search PubMed.
- J. Nicholson, Chem, 2022, 8, 17–19 Search PubMed.
- S. L. Singh, K. Chauhan, A. S. Bharadwaj, V. Kishore, P. Laux, A. Luch and A. V. Singh, Int. J. Mol. Sci., 2023, 24, 6153 Search PubMed.
- B. Domon and R. Aebersold, Science, 2006, 312, 212–217 Search PubMed.
- T. E. Angel, U. K. Aryal, S. M. Hengel, E. S. Baker, R. T. Kelly, E. W. Robinson and R. D. Smith, Chem. Soc. Rev., 2012, 41, 3912–3928 Search PubMed.
- W. Timp and G. Timp, Sci. Adv., 2020, 6, eaax8978 Search PubMed.
- P. Edman, Acta Chem. Scand., 1950, 4, 283–293 Search PubMed.
- Y. S. Xiao, M. M. Vecchi and D. Y. Wen, Anal. Chem., 2016, 88, 10757–10766 Search PubMed.
- D. S. Talaga and J. Li, J. Am. Chem. Soc., 2009, 131, 9287–9297 Search PubMed.
- J. F. Li, Y. F. Huang, Y. Ding, Z. L. Yang, S. B. Li, X. S. Zhou, F. R. Fan, W. Zhang, Z. Y. Zhou, D. Y. Wu, B. Ren, Z. L. Wang and Z. Q. Tian, Nature, 2010, 464, 392–395 CrossRef CAS PubMed.
- Y. He, S. Su, T. T. Xu, Y. L. Zhong, J. A. Zapien, J. Li, C. H. Fan and S. T. Lee, Nano Today, 2011, 6, 122–130 CrossRef CAS.
- J. Cao, H. L. Liu, J. M. Yang, Z. Q. Li, D. R. Yang, L. N. Ji, K. Wang and X. H. Xia, ACS Sens., 2020, 5, 2198–2204 CrossRef CAS PubMed.
- H. L. Liu, J. Cao, S. Hanif, C. E. Yuan, J. Pang, R. Levicky, X. H. Xia and K. Wang, Anal. Chem., 2017, 89, 10407–10413 CrossRef CAS PubMed.
- E. A. Vitol, Z. Orynbayeva, M. J. Bouchard, J. Azizkhan-Clifford, G. Friedman and Y. Gogotsi, ACS Nano, 2009, 3, 3529–3536 CrossRef CAS PubMed.
- J. F. Masson, J. Breault-Turcot, R. Faid, H. P. Poirier-Richard, H. Yockell-Lelievre, F. Lussier and J. P. Spatz, Anal. Chem., 2014, 86, 8998–9005 CrossRef CAS PubMed.
- W. Li, L. Guo, X. L. Ding, Y. Ding, L. N. Ji, X. H. Xia and K. Wang, ACS Nano, 2024, 18, 19200–19207 CrossRef CAS PubMed.
- S. Cabello-Aguilar, A. Abou Chaaya, F. Picaud, M. Bechelany, C. Pochat-Bohatier, S. Yesylevskyy, S. Kraszewski, M. C. Bechelany, F. Rossignol, E. Balanzat, J. M. Janot, P. Miele, P. Dejardin and S. Balme, Phys. Chem. Chem. Phys., 2014, 16, 17883–17892 RSC.
- K. J. Freedman, S. R. Haq, J. B. Edel, P. Jemth and M. J. Kim, Sci. Rep., 2013, 3, 1638 CrossRef PubMed.
- W. Si and A. Aksimentiev, ACS Nano, 2017, 11, 7091–7100 CrossRef CAS PubMed.
- M. Iarossi, D. Darvill, A. Hubarevich, J. A. Huang, Y. Q. Zhao, A. F. De Fazio, D. B. O'Neill, F. Tantussi and F. De Angelis, Adv. Funct. Mater., 2023, 33, 2301934 CrossRef CAS.
- K. Luo, T. Ala-Nissila, S. C. Ying and R. Metzler, EPL, 2009, 88, 68006 Search PubMed.
- V. V. Palyulin, T. Ala-Nissila and R. Metzler, Soft Matter, 2014, 10, 9016–9037 RSC.
- G. Hu, H. Yan, G. H. Xi, Z. W. Gao, Z. Q. Wu, Z. H. Lu and J. Tu, IET Nanobiotechnol., 2023, 17, 257–268 CrossRef PubMed.
- A. Balijepalli, J. W. F. Robertson, J. E. Reiner, J. J. Kasianowicz and R. W. Pastor, J. Am. Chem. Soc., 2013, 135, 7064–7072 Search PubMed.
- S. K. Kannam, S. C. Kim, P. R. Rogers, N. Gunn, J. Wagner, S. Harrer and M. T. Downton, Nanotechnology, 2014, 25, 155502 CrossRef PubMed.
- D. Di Marino, E. L. Bonome, A. Tramontano and M. Chinappi, J. Phys. Chem. Lett., 2015, 6, 2963–2968 CrossRef CAS PubMed.
- H. Chen, L. Li, T. Zhang, Z. W. Qiao, J. Tang and J. Zhou, J. Phys. Chem. C, 2018, 122, 2070–2080 CrossRef CAS.
- M. A. Nejad and H. M. Urbassek, Biomolecules, 2019, 9, 204 Search PubMed.
- M. A. Nejad and H. M. Urbassek, J. Mol. Graphics Modell., 2019, 86, 228–234 CrossRef CAS PubMed.
- S. Ohayon, A. Girsault, M. Nasser, S. Shen-Orr and A. Meller, PLoS Comput. Biol., 2019, 15, e1007067 Search PubMed.
- G. Mitscha-Baude, B. Stadlbauer, S. Howorka and C. Heitzinger, ACS Nano, 2021, 15, 9900–9912 CrossRef CAS PubMed.
- W. Si, H. J. Yang, G. S. Wu, Y. Zhang and J. J. Sha, Nanoscale, 2021, 13, 15352–15361 Search PubMed.
- S. C. Liu, Y. L. Ying, W. H. Li, Y. J. Wan and Y. T. Long, Chem. Sci., 2021, 12, 3282–3289 Search PubMed.
- M. MacKenzie and C. Argyropoulos, Micromachines, 2023, 14, 459 Search PubMed.
- C. Maffeo, S. Bhattacharya, J. Yoo, D. Wells and A. Aksimentiev, Chem. Rev., 2012, 112, 6250–6284 CrossRef CAS PubMed.
- D. Fologea, J. Uplinger, B. Thomas, D. S. McNabb and J. L. Li, Nano Lett., 2005, 5, 1734–1737 Search PubMed.
- S. W. Kowalczyk, D. B. Wells, A. Aksimentiev and C. Dekker, Nano Lett., 2012, 12, 1038–1044 Search PubMed.
- L. Z. Sun, W. P. Cao and M. B. Luo, J. Chem. Phys., 2009, 131, 194904 Search PubMed.
- B. N. Anderson, M. Muthukumar and A. Meller, ACS Nano, 2013, 7, 1408–1414 CrossRef CAS PubMed.
- M. A. Paun, V. A. Paun and V. P. Paun, Polymers, 2022, 14, 1166 Search PubMed.
- F. Mondaini and L. Moriconi, Phys. Lett. A, 2014, 378, 1767–1772 Search PubMed.
- V. V. Lehtola, R. P. Linna and K. Kaski, Phys. Rev. E: Stat., Nonlinear, Soft Matter Phys., 2010, 81, 031803 Search PubMed.
- V. V. Lehtola, K. Kaski and R. P. Linna, Phys. Rev. E: Stat., Nonlinear, Soft Matter Phys., 2010, 82, 031908 Search PubMed.
- J. Sarabadani and T. Ala-Nissila, J. Phys.: Condens. Matter, 2018, 30, 274002 CrossRef PubMed.
- H. W. de Haan, D. Sean and G. W. Slater, Phys. Rev. E, 2018, 98, 022501 CrossRef PubMed.
- P. Y. Hsiao, Polymers, 2018, 10, 1229 Search PubMed.
- A. Bhattacharya and K. Binder, Phys. Rev. E: Stat., Nonlinear, Soft Matter Phys., 2010, 81, 041804 Search PubMed.
- M. Rycenga, C. M. Cobley, J. Zeng, W. Y. Li, C. H. Moran, Q. Zhang, D. Qin and Y. N. Xia, Chem. Rev., 2011, 111, 3669–3712 CrossRef CAS PubMed.
- K. M. Mayer and J. H. Hafner, Chem. Rev., 2011, 111, 3828–3857 CrossRef CAS PubMed.
- A. A. Al Balushi, A. Kotnala, S. Wheaton, R. M. Gelfand, Y. Rajashekara and R. Gordon, Analyst, 2015, 140, 4760–4778 RSC.
- C. Chen, J. A. Hutchison, P. Van Dorpe, R. Kox, I. De Vlaminck, H. Uji-i, J. Hofkens, L. Lagae, G. Maes and G. Borghs, Small, 2009, 5, 2876–2882 CrossRef CAS PubMed.
- C. Chen, Y. Li, S. Kerman, P. Neutens, K. Willems, S. Cornelissen, L. Lagae, T. Stakenborg and P. Van Dorpe, Nat. Commun., 2018, 9, 1733 CrossRef PubMed.
- E. C. Le Ru, M. Meyer and P. G. Etchegoin, J. Phys. Chem. B, 2006, 110, 1944–1948 CrossRef CAS PubMed.
- M. P. Cecchini, A. Wiener, V. A. Turek, H. Chon, S. Lee, A. P. Ivanov, D. W. McComb, J. Choo, T. Albrecht, S. A. Maier and J. B. Edel, Nano Lett., 2013, 13, 4602–4609 Search PubMed.
- S. Kerman, C. Chen, Y. Li, W. Van Roy, L. Lagae and P. Van Dorpe, Nanoscale, 2015, 7, 18612–18618 RSC.
- A. Longato, M. Vanzan, E. Colusso, S. Corni and A. Martucci, Small, 2023, 19, e2205522 CrossRef PubMed.
- R. Brilmayer, C. Forster, L. Zhao and A. Andrieu-Brunsen, Curr. Opin. Biotechnol., 2020, 63, 200–209 CrossRef CAS PubMed.
- M. Nguyen, A. Lamouri, C. Salameh, G. Levi, J. Grand, L. Boubekeur-Lecaque, C. Mangeney and N. Felidj, Nanoscale, 2016, 8, 8633–8640 RSC.
- I. Tijunelyte, I. Kherbouche, S. Gam-Derouich, M. Nguyen, N. Lidgi-Guigui, M. L. de la Chapelle, A. Lamouri, G. Levi, J. Aubard, A. Chevillot-Biraud, C. Mangeney and N. Felidj, Nanoscale Horiz., 2018, 3, 53–57 RSC.
- E. S. A. Goerlitzer, L. E. Speichermann, T. A. Mirza, R. Mohammadi and N. Vogel, Nanoscale Adv., 2020, 2, 394–400 RSC.
- A. N. Koya, M. Romanelli, J. Kuttruff, N. Henriksson, A. Stefancu, G. Grinblat, A. De Andres, F. Schnur, M. Vanzan, M. Marsili, M. Rahaman, A. Viejo Rodríguez, T. Tapani, H. Lin, B. D. Dana, J. Lin, G. Barbillon, R. Proietti Zaccaria, D. Brida, D. Jariwala, L. Veisz, E. Cortes, S. Corni, D. Garoli and N. Maccaferri, Appl. Phys. Rev., 2023, 10, 021318 CAS.
- J. A. Moore and J. C. L. Chow, Nano Express, 2021, 2, 022001 CrossRef CAS.
- L. B. Wright, P. M. Rodger, S. Corni and T. R. Walsh, J. Chem. Theory Comput., 2013, 9, 1616–1630 CrossRef CAS PubMed.
- P. Charchar, A. J. Christofferson, N. Todorova and I. Yarovsky, Small, 2016, 12, 2395–2418 CrossRef CAS PubMed.
- T. R. Walsh and M. R. Knecht, Chem. Rev., 2017, 117, 12641–12704 CrossRef CAS PubMed.
- F. Iori, R. Di Felice, E. Molinari and S. Corni, J. Comput. Chem., 2009, 30, 1465–1476 CrossRef CAS PubMed.
- M. Vanzan, M. Rosa and S. Corni, Nanoscale Adv., 2020, 2, 2842–2852 RSC.
- S. Brown, Nat. Biotechnol., 1997, 15, 269–272 CrossRef CAS PubMed.
- M. Sarikaya, C. Tamerler, D. T. Schwartz and F. O. Baneyx, Annu. Rev. Mater. Res., 2004, 34, 373–408 CrossRef CAS.
- Y. Lyu, L. M. Becerril, M. Vanzan, S. Corni, M. Cattelan, G. Granozzi, M. Frasconi, P. Rajak, P. Banerjee, R. Ciancio, F. Mancin and P. Scrimin, Adv. Mater., 2024, 36, e2211624 CrossRef PubMed.
- V. Amendola, R. Pilot, M. Frasconi, O. M. Marago and M. A. Iati, J. Phys.: Condens. Matter, 2017, 29, 203002 CrossRef PubMed.
- R. Pilot, R. Signorini, C. Durante, L. Orian, M. Bhamidipati and L. Fabris, Biosensors, 2019, 9, 57 CrossRef CAS PubMed.
- M. Inagaki, T. Isogai, K. Motobayashi, K. Q. Lin, B. Ren and K. Ikeda, Chem. Sci., 2020, 11, 9807–9817 RSC.
- M. Inagaki, K. Motobayashi and K. Ikeda, J. Phys. Chem. Lett., 2017, 8, 4236–4240 CrossRef CAS PubMed.
- R. Moldovan, E. Vereshchagina, K. Milenko, B. C. Iacob, A. E. Bodoki, A. Falamas, N. Tosa, C. M. Muntean, C. Farca and E. Bodoki, Anal. Chim. Acta, 2022, 1209, 339250 CrossRef CAS PubMed.
- M. Z. Tien, D. K. Sydykova, A. G. Meyer and C. O. Wilke, PeerJ, 2013, 1, e80 Search PubMed.
- W. Humphrey, A. Dalke and K. Schulten, J. Mol. Graphics Modell., 1996, 14, 33–38 Search PubMed.
- M. Belkin, S. H. Chao, M. P. Jonsson, C. Dekker and A. Aksimentiev, ACS Nano, 2015, 9, 10598–10611 CrossRef CAS PubMed.
- Y. Zhao, A. Hubarevich, A. F. De Fazio, M. Iarossi, J. A. Huang and F. De Angelis, Nano Lett., 2023, 23, 4830–4836 CrossRef CAS PubMed.
- W. R. Forsyth, J. M. Antosiewicz and A. D. Robertson, Proteins, 2002, 48, 388–403 Search PubMed.
- D. Van Der Spoel, E. Lindahl, B. Hess, G. Groenhof, A. E. Mark and H. J. Berendsen, J. Comput. Chem., 2005, 26, 1701–1718 CrossRef CAS PubMed.
- M. J. Abraham, D. L. van der Spoel and E. B. Hess and Gromacs development team, GROMACS User Manual version 2018.4, https://www.gromacs.org Search PubMed.
- H. Bekker, H. J. C. Berendsen, E. J. Dijkstra, S. Achterop, R. van Drunen, D. van der Spoel, A. Sijbers and K. H., et al., in Physics computing, ed. R. A. de Groot and J. Nadrchal, World Scientific, Singapore, 1993, vol. 92, pp. 252–256 Search PubMed.
- H. J. C. Berendsen, D. van der Spoel and R. van Drunen, Comput. Phys. Commun., 1995, 91, 43–56 CrossRef CAS.
- E. Lindahl, B. Hess and D. van der Spoel, J. Mol. Model., 2001, 7, 306–317 CrossRef CAS.
- B. Hess, C. Kutzner, D. van der Spoel and E. Lindahl, J. Chem. Theory Comput., 2008, 4, 435–447 Search PubMed.
- S. Pronk, S. Páll, R. Schulz, P. Larsson, P. Bjelkmar, R. Apostolov, M. R. Shirts, J. C. Smith, P. M. Kasson, D. van der Spoel, B. Hess and E. Lindahl, Bioinformatics, 2013, 29, 845–854 Search PubMed.
- S. Páll, M. J. Abraham, C. Kutzner, B. Hess and E. Lindahl, in Solving software challenges for exascale, ed. S. Markidis and E. Laure, Springer International Publishing Switzerland, London, 2015, pp. 3–27 Search PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1–2, 19–25 Search PubMed.
- W. L. Jorgensen and J. Tiradorives, J. Am. Chem. Soc., 1988, 110, 1657–1666 Search PubMed.
- W. L. Jorgensen, D. S. Maxwell and J. TiradoRives, J. Am. Chem. Soc., 1996, 118, 11225–11236 Search PubMed.
- G. A. Kaminski, R. A. Friesner, J. Tirado-Rives and W. L. Jorgensen, J. Phys. Chem. B, 2001, 105, 6474–6487 Search PubMed.
- C. D. Berweger, W. F. van Gunsteren and F. Müller-Plathe, Chem. Phys. Lett., 1995, 232, 429–436 Search PubMed.
- B. Hess, H. Bekker, H. J. C. Berendsen and J. G. E. M. Fraaije, J. Comput. Chem., 1997, 18, 1463–1472 Search PubMed.
- G. Bussi, D. Donadio and M. Parrinello, J. Chem. Phys., 2007, 126, 014101 Search PubMed.
- M. Parrinello and A. Rahman, J. Appl. Phys., 1981, 52, 7182–7190 CrossRef CAS.
- D. Frenkel and B. Smit, Understanding Molecular Simulation, Academic Press, Cornwall, 2002 Search PubMed.
- A. J. Bard and L. R. Faulkner, Electrochemical methods: Fundamentals and Applications, John Wiley & Sons, Inc., New York, 2001, p. 550 Search PubMed.
- In order to test the adequacy of our theoretical-computational model, we used the solution of the Poisson–Boltzmann equation for the case of planar symmetry as an approximate expression for the radial potential variation from the internal surface of the nanopore to the center of the pore. Then, integration of the corresponding Poisson equation over the volume inside the nanopore gave 58 and 220 ions for Vt = 160 mV and Vt = 220 mV, respectively, in very reasonable agreement with the 260 and 204 ions used in our computations.
- K. Inoue, N. Baden and M. Terazima, J. Phys. Chem. B, 2005, 109, 22623–22628 CrossRef CAS PubMed.
- S. Costantini, G. Colonna and A. M. Facchiano, Biochem. Biophys. Res. Commun., 2006, 342, 441–451 CrossRef CAS PubMed.
- D. Zanuy and C. Alemán, Biomacromolecules, 2001, 2, 651–657 CrossRef CAS PubMed.
- J. Dzubiella, J. Phys. Chem. B, 2010, 114, 7098–7103 CrossRef CAS PubMed.
- B. H. Zimm and J. K. Bragg, J. Chem. Phys., 1959, 31, 526–535 CrossRef CAS.
- W. Zhang and S. Nilsson, Macromolecules, 1993, 26, 2866–2870 CrossRef CAS.
- I. M. Derrington, T. Z. Butler, M. D. Collins, E. Manrao, M. Pavlenok, M. Niederweis and J. H. Gundlach, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 16060–16065 CrossRef PubMed.
- D. V. Verschueren, S. Pud, X. Shi, L. De Angelis, L. Kuipers and C. Dekker, ACS Nano, 2019, 13, 61–70 CrossRef CAS PubMed.
- B. J. Bennion and V. Daggett, Proc. Natl. Acad. Sci. U. S. A., 2003, 100, 5142–5147 CrossRef CAS PubMed.
- F. Rashid, S. Sharma and B. Bano, Protein J., 2005, 24, 283–292 Search PubMed.
- A. Das and C. Mukhopadhyay, J. Phys. Chem. B, 2009, 113, 12816–12824 Search PubMed.
- M. Tarek, Biophys. J., 2005, 88, 4045–4053 Search PubMed.
- N. Basdevant, D. Dessaux and R. Ramirez, Sci. Rep., 2019, 9, 15740 CrossRef PubMed.
- Z. T. Jiang, L. You, W. H. Dou, T. T. Sun and P. Xu, Polymers, 2019, 11, 282 CrossRef PubMed.
- J. Wang and Z. L. Li, J. Chem. Phys., 2024, 161, 094305 Search PubMed.
- P. Raiteri, P. Kraus and J. D. Gale, J. Chem. Phys., 2020, 153, 164714 CrossRef CAS PubMed.
- M. Niyozaliev, J. Matyakubov, D. Abduvokhidov, P. Attri, Z. T. Chen and J. Razzokov, J. Phys. D: Appl. Phys., 2024, 57, 285202 Search PubMed.
- H. Kim, B. Fábián and G. Hummer, J. Chem. Theory Comput., 2023, 19, 8919–8929 Search PubMed.
- M. T. H. Nguyen, D. Biriukov, C. Tempra, K. Baxova, H. Martinez-Seara, H. Evci, V. Singh, R. Sachl, M. Hof, P. Jungwirth, M. Javanainen and M. Vazdar, Langmuir, 2022, 38, 11284–11295 CrossRef CAS PubMed.
- I. Şoldănescu, A. Lobiuc, O. A. Caliman-Sturdza, M. Covasa, S. Mangul and M. Dimian, Biosensors, 2025, 15, 540 CrossRef PubMed.
- W. C. K. Poon and D. Andelman, Soft Condensed Matter Physics in Molecular and Cell Biology, Taylor & Francis Group, LLC, Boca Raton, USA, 2006, ch. 6 Search PubMed.
- L. Restrepo-Pérez, S. John, A. Aksimentiev, C. Joo and C. Dekker, Nanoscale, 2017, 9, 11685–11693 RSC.
| This journal is © The Royal Society of Chemistry 2026 |





