Open Access Article
Open Access ArticleA comprehensive review of the structural diversity, biosynthesis and chemical synthesis of Lasiodiplodia spp. natural products
Maria Michela Salvatore
 *a,
Marco Masi
*a,
Marco Masi
 bc,
Marina DellaGreca
bc,
Marina DellaGreca
 b and
Anna Andolfi
b and
Anna Andolfi
 *bc
*bc
aDepartment of Veterinary Medicine and Animal Production, University of Naples Federico II, 80137 Naples, Italy. E-mail: mariamichela.salvatore@unina.it
bDepartment of Chemical Sciences, University of Naples Federico II, 80126 Naples, Italy. E-mail: andolfi@unina.it
cBAT Center-Interuniversity Center for Studies on Bioinspired Agro-Environmental Technology, University of Naples Federico II, 80055 Portici (NA), Italy
First published on 21st April 2026
Abstract
Covering: up to the end of 2025
The genus Lasiodiplodia belongs to the Botryosphaeriaceae family. Over the past few decades, it has proven to be a great source of interesting natural products, including depsidones, jasmonates, preussomerins, and β-resorcylic acid derivatives. This review provides an update on the secondary metabolites of 13 species of Lasiodiplodia and their bioactivities, covering the period up to the end of 2025. Further insights into their proposed biosynthetic pathways are discussed, and key examples of the synthetic pathways leading to their production are highlighted.
1 Introduction
Lasiodiplodia is one of the most important plant pathogenic genus of Botryosphaeriaceae, a genus-rich family in the Dothideomycetes class. Species of the genus Lasiodiplodia are associated with various host plants, acting as primary or secondary pathogens and even as endophytes that, under stress conditions of the host, can switch their lifestyle and become pathogens.1,2 Consequently, species under this genus have garnered considerable research attention due to their global distribution and involvement in diseases affecting several economically important plants, causing extensive damage to crops. This is a concern that has further increased with climate change, which is altering the distribution and impact of plant pathogens,3 including Lasiodiplodia spp.4,5 In fact, rising temperatures have been identified as the main abiotic factor exacerbating the impact of plant diseases by promoting fungal growth and colonization with economically relevant consequences.5Since its introduction in 1894, Lasiodiplodia has had an intricate taxonomic history. It was first introduced by Ellis and Everhart with Lasiodiplodia tubericola as the type species,6 although the genus was formally described in 1896 by Clendenin.7 Subsequently, prevailing over the epithet tubericola, Lasiodiplodia theobromae (Pat.) Griffon & Maubl. has been proposed and recognized as the type species of the Lasiodiplodia genus. The use of traditional morphology in fungal species identification for many years led to Lasiodiplodia being treated as a monotypic genus. This trend ended with the introduction of DNA sequence-based methods, which recognized L. theobromae as a complex of different cryptic species8 and enabled the identification of isolates related to but different from the known taxa.9,10 Hence, from that moment onwards, the widespread use of biomolecular markers for fungal identification has led to the proliferation of new species belonging to the genus Lasiodiplodia, such as Lasiodiplodia pseudotheobromae,8 Lasiodiplodia mediterranea,11 Lasiodiplodia venezuelensis9 and Lasiodiplodia brasiliensis.12 To complicate matters further, until recently, it was common practice in mycology to name both the asexual and sexual stages of the same fungus differently (i.e., dual nomenclature). This nomenclature system increased the confusion within the Lasiodiplodia genus because, for example, Botryosphaeria rhodina (Berk & M.A. Curtis) Arx was used as the teleomorphic name of L. theobromae. This practice is no longer acceptable according to the latest International Code of Nomenclature for algae, fungi, and plants.13
Although the employment of new molecular tools for fungal taxonomy has improved the classification, the phylogenetic tree of the genus Lasiodiplodia is continuously updated due to the detection of new taxonomic relationships among botryosphaeriaceous fungi that cause reconsiderations of the nomenclature.14
Despite the long list of species belonging to the genus Lasiodiplodia gradually emerging in recent decades,15 L. theobromae remains the most reported and studied species. This is also due to its great ability to infect a wide range of crops and trees, causing rotting and dieback in most infected plant hosts, thereby resulting in significant yield losses and reduced fruit quality.16 Furthermore, L. theobromae is also recognized as an emerging human pathogen associated with several diseases, including onychopathy,17 keratitis18 and rhinosinusitis.19
Species of Lasiodiplodia are prolific sources of secondary metabolites belonging to different classes of natural products and exhibiting a broad range of activities. The diverse secondary metabolism observed in the Lasiodiplodia genus may be related to some of the peculiar features of these fungi, such as their global distribution, different symbiotic lifestyles, host adaptability and capacity to cause diverse symptoms in infected hosts. Thus, secondary metabolites produced by Lasiodiplodia spp. may be responsible for the dynamic host–fungus–environment interaction, which is critical due to its ecological implications and biotechnological applications.
In this review, we present a comprehensive overview of the secondary metabolites produced by the fungi of the genus Lasiodiplodia, their bioactivities, selected biosynthetic pathways and examples of synthetic approaches.
To date, over 270 bioactive compounds have been reported from cultures of the species of Lasiodiplodia and many of them showed interesting bioactivities, such as phytotoxic, cytotoxic and antimicrobial activities.
PubMed, Google Scholar, Web of Science, SciFinder and Scopus were used to collect all the published articles about Lasiodiplodia spp. natural products. Data are organized in tables and figures according to their chemical structures and then the chemical and biological aspects are discussed.
2 Classification, structures and occurrence of Lasiodiplodia spp. natural products
Sixty years ago, Gupta et al.20 described the production, isolation and biological properties of a new secondary metabolite from an isolate of L. theobromae, which was named botryodiplodin. Since then, over 270 metabolites have been identified as products of Lasiodiplodia spp. and some of them have been reported exclusively from this fungal genus.The taxonomic and phylogenetic aspects of this fungal genus have long been a research subject, and this has inevitably affected the study of the metabolism of Lasiodiplodia spp. (Fig. 1A). In fact, the first decade of the 2000s was crucial for this genus, which, as mentioned in the previous section, experienced dramatic taxonomic changes following the advent of molecular identification to improve fungal characterization and the introduction of the principle “one species-one name” in mycology.13 Consequently, the study of metabolites produced by species other than L. theobromae began about 10 years ago, when the evolution of the publications related to Lasiodiplodia metabolites presented a considerable increase in total number (see Fig. 1A). This trend demonstrates a significant and growing interest by the scientific community in the detection of Lasiodiplodia metabolites, which is currently related to their structural diversity, bioactivity and implications of their occurrence. Thus far, 13 species belonging to the genus Lasiodiplodia have been investigated for the production of secondary metabolites.
 | ||
| Fig. 1 (A) Evolution of the number of publications on the metabolites of Lasiodiplodia spp. over the years. (B) Pie-of-pie chart of the metabolites detected per species. | ||
The pie charts in Fig. 1B clearly show that, among Lasiodiplodia spp., a higher number of metabolites is reported for L. theobromae, but we must not forget that this species is also more studied than the other 12 due to its early detection (Fig. 1A). Hence, it is quite possible that the imbalance in metabolite production by Lasiodiplodia spp. that emerges from Fig. 1B reflects what has occurred taxonomically for this genus since its first isolation.
Apart from that, L. theobromae is a very promising source of bioactive secondary metabolites and, even if less investigated, it seems that all Lasiodiplodia spp. share a good ability to produce bioactive compounds in vitro.
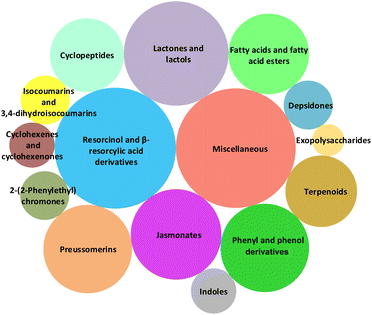
As can be seen in Fig. 2, compounds produced by fungi from the genus Lasiodiplodia can be collected in 15 classes according to their chemical structures. The bubble chart in Fig. 2 allows a quick and compact visualization of the compound abundance in each class, also allowing comparisons by bubble size. It can be easily seen that Lasiodiplodia spp. are good producers of compounds that can be classified as “resorcinol and β-resorcylic acid derivatives”, “lactones and lactols” and “jasmonates”.
For a more organized and clearer discussion, some metabolites have been included in a single class even if they exhibit the proper structural features of two or more classes. This is expected, given the structural complexity of some natural products, which makes a clear separation between the groups impossible.
For completeness, a few primary metabolites occasionally detected in Lasiodiplodia spp. cultures are mentioned (e.g., glycerol, succinic acid, and uracil), but they are treated briefly in the following sections to preserve the focus on the structural diversity of secondary metabolites.
It should be noted that manuscripts listing unnatural or frequently occurring compounds as contaminants have been deliberately excluded from the discussion.21–23
2.1 Cyclohexenols and cyclohexenones
Cyclohexenols and cyclohexenones are polyhydroxylated polyketides commonly found as natural products.24–27 Despite their frequent occurrence, to date, these compounds have been identified in the Lasiodiplodia genus only from L. theobromae, which, consequently, could mean that this is the only Lasiodiplodia species capable of producing compounds of this family (Table 1). The first identified compound of this class is an epoxycyclohexanediol named theobroxide (1), which owes its trivial name to its identification in cultures of L. theobromae IFO 31059.28 Although five novel related compounds have subsequently been identified (2–6), theobroxide is the only one containing an epoxide group (Fig. 3).| No. | Compound | Fungal producer (strain) | Source | Ref. |
|---|---|---|---|---|
| Cyclohexenols and cyclohexenones | ||||
| 1 | Theobroxide | Lasiodiplodia theobromae (IFO 31059) | — | 28 |
| L. theobromae (OCS71) | — | 29 | ||
| 2 | (4S,5S)-4,5-Dihydroxy-2-methyl-cyclohex-2-enone | L. theobromae (OCS71) | — | 29 |
| 3 | (4R,5R)-4,5-Dihydroxy-3-methylcyclohex-2-enone | L. theobromae (OCS71) | — | 29 |
| 4 | (4S,5S)-4,5-Dihydroxy-3-methylcyclohex-2-enone | L. theobromae (OCS71) | — | 29 and 31 |
| 5 | (3aR,4S,5R,7aS)-4,5-Dihydroxy-6-methyl-3a,4,5,7a-tetrahydrobenzo[d][1,3]dioxol-2-one | L. theobromae (OCS71) | — | 29 |
| L. theobromae (IFO 31059) | — | 30 | ||
| 6 | (3aS,4R,5S,7aR)-4,5-Dihydroxy-7-methyl-3a,4,5,7a-tetrahydrobenzo[1,3]dioxol-2-one | L. theobromae (OCS71) | — | 31 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Cyclopeptides | ||||
| 7 | Aldsulfin | Lasiodiplodia pseudotheobromae (FKI-4499) | Soil | 41 |
| 8 | Cyclo-(D-N-OH-Ala, D-Trp) | Lasiodiplodia chiangraiensis (MFLUCC21-0003) | Decaying wood | 36 |
| 9 | Cyclo-(Leu–Pro) | Lasiodiplodia iranensis (F0619) | Avicennia germinans | 42 |
| L. theobromae (AUMC 8903) | Dracaena draco | 43 | ||
| L. theobromae (SNFF) | Solanum nigrum | 44 | ||
| L. theobromae (LA-SOL3, LA-SV1) | Vitis vinifera | 45 | ||
| 10 | Cyclo-(Phe–Pro) | L. theobromae (SNFF) | S. nigrum | 44 |
| L. theobromae (LA-SOL3, LA-SV1) | V. vinifera | 45 | ||
| 11 | Cyclo-(Trp–Ala) | L. theobromae (CAA019 and CBS339.90) | Cocos nucifera and human | 46 |
| L. pseudotheobromae (F2) | Illigera rhodantha | 35 | ||
| L. chiangraiensis (MFLUCC21-0003) | Decaying wood | 36 | ||
| 12 | (3R,6R)-3-((1H-Indol-3-yl)methyl)-1-hydroxy-6-methylpiperazine-2,5-dione | L. chiangraiensis (MFLUCC21-0003) | Decaying wood | 36 |
| 13 | Lasiodipline A | L. pseudotheobromae (F2) | I. rhodantha | 35 |
| 14 | Lasiodipline B | L. pseudotheobromae (F2) | I. rhodantha | 35 |
| 15 | Lasiodipline C | L. pseudotheobromae (F2) | I. rhodantha | 35 |
| L. pseudotheobromae (FKI-4499) | Soil | 41 | ||
| 16 | Lasiodipline D | L. pseudotheobromae (F2) | I. rhodantha | 35 |
| 17 | Lasiodipline E | L. pseudotheobromae (F2) | I. rhodantha | 35 |
| 18 | Lasiodipline F | L. pseudotheobromae (F2) | I. rhodantha | 35 |
| 19 | Lasiodipline G | L. chiangraiensis (MFLUCC21-0003) | Decaying wood | 36 |
| 20 | Maculosin | L. theobromae (AUMC 8903) | D. draco | 43 |
| 21 | Clavatustide C | L. chiangraiensis | Decaying wood | 36 |
| 22 | Scopularide A | L. theobromae | Musa paradisiaca | 40 |
Theobroxide is known for inducing potato micro-tuber formation,28 and when tested for plant growth regulation on seedlings of Nicotiana tabacum, along with some related compounds (2–4), it showed inhibitory effects (Table S1).29 Compounds 5 and 6 possess a further subunit constituted by a five-membered cyclic carbonate, a structural feature not commonly found in secondary metabolites. These two novel theobroxide-related compounds have been isolated from the mycelia and culture filtrates of L. theobromae IFO 31059 and OCS71.29–31
2.2 Cyclopeptides
Diketopiperazines are the smallest cyclopeptides obtained by the condensation of two α-amino acids and characterized by a broad structural diversity. These compounds are abundant in nature as products of plants, animals and microorganisms.32–34 To date, fourteen diketopiperazines have been found in cultures of Lasiodiplodia spp. (Table 1 and Fig. 4). (3R,6R)-3-((1H-Indol-3-yl)methyl)-1-hydroxy-6-methylpiperazine-2,5-dione (12) and lasiodiplodines A–G (13–19, respectively) share the same framework obtained by the condensation of tryptophan and alanine residues, differing in some substituents. In particular, L. pseudotheobromae is a good producer of sulfureous diketopiperazines with a di- and tri-sulfide bridge (7, 16 and 19) or methylthio groups (13–15, 17 and 18). Lasiodipline F (18) is a diketopiperazine possessing an unusual skeleton including a characteristic tetracyclic indole unit.35 Recently, a new thiodiketopiperazine, named lasiodipline G (19), has been isolated from cultures of a decaying wood-derived strain of L. chiangraiensis.36 Furthermore, this new compound exhibited significant cytotoxic activities against several cell lines (Table S1). Promising data on the antimicrobial activities of lasiodipline E (17) were obtained by Wei et al.35 In fact, 17 was demonstrated to be antibacterial against the clinical strains Streptococcus sp., Bacteroides vulgatus, Peptostreptococcus sp. and Veillonella parvula (Table S1).Clavatustide C (21) and scopularide A (22) are cyclodepsipeptides, cyclic peptides made up of both proteinogenic and nonproteinogenic amino acids linked by amide and ester bonds.37 In particular, clavatustide C is a cyclo-(L-Ile–L-Ile–L-Leu–L-Lue–L-Leu) and identified as product of L. chiangraiensis. It must be noted that this compound was wrongly reported as clavatustide B in the original paper, but experimental data in the SI confirm the structure of 21. Thus, its correct trivial name and structure are reported in Table 1 and Fig. 4.36 Clavatustide C has the same amino acid composition (i.e., two L-Ile and three L-Leu), but a different sequence, as viscumamide, a metabolite produced by the fungus Paecilomyces sp.38 Viscumamide contains the most common cyclic sequence of these five peptide units in which two L-isoleucines are embedded between three L-leucines.39
Scopularide A (22) is also known as cyclo-(3-hydroxy-4-methyldecanoyl-Gly–L-Val–D-Leu–L-Ala–L-Phe) and has been identified in cultures of L. theobromae isolated from Musa paradisiaca.40
2.3 Depsidones
Depsidones are metabolites that share a conserved 11H-dibenzo[b,e][1,4]dioxepin-11-one ring substituted in different positions with various substituents. These compounds are mostly found in lichens47–49 and only a few of them have been isolated from non-lichen sources.50,51 Five previously undescribed depsidones, such as botryorhodines A–D and I (23–27), have been isolated from cultures of L. theobromae, along with 3,8-dihydroxy-4-(methoxymethyl)-1,6-dimethyl-11H-dibenzo[b,e][1,4]dioxepin-11-one (28) and simplicildone A (29) (Table 2), which had been previously reported from a marine derived fungus52 and from the endophytic fungus Simplicillium sp.,53 respectively.| No. | Compound | Fungal producer (strain) | Source | Ref. |
|---|---|---|---|---|
| Depsidones | ||||
| 23 | Botryorhodine A | Lasiodiplodia theobromae (TBRC 15112) | Achyranthes aspera | 54 |
| L. theobromae (BPPCA 144) | Aglaia argentea | 55 | ||
| L. theobromae | Bidens pilosa | 56 | ||
| L. theobromae (M4.2-2) | Mangrove sediment | 57 | ||
| 24 | Botryorhodine B | L. theobromae | B. pilosa | 56 |
| L. theobromae (M4.2-2) | Mangrove sediment | 57 | ||
| 25 | Botryorhodine C | L. theobromae | B. pilosa | 56 |
| 26 | Botryorhodine D | L. theobromae | B. pilosa | 56 |
| L. theobromae (M4.2-2) | Mangrove sediment | 57 | ||
| 27 | Botryorhodine I | L. theobromae (M4.2-2) | Mangrove sediment | 57 |
| 28 | 3,8-Dihydroxy-4-(methoxymethyl)-1,6-dimethyl-11H-dibenzo[b,e][1,4]dioxepin-11-one | L. theobromae (M4.2-2) | Mangrove sediment | 57 |
| 29 | Simplicildone A | L. theobromae (M4.2-2) | Mangrove sediment | 57 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Exopolysaccharides | ||||
| 30 | Lasiosan | Lasiodiplodia sp. (B2) | Musa paradisiaca | 61 |
| 31 | Lasiodiplodan | L. theobromae (CCT3966) | — | 67 |
| L. theobromae (MMPI) | Annona squamosa | 62, 63 and 68 | ||
| L. theobromae (MMBJ) | — | 69 | ||
| 32 | Botryosphaerans | L. theobromae (MAMB-05) | — | 66 and 70–72 |
| L. theobromae (RCYU 30101) | — | 73 | ||
| L. theobromae | — | 65 | ||
| L. theobromae (DABAC-P82) | — | 74 | ||
As can be seen in Fig. 5, depsidones produced by L. theobromae share the same basic framework but differ in some substituents (i.e. methyl and hydroxyl groups) on the phenyl moieties (23–27).54–57
2.4 Exopolysaccharides
Bacteria and fungi are known to produce extracellular polysaccharides with different structural complexities.58,59 Exopolysaccharides exhibit peculiar structural features, including monosaccharide composition, type and configuration of glycosidic linkages, number of residues, and degree of branching. These biopolymers play diverse roles in the life cycle of producing microorganisms. In particular, exopolysaccharides provide protection to microbial cells during infection, confer tolerance to various antimicrobial agents and are involved in biofilm formation.60Some Lasiodiplodia spp. have been identified as exopolysaccharide producers (Table 2). Lasiosan (30) is a glucomannan produced by a strain of Lasiodiplodia sp. isolated from a spoiled banana sample. The component sugars of lasiosan are β-(1→4)-linked D-mannose and D-glucose in a ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.61 Some strains of L. theobromae biosynthesize lasiodiplodan, an exocellular (1→6)-β-glucan existing in a triple-helix conformation (Fig. 6).62,63 Lasiodiplodan (31) is an unusual glucan, as it is a linear (1→6)-β-D-glucan, whereas glucans are mainly of the (1→3)-β type (Fig. 6).64 A comparison of the fermentative parameters of L. theobromae MMPI to evaluate the lasiodiplodan yield showed that the maximum production can be obtained in agitated flasks with urea as the nitrogen source and glucose or maltose as the carbon source.62
1.61 Some strains of L. theobromae biosynthesize lasiodiplodan, an exocellular (1→6)-β-glucan existing in a triple-helix conformation (Fig. 6).62,63 Lasiodiplodan (31) is an unusual glucan, as it is a linear (1→6)-β-D-glucan, whereas glucans are mainly of the (1→3)-β type (Fig. 6).64 A comparison of the fermentative parameters of L. theobromae MMPI to evaluate the lasiodiplodan yield showed that the maximum production can be obtained in agitated flasks with urea as the nitrogen source and glucose or maltose as the carbon source.62
Unlike 30 and 31, botryosphaerans (32) are branched exopolysaccharides produced by L. theobromae. The structure of the first botryosphaeran isolated was characterised as a (1→3; 1→6)-β-D-glucan, with approximately 22% side branching comprising single (1→6)-β-linked glucosyl and (1→6)-β-linked diglucosyl residues (Fig. 6). Comparing botryosphaeran production by L. theobromae on diverse carbon sources, it was observed that the carbon source affected the side chain structures, but not the main chain constitution of 32. Indeed, botryosphaerans have the same backbone composed of glucopyranosyl units (1→3)-linked and substituted at O-6 by glucosyl and diglucosyl units, but when the fungus was grown on fructose, the produced exopolysaccharide had 31% side-branching, a branch point to every three glucose units in the main chain, while when the carbon source was sucrose, the obtained exopolysaccharide was less branched (21%), with a branch point for every five glucosyl residues in the main chain. The result obtained from L. theobromae grown on sucrose is very similar to the one obtained when the fungus was grown on glucose (22% side branching).65,66 The production of exopolysaccharides by Lasiodiplodia spp. is particularly relevant because these compounds were found to be significantly active during in vivo assays (Fig. S1).
2.5 Fatty acids and fatty acid esters
A variety of free fatty acids and naturally occurring esters of fatty acids was identified in cultures of Lasiodiplodia spp. (Table 3). Some of them are very common, such as palmitic acid (33) and oleic acid (41), which are the most common saturated and unsaturated fatty acids, respectively, found in animals, plants and microorganisms (Fig. 7).75–77 Fatty acids are typically considered primary metabolites, as they are essential for the growth, development, and reproduction of living organisms.78 However, in Lasiodiplodia spp., they may also act as secondary metabolites, particularly when involved in plant–fungus interactions. In fact, the high quantities and the wide variety of fatty acids produced by Lasiodiplodia spp. might be linked to the capacity of these fungi to infect a broad host range. Fatty acids and modified fatty acids play a pivotal role in plant–microbe interactions, serving diverse functions, such as signalling and virulence factors.79 Furthermore, fatty acids represent the starting material for many secondary metabolites involved in fungal virulence, such as jasmonates.80| No. | Compound | Fungal producer (strain) | Source | Ref. |
|---|---|---|---|---|
| Fatty acids and fatty acids esters | ||||
| 33 | Hexadecanoic acid (palmitic acid) | Lasiodiplodia citricola (ALG111 and ALG34) | Citrus sinensis | 84 |
| Lasiodiplodia theobromae (CBS 122127 and 2334) | — | 96 | ||
| 34 | 9-Hydroxy-10-oxo-12Z,15Z-octadecadienoic acid | Lasiodiplodia theobromae (CBS 122127 and 2334) | — | 96 |
| 35 | Methyl hexadecanoate | L. theobromae (UCD256Ma and MXL28) | Vitis vinifera | 81 |
| Lasiodiplodia pseudotheobromae (IBRL OS-64) | Ocimum sanctum | 97 | ||
| 36 | Ethyl hexadecanoate | L. theobromae (UCD256Ma and MXL28) | V. vinifera | 81 |
| 37 | 2-Methylpropyl hexadecanoate | L. theobromae (UCD256Ma and MXL28) | V. vinifera | 81 |
| 38 | Bis 2-ethylhexyl hexanedioate | L. pseudotheobromae (IBRL OS-64) | O. sanctum | 97 |
| 39 | Octadecanoic acid (stearic acid) | L. theobromae (CBS 122127) | — | 96 |
| L. pseudotheobromae (APR5) | Andrographis paniculata | 98 | ||
| 40 | Methyl octadecanoate | L. pseudotheobromae (IBRL OS-64) | O. sanctum | 97 |
| 41 | (Z)-9-Octadecenoic acid (oleic acid) | L. theobromae (CBS 122127) | — | 96 |
| L. pseudotheobromae (APR5) | A. paniculata | 98 | ||
| 42 | 9-Methyl (Z)-9-octadecenoate | L. theobromae (UCD256Ma and MXL28) | V. vinifera | 81 |
| 43 | Ethyl octadecanoate | L. theobromae (UCD256Ma and MXL28) | V. vinifera | 81 |
| 44 | Ethyl (Z)-9-octadecenoate | L. theobromae (UCD256Ma and MXL28) | V. vinifera | 81 |
| 45 | Ethyl (E)-9-octadecenoate | L. theobromae (UCD256Ma) | V. vinifera | 81 |
| 46 | 9,12-Octadecadienoic acid (linoleic acid) | L. theobromae (CBS 122127 and 2334) | — | 96 |
| L. citricola (ALG111, ALG39, ALG81 and ALG34) | C. sinensis | 84 | ||
| 47 | Methyl (Z,Z)-9,12-octadecadienoate | L. theobromae (UCD256Ma and MXL28) | V. vinifera | 81 |
| 48 | Ethyl-(Z,Z)-9,12-octadecadienoate | L. theobromae (UCD256Ma and MXL28) | V. vinifera | 81 |
| 49 | (Z,Z,Z)-9,12,15-Octadecatrienoic acid (linolenic acid) | L. citricola (ALG111 and ALG39) | C. sinensis | 84 |
| 50 | Eicosanoic acid | L. theobromae (CBS 122127) | — | 96 |
| 51 | Ethyl-(Z,Z,Z)-9,12,15-octadecatrienoate | L. theobromae (UCD256Ma and MXL28) | V. vinifera | 81 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Indoles | ||||
| 52 | Indole | L. theobromae (LA-SOL3, LA-SV1) | V. vinifera | 45 |
| 53 | Indole-3-acetic acid | L. theobromae (2334, 1517 and 83) | C. cinensis and wood | 85 |
| L. citricola (ALG111) | C. sinensis | 84 | ||
| 54 | Indole-3-butyric acid | L. theobromae (2334, 1517 and 83) | C. cinensis and wood | 85 |
| 55 | Indole-3-carbaldehyde | L. pseudotheobromae (LPS-1) | Ilex cornuta | 99 |
| L. theobromae (LA-SOL3, LA-SV1) | V. vinifera | 45 | ||
| L. theobromae | — | 100 | ||
| L. theobromae (SNFF) | Solanum nigrum | 44 | ||
| 56 | Indole-3-carboxylic acid | L. theobromae (CAA019, CBS339.90) | Cocos nucifera and human | 46 |
| L. theobromae (LA-SOL3, LA-SV1) | V. vinifera | 45 and 101 | ||
| L. pseudotheobromae (LPS-1) | I. cornuta | 99 | ||
| L. theobromae | Bidens pilosa | 56 | ||
| L. theobromae | — | 100 | ||
| L. theobromae (CSS01s) | V. vinifera | 102 | ||
| Lasiodiplodia chiangraiensis (MFLUCC21-0003) | Decaying wood | 36 | ||
| L. citricola (ALG111 and ALG39) | C. sinensis | 84 | ||
| Lasiodiplodia sp. | Handroanthus impetiginosus | 103 | ||
| 57 | Indole-3-propionic acid | L. theobromae (2334, 1517 and 83) | C. cinensis and wood | 85 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Isocoumarins and 3,4-dihydroisocoumarins | ||||
| 58 | (−)-Mellein | L. theobromae (BPPCA 144) | Aglaia argentea | 55 |
| Lasiodiplodia laeliocattleyae (CMM0206) | V. vinifera | 104 | ||
| L. theobromae (MJ2211) | Vitex pinnata | 105 | ||
| L. theobromae (LA-SOL3, LA-SV1) | V. vinifera | 45 and 101 | ||
| L. theobromae (GK-1) | C. nucifera | 106 | ||
| L. theobromae (M4.2-2) | Mangrove sediment | 57 | ||
| Lasiodiplodia euphorbiaceicola (CMM0181) | V. vinifera | 107 | ||
| L. theobromae (PSU-M35) | Garcinia mangostana | 108 | ||
| L. theobromae | — | 100 | ||
| L. theobromae (IFO 31059) | — | 28 | ||
| L. pseudotheobromae (C1136) | Tridax procumbus | 109 | ||
| L. venezuelensis (A02EtM) | Astrocaryum sciophilum | 95 | ||
| L. theobromae (SJF-1) | Syzygium cumini | 93 | ||
| L. theobromae (TBRC 15112) | A. aspera | 54 | ||
| L. citricola (ALG111, ALG39, ALG81 and ALG34) | C. sinensis | 84 | ||
| L. theobromae (NSTRU-PN1.4) | Soil | 110 | ||
| 59 | (−)-(3R,4R)-4-Hydroxymellein | L. theobromae (CAA019 and CBS339.90) | C. nucifera and human | 46 |
| L. theobromae (LA-SOL3 and LA-SV1) | V. vinifera | 45 and 101 | ||
| L. theobromae (#009) | Psidium guajava | 111 | ||
| L. euphorbiaceicola (CMM0181) | V. vinifera | 107 | ||
| L. theobromae (PSU-M35) | G. mangostana | 108 | ||
| L. theobromae | — | 100 | ||
| L. venezuelensis (A02EtM) | A. sciophilum | 95 | ||
| L. citricola (ALG111, ALG39, ALG81 and ALG34) | C. sinensis | 84 | ||
| 60 | (−)-(3R,4S)-4-Hydroxymellein | L. theobromae (CAA019 and CBS339.90) | C. nucifera and human | 46 |
| L. euphorbiaceicola (CMM0181) | V. vinifera | 107 | ||
| L. theobromae (PSU-M35) | G. mangostana | 108 | ||
| L. venezuelensis (A02EtM) | A. sciophilum | 95 | ||
| Lasiodiplodia sp. | H. impetiginosus | 103 | ||
| 61 | (−)-(3R)-5-Hydroxymellein | L. theobromae (PSU-M35) | G. mangostana | 108 |
| 62 | (+)-(3R,4S)-4,5-Dihydroxymellein | L. venezuelensis (A02EtM) | A. sciophilum | 95 |
| 63 | Orthosporin | L. theobromae | Musa paradisiaca | 112 |
| 64 | Citreoisocoumarinol | L. theobromae | M. paradisiaca | 40 |
Investigations conducted on cultures of L. theobromae strains isolated in California, Mexico showed a high production and a wide variety of fatty acids and their esters with significant effects on tobacco plants.81 In fact, fatty acids could be considered plant growth regulators due to their ability to affect tobacco germination and early growth (Table S1).
The lipid metabolism of L. theobromae seems to be affected by nutrient availability. Indeed, when testing different carbon sources, the production of different fatty acids was observed, which could have implications for pathogenicity.81,82
2.6 Indoles
Indole represents the scaffold of a large family of organic compounds comprising over a thousand members. In fact, several natural products exhibit highly complex structures characterised by an indole nucleus.83 Six indoles, including the parent compound of this class of metabolites, have been identified in cultures of Lasiodiplodia spp. (Table 3 and Fig. 8). Although indoles have wide structural diversity, those produced by Lasiodiplodia spp. are very simple, featuring an additional group at C-3 of the pyrrole ring. Among them, indole 3-carboxylic acid (56) is the most common in Lasiodiplodia spp. cultures. Besides this compound, Lasiodiplodia citricola and L. theobromae also produce indole 3-acetic acid (53), the most studied plant hormone belonging to the auxin class.84,852.7 Isocoumarins and 3,4-dihydroisocoumarins
Isocoumarin is a heterocyclic compound and a key structural motif of a large group of natural products. These compounds comprise a α-pyranone condensed with one aromatic ring and are structural isomers of coumarins (Table 3).3,4-Dihydroisocoumarins are a subgroup of isocoumarins, also named melleins like their parent compound, i.e. (−)-mellein. (−)-Mellein (58) was first reported in 1933 as a product of Aspergillus melleus,86 and subsequently this compound and its derivatives have been frequently found in fungal species such as Cladosporium,87 Fusarium88 and Penicillium89,90 as well as several botryosphaeriaceous species.26,27,91,92 (−)-Mellein was extensively studied for its bioactivities, showing promising results as an antimicrobial agent (Table S1).93
(+)-(3R,4S)-4,5-Dihydroxymellein (62) has been reported from L. venezuelensis isolated from Astrocaryum sciophilum palm tree leaves.94,95 However, there is an inaccuracy in the study by Pellissier et al.94 in 2021, because the authors reported in their paper the structure of its diastereomer (i.e. (+)-(3R,4R)-4,5-dihydroxymellein). After checking the data reported in the manuscript,94 we can conclude that (+)-(3R,4S)-4,5-dihydroxymellein was detected and the correct structure is reported in Fig. 9.
2.8 Jasmonates
Jasmonates are an important subgroup of oxylipins derived from linolenic acid, including the free acid and several conjugates, which are signalling molecules produced by plants and fungi. (−)-Jasmonic acid (65, 3-oxo-2-(2′-pentenyl)-cyclopentaneacetic acid) is the basic structure of this naturally occurring family of compounds, which is characterized by a cyclopentanone ring with a pentenyl and a carboxylic acid side chain. This compound occurs in the essential oils of Jasminum grandiflorum along with its methyl ester (66).113 Even if jasmonates are particularly known as plant hormones involved in development processes, Aldridge et al. in 1971 first isolated (−)-jasmonic acid from L. theobromae.100 Subsequently, several findings have confirmed that numerous species of Lasiodiplodia are capable of synthesizing compounds belonging to this class (Table 4). This could be the result of a phenomenon frequently observed in the genomes of prokaryotes and eukaryotes named horizontal gene transfer. It is defined as the exchange and stable integration of genetic material among phylogenetically distant species, which could significantly impact their metabolic capabilities.114,115 Indeed, comparative genomic and transcriptomic studies on genome evolution and the origin of pathogenicity in botryosphaeriaceous fungi demonstrated the role of genome expansion and horizontal gene transfer in the evolutionary adaptation of Botryosphaeriaceae.116,117| No. | Compound | Fungal producer (strain) | Source | Ref. |
|---|---|---|---|---|
| Jasmonates | ||||
| 65 | (−)-Jasmonic acid | Lasiodiplodia theobromae | Mangifera indica, Cocos nucifera, Passiflora edulis, Carica papaya, Paullinia cupana, and Theobroma cacao | 122 |
| L. theobromae (2334, 1517 and 83) | Citrus cinensis, wood | 85 | ||
| L. theobromae | Rosa species | 132 | ||
| L. theobromae (CAA019 and CBS339.90) | C. nucifera and human | 46 | ||
| L. theobromae (LA-SOL3, LA-SV1) | Vitis vinifera | 45 and 101 | ||
| Lasiodiplodia mediterranea (B6) | V. vinifera | 131 | ||
| Lasiodiplodia sp. (BL 101) | V. vinifera | 129 | ||
| Lasiodiplodia pseudotheobromae (LPS-1) | Ilex cornuta | 99 | ||
| Lasiodiplodia brasiliense (CMM0418) | V. vinifera | 107 | ||
| Lasiodiplodia crassispora (CMM0390) | V. vinifera | 107 | ||
| Lasiodiplodia iranensis (CMM0840) | V. vinifera | 107 | ||
| Lasiodiplodia pseudotheobromae (CMM0204) | V. vinifera | 107 | ||
| L. theobromae (D 7/2) | Citrus sinensis | 123–125 | ||
| L. theobromae | — | 100 | ||
| L. theobromae (IFO 31059) | — | 28 | ||
| L. theobromae (CSS01s) | V. vinifera | 102 | ||
| L. iranensis (CCTCC no. M2017288) | Barringtonia racemosa | 133 and 134 | ||
| 66 | Methyljasmonate | Lasiodiplodia sp. (BL 101) | V. vinifera | 129 |
| 67 | cis-Jasmone | L. theobromae (MAFF no. 306027) | — | 135 |
| 68 | (11R)-(−)-Hydroxyjasmonic acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| 69 | (11S)-(−)-Hydroxyjasmonic acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| L. theobromae (2334) | Helianthus annuus | 119 | ||
| 70 | 8-Hydroxy-jasmonic acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| 71 | 12-Hydroxy-jasmonic acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| L. theobromae (2334) | H. annuus | 119 | ||
| 72 | 3-Oxo-2-(1-hydroxy-2Z-pentenyl)cyclopent-1-yl-butyric acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| 73 | 3-Oxo-2-(4-hydroxy-2Z-pentenyl)cyclopent-1-yl-butyric acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| 74 | Jasmonic acid–glycine | L. theobromae (2334, 1517 and 83) | C. cinensis and wood | 85 |
| 75 | Jasmonic acid–isoleucine | L. theobromae (2334, 1517 and 83) | C. cinensis and wood | 85 |
| 76 | Jasmonic acid–serine | L. theobromae (2334, 1517 and 83) | C. cinensis and wood | 85 |
| 77 | Jasmonic acid–threonine | Lasiodiplodia iranensis (F0619) | Avicennia germinans | 42 |
| L. theobromae (2334 and 1517) | C. cinensis and wood | 85 | ||
| 78 | (+)-7-iso-Jasmonic acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| 79 | Ethyl (+)-7-iso-jasmonate | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| 80 | (+)-9,10-Dihydro-7-iso-jasmonic acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| 81 | 4,5-Didehydro-7-iso-jasmonic acid | L. iranensis (F0619) | A. germinans | 42 |
| L. theobromae (D 7/2) | C. sinensis | 123–125 | ||
| 82 | 11,12-Didehydro-7-iso-jasmonic acid | L. iranensis (F0619) | A. germinans | 42 |
| L. theobromae (D 7/2) | C. sinensis | 123–125 | ||
| 83 | (1S,2S)-[3-Oxo-2-(2Z-pentenyl)-cyclopentyl]butanoic acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| L. theobromae (2334) | H. annuus | 119 | ||
| 84 | (1R,2S)-[3-Oxo-2-(2Z-pentenyl)-cyclopentyl]propanoic acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| 85 | (+)-Cucurbic acid | L. theobromae (D 7/2) | C. sinensis | 123–125 |
| 86 | Lasiojasmonate A | Lasiodiplodia sp. (BL 101) | V. vinifera | 129 |
| 87 | Lasiojasmonate B | Lasiodiplodia sp. (BL 101) | V. vinifera | 129 |
| 88 | Lasiojasmonate C | Lasiodiplodia sp. (BL 101) | V. vinifera | 129 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Lactones and lactols | ||||
| 89 | (−)-Botryodiplodin | L. theobromae | 20 | |
| L. theobromae (LA-SOL3 and LA-SV1) | V. vinifera | 45 and 101 | ||
| L. theobromae | C. nucifera | 136 | ||
| Lasiodiplodia sp. (BL 101) | V. vinifera | 129 | ||
| 90 | 3-epi-Botryodiplodin | L. theobromae (LA-SOL3 and LA-SV1) | V. vinifera | 101 |
| 91 | (3R,4S)-4-Acetyl-3-methyldihydrofuran-2(3H)-one | L. theobromae (LA-SOL3 and LA-SV1) | V. vinifera | 101 |
| 92 | (3S,4S)-4-Acetyl-3-methyldihydrofuran-2(3H)-one | L. theobromae (CAA019 and CBS339.90) | C. nucifera, and human | 46 |
| L. theobromae (LA-SOL3 and LA-SV1) | V. vinifera | 101 | ||
| L. theobromae (PSU-M35) | Garcinia mangostana | 108 | ||
| 93 | (3R,4R)-4-Acetyl-3-methyl-2(3H)-dihydrofuranone | L. theobromae (NSTRU-PN1.4) | Soil | 110 |
| 94 | Nigrosphaerilactone (3S,4R,5R)-4-(hydroxymethyl)-3,5-dimethyldihydrofuran-2(3H)-one | L. theobromae (CAA019 and CBS339.90) | C. nucifera and human | 46 |
| L. theobromae (LA-SOL3 and LA-SV1) | V. vinifera | 101 | ||
| L. mediterranea (B6) | V. vinifera | 131 | ||
| Lasiodiplodia sp. (BL 101) | V. vinifera | 129 | ||
| L. pseudotheobromae (#1048AMSTYEL) | Aegle marmelos | 137 | ||
| L. theobromae (PSU-M35) | G. mangostana | 108 | ||
| L. theobromae (NSTRU-PN1.4) | Soil | 110 | ||
| 95 | Botryosphaerilactone A | L. theobromae (CAA019 and CBS339.90) | C. nucifera and human | 46 |
| Lasiodiplodia sp. (BL 101) | V. vinifera | 129 | ||
| L. theobromae (PSU-M35) | G. mangostana | 108 | ||
| L. theobromae (NSTRU-PN1.4) | Soil | 110 | ||
| 96 | Botryosphaerilactone B | L. theobromae (PSU-M35) | G. mangostana | 108 |
| L. theobromae (NSTRU-PN1.4) | Soil | 110 | ||
| 97 | Botryosphaerilactone C | L. theobromae (PSU-M35) | G. mangostana | 108 |
| L. theobromae (NSTRU-PN1.4) | Soil | 110 | ||
| 98 | Botryosphaerilactone D | L. theobromae (NSTRU-PN1.4) | Soil | 110 |
| 99 | Botryosphaerilactone E | L. theobromae (NSTRU-PN1.4) | Soil | 110 |
| 100 | 16-O-Acetylbotryosphaerilactone A | Lasiodiplodia sp. (BL 101) | V. vinifera | 129 |
| 101 | 16-O-Acetylbotryosphaerilactone C | Lasiodiplodia sp. (BL 101) | V. vinifera | 129 |
| 102 | Lasiolactol A | L. theobromae (LA-SOL3 and LA-SV1) | V. vinifera | 101 |
| L. mediterranea (B6) | V. vinifera | 131 | ||
| 103 | Lasiolactol B | L. theobromae (LA-SOL3 and LA-SV1) | V. vinifera | 101 |
| L. mediterranea (B6) | V. vinifera | 131 | ||
| 104 | (3S,4R)-3-Carboxy-2-methylene-heptan-4-olide | L. theobromae | M. indica | 138 |
| L. venezuelensis (A02EtM) | Astrocaryum sciophilum | 95 | ||
| Lasiodiplodia sp. | Handroanthus impetiginosus | 103 | ||
| 105 | Decumbic acid | L. theobromae | M. indica | 138 |
| L. venezuelensis (A02EtM) | A. sciophilum | 95 | ||
| Lasiodiplodia sp. | H. impetiginosus | 103 | ||
| 106 | Decumbic acid B | L. venezuelensis (A02EtM) | A. sciophilum | 95 |
| 107 | Lasiolactone (R)-(−)-2-octeno-D-lactone | L. theobromae (GK-1) | C. nucifera | 106 |
| L. theobromae (PSU-M114) | G. mangostana | 108 | ||
| 108 | Lasdiplactone | L. pseudotheobromae (#1048AMSTYEL) | Aegle marmelos | 137 |
| 109 | Massoialactone | L. theobromae (GK-1) | C. nucifera | 106 |
| 110 | Tetrahydro-4-hydroxy-6-propylpyran-2-one | L. theobromae (PSU-M114) | G. mangostana | 108 |
| 111 | (4R*,6R*)-4-Hydroxy-6-N-propyl-1-oxacyclo-hexan-2-one | L. theobromae (BPPCA 144) | Aglaia argentea | 55 |
| 112 | (5S,6S)-6-((3′S,4′S,Z)-3′,4′-Dihydroxypent-1-en-1-yl)-5-hydroxy-5,6-dihydro-2H-pyran-2-one | L. venezuelensis | A. sciophilum | 94 |
| 113 | Pantolactone | L. pseudotheobromae (APR5) | A. paniculata | 98 |
| 114 | Monocerin | L. theobromae (AUMC 8903) | D. draco | 43 |
| 115 | Dihydrocumambrin A | L. theobromae (AUMC 8903) | D. draco | 43 |
| 116 | Lasiodione A | Lasiodiplodia sp. (AD-2102) | Artemisia desertorum | 139 |
| 117 | 3-Methyl-3,4-dihydro-1H-isochromene-1,8(7H)-dione | L. theobromae | Peronema canescens | 140 |
| 118 | 3-Hydroxy-4(hydroxy(4-hydroxyphenyl)methyl)-γ-butyrolactone | L. theobromae | P. canescens | 140 |
| 119 | Diplobifuranylone B | L. venezuelensis | A. sciophilum | 94 |
| 120 | 3ξ-(1ξ-Hydroxyethyl)-7-hydroxy-1-isobenzofuranone | L. venezuelensis | A. sciophilum | 94 |
The production of (−)-jasmonic acid by numerous strains of L. theobromae has been optimized, showing promising high yields of this compound in fermentation medium.85,118–122
As reported in Fig. 10, 4,5-didehydro-7-iso-jasmonic acid (81) is the only jasmonate produced by Lasiodiplodia spp. presenting a cyclopentenone ring. In general, jasmonic acid derivatives exhibit modifications on their side chains. For instance, six hydroxylated derivatives on the pentenyl side chain have been reported from L. theobromae.119,123–125
In plants, (−)-jasmonic acid frequently occurs conjugated with sugars and with a variety of amino acids (e.g. isoleucine, leucine, valine, alanine, tyrosine, and phenylalanine). Isoleucine is the principal amino acid linked to (−)-jasmonic acid, while other amino acids rarely occur in conjugated form.126–128 The production of glucose or gentiobiose esters of jasmonic acid by Lasiodiplodia spp. has not yet been reported, while the presence of jasmonic acid–glycine, jasmonic acid–isoleucine, jasmonic acid–serine and jasmonic acid–threonine conjugates (74–77) has been reported in fermentation broths of L. theobromae isolated from Citrus cinensis and wood of Brazilian Amazonia (Table 4).85
Interestingly, although jasmonates are primarily plant metabolites, some of them have not been isolated from plants before but have only been detected as fungal products. This is the case of lasiojasmonates, three (−)-jasmonic acids esterified with lactone/lactol subunits (86–88), isolated from culture filtrates of a pathogenic strain of Lasiodiplodia sp.129
2.9 Lactones and lactols
Lactones and lactols, as reported in Table 4, are a large group of natural polyketides. These compounds are γ- and δ-lactones and lactols.(−)-Botryodiplodin (89) is the first compound belonging to this series isolated from Lasiodiplodia spp. and as reported above, it is also the first secondary metabolite isolated from fungi belonging to this genus. This compound attracted the attention of researchers due to its good antimicrobial activity against several fungal and bacterial strains (Table S1).20 (−)-Botryodiplodin is a hemiacetal and it is reported as epimeric equilibrium at carbon one.92 Furthermore, epi-botryodiplodin (90) is the epimeric form of 89 on C-3 isolated for the first time as a natural product from L. theobromae associated with grapevine.101
Some compounds reported in Fig. 11 are dimeric structures constituted by two subunits of lactones or lactols. Botryosphaerilactones (95–99) are five dimeric γ-lactones isolated from strains of L. theobromae.108,110 Furthermore, the acetyl derivatives of botryosphaerilactones A and C (95 and 97, respectively) were detected from a strain of Lasiodiplodia sp. isolated from grapevine.129 Nigrosphaerilactone (94) is produced by several fungi130 including different Lasiodiplodia spp. (Table 4). This compound has been recently named nigrosphaerilactone, following its discovery as a product of Nigrospora sphaerica.130 For this reason, in many papers it appears with its IUPAC name: (3S,4R,5R)-4-(hydroxymethyl)-3,5-dimethyldihydrofuran-2(3H)-one.
Lasiolactols (102 and 103) are two dimeric γ-lactols identified in cultures of L. mediterranea131 and closely structurally related to botryosphaerilactones A and C (95 and 97, respectively).
2.10 2-(2-Phenylethyl)chromones
2-(2-Phenylethyl)chromones consist of an oxygen-containing heterocycle with a benzoannelated γ-pyrone moiety linked to a phenylethyl residue. These compounds present various substituents, such as hydroxy, methoxy, and chloro groups, on different positions, which endow them with extensive structural variability.141,142 2-(2-Phenylethyl)chromones are key components in agarwood, a resinous material collected from Aquilaria trees and used as incense and in Asian traditional medicines.141 In fact, more than two hundred 2-(2-phenylethyl)chromone derivatives have been isolated and identified from Aquilaria spp. Interestingly, seven known 2-(2-phenylethyl)chromone analogues (121–127) have been isolated from a solid culture of an endophytic strain of L. theobromae from Aquilaria sinensis (Table 5 and Fig. 12).143| No. | Compound | Fungal producer (strain) | Source | Ref. |
|---|---|---|---|---|
| 2-(2-Phenylethyl)chromones | ||||
| 121 | 6-Hydroxy-7-methoxy-2-(2-phenylethyl)chromone | Lasiodiplodia theobromae (A13) | Aquilaria sinensis | 143 |
| 122 | 6,7-Dimethoxy-2-(2-phenylethyl)chromone | L. theobromae (A13) | A. sinensis | 143 |
| 123 | (5S,6R,7S,8R)-2-(2-Phenylethyl)-5,6,7,8-tetrahydrochromone | L. theobromae (A13) | A. sinensis | 143 |
| 124 | 6-Hydroxy-2-(2-phenylethyl)chromone | L. theobromae (A13) | A. sinensis | 143 |
| 125 | 4-Hydroxy-2-(2-phenylethyl)chromone | L. theobromae (A13) | A. sinensis | 143 |
| 126 | 6-Methoxy-2-phenethyl-4H-chromen-4-one | L. theobromae (A13) | A. sinensis | 143 |
| 127 | 6-Methoxy-2-(4-methoxyphenethyl)-4H-chromen-4-one | L. theobromae (A13) | A. sinensis | 143 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Phenyl and phenol derivatives | ||||
| 128 | Phenol | Lasiodiplodia pseudotheobromae (APR5) | Andrographis paniculata | 98 |
| 129 | Tyrosol | L. theobromae (BPPCA 144) | Aglaia argentea | 55 |
| Lasiodiplodia laeliocattleyae (CMM0206) | Vitis vinifera | 104 | ||
| L. theobromae (LA-SOL3 and LA-SV1) | V. vinifera | 45 and 101 | ||
| Lasiodiplodia sp. | Handroanthus impetiginosus | 154 | ||
| Lasiodiplodia euphorbiaceicola (CMM0181) | V. vinifera | 107 | ||
| Lasiodiplodia hormozganensis (CMM0126) | V. vinifera | 107 | ||
| L. theobromae (TBRC 15112) | Achyranthes aspera | 54 | ||
| Lasiodiplodia citricola (ALG111 and ALG81) | Citrus sinensis | 84 | ||
| 130 | 2-Phenylethanol | L. theobromae (BPPCA 144) | A. argentea | 55 |
| L. theobromae (GK-1) | Cocos nucifera | 106 | ||
| L. pseudotheobromae (APR5) | Andrographis paniculata | 98 | ||
| L. citricola (ALG111 and ALG39) | C. sinensis | 84 | ||
| 131 | 2-(4-Hydroxyphenyl)acetic acid | L. theobromae (BPPCA 144) | A. argentea | 55 |
| 132 | 3-Hydroxyphenylacetic acid | L. citricola (ALG111) | C. sinensis | 84 |
| 133 | 4-Hydroxyphenylacetic acid | L. citricola (ALG111) | C. sinensis | 84 |
| L. theobromae (TBRC 15112) | A. aspera | |||
| 134 | 6-Methylsalicylic acid | L. theobromae (PSU-M35) | Garcinia mangostana | 108 |
| 135 | Scytalone | L. theobromae (CAA019 and CBS339.90) | C. nucifera, and human | 46 |
| 136 | 3-Hydroxybenzoic acid | L. citricola (ALG111) | C. sinensis | 84 |
| 137 | 4-Hydroxybenzoic acid | L. theobromae | Psidium guajava | 155 |
| L. hormozganensis (CMM0126) | V. vinifera | 107 | ||
| L. citricola (ALG111) | C. sinensis | 84 | ||
| Lasiodiplodia sp. | H. impetiginosus | 103 | ||
| 138 | 3ξ-(1ξ-Hydroxyethyl)-7-hydroxy-1-isobenzofuranone | Lasiodiplodia venezuelensis (A02EtM) | Astrocaryum sciophilum | 95 |
| 139 | t-Butylhydroquinone | L. pseudotheobromae (APR5) | A. paniculata | 98 |
| 140 | p-Cresol | L. pseudotheobromae (APR5) | A. paniculata | 98 |
| 141 | N-(4-Hydroxyphenyl)acetamide | L. theobromae (TBRC 15112) | A. aspera | 54 |
| 142 | 4-Hydroxyphenylacetamide | Lasiodiplodia sp. | H. impetiginosus | 103 |
| 143 | Phenylacetic acid | L. citricola (ALG111) | C. sinensis | 84 |
| 144 | Cinnamic acid | L. citricola (ALG111) | C. sinensis | 84 |
| 145 | Protocatechuic acid | L. citricola (ALG111) | C. sinensis | 84 |
| L. theobromae | Musa paradisiaca | 40 | ||
| 146 | Syringic acid | L. citricola (ALG111) | C. sinensis | 84 |
| 147 | Vanillic acid | L. citricola (ALG111) | C. sinensis | 84 |
| 148 | O-Methyl alboatrin | L. theobromae (NSTRU-PN1.4) | Soil | 110 |
| 149 | Salicylic acid | L. theobromae (2334, 1517 and 83) | C. cinensis and wood | 85 |
Given the economic importance of agarwood constituents, the production of Aquilaria metabolites by an endophytic strain represents a valuable discovery.
2.11 Phenyl and phenol derivatives
The list of phenyl and phenol derivatives produced by Lasiodiplodia spp. includes some well-known natural compounds. Phenyl and phenol derivatives are characterized by having at least one aromatic ring which, in the case of phenol derivatives, is functionalized to one or more hydroxyl groups. These compounds range from simple, low molecular-weight compounds to larger, more complex structures.144,145 These compounds are widespread throughout the plant kingdom,146,147 but several reports describe their occurrence as fungal products. As can be seen in Table 5 and Fig. 13, Lasiodiplodia spp. produce mostly phenol derivatives with simple structures that include a single aromatic group linked to diverse substituents. Due to its wide spectrum of bioactivities, tyrosol (129) holds a place of particular relevance among phenolic compounds. From a structural viewpoint, 129 is characterized by a hydroxyethyl chain at the para position of the phenol ring. It is the most abundant phenolic compound in olive oils and wine and described as a health-promoting compound due to its beneficial properties.148–151 Several species of Lasiodiplodia biosynthesize tyrosol and some of them are fungi associated with grapevine.45,101,104,107 From a strain of L. theobromae isolated from Achyranthes aspera,54 tyrosol has been isolated together with two other phenol derivatives, 4-hydroxyphenylacetic acid (133) and N-(4-hydroxyphenyl)acetamide (141). 4-Hydroxyphenylacetic acid has been wrongly reported by the authors as N-(4-hydroxyphenyl)acetic acid, hence its correct structure and name have been added in Table 5 and Fig. 13.A strain of L. citricola (previously known as Lasiodiplodia mitidjana) isolated from C. sinensis turned out to be a good producer of phenol compounds, including protocatechuic acid, syringic acid and vanillic acid (145–147).84 Lasiodiplodia citricola also produces phenylacetic acid (143), a phenyl derivative that has been shown to be an active auxin.84 However, its effect is much weaker than the effect of indole-3-acetic acid, the other auxin produced by Lasiodiplodia spp.152,153
2.12 Preussomerins
Preussomerins are natural products having an epoxynaphthoquinone structure linked to a bisnaphthoquinone spiroketal moiety as a common structural unit. Based on their structural features, these natural products can be divided into three subclasses: two-oxygen-bridge-type, three-oxygen-bridge-type and those with two oxygen bridges and one C–C bridge.156 Nine preussomerins with two oxygen bridges (162, 163, and 165–171) and fifteen preussomerins with three oxygens bridges (150–161, 164, 172 and 173) have been isolated from this fungal genus (Table 6 and Fig. 14). Most of them are well-known fungal secondary metabolites, including the first preussomerin, named preussomerin A (152), identified in 1990 from the coprophilous fungus Preussia isomera.157 Two new preussomerins derivatives, i.e., lasiodiplodiapyrones A and B (172 and 173, respectively), belonging to the subclass with three oxygen bridges, have been isolated from L. pseudotheobromae. Compared to the common preussomerins, these compounds possess an unexpected 6-methyl-4H-furo[3,2-c]pyran-4-one moiety, a highly functionalized conjoint and a complicated polycyclic ring system. These metabolites were isolated and characterized via spectroscopic techniques, along with two known congeners, preussomerin B (153) and preussomerin SA1 (164).158 A novel two-oxygen-bridge-type preussomerin named mitidjospirone (162) was isolated and identified from the mycelial extract of L. citricola, together with several known compounds including palmarumycin JC1 (163). Mitidjospirone owes its trivial name to its fungal producer, formerly known as L. mitidjana.84 Chloropreussomerins A and B (150 and 151, respectively) are two new chlorinated preussomerins isolated from a mangrove endophytic strain of L. theobromae. Other known preussomerins have been isolated from the same strain and identified as preussomerin A (152), preussomerin C (154), preussomerin D (155), preussomerin F (156), preussomerin G (157), preussomerin H (158), preussomerin K (159), and Ymf 1029 E (161).159| No. | Compound | Fungal producer (strain) | Source | Ref. |
|---|---|---|---|---|
| Preussomerins | ||||
| 150 | Chloropreussomerin A | L. theobromae (ZJ-HQ1) | Acanthus ilicifolius | 159 |
| 151 | Chloropreussomerin B | L. theobromae (ZJ-HQ1) | A. ilicifolius | 159 |
| 152 | Preussomerin A | L. theobromae (ZJ-HQ1) | A. ilicifolius | 159 |
| 153 | Preussomerin B | L. pseudotheobromae (414-JZ-40) | Soil | 158 |
| 154 | Preussomerin C | L. theobromae (ZJ-HQ1) | A. ilicifolius | 159 |
| 155 | Preussomerin D | L. theobromae (ZJ-HQ1) | A. ilicifolius | 159 |
| 156 | Preussomerin F | L. theobromae (ZJ-HQ1) | A. ilicifolius | 159 |
| 157 | Preussomerin G | L. theobromae (ZJ-HQ1) | A. ilicifolius | 159 |
| 158 | Preussomerin H | L. theobromae (ZJ-HQ1) | A. ilicifolius | 159 |
| 159 | Preussomerin K | L. theobromae (ZJ-HQ1) | A. ilicifolius | 159 |
| 160 | Preussomerin M | L. theobromae (ZJ-HQ1) | A. ilicifolius | 159 |
| 161 | Ymf 1029 E | L. theobromae (ZJ-HQ1) | A. ilicifolius | 159 |
| 162 | Mitidjospirone | L. citricola (ALG111) | C. sinensis | 84 |
| 163 | Palmarumycin JC1 | L. citricola (ALG111) | C. sinensis | 84 |
| 164 | Preussomerin SA1 | L. pseudotheobromae (414-JZ-40) | Soil | 158 |
| 165 | (+)-(R)-CJ-12372 | Lasiodiplodia venezuelensis (A02EtM) | Astrocaryum sciophilum | 95 |
| 166 | (+)-(R)-Palmarumycin EG1 | L. venezuelensis (A02EtM) | A. sciophilum | 95 |
| 167 | (R)-4-Methoxy-3,4-dihydro-2H-spiro[naphthalene-1,2′-naphtho[1,8-de][1,3]dioxin]-6-ol | L. venezuelensis (A02EtM) | A. sciophilum | 95 |
| 168 | Palmarumycin LP1 | L. pseudotheobromae (XSZ-3) | Camptotheca acuminata | 160 |
| 169 | Ascochytatin | L. pseudotheobromae (XSZ-3) | C. acuminata | 160 |
| 170 | Sch 50676 | L. pseudotheobromae (XSZ-3) | C. acuminata | 160 |
| 171 | Cladospirone B | L. theobromae (MJ2211) | Vitex pinnata | 105 |
| L. pseudotheobromae (XSZ-3) | C. acuminata | 160 | ||
| 172 | Lasiodiplodiapyrone A | L. pseudotheobromae (414-JZ-40) | Soil | 158 |
| 173 | Lasiodiplodiapyrone B | L. pseudotheobromae (414-JZ-40) | Soil | 158 |
2.13 Resorcinol and β-resorcylic acid derivatives
Resorcinol derivatives possess a 1,3-dihydroxybenzene core, which is a scaffold widespread in nature.161–164 With the exception of 3-(2-ethoxycarbonyl-3,5-dihydroxyphenyl)propionic acid,165 novel resorcinol derivatives have been found in cultures of Lasiodiplodia spp. (Table 7 and Fig. 15).154,166| No. | Compound | Fungal producer (strain) | Source | Ref. |
|---|---|---|---|---|
| Resorcinol and β-resorcylic acid derivatives | ||||
| 174 | Adeninealkylresorcinol | Lasiodiplodia sp. | Handroanthus impetiginosus | 154 |
| 175 | 3-O-Methyl-5-(8-methoxyl-6-oxononyl)-resorcinol | Lasiodiplodia sp. | H. impetiginosus | 154 |
| 176 | 3-O-Methyl-5-[(7E)-6-oxo-7-nonenyl]-resorcinol | Lasiodiplodia sp. | H. impetiginosus | 154 |
| 177 | (R)-Lasiodiplodin | Lasiodiplodia theobromae (IFO 31059) | — | 174 and 175 |
| L. theobromae | — | 100 | ||
| L. theobromae (CSS01s) | V. vinifera | 102 | ||
| L. pseudotheobromae (J-10) | Sarcandra glabra | 176 | ||
| L. theobromae (CAA019) | C. nucifera | 46 | ||
| Lasiodiplodia sp. (318#) | Excoecaria agallocha | 168 and 169 | ||
| L. citricola (ALG111 and ALG39) | C. sinensis | 84 | ||
| 178 | (S)-Lasiodiplodin | L. theobromae (PSU-M35) | Garcinia mangostana | 108 |
| L. theobromae (TBRC 15112) | Achyranthes aspera | 54 | ||
| 179 | (3R,4S)-4-Hydroxy-lasiodiplodin | L. theobromae (IFO 31059) | — | 177 |
| 180 | (3R,5R)-5-Hydroxy-lasiodiplodin | L. theobromae (IFO 31059) | — | 177 |
| L. mediterranea (B6) | V. vinifera | 131 | ||
| L. theobromae | S. tuberosum | 178 | ||
| L. theobromae (PSU-M35) | G. mangostana | 108 | ||
| Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 | ||
| 181 | (3R,5S)-5-Hydroxy-lasiodiplodin | L. theobromae (IFO 31059) | — | 174, 175, 177 and 178 |
| L. theobromae (PSU-M35) | G. mangostana | 108 | ||
| Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 | ||
| 182 | (3R,6S)-6-Hydroxy-lasiodiplodin | L. theobromae (Shimokita 2) | M. indica | 179 |
| 183 | (12E,15R)-5-Hydroxy-3-methoxy-16-methyl-8,9,10,11,14,15-hexahydro-1H-benzo[c][1]oxacyclodocecin-1-one | Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 |
| 184 | Botryosphaeriodiplodin | L. mediterranea (B6) | V. vinifera | 131 |
| L. theobromae (PSU-M35) | G. mangostana | 108 | ||
| 185 | (3R)-De-O-methyl-lasiodiplodin | L. theobromae | Pyrenula bahiana | 180 |
| L. theobromae (3PR3) | Mapania kurzii | 181 | ||
| L. theobromae | — | 100 | ||
| Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 | ||
| L. theobromae (MJ2211) | Vitex pinnata | 105 | ||
| L. theobromae (M4.2-2) | Mangrove sediment | 57 | ||
| 186 | 4-Hydroxy-de-O-methyl-lasiodiplodin | L. theobromae (3PR3) | M. kurzii | 181 |
| 187 | (3R,5R)-5-Hydroxy-de-O-methyllasiodiplodin | L. theobromae | — | 177 |
| 188 | (6R)-6-Hydroxy-de-O-methyllasiodiplodin | L. theobromae (3PR3) | M. kurzii | 181 |
| L. theobromae (IFO 31059) | — | 177 | ||
| 189 | 5-Oxo-lasiodiplodin | L. theobromae (IFO 31059) | — | 177 and 178 |
| 190 | (3R)-7-Oxo-lasiodiplodin | Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 |
| 191 | (3S)-7-Oxo-de-O-methyllasiodiplodin | L. theobromae (GI-1005) | Rhizophora mucronata | 165 |
| 192 | (3R,5E)-5-Etheno-lasiodiplodin | Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 |
| 193 | (3S,7R)-9-Etheno-7-hydroxy-13-O-methyl-de-O-methyllasiodiplodin | L. theobromae (GI-1005) | R. mucronata | 165 |
| 194 | (E)-9-Etheno-de-O-methyl-lasiodiplodin | L. theobromae (3PR3) | M. kurzii | 181 |
| 195 | (R)-14-Methoxy-3-methyl-3,4,5,6,7,8,9,10-octahydro-1H-benzo[c][1]oxacyclododecine-1,11,12-trione | Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 |
| 196 | Lasiodiplactone A | L. theobromae (ZJ-HQ1) | Acanthus ilicifolius | 170 |
| 197 | epi-8,9-Dihydrogreensporone C | Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 |
| 198 | (−)-(R)-Nordinone | L. theobromae (M4.2-2) | Mangrove sediment | 57 |
| 199 | (R)-Zearalenone | Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 |
| 200 | 2,4-Dihydroxy-6-nonylbenzoate | Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 |
| 201 | Ethyl (S)-2,4-dihydroxy-6-(8-hydroxynonyl)benzoate | L. theobromae (GI-1005) | R. mucronata | 165 |
| L. theobromae (GC-22) | Xylocarpus granatum | 171 | ||
| Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 | ||
| 202 | Ethyl 2,4-dihydroxy-6-(8-hydroxyheptyl)benzoate | L. theobromae (GI-1005) | R. mucronata | 165 |
| L. theobromae (GC-22) | X. granatum | 171 | ||
| 203 | Ethyl 2,4-dihydroxy-6-(4-methoxycarbonylbutyl)benzoate | L. theobromae (GC-22) | X. granatum | 171 |
| 204 | Isobutyl (S)-2,4-dihydroxy-6-(8-hydroxynonyl)benzoate | L. theobromae (GC-22) | X. granatum | 171 |
| Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 | ||
| 205 | Ethyl 2,4-dihydroxy-6-(8-oxononyl)benzoate | L. theobromae (GI-1005) | R. mucronata | 165 |
| Lasiodiplodia sp. (318#) | E. agallocha | 168 and 169 | ||
| L. theobromae (GC-22) | X. granatum | 171 | ||
| 206 | (3S,7R)-7-Hydroxy-13-O-methyl-de-O-methyllasiodiplodin | L. theobromae (GI-1005) | R. mucronata | 165 |
| 207 | (15S)-De-O-methyllasiodiplodin | L. theobromae (GC-22) | X. granatum | 171 |
| L. brasiliensis (WS-TS-A1) | Cannabis sativa | 182 | ||
| 208 | (13S,15S)-13-Hydroxy-de-O-methyllasiodiplodin | L. theobromae (GC-22) | X. granatum | 171 |
| 209 | (14S,15S)-14-Hydroxy-de-O-methyllasiodiplodin | L. theobromae (GC-22) | X. granatum | 171 |
| 210 | (13R,14S,15S)-13,14-Dihydroxy-de-O-methyllasiodiplodin | L. theobromae (GC-22) | X. granatum | 171 |
| 211 | (E)-9-Etheno-de-O-methyl-lasiodiplodin | L. theobromae (3PR3) | M. kurzii | 181 |
| 212 | 6-Oxo-de-O-methyl-lasiodiplodin | L. theobromae (3PR3) | M. kurzii | 181 |
| 213 | Ethyl (6′R)-2,4-hydroxy-6-(6′-hydroxyheptyl)-benzoate | L. theobromae (IFO 31059) | — | 166 |
| 214 | Isobutyl (6′R)-2,4-hydroxy-6-(6′-hydroxyheptyl)-benzoate | L. theobromae (IFO 31059) | — | 166 |
| 215 | 3-(2-Ethoxycarbonyl-3,5-dihydroxyphenyl)propionic acid | L. theobromae (GC-22) | X. granatum | 171 |
| L. theobromae (GI-1005) | R. mucronata | 165 | ||
| 216 | (3S)-3-[(R)-8-Hydroxynonyl]-6-hydroxy-8-methoxy-3,4-dihydroisochroman-1-one | Lasiodiplodia sp. | H. impetiginosus | 154 |
Three novel alkylresorcinols (174–176) have been isolated from an endophytic strain of Lasiodiplodia sp. obtained from a traditional Chinese medical plant Houttuynia cordata. Alkylresorcinols, also known as resorcinolic lipids, are characterized by an alkyl or an alkenyl chain at 5-position of the aromatic ring linked to the 1,3-dihydroxybenzene skeleton.161 Among the three new compounds identified, adeninealkylresorcinol (174) is an unusual alkylresorcinol with an adenine-alkyl resorcinol conjoined skeleton.154
β-Resorcylic acid derivatives are members of a unique fungal metabolites class characterized by a 12- and 14-membered macrolactone ring linked to a 2,4-dihydroxybenzoic acid residue. Since their first identification, over 200 β-resorcylic acid derivatives have been reported from different genera of fungi, including Aigialus, Curvularia, Lasiodiplodia, Penicillium, and Pochonia.167
Lasiodiplodia theobromae was found to be a remarkable producer of these compounds, which are often named lasiodiplodins (Table 7 and Fig. 15). (R)-Lasiodiplodin (177) and (R)-de-O-methyllasiodiplodin (185) are the first members of the β-resorcylic acid derivatives class to be found in the cultural filtrate of Lasiodiplodia spp. A series of compounds belonging to this class of natural products has been isolated from mangrove endophytic strains of Lasiodiplodia spp. (Table 7).168–171
Among them, several novel lasiodiplodins (180, 181, 183, 185, 190, 192, 195, 197, 199–201, 204, and 205) have been identified from Lasiodiplodia sp. 318# isolated from Excoecaria agallocha.168,169
A mangrove endophytic strain of L. theobromae produced an unprecedented β-resorcylic acid derivative named lasiodiplactone A (196) possessing a pyran ring and a furan ring.170
A strain of L. theobromae isolated from the inner tissue of a dead branch of the mangrove plant Xylocarpus granatum produced ten β-resorcylic acid derivatives, including five lasiodiplodins, in which their macrocyclic ring is affected by modifications that cause the opening of the ring and diverse substituents in the ortho to the carboxylic group (i.e. hydroxyheptyl or hydroxynonyl moieties).171
Due to their abundant production by Lasiodiplodia spp., the employment of these compounds for fungal chemotaxonomy has been proposed.92 In fact, secondary metabolites could be very useful chemotaxonomic markers and have already been used successfully in large genera such as Alternaria, Aspergillus, Fusarium, Penicillium, and Xylaria.172 In this respect, the exclusive production of lasiodiplodins by the unidentified endophytic fungus ZZF3 suggested that this isolate possibly belongs to the Lasiodiplodia genus,173 which is why this strain is included in the present review.
2.14 Terpenoids
Terpenoids are the most abundant and structurally diverse plant secondary metabolites derived from a five-carbon isoprene unit.183–185 Steroids are complex four-ring organic compounds obtained from terpenoid precursors, which play many roles and functions in microorganisms and organisms.186,187 The well-known sterols ergosterol, ergosterol peroxide and stigmasterol (217–219, respectively) have been isolated from the endophytic fungus L. theobromae derived from the root of Aglaia argentea (Table 8).188 Although this is the first report on the isolation of these steroids from species of Lasiodiplodia, this detection is not surprising because ergosterol represents the most abundant sterol in fungal cell membranes and it is critical for defining membrane fluidity and regulating cellular processes.189| No. | Compound | Fungal producer (strain) | Source | Ref. |
|---|---|---|---|---|
| Terpenoids | ||||
| 217 | Ergosterol | Lasiodiplodia theobromae (#009) | Psidium guajava | 111 |
| L. theobromae (Arg-8) | Aglaia argentea | 188 | ||
| 218 | Ergosterol peroxide | L. theobromae (Arg-8) | A. argentea | 188 |
| 219 | Stigmasterol | L. theobromae (Arg-8) | A. argentea | 188 |
| 220 | (22E,24R)-4α,5α-Epoxyergosta-9α,14β-dihydroxy-7,22-diene-3,6-dione | Lasiodiplodia pseudotheobromae (414-JZ-40) | Soil | 158 |
| 221 | (22E,24R)-4α,5α-Epoxyergosta-9α,14α-dihydroxy-7,22-diene-3,6-dione | L. pseudotheobromae (414-JZ-40) | Soil | 158 |
| 222 | 12α-Hydroxyergosta-7,22,24(28)-triene-3-one | L. pseudotheobromae (414-JZ-40) | Soil | 158 |
| 223 | 3β,12α-Dihydroxyergosta-7,24(28)-diene | L. pseudotheobromae (414-JZ-40) | Soil | 158 |
| 224 | (22E,24R)-9α,14β-Dihydroxyergosta-4,7,22-triene-3,6-dione | L. pseudotheobromae (414-JZ-40) | Soil | 158 |
| 225 | 3β,14β-Dihydroxy-6-oxo-A-nor-ergosta-7,22-diene-4-oic acid δ-lactone | L. pseudotheobromae (414-LY-1) | Soil | 190 and 191 |
| 226 | 2β,3β,12α-Trihydroxyergosta-7,22-diene | L. pseudotheobromae (414-LY-1) | Soil | 190 and 191 |
| 227 | 2β,3β,12α-Trihydroxyergosta-7,24(28)-diene | L. pseudotheobromae (414-LY-1) | Soil | 190 and 191 |
| 228 | 9α,11α,15α-Trihydroxyergosta-4,6,8(14),22-tetraen-3-one | L. pseudotheobromae (414-LY-1) | Soil | 190 and 191 |
| 229 | Cholestanol glucoside | L. theobromae | Saraca asoca | 194 and 195 |
| 230 | Gibberellic acid (GA3) | Lasiodiplodia theobromae (2334, 1517 and 83) | Citrus cinensis and wood | 85 |
| 231 | Abscisic acid | L. theobromae (2334, 1517 and 83) | C. cinensis and wood | 85 |
| Lasiodiplodia iranensis (F0619) | Avicennia germinans | 42 | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||||
| Miscellaneous | ||||
| 232 | Glycerol | Lasiodiplodia citricola (ALG11, ALG39 and ALG81) | Citrus sinensis | 84 |
| Lasiodiplodia avicenniae (P2P4) | Avicennia alba | 196 | ||
| 233 | Eugenol | L. citricola (ALG11) | C. sinensis | 84 |
| 234 | Colletopeptide B | L. theobromae (BPPCA 144) | A. argentea | 55 |
| 235 | Taxol | L. theobromae | Morinda citrifolia | 197 and 198 |
| L. theobromae (BT 115) | Taxus baccata | 199 | ||
| 236 | 2,4,6-Trimethyloct-2-enoic acid, 1,2,6,8a-tetrahydro-7-hydroxy-1,8a-dimethyl-6-oxo-2-naphtalenyl ester | L. theobromae (#009) | Psidium guajava | 111 |
| 237 | Aranorosinol B | L. theobromae | P. guajava | 155 |
| 238 | Lasiodiplin | L. theobromae (TBRC 15112) | Achyranthes aspera | 54 |
| 239 | 5-Hydroxymethyl-2-furancarboxylic acid | Lasiodiplodia venezuelensis | Astrocaryum sciophilum | 94 |
| 240 | (Z)-3-((2R,3R,6R)-3-Hydroxy-6-((R)-1-hydroxyethyl)-3,6-dihydro-2H-pyran-2-yl)acrylamide | L. venezuelensis | A. sciophilum | 94 |
| 241 | (2Z,4Z,8E)-6,7-Dihydroxydeca-2,4,8-trienoic acid | L. venezuelensis | A. sciophilum | 94 |
| 242 | Uridine | L. venezuelensis | A. sciophilum | 94 |
| 243 | Norharman | L. theobromae (AUMC 8903) | Dracaena draco | 43 |
| 244 | Bergapten | L. theobromae (AUMC 8903) | D. draco | 43 |
| 245 | Meranzin | L. theobromae (AUMC 8903) | D. draco | 43 |
| 246 | Formyl indanone | L. theobromae (AUMC 8903) | D. draco | 43 |
| 247 | Halaminol A | L. theobromae (AUMC 8903) | D. draco | 43 |
| 248 | Palmitoleamide | L. theobromae (AUMC 8903) | D. draco | 43 |
| 249 | Palmitic amide | L. theobromae (AUMC 8903) | D. draco | 43 |
| 250 | Capsi amide | L. theobromae (AUMC 8903) | D. draco | 43 |
| 251 | Lasiodione B | Lasiodiplodia sp. (AD-2102) | Artemisia desertorum | 139 |
| 252 | Lasdiploic acid | L. pseudotheobromae (#1048AMSTYEL) | Aegle marmelos | 137 |
| 253 | Botryosphaeridione | L. theobromae (PSU-M35) | Garcinia mangostana | 108 |
| L. theobromae (NSTRU-PN1.4) | Soil | 110 | ||
| 254 | Botryosphaerihydrofuran | L. theobromae (PSU-M35) | G. mangostana | 108 |
| 255 | Botryosphaerinone | L. theobromae (PSU-M35) | G. mangostana | 108 |
| 256 | Succinic acid | L. theobromae (LA-SOL3 and LA-SV1) | Vitis vinifera | 45 |
| L. citricola (ALG111 and ALG81) | C. sinensis | 84 | ||
| 257 | (Z)-3-((2R,3R,6R)-3-Hydroxy-6-((R)-1-hydroxyethyl)-3,6-dihydro-2H-pyran-2-yl)acrylamide | L. venezuelensis (A02EtM) | A. sciophilum | 95 |
| 258 | Aconitate B | L. venezuelensis (A02EtM) | A. sciophilum | 95 |
| 259 | (3R,4R,Z)-4-Hydroxy-1-((2S,3S)-3-hydroxy-6-oxo-3,6-dihydro-2H-pyran-2-yl)pent-1-en-3-yl acetate | L. venezuelensis (A02EtM) | A. sciophilum | 95 |
| 260 | (2Z,6Z)-4,5,8,9-Tetrahydroxydeca-2,6-dienamide | L. venezuelensis (A02EtM) | A. sciophilum | 95 |
| 261 | (2Z,4Z,8E)-6,7-Dihydroxydeca-2,4,8-trienoic acid | L. venezuelensis (A02EtM) | A. sciophilum | 95 |
| 262 | (2R)-Butylitaconic acid | L. venezuelensis (A02EtM) | A. sciophilum | 95 |
| 263 | Erythritol | L. pseudotheobromae (APR5) | Andrographis paniculata | 98 |
| 264 | Niacin | L. pseudotheobromae (APR5) | A. paniculata | 98 |
| 265 | Uracil | L. theobromae (TBRC 15112) | A. aspera | 54 |
| 266 | 1,8-Dihydroxyantraquinone | L. citricola (ALG111) | C. sinensis | 84 |
| 267 | Vitamin B6 (pyridoxine) | L. citricola (ALG111, ALG39, ALG81 and ALG34) | C. sinensis | 84 |
| 268 | 1,3-Butanediol | L. avicenniae (P2P4) | A. alba | 196 |
| 269 | 2,3-Butanediol | L. avicenniae (P2P4) | A. alba | 196 |
| 270 | 4,5,6-Trimethyl-2(1H)-pyrimidinone | L. theobromae (NSTRU-PN1.4) | Soil | 110 |
| 271 | L-Isoleucinamide | L. theobromae (NSTRU-PN1.4) | Soil | 110 |
| 272 | Zeatin | L. theobromae (2334, 1517 and 83) | C. cinensis and wood | 85 |
| 273 | Zeatin riboside | L. theobromae (2334, 1517 and 83) | C. cinensis and wood | 85 |
Eight new ergostene-type steroids (220–223 and 225–228), along with a known congener (224), have been isolated and identified as products of L. pseudotheobromae from the soil of Hainan wetland park. Their structures have been elucidated based on spectroscopic methods and single crystal X-ray diffraction analysis (Table 8 and Fig. 16).158,190,191
HPLC-ESI-MS/MS investigations of the metabolite profiles of L. theobromae strains, isolated from Cuban Citrus cinensis and from the wood of Brazilian Amazonia,85 revealed two other phytohormones belonging to the class of terpenoids: gibberellic acid and abscisic acid (230 and 231, respectively). These compounds are commonly produced by plants, influencing their growth and development, but are quite rare as fungal products.192 Several phytohormones have been simultaneously identified in the fermentation broth of strains of Lasiodiplodia spp. Some of them (e.g., indole acetic acid, jasmonic acid and its amino acid conjugates) have been discussed in the sections dedicated to jasmonates and indoles. Little is known about their role and importance beyond plant systems. Plant-associated fungi appear to be more likely to produce phytohormones to alter their flow in host plants.193 The detection of some phytohormones in cultures of L. theobromae85 and L. iranensis42 supports this view considering that these strains were isolated from plants.
2.15 Miscellaneous
This category includes products of Lasiodiplodia spp. that have no structural affinity with previous classes. As can be seen in Table 8 and Fig. 17, the members of this group present heterogenous structures ranging from simple compounds, e.g., glycerol (232) and 2,3-butanediol (269), to very complex products, e.g. colletopeptide B (234) and taxol (235). Some compounds listed in this section (i.e., glycerol, succinic acid, and uracil) are primary metabolites and, for this reason, are commonly found in natural sources. However, Table 8 presents metabolites exclusively produced by Lasiodiplodia spp. This is the case of a new dihydronaphthalene-2,6-dione derivative named botryosphaeridione, which is produced by strains of L. theobromae.108,1103 Biological activities of Lasiodiplodia spp. secondary metabolites
Secondary metabolites are important mediators of biological interactions and represent promising sources of drugs. Given their relevance in many fields of research, the biological properties of secondary metabolites have been investigated in most papers on the metabolism of Lasiodiplodia spp. (Table S1).Since some of these compounds are well-known from various natural sources and have been extensively studied in dedicated articles, the present section focuses exclusively on bioactivities documented for metabolites isolated from Lasiodiplodia spp., including antimicrobial, cytotoxic, immunomodulatory, phytotoxic (Fig. 18).
 | ||
| Fig. 18 Main bioactivities of the Lasiodiplodia spp. natural products. Metabolites in this figure are indicated by numerical codes. | ||
Several Lasiodiplodia spp. are associated with plant diseases, with the possible involvement of toxic secondary metabolites in various symptoms. In fact, the association of Lasiodiplodia spp. with economically and ecologically important plants has encouraged researchers to conduct phytotoxicity tests on fungal metabolites to better understand their role in symptom expression. Experimental data (Table S1) from phytotoxicity assays have confirmed that some secondary metabolites produced by pathogenic strains of Lasiodiplodia spp. can be considered phytotoxins harmful to plants even at low concentrations and may be involved in the infection mechanisms. For instance, Lasiodiplodia spp. are known pathogens of grapevine, causing aggressive diseases, such as canker and dieback, which symptoms, particularly the foliar ones, can be attributed to the fungal production of toxic secondary metabolites.101,200 In fact, metabolites (i.e., (−)-jasmonic acid (65), nigrosphaerilactone (94), lasiolactols A and B (102 and 103, respectively), and botryosphaeriodiplodin (184)) from a grapevine strain of L. mediterranea showed moderate activity in tests conducted on grapevine cv. Inzolia leaves (Table S1). The phytotoxicity of these compounds increased with increasing concentrations; moreover, 65 was the most active compound (biggest necrotic spots on detached grapevine leaves).131 However, some metabolites exert phytotoxic effects not exclusively on their hosts but also on a wide range of plants. This is exemplified by (−)-mellein (58), which exhibits toxicity toward several plants, including Bromus sp., Cynodon dactylon, Loietto perenne, Setaria italica and members of the Valerianaceae family (Table S1). When tested with a leaf puncture assay on its host plant (Tridax procumbus), (−)-mellein induced necrotic circular lesions within 2 days, closely resembling those caused by the pathogen (L. pseudotheobromae). In contrast, assays on cultivated species showed no effect on Solanaceous species (red pepper and potato), Cucurbitaceae (melon and cucumber), and Leguminosae (cowpea), whereas severe necrosis was observed on monocotyledon Poaceae weeds. These results indicated that (−)-mellein exhibits selective bioherbicidal activity, with a stronger impact on monocotyledons than dicotyledons.109 Another important phytotoxic metabolite of Lasiodiplodia spp. is (−)-jasmonic acid (65), which demonstrated significant phytotoxicity against several plants (Table S1). In particular, this phytotoxin induced the development of brown necrotic lesions on leaves of Rosa sp.,132 Quercus suber,129 Vitis vinifera129 and Solanum lycopersicum.46 As reported in Section 2.8, (−)-jasmonic acid is primarily a phytohormone and its fungal production could be important to manipulate the host physiology for survival, colonization, and nutrient acquisition. In addition to (−)-jasmonic acid, Lasiodiplodia spp. produce other compounds well-known as phytohormones, e.g., salicylic acid (149), gibberellic acid (230) and abscisic acid (231). Therefore, phytotoxic metabolites are not the only compounds that might be involved in the dynamic plant–fungus interactions. In fact, it was also observed that several fatty acid esters produced by L. theobromae showed an effect of plant growth regulation in the model plant Nicotiana tabacum.81,82 In addition, theobroxide (1), its related compounds (4–6) and some β-resorcylic acid derivatives (179–182) demonstrated the ability to induce micro-tuber formation in potatoes (Table S1).29,31 These results proved that compounds produced by Lasiodiplodia spp. could mimic or interact with plant metabolites influencing developmental processes.
In recent years, research has focused on climate change, observing that the phytotoxin-induced pathogenicity of Lasiodiplodia spp. responds to environmental shifts altering the microbial pathogen–host interactions. The adaptability of these pathogens to new environmental conditions has raised concerns in view of a possible dramatic scenario characterized by new host colonization.16 In fact, it was reported that higher temperatures and drought conditions are factors that promote the expansion of the host range and intensify the threat to agriculture and forestry.5,45,46,101,201 We should not forget that Lasiodiplodia spp., especially L. theobromae, are recognized as opportunistic human pathogens. Consequently, the expansion of the host range and the increasing severity of infections pose a critical cross-kingdom risk to global health.
The adaptability of Lasiodiplodia spp. to their hosts is linked to the capacity to produce various metabolites under differing environmental conditions. Although the previously described phytotoxicity of these compounds undoubtedly plays a primary role during plant infection, the broad host range suggests that further bioactivities of Lasiodiplodia spp. metabolites should also be considered because these are not simple by-products, but tools employed as a key adaptive strategy. Indeed, the antimicrobial activity of Lasiodiplodia spp. secondary metabolites is particularly important for host adaptability because the fungus can use these compounds as weapons to compete with other microorganisms during host infection. Table S1 shows that several secondary metabolites have been evaluated for their antimicrobial effects. Among them, (−)-mellein (58) and (−)-botryodiplodin (89) exhibit valuable antimicrobial activities, highlighting their potential as bioactive lead compounds.20,93 (−)-Mellein produced by L. theobromae isolated from the medicinal plant Syzygium cumini showed potent anti-Xanthomonas activity, with MIC values ranging from 1.9 to 62.5 µg mL−1 against 11 Xanthomonas strains, along with broad-spectrum antibacterial and antifungal effects with the MIC of 7.8–31.25 µg mL−1 and 1.9–31.25 µg mL−1, respectively (Table S1). Molecular docking studies further supported its activity, revealing favourable binding interactions with proteins involved in Xanthomonas sp. pathogenicity. In addition, in silico absorption, distribution, metabolism, and excretion (ADME) studies indicated that this 3,4-dihydroisocoumarin possesses suitable oral bioavailability, balanced lipophilicity, good water solubility, favourable pharmacokinetics and overall drug-likeness profiles. (−)-Mellein complies with Lipinski's rule of five with zero violations and shows high predicted gastrointestinal absorption and blood–brain barrier permeation properties. Thus, from a medicinal chemistry perspective, no structural alerts were detected, reinforcing the potential of (−)-mellein as a promising drug candidate.93 Even (−)-botryodiplodin displayed broad-spectrum antimicrobial properties, with well-documented inhibitory effects against both fungi and bacteria (Table S1). Therefore, its hemiacetalic γ-lactone core represents a promising scaffold for future drug discovery.20
The ecological success of Lasiodiplodia spp. could be also related to the cytotoxic and immunomodulatory activities of secondary metabolites. Some preussomerins (150–152, 155–161, 164, 168, and 170) displayed cytotoxicity against human cell lines. Chlorinated preussomerins (150 and 151) showed potent cytotoxicity against A549 and MCF-7 human cancer cell lines, with IC50 values ranging from 5.9 to 8.9 µM.159 Furthermore, some preussomerins (150–152 and 156–159) that showed cytotoxicity were also active in antimicrobial assays against Staphylococcus aureus and Bacillus subtilis,159 while other preussomerins (153, 164, 172, and 173) showed suppressive effects on the production of NO.
The biological activity data reported in Table S1 highlight the dual role of Lasiodiplodia spp. metabolites, where they are responsible for fungal fitness and niche adaptation, but they also represent promising compounds for innovation in the agricultural, pharmaceutical, and biotechnological fields. In this respect, in addition to in vitro assays, reports describe some interesting in vivo bioactivities of exopolysaccharides produced by L. theobromae (Table S1). The effects of botryosphaeran (32) have been investigated in streptozotocin-induced diabetic rats and in high-fat diet-fed hyperlipidemic Wistar rats, observing that this exopolysaccharide possess hypoglycaemic and hypocholesterolemic properties in conditions of diabetes mellitus and hyperlipidaemia, respectively, and may be used as an oral anti-diabetic agent.71 Miranda et al.70 evaluated the in vivo genotoxic activity of botryosphaeran and its effect on the clastogenicity induced by cyclophosphamide in mice using the micronucleus test in bone marrow and peripheral blood cells. Data showed that this exopolysaccharide has a protective effect of cyclophosphamide-induced clastogenicity in mice. Furthermore, an interesting study showed that when sulfonated, botryosphaeran induces anticoagulant activity in activated partial thromboplastin time and thrombin time tests, while botryosphaeran did not inhibit any of the coagulation tests. Hence, the derivatization of botryosphaeran enhanced its biological activity.72
4 Structure–activity relationship (SAR)
The exploration of structure–activity relationships (SARs) is supported by the idea that specific structural components represent requirements for biological activities. Although Lasiodiplodia spp. produce structurally diverse natural products, only a limited number of metabolites has been investigated through systematic biological screening suitable for SAR analysis. As a result, significant gaps remain in the literature, limiting a comprehensive SAR interpretation and obscuring the effect of minor structural modifications on the biological activity.Nevertheless, several targeted studies have provided SAR insights, which are summarized below to illustrate the current state of knowledge.
As mentioned, (−)-jasmonic acid (65) is the parent compound of a large family of natural compounds named jasmonates (Fig. 10). (−)-Jasmonic acid is known for its role in plant growth and it is often conjugated to amino acids, particularly with isoleucine. Jasmonic acid–isoleucine has been also found as product of L. theobromae85 and previous studies reported that this compound is the most biologically active plant jasmonate.127 Three new jasmonic acid conjugates, named lasiojasmonates A–C, have been isolated from Lasiodiplodia sp., along with (−)-jasmonic acid and other known compounds. These compounds have been tested for phytotoxicity in a leaf-puncture assay on cork oak and grapevine leaves and on a tomato cutting assay. Interestingly, in these bioassays, only (−)-jasmonic acid caused vein necrosis or plant withering, suggesting that esterification with the lactone/lactol moiety affected the phytotoxic activity.129 However, because only few jasmonates have been screened, the contribution of side chain substitutions, stereochemical configurations or chain length variations remains unexplored.
A remarkable evaluation of the relationships between structure and bioactivity was conducted on a series of β-resorcylic acid derivatives (Fig. 15) isolated from Lasiodiplodia spp. 318#, which were tested for their cytotoxic activity against THP-1, MDA-MB-435, A549, HepG2 and HCT-116 cell lines. Among the isolated compounds, only ethyl (S)-2,4-dihydroxy-6-(8-hydroxynonyl)benzoate (201), an open-ring lasiodiplodin, showed moderate cytotoxic activities, while the other compounds showed no notable cytotoxic activities, which indicated that the hydroxylation of C-3 or the open-ring structure increased the flexibility in the macrocyclic lactone ring and might contribute to cytotoxic activities.168 Subsequently, cytotoxic activity investigations of five other lasiodiplodins, i.e., (12E,15R)-5-hydroxy-3-methoxy-16-methyl-8,9,10,11,14,15-hexahydro-1H-benzo[c][1]oxacyclodocecin-1-one (183), ethyl 2,4-dihydroxy-6-(8-oxononyl)benzoate (205), (R)-zearalenone (199), 2,4-dihydroxy-6-nonylbenzoate (200) and (R)-de-O-methyllasiodiplodin (185), against MMQ and GH3 cell lines confirmed the importance of the hydroxyl group at the C-3 position. In fact, (12E,15R)-5-hydroxy-3-methoxy-16-methyl-8,9,10,11,14,15-hexahydro-1H-benzo[c][1]oxacyclodocecin-1-one (183), which is characterized by a methoxy moiety instead of a hydroxyl group at the C-3 position, was inactive. Furthermore, comparing the activity of ethyl 24-dihydroxy-6-(8-oxononyl)benzoate and (R)-zearalane, it seems that carbonylation of position C-15 reduced the cytotoxic activities (Fig. 16).168
Some insights into the SAR of β-resorcylic acid derivatives have been provided studying the phytotoxicity of ten compounds obtained from L. theobromae GC-22. (15S)-De-O-methyllasiodiplodin (207), (14S,15S)-14-hydroxy-de-O-methyllasiodiplodin (209) and ethyl 2,4-dihydroxy-6-(8-hydroxyheptyl)benzoate (202) showed phytotoxic effects against Digitaria ciliaris in a dose-dependent manner, while (13R,14S,15S)-13,14-dihydroxy-de-O-methyllasiodiplodin had a weak phytotoxic effect, which suggested that a hydroxyl group at C-14 had a significant impact on the phytotoxic activity and the presence of an additional hydroxyl group at C-13 of the macrocyclic lactone ring resulted in diminished phytotoxicity. Similarly, the hydroxyl or carbonyl group on the side chain attached to C-7 plays an important role in plant growth activity because ethyl 2,4-dihydroxy-6-(4-methoxycarbonylbutyl)benzoate (203), isobutyl (S)-2,4-dihydroxy-6-(8-hydroxynonyl)-benzoate (204) and ethyl 2,4-dihydroxy-6-(8-oxononyl)benzoate (205) displayed enhanced elongation activity toward D. ciliaris.171 This result has also been confirmed by testing the phytotoxic effect of (3S,7R)-7-hydroxy-13-O-methyl-de-O-methyllasiodiplodin (206) and (3S,7R)-9-etheno-7-hydroxy-13-O-methyl-de-O-methyllasiodiplodin (193). In fact, these compounds share the same skeleton but the double bond and a hydroxyl group at C-7 of (3S,7R)-9-etheno-7-hydroxy-13-O-methyl-de-O-methyllasiodiplodin (193) reduced the phytotoxicity.165
Another interesting correlation between chemical structure and bioactivity arises from preussomerins (Fig. 14) isolated from L. theobromae ZJ-HQ1, which have been evaluated for their cytotoxicity on A549, HepG2, HeLa, MCF-7 and HEK293T human cell lines (Table S1). Chloropreussomerins A (150) and B (151) and preussomerin D (155) showed significant activity against the A549 and MCF-7 cell lines, while preussomerins F, G, H and K (156–159, respectively) exhibited promising growth-inhibitory effects on the A549, HepG2, and MCF-7 cell lines. According to these data, it seems that the cytotoxicity of preussomerins strictly depends on the substitution of ring A. In fact, preussomerins F, G, H and K showed the same level of cytotoxicity because they all contain a ketone carbonyl and epoxide at ring A, whereas preussomerins A (152) and C (153) and Ymf 1029 E (161), presenting a hydroxyl group at C-1 of ring A instead of a ketone carbonyl group, displayed weak or no cytotoxic activity against all five cell lines. Furthermore, chlorinated preussomerins showed stronger inhibitory effects against the five human cell lines, indicating that the chlorine group at C-2 was important for cytotoxicity.159 The same researchers also tested some isolated preussomerins (Fig. 14) for their antibacterial activities, observing different levels of activity against Gram-positive bacteria and no activity against Gram-negative bacteria (Table S1). This result suggests that these compounds possess selective inhibition against Gram-positive bacteria.159 Overall, the existing SAR data for preussomerins are insightful but far from comprehensive because these class features are highly complex and only a small subset has been investigated.
Steroidal metabolites from L. pseudotheobromae reveal a straightforward SAR framework. The new steroids (22E,24R)-4α,5α-epoxyergosta-9α,14β-dihydroxy-7,22-diene-3,6-dione (220), (22E,24R)-4α,5α-epoxyergosta-9α,14α-dihydroxy-7,22-diene-3,6-dione (221), 12α-hydroxyergosta-7,22,24(28)-triene-3-one (222) and 3β,12α-dihydroxyergosta-7,24(28)-diene (223), along with a known congener (22E,24R)-9α,14β-dihydroxyergosta-4,7,22-triene-3,6-dione (224) (Fig. 16) have been tested for their cytotoxicity against five human cancer cell lines (RKO, A549, HepG2, SU-DHL-2, and HL-60). (22E,24R)-9α,14β-Dihydroxyergosta-4,7,22-triene-3,6-dione (224) showed remarkable activity, whereas (22E,24R)-4α,5α-epoxyergosta-9α,14β-dihydroxy-7,22-diene-3,6-dione (220) only exhibited moderate activities. Comparing the structure and the cytotoxic activities of these compounds, it can be deduced that the epoxidation of the double bond at C-4 reduces the cytotoxic activity of these steroids, while the α-oriented OH at C-14 is essential for their cytotoxicity. In the same study, these compounds have been also evaluated for neuroprotective activity and 3β,12α-dihydroxyergosta-7,24(28)-diene (223) was found to be the most active in vitro on glutamate-treated SH-SY5Y cells.191 The current SAR landscape is built on relatively small analogue sets, clearly indicating that future efforts should prioritize studies creating a bridge between chemical structure and biological activity. Hence, the SAR insights outlined in this section do not mark a definitive endpoint but rather provide the preliminary basis for a more focused chemical–biological framework of Lasiodiplodia secondary metabolism.
5 Biosynthesis
Secondary metabolites are produced by a sequence of chemical transformations catalysed by enzymes. Biosynthetic pathways can differ according to the (micro)organism. In fact, there are alternative routes that could lead to the production of the same or similar compounds.202Biosynthetic research on Lasiodiplodia spp. metabolites has delineated that they are essentially produced by polyketide and shikimate pathways. Not all pathways are experimentally determined in Lasiodiplodia spp., but possible biosynthetic origins can be hypothesized based on known chemical reactions.
Most compounds produced by Lasiodiplodia spp. are polyketides, which, while structurally diverse, are obtained by successive rounds of decarboxylative Claisen condensations between a thioesterified malonate derivative and an acyl thioesterconstitute.202
The biosynthesis of some metabolites via the polyketide biosynthetic pathway has been demonstrated by the administration of 13C/2H-labeled acetates to Lasiodiplodia spp. A possible biosynthetic pathway has been described for theobroxide (1) and its related compounds through the administration of 13C-labeled acetates to L. theobromae to establish the tetraketide origins. By administering 2H and 13C-labeled acetates, the origin of the carbonate carbon of the theobroxide derivative was also determined (Fig. 19).203
Similar studies have been conducted to investigate lasiodiplodin (177) and its (5S)-5-hydroxylated derivative (181) by the administration of 13C-labeled acetates to L. theobromae.174,175 It has been demonstrated that these metabolites are biosynthesized via highly reduced acyl intermediates in the same manner as other resorcylic acid derivatives (Fig. 20). An extension of this biosynthetic pathway has been proposed to generate lasiodiplactone A (196).170
Based on biosynthetic knowledge of preussomerins, Liang et al.158 hypothesized the biosynthesis of lasiodiplodiapyrones A and B (172 and 173), two adducts of an α-pyrone and a polyketone obtained from L. pseudotheobromae 414-JZ-40, respectively, along with some congeners (Fig. 21).
There is extensive literature on the detailed mechanism of (−)-jasmonic acid (65) biosynthesis in plants starting from the oxygenation of α-linolenic acid. It is interesting that the cyclization mechanism in L. theobromae appears to be identical to that in plants. In fact, it is demonstrated that (−)-jasmonic acid is synthesized via a fatty acid synthetic pathway in L. theobromae, which is supported by 13C labeling experiments. The incorporation of a synthetic 2H-labeled linolenic acid into iso-jasmonic acid indicates that (−)-jasmonic acid biosynthesis in L. theobromae is similar to that of plants, differing only in the facial selectivity of the cyclopentenone reduction (Fig. 22).204 It was reported that the biosynthetic pathway of cis-jasmone (67) in plants proceeded using (−)-jasmonic acid as a biosynthetic intermediate (Fig. 22). By using a combined approach involving feeding the fungus deuterium-labeled compounds and GC-MS analysis, the cis-jasmone biosynthetic pathway in L. theobromae was elucidated.135
The shikimate pathway is the metabolic process responsible for the biosynthesis of the aromatic amino acids: phenylalanine, tyrosine, and tryptophan. Microorganisms have evolved to assemble various secondary metabolites using these amino acids as building blocks.202
Indole-3-carbaldehyde (55) and indole-3-carboxylic acid (56) are produced by several microorganisms including Lasiodiplodia spp. It has been suggested that tryptophan was converted to indole-3-carboxylic acid via indole acetic acid in microorganisms, while some plants can directly metabolise L-tryptophan to indole-3-carboxaldehyde, which is further converted to indole-3-carboxylic acid. By analysing the metabolic intermediates of Lasiodiplodia sp. ME4-2, an indole acetic acid-independent route has been hypothesized,205 while based on the metabolic investigation of cultures of L. citricola ALG 111, indole-3-carboxylic acid was detected along with indole acetic acid, suggesting that it was biosynthesised from tryptophan via the indole acetic acid pathway.84
6 Synthetic strategies
Owing to their unique structural features and interesting bioactivities, Lasiodiplodia spp. secondary metabolites have attracted attention from organic chemists to develop strategies for their synthesis. Many papers are available for a detailed description of synthetic approaches and thus we will focus on recent advancements as examples of the huge work done on the synthesis of these natural products. The supply of fungal compounds could be very low, as in the case of theobroxide (1), an epoxy cyclohexene compound isolated from cultures of Lasiodiplodia spp. showing potato micro-tuber-inducing activity and phytotoxicity (Table S1). Hence, an efficient synthesis of theobroxide has been implemented starting with the commercially available 1-methylcyclohexa-1,4-diene and the singlet oxygen oxidation of arene oxide as the key step.206 The optical resolution of racemic theobroxide was also achieved with lipase (Fig. 23). Practically, this concise four-step approach is advantageous because it reduces the number of synthetic steps compared to previous, more laborious strategies for the synthesis of theobroxide.207,208As reported in the previous sections, Lasiodiplodia spp. produce several β-resorcylic acid derivatives characterized by a scaffold that has been the target of several synthetic studies over the past 50 years.209–212 The synthesis of (R)-(+)-lasiodiplodin (177) has been reported by several researchers, where most of them synthesised this compound starting from chiral materials (e.g., enantiopure epoxides and alcohols), while some researchers synthesized the racemic form. Huang et al.213 described the total synthesis of (R)-(+)-lasiodiplodin using catalytic asymmetric allylic substitution, sp3–sp2 Suzuki coupling and alkene ring-closing metathesis as key steps (Fig. 24). Recently, the protecting group-free synthesis of (±)-de-O-methyllasiodiplodin was reported.214 This five-step synthesis proceeds in 42% yield from affordable starting materials (i.e. 9-decenoic acid). This efficient synthesis marks a significant improvement over previous strategies, which were limited by low yields.212,215,216
Substances with the 2-(2-phenylethyl)chromone scaffold from natural sources, including from Lasiodiplodia spp., have restricted availability, which has stimulated the development of synthetic approaches. Fig. 25 shows the synthesis of 2-(2-phenylethyl)chromones with hydroxylation and methoxylation as key mechanisms that contribute to their structural diversity.217
Due to their original chemical structures, preussomerins have stimulated many creative synthetic approaches.156 The total syntheses of members of the preussomerin family have been achieved with the identification of the absolute stereochemistry of some of these natural products.218,219 Fig. 26 shows the total synthesis of (±)-preussomerin G (157) leading to the bis-acetal ring system. In this elegant approach, the preussomerin skeleton is synthesized by a direct acetalization method as the first step, followed by additional oxidation, which is a possible biomimetic route. However, (−)-preussomerin G was synthesized by Barrett et al.220 in 2002, achieving the first enantioselective route for accessing this family of natural products.
7 Concluding remarks
This review highlighted compounds discovered through the research activities conducted worldwide on Lasiodiplodia spp. The enormous chemical diversity of Lasiodiplodia metabolites (273 chemically defined compounds) and their bioactivities confirm the role of these compounds in fungal associations and adaptability. These compounds were classified by structure into fourteen distinct groups and an additional miscellaneous group containing compounds that show no structural affinity to the established classes. The biosynthetic capabilities of these fungi are quite original, as they constitute the only known source of certain unique natural products, such as theobroxide, lasiosan, lasiodiplodan, botryosphaeran, and lasiodiplodiapyrones A and B.The ecological success of these fungi seems to be driven by their ability to produce secondary metabolites that enhance both host adaptability and infection mechanisms. This is further evidenced by the broad spectrum of bioactivities observed in Lasiodiplodia spp. metabolites, ranging from phytotoxic to antimicrobial and cytotoxic effects. However, some metabolites require an in-depth study to clarify their roles in fungal infection. In particular, Lasiodiplodia spp. are prolific producers of jasmonates, which warrant attention due to their roles as plant hormones. Further research is necessary to elucidate the mechanisms of fungal production for these established plant compounds.
When investigated, the biosynthetic pathways leading to the production of secondary metabolites in Lasiodiplodia spp. are similar to those of plants and other microorganisms, differing only in few details.
These metabolites have shown significant bioactivities suggesting the potential use of Lasiodiplodia spp. in biotechnological applications. Therefore, these fungi should be considered as biofactories with the potential to significantly expand the bioactive products currently known for use in various industrial sectors. For this reason, the synthesis of these compounds has been challenging since their initial discovery and continues to stimulate the creativity of organic chemists.
8 Author contributions
M. M. S.: conceptualization, writing – original draft, and writing – review and editing; M. M.: writing – review and editing and project administration; M. D. G.: writing – review and editing and project administration; and A. A.: writing – review and editing and project administration.9 Conflicts of interest
There are no conflicts to declare.10 Data availability
No primary research, software or code have been included and no new data were generated or analysed as part of this manuscript.The data supporting this article (Table S1) have been included as part of the supplementary information (SI). See DOI: https://doi.org/10.1039/d5np00090d.
11 Acknowledgements
This work was carried out within the Program for the Finanziamento della Ricerca di Ateneo (FRA) 2022 dell’Università degli Studi di Napoli Federico II and by the funding from the Progetto PRIN: PROGETTI DI RICERCA DI RILEVANTE INTERESSE NAZIONALE: Bando 2022 PNRR Prot. P2022WXE4T, financed by the European Union Next-Generation EU (Piano Nazionale di Ripresa e Resilienza, PNRR).12 References
- B. Slippers and M. J. Wingfield, Fungal Biol. Rev., 2007, 21, 90–106 CrossRef.
- M. M. Salvatore, A. Andolfi and R. Nicoletti, Agriculture, 2020, 10, 488 CrossRef CAS.
- M. M. Raza and D. P. Bebber, Curr. Opin. Microbiol., 2022, 70, 102233 CrossRef CAS PubMed.
- O. Z. Dianda, I. Wonni, L. Ouédraogo, P. Sankara, C. Tollenaere, E. M. Del Ponte and D. Fernandez, Physiol. Mol. Plant Pathol., 2023, 126, 102041 CrossRef.
- C. Félix, A. S. Duarte, R. Vitorino, A. C. L. Guerreiro, P. Domingues, A. C. M. Correia, A. Alves and A. C. Esteves, Front. Plant Sci., 2016, 7, 1096 Search PubMed.
- J. B. Ellis and B. M. Everhart, Proc. Acad. Nat. Sci. Philadelphia, 1894, 46, 322–386 Search PubMed.
- I. Clendenin, Bot. Gaz., 1896, 21, 92–93 CrossRef PubMed.
- A. Alves, P. W. Crous, A. Correia and A. J. L. Phillips, Fungal Diversity, 2008, 28, 1–13 Search PubMed.
- T. I. Burgess, P. A. Barber, S. Mohali, G. Pegg, W. De Beer and M. J. Wingfield, Mycologia, 2006, 98, 423–435 CrossRef CAS PubMed.
- D. Pavlic, B. Slippers, T. A. Coutinho, M. Gryzenhout and M. J. Wingfield, Stud. Mycol., 2004, 50, 313–322 Search PubMed.
- B. T. Linaldeddu, A. Deidda, B. Scanu, A. Franceschini, S. Serra, A. Berraf-Tebbal, M. Zouaoui Boutiti, M. L. Ben Jamâa and A. J. L. Phillips, Fungal Diversity, 2015, 71, 201–214 CrossRef.
- M. S. B. Netto, I. P. Assunção, G. S. A. Lima, M. W. Marques, W. G. Lima, J. H. A. Monteiro, V. de Queiroz Balbino, S. J. Michereff, A. J. L. Phillips and M. P. S. Câmara, Fungal Diversity, 2014, 67, 127–141 CrossRef.
- D. L. Hawksworth, P. W. Crous, S. A. Redhead, D. R. Reynolds, R. A. Samson, K. A. Seifert, J. W. Taylor, M. J. Wingfield, Ö. Abaci, C. Aime, A. Asan, F. Y. Bai, Z. W. de Beer, D. Begerow, D. Berikten, T. Boekhout, P. K. Buchanan, T. Burgess, W. Buzina, L. Cai, P. F. Cannon, J. L. Crane, U. Damm, H. M. Daniel, A. D. van Diepeningen, I. Druzhinina, P. S. Dyer, U. Eberhardt, J. W. Fell, J. C. Frisvad, D. M. Geiser, J. Geml, C. Glienke, T. Gräfenhan, J. Z. Groenewald, M. Groenewald, J. de Gruyter, E. Guého-Kellermann, L. D. Guo, D. S. Hibbett, S. B. Hong, G. S. de Hoog, J. Houbraken, S. M. Huhndorf, K. D. Hyde, A. Ismail, P. R. Johnston, D. G. Kadaifciler, P. M. Kirk, U. Kõljalg, C. P. Kurtzman, P. E. Lagneau, C. André Lévesque, X. Liu, L. Lombard, W. Meyer, A. Miller, D. W. Minter, M. J. Najafzadeh, L. Norvell, S. M. Ozerskaya, R. Öziç, S. R. Pennycook, S. W. Peterson, O. V. Pettersson, W. Quaedvlieg, V. A. Robert, C. Ruibal, J. Schnürer, H. J. Schroers, R. Shivas, B. Slippers, H. Spierenburg, M. Takashima, E. Taskin, M. Thines, U. Thrane, A. H. Uztan, M. van Raak, J. Varga, A. Vasco, G. Verkley, S. I. R. Videira, R. P. de Vries, B. S. Weir, N. Yilmaz, A. Yurkov and N. Zhang, IMA Fungus, 2011, 2, 105–111 CrossRef PubMed.
- MycoBank, https://www.mycobank.org, accessed 12 December 2025.
- N. Amaresan and K. Kumar, Compendium of Phytopathogenic Microbes in Agro-Ecology, 2025, vol. 1 Search PubMed.
- S. Jiang, L. Hu, Z. Rui, D. Wang, L. Zhang, J. J. Zhou and Z. Chen, J. Agric. Food Chem., 2025, 73, 21223–21234 CrossRef CAS.
- P. Borgohain, P. Barua, J. Mahanta, L. R. Saikia, D. Shaw and S. M. Rudramurthy, J. Med. Mycol., 2021, 31, 101167 CrossRef PubMed.
- P. D. da Rosa, C. Locatelli, K. Scheid, D. Marinho, L. Kliemann, A. Fuentefria and L. Z. Goldani, Mycopathologia, 2018, 183, 565–571 CrossRef.
- A. K. Maurya, S. Kumari, G. Behera, A. Bhadade and K. Tadepalli, Med. Mycol. Case Rep., 2023, 40, 22–24 CrossRef PubMed.
- S. R. Gupta, R. R. Chandran and P. V. Divekar, Indian J. Exp. Biol., 1966, 4, 152–153 Search PubMed.
- D. C. Nwobodo, M. C. Ugwu, P. M. Eze, U. M. Okezie, F. B. Okoye and C. O. Esimone, Not. Sci. Biol., 2022, 14, 11284 CrossRef CAS.
- C. dos Reis Feliciano, H. S. de Souza, V. C. Costa, O. C. Gómez, J. H. H. Luiz, L. F. Gorup and M. G. Santos, Diamond Relat. Mater., 2024, 142, 110763 Search PubMed.
- K. Kavitha, U. Yuvaraj, A. Rajalakshmi, G. Suresh, M. Harini, V. Prabakaran, S. Bharathi, R. Puvanakrishnan and B. Ramesh, Chem. Biodiversity, 2025, 22, 1–13 CrossRef PubMed.
- G. Maeda, J. Van Der Wal, A. K. Gupta, J. J. E. Munissi, A. Orthaber, P. Sunnerhagen, S. S. Nyandoro and M. Erdélyi, J. Nat. Prod., 2020, 83, 210–215 CrossRef CAS PubMed.
- M. T. J. Quimque, R. J. Y. Magsipoc, L. C. J. Llames, A. I. G. Flores, K. Y. M. Garcia, A. Ratzenböck, H. Hussain and A. P. G. MacAbeo, ACS Omega, 2022, 7, 36856–36864 CrossRef CAS PubMed.
- M. M. Salvatore, A. Alves and A. Andolfi, Agriculture, 2021, 11, 149 CrossRef CAS.
- M. M. Salvatore, M. Masi and A. Andolfi, Phytochem. Rev., 2025, 24, 1565–1589 CrossRef CAS.
- K. Nakamori, H. Matsuura, T. Yoshihara, A. Ichihara and Y. Koda, Phytochemistry, 1994, 35, 835–839 CrossRef CAS.
- K. Naoki, K. Nabeta and H. Matsuura, Biosci., Biotechnol., Biochem., 2009, 73, 1890–1892 CrossRef PubMed.
- H. Matsuura, N. Obara, N. Chisaka, A. Ichihara and T. Yoshihara, Biosci., Biotechnol., Biochem., 1998, 62, 2460–2462 CrossRef CAS PubMed.
- R. Takei, K. Takahashi, H. Matsuura and K. Nabeta, Biosci., Biotechnol., Biochem., 2008, 72, 2069–2073 CrossRef CAS PubMed.
- J. Jia, J. Yao, J. Kong, A. Yu, J. Wei, Y. Dong, R. Song, D. Shan, X. Zhong, F. Lv and Q. Fan, Curr. Med. Chem., 2023, 306, 1060–1085 CrossRef PubMed.
- A. Ortiz and E. Sansinenea, Curr. Med. Chem., 2017, 24, 2773–2780 CrossRef CAS PubMed.
- R. M. Huang, X. X. Yi, Y. Zhou, X. Su, Y. Peng and C. H. Gao, Mar. Drugs, 2014, 12, 6213–6235 CrossRef PubMed.
- W. Wei, N. Jiang, Y. N. Mei, Y. L. Chu, H. M. Ge, Y. C. Song, S. W. Ng and R. X. Tan, Phytochemistry, 2014, 100, 103–109 CrossRef CAS PubMed.
- S. Khruengsai, P. Pripdeevech, C. Tanapichatsakul, W. C. Sum, M. A. A. Ibrahim, M. Stadler and S. S. Ebada, RSC Adv., 2023, 13, 19373–19378 RSC.
- S. Liuu, A. Damont, A. Perret, O. Firmesse, F. Becher, G. Lavison-Bompard, A. Hueber, A. S. Woods, E. Darii, F. Fenaille and J.-C. Tabet, Mass Spectrom. Rev., 2025, 1013–1098 CrossRef CAS PubMed.
- Z. Y. Guo, Z. J. Huang, L. Wen, Q. Wan, F. Liu, Z. G. She, Y. C. Lin and S. N. Zhou, J. Chin. Med. Mater., 2007, 30, 1526–1529 CAS.
- P. Ye, L. Shen, W. Jiang, Y. Ye, C. A. Chen, X. Wu, K. Wang and B. Wu, Mar. Drugs, 2014, 12, 3203–3217 CrossRef PubMed.
- U. M. Okezie, C. I. Chimezie, N. J. Okonkwo-Uzor, V. U. Chigozie, O. G. Tochukwu, I. N. Okpoli, M. G. U. Nwaneri, J. E. Achilonu, O. B. Ifeagwu, C. J. Ikem, C. C. Onwuzuligbo and S. H. Buzugbe, Asian J. Biochem., Genet. Mol. Biol., 2025, 17, 58–68 CrossRef.
- K. Sakai, M. Iwatsuki, M. Iizuka, Y. Asami, K. Nonaka, R. Masuma, M. Takizawa, T. Nakashima, T. Tokiwa, K. Shiomi and S. Ōmura, J. Antibiot., 2021, 74, 363–369 CrossRef CAS PubMed.
- L. M. Delgado Gómez, D. Torres-Mendoza, K. Hernández-Torres, H. E. Ortega and L. Cubilla-Rios, Metabolites, 2023, 13, 912 CrossRef PubMed.
- A. M. Zaher, A. M. Moharram, R. Davis, P. Panizzi, M. A. Makboul and A. I. Calderón, Nat. Prod. Res., 2015, 29, 2275–2281 CrossRef CAS PubMed.
- S. S. El-Hawary, A. M. Sayed, M. E. Rateb, W. Bakeer, S. F. AbouZid, R. Mohammed, A. M. Sayed and M. E. Rateb, Nat. Prod. Res., 2017, 31, 2568–2571 CrossRef CAS PubMed.
- M. M. Salvatore, C. Félix, F. Lima, V. Ferreira, A. S. Duarte, F. Salvatore, A. Alves, A. C. Esteves and A. Andolfi, Molecules, 2020, 25, 3833 CrossRef CAS PubMed.
- C. Félix, M. M. Salvatore, M. Dellagreca, R. Meneses, A. S. Duarte, F. Salvatore, D. Naviglio, M. Gallo, J. V. Jorrín-Novo, A. Alves, A. Andolfi and A. C. Esteves, Mycologia, 2018, 110, 642–653 CrossRef PubMed.
- I. Ureña-Vacas, E. González-Burgos, P. K. Divakar and M. P. Gómez-Serranillos, Planta Med., 2022, 88, 855–880 CrossRef PubMed.
- G. Stojanovic, I. Stojanovic and A. Smelcerovic, Mini-Rev. Org. Chem., 2012, 9, 178–184 CrossRef CAS.
- S. R. M. Ibrahim, G. A. Mohamed, R. A. Al Haidari, A. A. El-Kholy, M. F. Zayed and M. T. Khayat, Fitoterapia, 2018, 129, 317–365 CrossRef CAS PubMed.
- N. Thi Hoang Anh, N. Mai Anh, V. Thi Thu Huyen, P. Thi Dao, D. Thi Mai Huong, P. Van Cuong, D. Thanh Xuan, B. Huu Tai, L. Thi Hong Minh and P. Van Kiem, Chem. Biodiversity, 2023, 20, e202301660 CrossRef CAS PubMed.
- P. Phainuphong, V. Rukachaisirikul, S. Phongpaichit, J. Sakayaroj, P. Kanjanasirirat, S. Borwornpinyo, N. Akrimajirachoote, C. Yimnual and C. Muanprasat, Tetrahedron, 2018, 74, 5691–5699 CrossRef CAS.
- S. Zhigang, C. Senhua, L. Zhaoming, L. Yayue, T. Chunbing, L. Yongjun and H. Lei, CN 104987319A, 2015, 20151021, Patent written in Chinese Application: CN 2015-10205871.
- P. Saetang, V. Rukachaisirikul, S. Phongpaichit, S. Preedanon, J. Sakayaroj, S. Borwornpinyo, S. Seemakhan and C. Muanprasat, Phytochemistry, 2017, 143, 115–123 CrossRef CAS PubMed.
- W. Jampanya, C. Boonlarppradab, C. Srisuksam, S. Samipak and A. Amnuaykanjanasin, Crop Prot., 2025, 188, 107029 CrossRef CAS.
- S. Purbaya, S. E. Sinaga, W. Safriansyah, A. P. Wulandari, D. Harneti, Y. Mulyani, A. Azhari, K. Farabi, F. F. Abdullah and U. Supratman, J. Biol. Act. Prod. Nat., 2024, 14, 443–454 CAS.
- R. Abdou, K. Scherlach, H. Dahse, I. Sattler and C. Hertweck, Phytochemistry, 2010, 71, 110–116 CrossRef CAS PubMed.
- B. O. Umeokoli, W. Ebrahim, M. El-Neketi, W. E. G. Müller, R. Kalscheuer, W. Lin, Z. Liu and P. Proksch, Nat. Prod. Res., 2019, 33, 2215–2222 CrossRef CAS PubMed.
- S. J. Lee, H. S. Jeon, J. Y. Yoo and J. H. Kim, Foods, 2021, 10, 2148 CrossRef CAS PubMed.
- O. O. Osemwegie, C. O. Adetunji, E. A. Ayeni, O. I. Adejobi, R. O. Arise, C. O. Nwonuma and A. O. Oghenekaro, Heliyon, 2020, 6, e04205 CrossRef PubMed.
- F. Salimi and P. Farrokh, World J. Microbiol. Biotechnol., 2023, 39, 213 CrossRef CAS PubMed.
- C. G. Kumar, P. Mongolla and S. Pombala, Process Biochem., 2018, 72, 162–169 CrossRef CAS.
- M. A. Alves Da Cunha, J. A. Turmina, R. C. Ivanov, R. R. Barroso, P. T. Marques, E. A. I. Fonseca, Z. B. Fortes, R. F. H. Dekker, N. Khaper and A. M. Barbosa, J. Ind. Microbiol. Biotechnol., 2012, 39, 1179–1188 CrossRef CAS PubMed.
- A. F. D. Vasconcelos, N. K. Monteiro, R. F. H. Dekker, A. M. Barbosa, E. R. Carbonero, J. L. M. Silveira, G. L. Sassaki, R. da Silva and M. de Lourdes Corradi da Silva, Carbohydr. Res., 2008, 343, 2481–2485 CrossRef CAS PubMed.
- J. Ruiz-Herrera and L. Ortiz-Castellanos, Cell Surf., 2019, 5, 100022 CrossRef CAS PubMed.
- M. De Lourdes Corradi Da Silva, N. L. Izeli, P. F. Martinez, I. R. Silva, C. J. L. Constantino, M. S. Cardoso, A. M. Barbosa, R. F. H. Dekker and G. V. J. Da Silva, Carbohydr. Polym., 2005, 61, 10–17 CrossRef.
- A. M. Barbosa, R. M. Steluti, R. F. H. Dekker, M. S. Cardoso and M. L. Corradi Da Silva, Carbohydr. Res., 2003, 338, 1691–1698 CrossRef CAS PubMed.
- P. Abdeshahian, R. R. Philippini, F. Antonio, F. Antunes, A. P. Ingle, M. Abdeshahian and S. Silv, Sustainability, 2021, 13, 9697 CrossRef CAS.
- M. L. K. Marchioro, G. A. P. B. Candeia, L. M. Bertoleti, A. M. Barbosa-Dekker, R. F. H. Dekker and M. A. A. da Cunha, Fermentation, 2025, 11, 166 CrossRef CAS.
- M. C. Pires, N. de Gois Andriolo, B. R. P. Lopes, A. L. T. G. Ruiz, V. M. G. do Nascimento, K. A. Toledo and C. dos Santos, BMC Complementary Med. Ther., 2023, 23, 356 CrossRef CAS PubMed.
- C. C. B. O. Miranda, R. F. H. Dekker, J. M. Serpeloni, E. A. I. Fonseca, I. M. S. Cólus and A. M. Barbosa, Int. J. Biol. Macromol., 2008, 42, 172–177 CrossRef CAS PubMed.
- C. C. B. O. Miranda-Nantes, E. A. I. Fonseca, C. T. B. V. Zaia, R. F. H. Dekker, N. Khaper, I. A. Castro and A. M. Barbosa, Mycobiology, 2011, 39, 187–193 CrossRef CAS PubMed.
- S. F. Mendes, O. dos Santos, A. M. Barbosa, A. F. D. Vasconcelos, G. Aranda-Selverio, N. K. Monteiro, R. F. H. Dekker, M. S. Pereira, A. M. F. Tovar, P. A. de Souza Mourão and M. de Lourdes Corradi da Silva, Int. J. Biol. Macromol., 2009, 45, 305–309 CrossRef CAS PubMed.
- B. B. C. Weng, Y. C. Lin, C. W. Hu, M. Y. Kao, S. H. Wang, D. Y. Lo, T. Y. Lai, L. S. Kan and R. Y. Y. Chiou, Food Chem. Toxicol., 2011, 49, 910–916 CrossRef CAS PubMed.
- L. Selbmann, F. Stingele and M. Petruccioli, Antonie van Leeuwenhoek, 2003, 84, 135–145 CrossRef CAS PubMed.
- G. Carta, E. Murru, S. Banni and C. Manca, Front. Physiol., 2017, 8, 902 CrossRef PubMed.
- R. A. Sidorov, A. V. Zhukov, V. P. Pchelkin and V. D. Tsydendambaev, in Palmitic Acid: Occurrence, Biochemistry and Health Effects, NOVA Publishers, New York, NY, USA, 2018, pp. 125–144 Search PubMed.
- A. C. Rustan and C. A. Drevon, Fatty acids: structures and properties, in Encycl. Life Sci., 2005, DOI:10.1038/npg.els.0003894.
- C. C. C. R. De Carvalho and M. J. Caramujo, Molecules, 2018, 23, 2583 CrossRef PubMed.
- J. W. Walley, D. J. Kliebenstein, R. M. Bostock and K. Dehesh, Curr. Opin. Plant Biol., 2013, 16, 520–526 CrossRef CAS PubMed.
- A. Schaller and A. Stintzi, Phytochemistry, 2009, 70, 1532–1538 CrossRef CAS PubMed.
- C. C. Uranga, J. Beld, A. Mrse, I. Córdova-Guerrero, M. D. Burkart and R. Hernández-Martínez, Biochem. Biophys. Res. Commun., 2016, 472, 339–345 CrossRef CAS PubMed.
- C. C. Uranga, J. Beld, A. Mrse, I. Córdova-Guerrero, M. D. Burkart and R. Hernández-Martínez, Data Brief, 2016, 8, 31–39 CrossRef PubMed.
- T. V. Sravanthi and S. L. Manju, Eur. J. Pharm. Sci., 2016, 91, 1–10 CrossRef CAS PubMed.
- M. M. Salvatore, M. DellaGreca, R. Nicoletti, F. Salvatore, A. Tuzi, G. De Tommaso, A. Alves, A. E. Mahamedi, A. Berraf-Tebbal and A. Andolfi, Nat. Prod. Res., 2023, 37, 424–433 CrossRef CAS PubMed.
- G. Castillo, A. Torrecillas, C. Nogueiras, G. Michelena, J. Sánchez-Bravo and M. Acosta, World J. Microbiol. Biotechnol., 2014, 30, 1937–1946 CrossRef CAS PubMed.
- H. Nishikawa, Bull. Agric. Chem. Soc. Jpn., 1933, 9, 10–12 Search PubMed.
- M. M. Salvatore, A. Andolfi and R. Nicoletti, Molecules, 2021, 26, 3959 CrossRef CAS PubMed.
- J. Wei and B. Wu, Fitoterapia, 2020, 146, 104638 CrossRef CAS PubMed.
- R. Nicoletti, A. Andolfi, A. Becchimanzi and M. M. Salvatore, Microorganisms, 2023, 11, 1–32 CrossRef PubMed.
- E. S. Elkhayat and A. M. Goda, Bull. Fac. Pharm., 2017, 55, 85–89 Search PubMed.
- M. M. Salvatore, M. Masi, M. DellaGreca and A. Andolfi, Phytochem. Rev., 2026, 25, 1431–1464 CrossRef CAS.
- M. M. Salvatore, A. Alves and A. Andolfi, Toxins, 2020, 12, 457 CrossRef CAS PubMed.
- M. Saraswathi, S. H. Meshram, B. Siva, S. Misra and K. Suresh Babu, Lett. Appl. Microbiol., 2022, 75, 1475–1485 CrossRef CAS PubMed.
- L. Pellissier, S. Leoni, L. Marcourt, E. F. Queiroz, N. Lecoultre, L. M. Quiros-Guerrero, M. Barthélémy, V. Eparvier, J. Chave, D. Stien, K. Gindro, K. Perron and J. L. Wolfender, Microorganisms, 2021, 9, 1807 CrossRef CAS PubMed.
- L. Pellissier, A. Koval, L. Marcourt, E. Ferreira Queiroz, N. Lecoultre, S. Leoni, L. M. Quiros-Guerrero, M. Barthélémy, B. L. Duivelshof, D. Guillarme, S. Tardy, V. Eparvier, K. Perron, J. Chave, D. Stien, K. Gindro, V. Katanaev and J. L. Wolfender, Front. Chem., 2021, 9, 664489 CrossRef CAS PubMed.
- F. Jernerén, F. Eng, M. Hamberg and E. H. Oliw, Lipids, 2012, 47, 65–73 CrossRef PubMed.
- M. T. M. Jalil and D. Ibrahim, Hayati J. Biosci., 2022, 29, 570–585 CrossRef.
- G. Segaran and M. Sathiavelu, Front. Plant Sci., 2023, 14, 1125630 CrossRef CAS PubMed.
- Y. Que, D. Huang, S. Gong, X. Zhang, B. Yuan, M. Xue, W. Shi, F. Zeng, M. Liu, T. Chen, D. Yu, X. Yan, Z. Wang, L. Yang and L. Xiang, Front. Cell. Infect. Microbiol., 2022, 12, 898500 CrossRef CAS PubMed.
- D. C. Aldridge, S. Galt, D. Giles and W. B. Turner, J. Chem. Soc. C, 1971, 1623–1627 RSC.
- C. Félix, M. M. Salvatore, M. DellaGreca, V. Ferreira, A. S. Duarte, F. Salvatore, D. Naviglio, M. Gallo, A. Alves, A. C. Esteves and A. Andolfi, Mycologia, 2019, 111, 466–476 CrossRef PubMed.
- J. V. S. Aluthmuhandiram, K. W. T. Chethana, W. Zhang, J. Peng, E. Zhao, X. H. Li, N. Saichana and J. Yan, J. Phytopathol., 2021, 169, 716–723 CrossRef CAS.
- L. Silva, J. Mateo, R. Marin, O. C. Gómez, A. Gilberto, J. Schripsema, J. Honorata and H. Luiz, Fitoterapia, 2026, 188, 107031 CrossRef PubMed.
- P. Reveglia, M. Masi, A. Cimmino, S. Michereff, T. Cinelli, L. Mugnai and A. Evidente, Phytopathol. Mediterr., 2019, 58, 207–211 CAS.
- N. Kamal, C. V. Viegelmann, C. J. Clements and R. A. Edrada-Ebel, Planta Med., 2016, 83, 565–573 CrossRef PubMed.
- M. Matsumoto and H. Nago, Biosci., Biotechnol., Biochem., 1994, 58, 1262–1266 CrossRef CAS.
- A. Cimmino, T. Cinelli, M. Masi, P. Reveglia, M. A. Da Silva, L. Mugnai, S. J. Michereff, G. Surico and A. Evidente, J. Agric. Food Chem., 2017, 65, 1102–1107 CrossRef CAS PubMed.
- V. Rukachaisirikul, J. Arunpanichlert and Y. Sukpondma, Tetrahedron, 2009, 65, 10590–10595 CrossRef CAS.
- C. O. Adetunji, J. K. Oloke, M. Pradeep, A. P. Oluyori, R. S. Jolly and O. M. Bello, Beni-Suef Univ. J. Basic Appl. Sci., 2018, 7, 505–510 Search PubMed.
- J. Arunpanichlert, V. Rukachaisirikul, T. Chaiwarin, Y. Tantirungrotechai, N. Khamthong, S. Phongpaichit, S. Liamthong and J. Sakayaroj, Nat. Prod. Res., 2022, 36, 1948–1958 CrossRef CAS PubMed.
- F. M. Nunes, M. D. C. F. De Oliveira, Â. M. C. Arriaga, T. L. G. Lemos, M. Andrade-Neto, M. C. De Mattos, J. Mafezoli, F. M. P. Viana, V. M. Ferreira, E. Rodrigues-Filho and A. G. Ferreira, J. Braz. Chem. Soc., 2008, 19, 478–482 CrossRef CAS.
- U. Okezie, P. M. Eze, F. Okoye and C. O. Esimone, Not. Sci. Biol., 2022, 14, 11084 CrossRef CAS.
- E. Demole, E. Lederer and D. Mercier, Helv. Chim. Acta, 1962, 45, 675 CrossRef CAS.
- D. A. Fitzpatrick, FEMS Microbiol. Lett., 2012, 329, 1–8 CrossRef CAS PubMed.
- P. Tiwari and H. Bae, Plants, 2020, 9, 305 CrossRef CAS PubMed.
- X. Wang, W. Zhang, J. Peng, I. S. Manawasinghe, L. Wu, Y. Li, Q. Xing, X. Li and J. Yan, Fungal Diversity, 2024, 125, 221–241 CrossRef CAS.
- Z. L. Deng, A. J. Dissanayake, J. T. Zhu, N. Wu, J. Deng, H. Z. Du, W. L. Li, Y. H. Lu, X. Tang, J. Xu, Y. Zhang and J. K. Liu, Fungal Diversity, 2025, 134, 1–18 CrossRef CAS.
- K. M. Jackson, M. Ponnusamy and S. Uthandi, Int. J. Curr. Microbiol. Appl. Sci., 2017, 6, 1635–1639 CrossRef CAS.
- F. Eng, S. Haroth, K. Feussner, D. Meldau, D. Rekhter, T. Ischebeck, F. Brodhun and I. Feussner, PLoS One, 2016, 11, e0167627 CrossRef PubMed.
- P. C. Dhandhukia and V. R. Thakkar, Afr. J. Biotechnol., 2007, 6, 707–712 CAS.
- P. C. Dhandhukia and V. R. Thakkar, J. Appl. Microbiol., 2008, 105, 636–643 CrossRef CAS PubMed.
- E. I. Laredo-Alcalá, J. L. Martínez-Hernandez, L. Guillen-Cisneros and F. D. Hernández-Castillo, Agrociencia, 2017, 51, 885–893 Search PubMed.
- O. Miersch, G. Schneider and G. Sembdner, Phytochemistry, 1991, 30, 4049–4051 CrossRef CAS.
- O. Miersch, A. Preiss, G. Sembdner and K. Schreiber, Phytochemistry, 1987, 26, 1037–1039 CrossRef CAS.
- O. Miersch, J. Schmidt, G. Sembdner and K. Schreiber, Phytochemistry, 1989, 28, 1303–1305 CrossRef CAS.
- P. E. Staswick and I. Tiryaki, Plant Cell, 2004, 16, 2117–2127 CrossRef CAS PubMed.
- M. Ghorbel, F. Brini, A. Sharma and M. Landi, Plant Cell Rep., 2021, 40, 1471–1494 CrossRef CAS PubMed.
- A. Piotrowska and A. Bajguz, Phytochemistry, 2011, 72, 2097–2112 CrossRef CAS PubMed.
- A. Andolfi, L. Maddau, A. Cimmino, B. T. Linaldeddu, S. Basso, A. Deidda, S. Serra and A. Evidente, Phytochemistry, 2014, 103, 145–153 CrossRef CAS PubMed.
- M. M. Salvatore, M. T. Russo, S. Meyer, A. Tuzi, M. Della Greca, M. Masi and A. Andolfi, Molecules, 2024, 29, 438 CrossRef CAS PubMed.
- A. Andolfi, S. Basso, S. Giambra, G. Conigliaro, S. Lo Piccolo, A. Alves and S. Burruano, Chem. Biodiversity, 2016, 13, 395–402 CrossRef CAS PubMed.
- A. Husain, A. Ahmad and K. Agrawal, J. Nat. Prod., 1993, 56, 2008–2011 CrossRef CAS.
- Z. Shen, P. Zheng, R. Li, X. Sun, P. Chen and D. Wu, Biotechnol. J., 2022, 17, 2100550 CrossRef CAS.
- R. Li, X. Sun, Y. Fu, D. Wu, P. Chen and P. Zheng, J. Agric. Food Chem., 2024, 72, 23379–23388 CrossRef CAS PubMed.
- R. Matsui, N. Amano, K. Takahashi, Y. Taguchi, W. Saburi, H. Mori, N. Kondo, K. Matsuda and H. Matsuura, Sci. Rep., 2017, 7, 6688 CrossRef PubMed.
- O. Ekhorutomwen and C. Nnamdi, Proceed. Niger. Acad. Sci., 2024, 16, 18–28 Search PubMed.
- S. Kumar, A. D. Pagar, F. Ahmad, V. Dwibedi, A. Wani, P. V. Bharatam, M. Chhibber, S. Saxena and I. Pal Singh, Bioorg. Chem., 2019, 87, 851–856 CrossRef CAS PubMed.
- G. He, H. Matsuura and T. Yoshihara, Phytochemistry, 2004, 65, 2803–2807 CrossRef CAS PubMed.
- W. Dong, W. B. Gao, F. W. Liu, J. Y. Jin-Ye, Y. Chen, H. L. Zhang, Z. Lv, L. N. Guo and B. Song, Phytochem. Lett., 2023, 56, 81–83 CrossRef CAS.
- E. Elfita, R. Oktiansyah, M. Mardiyanto, H. Widjajanti, A. Setiawan and S. S. A. Nasution, Biointerface Res. Appl. Chem., 2023, 13, 530 Search PubMed.
- M. Yu, Q. Q. He, X. Q. Chen, J. Feng, J. H. Wie and Y. Y. Liu, Chem. Biodiversity, 2022, 19, e202200490 CrossRef CAS PubMed.
- S. R. M. Ibrahim and G. A. Mohamed, Nat. Prod. Res., 2015, 29, 1489–1520 CrossRef CAS PubMed.
- Y. Zhang, H. Liu, W. Li, M. Tao, Q. Pan, Z. Sun, W. Ye, H. Li and W. Zhang, Chin. Herb. Med., 2017, 9, 58–62 Search PubMed.
- D. A. Whiting, Nat. Prod. Rep., 2001, 583–606 CAS.
- K. A. Scott, P. B. Cox and J. T. Njardarson, J. Med. Chem., 2022, 65, 7044–7072 CrossRef CAS PubMed.
- C. Ve, Phytochem. Rev., 2012, 153–177 Search PubMed.
- L. M. Babenko, O. E. Smirnov, K. O. Romanenko and O. K. Trunova, Ukr. Biochem. J., 2019, 91 DOI:10.15407/ubj91.03.005.
- M. M. Salvatore, A. Maione, A. Buonanno, M. Guida, A. Andolfi, F. Salvatore and E. Galdiero, Food Chem., 2025, 470, 142657 CrossRef CAS PubMed.
- A. K. Marković, J. Torić, M. Barbarić and C. J. Brala, Molecules, 2019, 24, 2001 CrossRef.
- T. C. Yadav, N. Kumar, U. Raj, N. Goel, P. K. Vardawaj, R. Prasad and V. Pruthi, J. Biomol. Struct. Dyn., 2020, 38, 382–397 CrossRef CAS PubMed.
- P. Pacheco-lin, E. Martı, E. Siles and A. Miranda-vizuete, Mech. Ageing Dev., 2012, 133, 563–574 CrossRef PubMed.
- V. C. Perez, H. Zhao, M. Lin and J. Kim, Plants, 2023, 12, 266 CrossRef CAS PubMed.
- S. D. Cook, Plant Cell Physiol., 2019, 60, 243–254 CrossRef CAS PubMed.
- L. Silva Tironi, D. M. Barbosa Moreira, R. F. Dias Bruzadelli, A. Gilberto Ferreira, J. Schripsema and J. H. Hortolan Luiz, Chem. Biodiversity, 2025, 22, e202401649 CrossRef CAS PubMed.
- N. T. Ujam, P. M. Eze, C. Ejikeugwu, F. B. C. Okoye and C. O. Esimone, Int. J. Innovative Sci. Eng. Technol., 2020, 7, 123–129 Search PubMed.
- Y. S. Cai, Y. W. Guo and K. Krohn, Nat. Prod. Rep., 2010, 27, 1840–1870 RSC.
- H. A. Weber, N. C. Baenziger and J. B. Gloer, J. Am. Chem. Soc., 1990, 112, 6718–6719 CrossRef CAS.
- Y. Liang, Q. Li, Y. Li, Y. Zheng, Y. Shen, H. Yang, Y. Lu, J. Liu and Q. Zhou, J. Nat. Prod., 2023, 86, 18–23 CrossRef CAS PubMed.
- S. Chen, D. Chen, R. Cai, H. Cui, Y. Long, Y. Lu, C. Li and Z. She, J. Nat. Prod., 2016, 79, 2397–2402 CrossRef CAS.
- X. Lü, G. Chen, Z. Li, Y. Zhang, Z. Wang, W. Rong, Y. Pei, H. Pan, H. Hua and J. Bai, Helv. Chim. Acta, 2014, 97, 1289–1294 CrossRef.
- T. P. Martins, C. Rouger, N. R. Glasser, S. Freitas, N. B. De Fraissinette, E. P. Balskus, D. Tasdemir and P. N. Leão, Nat. Prod. Rep., 2019, 36, 1437–1461 RSC.
- Z. Wu, Y. Li, D. Liu, M. Ma, J. Chen and W. Lin, Chem. Biodiversity, 2017, 14, e1700059 CrossRef PubMed.
- Y. K. Hu, L. Wang, J. H. Wang, M. J. Li, F. Li, J. Yang and Y. Zhao, Nat. Prod. Res., 2021, 35, 5948–5953 CrossRef CAS PubMed.
- O. F. Iscosa, A. F. Barrero, J. F. Sanchez, F. Reyes, I. Rodriguez, D. D. Q. Orgcnica, F. De Ciencias and U. De Granada, Phytochemistry, 1991, 30, 641–643 CrossRef.
- Y. Shiono, S. Sato, F. F. Sofian, T. Koseki, F. F. Abdullah, S. Salam, D. Harneti, R. Maharani and U. Supratman, Phytochem. Lett., 2021, 44, 1–6 CrossRef CAS.
- Q. Yang, M. Asai and T. Yoshihara, Z. Naturforsch., C: J. Biosci., 2000, 55, 546–551 CrossRef CAS.
- W. Shen, H. Mao, Q. Huang and J. Dong, Eur. J. Med. Chem., 2015, 97, 747–777 CrossRef CAS PubMed.
- J. Huang, J. Xu, Z. Wang, D. Khan, S. I. Niaz, Y. Zhu, Y. Lin, J. Li and L. Liu, Nat. Prod. Res., 2017, 31, 326–332 CrossRef CAS PubMed.
- J. Li, Y. Xue, J. Yuan, Y. Lu, X. Zhu, Y. Lin and L. Liu, Nat. Prod. Res., 2016, 30, 755–760 CrossRef CAS PubMed.
- S. Chen, Z. Liu, H. Liu, Y. Long, D. Chen, Y. Lu and Z. She, Org. Biomol. Chem., 2017, 15, 6338–6341 RSC.
- S. Sato, F. F. Sofian, W. Suehiro, D. Harneti, R. Maharani, U. Supratman, F. F. Abdullah, S. Salam, T. Koseki and Y. Shiono, Chem. Biodiversity, 2021, 18, 1–14 Search PubMed.
- J. C. Frisvad, B. Andersen and U. Thrane, Mycol. Res., 2008, 112, 231–240 CrossRef CAS PubMed.
- R. Y. Yang, C. Y. Li, Y. C. Lin, G. T. Peng, Z. G. She and S. N. Zhou, Bioorg. Med. Chem. Lett., 2006, 16, 4205–4208 CrossRef CAS PubMed.
- T. Kashima, K. Takahashi, H. Matsuura and K. Nabetay, Biosci., Biotechnol., Biochem., 2009, 73, 1118–1122 CrossRef CAS PubMed.
- T. Kashima, K. T. Akahashi, H. M. Atsuura and K. N. Abeta, Biosci., Biotechnol., Biochem., 2009, 73, 2522–2524 CrossRef CAS PubMed.
- H. Luo, S. Meng, Y. Deng, Z. Deng and H. Shi, Arch. Microbiol., 2023, 205, 140 CrossRef CAS PubMed.
- Q. Yang, M. Asai, H. Matsuura and T. Yoshihara, Phytochemistry, 2000, 54, 489–494 CrossRef CAS PubMed.
- H. Matsuura, K. Nakamori, E. A. Omer, C. Hatakeyama, T. Yoshihara and A. Ichihara, Phytochemistry, 1998, 49, 579–584 CrossRef CAS.
- P. Li, K. Takahashi, H. Matsuura and T. Yoshihara, Biosci., Biotechnol., Biochem., 2005, 69, 1610–1612 CrossRef CAS PubMed.
- P. Paranagama, S. Santhirasegaram, S. Wickramarachchi and R. N. Attanayake, Asian J. Chem., 2022, 10–14 Search PubMed.
- S. Sultan, L. Sun, J. W. Blunt, A. L. J. Cole, M. H. G. Munro, K. Ramasamy and J.-F. F. Weber, Tetrahedron Lett., 2014, 55, 453–455 CrossRef CAS.
- W. Seemakram, J. Paluka, T. Suebrasri, C. Lapjit, S. Kanokmedhakul, F. T. Schevenels and S. Boonlue, Rhizosphere, 2024, 29, 100853 CrossRef.
- J. Schrader and J. Bohlmann, Biotechnology of Isoprenoids, Springer International Publishing, 2015 Search PubMed.
- D. A. T. Boncan, S. S. K. Tsang, C. Li, I. H. T. Lee, H. Lam, T. Chan and J. H. L. Hui, Int. J. Mol. Sci., 2020, 21, 7382 CrossRef CAS PubMed.
- P. M. Dewick, Medicinal Natural Products – A Biosynthetic Approach, John Wiley & Sons, 3rd edn, 2009 Search PubMed.
- E. Maser and T. Lanišnik Rižner, J. Steroid Biochem. Mol. Biol., 2012, 129, 1–3 CrossRef CAS PubMed.
- T. J. Cole, K. L. Short and S. B. Hooper, Semin. Fetal Neonatal Med., 2019, 24, 170–175 CrossRef PubMed.
- S. Purbaya, D. Harneti, A. P. Wulandari, Y. Mulyani, A. Azhari, A. P. Sari and U. Supratman, Molekul, 2023, 18, 396–403 CrossRef CAS.
- H. L. Choy, E. A. Gaylord and T. L. Doering, mBio, 2023, 14, e0135323 CrossRef PubMed.
- Y. Liang, M. Zhang, M. Yu, J. Wang, H. Zhu and C. Chen, Tetrahedron Lett., 2020, 61, 151737 CrossRef CAS.
- Y. Liang, L. Li, Y. Shen, Y. Zheng, Q. Li, Q. Tong, Q. Zhou, X. N. Li, D. Li, H. Zhu, W. Sun, C. Chen and Y. Zhang, Phytochemistry, 2022, 201, 113248 CrossRef CAS PubMed.
- E. N. Morrison, S. Knowles, A. Hayward, R. Greg Thorn, B. J. Saville and R. J. N. Emery, Mycologia, 2015, 107, 245–257 CrossRef CAS PubMed.
- A. M. Ashby, in Physiological and Molecular Plant Pathology, 2000, pp. 147–158 Search PubMed.
- J. M. Valayil and C. Jayabaskaran, Eur. J. Exp. Biol., 2016, 5, 28–36 Search PubMed.
- M. J. Valayil, G. C. Kuriakose, C. Jayabaskaran, C. Kuriakose and C. G. Jayabaskaran, Anticancer Agents Med. Chem., 2016, 16, 865–874 CrossRef PubMed.
- A. Hartanto, E. Munir, M. Basyuni, M. N. Saleh, L. D. S. Hastuti, Y. Yurnaliza, K. Nurtjahja and A. Lutfia, Rasayan J. Chem., 2023, 16, 182–187 CrossRef CAS.
- M. Pandi, R. Manikandan and J. Muthumary, Biomed. Pharmacother., 2010, 64, 48–53 CrossRef CAS PubMed.
- M. Pandi, R. S. Kumaran, Y. K. Choi, H. J. Kim and J. Muthumary, Afr. J. Biotechnol., 2011, 10, 1428–1435 CAS.
- R. Venkatachalam, K. Subban and M. J. Paul, J. Biotechnol., 2008, 136, S189–S190 CrossRef.
- A. Andolfi, L. Mugnai, J. Luque, G. Surico, A. Cimmino and A. Evidente, Toxins, 2011, 3, 1569–1605 CrossRef CAS PubMed.
- C. Félix, R. Meneses, M. F. M. Gonçalves, L. Tilleman, A. S. Duarte, J. V. Jorrín-Novo, Y. Van de Peer, D. Deforce, F. Van Nieuwerburgh, A. C. Esteves and A. Alves, Sci. Rep., 2019, 9, 1–12 Search PubMed.
- P. M. Dewick, Medicinal Natural Products: A Biosynthetic Approach, John Wiley & Sons, Hoboken, NJ, USA, 3rd edn, 2016 Search PubMed.
- P. Li, R. Takei, K. Takahashi and K. Nabeta, Phytochemistry, 2007, 68, 819–823 CrossRef CAS PubMed.
- K. Tsukada, K. Takahashi and K. Nabeta, Phytochemistry, 2010, 71, 2019–2023 CrossRef CAS PubMed.
- C. D. Qian, Y. H. Fu, F. S. Jiang, Z. H. Xu, D. Q. Cheng, B. Ding, C. X. Gao and Z. S. Ding, BMC Microbiol., 2014, 14, 297 CrossRef PubMed.
- H. Arimoto, T. Shimano and D. Uemura, J. Agric. Food Chem., 2005, 53, 3863–3866 CrossRef CAS PubMed.
- M. T. Barros, C. D. Maycock and M. R. Ventura, Chem.–Eur. J., 2000, 6, 3991–3996 CrossRef CAS PubMed.
- T. Kamikubo and K. Ogasawara, Tetrahedron Lett., 1995, 36, 1685–1688 CrossRef CAS.
- M. Fink, H. Gaier and H. Gerlacho, Helv. Chim. Acta, 1982, 65, 2563–2569 CrossRef CAS.
- M. Braun, U. Mahler, R. Botrysdiplodia and I. Kulturfiltraten, Liebigs Ann. Chem., 1990, 6, 513–517 CrossRef.
- F. Bracher and B. Schulte, J. Chem. Soc., Perkin Trans. 1, 1996, 2619–2622 RSC.
- G. Solladié, A. Rubio, M. C. Carreño and J. L. García Ruano, Tetrahedron: Asymmetry, 1990, 1, 187–198 CrossRef.
- Y. Huang, A. J. Minnaard and B. L. Feringa, Synthesis, 2011, 1055–1058 Search PubMed.
- J. M. Madrigal Lombera and I. B. Seiple, Tetrahedron Lett., 2024, 147, 1–4 CrossRef PubMed.
- C. Jiang, R. Zhou, J. Gong, L. Chen, T. Kurtán, X. Shen and Y. Guo, Bioorg. Med. Chem. Lett., 2011, 21, 1171–1175 CrossRef CAS PubMed.
- S. K. Dey, A. Rahman and A. Alghamdi, Eur. J. Org. Chem., 2016, 1684–1692 Search PubMed.
- M. Zhang, M. Niu, J. Fan, Z. Lu, Z. Zhu, B. Gao and S. P. Shi, Fitoterapia, 2025, 180, 106296 CrossRef CAS PubMed.
- X. Liu, S. Li, X. Wei, Y. Zhao, D. Lai, L. Zhou and M. Wang, RSC Adv., 2020, 10, 1588–1594 RSC.
- S. Chi and C. H. Heathcock, Org. Lett., 2000, 2, 4103–4105 CrossRef PubMed.
- A. G. M. Barrett, F. Blaney, A. D. Campbell, D. Hamprecht, T. Meyer, A. J. P. White, D. Witty and D. J. Williams, J. Org. Chem., 2002, 67, 2735–2750 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |