 Open Access Article
Open Access Articleβ-Lactam antibiotics and β-lactamases: historical perspectives and a review of β-lactamase inhibitors derived from natural products
Phanankosi Moyo†
a,
Ikhane O. Albert†
b,
Neo Hlungwanic,
Thulani Sibandad,
Perfoy Lumub,
Ndivhuwo Kevin Khorommbic,
Gurleen Kaure,
Karina Calvopina Tapiae,
George Siegwartf,
Nana Kwaku Buabengg,
Cynthia A. Danquah h,
Lyndy J. McGaw
i,
Sekelwa Cosa
b,
Christopher J. Schofield
h,
Lyndy J. McGaw
i,
Sekelwa Cosa
b,
Christopher J. Schofield
 e and
Vinesh J. Maharaj
e and
Vinesh J. Maharaj
 *c
*c
aDepartment of Plant and Soil Sciences, Faculty of Natural and Agricultural Sciences, University of Pretoria, Pretoria, Gauteng, South Africa
bDepartment of Biochemistry, Genetics and Microbiology, University of Pretoria, Hatfield, South Africa
cBiodiscovery Centre, Department of Chemistry, University of Pretoria, Pretoria 0028, South Africa. E-mail: vinesh.maharaj@up.ac.za; Tel: +27 824665466
dDepartment of Biology, National University of Lesotho, Maseru, Lesotho
eChemistry Research Laboratory, Department of Chemistry, The Ineos Oxford Institute for Antimicrobial Research, University of Oxford, Oxford, UK
fSir William Dunn School of Pathology, The Ineos Oxford Institute for Antimicrobial Research, University of Oxford, Oxford, UK
gUniversity of North-Texas Health at Fort Worth, USA
hDepartment of Pharmacology, Faculty of Pharmacy and Pharmaceutical Sciences, College of Health Sciences, Kwame Nkrumah University of Science and Technology, PMB, Kumasi, Ghana
iPhytomedicine Programme, Department of Paraclinical Sciences, Faculty of Veterinary Science, University of Pretoria, Private Bag X04, Onderstepoort, Pretoria 0110, South Africa
First published on 23rd April 2026
Abstract
Covering: up to the end of November 2025
The history of β-lactam antibiotics and, subsequently, β-lactamase inhibitors highlight the indispensable role of natural products in modern medicine. The isolation and testing of β-lactam bearing natural products led to world-changing therapeutic breakthroughs, yielding efficacious, safe, and cost-effective medicines that are still widely used today. The scientific triumph of β-lactams was, however, followed by a period of reduced research into new antibiotics, during which time bacterial pathogens acquired multiple mechanisms of resistance. The available evidence suggests that this situation is not irretrievable – whereas insufficient medicinal chemistry is currently being carried out to enable a renaissance in antibiotic development, our molecular understanding of antimicrobial modes of action and resistance mechanisms has improved dramatically. The history of β-lactams and many other drug classes suggests that natural product-based approaches will be critical in the future. Here, we summarise the history of β-lactams, focusing on natural product science. We then examine historical and recent efforts to identify new types of β-lactamase inhibitors using natural product-based screening approaches. We conclude by providing a perspective on how we can most efficiently discover β-lactamase inhibitors from microbial and plant-derived natural products.
1. Introduction
The rise of bacterial antimicrobial resistance (AMR) presents major global health and economic challenges.1–7 According to estimates for 2021, bacterial AMR was directly responsible for ∼1.14 million fatalities and was associated with an additional ∼4.71 million fatalities globally. Amongst the World Health Organisation regions, the South Asian and sub-Saharan African regions were the most affected, accounting for ∼47% of global fatalities due to bacterial AMR in 2021.3 If left unchecked, AMR is predicted to cause 8.2 million global fatalities annually by 2050.3 In addition to its toll in human lives, the financial costs of AMR are substantial. In China, the economic cost attributed to AMR was estimated to be US$77 billion in 2017, while the United States of America is reported to be losing US$55 billion annually due to AMR.4,6,7 The World Bank projects that AMR will lead to a 1.1–3.8% reduction in annual global gross domestic product by 2050.5 The adverse economic effects of AMR will most likely disproportionately impact low-to middle-income countries, further widening the inequality gap between developed and developing countries.The increase in antibiotic resistance rates over time has been a major driver of the current AMR crisis.8 Of particular concern are the ESKAPE pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and Enterobacter species), which, along with Mycobacterium tuberculosis, present a significant public health challenge.3,9,10 The ESKAPE pathogens now possess a plethora of intrinsic and acquired mechanisms that enable resistance to multiple antibiotic classes. The increasing resistance to carbapenems (e.g., imipenem, meropenem, tebipenem, and ertapenem), which have often been considered the last line of defence against infections caused by multidrug-resistant pathogens, highlights the AMR crisis. Carbapenem-resistant strains of K. pneumoniae, A. baumannii, and P. aeruginosa are now widespread globally.11 Many ESKAPE pathogens have also developed resistance to extended-spectrum cephalosporins, fluoroquinolones, and aminoglycosides.3 The pan-resistance of many ESKAPE pathogens contributes significantly to AMR by substantially limiting treatment options against them, compromising infection management, and increasing the risk of morbidity and mortality.3,9,10 The severity of the AMR threat is evident: in 2021, an estimated 683![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 deaths were attributed to infections caused by antibiotic-resistant ESKAPE pathogens alone.3
000 deaths were attributed to infections caused by antibiotic-resistant ESKAPE pathogens alone.3
Since the mid-20th century, when the major antibiotic classes were discovered, there has been a dearth in the discovery of new antibiotic classes.10,12 Contributing factors to this lack of innovation include scientific and technical challenges, limited economic incentives, the efficacy and low costs of existing drugs, and regulatory hurdles.1,13 The recent lack of new antibiotic classes has exacerbated the global AMR crisis, underscoring the need for a multifaceted solution that extends beyond developing novel antibiotics to include agents that can reverse existing drug resistance. The efficacy of agents targeting the underlying mechanisms of resistance is demonstrated by pioneering 20th-century research that led to β-lactamase inhibitors that protect β-lactam antibiotics, an approach that has the potential to be more widely employed.14 There is, however, a need for a deeper understanding of both direct and indirect resistance mechanisms to develop new medicines best suited to address the challenge of AMR.
The development and spread of antibiotic resistance results from multiple processes operating at different levels. It is important to distinguish between intrinsic insensitivity to antibiotics, wherein certain bacteria are naturally shielded from antibiotics, for example due to innate cell wall impermeability, and acquired resistance mechanisms that emerge under selective pressure. At the cellular level four major direct mechanisms of acquired resistance have been identified: limiting drug uptake, modifying drug targets, drug efflux, and drug inactivation.15,16 In Gram-negative Enterobacter species, the reduced uptake of imipenem and cephalosporins is achieved by changes in the structures and numbers of porin channels in the outer-membrane, thereby limiting antibiotic entry into cells.15,16 In Gram-positive bacteria, target modification is a widely used mechanism of drug resistance. For example, mutations or changes in the number of penicillin-binding proteins (PBP) reduce the effectiveness of β-lactam antibiotics. Similarly, drugs that disrupt nucleic acid synthesis, such as fluoroquinolones, are rendered ineffective by target modification.9,15,16 Most bacteria intrinsically encode (or can acquire) genes for efflux pumps that expel toxic substances out of cells, including antibiotics. Lastly, bacteria can acquire resistance by inactivating drugs via their enzyme catalysed degradation or modification. The latter mechanism is employed by a broad spectrum of bacteria, most famously exemplified by the β-lactamases that inactivate β-lactams by catalysing their hydrolysis, yielding inactive β-amino acids.16 Enzymes that degrade other classes of antibiotics, including tetracyclines, colistin, and fosfomycin, have been identified.17–20
Amongst the available antibiotic classes, the β-lactams are arguably the most important currently in use, constituting approximately 60–65% of the total antibiotic market.21 Increasing resistance to β-lactams, primarily due to β-lactamase activity, poses a critical challenge to these once-considered miracle drugs.3 To combat β-lactamase-mediated resistance, one established strategy is to use compounds that inhibit β-lactamases. Natural products are a reliable source of lead compounds for drug discovery,22 including clinically relevant β-lactamase inhibitors. This statement is strikingly exemplified by the discovery and development of the bacteria-derived compound clavulanic acid ∼50 years ago. Use of clavulanic acid in combination with a β-lactam antibiotic, in particular amoxicillin, is still increasing.22
In this review, we provide a perspective on β-lactams and β-lactamases, with an emphasis on the past and likely future roles of natural product science in both discovering new antibiotics and combating their emerging resistance. We conclude by offering thoughts on future avenues to find novel β-lactamase inhibitors from natural product sources.
2. β-Lactams and β-lactamases
2.1 Discovery of β-lactams
The serendipitous observation of the antibacterial activity of penicillin(s) by Alexander Fleming at St. Mary's Hospital, London, heralded the dawn of the modern antibiotic era (Fig. 1 and 2). Fleming observed that alcohol extracts of Penicillium mould possessed antibacterial activity against Staphylococcus cultures.23 Fleming, however, failed to isolate the active ingredient from the crude extracts, a challenge subsequently addressed in part by Ernst Chain, Guy Newton, Edward Abraham and others working at the University of Oxford in the 1940s. Abraham's proposal that the core penicillin structure comprised a 4-membered cyclic amide fused to a thiazolidine ring was subsequently validated by crystallography performed by Dorothy Crowfoot Hodgkin (Fig. 1).24 | ||
| Fig. 1 Structural classes of clinically used β-lactams and their interaction with penicillin-binding proteins (PBP). (a) Core structures of the major β-lactam classes: penams, penems, cephems, carbapenems, and monobactams. (b) Mechanistic comparison of PBP-mediated transpeptidation and inhibition: (i) penicillin-binding proteins catalyse the reaction of a C-terminal D-Ala–D-Ala unit with a PBP nucleophilic serine to produce D-Ala and an acyl–enzyme complex, which undergoes cross-linking with a Lys-sidechain; (ii) β-lactams react with the nucleophilic serine of the PBPs, forming a stable acyl–enzyme complex that undergoes slow hydrolysis, rendering it functionally irreversible. Nucleophilic serine-β-lactamases employ a related mechanism, though their acyl–enzyme complexes normally undergo efficient hydrolysis (see Fig. 7). | ||
Clinical trials using partially purified penicillin began in late 1941, though were hampered by the low quality of (impure) penicillin obtained at the time. The apparent low toxicity of the purified molecule(s) encouraged further trials. By 1945, large-scale production of penicillin in the United States of America and elsewhere had begun, primarily through batch fermentation involving Penicillium chrysogenum; the addition of corn-steep liquor resulted in the production of the breakthrough drug benzylpenicillin (penicillin G),25,26 which was widely used across war-torn Europe.27 Notwithstanding the success of penicillin, bacterial resistance emerged in the late 1940s (Fig. 2). Soon thereafter, Chain and Abraham discovered the first β-lactamase and purified the enzyme from E. coli, highlighting the role of enzymes in mediating antibiotic resistance.28 By the early 1950s, clinical reports began to emerge on streptococcal resistance to penicillin, prompting research into β-lactamase-stable penicillin analogues as well as new classes of β-lactams and β-lactamase inhibitors.29
In the quest for new antibiotics to combat resistance, the discovery of cephalosporins was a significant breakthrough. Giuseppe Brotzu, at the Hygiene Institute in Cagliari, Italy, isolated an antibiotic-producing strain of Cephalosporium acremonium from seawater near the outlet of a sewage pipe.30 Importantly, Brotzu demonstrated the antibacterial activity of fermentation extracts of C. acremonium in patients with a variety of infections, including typhoid fever, where penicillin had failed. Brotzu sent his strain to Oxford, where Abraham and Newton purified two β-lactams: the first was a penicillin and the second was cephalosporin C (Fig. 2 and 3).31,32 They assigned the structure of cephalosporin C through chemical analysis,31,33 a proposal validated through X-ray crystallography in 1961.34 This work marked the first report of a natural β-lactam with a fused ring structure different from that of the penicillins: in cephalosporins, the β-lactam is fused to a six-membered dihydrothiazine ring (Fig. 1).
 | ||
| Fig. 3 Evolution of cephalosporins. (a) Development of cephalosporins from natural cephalosporin C, which is converted to 7-aminocephalosporinic acid (7-ACA), a precursor for semisynthesis. (b) 1st–5th generation semisynthetic derivatives with expanding Gram-negative coverage and improved β-lactamase stability. (i) 1st generation cephalosporins: spectrum of activity – excellent Gram-positive activity, but limited activity against Gram-negative bacteria; β-lactamase stability – poor stability against extended serine-β-lactamases (ESBLs) and many other β-lactamases. (ii) 2nd generation cephalosporins: spectrum of activity – improved Gram-negative activity; β-lactamase stability – increased stability against some β-lactamases. (iii) 3rd generation cephalosporins: spectrum of activity – enhanced Gram-negative and reduced Gram-positive activity with some central nervous system penetration; β-lactamase stability – stable against many serine-β-lactamases but not ESBLs or AmpC. (iv) 4th generation cephalosporins: spectrum of activity – broader Gram-negative and Gram-positive activity; β-lactamase stability – enhanced penetration and stability against AmpC and many ESBLs. (v) 5th generation cephalosporins: spectrum of activity – retains broad Gram-negative coverage and is the first cephalosporin active against methicillin-resistant S. aureus; β-lactamase stability – some activity against ESBL producers, however, not stable against all carbapenemases.42 (c) Cefiderocol is a recently developed siderophore-bearing broad-spectrum cephalosporin.43,44 Cefiderocol possesses a cephalosporin core with the C7 sidechain of ceftazidime and a C3′ sidechain, related to that of cefepime, but modified by conjugation to a catechol.43 The catechol group enables cefiderocol to exploit the iron/siderophore transport system to penetrate the bacterial outer membrane and reach the periplasmic space of many Gram-negative bacteria.45 | ||
Although displaying less pronounced activity than some penicillins, cephalosporin C was found to be relatively stable to contemporary serine-β-lactamases, consistent with Brotzu's microbiological observations.35,36 Subsequently, the core scaffold of the cephalosporins, 7-aminocephalosporanic acid, was prepared from cephalosporin C (Fig. 3),37,38 thereby enabling the development of semisynthetic cephalosporins with modified C7 sidechains. There are two positions in the cephalosporin C molecule from which extensive modifications have been made, namely the C7 amide derivatives (prepared via C7 amines) and variation of the C3′ group. The production of many semisynthetic derivatives over the years has led to the development of the different generations of cephalosporins (Fig. 3).39,40 The most recently approved cephalosporin is cefiderocol (approved in 2019), for which initial resistance has, unfortunately, already been reported in carbapenem-resistant Gram-negative pathogens.41 Overall, the semisynthesis approach with cephalosporins effectively demonstrates the power of combining natural product science, medicinal chemistry, and clinically relevant microbiology to combat AMR.
Although new antibiotic classes emerged (Fig. 2), the penicillins and cephalosporins remained the most important antibiotic classes through the 1960s and early 1970s. It was during this period that insight into the mechanism of action of penicillins emerged. Tipper and Strominger proposed that penicillin inhibits bacterial cell wall biosynthesis by covalently binding to the nucleophilic active site serine residue of bacterial transpeptidases, commonly referred to as PBPs. Penicillin was proposed to structurally mimic the D-Ala–D-Ala moiety of peptidoglycan precursors; however, unlike the acyl–enzyme complex derived from the reaction of the transpeptidase nucleophilic serine with D-Ala–D-Ala, the complex formed from the reaction of penicillin is inert to transpeptidation (it can undergo slow hydrolysis), thus preventing crosslinking of peptidoglycans.46,47 Such inhibition of transpeptidase compromises the integrity of bacterial cell walls, leading to osmotic lysis (Fig. 1).
By the late 1960s, researchers at Beecham Pharmaceuticals had identified a series of β-lactamase inhibitors and, at least partially, β-lactamase-resistant β-lactams. Amongst these were the olivanic acids, a type of carbapenem, which were isolated from fermentation broths of Streptomyces olivaceus.48 It is important to note that by this time, it had been revealed that multiple bacteria (and fungi) produce multiple β-lactams,49,50 especially those of the Streptomyces genus.51 About the same time, progress on β-lactamase-resistant β-lactams was being made by the Merck group, which was screening fermentation broths of Streptomyces cattleya for inhibitors of cell wall biosynthesis, leading to the discovery of several representatives of the thienamycin family of carbapenems (Fig. 1, 2, and 4).52 The carbapenems were shown to contain a β-lactam ring fused to an unsaturated five-membered ring in which a CH2 group replaces the sulphur present in the thiazolidine ring of penicillin (Fig. 1 and 4).53 The carbapenems also differ from penicillin and cephalosporins in the nature and stereochemistry of the sidechains linked to the β-lactam ring, in particular by the replacement of the C6/C7 penicillin amides with the C6 hydroxyethyl group of the clinically used carbapenems.54
Thienamycin was found to display exceptionally high in vitro activity against a wide range of bacteria, including some strains of P. aeruginosa and, importantly, manifested substantial resistance to hydrolysis by contemporary serine-β-lactamases.55 However, its instability both in vitro and in vivo rendered thienamycin unsuitable for clinical use.56 To improve on thienamycin, chemists at Merck synthesised an N-formimidoyl derivative (imipenem) with improved in vitro and in vivo shelf life (Fig. 4). However, like all studied natural carbapenems, imipenem is a substrate for renal dehydropeptidase (DHP-I) which limited its in vivo activity.57 Cilastatin is a competitive inhibitor of DHP-I; in 1985, a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 combination of imipenem and cilastatin was launched under the brand name Primaxin®.58
1 combination of imipenem and cilastatin was launched under the brand name Primaxin®.58
Subsequent work at Merck revealed that synthetic carbapenems with a 1-β-methyl substituent were stable towards DHP-I and had a potency similar to that of imipenem, paving the way for a second generation of semisynthetic carbapenems such as meropenem and ertapenem (Fig. 5).59 The carbapenems became a commonly used last line of defence against bacterial infections where other β-lactams had failed. By the late 1990s, a few reports of carbapenem resistance in K. pneumoniae emerged; however, it was not until the 2000s that reports of widespread, major carbapenem resistance emerged, involving different Gram-negative bacteria (Fig. 3).60,61
The search for β-lactamase-resistant β-lactam compounds led to the discovery of another important β-lactam class, the monocyclic monobactams (Fig. 1, 2 and 5). The nocardicins were the first monocyclic β-lactam antibiotics to be discovered, being identified from fermentation broths of Nocardia uniformis subsp. tsuyamanensis by the Takeda company.62 Subsequently, the monobactams were isolated from Agrobacterium radiobacter and characterised by researchers at Squibb.63 They were found to inhibit certain nucleophilic serine-β-lactamases and were subsequently found to be surprisingly stable against the class B metallo-β-lactamases. The efficiency and low toxicity of the monobactams, in particular aztreonam, means that they are now widely used in human and animal medicine.64 However, as previously alluded to, bacterial pathogens have responded to the challenges posed by β-lactams by evolving β-lactamases that circumvent their lethality, including those targeting monobactams.
Like the carbapenems, the clinically used monobactams, of which aztreonam is the most important, are produced by total synthesis rather than direct fermentation or semisynthesis, as is the case for penicillins and cephalosporins. The carbapenem and monobactam classes of β-lactams thus serve as excellent examples of how natural product structures can inspire the development of synthetic derivatives suited for clinical use.
2.2 Discovery of β-lactamases
Although other β-lactam resistance mechanisms (e.g., efflux pumps, PBP modification) are important, from a clinical perspective, the most prevalent response to date by bacteria to β-lactam antibiotic pressure is likely through the production of β-lactamases (Fig. 6).65,66 Following the pioneering work of Chain and Abraham,28 >2000 unique β-lactamases have been identified (Fig. 6).67 β-Lactamases are localised within the periplasmic space,46,68 though some β-lactamases act extracellularly.69,70 | ||
| Fig. 6 Historical timeline showing milestones in the discovery of β-lactamases and β-lactamase inhibitors. Note that not all the β-lactamase inhibitors are β-lactams. | ||
β-Lactamases are classified using two principal systems, namely (i) the activity-based Bush–Jacoby–Medeiros system and (ii) the Ambler system.71 The Ambler classification system is the most widely adopted, including in this review. The Ambler system categorises β-lactamases into four classes, A, B, C, and D (Table 1), primarily based on specific amino acid sequences and their β-lactam hydrolysis mechanisms.72,73 The Ambler class A, C, and D β-lactamases employ a nucleophilic serine residue in catalysis and are hence termed serine-β-lactamases. By contrast, the Ambler class B β-lactamases are metallo-hydrolases and hence are termed metallo-β-lactamases.73 Serine-β-lactamases are structurally related to the PBPs and share a Ser–xxx–xxx–Lys active site motif.74 β-Lactam inactivation by serine-β-lactamases proceeds via nucleophilic attack by the serine residue, forming an acyl-enzyme (ester) intermediate, which undergoes hydrolysis. In contrast, metallo-β-lactamases instead utilise a metal-activated (likely Zn2+) water nucleophile to drive the hydrolytic reaction (Fig. 7).75
| Class | Mechanism | Clinically relevant example | Clinically relevant inhibitorsa |
|---|---|---|---|
| a Note that in most, but not all, cases β-lactamase inhibitors have no or little intrinsic antibacterial activity, with carbapenems being both antibiotics and inhibitors of some serine-β-lactamases. | |||
| Class A | Serine-β-lactamase | KPC, CTX-M, SHV, and TEM | Clavulanic acid, sulbactam, tazobactam, avibactam, relebactam, and vaborbactam |
| Class B | Metallo-β-lactamase | NDM, VIM, and IMP | None |
| Class C | Serine-β-lactamase | AmpC and CMY | Avibactam |
| Class D | Serine-β-lactamase | OXA-10, OXA-23, and OXA-48 | Avibactam and relebactam |
The active site of class A serine-β-lactamases is bordered by an α-helical domain, an α/β domain, and an Ω-loop.78 The hydrolytic activity of class A serine-β-lactamases ranges from narrow-spectrum to broad-spectrum and includes carbapenem-hydrolysing activity, making this class particularly concerning.79 Serine-β-lactamase catalysis proceeds via a canonical two-step acylation–deacylation process, which involves the nucleophilic serine (Ser70), a hydrolytic water, and an active site lysine (Lys73) and glutamic acid (Glu166) residues, the latter of which act as the general acid/base machinery for catalysis.65 Class A serine-β-lactamases such as TEM and SHV can be potently inhibited by the ‘classical’ serine-β-lactamase inhibitors, namely clavulanic acid, tazobactam, and sulbactam.80,81 Other class A serine-β-lactamases, especially the highly evolved and disseminated KPCs, evade such inhibitors. These, however, are susceptible to new generation serine-β-lactamase inhibitors such as avibactam, relebactam, and vaborbactam.82,83
Class C β-lactamases mainly comprise chromosomally encoded cephalosporinases and are widespread in Gram-negative bacteria. The best characterised class C serine-β-lactamases are the AmpC enzymes from Enterobacteriaceae, whose expression is induced by β-lactams. Class C β-lactamases are widely disseminated across the chromosomes of bacterial pathogens, including Citrobacter freundii (CMY enzymes) and A. baumannii (ADC enzymes).84–87 Class C serine-β-lactamases also occur in plasmids harboured, inter alia, in K. pneumoniae (ACT-1) and Salmonella Enteritidis (DHA-1). Their three-dimensional structures are related to class A serine-β-lactamases, and they follow the two-stage acylation–deacylation mechanism. Unlike the class A active sites, those of the class C enzymes appear more open, and the general acid/base residues differ, involving Tyr150 and Lys67.65,87
Resistance mediated by the class D serine-β-lactamases, also known as oxacillinases (OXAs), is particularly concerning, especially with respect to carbapenem (and cephalosporin) resistance in Gram-negative bacteria, including A. baumannii and Enterobacteriaceae.88,89 From a mechanistic perspective, the OXAs differ from the class A and C serine-β-lactamases in that they employ a carboxylated lysine residue as the general base for the acylation and deacylation steps. Additionally, they also appear to have a greater number of hydrophobic residues at their active sites, which distinguishes them structurally to class A and C serine-β-lactamases.65 The OXAs are both chromosomally and plasmid encoded, with the OXA-48 group being reported in multiple plasmids and in some Gram-positive pathogens.90 They are also strongly resistant to inhibitory molecules. This is especially critical, as the increasing prevalence of OXAs amongst opportunistic pathogens renders clinical therapies ineffective.91 There is an urgent need to develop effective inhibitors of class D serine-β-lactamases.
Metallo-β-lactamases are characterised by a His/Asn116–xxx117–His118–xxx119–Asp120–His/xxx121 motif that forms a metal-binding centre located at the interface of the two β-sheets that comprise the core of the protein.97 Class B1 metallo-β-lactamases (NDM, VIM, IMP) possess a binuclear Zn2+ centre made up of tri-His (3H) (Zn2+1) and Cys–His–Asp (Zn2+2) metal-binding sites. The two Zn2+ ions at the metal sites are bridged by the ‘hydrolytic’ water molecule/hydroxide ion, with an additional water molecule bound to Zn2+2.
In the presently small B2 metallo-β-lactamase sub-group, which utilises one Zn2+ ion for catalysis, the first conserved His of the defining motif is replaced by an Asn. This substitution disrupts coordination at one of the two potential Zn2+ binding sites, resulting in binding of only a single metal ion. The B3 metallo-β-lactamases, like the B1 metallo-β-lactamases, utilise two Zn2+ ions. However, in the B3 metallo-β-lactamases, the Zn2+2 coordinating Cys residue is replaced by another residue, which varies amongst B3 enzymes, with a His121 acting as the metal co-ordinating residue in some metallo-β-lactamases.65,96,98,99
The differences in structures and detailed mechanisms of the metallo-β-lactamases is striking. Moreover, these enzymes are resistant to all currently available serine-β-lactamase inhibitors,95 something that is a cause for serious concern. A positive note, however, is that bicyclic boronates, including compounds in clinical development, inhibit both serine-β-lactamases and, albeit less efficiently, metallo-β-lactamases.96,97
The history of β-lactams offers a clear example of how the discovery of a groundbreaking new therapy is followed by a period of intense effort and productivity, leading to multiple widely used medicines. This remarkably productive period was followed by a phase of reduced innovation in antibiotic research, during which time bacterial pathogens acquired multiple resistance mechanisms.99–102 The available evidence suggests that this situation is not irretrievable. Although relatively little recent medicinal chemistry has been applied to the rational improvement of β-lactams, with the work of Shionogi in Japan being an exception, our molecular understanding of resistance has improved dramatically as a consequence of new methods.81,97,98,103–108 The history of β-lactams as outlined above suggests that natural product-based approaches will be critical in the future. Below, we summarise work, both historical and recent studies aimed towards the identification of new types of β-lactamase inhibitors, beginning with an overview of the evolution of screening approaches used in the discovery of natural product-based inhibitors.
In parallel with the industry efforts at Beecham Pharmaceuticals, academic researchers at the Kitasato Institute in Japan, developed a systematic screening program to identify penicillinase inhibitors from microbial metabolites.110,111 This approach was significant as it represented an early example of a target-based screen, i.e. specifically isolating the enzyme of interest rather than relying solely on whole-cell interactions. By focusing on enzyme inhibition as a primary selection criterion, the Kitasato programme expanded the conceptual framework for inhibitor discovery and enabled the identification of microbial metabolites with direct β-lactamase inhibitory activity.110,111
Through the late 20th century and into the 21st century, screening approaches for β-lactamases inhibitors evolved, shifting towards high throughput screening. Leading this transition was Pfizer, which sought to efficiently discover novel antibiotics from both fermentation extracts and synthetic libraries in a cost-effective manner.112 This strategy has subsequently been adopted across both industry and academic research programs, in a manner enhanced by the availability of purified recombinant β-lactamases enabling screening against multiple targets.
As metallo-β-lactamases emerged as an increasing clinical threat, screening approaches required further adaptation to address the unique requirements of their zinc-dependent catalysis. In this context, academic researchers, have played pivotal roles in developing medium-to-high throughput phenotypic and target-based assays tailored to identifying metallo-β-lactamase inhibitors.113,114 Contributions from small companies and academics have been important at a time where the major pharmaceutical industry has largely exited the discovery space for new antibiotics and antibiotic potentiators.
β-Lactamase inhibitor screening has increasingly been informed by high-resolution structural and computational approaches.115–117 Techniques such as X-ray crystallography, complemented by molecular dynamics simulations, have enabled direct visualisation of enzyme–inhibitor complexes, providing detailed insights into binding modes, metal coordination, and conformational dynamics.115–117 This structure-guided paradigm supports rational inhibitor design, allowing for the systematic modification of molecular scaffolds to optimise binding affinity, pharmacokinetic properties, and stability against enzymatic degradation. These approaches collectively have been particularly instrumental, as exemplified by the discovery of vaborbactam118 and other cyclic boronates,119,120 as well as the identification of the aspergillomarasmine A113 and indole-2-carboxylate121 scaffolds as potent metallo-β-lactamase inhibitors.
In summary, β-lactamase inhibitor screening has evolved from foundational phenotypic assays into a sophisticated, often structure-guided, paradigm. Modern discovery pipelines now rely on integrated assay cascades that can systematically filter vast natural product libraries, moving from high-throughput primary screens to precise hit validation and rational optimisation. This structured approach efficiently manages the complexity of screening campaigns, allowing researchers to prioritise the most promising scaffolds for further development. Having summarised the methodological frameworks, the following section reviews the landmark and recent discoveries of natural product β-lactamase inhibitors identified from microbial and plant-based sources.
3. β-Lactamase inhibitors from natural products
Natural products, in particular secondary metabolites,122 have had a central role in the development of modern antibiotics and β-lactamase inhibitors. This section further describes the demonstrated and potential contribution of microbial and plant-derived natural products to the discovery of β-lactamase inhibitors (Tables 2 and 3).| Compound | Source organism | Target β-lactamase | IC50b (µM) | pIC50c |
|---|---|---|---|---|
a Plant species indicated as “Source organisms” that have been identified through literature searches as either the original sources from which the compounds were first isolated or as well-established producers of these compounds.b IC50 values in the table have been harmonised to a consistent unit, µM. Those previously reported as µg mL−1 were converted to µM units using the following formula 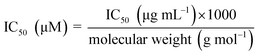 .c IC50 values were converted to pIC50 using the following formula: pIC50 = −log10(IC50[M]). .c IC50 values were converted to pIC50 using the following formula: pIC50 = −log10(IC50[M]). |
||||
| Clavulanic acid | S. clavuligerus | TEM-1 | 0.09 | 7.05 (ref. 123) |
| SHV-1 | 0.03 | 7.52 (ref. 123) | ||
| OXA-1 | 1.8 | 5.74 (ref. 123) | ||
| OXA-4 | 8.4 | 5.08 (ref. 123) | ||
| MM4550 | S. olivaceus | R-TEM | 0.001 | 9.00 (ref. 124) |
| MM13902 | S. olivaceus | R-TEM | 0.05 | 7.30 (ref. 124) |
| MM17880 | S. olivaceus | R-TEM | 0.03 | 7.52 (ref. 124) |
| SB236049 | C. funicola | B. cereus II | 0.7 | 6.15 (ref. 125) |
| CfiA | 2.0 | 5.70 (ref. 125) | ||
| IMP-1 | 151 | 3.82 (ref. 125) | ||
| SB236050 | C. funicola | B. cereus II | 256 | 3.59 (ref. 125) |
| SB238569 | C. funicola | B. cereus II | 19 | 4.72 (ref. 125) |
| AMA | A. versicolor | NDM-1 | 4.0 | 5.40 (ref. 113) |
| VIM-2 | 9.6 | 5.02 (ref. 113) | ||
| 9-HHIA | Aspergillus sp. | IMP-1 | 50.5 | 4.30 (ref. 126) |
| 10-HHIA | Aspergillus sp. | IMP-1 | 31.6 | 4.50 (ref. 126) |
| Emerione A | E. nidulans | NDM-1 | 12.1 | 4.92 (ref. 127) |
| Asperfunolone A | Aspergillus sp. | NDM-1 | 36.2 | 4.44 (ref. 127) |
| Holomycin | S. clavuligerus | NDM-1 (nitrocefin) | 0.153 | 6.81 (ref. 128) |
| NDM-1 (imipenem) | 0.11 | 6.96 (ref. 128) | ||
| Compound | Source organism | Target β-lactamase | IC50b (µM) | pIC50c |
|---|---|---|---|---|
a Plant species indicated as “Source organisms” that have been identified through literature searches as either the original sources from which the compounds were first isolated or as well-established producers of these compounds.b IC50 values in the table have been harmonised to a consistent unit, µM. Those previously reported as µg mL−1 were converted to µM units using the following formula 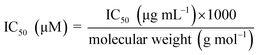 .c IC50 values were converted to pIC50 using the following formula: pIC50 = −log10(IC50[M]). .c IC50 values were converted to pIC50 using the following formula: pIC50 = −log10(IC50[M]). |
||||
| SB-202742 | S. mombin | TEM-1 | 13.5 | 4.87 (ref. 129) |
| OXA-1 | 215.25 | 3.67 (ref. 129) | ||
| P99 | 44.56 | 4.35 (ref. 129) | ||
| Rosmarinic acid | R. officinalis | VIM-2 | 41.5 | 4.38 (ref. 130) |
| Salvianolic acid A | S. miltiorrhiza | VIM-2 | 27.2 | 4.57 (ref. 130) |
| NDM-1 | 160.3 | 3.79 (ref. 130) | ||
| Salvianolic acid | S. miltiorrhiza | TEM-1 | 189 | 3.72 (ref. 130) |
| Hesperidin | Citrus sp. | NDM-1 | 3.35 | 5.48 (ref. 131) |
| Diosmin | Citrus sp. | NDM-1 | 20.7 | 4.68 (ref. 131) |
| Rutin | S. japonica | NDM-1 | 15.5 | 4.81 (ref. 131) |
| Stevioside | Stevia rebaudiana | NDM-1 | 214.1 | 3.67 (ref. 131) |
| Magnolol | Magnolia officinalis | NDM-1 | 24.3 | 4.61 (ref. 132) |
| Pterostilbene | Pterocarpus sp. | NDM-1 | 60 | 4.22 (ref. 133) |
| Embelin | E. ribes | NDM-1 | 2.1 | 5.68 (ref. 134) |
| VIM-1 | ∼200 | 3.70 (ref. 134) | ||
| IMP-1 | ∼100 | 4.00 (ref. 134) | ||
| Carnosic acid | R. officinalis | NDM-1 | 27.07 | 4.57 (ref. 135) |
| α-Mangostin | G. mangostana | OXA-48 | 0.52 | 6.28 (ref. 136) |
| NDM-1 | 25.43 | 4.59 (ref. 136) | ||
| Quercetin | S. japonica | OXA-48 | 1.47 | 5.83 (ref. 137 and 138) |
| NDM-1 | 5–10 | 5.00–5.30 (ref. 137 and 138) | ||
| Fisetin | Fragaria × ananassa | OXA-48 | 0.47 | 6.33 (ref. 138) |
| Luteolin | Reseda luteola | OXA-48 | 0.55 | 6.26 (ref. 138) |
| 3′,4′,7-Trihydroxyflavone | Medicago sativa | OXA-48 | 1.89 | 5.72 (ref. 138) |
| Apigenin | Matricaria chamomilla | OXA-48 | 3.64 | 5.44 (ref. 138) |
| Kaempferol | Delphinium consolida | OXA-48 | 4.54 | 5.34 (ref. 138) |
| Isoquercitrin | Cercis canadensis | OXA-48 | 11.55 | 4.94 (ref. 138) |
| Chrysin | Camellia sinensis | OXA-48 | 108 | 3.97 (ref. 138) |
| Galangin | Alpinia officinarum | OXA-48 | 216 | 3.67 (ref. 138) |
| Taxifolin | Pseudotsuga taxifolia | OXA-48 | 3.22 | 5.49 (ref. 138) |
| Naringenin | Citrus × paradisi | OXA-48 | 68.1 | 4.17 (ref. 138) |
3.1 β-Lactamase inhibitors from microorganisms
An influential search for β-lactamase inhibitors was conducted by researchers at the Kitasato University in Japan during the early 1970s.110,111 Recognising penicillinases as a key driver of penicillin resistance, they developed an innovative screening method to identify microbial metabolites that inhibit these enzymes. Through systematic screening of bacterial and fungal filtrates, they identified four Streptomyces strains with penicillinase-inhibition activity, including Streptomyces gedanensis ATCC 4880, which produced a potent inhibitor designated KA-107. KA-107 was isolated and, whilst lacking intrinsic antibacterial activity, was found to synergistically restore the efficacy of penicillin G, ampicillin, and phenethicillin against a resistant S. aureus strain (FS-1277). Notably, growth inhibition was only observed when KA-107 was co-administered with a penicillin, supporting its mode of action as a β-lactamase inhibitor, a proposal supported by optical density-based growth curves, which demonstrated prolonged suppression of bacterial growth when both agents were present.110,111Shortly after the groundbreaking work in Japan, studies on clavulanic acid, isolated from fermentation broths of S. clavuligerus,139 were reported (as described in Section 2.2.3; Fig. 8a). Clavulanic acid belongs to the clavam family of β-lactams, which are characterised by a fused bicyclic β-lactam and oxazolidine ring system, with the (3R,5R) stereochemistry, which confers potent inhibition of some serine-β-lactamases, while exhibiting weak antibacterial activity. In contrast, clavams (such as clavaminic acid, a biosynthetic precursor of clavulanic acid) with the (3S,5S)-stereochemistry, can exhibit intrinsic antibacterial activity.140 The structure of clavulanic acid was elucidated using spectrophotometric methods and X-ray crystallography.141 Its discovery was revolutionary because at the time of its clinical introduction, there was no available agent that restored the activity of any antibiotic against drug-resistant bacteria.
 | ||
| Fig. 8 Structural classes and mechanisms of β-lactamase inhibitors. (a) (i) IC50 values (µM) of classical β-lactamase inhibitors, clavulanic acid and related compounds tazobactam and sulbactam, against selected class A (TEM and SHV variants) and class D (OXA-type) serine-β-lactamases (data are from Brown et al.139). Clavulanic acid and tazobactam exhibit potent inhibition against class A enzymes with sub-micromolar IC50 values. Tazobactam is somewhat more active than clavulanic acid for a few class D β-lactamases, notably the OXA-2 and OXA-5. Note sulbactam and tazobactam are not naturally occurring compounds and are prepared from penicillin precursors. (a) (ii) Schematic illustration of the mechanism of clavulanic acid against serine-β-lactamase: the initially formed acyl–enzyme complex either undergoes hydrolysis or fragmentation of its oxazolidine ring to give stable acyl–enzyme complexes, including E/Z-enamines (with trans/E-enamines being observed). (b) Structures of more recently developed non-β-lactam β-lactamase inhibitors. Representative structures of diazabicyclooctane inhibitors (avibactam, relebactam) and boronic acid-based inhibitors (vaborbactam, taniborbactam), the latter of which react with both serine-β-lactamases and metallo-β-lactamases to give complexes proposed to mimic the tetrahedral transition state(s). | ||
Clavulanic acid works as a “mechanism based” inhibitor of serine-β-lactamase: it initially reacts to form an acyl–enzyme complex, which can either undergo hydrolysis (turnover) or oxazolidine ring-opening, leading to stable acyl–enzyme complexes (Fig. 8a).142 Clavulanic acid was used clinically for the first time in combination with the penicillin antibiotic amoxicillin (marketed as Augmentin), thereby protecting the latter against β-lactamases.109 Clavulanic acid shows strong potency against the class A serine-β-lactamases such as TEM-1, TEM-2, and SHV-1, as well as some ESBLs, including TEM-3 and SHV-2 (Fig. 8a). It has mixed activity against class C and D serine-β-lactamases, and it is inactive against class B metallo-β-lactamases. Despite these limitations, clavulanic acid remains widely used in clinical practice. Moreover, it has served as a structural scaffold that inspired the development of other clinically important, mechanistically related β-lactamase inhibitors, namely, tazobactam and sulbactam, both of which are prepared by semisynthesis from penicillin precursors (Fig. 8a).123
More recently, new classes of synthetic serine-β-lactamase inhibitors have been developed that do not contain a β-lactam, but which react with the nucleophilic serine (Fig. 8b). Although synthetic, these inhibitors mimic the mechanisms used by natural products to block β-lactamases. Avibactam was the first member of the diazabicyclooctane class of reversibly reacting broad-spectrum serine-β-lactamase inhibitors to be approved for clinical use. Avibactam is clinically used in combination with the cephalosporin ceftazidime.143 Its development was groundbreaking because it demonstrated the viability of β-lactamase inhibition by a non-β-lactam scaffold. Variants of avibactam have been developed, some of which exhibit antibacterial activity.143 After a long development period, boronates have also emerged as clinically used inhibitors of serine-β-lactamases and, in some cases, metallo-β-lactamases (Fig. 8b). These compounds react with serine-β-lactamases and bind to metallo-β-lactamase to give complexes that mimic the high-energy tetrahedral intermediates during β-lactam catalysis.144,145 To date, vaborbactam, a monocyclic boronate, is the only clinically approved β-lactamase inhibitor of this class. However, bicyclic boronates, such as taniborbactam, with a broader spectrum of activity, will likely be approved in the near future.119,146
The olivanic acids, a carbapenem subclass, were discovered alongside work that led to the discovery of clavulanic acid.139 Three compounds, designated MM 4550, MM 13902, and MM 17880, were isolated from S. olivaceus ATCC 31126 (Fig. 9a)124,139 and share a common (5R),(6R),(8S)-stereochemistry. In in vitro antibacterial studies, all three compounds displayed marked activity against both Gram-positive and Gram-negative bacteria, including S. aureus, E. coli, K. aerogenes, Proteus spp., and P. aeruginosa, many of which were β-lactamase producers. MIC values ranged from 0.05 to 500 µg mL−1, depending on the strain and compound, with MM 13902 and MM 17880 consistently demonstrating lower MICs across several isolates. In β-lactamase inhibition assays, the olivanic acids inhibited a wide range of serine-β-lactamases in cell-free systems (with IC50 values ranging from 0.5 to 75 ng mL−1; with pIC50 values ranging from 6.72 to 9.01), supporting dual roles as antibiotics and serine-β-lactamase inhibitors. In these β-lactamase inhibitory assays, MM 4550 consistently emerged as the most potent inhibitor, particularly where penicillin was used as the substrate.124 Mechanistic studies have revealed that the carbapenems, including the olivanic acids, initially form a Δ2-pyrroline acyl–enzyme complex, which can either undergo hydrolysis or tautomerisation to give an epimeric Δ1-pyrroline form(s), which is inhibitory (Fig. 9a).147
 | ||
Fig. 9 Structures of microbial-derived olivanic acids, a carbapenem sub-type of serine-β-lactamase inhibitors, and the proposed outline mechanism of action of carbapenems. (a) (i) The olivanic acid compounds MM 4550, MM 13902, and MM 17880, which were isolated from S. olivaceus. MM 4550 was also isolated from S. fulvoviridis and named MC696-SY2-A. (a) (ii) Proposed pathway for inhibition of serine-β-lactamases by carbapenems. The acyl-enzyme initially forms as a Δ2-pyrroline, which can tautomerise (that is, the double bond migrates from C2![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C3 to C2 C3 to C2![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) to give a more stable Δ1-pyrroline (imine) complex, which is proposed to be the inhibited enzyme species.147 (a) (iii) Note that in the case of the OXA serine-β-lactamases (class D SBLs), deacylation of carbapenem-derived acyl–enzyme complexes to give tautomeric lactone products can occur.148,149 Not all potential products are shown. (b) Structure of the serine-β-lactamase inhibitor PS-5 isolated from a Streptomyces species. N) to give a more stable Δ1-pyrroline (imine) complex, which is proposed to be the inhibited enzyme species.147 (a) (iii) Note that in the case of the OXA serine-β-lactamases (class D SBLs), deacylation of carbapenem-derived acyl–enzyme complexes to give tautomeric lactone products can occur.148,149 Not all potential products are shown. (b) Structure of the serine-β-lactamase inhibitor PS-5 isolated from a Streptomyces species. | ||
Olivanic acids were also isolated from a different microbial species: Umezawa et al.150 discovered two β-lactamase inhibitors, MC696-SY2-A and MC696-SY2-B, produced by cultures of S. fulvoviridis MC696-SY2; the former of these was shown to be identical or very similar to MM 4550 (Fig. 9a).151 The compounds were identified using iodometric titration, an assay whereby β-lactamase-mediated hydrolysis of penicillin releases penicilloic acid, which reduces iodine to iodide, thereby decreasing the intensity of the characteristic blue starch–iodine complex.152
Okamura et al.153 isolated a carbapenem with a C6 ethyl group (PS-5) (Fig. 9b), in studies searching for antibiotics by screening filtrates from soil-isolated Streptomyces species. PS-5 was isolated and its structure was elucidated using spectroscopic methods.153 PS-5 demonstrated notable intrinsic antibacterial activity against S. aureus (MIC = 0.16 µg mL−1) and Diplococcus pneumoniae (MIC 0.02 µg mL−1). Importantly, at low concentrations PS-5 was shown to enhance the activity of ampicillin and cephaloridine against a β-lactamase producing Proteus vulgaris strain.153 Mechanistic studies by Fukagawa et al.154 established PS-5's mode of β-lactamase inhibition through the application of a newly developed computer-assisted UV-spectrophotometric assay, which analysed reaction progress curves using integrated Michaelis–Menten kinetics rather than traditional initial-velocity methods. These studies revealed that PS-5 was highly resistant to hydrolysis by the Bacillus licheniformis 749/C β-lactamase, and that its inhibition followed first-order kinetics. The rate of enzyme inactivation was shown to be both concentration- and time-dependent, although a small fraction of enzymatic activity invariably remained, suggesting that PS-5 may act through a reversible mechanism.154
Proteins can also be β-lactamase inhibitors: the β-lactamase inhibitory protein (BLIP), was isolated from S. clavuligerus by Doran et al.155 and was shown to inhibit β-lactamases through non-covalent interactions. Mechanistically, BLIP acts as a tight-binding competitive inhibitor, occluding the β-lactamase active site.156 Following this foundational discovery, Kang et al.157 isolated a homologous protein, BLIP-I, from Streptomyces exfoliatus SMF19. BLIP-I exhibits 38% sequence identity with BLIP and is a potent inhibitor of the TEM-1 β-lactamase with a Ki of 0.047 nM. Site-directed mutagenesis studies, particularly at residue Asp49, showed that specific sidechain interactions were critical for maintaining tight binding to TEM-1, with the D49A mutation resulting in a 200-fold loss of inhibition potency.157 Further expanding the BLIP-family, Lim et al.158 reported on the discovery of BLIP-II, which is structurally distinct from both BLIP and BLIP-I. Despite lacking sequence similarity to BLIP, BLIP-II binds to TEM-1 with sub-nanomolar affinity, primarily through hydrophobic contacts at its apical β-turn regions, forming a sterically occlusive interaction that blocks substrate access to the TEM-1 active site. Kinetic analyses revealed that BLIP-II binding was tight and slowly reversible, requiring approximately 1 hour to achieve complete inhibition, indicating a structurally rigid yet adaptable inhibitory interface.158
The emergence of metallo-β-lactamases has prompted investigations aimed at discovering natural product-based inhibitors for them. A pioneering study was conducted by Payne et al.,125 who identified metallo-β-lactamase inhibitors from the fungal strain Chaetomium funicola. Through targeted natural product screening, three structurally related tricyclic metabolites, SB236049, SB236050, and SB238569, were purified (Fig. 10a). Amongst these, SB236049 demonstrated the most potent activity against the Bacteroides fragilis CfiA (IC50 = 0.7 µM; pIC50 = 6.15) and B. cereus II (IC50 = 0.3 µM; pIC50 = 6.52) metallo-β-lactamases, while also inhibiting P. aeruginosa IMP-1 (IC50 = 151 µM; pIC50 = 3.82). Comparative analysis confirmed that all three compounds were inactive against the serine-β-lactamase P99, highlighting their selectivity towards metallo-β-lactamases.
Mechanistic studies revealed that SB236049, SB236050, and SB238569 are competitive inhibitors of the CfiA, B. cereus II, and IMP-1 metallo-β-lactamases.125 Importantly, their inhibition activities remained constant across a range of Zn2+ concentrations, excluding nonspecific Zn2+ chelation as their mode of action. Detailed kinetic evaluation of SB238569 provided Ki values of 3.4 µM for CfiA, 17 µM for IMP-1, and 79 µM for B. cereus II. Crystallographic studies of SB236050 bound to CfiA showed that the inhibitor forms polar interactions with Lys184, Asn193, and His162, and makes a critical π-stacking interaction with Trp49 in the “flap” domain, which adopts a closed conformation over the active site upon inhibitor binding.125
When combined with meropenem, SB236050 and SB238569 exhibited promising antibacterial synergy.125 Use of 8 µg mL−1 of these inhibitors resulted in an eight to sixteen-fold reduction in the meropenem MIC (<4 µg mL−1) against clinical isolates of B. fragilis producing CfiA. However, no synergy was observed against IMP-1-producing P. aeruginosa, likely due to the inhibitors' poor outer membrane penetration rather than a lack of enzymatic inhibition.125 Collectively, this study demonstratied that selective, competitive inhibition of metallo-β-lactamases by natural products is feasible.
King et al.113 screened 500 natural product extracts combined with a sub-lethal concentration of meropenem against an efflux-deficient E. coli strain engineered to express NDM-1 and exhibit enhanced permeability to small molecules. From this cell-based screen, one hit extract from the fungus Aspergillus versicolor was identified that demonstrated reproducible activity in restoring meropenem activity against an NDM-1 producing E. coli strain. Using a bioactivity-guided isolation approach, the active compound was identified as the non-ribosomal peptide aspergillomarasmine A (AMA) (Fig. 10b).113 Aspergillomarasmine A demonstrated potent in vitro concentration-dependent inhibition of NDM-1, with an IC50 of 4.0 µM (pIC50 = 5.40), and an IC50 of 9.6 µM (pIC50 = 5.02) for VIM-2 inhibition. However, AMA was poorly active against IMP-7 and inactive against serine-β-lactamases such as TEM-1, CTX-M-15, KPC-2, and OXA-48. Mechanistic studies indicated that AMA acts as a selective Zn2+ chelator, removing the essential metal ions from the metallo-β-lactamase active site rather than directly interacting with the enzyme–substrate complex. This metal depletion mechanism was validated through independent approaches, including Zn2+ titration, inductively coupled plasma mass spectrometry, and time-dependent kinetic assays.159,160 Detailed equilibrium dialysis and NMR analyses further confirmed that AMA efficiently removes Zn2+ from both high- and low-affinity binding sites of NDM-1 and VIM-2, whereas IMP-7 exhibited partial resistance likely due to its tighter Zn2+ binding.113,159 Aspergillomarasmine A counteracted resistance and fully restored the efficacy of meropenem, both in vitro and in vivo. This study marked the first report of a compound with in vitro and in vivo activity against metallo-β-lactamases.113 Combination therapies of AMA with meropenem and the serine-β-lactamase inhibitor avibactam have shown promise for treating infections caused by bacterial strains co-expressing multiple β-lactamases, thereby extending the clinical potential of AMA.161
Another Aspergillus species, Aspergillus sp. OPMF00815, has served as a source of metallo-β-lactamase inhibitors, namely the hydroxyhexylitaconic acids.126 Wachino et al.126 conducted an extensive cell-based screen of 5488 microbial extracts leading to the identification of 16 samples. The 16 active extracts were further evaluated against IMP-1, with five showing >50% inhibition. Two extracts, F2400 and F1765, derived from an Aspergillus species, emerged as the most active (>70% inhibition) against IMP-1. These two extracts synergistically enhanced meropenem activity in cell-based assays, resulting in a 32-fold reduction in MIC. Focusing on sample F2400, the researchers successfully isolated two active compounds, 9-hydroxyhexylitaconic acid (9-HHIA) and 10-hydroxyhexylitaconic acid (10-HHIA) (Fig. 10c). Both these compounds demonstrated moderate inhibition of IMP-1, with IC50 values of 50.5 µM (pIC50 = 4.30) and 31.6 µM (pIC50 = 4.50), respectively; they showed moderate activity against VIM-2, but no activity against NDM-1 and SMB-1.126 Encouragingly, both compounds restored meropenem efficacy in a dose-dependent manner against an IMP-1-producing E. coli transformant strain and two clinical isolates of E. coli and K. pneumoniae. Kinetic studies revealed the mechanism of action to be substrate-competitive, rather than due to Zn2+ chelation. Structural studies showed that these compounds interacted with the Zn2+ ions in the active site, forming a stable non-covalent complex. In silico docking studies further supported these findings, implying hydrophobic interactions between the elongated methylene sidechains of HHIAs and the hydrophobic loop regions surrounding the IMP-1 active site, which are absent in less selective inhibitors.126
Fungal compounds have proven to be worthy candidates in the search for metallo-β-lactamase inhibitors. From a virtual docking screen of ∼1000 fungal compounds, four promising candidates, asperfunolone A, aspoquinolone E, emervaridone A, and emerione A, were identified with good docking scores against NDM-1 (Fig. 10d).127 Of these, emerione A exhibited the best docking score (−39.5 kcal mol−1) and the most potent in vitro activity (IC50 = 12.1 µM; pIC50 = 4.92) against NDM-1, closely followed by asperfunolone A (IC50 = 36.2 µM; pIC50 = 4.44), while the other two compounds did not demonstrate significant inhibition. Emerione A, a methylated polyketide, was observed to form interactions within the NDM-1 active site, including π–π stacking with Phe70 and direct coordination with the catalytic Zn2+ ions, suggesting a mixed inhibition mechanism involving both direct enzyme binding and partial metal chelation in solution. Emerione A and asperfunolone A showed weak antibacterial activity against E. coli and K. pneumoniae, with MIC values of 32 and 64 µM, respectively. Importantly, both emerione A and asperfunolone A restored the activity of meropenem against an NDM-1 producing E. coli strain. Emerione was the only compound able to resensitise NDM-1 producing K. pneumoniae strains to meropenem in vitro.127
In an extensive study, 6130 microbial broths were screened using a paper disk method. An extract of the fungus Paecilomyces (FKI-6801) was found to resensitise an IMP-1 producing E. coli clinical isolate to meropenem in vitro.162 The active compound in the extract was isolated and identified as 3Z,5E-octa-3,5-diene-1,3,4-tricarboxylic acid-3,4-anhydride (ODTAA) (Fig. 10e). ODTAA was weakly active against IMP-1 with an estimated IC50 between 67 and 224 µg mL−1 (pIC50 between 3.00 and 3.52). Recognising the need for potency optimisation, the authors successfully accomplished the total synthesis of ODTAA in nine steps. Subsequent medicinal chemistry efforts introduced a catechol moiety to the maleic anhydride group and another in which the maleic anhydride was converted to a monamide. These efforts improved inhibition of IMP-1 with IC50 values of 0.55 and 0.56 µg mL−1 (pIC50 values of 5.73 and 5.86) being obtained. Moreover, synthetic derivatives (such as i, ii, iii, iv, and v in Fig. 10e) exhibited improved membrane permeability and restored the antibacterial activity of meropenem against IMP-1-producing E. coli and P. aeruginosa. This study highlights how integrating natural product discovery with strategic medicinal chemistry can significantly improve lead molecule efficacy, thus opening the prospect of developing clinically viable metallo-β-lactamase inhibitors.162
In addition to having served as a source of antibiotics and serine-β-lactamase inhibitors, S. clavuligerus has also yielded a metallo-β-lactamase inhibitor.128 Holomycin, a dithiolopyrrolone antibiotic, and more specifically its reduced form (Fig. 10f), inhibits metallo-β-lactamase NDM-1 with an IC50 of 110 nM (pIC50 = 6.94). Mechanistic studies revealed that holomycin acts as a prodrug: it enters bacterial cells as the disulfide form and undergoes intracellular reduction to form the active ene-dithiol, reduced-holomycin, which exhibits strong Zn2+ binding properties, thereby disrupting metal homeostasis by chelating essential Zn2+ ions from bacterial metalloenzymes.128 Spectrophotometric and mass spectrometric assays show reduced-holomycin binds zinc ions with high affinity, forming a stable 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ligand-to-metal complex. The chelation mechanism was shown to be reversible as NDM-1 activity could be restored by the addition of excess Zn2+.128
1 ligand-to-metal complex. The chelation mechanism was shown to be reversible as NDM-1 activity could be restored by the addition of excess Zn2+.128
The work on holomycin is further evidence that S. clavuligerus stands out as a natural source of antibiotics as well as serine-β-lactamase and metallo-β-lactamase inhibitors, including both small molecules and proteins. It is reasonable to hypothesise that undiscovered organisms possess similar biosynthetic capabilities and could contribute to the discovery of new types of β-lactamase inhibitors.
3.2 β-Lactamase inhibitors from plants
The successful discovery of β-lactamase inhibitors from microorganisms has motivated the search for similar compounds in plants. Both past and ongoing plant-derived natural product screening efforts, using enzyme- and phenotypic-based assays to find extracts, fractions, and compounds that inhibit β-lactamases.Coates et al.129 reported on the bioassay-guided isolation of SB-202742 (Fig. 11), a serine-β-lactamase inhibitor identified from Spondias mombin, making it one of the first purified plant-derived β-lactamase inhibitors. This anacardic acid displayed reversible, non-competitive inhibition against several serine-β-lactamases, including TEM-1 (IC50 = 5 µg mL; pIC50 = 4.87), OXA-1 (IC50 = 79.7 µg mL; pIC50 = 3.67), and P99 (IC50 = 16.5 µg mL; pIC50 = 4.34), amongst others.129 However, SB-202742 was inactive against a S. aureus Russell produced β-lactamase (IC50 > 567 µg mL; pIC50 < 2.82). Disappointingly, SB-202742 was unable to restore amoxicillin activity against various β-lactamase-producing bacteria, despite encouraging enzyme activity. The authors hypothesised the lack of cellular activity could be due to the inability of SB-202742 to traverse the bacterial cell wall. Interestingly, SB-202742 showed promising activity against a Staphylococcus species (MIC ranging from 2 to 32 µg mL−1) and exceptional activity against Streptococcus faecalis (MIC = 0.25 µg mL−1).129 Building on this study, analogues of SB-202742 (Fig. 11, compounds a to d) were prepared and screened against a range of serine-β-lactamases providing interesting SAR data.163 Their results showed that the length of the alkyl sidechain strongly influenced activity, with longer chains conferring greater activity. It was observed that saturated sidechains conferred better activity in comparison to unsaturated ones. Many of the analogues showed improved activity compared to SB-202742, except against S. aureus Russell where they were also inactive. While the analogues were orders of magnitude less active than clavulanic acid for most β-lactamases, they were comparatively more active against the class C β-lactamases. As with SB-202742, none of the analogues showed any synergistic antibacterial activity with amoxicillin.163
Denny et al.164 showed the inhibition of a partially purified L-1 metallo-β-lactamase from Stenotrophomonas maltophilia by the flavonoid galangin (Fig. 12a). Activity of galangin against L1 was concentration dependent between 5 and 50 µg mL−1. The inhibition of the L-1 metallo-β-lactamase was not reversed by the addition of excess Zn2+, suggesting a non-chelating mechanism. Quercetin was also screened and showed inhibition of the L-1 metallo-β-lactamase, although this was less pronounced than that of galangin. In phenotypic assays, galangin did not have synergistic activity with imipenem, with the authors speculating that this could be due to limited cell-penetration of the compound.164 Interestingly, eight years following this study, galangin was isolated from rhizomes of the plant Alpinia officinarum along with two other flavonoid compounds, namely kaempferide and kaempferide-3-O-β-D-glucoside (Fig. 12a).165 While galangin showed inherent antibacterial activity, it also inhibited a penicillinase I from B. cereus and another penicillinase IV from E. cloacae. Galangin and two other test polyphenols, quercetin and baicalein (Fig. 12a), were shown to have synergistic activity with a wide range of β-lactams against clinical isolates of a penicillin-resistant S. aureus strain.165
Further reports have emerged regarding the metallo-β-lactamase-inhibiting properties of polyphenols, particularly flavonoids (Fig. 12–14). Shi et al.131 used an in silico strategy to evaluate polyphenols against NDM-1. From a list of 22 prioritised compounds, six were screened in vitro against NDM-1. The flavonoid hesperidin was the most active, with an IC50 of 3.4 µM (pIC50 = 5.47). This was followed by the compounds diosmin (IC50 = 20.7 µM; pIC50 = 4.68) and rutin (IC50 = 15.5 µM; pIC50 = 4.81) (Fig. 12a). Stevioside had the weakest activity (IC50 = 214.1 µM; pIC50 = 3.67), and one compound, ginsenoside (Rg3), was inactive (Fig. 12b).131 Using the virtual tool IFPTarget, Yu et al.130 identified the polyphenol rosmarinic acid as a putative inhibitor of VIM-2. In vitro enzymatic assays confirmed its competitive reversible inhibition of VIM-2, with an IC50 of 41.5 µM (pIC50 = 4.38), and no Zn2+ chelation mechanism was implicated.130 However, rosmarinic acid was weakly active against NDM-1 and TEM-1 at 600 µM (Fig. 13a). Nonetheless, three structurally related compounds, namely salvianic acid, caffeic acid, and salvianolic acid A (Fig. 13a), were evaluated, with the latter compound emerging as the most promising, showing activity against VIM-2 (IC50 = 27.2 µM; pIC50 = 4.57), NDM-1 (IC50 = 160.3 µM; pIC50 = 3.80), and TEM-1 (IC50 = 34.3 µM; pIC50 = 4.47).130 In a cell-based assay using an E. coli strain expressing VIM-2, salvianolic acid A and rosmarinic acid were able to reduce the MIC of meropenem by 2 to 4-fold at non-lethal concentrations.130
Magnolol, a biphenolic natural product isolated from the bark of Magnolia officinalis, was identified as an NDM-1 inhibitor (Fig. 13b). Liu et al.132 demonstrated that magnolol moderately inhibits NDM-1 activity (IC50 = 6.47 µg mL; pIC50 = 4.61) and restored the efficacy of meropenem against an NDM-1-producing E. coli strain, reducing MIC values by up to four-fold. While magnolol showed no intrinsic antibacterial activity, its combination with meropenem resulted in complete bacterial clearance within hours in vitro. Molecular modelling and mutagenesis studies indicated that magnolol binds directly to the active site (residues Val73, Lys211, Gly219, and His250), primarily via hydrophobic interactions and a stabilising hydrogen bond with Ser217, thereby preventing substrate access.132 Importantly, unlike classical metal chelators, magnolol did not disrupt Zn2+ coordination,132 possibly reducing the risk of off-target toxicity. Previous toxicological evaluations suggest that the magnolia bark extract (containing 94% magnolol) is well tolerated in vivo.166,167 These findings highlight magnolol as a safe, natural product-derived lead with potential for development as an adjuvant β-lactamase inhibitor.
The polyphenols ellagic acid and urolithin were also evaluated against the serine-β-lactamase CTX-M-15 (Fig. 13b).168 Ellagic acid inhibited CTX-M-15, while urolithin showed only weak activity. The authors hypothesised that ellagic acid's mechanism involved reaction with the nucleophilic serine residue. Ellagic acid was inactive against NDM-1 at concentrations up to 10 mM.168
The polyphenol pterostilbene (Fig. 13b) also shows activity against NDM-1,133 with an IC50 of 15.4 µg mL−1 (pIC50 = 4.22).133 Cell-based assays revealed that pterostilbene alone has minimal antibacterial activity against NDM positive E. coli and K. pneumoniae strains; in combination with meropenem, pterostilbene showed synergy against the NDM positive isolates, significantly lowering its MIC. Encouragingly, synergistic activity of pterostilbene and meropenem was observed in in vivo models: a murine thigh model of E. coli infection and a mouse pneumonia model of K. pneumoniae infection. In both cases, the pterostilbene–meropenem combination significantly reduced the bacterial burden and improved survival rates compared to the single agent treatments.133 Embelin (Fig. 13b), a hydroxyl-substituted benzoquinone from Embelia ribes, along with curcumin and resveratrol, emerged as promising hits from a primary screen of an in house collection of natural product extracts and chemicals.134 Embelin emerged as the most active of these compounds with an IC50 of 2 µM (pIC50 = 5.7) against NDM-1 and further showed moderate activity against VIM-1 (IC50 = 200 µM; pIC50 = 3.70) and IMP-1 (IC50 = 100 µM; pIC50 = 4.00). Embelin restored the activity of various antibiotics, namely meropenem, imipenem, biapenem, cefepime, ceftazidime, and cefradine, in a dose-dependent manner by 2 to 512-fold against an NDM-1-producing K. pneumoniae strain, with the most significant improvement observed with the carbapenems.134 Alone, embelin did not exhibit activity against K. pneumoniae at concentrations up to 128 µg mL−1. Furthermore, embelin was able to restore meropenem sensitivity against NDM-1 positive clinical isolates of E. coli (17 isolates), K. pneumoniae (15 isolates), and A. baumannii (14 isolates). This was at concentrations at which embelin alone did not show antibacterial activity against these isolates.134
Carnosic acid, a natural diterpene found in Salvia rosmarinus, was identified as a promising allosteric inhibitor of NDM-1 (Fig. 13b).135 The identification of carnosic acid, made through virtual screening, was validated using enzymatic assays, demonstrating an IC50 of 27.1 µM (pIC50 = 4.57) against NDM-1. Although carnosic acid itself lacked intrinsic antibacterial activity, it partially restored the efficacy of meropenem against an NDM-1 producing E. coli strain, resulting in a 4-fold reduction of the antibiotic's MIC. Mechanistic investigations, including molecular docking and 100 ns molecular dynamics simulations, indicated that carnosic acid may bind to a novel allosteric site on NDM-1, involving Phe46, Tyr64, Leu65, Asp66, and Thr94, and both hydrophobic interactions and hydrogen bonds. Carnosic acid binding is proposed to induce conformational changes in the active site region, leading to a loss of catalytic activity without direct chelation of the Zn2+ ions. Fluorescence quenching assays and binding energy calculations supported these findings. Mutagenesis of key residues (F46A, L65A, T94A) diminished binding affinity and inhibitory potency, confirming the specificity of the allosteric interaction. This study introduced an innovative paradigm in which a natural molecule achieves non-competitive, allosteric inhibition of NDM-1, contrasting with earlier active-site-binding inhibitors and potentially reducing the likelihood of resistance development. Given its low toxicity and broad availability, carnosic acid represents a promising lead scaffold for the development of allosterically binding metallo-β-lactamase inhibitors.135
Building on the success of their earlier study, Cheng et al.136 evaluated 500 flavonoids for inhibition of the class D serine-β-lactamase OXA-48. Six compounds emerged as promising candidates (IC50 ranging from 0.52 to 52.7 µM; pIC50 ranging from 4.28 to 6.28). Amongst them, α-mangostin, a tricyclic xanthone (a sub-group of polyphenols), emerged as the most active and was subsequently prioritised for further studies (Fig. 13b). The authors screened α-mangostin against a panel of β-lactamases (at 50 µM) followed by dose response investigations against NDM-1, giving an IC50 value of 25.3 µM (pIC50 = 4.60).136 Thus, α-mangostin dual activity against both the serine-β-lactamases and metallo-β-lactamases with selectivity towards OXA-48. Kinetic studies showed that α-mangostin was a non-competitive inhibitor of OXA-48. Using jump dilution and equilibrium distillation assays, the authors demonstrated that α-mangostin is a reversible inhibitor of OXA-48. Thermal shift assays showed that α-mangostin binds to and destabilise OXA-48, resulting in a decrease in the melting temperature.136
Quercetin is a flavonoid that has gained significant interest over the years for its β-lactamase inhibition properties (Fig. 14). In one study, it was observed that phenols with a keto group on the ortho-position were effective inhibitors of NDM-1: inhibition was improved by the presence of a second ketone substituent, particularly if it was either an aromatic or enolic ketone.137 Using a chemoinformatic approach, 222 molecules bearing the minimum required moieties were identified and screened in vitro against NDM-1 using a miniaturised assay. Quercetin (71% inhibition at 50 µM), along with two other flavonoids, morin (78% inhibition at 80 µM) and myricetin (85% inhibition at 50 µM), were identified as the most active inhibitors against NDM-1. Myricetin (IC50 = 3.3 µM; pIC50 = 5.48) emerged as the most active from full-dose response studies, closely followed by quercetin, whose IC50 could only be estimated to be between 5 and 10 µM (pIC50 between 5.00 and 5.30) due to its limited solubility.137 To address solubility issues, quercetin analogues were synthesised by adding either a methyl or an isopropyl group to increase their hydrophobicity. The addition of the latter functional groups resulted in a mixture of two analogues with improved activity (86% inhibition at 50 µM) compared to their parent, quercetin. Both quercetin and myricetin (at 100 µg mL−1) showed synergistic activity with imipenem, significantly reducing its MIC by 4-fold (quercetin) and 16-fold (myricetin) (Fig. 14) against an NDM-1-expressing E. coli strain.137 However, only myricetin showed synergy with imipenem against K. pneumoniae, reducing the MIC of the carbapenem 16-fold. NMR studies showed that the three flavonoids are specific inhibitors of NDM-1. Active site residues involved in binding with the flavonoids were identified as tryptophan 93 (W93), histidine 122 (H122), glutamine 123 (Q123), aspartic acid 124 (D124), asparagine 220 (N220), and histidine 250 (H250), as well as the two Zn2+ ions.137
Another study138 employed an OXA-48 producing, efflux-deficient, and outer-membrane permeable E. coli construct to screen 150 polyphenols (at 40 µg mL−1) for their ability to enhance the activity of piperacillin (at 128 µg mL−1) against the construct strain. Fifteen percent of the compounds were excluded because they showed antibacterial activity against the construct in the absence of piperacillin. Quercetin emerged as the only compound that restored antibiotic activity without showing bactericidal activity alone. Motivated by these findings, the authors investigated 15 other flavonoids together with quercetin in an enzyme-based assay against OXA-48. Eight out of the 16 compounds showed >50% inhibition against OXA-48 (at 50 µM). The eight compounds were fisetin (90% inhibition), quercetin (87%), luteolin (81%), 3′,4′,7-trihydroxyflavone (80%), apigenin (77%), kaempferol (76%), taxifolin (76%), and isoquercitrin (61%) (Fig. 14). The three most active compounds were fisetin (IC50 = 0.47 µM; pIC50 = 6.33), luteolin (IC50 = 0.55 µM; pIC50 = 6.26) and quercetin (IC50 = 1.47 µM; pIC50 = 5.83).138 Structure–activity relationship studies showed that hydroxylation in the core structure of flavonoids is important for inhibition of OXA-48. It was observed that the presence of a hydroxyl group in the A and B rings improved potency. Kinetic studies showed quercetin to be a non-competitive and reversible inhibitor of OXA-48. Using combination assays, quercetin (64 µg mL−1) reduced the MIC of piperacillin 8-fold against an E. coli construct. Comparatively, quercetin was only able to reduce the MIC of imipenem 2-fold against the same E. coli strain.138 Synergy between piperacillin and quercetin was further confirmed in an in vivo study, in which the drug combination significantly reduced bacterial load in the spleen and liver of mice infected with an OXA-48-producing E. coli strain. The two compounds alone did not significantly reduce the bacterial burden in these organs. Overall, the results of these two studies showed that quercetin has dual activity against both serine-β-lactamases and metallo-β-lactamases, a trait that few compounds have been observed to possess thus far,138 with bicyclic boronates, e.g. taniborbactam, being another example.119
Recent work has focused on transforming quercetin into a potent medicinal chemistry lead. In 2024, Lee et al.169 reported on the development of 3-O-substituted quercetin derivatives designed to simultaneously inhibit two critical resistance mechanisms in Gram-negative Enterobacteriaceae, namely RND-type efflux pumps (such as AcrAB-TolC) and broad-spectrum carbapenemases. Molecular docking studies revealed that the introduction of an ethylene-linked sterically demanding group at the 3-OH position allows these molecules to occupy novel binding sites in both targets. Notably, compound 23, featuring a tert-butyl group, exhibited pan-carbapenemase inhibitory activity against KPC-2, OXA-48, VIM-2, and NDM-1, while also potently inhibiting efflux. This dual action successfully rescued the activity of meropenem in clinical multidrug-resistant strains and showed efficacy in a CRE-infected mouse model.169
This optimisation strategy was further refined in 2025 by addressing the challenges of periplasmic accumulation and outer membrane penetration in P. aeruginosa. Recognising that the highly hydrophobic nature of the earlier 3-O-alkyl derivatives limited their efficacy to toxic concentrations, the researchers introduced polar substituents at the 7-OH position.170 This led to the identification of 7-O-carboxylic acid-substituted 3-O-alkyl difluoroquercetin, which enhanced periplasmic concentrations through better utilisation of porin pathways. This compound demonstrated potent synergism with aztreonam, a monobactam typically stable to metallo-β-lactamases but vulnerable to other β-lactamases and efflux. By simultaneously inhibiting NDM-1 and the mexAB-oprM efflux system, this compound sensitized 96% of carbapenemase-producing P. aeruginosa isolates to aztreonam at clinically relevant concentrations, marking a significant advancement in the development of non-toxic multidrug-resistant reversal agents.170
The chromone scaffold is ubiquitous in nature, occurring predominantly in plants and to a lesser extent in other natural sources.171,172 It has recently been recognised as a promising pharmacophore for metallo-β-lactamase inhibition (Fig. 15). Christopeit et al.173 reported 3-formylchromone [Fig. 15a(i)] as a reversible covalent inhibitor of NDM-1, forming a Schiff base with Lys224 in the active site [Fig. 15a(ii)]. Although its potency was moderate (KI = 580 nM),173 this study established chromones as viable warheads for metallo-β-lactamase inhibition, providing mechanistic validation through mass spectrometry, mutagenesis, and covalent docking studies. Subsequent optimisation by Liang et al.174 led to 7-substituted 3-formylchromones, including the benzo[b]thiophene derivative, CS-23, which irreversibly inhibited NDM-1 via Lys211 modification and displayed nanomolar potency (IC50 = 36 nM; pIC50 = 7.44; KI = 4.7 nM) [Fig. 15a(iii)]. Importantly, CS-23 synergises strongly with meropenem, reducing its MIC value against an NDM-1-producing E. coli strain by 32-fold, resulting in a full restoration of the carbapenem's efficacy.174
The volatile oil component houttuynin, a naturally occurring aldehyde from Houttuynia cordata, and its sodium houttuyfonate derivatives have been developed as partially reversible covalent NDM-1 inhibitors (Fig. 15b.).175 These compounds couple covalent lysine reaction with zinc coordination and hydrophobic interactions, yielding low micromolar activity (IC50s 1–14.2 µM; pIC50s 4.85–6.00) and potentiating meropenem activity against resistant strains (Fig. 15b).175 Collectively, these findings reveal that natural products not only provide privileged scaffolds, but can also be harnessed to generate covalent inhibitors of metallo-β-lactamases, representing a novel paradigm in the fight against these enzymes.
4. Future perspectives
Collectively, the foundational and subsequent studies reviewed here highlight the rich potential of microorganisms and plants as reservoirs of β-lactamase inhibitors. Systematic screening, bioassay-guided fractionation, and medicinal chemistry-driven optimisation have yielded promising scaffolds, however, persistent challenges, including bacterial cell penetration, metabolic stability, and target selectivity, continue to hinder the translational progress of many natural product classes. Historically, natural product discovery efforts progressed from actinomycete-derived serine-β-lactamase inhibitors to fungal113,125 and plant-derived metabolites with activity against metallo-β-lactamases. Building on this trajectory, the field is now converging on the ambitious goal of identifying broad-spectrum inhibitors capable of simultaneously neutralising multiple β-lactamase classes, a critical objective in remedying antimicrobial resistance.Within this context, a careful examination of the history of β-lactamase inhibitor discovery provides valuable guidance for future research directions, highlighting a vast but largely unexplored chemical space. For instance, although >400![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 naturally occurring compounds are catalogued,176,177 only a small fraction have been investigated for β-lactamase inhibition. Accordingly, more strategic prioritisation of natural compounds is warranted, particularly guided by historical insights into chemical classes that have repeatedly demonstrated efficacy. Amongst these, flavonoids represent a promising class of β-lactamase inhibitors, especially highly hydroxylated derivatives containing catechol moieties. However, their polarity limits membrane permeability, particularly in Gram-negative bacteria, and some are substrates of bacterial efflux pumps. Medicinal chemistry approaches, such as methylation of hydroxyl groups, prenylation, or halogenation, can enhance lipophilicity and membrane permeability.178–180 Alternative strategies include glycosylation to improve solubility and hybridisation with antibiotic warheads, to leverage on the dual mechanisms of action. Moreover, to avoid altering the physicochemical profile of the flavonoid scaffold, nanotechnology-based delivery systems warrant exploration to improve cellular uptake.179
000 naturally occurring compounds are catalogued,176,177 only a small fraction have been investigated for β-lactamase inhibition. Accordingly, more strategic prioritisation of natural compounds is warranted, particularly guided by historical insights into chemical classes that have repeatedly demonstrated efficacy. Amongst these, flavonoids represent a promising class of β-lactamase inhibitors, especially highly hydroxylated derivatives containing catechol moieties. However, their polarity limits membrane permeability, particularly in Gram-negative bacteria, and some are substrates of bacterial efflux pumps. Medicinal chemistry approaches, such as methylation of hydroxyl groups, prenylation, or halogenation, can enhance lipophilicity and membrane permeability.178–180 Alternative strategies include glycosylation to improve solubility and hybridisation with antibiotic warheads, to leverage on the dual mechanisms of action. Moreover, to avoid altering the physicochemical profile of the flavonoid scaffold, nanotechnology-based delivery systems warrant exploration to improve cellular uptake.179
While flavonoids offer a strong foundation, the discovery landscape of naturally occurring β-lactamase inhibitors is expanding beyond this class of molecules to include tricyclic phenolic compounds and proteins. Tricyclic phenolic compounds, whether fungal or plant-derived,125 have demonstrated notable activity, suggesting that these frameworks represent an underexplored chemical space for further β-lactamase inhibitor development. In parallel, naturally produced protein β-lactamase inhibitors (BLIP, BLIP-I, and BLIP-II), demonstrate exceptional potency against β-lactamases, but their application is limited by their size, poor uptake, proteolytic instability, and the potential for adverse immune reactions. Consequently, this creates an opportunity to explore engineered protein–protein interaction inhibitors and novel peptide mimics of the BLIPs. In this context, cyclic peptides, potentially identified through phage-display or RNA display type approaches,181 represent a compelling yet underexplored frontier for β-lactamase inhibitors. Unlike linear peptides, cyclic structures offer enhanced stability and high selectivity, a strategy already showing success in targeting undruggable sites in oncology.182 The advantage of protein mimicking cyclic peptides is that they will retain binding specificity with improved pharmacokinetics and synthetic tractability, thereby enhancing their viability as drug candidates. However, this increased stability of cyclic peptides must be balanced against potential toxicity risks associated with prolonged clearance half-lives.
Complementing these molecular design strategies is the need for broader bioprospecting, with relatively few plant and microbial species having been systematically explored as sources of β-lactamase inhibitors. In this regard, priority groups that merit further investigation include taxa such as Brassicaceae lineage183 and the Salvia genus, which have yielded compounds such as rosmarinic acid and carnosic acid.130,135 Similarly, from a microbial perspective, although Streptomyces and various fungi are prolific sources of natural products, many species remain unexplored, thereby motivating broader screening of Actinomycetes and bioremediation-associated fungi,184,185 given their capacity to produce metal-chelating secondary metabolites relevant to metallo-β-lactamase inhibition. To further expand this chemical space, attention should turn to marine derived natural products, where the marine environment harbours an extraordinary biodiversity, yielding structurally unique metabolites.186 However, translating these discoveries is often constrained by production challenges, as large-scale synthesis is often seen as a bottleneck in advancing these complex molecules through drug discovery pipelines. In this context, advances in biocatalysis hold promise in enabling exploitation of the biosynthesis of complex natural products,187,188 including those of marine origin.189 In parallel, the integration of synthetic biology and natural biosynthesis offers an opportunity to enable production of modified natural products with enhanced pharmacological properties. While synthetic biology tools have been developed for engineering microbial biosynthetic pathways, their application to plants and other producer organisms is an emerging frontier with potential to expand the arsenal of accessible β-lactamase inhibitor chemotypes. Beyond marine systems, underexplored terrestrial organisms such as lichens, which produce unique compounds,190 warrant systematic evaluation.
To effectively operationalise these discovery efforts, innovative methodologies for the isolation and identification of β-lactamase inhibitors from complex natural product extracts are required. Integration of metabolomics tools such as Global Natural Products Social Molecular Networking (GNPS),191 alongside advanced annotation tools such as SIRIUS,192 DreaMS,193 and ChemEmbed,194 is significantly accelerating the annotation and prioritisation of bioactive metabolites. These modern techniques enable the rapid distinction of novel compounds from known chemical space, reduce redundancy in discovery, and facilitate a more targeted approach to natural product drug discovery.
Building further on these integrated approaches, the introduction of in silico tools has substantial potential to advance the field, including by enabling the efficient identification and optimisation of natural product β-lactamase inhibitor hit compounds. To date, most virtual screening efforts have focused on targeting the active sites of β-lactamases; expanding into allosteric site targeting may be a productive approach. Allosteric sites in β-lactamases have been recognised since 2004,195 yet relatively few inhibitors have been identified that exploit such sites.196,197 Recent studies have demonstrated the feasibility of such an approach, with carnosic acid emerging as the first reported natural product to inhibit NDM-1 via an allosteric mechanism.135 Given this proof of concept, there is substantial merit in pursuing the discovery of additional natural allosteric inhibitors to diversify therapeutic strategies against β-lactamase-mediated resistance further. Combination therapies of traditional and allosteric inhibitors could have a synergistic effect, helping to overcome antibiotic resistance.198 One important issue that should be considered with respect to binding at non-essential allosteric sites is whether resistance will efficiently emerge, something that has been an issue with allosteric drugs used for cancer treatment.199
5. Conclusion
In conclusion, the history of the development of β-lactam antibiotics and β-lactamase inhibitors highlights the value of natural products as a fertile source of bioactive scaffolds. Natural products have yielded structurally diverse transpeptidase and β-lactamase inhibitors operating via a range of inhibition mechanisms, including competitive and non-competitive inhibition, Zn2+ ion chelation, covalent reaction, and allosteric modulation. Despite decades of progress, however, the vast diversity of plant and microbial taxa remains largely unexplored in the search for novel antibiotics and β-lactamase inhibitors. We have outlined a framework for prioritising and investigating such resources to identify β-lactamase inhibitors. When integrated with medicinal chemistry and structural biology approaches, natural product-derived candidates hold significant promise for advancement along the drug discovery pipeline toward preclinical development.6. Author contributions
Phanankosi Moyo – conceptualisation, writing – original draft, supervision, and writing – review and editing. Ikhane O. Albert, Neo Hlungwani, Perfoy Lumu, and Ndivhuwo Kevin Khorommbi – writing – original draft, and writing – review and editing. Thulani Sibanda, Karina Calvopina Tapia, Gurleen Kaur, Nana Kwaku Buabeng, Cynthia A. Danquah, Sekelwa Cosa, George Siegwart, and Lyndy J. McGaw – writing – review and editing. Christopher J. Schofield – conceptualisation, supervision, and writing – review and editing. Vinesh J. Maharaj – conceptualisation, supervision, and writing – review and editing.7. Conflicts of interest
There are no conflicts of interest to declare.8. Data availability
No primary research results, software or code are included, and no new data were generated or analysed as part of this review.9. References
- C. J. Murray, K. S. Ikuta, F. Sharara, L. Swetschinski, G. R. Aguilar, A. Gray, C. Han, C. Bisignano, P. Rao and E. Wool, Lancet, 2022, 399, 629–655 CrossRef CAS PubMed.
- I. N. Okeke, M. E. A. de Kraker, T. P. Van Boeckel, C. K. Kumar, H. Schmitt, A. C. Gales, S. Bertagnolio, M. Sharland and R. Laxminarayan, Lancet, 2024, 403, 2426–2438 CrossRef CAS PubMed.
- M. Naghavi, S. E. Vollset, K. S. Ikuta, L. R. Swetschinski, A. P. Gray, E. E. Wool, G. R. Aguilar, T. Mestrovic, G. Smith and C. Han, Lancet, 2024, 404, 1199–1226 CrossRef PubMed.
- A. N. Poudel, S. Zhu, N. Cooper, P. Little, C. Tarrant, M. Hickman and G. Yao, PLoS One, 2023, 18, e0285170 CrossRef CAS PubMed.
- O. Jonas, A. Irwin, F. Berthe, F. Le Gall, P. V. Marquez, I. Nikolic, C. Plante, M. Schneidman, D. Shriber and A. Thiebaud, Drug-resistant infections: a threat to our economic future, The World Bank, Washington DC, 2017 Search PubMed.
- X. Zhen, C. Stålsby Lundborg, X. Sun, N. Zhu, S. Gu and H. Dong, Antimicrob. Resist. Infect. Control, 2021, 10, 5 CrossRef PubMed.
- P. Dadgostar, Infect. Drug Resist., 2019, 12, 3903–3910 CrossRef CAS PubMed.
- A. McDonnell, K. Klemperer, M. Pincombe, R. S. Bonnifield, P. Yadav and J. Guzman, A New Grand Bargain to Improve the Antimicrobial Market for Human Health, Center for Global Development (CGD), Washington DC, USA and London, UK, 2023 Search PubMed.
- D. M. De Oliveira, B. M. Forde, T. J. Kidd, P. N. Harris, M. A. Schembri, S. A. Beatson, D. L. Paterson and M. J. Walker, Clin. Microbiol. Rev., 2020, 33, 1–49 CrossRef PubMed.
- W. R. Miller and C. A. Arias, Nat. Rev. Microbiol., 2024, 22, 598–616 CrossRef CAS PubMed.
- WHO, Guidelines for the prevention and control of carbapenem-resistant Enterobacteriaceae, Acinetobacter baumannii and Pseudomonas aeruginosa in health care facilities, Report 978-92-4-155017-8, World Health Organisation, Geneva, Switzerland, 2017 Search PubMed.
- M. A. Cook and G. D. Wright, Sci. Transl. Med., 2022, 14, eabo7793 CrossRef CAS PubMed.
- L. L. Silver, Clin. Microbiol. Rev., 2011, 24, 71–109 CrossRef CAS PubMed.
- H. Venter, Biosci. Rep., 2019, 39, BSR20180474 CrossRef CAS PubMed.
- W. C. Reygaert, AIMS Microbiol., 2018, 4, 482–501 CAS.
- E. M. Darby, E. Trampari, P. Siasat, M. S. Gaya, I. Alav, M. A. Webber and J. M. Blair, Nat. Rev. Microbiol., 2023, 21, 280–295 CrossRef CAS PubMed.
- L. L. Silver, Cold Spring Harbor Perspect. Med., 2017, 7, a025262 CrossRef PubMed.
- S. Shahzad, M. D. P. Willcox and B. Rayamajhee, Antibiotics, 2023, 12, 1597 CrossRef CAS PubMed.
- J. L. Markley and T. A. Wencewicz, Front. Microbiol., 2018, 9, 1058 CrossRef PubMed.
- M. J. Beech, E. C. Toma, H. G. Smith, M. M. Trush, J. H. J. Ang, M. Y. Wong, C. H. J. Wong, H. S. Ali, Z. Butt, V. Goel, F. Duarte, A. J. M. Farley, T. R. Walsh and C. J. Schofield, Chem. Sci., 2025, 16, 9691–9704 RSC.
- R. P. Elander, Appl. Microbiol. Biotechnol., 2003, 61, 385–392 CrossRef CAS PubMed.
- D. J. Newman and G. M. Cragg, J. Nat. Prod., 2020, 83, 770–803 CrossRef CAS PubMed.
- J. W. Bennett and K.-T. Chung, Adv. Appl. Microbiol., 2001, 49, 163–184 CAS.
- E. Chain, H. W. Florey, A. D. Gardner, N. G. Heatley, M. A. Jennings, J. Orr-Ewing and A. G. Sanders, Lancet, 1940, 236, 226–228 CrossRef.
- V. Davey and M. J. Johnson, Appl. Environ. Microbiol., 1953, 1, 208–211 CrossRef CAS PubMed.
- A. Ball, J. Gray and J. M. Murdoch, in Antibacterial Drugs Today, ed. A. Ball, J. Gray and J. M. Murdoch, Springer Netherlands, Dordrecht, 1 edn, 1978, pp. 6–18 Search PubMed.
- R. Gaynes, Emerging Infect. Dis., 2017, 23, 849 CrossRef.
- E. P. Abraham and E. Chain, Nature, 1940, 146, 837 CrossRef CAS.
- M. Finland, Rev. Infect. Dis., 1979, 1, 4–21 CrossRef CAS PubMed.
- G. Bo, Clin. Microbiol. Infect., 2000, 6, 6–8 CrossRef PubMed.
- G. G. Newton and E. Abraham, Nature, 1955, 175, 548 CrossRef CAS PubMed.
- R. B. Morin and M. Gorman, Penicillins and Cephalosporins, Academic Press, New York, 1982 Search PubMed.
- E. Abraham, Bioessays, 1990, 12, 601–606 CrossRef CAS PubMed.
- D. C. Hodgkin and E. Maslen, Biochem. J., 1961, 79, 393–402 CrossRef CAS PubMed.
- J. A. Jeffery, E. Abraham and G. Newton, Biochem. J., 1961, 81, 591 CrossRef CAS PubMed.
- A. H. Araten, R. S. Brooks, S. D. Choi, L. L. Esguerra, D. Savchyn, E. J. Wu, G. Leon, K. J. Sniezek and M. P. Brynildsen, J. Antibiot., 2024, 77, 135–146 CrossRef CAS PubMed.
- K. Matsumoto, Bioprocess Technol., 1993, 16, 67–88 CAS.
- X. Li, J. Wang, W. Su, C. Li, G. Qu, B. Yuan and Z. Sun, Mol. Catal., 2023, 550, 113595 CAS.
- G. Volpato, R. C. Rodrigues and R. Fernandez-Lafuente, Curr. Med. Chem., 2010, 17, 3855–3873 CrossRef CAS PubMed.
- J. Hamilton-Miller, Int. J. Antimicrob. Agents, 2008, 31, 189–192 CrossRef CAS PubMed.
- S. Karakonstantis, M. Rousaki and E. I. Kritsotakis, Antibiotics, 2022, 11, 723 CrossRef CAS PubMed.
- X. Lin and U. Kück, Appl. Microbiol. Biotechnol., 2022, 106, 8007–8020 CrossRef CAS PubMed.
- T. Aoki, H. Yoshizawa, K. Yamawaki, K. Yokoo, J. Sato, S. Hisakawa, Y. Hasegawa, H. Kusano, M. Sano, H. Sugimoto, Y. Nishitani, T. Sato, M. Tsuji, R. Nakamura, T. Nishikawa and Y. Yamano, Eur. J. Med. Chem., 2018, 155, 847–868 CrossRef CAS PubMed.
- H. G. Smith, S. Basak, V. Aniebok, M. J. Beech, F. M. Alshref, M. D. Allen, A. J. M. Farley and C. J. Schofield, Chem. Sci., 2024, 15, 16928–16937 RSC.
- A. Ito, T. Nishikawa, S. Matsumoto, H. Yoshizawa, T. Sato, R. Nakamura, M. Tsuji and Y. Yamano, Antimicrob. Agents Chemother., 2016, 60, 7396–7401 CrossRef CAS PubMed.
- D. J. Tipper and J. L. Strominger, Proc. Natl. Acad. Sci. U. S. A., 1965, 54, 1133–1141 CrossRef CAS PubMed.
- B. Lee, J. Mol. Biol., 1971, 61, 463–469 CrossRef CAS PubMed.
- D. Butterworth, M. Cole, G. Hanscomb and G. Rolinson, J. Antibiot., 1979, 32, 287–294 CrossRef CAS PubMed.
- J. S. Wells, J. C. Hunter, G. L. Astle, J. C. Sherwood, C. M. Ricca, W. H. Trejo, D. P. Bonner and R. B. Sykes, J. Antibiot., 1982, 35, 814–821 CrossRef CAS PubMed.
- R. Sykes, C. Cimarusti, D. Bonner, K. Bush, D. Floyd, N. Georgopapadakou, W. Koster, W. Liu, W. Parker and P. Principe, Nature, 1981, 291, 489–491 CrossRef CAS PubMed.
- R. Nagarajan, L. D. Boeck, M. Gorman, R. L. Hamill, C. E. Higgens, M. M. Hoehn, W. M. Stark and J. G. Whitney, J. Am. Chem. Soc., 1971, 93, 2308–2310 CrossRef CAS PubMed.
- K. E. Wilson, A. J. Kempf, J. M. Liesch and B. H. Arison, J. Antibiot., 1983, 36, 1109–1117 CrossRef CAS PubMed.
- K. M. Papp-Wallace, A. Endimiani, M. A. Taracila and R. A. Bonomo, Antimicrob. Agents Chemother., 2011, 55, 4943–4960 CrossRef CAS PubMed.
- G. G. Zhanel, R. Wiebe, L. Dilay, K. Thomson, E. Rubinstein, D. J. Hoban, A. M. Noreddin and J. A. Karlowsky, Drugs, 2007, 67, 1027–1052 CrossRef CAS PubMed.
- J. Kahan, F. Kahan, R. Goegelman, S. Currie, M. Jackson, E. Stapley, T. Miller, A. Miller, D. Hendlin and S. Mochales, J. Antibiot., 1979, 32, 1–12 CrossRef CAS PubMed.
- S. S. Weaver, G. P. Bodey and B. M. LeBlanc, Antimicrob. Agents Chemother., 1979, 15, 518–521 CrossRef CAS PubMed.
- F. M. Kahan, H. Kropp, J. G. Sundelof and J. Birnbaum, J. Antimicrob. Chemother., 1983, 12, 1–35 CrossRef CAS PubMed.
- R. Forsyth and D. Ip, J. Pharm. Biomed. Anal., 1994, 12, 1243–1248 CrossRef CAS PubMed.
- J. Edwards, J. Antimicrob. Chemother., 1995, 36, 1–17 CrossRef CAS PubMed.
- F. S. Codjoe and E. S. Donkor, Med. Sci., 2017, 6, 1 Search PubMed.
- T. Hong, E. Smith Moland, B. Abdalhamid, N. D. Hanson, J. Wang, C. Sloan, D. Fabian, A. Farajallah, J. Levine and K. S. Thomson, Clin. Infect. Dis., 2005, 40, e84–e86 CrossRef PubMed.
- R. Sykes and D. Bonner, Rev. Infect. Dis., 1985, 7, S579–S593 CrossRef CAS PubMed.
- H. C. Neu, Med. Clin. North Am., 1988, 72, 555–566 CrossRef CAS PubMed.
- P. O. Madsen, K. T. Nielsen and P. H. Graversen, J. Urol., 1988, 140, 925–932 CrossRef CAS PubMed.
- C. L. Tooke, P. Hinchliffe, E. C. Bragginton, C. K. Colenso, V. H. Hirvonen, Y. Takebayashi and J. Spencer, J. Mol. Biol., 2019, 431, 3472–3500 CrossRef CAS PubMed.
- D. A. Chiang and J. P. Dekker, Commun. Med., 2024, 4, 170 CrossRef PubMed.
- K. Bush and P. A. Bradford, Clin. Microbiol. Rev., 2020, 33, e00047 CrossRef CAS PubMed.
- D. J. Waxman and J. L. Strominger, Annu. Rev. Biochem., 1983, 52, 825–869 CrossRef CAS PubMed.
- E. Deák, A. I. Szabó, A. Kálmáczhelyi, Z. Gál, G. Barabás and A. Penyige, Microbiol., 1998, 144(Pt 8), 2169–2177 CrossRef PubMed.
- J. Lee, E. Y. Lee, S. H. Kim, D. K. Kim, K. S. Park, K. P. Kim, Y. K. Kim, T. Y. Roh and Y. S. Gho, Antimicrob. Agents Chemother., 2013, 57, 2589–2595 CrossRef CAS PubMed.
- A. Akhtar, N. Fatima and H. M. Khan, in β-Lactam Resistance in Gram-Negative Bacteria: Threats and Challenges, ed. M. Shahid, A. Singh and H. Sami, Springer Singapore, Singapore 2022, pp. 25–33 Search PubMed.
- K. Bush, Expert Rev. Anti-Infect. Ther., 2023, 21, 513–522 CrossRef CAS PubMed.
- R. M. Joji, A. Al-Mahmeed, F. K. Dar and M. Shahid, in β-Lactam Resistance in Gram-Negative Bacteria: Threats and Challenges, ed. M. Shahid, A. Singh and H. Sami, Springer Singapore, Singapore, 2022, pp. 17–24 Search PubMed.
- A. Philippon, H. Jacquier, E. Ruppé and R. Labia, Curr. Res. Transl. Med., 2019, 67, 115–122 Search PubMed.
- P. Salahuddin, A. Kumar and A. U. Khan, Curr. Protein Pept. Sci., 2018, 19, 130–144 CAS.
- A. Singh, M. Shahid, H. Sami, M. Shadab and H. M. Khan, in β-Lactam Resistance in Gram-Negative Bacteria: Threats and Challenges, ed. M. Shahid, A. Singh and H. Sami, Springer Singapore, Singapore, 1 edn, 2022, pp. 35–80, DOI:10.1007/978-981-16-9097-6.
- T. Palzkill, Front. Mol. Biosci., 2018, 5, 16 CrossRef PubMed.
- A. Egorov, M. Rubtsova, V. Grigorenko, I. Uporov and A. Veselovsky, Biomolecules, 2019, 9, 854 CrossRef CAS PubMed.
- M. Tilahun, Y. Kassa, A. Gedefie and M. Ashagire, Infect. Drug Resist., 2021, 14, 4363–4374 CrossRef CAS PubMed.
- K. Bush and P. A. Bradford, Nat. Rev. Microbiol., 2019, 17, 295–306 CrossRef CAS PubMed.
- P. G. Ambrose, O. Lomovskaya, D. C. Griffith, M. N. Dudley and B. VanScoy, Curr. Opin. Pharmacol., 2017, 36, 86–93 CrossRef CAS PubMed.
- D. A. Hayden, B. P. White and K. K. Bennett, J. Pharm. Technol., 2020, 36, 202–210 CrossRef CAS PubMed.
- I. Karaiskos, I. Galani, G. L. Daikos and H. Giamarellou, Antibiotics, 2025, 14, 528 CrossRef CAS PubMed.
- A. Beceiro and G. Bou, Rev. Res. Med. Microbiol., 2004, 15, 141–152 CrossRef.
- V. Zdarska, M. Kolar and P. Mlynarcik, Infect., Genet. Evol., 2024, 122, 105610 CrossRef CAS PubMed.
- L. Xiao, X. Wang, N. Kong, L. Zhang, M. Cao, M. Sun, Q. Wei and W. Liu, Front. Microbiol., 2019, 10, 2175 CrossRef PubMed.
- A. Philippon, G. Arlet, R. Labia and B. I. Iorga, Clin. Microbiol. Rev., 2022, 35, e00150 CrossRef PubMed.
- S. E. Boyd, A. Holmes, R. Peck, D. M. Livermore and W. Hope, Antimicrob. Agents Chemother., 2022, 66, e00216–e00222 CrossRef PubMed.
- L. Capodimonte, F. A. Meireles, G. Bahr, R. A. Bonomo, M. Dal Peraro, C. Lopez and A. J. Vila, BioRxiv, 2024, DOI:10.1101/2024.11.04.622015.
- V. Stojanoski, L. Hu, B. Sankaran, F. Wang, P. Tao, B. V. Prasad and T. Palzkill, ACS Infect. Dis., 2021, 7, 445–460 CrossRef CAS PubMed.
- D. Rajalingam, L. Piszkin, A. Rodriguez-Medina and J. W. Peng, J. Am. Chem. Soc., 2024, 146, 28648–28652 CrossRef CAS PubMed.
- Z. Cheng, P. W. Thomas, L. Ju, A. Bergstrom, K. Mason, D. Clayton, C. Miller, C. R. Bethel, J. VanPelt, D. L. Tierney, R. C. Page, R. A. Bonomo, W. Fast and M. W. Crowder, J. Biol. Chem., 2018, 293, 12606–12618 CrossRef CAS PubMed.
- L. Dortet, L. Poirel and P. Nordmann, BioMed Res. Int., 2014, 2014, 249856 Search PubMed.
- S. E. Boyd, D. M. Livermore, D. C. Hooper and W. W. Hope, Antimicrob. Agents Chemother., 2020, 64, e00397 CrossRef CAS PubMed.
- M. F. Mojica, M. Rossi, A. J. Vila and R. A. Bonomo, Lancet Infect. Dis., 2022, 22, e28–e34 CrossRef CAS PubMed.
- G. Bahr, L. J. Gonzalez and A. J. Vila, Chem. Rev., 2021, 121, 7957–8094 CrossRef CAS PubMed.
- Y. Yang, Y.-H. Yan, C. J. Schofield, A. McNally, Z. Zong and G.-B. Li, Trends Microbiol., 2023, 31, 735–748 CrossRef CAS PubMed.
- S. Alfei and A. M. Schito, Pharmaceuticals, 2022, 15, 476 CrossRef CAS PubMed.
- L.-C. Ju, Z. Cheng, W. Fast, R. A. Bonomo and M. W. Crowder, Trends Pharmacol. Sci., 2018, 39, 635–647 CrossRef CAS PubMed.
- R. A. Bonomo, Cold Spring Harbor Perspect. Med., 2017, 7, a025239 CrossRef PubMed.
- M. Castanheira, P. J. Simner and P. A. Bradford, J. Antimicrob. Chemother., 2021, 3, dlab092 Search PubMed.
- A. O. Ikhane, S. Z. Sithole, N. D. Cele, F. O. Osunsanmi, R. A. Mosa and A. R. Opoku, Antioxidants, 2024, 13, 608 CrossRef CAS PubMed.
- R. L. Antipin, D. A. Beshnova, R. A. Petrov, A. S. Shiryaeva, I. P. Andreeva, V. G. Grigorenko, M. Y. Rubtsova, A. G. Majouga, V. S. Lamzin and A. M. Egorov, Bioorg. Med. Chem. Lett., 2017, 27, 1588–1592 CrossRef CAS PubMed.
- C. Avery, L. Baker and D. J. Jacobs, Entropy, 2022, 24, 729 CrossRef CAS PubMed.
- R. P. McGeary, D. T. Tan and G. Schenk, Future Med. Chem., 2017, 9, 673–691 CrossRef CAS PubMed.
- X. Tan, H. S. Kim, K. Baugh, Y. Huang, N. Kadiyala, M. Wences, N. Singh, E. Wenzler and Z. P. Bulman, Infect. Drug Resist., 2021, 14, 125–142 CrossRef CAS PubMed.
- Z. Yang, Y. Niu, Y. Le, X. Ma and C. Qiao, Phytomedicine, 2010, 17, 139–141 CrossRef CAS PubMed.
- J. P. Werner, J. M. Mitchell, M. A. Taracila, R. A. Bonomo and R. A. Powers, Protein Sci., 2017, 26, 515–526 CrossRef CAS PubMed.
- A. G. Brown, Drug Des. Delivery, 1986, 1, 1–21 CAS.
- T. Hata, S. Omura, Y. Iwai, H. Ohno, H. Takeshima and N. Yamaguchi, J. Antibiot., 1972, 25, 473–474 CrossRef CAS PubMed.
- Y. Iwai, H. Ohno, H. Takeshima, N. Yamaguchi, S. Ōmura and T. Hata, Antimicrob. Agents Chemother., 1973, 4, 222–225 CrossRef CAS PubMed.
- D. A. Pereira and J. A. Williams, Br. J. Pharmacol., 2007, 152, 53–61 CrossRef CAS PubMed.
- A. M. King, S. A. Reid-Yu, W. Wang, D. T. King, G. De Pascale, N. C. Strynadka, T. R. Walsh, B. K. Coombes and G. D. Wright, Nature, 2014, 510, 503–506 CrossRef CAS PubMed.
- S. S. van Berkel, J. r. Brem, A. M. Rydzik, R. Salimraj, R. Cain, A. Verma, R. J. Owens, C. W. Fishwick, J. Spencer and C. J. Schofield, J. Med. Chem., 2013, 56, 6945–6953 CrossRef CAS PubMed.
- I. Heinze-Krauss, P. Angehrn, R. L. Charnas, K. Gubernator, E. M. Gutknecht, C. Hubschwerlen, M. Kania, C. Oefner, M. G. Page, S. Sogabe, J. L. Specklin and F. Winkler, J. Med. Chem., 1998, 41, 3961–3971 CrossRef CAS PubMed.
- N. C. Strynadka, R. Martin, S. E. Jensen, M. Gold and J. B. Jones, Nat. Struct. Biol., 1996, 3, 688–695 CrossRef CAS PubMed.
- R. A. Powers, F. Morandi and B. K. Shoichet, Structure, 2002, 10, 1013–1023 CrossRef CAS PubMed.
- S. J. Hecker, K. R. Reddy, M. Totrov, G. C. Hirst, O. Lomovskaya, D. C. Griffith, P. King, R. Tsivkovski, D. Sun and M. Sabet, J. Med. Chem., 2015, 58, 3682–3692 CrossRef CAS PubMed.
- B. Liu, R. E. L. Trout, G.-H. Chu, D. McGarry, R. W. Jackson, J. C. Hamrick, D. M. Daigle, S. M. Cusick, C. Pozzi, F. De Luca, M. Benvenuti, S. Mangani, J.-D. Docquier, W. J. Weiss, D. C. Pevear, L. Xerri and C. J. Burns, J. Med. Chem., 2020, 63, 2789–2801 CrossRef CAS PubMed.
- J. Brem, R. Cain, S. Cahill, M. A. McDonough, I. J. Clifton, J. C. Jiménez-Castellanos, M. B. Avison, J. Spencer, C. W. Fishwick and C. J. Schofield, Nat. Commun., 2016, 7, 12406 CrossRef CAS PubMed.
- J. Brem, T. Panduwawala, J. U. Hansen, J. Hewitt, E. Liepins, P. Donets, L. Espina, A. J. Farley, K. Shubin and G. G. Campillos, Nat. Chem., 2022, 14, 15–24 CrossRef CAS PubMed.
- K. B. Louie, S. M. Kosina, Y. Hu, H. Otani, M. de Raad, A. N. Kuftin, N. J. Mouncey, B. P. Bowen and T. R. Northen, in Comprehensive Natural Products III, ed. H.-W. Liu and T. P. Begley, Elsevier, Oxford, 2020, vol. 3, pp. 263–306 Search PubMed.
- D. J. Payne, R. Cramp, D. J. Winstanley and D. Knowles, Antimicrob. Agents Chemother., 1994, 38, 767–772 CrossRef CAS PubMed.
- J. Hood, S. Box and M. Verrall, J. Antibiot., 1979, 32, 295–304 CrossRef CAS PubMed.
- D. J. Payne, J. A. Hueso-Rodríguez, H. Boyd, N. O. Concha, C. A. Janson, M. Gilpin, J. H. Bateson, C. Cheever, N. L. Niconovich and S. Pearson, Antimicrob. Agents Chemother., 2002, 46, 1880–1886 CrossRef CAS PubMed.
- J.-I. Wachino, W. Jin, C. Norizuki, K. Kimura, M. Tsuji, H. Kurosaki and Y. Arakawa, Microbiol. Spectrum, 2024, 12, e0234423 CrossRef PubMed.
- Y. He, S. Zhou, W. Sun, Q. Li, J. Wang, J. Zhang and J. Glob, Antimicrob. Resist., 2022, 28, 216–222 CrossRef CAS PubMed.
- A. N. Chan, A. L. Shiver, W. J. Wever, S. Z. A. Razvi, M. F. Traxler and B. Li, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 2717–2722 CrossRef CAS PubMed.
- N. J. Coates, M. L. Gilpin, M. N. Gwynn, D. E. Lewis, P. H. Milner, S. R. Spear and J. W. Tyler, J. Nat. Prod., 1994, 57, 654–657 CrossRef CAS PubMed.
- Z.-J. Yu, S. Liu, S. Zhou, H. Li, F. Yang, L.-L. Yang, Y. Wu, L. Guo and G.-B. Li, Bioorg. Med. Chem. Lett., 2018, 28, 1037–1042 CrossRef CAS PubMed.
- C. Shi, J. Chen, B. Xiao, X. Kang, X. Lao, H. Zheng and J. Glob, Antimicrob. Resist., 2019, 18, 80–87 CrossRef PubMed.
- S. Liu, Y. Zhou, X. Niu, T. Wang, J. Li, Z. Liu, J. Wang, S. Tang, Y. Wang and X. Deng, Cell Death Discovery, 2018, 4, 28 CrossRef PubMed.
- S. Liu, J. Zhang, Y. Zhou, N. Hu, J. Li, Y. Wang, X. Niu, X. Deng and J. Wang, Br. J. Pharmacol., 2019, 176, 4548–4557 CrossRef CAS PubMed.
- N.-Z. Ning, X. Liu, F. Chen, P. Zhou, L. Hu, J. Huang, Z. Li, J. Huang, T. Li and H. Wang, Front. Microbiol., 2018, 9, 71 CrossRef PubMed.
- Y. Yang, Y. Guo, Y. Zhou, Y. Gao, X. Wang, J. Wang and X. Niu, Front. Pharmacol, 2020, 11, 581001 CrossRef CAS PubMed.
- W. Cheng, Y. Zhang, C. Chen, L. Gao, Y. Lv, D. Yu, B. Chen and Y. Wan, Med. Chem. Res., 2024, 33, 314–323 CrossRef CAS.
- G. Rivière, S. Oueslati, M. Gayral, J.-B. Créchet, N. Nhiri, E. Jacquet, J.-C. Cintrat, F. Giraud, C. Van Heijenoort and E. Lescop, ACS Omega, 2020, 5, 10466–10480 CrossRef PubMed.
- Y. Zhang, C. Chen, B. Cheng, L. Gao, C. Qin, L. Zhang, X. Zhang, J. Wang and Y. Wan, Front. Pharmacol, 2022, 13, 926104 CrossRef CAS PubMed.
- A. Brown, D. Butterworth, M. Cole, G. Hanscomb, J. Hood, C. Reading and G. Rolinson, J. Antibiot., 1976, 29, 668–669 CrossRef CAS PubMed.
- K. H. Baggaley, A. G. Brown and C. J. Schofield, Nat. Prod. Rep., 1997, 14, 309–333 RSC.
- T. T. Howarth, A. G. Brown and T. J. King, J. Chem. Soc., Chem. Commun., 1976, 266b–267 RSC.
- P. A. Lang, M. de Munnik, A. O. Oluwole, T. D. Claridge, C. V. Robinson, J. Brem and C. J. Schofield, ChemBioChem, 2024, 25, e202400280 CrossRef CAS PubMed.
- M. S. Butler, I. R. Henderson, R. J. Capon and M. A. T. Blaskovich, J. Antibiot., 2023, 76, 431–473 CrossRef CAS PubMed.
- A. Krajnc, P. A. Lang, T. D. Panduwawala, J. Brem and C. J. Schofield, Curr. Opin. Chem. Biol., 2019, 50, 101–110 CrossRef CAS PubMed.
- S. T. Cahill, R. Cain, D. Y. Wang, C. T. Lohans, D. W. Wareham, H. P. Oswin, J. Mohammed, J. Spencer, C. W. Fishwick, M. A. McDonough, C. J. Schofield and J. Brem, Antimicrob. Agents Chemother., 2017, 61, e02260 CrossRef CAS PubMed.
- E. Lence and C. González-Bello, Adv. Ther., 2021, 4, 2000246 CrossRef CAS.
- C. J. Easton and J. R. Knowles, Biochemistry, 1982, 21, 2857–2862 CrossRef CAS PubMed.
- K. M. J. Aertker, H. T. H. Chan, C. T. Lohans and C. J. Schofield, J. Biol. Chem., 2020, 295, 16604–16613 CrossRef CAS PubMed.
- C. T. Lohans, E. van Groesen, K. Kumar, C. L. Tooke, J. Spencer, R. S. Paton, J. Brem and C. J. Schofield, Angew. Chem., Int. Ed., 2018, 57, 1282–1285 CrossRef CAS PubMed.
- H. Umezawa, S. Mitsuhashi, M. Ham-Ada, S. Iyobe, S. Takahashi, R. Utahara, Y. Osato, S. Yamazaki, H. Ogawara and K. Maeda, J. Antibiot., 1973, 26, 51–54 CrossRef CAS PubMed.
- K. Maeda, S. Takahashi, M. Sezaki, K. Iinuma, H. Naganawa, S. Kondo, M. Ohno and H. Umezawa, J. Antibiot., 1977, 30, 770–772 CrossRef CAS PubMed.
- R. P. Novick, Biochem. J., 1962, 83, 236–240 CrossRef CAS PubMed.
- K. Okamura, S. Hirata, Y. Okumura, Y. Fukagawa, Y. Shimauchi, K. Kouno and T. Ishikura, J. Antibiot., 1978, 31, 480–482 CrossRef CAS PubMed.
- Y. Fukagawa, T. Takei and T. Ishikura, Biochem. J., 1980, 185, 177–185 CrossRef CAS PubMed.
- J. L. Doran, B. Leskiw, S. Aippersbach and S. E. Jensen, J. Bacteriol., 1990, 172, 4909–4918 CrossRef CAS PubMed.
- D. Reichmann, M. Cohen, R. Abramovich, O. Dym, D. Lim, N. C. J. Strynadka and G. Schreiber, J. Mol. Biol., 2007, 365, 663–679 CrossRef CAS PubMed.
- S. G. Kang, H. U. Park, H. S. Lee, H. T. Kim and K. J. Lee, J. Biol. Chem., 2000, 275, 16851–16856 CrossRef CAS PubMed.
- D. Lim, H. U. Park, L. De Castro, S. G. Kang, H. S. Lee, S. Jensen, K. J. Lee and N. C. Strynadka, Nat. Struct. Biol., 2001, 8, 848–852 CrossRef CAS PubMed.
- D. Sychantha, C. M. Rotondo, K. H. Tehrani, N. I. Martin and G. D. Wright, J. Biol. Chem., 2021, 297, 100918 CrossRef CAS PubMed.
- A. Bergstrom, A. Katko, Z. Adkins, J. Hill, Z. Cheng, M. Burnett, H. Yang, M. Aitha, M. R. Mehaffey, J. S. Brodbelt, K. H. M. E. Tehrani, N. I. Martin, R. A. Bonomo, R. C. Page, D. L. Tierney, W. Fast, G. D. Wright and M. W. Crowder, ACS Infect. Dis., 2018, 4, 135–145 CrossRef CAS PubMed.
- C. M. Rotondo and G. D. Wright, Antimicrob. Agents Chemother., 2024, 68, e00272 CrossRef PubMed.
- A. Ikeda, Y. Ikegaya, M. Honsho, H. Matsui, K. Nonaka, T. Ishii, Y. Asami, H. Hanaki, T. Hirose and T. Sunazuka, Bioorg. Med. Chem., 2023, 78, 117109 CrossRef CAS PubMed.
- N. W. Hird and P. H. Milner, Bioorg. Med. Chem. Lett., 1994, 4, 1423–1428 CrossRef CAS.
- B. J. Denny, P. A. Lambert and P. W. West, FEMS Microbiol. Lett., 2002, 208, 21–24 CAS.
- G. Eumkeb, S. Sakdarat and S. Siriwong, Phytomedicine, 2010, 18, 40–45 CrossRef CAS PubMed.
- Z. Liu, X. Zhang, W. Cui, X. Zhang, N. Li, J. Chen, A. W. Wong and A. Roberts, Regul. Toxicol. Pharmacol., 2007, 49, 160–171 CrossRef CAS PubMed.
- N. Li, Y. Song, W. Zhang, W. Wang, J. Chen, A. W. Wong and A. Roberts, Regul. Toxicol. Pharmacol., 2007, 49, 154–159 CrossRef CAS PubMed.
- N. Talbot, N. T. Powles and M. I. Page, RSC Adv., 2019, 9, 30637–30640 RSC.
- T. Lee, S. Lee, M. K. Kim, J. H. Ahn, J. S. Park, H. W. Seo, K.-H. Park and Y. Chong, ACS Infect. Dis., 2024, 10, 1624–1643 CrossRef CAS PubMed.
- S. Lee, T. Lee, M. K. Kim, J. H. Ahn, S. Jeong, K.-H. Park and Y. Chong, Antibiotics, 2024, 13, 1202 CrossRef CAS PubMed.
- A. Gaspar, E. M. P. J. Garrido, F. Borges and J. M. P. J. Garrido, ACS Omega, 2024, 9, 21706–21726 CrossRef CAS PubMed.
- Y. Amen, M. Elsbaey, A. Othman, M. Sallam and K. Shimizu, Molecules, 2021, 26, 7646 CrossRef CAS PubMed.
- T. Christopeit, A. Albert and H.-K. S. Leiros, Bioorg. Med. Chem., 2016, 24, 2947–2953 CrossRef CAS PubMed.
- Y. Liang, M. Huang, X. Zhu, L. Wang and Y. Wan, Bioorg. Chem., 2025, 164, 108902 CrossRef CAS PubMed.
- L. Wang, Y. Liang, P. Luo, M. Huang and Y. Wan, Bioorg. Chem., 2024, 147, 107328 CrossRef CAS PubMed.
- M. Sorokina and C. Steinbeck, J. Cheminf., 2020, 12, 20 CAS.
- M. Sorokina, P. Merseburger, K. Rajan, M. A. Yirik and C. Steinbeck, J. Cheminf., 2021, 13, 2 Search PubMed.
- C. Li, D. Taotao, C. Jun, C. Mingshun, L. Ruihong, L. Chengmei, D. Liqing and D. J. McClements, Crit. Rev. Food Sci. Nutr., 2023, 63, 10637–10658 CrossRef CAS PubMed.
- Y. Liu, J. Zhu, Z. Liu, Y. Zhi, C. Mei and H. Wang, Int. J. Mol. Sci., 2025, 26, 2455 CrossRef CAS PubMed.
- M.-Z. Wei, Y.-Y. Zhu, W.-B. Zu, H. Wang, L.-Y. Bai, Z.-S. Zhou, Y.-L. Zhao, Z.-J. Wang and X.-D. Luo, Eur. J. Med. Chem., 2024, 271, 116401 CrossRef CAS PubMed.
- Y. Huang, M. M. Wiedmann and H. Suga, Chem. Rev., 2019, 119, 10360–10391 CrossRef CAS PubMed.
- Á. Roxin and G. Zheng, Future Med. Chem., 2012, 4, 1601–1618 CrossRef PubMed.
- M. Jimenez-Valera, A. Ruiz-Bravo and A. Ramos-Cormenzana, J. Antimicrob. Chemother., 1987, 19, 31–37 CrossRef CAS PubMed.
- Y. Dinakarkumar, G. Ramakrishnan, K. R. Gujjula, V. Vasu, P. Balamurugan and G. Murali, Environ. Chem. Ecotoxicol., 2024, 6, 293–302 CrossRef CAS.
- L. Dusengemungu, G. Kasali, C. Gwanama and B. Mubemba, Environ. Adv., 2021, 5, 100083 CrossRef CAS.
- H. Malve, J. Pharm. BioAllied Sci., 2016, 8, 83–91 CrossRef CAS PubMed.
- B. S. Moore and T. A. Gulder, Nat. Prod. Rep., 2020, 37, 1292–1293 RSC.
- S. Chakrabarty, E. O. Romero, J. B. Pyser, J. A. Yazarians and A. R. H. Narayan, Acc. Chem. Res., 2021, 54, 1374–1384 CrossRef CAS PubMed.
- B.-S. Chen, D. Zhang, F. Z. R. de Souza and L. Liu, Mar. Drugs, 2022, 20, 368 CrossRef CAS PubMed.
- M. Ren, S. Jiang, Y. Wang, X. Pan, F. Pan and X. Wei, Front. Microbiol., 2023, 14, 1177123 CrossRef PubMed.
- M. Wang, J. J. Carver, V. V. Phelan, L. M. Sanchez, N. Garg, Y. Peng, D. D. Nguyen, J. Watrous, C. A. Kapono, T. Luzzatto-Knaan, C. Porto, A. Bouslimani, A. V. Melnik, M. J. Meehan, W. T. Liu, M. Crüsemann, P. D. Boudreau, E. Esquenazi, M. Sandoval-Calderón, R. D. Kersten, L. A. Pace, R. A. Quinn, K. R. Duncan, C. C. Hsu, D. J. Floros, R. G. Gavilan, K. Kleigrewe, T. Northen, R. J. Dutton, D. Parrot, E. E. Carlson, B. Aigle, C. F. Michelsen, L. Jelsbak, C. Sohlenkamp, P. Pevzner, A. Edlund, J. McLean, J. Piel, B. T. Murphy, L. Gerwick, C. C. Liaw, Y. L. Yang, H. U. Humpf, M. Maansson, R. A. Keyzers, A. C. Sims, A. R. Johnson, A. M. Sidebottom, B. E. Sedio, A. Klitgaard, C. B. Larson, C. A. B. P, D. Torres-Mendoza, D. J. Gonzalez, D. B. Silva, L. M. Marques, D. P. Demarque, E. Pociute, E. C. O'Neill, E. Briand, E. J. N. Helfrich, E. A. Granatosky, E. Glukhov, F. Ryffel, H. Houson, H. Mohimani, J. J. Kharbush, Y. Zeng, J. A. Vorholt, K. L. Kurita, P. Charusanti, K. L. McPhail, K. F. Nielsen, L. Vuong, M. Elfeki, M. F. Traxler, N. Engene, N. Koyama, O. B. Vining, R. Baric, R. R. Silva, S. J. Mascuch, S. Tomasi, S. Jenkins, V. Macherla, T. Hoffman, V. Agarwal, P. G. Williams, J. Dai, R. Neupane, J. Gurr, A. M. C. Rodríguez, A. Lamsa, C. Zhang, K. Dorrestein, B. M. Duggan, J. Almaliti, P. M. Allard, P. Phapale, L. F. Nothias, T. Alexandrov, M. Litaudon, J. L. Wolfender, J. E. Kyle, T. O. Metz, T. Peryea, D. T. Nguyen, D. VanLeer, P. Shinn, A. Jadhav, R. Müller, K. M. Waters, W. Shi, X. Liu, L. Zhang, R. Knight, P. R. Jensen, B. O. Palsson, K. Pogliano, R. G. Linington, M. Gutiérrez, N. P. Lopes, W. H. Gerwick, B. S. Moore, P. C. Dorrestein and N. Bandeira, Nat. Biotechnol., 2016, 34, 828–837 CrossRef CAS PubMed.
- K. Dührkop, M. Fleischauer, M. Ludwig, A. A. Aksenov, A. V. Melnik, M. Meusel, P. C. Dorrestein, J. Rousu and S. Böcker, Nat. Methods, 2019, 16, 299–302 CrossRef PubMed.
- R. Bushuiev, A. Bushuiev, R. Samusevich, C. Brungs, J. Sivic and T. Pluskal, Nat. Biotechnol., 2025, 44, 630–640 CrossRef PubMed.
- M. Faizan-Khan, R. Giné, J. M. Badia, M. Pérez-Ribera, M. Ruiz-Botella, A. Junza, J. Capellades, I. Pérez-López, S. Xing, A. Patan, L. Brugnara, A. Novials, J.-M. Servitja, M. Vinaixa, P. C. Dorrestein, M. Sales-Pardo, R. Guimerà and O. Yanes, Briefings Bioinf., 2026, 27, 1–12 Search PubMed.
- J. R. Horn and B. K. Shoichet, J. Mol. Biol., 2004, 336, 1283–1291 CrossRef CAS PubMed.
- P. W. Thomas, E. J. Cho, C. R. Bethel, T. Smisek, Y.-C. Ahn, J. M. Schroeder, C. A. Thomas, K. N. Dalby, J. T. Beckham, M. W. Crowder, R. A. Bonomo and W. Fast, ACS Infect. Dis., 2022, 8, 811–824 CrossRef CAS PubMed.
- F. G. Avci, F. E. Altinisik, I. Karacan, D. Senturk Karagoz, S. Ersahin, A. Eren, N. A. Sayar, D. Vardar Ulu, E. Ozkirimli and B. Sariyar Akbulut, J. Mol. Graphics Modell., 2018, 84, 125–133 CrossRef CAS PubMed.
- E. Gianquinto, D. Tondi, G. Arrigo, L. Lazzarato and F. Spyrakis, Antibiotics, 2020, 9, 833 CrossRef CAS PubMed.
- R. Reinbold, I. C. Hvinden, P. Rabe, R. A. Herold, A. Finch, J. Wood, M. Morgan, M. Staudt, I. J. Clifton, F. A. Armstrong, J. S. O. McCullagh, J. Redmond, C. Bardella, M. I. Abboud and C. J. Schofield, Nat. Commun., 2022, 13, 4785 CrossRef CAS PubMed.
Footnote |
| † Equal contributions. |
| This journal is © The Royal Society of Chemistry 2026 |
















