 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Highlights of cyanobacterial metabolites reported between 2021–2024
Simon Sieber *a and
Elisabeth M.-L. Janssen
*a and
Elisabeth M.-L. Janssen *b
*b
aUniversity of Zürich, Department of Chemistry, Winterthurerstrasse 190, 8057 Zürich, Switzerland. E-mail: simon.sieber@uzh.ch
bSwiss Federal Institute of Aquatic Science and Technology (EAWAG), Überlandstrasse 133, 8600, Dübendorf, Switzerland. E-mail: elisabeth.janssen@eawag.ch
First published on 16th March 2026
Abstract
Covering: 2021 to 2024
This review highlights cyanobacteria secondary metabolites reported between 2021 and 2024. Those compounds are categorised into major classes, including peptides, alkaloids, lipids, polyketides, terpenes/terpenoids, and natural products from other classes. For selected compounds, the review provides details of advanced methodologies employed in their discovery, isolation and structure elucidation. Particular attention is placed on techniques for determining the relative and absolute configurations of stereocentres, including computational approaches, chemical derivatisation, and total synthesis. Furthermore, the review outlines the biosynthetic pathway of several newly identified cyanobacterial natural products.
1. Introduction
Cyanobacteria are a well-recognised source of bioactive metabolites, and their chemical diversity and biological potential continue to attract substantial attention, as reflected by the numerous reports published each year.1–14 In this review, we summarised novel natural products isolated from cyanobacteria over a recent time frame (2021 to 2024), allowing for a more in-depth discussion of a selected set of these compounds. Particular focus is placed on the methodologies employed for the structure elucidation, a process that is often complex and requires the integration of complementary techniques. In most cases, nuclear magnetic resonance (NMR) and mass spectrometry (MS) provide sufficient information to define the core structure and relative configuration of the natural products. However, assigning the absolute configuration is typically more challenging and frequently requires additional methodologies, such as computational approaches, chemical derivatisation, or total synthesis.15–24 In some cases, biosynthetic insights derived from metagenomic analysis greatly facilitate both structural assignment and stereochemical determination. This review highlights representative examples illustrating the application of these methodologies to the structural characterisation of cyanobacterial metabolites.Novel cyanobacterial metabolites are constantly being discovered, with 3162 entries (Fig. 1) as of the end of 2024, of which 3084 are listed in the open-access structural database CyanoMetDB (Version 03, 2024). This includes 110 compounds that have not been unequivocally associated with cyanobacteria, are of synthetic origin, or resemble degradation products.25 In the years 2021 through 2024, a total of 335 metabolites have been newly reported, with 58, 99, 108, and 70 each respective year, highlighting the importance of these organisms as a source of novel natural products.
Herein, we report selected newly discovered structures and order them by compound class, including peptides, alkaloids, lipids, polyketides, terpenes/terpenoids, and natural products from other classes.
2. Peptides
Peptides can have two different biosynthetic origins; they can be produced by ribosomes or by large non-ribosomal peptide synthetases (NRPS) complexes.26 Structural diversity in ribosomally produced peptides is greatly increased by the action of tailoring enzymes, giving rise to the so-called ribosomally synthesised and post-translationally modified peptides (RiPPs).27 Cyanobacteria are known to produce a great variety of peptides from NRPS and many families of RiPPs,28 such as, for example, graspetides, cyanobactins, and proteusins.29–32 The significance of these peptide classes is highlighted by the observation that more than 60 per cent of known cyanobacterial secondary metabolites contain at least one amide bond.25 Between 2021 and 2024, a total of 253 new peptide-based metabolites were described, representing 75 per cent of all new metabolites in this time span. Many of these belong to familiar cyanopeptide classes, including 86 cyanopeptolins, 19 microginins, 8 anabaenopeptins, 8 microcystins, and 3 microviridins.2.1. Ahp-cyclodepsipeptides
This class of cyclic peptides includes all compounds that possess the unusual 3-amino-6-hydroxy-2-piperidone (Ahp) moiety. Compounds from this class have recently been reviewed, covering 1989 to 2019.33 Names vary significantly among members, including symplocamide, micropeptins, lyngbyastatins, and cyanopeptolins, which constitute most of the reported compounds. Between 2021 and 2024, 86 new cyanopeptolins have been identified, representing nearly 30% of the 291 known variants of the cyanopeptide class. Of those, 79 new cyanopeptolins alone were identified from Nostoc edaphicum CCNP1411 isolated from coastal waters in the Gulf of Gdánsk.34 This Nostoc strain produces a total of 93 cyanopeptolins,34–36 which is arguably the highest number of cyanopeptides ever recorded in a single strain. This high number is an impressive demonstration of how NRPS can lead to a wealth of structural modifications around a core scaffold of cyanopeptides. Mazur-Marzec and colleagues present an overview not only of the structural diversity of cyanopeptolins but also of the similarly diverse naming convention.34 The characteristic features of cyanopeptolins are a cyclic structure of six amino acids, linked together through an ester bond (depsipeptide) and with the residue Ahp. The team detected 67 of the 79 cyanopeptolins from Nostoc edaphicum from liquid chromatography-high-resolution tandem mass spectrometry (LC-HRMS/MS) data while noting the limitations of structure elucidation based on MS/MS alone. Six cyanopeptides (CP999, CP990, CP983, CP949, CP941, and CP919) were isolated in sufficient quantities to confirm the structure assignments by 1D and 2D NMR analyses (500 MHz and 700 MHz, respectively). Additional NMR analysis of a previously published cyanopeptolin, CP999, led to a building block re-assignment to N,O-di-methyl tyrosine instead of methyl-homotyrosine formerly assigned by HRMS/MS,37 emphasising the need to require a second line of spectroscopic evidence for structure elucidation. The bioactivity of 34 cyanopeptolins was further assessed, which allowed the proposal of a structure–activity relationship (SAR).34 Inhibitory activity against trypsin was present when arginine is in position 2, while amino acids tyrosine, phenylalanine and leucine in position 2 inflicted inhibition of chymotrypsin, and elastase inhibition was present with leucine in position 2 and enhanced by 2-amino-2-butenoic acid at the N-terminus of the side chain (Fig. 2). The study also found that the viability of human cervical cancer (HeLa) cells was significantly decreased (at least 60%) by cyanopeptolins with arginine or leucine at position 2, making them potentially interesting compounds for pharmaceutical applications industries. Furthermore, additional Ahp-cyclodepsipeptides, named micropeptins LOF941, LOF925, and LOF953, have been isolated from a benthic mat dominated by a Microcoleus autumnalis strain.38 The potent neurotoxin anatoxin-a was also detected in this extract, which resonates well with several recent studies reporting the production of anatoxin-a/dihydroanatoxin-a from Microcoleus strains around the world.39–47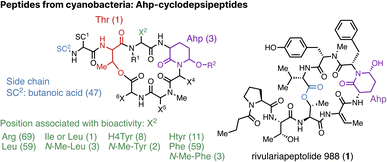 | ||
| Fig. 2 General structure of cyanopeptolins listing common variants at position 2 (X2) associated with bioactivity and butanoic acid (BA) at the terminus of the side chain (SC2). The number of cyanopeptide variants with the respective variation is given in brackets based on CyanoMetDB (Version 03, 2024).25 Potent protease inhibitor rivulariapeptolide 988 (1) isolated from Rivularia spp. PCC 7116.48 | ||
In 2021, five novel cyclic peptides were identified by functional metabolomics using native mass spectrometry, which detected ligands, i.e. metabolites, directly bound to an enzyme.48 To improve the identification of specific protein-metabolite interactions, Ultra-high performance liquid chromatography (UHPLC) separation was performed, followed by post-column pH and water content adjustment to make the conditions suitable for native MS, and the addition of a protein-binding partner, i.e., chymotrypsin. By continuously infusing chymotrypsin post-column, a mass shift indicated when metabolites bind to this serine protease at a given retention time. Extracted metabolites from Rivularia spp. PCC 7116 were analysed by this technique, and a total of 30 chymotrypsin ligands were tentatively identified. Comparison with structural and spectral databases revealed that most were unknown at the time. Five such binding metabolites were isolated, their structures were elucidated by HRMS/MS, chemical derivatisation and NMR, and they were termed rivulariapeptolides, including rivulariapeptolide 988 (1, Fig. 2). Rivulariapeptolides can be considered a new group of cyanopeptides belonging to the class of Ahp-cyclodepsipeptides.33
2.2. Microginins
Those compounds are characterised by their linear peptidic scaffold terminating with 3-amino-2-hydroxydecanoic acid (Ahda) residue at the N-terminus.49,50 Their structural diversity comprises variation in the number and identity of amino acids and in the modification of the aliphatic side chain by chlorination or deoxygenation, yielding the 3-aminodecanoic acid (Ada) moiety,51 or further variation such as 2-amino-5-(4-hydroxyphenyl)pentanoic acid (Ahppa).52 A wide variety of microginins have been newly reported, with 19 new members representing 15% of the total of variants identified to date.51–53 Recent success relied on knowledge and characterisation of biosynthetic gene clusters (BGCs). For example, a metagenomic screen targeting homologues of the dimetal-carboxylate halogenase CylC54 identified several microginin-producing strains.55 The corresponding biosynthetic gene cluster, a hybrid polyketide synthase(PKS)/NRPS (PKS/NRPS) assembly, was designated mic.51 The mic gene cluster from Microcystis aeruginosa LEGE 91341 was characterised in detail, and its heterologous expression in Escherichia coli using direct pathway cloning (DiPaC) technology led to the production of 12 new microginin congeners (Fig. 3).51 As demonstrated, identification of the BGC for a natural product class and heterologous expression in model organisms can serve as a second line of evidence and allow for the production of larger quantities for novel compound identification. | ||
| Fig. 3 General structure of 12 newly identified microginins based on the Ada, Pro/mPro, Hphe/Htyr, Pro, and Tyr residues containing non, mono or dichlorinated side chains with three variants missing the C-terminal Tyr.51 Ada: 3-aminodecanoic acid; mPro: methyl proline; Hphe: homophenylalanine; Htyr: homotyrosine. | ||
2.3. Microviridins
The common scaffold of those RiPPs consists of a tricyclic peptide containing two ω-esters and one ω-amide bond (Fig. 4).56,57 Most microviridins have been found to possess potent protease–inhibitory activity, with values reaching the nanomolar range.58,59 The success of employing genome mining to identify novel metabolite variants was also demonstrated by the discovery of three new microviridins from the mat-forming cyanobacterium Nostoc sp. TH1SO1.60 Genome mining of a single isolated filament of the cyanobacterium Nostoc sp. TH1SO1 led to the identification of three microviridin BGCs, which collectively encode five mdnA precursor peptide genes, within a high-quality metagenome-assembled genome (MAG). LC-HRMS/MS analysis of the extract enabled the identification of microviridin-1688 (2), microviridin-1739 (3) and microviridin-1748 (4) originating from the first of these BGCs (Fig. 4). No microviridins were predicted to be produced by the two other gene clusters, suggesting that these BGCs are not expressed under the culture conditions. Following the hypothesis that quorum-sensing molecules may trigger activation, the authors also investigated the BGCs of epibiont MAGs recovered from the same isolated Nostoc strain. One of the four autoinducer synthases (SGBI from Sphingobium) was heterologously expressed in E. coli, and the production of homoserine lactones (HSLs) was evident, previously associated with biofilm development of the cyanobacterium Gloeothece.61 When Nostoc sp. TH1SO1 was cultured with HSLs, up to a two-fold increase in microviridin-1688 production was observed, suggesting HSL-mediated quorum sensing. | ||
| Fig. 4 Microviridins 1688 (2), 1739 (3), and 1748 (4) isolated from Nostoc sp. Th1SO1.60 | ||
2.4. Anabaenopeptins and microcystins
Both are cyclic peptides produced by NRPS modules, with anabaenopeptins characterised by a conserved ureido group and microcystins by the unique 3-amino-9-methoxy-2,6,8-trimethyl-10-phenyldeca-4,6-dienoic acid (Adda) residue.62–65 In recent years, eight new anabaenopeptins were identified52,66–68 and the most novel finding was a glutamic acid Cα ethyl ester.66 This modified amino acid at this position had been proposed for only one variant before,69 and none of the previously 126 known anabaenopeptin variants contained this residue at the C-terminus thus far, which is most often occupied by leucine, isoleucine, phenylalanine, arginine, or tyrosine.Among the eight newly discovered microcystins,53,70–73 two contained S-propylhomocysteine and its sulfoxide variant at position 2 (comparable to the position of leucine in MC-LR), namely MC-PrhcysR and MC-Prhcys(O)R, respectively (Fig. 5).53 Cysteines are rare in microcystins, as they are present in only 8 of the 332 reported microcystins.25 Two additional microcystins contain a glutamic acid methyl ester, where it is possible that the esterification is also an artefact resulting from the extraction process.70
 | ||
| Fig. 5 O-Ethylation of glutamic acids and thioether oxidation to sulfoxide products are debated to be natural building blocks of secondary metabolites or artefacts formed during sample handling.53,70 | ||
2.5. Prenylated cyanobactins
Prenylation, catalysed by prenyltransferases (PTases), is a post-translational modification found in several organisms, including cyanobacteria.74–76 The modification can occur at two positions of the prenyl group; attachment at the C-1 position is termed forward prenylation, and attachment at the C-3 position is called reverse prenylation.77 The identification of the novel cyanobactin tolypamide (5, Fig. 6) from the cyanobacterium Tolypothrix sp. PCC 7601 revealed forward O-prenylation of the threonine residue, a previously unknown modification in cyanobacterial metabolites.78 The corresponding TolF PTase shared only low sequence coverage with known PTases, and further characterisation demonstrated its selectivity for serine and threonine forward prenylation. | ||
| Fig. 6 Tolypamide (5) prenylated at threonine position isolated from Tolypothrix sp. PCC 7601 (ref. 78) and argicyclamides (6–8) mono- and diprenylated at the guanidine moieties of arginine from Microcystis aeruginosa NIES-88.79 Steromaze (9) containing the posttranslational modified heptophan amino acid (10) from Nostoc calcicola FACHB-389.80 | ||
Mono- and bis-prenylations of the guanidine moiety of arginine have also been reported in three new natural products called argicyclamides A–C (6–8, Fig. 6).79 The argicyclamides were isolated from the cyanobacterium Microcystis aeruginosa NIES-88, and the authors demonstrated that prenylation occurred at either one or both Nω atoms of the guanidinium group of arginine. The structures of the compounds were elucidated by HRMS and NMR analyses and further supported by the total synthesis and heterologous expression of the unprenylated argicyclamide C (8). Furthermore, biosynthetic studies revealed that the precursor peptide is matured by an enzyme encoded in the BGC of another RiPP (kawaguchipeptins) produced by the same strain.81–83 Additionally, the predicted PTase AgcF was identified in the argicyclamide BGC and shows distinct sequence homology compared with previously identified guanidine PTases from cyanobacteria.
A more complex transformation was observed in the case of steromaze (9), with the formation of the heptophan amino acid (10) catalysed by a PTase-terpene cyclase (Fig. 6).80 Metagenomic analysis targeting putative α-β-β-α architecture (ABBA-fold) PTases84,85 led to the identification of a BGC named nct from Nostoc calcicola FACHB-389.This BGC contains two precursor peptides (NctA1 and NctA2), the predicted prenyltransferase-terpene cyclase (NctPC), a type II lanthionine synthetase (NctM) and an FeII/2-oxoglutarate-dependent oxygenase (NctI). Heterologous expression in E. coli was used to produce steromaze (9) from a modified NctA2 precursor peptide and the three maturation enzymes NctI, NctPC, and NctM. Its structure was elucidated using several analytical techniques, including NMR, MS, and chemical derivatisation. An extended version of steromaze, with five additional amino acids, was tested due to stability issues encountered during the final large-scale preparation step of 9. This compound showed no cytotoxicity against HeLa cells at concentrations up to 125 µM, no antibacterial activity against several strains, including ESKAPE strains, at concentrations up to 134 µM, and exhibited anti-cyanobacterial activity against Synechococcus elongatus PCC 7942, with an IC50 of 61.2 µM. Interestingly, the non-prenylated analogue was also tested and found to be inactive against this cyanobacterium, highlighting the importance of the heptophan amino acid for bioactivity.
2.6. Other lipopeptides
Lipopeptides constitute a diverse class of natural products characterised by fatty acid side chains. Between 2021 and 2024, 16 new linear lipopeptides have been described–caciqueamide,86 cyanochelin A,87 komesuamide,88 odopenicillatamide,88 phormidepistatin,89 wenchangamide A,90 amantamide B,91 odookeanynes A and B,92 pemuchiamides A and B,93 amantamide C,94 kavaratamides A-C,95 N-desmethylmajusculamide B,96 as well as 7 short-chain linear lipopeptides termed luquilloamides A-G.97 The structural diversity and antifungal properties of cyclic lipopeptides have been presented in a recent review.98 A group of eight new laxaphycins were discovered from Nostoc sp. UHCC 0702, adding to 28 previously published variants in this group of macrocyclic lipopeptides.99 Among the 12 building blocks, a modified amino acid unusual among secondary metabolites in cyanobacteria was evident, 3-hydroxy-4-methylproline (OHMePro). The new compounds termed heinamides were shown to have antifungal properties identified by antimicrobial bioactivity screening against 19 strains of fungi and bacteria.99 The complete genome (8.6 Mb) was assembled, harbouring the laxaphycin BGC (lxa, 93 kb) with 13 open reading frames and an unusual branch, as verified by heterologous expression. A review of microbial laxaphycins has also been published recently.100 Six cyclic lipopeptides, each with a macrocycle of 10 or 12 amino acids, have been isolated from the cyanobacterium strain Nodularia sp. NIES-3585 and named noducyclamides A1-4 and B1-2.101 Their bioactivity was evaluated, and noducyclamides A1, A2, and B1 were found to be active at low micromolar concentrations against the MCF7 breast cancer cell line. In addition, several novel aeruginosins (525,102 KT688,103 KT718,103 and KT575) have recently been characterised and tested for protease-inhibitory activity.103 Aeruginosin 525 is produced by the strain Aphanizomenon sp. KUCC C2, isolated from the Curonian Lagoon, Southeastern Baltic Sea.102,104 The compound exhibits inhibitory activity against trypsin (IC50: 71.7 µM), carboxypeptidase A (IC50: 89.7 µM), and thrombin (IC50: 0.59 µM). Aeruginosins KT688, KT718, and KT575 were isolated from Microcystis biomass collected in Lake Kinneret, Israel, during the 2016 spring bloom.103 Aeruginosin KT688 inhibited trypsin with an IC50 value of 2.38 µM, and aeruginosin KT718 with an IC50 value of 1.43 µM. Furthermore, PM170453, which possesses a terminal alkyne functional group, was isolated from Lyngbya sp., and its structure features a depsipeptide macrocyclic core.1052.7. Other peptides
In this section, natural products with peptidic scaffolds that do not fall into the preceding categories are described. Cyanobacterial blooms composed of Lyngbya cf. confervoides were collected during bloom seasons near the coast of Fort Lauderdale in Florida.106 The cyclic peptide gatorbulin-1 (11, Fig. 7) was isolated via a bioassay-guided fractionation targeting anticancer compounds. The structure was elucidated by extensive 1D and 2D NMR analyses, together with Marfey's method107 and chemical transformation to determine the absolute configuration. Gatorbulin-1 (11) is a cyclodepsipeptide containing unusual amino acid residues, including 4-methylproline, and featuring a hydroxamate group at the N terminus of alanine. To further evaluate the activity of the compound and confirm its structure, a total synthetic route was developed, providing 11 with an overall yield of 5.6% over 20 steps. The cytotoxic profile of gatorbulin-1 (11) was evaluated, and the results indicated a potential role as a tubulin polymerisation inhibitor. This hypothesis was confirmed by further experiments, and the precise mode of action was revealed through successful co-crystallisation of the compound with αβ-tubulin-DARPin complex.108 | ||
| Fig. 7 Gatorbulin-1 (11), a highly modified tubulin binder, identified by activity-guided isolation,106 the metallophore leptochelin A (12),109 and three linear and cyclic depsipeptides: hectoramide B (13),110 wajeepeptin (14),111 and tumonolide (15).112 | ||
Cyanobacteria from the genus Leptothoe were isolated from three different locations (Indonesia in 1994, Egypt in 2007, and the Republic of Cape Verde in 2018), and all strains were found to produce novel metallophores leptochelin A-C (Fig. 7, leptochelin A (12)).109 The metal-chelating properties were discovered during a detailed investigation of a minor metabolite, which was identified as the zinc complex of leptochelin A (12).
Finally, several linear and cyclic depsipeptides have been discovered in Moorena species110,111 and in a cyanobacterium collected in Tumon Bay, Guam.111,112 Hectoramide B (13) was isolated from a coculture composed of a cyanobacterium and the fungi Candida albicans.110 The macrocycle wajeepeptin (14) was identified in a Moorena sp. and displayed cytotoxicity against HeLa cells and the parasite Trypanosoma brucei rhodesiense.111 The enamide tumonolide (15) was isolated alongside its aldehyde derivative (tumonolide aldehyde), in which the enamide is converted to an aldehyde and a terminal amide.112 Tumonolide (15) was screened against a selection of GPCRs and identified as a potent, selective antagonist of the tachykinin receptor TACR2.
3. Alkaloids
The primary alkaloid discovered in cyanobacteria in recent years is the neurotoxin aetokthonotoxin (AETX, 16, Fig. 8), characterised by a pentabrominated biindole scaffold.113 This study originated from investigations into the unexplained deaths of bald eagles suffering from a lethal neurological disease, vacuolar myelinopathy.114 It was hypothesised that the cyanobacterium Aetokthonos hydrillicola,115,116 growing on the invasive aquatic plant Hydrilla verticillata, was producing a potent toxin responsible for the bird deaths. The cyanobacterial strain was successfully isolated and cultured, and it was eventually found that bromine supplementation in the culture medium was necessary for toxin production.113 After optimising culture conditions, AETX was isolated and its structure elucidated. Additionally, a metagenomic approach identified the AETX gene cluster, comprising six genes (aetA to aetF). AetF was predicted to function as a halogenase, and in vitro experiments confirmed its ability to brominate tryptophan at positions 5 and 7, corresponding to the left side of AETX.113,117 Subsequently, the remaining enzymes from the gene cluster were expressed and purified.117 In vitro experiments indicated that AetE is a tryptophanase that converts tryptophan to indole; AetA is a second halogenase that brominates indole at positions 2 and 3; AetD facilitates nitrile formation; and AetB, a cytochrome P450 enzyme, catalyses the C–N coupling (Fig. 8).117 Crystallisation of AetF118 and AetD119,120 provided crucial insights into their enzymatic mechanisms. | ||
| Fig. 8 Structure and proposed biosynthesis of the alkaloid aetokthonotoxin (16).113,117 Figure adapted with permission from Adak et al.117 | ||
Since its discovery, research on AETX has expanded to include the development of a total synthetic route to the natural product in five steps with an overall yield of 29%,121 and the creation of a PCR-based strategy for detecting AETX-producing cyanobacteria.122 Furthermore, Aetokthonos hydrillicola was further investigated for bioactive metabolites and novel linear peptides, named aetokthonostatins, were discovered.123
4. Lipids
Glasser and coworkers developed a novel protocol to accelerate the discovery of such compounds using a halide depletion strategy.124 This approach enabled the discovery of a new class of chlorinated glycolipids, nostochlorosides A–G (17–23, Fig. 9), from Nostoc punctiforme ATCC 29133. Those natural products consist of a hydroxylated lipid scaffold functionalised with a rare O-methylated gulose and chlorinated at the 12-position. The identity of the carbohydrate moiety was determined by GC-MS after hydrolysis and permethylation. In addition, the position of the chlorine atom was established via feeding experiments with D2-labelled palmitic acid. Furthermore, the BGC ngl was identified as responsible for producing those compounds. One of the enzymes of interest is NglO, a homolog of CylK, which was found to catalyse a reaction at the halogenated carbon.54,125,126 The biosynthetic role of NglO was elucidated as mediating the oligomerisation of nostochloroside A (17) via a nucleophilic chloride displacement by one of its hydroxyl groups. Evidence of this unique transformation was obtained by computational analysis and mutagenesis. | ||
| Fig. 9 Nostochloroside family class discovered in Nostoc punctiforme ATCC 29133 following a halide depletion strategy.124 | ||
Another class of chlorinated glycolipids is the bartoloside family,127,128 whose structural features include a dialkylresorcinol core bearing a xylose residue and chlorinated aliphatic side chains (Fig. 10; structures of bartoloside A (24) and H (25)). In 2024, Reis and coworkers reported a combined PCR- and metabolomics-based strategy to discover strains producing these compounds and to identify new derivatives.129 This approach consisted of a PCR screening to detect the presence of brtB and brtD, which encode enzymes responsible for the O-alkylation of the side chain130 and the formation of the dialkylresorcinol core, respectively. Selected cyanobacterial strains of the genus Synechocystis were cultured, their extracts analysed, and the data subjected to a molecular networking analysis. Using this strategy, derivatives of bartolosides A and H were identified that lacked the xylose unit or featured the replacement of the chlorine atom with hydrogen or hydroxyl groups (Fig. 10). Furthermore, additional derivatives of bartoloside B and bartoloside esters were also detected.
 | ||
| Fig. 10 Derivatives of the glycolipid bartolosides identified following a PCR screening and molecular network analysis. Bartoloside A (24) and H (25) are represented in this figure.129 | ||
5. Polyketides
Some cyanobacteria have a significant capacity to produce secondary metabolites. For instance, it was reported in 2019 that Nostoc punctiforme PCC 73102 harbours 21 BGCs identified by antiSMASH, with a few reported to the production of a natural product.131–133 Two of these BGCs, pks1 and pks2, were silent and pks1 was found to be significantly activated under high-density cultivation with high light intensity and CO2 concentration.131,132 It was hypothesised that pks1 produced quorum-sensing (QS) mediators as the gene cluster was highly transcribed at high cell density. An analysis of the pks1 BGC revealed that it is constituted of two subclusters, most likely positively regulated by an AraC-type transcription factor.132 To identify those putative QS mediators, a mutant that overexpresses the AraC-type protein was constructed, and metabolite analysis led to the isolation of four novel polyketides named nostoclides N1 (26) and N2 (27), nostovalerolactone (28) and 9-dehydronostovalerolactone (29, Fig. 11). The nostoclides and nostovalerolactones were determined to be signalling messenger molecules, as they positively regulate seven BGCs from Nostoc punctiforme PCC 73102. The biosynthesis of nostovalerolactone (28) and its congener 29 was further investigated to elucidate the formation of the ∂-valerolactone ring fused to the oxepane-like unit (Fig. 11).132 The bicyclic core structure 30 is proposed to be formed by NvlD, a FabH-like 3-oxoacyl-ACP synthase in concert with an oxygenase, which could be NvlQ or NvlH, two putative Baeyer-Villiger monooxygenases. | ||
| Fig. 11 (Top) nostoclides N1 (26) and N2 (27), nostovalerolactone (28) and 9-dehydronostovalerolactone (29) isolated from Nostoc punctiforme PCC 73102 and found as potential chemical mediators. (bottom) Key biosynthesis steps of nostovalerolactone (28).132 | ||
Natural product discovery faces numerous challenges, with the limited material availability being particularly notable.134 Fortunately, advances in instrumentation, such as high-performance NMR spectrometers, now enable characterisation of compounds at the nanomole scale.135–137 The discovery of the polyketide beru'amide (31, Fig. 12) underscores the transformative potential of these technologies in uncovering and elucidating the structures of scarce natural products.138 Beru'amide (31) was isolated from the marine cyanobacterium Okeania sp., collected along the Japanese coastline, using a 1H-NMR-guided approach targeting the characteristic chemical shift of a tert-butyl group. Despite the small yield (68 µg), structure elucidation relied on HRMS and NMR spectra recorded on an 800 MHz spectrometer. This approach revealed that beru'amide contains two trans-olefins, a trans-chloroolefin, an amide group, and a tert-butyl group (Fig. 12). To further elucidate the structure, the authors employed computational methods139,140 to calculate and compare chemical shifts using an optimised protocol developed by Hehre and coworkers141 using DP4 probability.142 The computational analysis suggested a syn relationship between the methyl and O-acetylated groups. A larger quantity of the compound was required to determine the absolute configuration of its stereocentres, and accordingly, a synthetic route to beru'amide (31, Fig. 12) was developed. This example highlights the importance of synthetic organic chemistry in the structural elucidation of natural products. The total synthesis began with the conversion of 3-butyn-1-ol (32) to the trans-chloroolefin 33 in two steps: chlorination of the terminal alkyne with N-chlorosuccinimide (NCS), followed by its reduction with lithium aluminium hydride (LAH) and AlCl3. The intermediate alcohol 33 was oxidised with Dess-Martin periodinane (DMP), then subjected to an Evans-syn aldol reaction with 34. The auxiliary was removed with H2O2, and the resulting hydroxy group was acetylated with acetic anhydride in pyridine, yielding the acetylated intermediate 35. The second fragment was synthesised by coupling carboxylic acid 36 with 3-butyn-1-amine (37) to form an amide bond, followed by hydrostannylation to obtain the intermediate 38. The final Stille coupling was performed by first activating the carboxylic acid 35 to its acyl chloride using Ghosez's reagent, then adding 38 and the catalyst Pd(t-Bu3P)2, which yielded beru'amide (31). The enantiomer was synthesised using the same strategy. Biological evaluation of beru'amide revealed activity in the low micromolar range, with IC50 values of 8.0 µM against HeLa cells and 1.2 µM against Trypanosoma brucei rhodesiense (T. b. rhodesiense), a parasite causing life-threatening sleeping sickness.
 | ||
| Fig. 12 Total synthesis of the polyketides beru'amide (31).138 | ||
In 2023, glycosylated macrolides named akunolides A–D (39–42, Fig. 13) were isolated from a species of Okeania sp. cyanobacterium collected at Akuna Beach, Okinawa, Japan.143 Structural elucidation by HRMS and NMR revealed a 16-membered macrolide core glycosylated with either a mono- or dimethylated pyranose. For akunolide A (39), the structural features include a tetrahydropyran (THP) ring, a dimethylated pyranose, a terminal alkyne, two gem-dimethyl groups, and an s-trans diene (Fig. 13). The relative configuration of akunolide A was determined by analysing proton coupling constants and 1H–1H NOESY correlations. Akunolide B (40) differs from akunolide A by the substitution of the terminal alkyne with an alkene. Both compounds (39 and 40) were subjected to catalytic hydrogenation, yielding the same saturated derivative 43, thereby confirming the proposed structure of akunolide B (40). The absolute configuration of akunolide B was further investigated using a modified Mosher's method.144 Mono- (C-9) and bis- (C-9 and C-2′) esterification were carried out using the R- and S- enantiomers of α-methoxy-α-trifluoromethylphenylacetic acid (MTPA). Subsequent NMR analysis enabled the assignment of all stereocentres. The relative and absolute configurations of akunolides C (41) and D (42) were determined by comparing proton coupling constants and 13C-NMR chemical shifts. The biological activity of akunolides A–D (39–42) was assessed against the parasite T. b. rhodesiense, and their cytotoxicities was evaluated using the normal human cell line WI-38. Akunolides A–D exhibited IC50 values of 11, 11, 12, and 14 µM, respectively, against T. b. rhodesiense, and >150, 55, >150, and 49 µM, respectively, against WI-38. Furthermore, the toxicity of akunolides A–C (39–41) was evaluated in mice via intraperitoneal injection, with no adverse effects observed at up to 200 µg per injection.145 The therapeutic significance of this class of compounds is further highlighted by the discovery of a structurally related polyketide, moorenaside, reported in 2024.146 This natural product exhibits notable anti-inflammatory properties by acting through the Keap1/Nrf2 oxidative stress pathway.
 | ||
| Fig. 13 Macrolide akunolide A–D (39–42) isolated from an Okeania sp. with their biological activities.143 THP: tetrahydropyran, T. b. rhodesiense: Trypanosoma brucei rhodesiense. | ||
The polyketide caldorazole (44) was isolated from the cyanobacterium Caldora sp., found along the coastline of Ishigaki Island, Japan (Fig. 14).147 Its structure was elucidated by HRMS and NMR spectroscopy. The individual components of the molecule were deduced from NMR correlations, in particular using the 1H–1H COSY and 1H–1H TOCSY spectra, and their connections were established via 1H–13C HMBC correlations. The configuration of the olefins was determined by 1H–1H NOESY, coupling constants analysis, and comparative 13C-NMR analysis. Caldorazole (44) features notable structural elements, including two thiazole heterocycles, six trans-olefins, and an O-methylenolpyruvamide group. The biological activity of caldorazole was evaluated, revealing strong potency against several tumour cell lines, with IC50 values of 23 nM for HeLa, 68 nM for CaSki, and 74 nM for HT1080 cells. Initially, it was hypothesised that its activity was linked to the O-methylenolpyruvamide group and the connected conjugated diene, potentially acting as an alkylating agent. To test this hypothesis, a truncated mimic of caldorazole 45 was synthesised from pyruvic acid and 3-phenylpropanal. When tested against the HeLa cell line, this derivative exhibited at least 1000-fold lower potency than caldorazole (44), indicating that cytotoxicity is not solely dependent on those groups. Further investigations into the mode of action suggested that caldorazole induces apoptosis by depleting ATP. Tumour cells rely on two primary sources of ATP: glycolysis and mitochondrial oxidative phosphorylation (respiration). To explore these mechanisms, caldorazole was tested in the presence of a glycolysis inhibitor, and its cytotoxicity was found to be selective under glucose-restricted conditions. Moreover, it demonstrated potent inhibition of mitochondrial complex I, with an IC50 of 56 nM. A total synthesis of caldorazole was also achieved, employing a linear sequence of 15 steps.148 Key transformations in the synthesis included a Stille cross-coupling, a sulfone coupling, and an amide coupling for the incorporation of the O-methylenolpyruvamide moiety.
 | ||
| Fig. 14 Caldorazole (44) discovered from the cyanobacterium Caldora sp., its derivative 45 and its mode of action.147 | ||
In 2021, Figueiredo and coworkers developed a targeted method to identify cyanobacterial metabolites containing a fatty acid unit.149 They supplemented cyanobacterial cultures with stable isotopically labelled hexanoic acid (d11-C6, 46, Fig. 15), hypothesising that the cyanobacteria would incorporate this precursor into secondary metabolites. This approach was applied to Nodularia sp. LEGE 06071 and nocuolactylates A–C (47–49) were identified through comparative metabolomic analysis (Fig. 15). The structures of these natural products were determined by NMR and HRMS/MS. A detailed analysis of the MS spectrum of nocuolactylate A (47) revealed an in-source fragment of the cytotoxic natural product nocuolin A (50, Fig. 15).150 Additionally, the second part of the compound was found to derive from the chlorosphaerolactylate class of natural products.151 The cytotoxic activity of nocuolactylates A (47) and B (48) was tested against multiple cell lines, showing lower potency than nocuolin A (50), suggesting a possible prodrug mode of action for this new class of compounds. Furthermore, other natural products, members of the hapalosin group, have been discovered using the same methodology by supplementing d11-C6 to the cyanobacterium Fischerella sp. PCC 9431.149
 | ||
| Fig. 15 Nocuolactylates (47–49): cyanobacterial polyketide metabolites isolated from Nodularia sp. LEGE 06071 following a stable isotopically labelling methodology.149 | ||
6. Terpenes/terpenoids
Two cyanobacteria strains, Calothrix sp. R-3-1 and Scytonema sp. U-3-3, were selected during a biological assay screening targeting the inhibition of the transient receptor potential melastatin 7 (TRPM7).152 After approximately 45 days of culture, compounds present in the medium were extracted using the highly porous adsorbent Diaion™ HP20 resin. The analysis of the extract by HPLC-UV and MS led to the isolation of six novel terpenes, including spironostoic acid (51), 11,12-didehydrospironostoic acid (52), and 12-hydroxy-2-oxo-11-epi-hinesol (53) from the strain R-3-1, and stigolone (54), 11R,12-dihydroxystigolone (55), and 11S,12-dihydroxystigolone (56, Fig. 16) from the strain U-3-3. The core structures of these natural products were elucidated using a combination of HRMS, infrared (IR), and 1D and 2D NMR spectroscopy. The challenge was to assign the relative and absolute configurations given the number of possible conformers. A systematic methodology was developed to determine the structures of the spirovetivane-type compounds 51–53. Initially, the strategy focused on determining the stereocentres at C-4 and C-5. Conformational analysis indicated the possibility of eight conformers for those four diastereomers (4R, 5R), (4R, 5S), (4S, 5R), and (4S, 5S). Analysis of the coupling constants indicated whether the methyl group attached to C-4 was in an axial or equatorial position, and 1H–1H NOESY key correlations indicated the relative configuration at position C-5, resulting in two possible conformers. Finally, electron circular dichroism (ECD) experiments could differentiate between the two. The configuration at C-7 was assigned through analysis of 1H–1H NOESY correlations, and computational analyses determined the configuration of the final stereocentre at C-11. A similar approach was used to determine the relative and absolute configurations of the eudesmane-type natural products isolated from strain U-3-3 54–56. The relative configuration was based on key 1H–1H NOESY correlations, coupling constants, and conformational analysis of the trans-decalin system. The absolute configuration was determined by comparative ECD analysis. The described compounds showed only slight activity at 50 µM against TRPM7, a transient receptor potential channel whose expression correlates with in vitro migration in cell lines, including tumour cell lines. | ||
| Fig. 16 Spirovetivate-like compounds isolated from Calothrix sp. R-3-1 and eudesmane-type compounds isolated from Scytonema sp. U-3-3.152 | ||
Research into natural products has undergone a significant transformation in recent years, driven by rapid advances in (meta)genomics and the availability of sophisticated tools for analysing BGCs.153–161 However, the effective discovery of novel natural products using these tools critically depends on well-annotated and experimentally validated BGCs. In the case of cyanobacteria, terpene natural products are well-documented;162 however, only a limited number of terpene gene clusters have been identified, e.g. the merosterol BGC in Scytonema sp. PCC 10023.85 Recently, the cyanobacterium Brasilonema HT-58-2 was found to produce the terpenoid tolypodiol (57) along with some derivatives (Fig. 17).163 In 2023, further investigation of this strain revealed two additional compounds, 1-oxo-tolypodiol (58) and 6-oxo-tolypodiol (59, Fig. 17).164 The tyl BGC (tolypodiol) was identified through BLAST analysis of the Brasilonema HT-58-2 genome165 and corroborated by previous literature.166 To validate these findings, a heterologous expression approach was employed. The entire tyl BGC, comprising 13 enzymes with predicted functions, was expressed in the cyanobacterial host Anabaena sp. PCC 7120 (UTEX 2576). Tolypodiol and its derivatives were successfully produced and identified using NMR, MS, and computational analysis. Further analysis of the tyl gene cluster revealed that TylF is likely a member of a novel terpene cyclase family, predicted to recognise the carboxylate 60 as its substrate (Fig. 17). Additionally, TylH was identified as an S-adenosylmethionine (SAM)-dependent methyltransferase. To test its activity, the purified enzyme was assayed, and successful production of tolypodiol (57) from its desmethylated precursor was observed. This result supports the biosynthetic hypothesis that the carboxylate 61 is methylated by TylH to form the methyl ester 62.
 | ||
| Fig. 17 Tolypodiol derivatives (58 and 59) from the cyanobacterium Brasilonema HT-58-2 and part of their biosynthesis.164 | ||
Kagimminols A (63) and B (64) are two novel cembrene-type natural products isolated from the cyanobacterium Okeania sp. collected near Kagimmi beach in Japan (Fig. 18).167 The compounds were purified by preparative HPLC with trifluoroacetic acid (TFA) as an additive. However, kagimminol B (64) proved sensitive to the strongly acidic conditions generated during solvent evaporation of the HPLC fractions. To mitigate this effect, the HPLC fractions were neutralised with NaHCO3 before concentration. The structure and relative configuration of kagimminol A (63) were elucidated by HRMS, and 1D and 2D NMR analyses. Key 1H–1H NOESY correlations and 13C-NMR chemical shift comparisons were crucial for determining the configurations of the three olefins (C-7, C-11, and C-13). Moreover, a syn relationship between C-1 and C-3 was predicted from calculated NMR chemical shifts. The absolute configuration was determined by comparing theoretical and experimental ECD spectra and further confirmed by Mosher's ester analysis of the hydroxyl group at C-1. A similar approach was applied to the structure elucidation of kagimminol B (64). However, difficulties arose due to the β-diketone, which can exist in four tautomeric forms (Fig. 18), and an additional stereocentre at C-7. The relative configuration was determined by DP4 probability analysis,142 allowing assignment of 1S, 3S, and 7R, and a cis configuration for the double bond between C-13 and C-14. The absolute configuration was established by comparing calculated and experimental ECD spectra. Both compounds were found inactive against HeLa cells at concentrations up to 30 µM but active at low micromolar concentrations against the parasite T. b. rhodesiense.
 | ||
| Fig. 18 Cembrene-type natural products isolated from cyanobacterium Okeania sp.167 | ||
7. Natural products belonging to other classes
Artificial intelligence tools have also been employed to determine the structure of novel metabolites.21,168 The cyanobacterium Salileptolyngbya sp. was collected from Ie Island, Okinawa, Japan.169 Compounds exhibiting strong UV absorption were targeted first (bromoiesol A sulfate (65) and B sulfate (66)), and subsequent purification also revealed bromoiesol A (67) and B (68, Fig. 19). The low hydrogen atom content posed significant challenges for structural elucidation through standard NMR and HRMS techniques. To overcome this, an artificial intelligence tool, Small Molecule Accurate Recognition Technology (SMART),170 was employed to propose a structural motif by analysing correlations observed in the 1H–13C HSQC spectrum. The analysis suggested the presence of aryl ethers, and the structure of bromoiesol A (67) and bromoiesol B (68) were unambiguously determined by single-crystal X-ray diffraction. Refined purification enabled the isolation of bromoiesol A and B sulfates (65 and 66). To confirm the structure of bromoiesol A sulfate, a total synthesis was undertaken (Fig. 19). The reported phenol 69 was 171 reacted with the iodonium salt 70, and the resulting product was deprotected with BBr3 to yield the phenol 71. Repeating the same coupling/deprotection strategy with a different iodonium salt 72 facilitated the synthesis of bromoiesol A (67), and subsequent treatment with a sulphur trioxide-pyridine complex yielded bromoiesol A sulfate (65). The biological activity of these natural products was evaluated, revealing IC50 values in the low micromolar range against the parasite T. b. rhodesiense: 1.2 µM for 67, 0.7 µM for 68, 8.8 µM for 65, and 7.9 µM for 66. | ||
| Fig. 19 Discovery of polyhalogenated aryl ethers from Salileptolyngbya sp.: bromoiesol A sulfate (65), bromoiesol B sulfate (66), bromoiesol A (67), and bromoiesol B (68).169 | ||
In this section, we included new natural products with mixed scaffolds, such as iezoside (73, Fig. 20), which features peptide, polyketide, and glycosylated moieties.172 During a screening for antitumor agents, the fractionated extract of the cyanobacterium Leptochromothrix valpauliae was selected for its potent cytotoxicity against HeLa cells, with an IC50 value of 79 ng mL−1. The active compound, iezoside (73), was isolated and purified via a bioassay-guided fractionation approach. Its relative configuration was determined using a combination of HRMS, and 1D and 2D NMR analyses, with SMART170 initially used to predict the structural core motif. The absolute configuration of iezoside was resolved using analytical techniques tailored to each moiety. For the peptide moiety, ozonolysis and hydrolysis were performed to release the amino acids, and chiral phase HPLC and Marfey's method were applied to assign the configurations at C-4 and C-10 107 The stereocentres of the modified rhamnose were elucidated by derivatising the hydroxy group at C-33 with Mosher's acid reagent, followed by NMR analysis.144 Elucidation of the configurations of the remaining stereocentres at C-18 and C-19 posed a greater challenge. Conformational and computational studies were employed to calculate the proton and carbon chemical shifts of a truncated model system. A comparison of the predicted values indicated either a (18R, 19R) or (18S, 19R) configuration, with the former more probable. Additionally, calculated ECD spectra supported this assignment. A total synthesis was then developed to differentiate the two diastereomers and fully establish the absolute configuration of iezoside (73). This highly convergent synthetic strategy was based on coupling the peptide 74, polyketide 75, and glycoside 76 moieties (Fig. 20). The polyketide side chain was synthesised in 12 linear steps utilising the diastereoselective Evans aldol reaction.173 The modified glycoside was prepared from rhamnose following a modified version of a previously reported route,174 while the peptide moiety was synthesised in six steps starting from protected L-leucine. This strategy yielded both diastereomers, (18R, 19R)-iezoside and (18S, 19R)-iezoside, and comparison of the data enabled unambiguous assignment of the polyketide configuration as 18R and 19R.
 | ||
| Fig. 20 The potent antiproliferative agents iezoside (73), iezoside B (77), the key intermediates for its total synthesis approach and some derivatives synthesised for the SAR study.172,175,176 | ||
With the structure elucidated, the cytotoxicity of iezoside (73) against HeLa cells was evaluated, revealing a high potency with an IC50 value of 6.8 nM. The mode of action was investigated using the anticancer drug screening platform of the Japanese Foundation for Cancer Research 39 (JFCR39).177 Comparative growth inhibition profiling across multiple cancer cell lines suggested that iezoside (73) targets a Ca2+ transporter. Subsequent experiments identified the sarco/endoplasmic reticulum Ca2+-ATPase (SERCA) as the target. Furthermore, iezoside (73) and its desmethylated analogue, iezoside B (77 Fig. 20), have also been isolated from the cyanobacterium Dichothrix sp., and iezoside B exhibited lower activity than iezoside.175
A structure–activity relationship (SAR) study was initiated to assess the significance of various structural features for biological activity.176 First, the two remaining diastereomers of the polyketide side chain, (18S, 19S)-iezoside and (18R, 19S)-iezoside, were synthesised and evaluated for antiproliferative activity against HeLa cells and for SERCA inhibitory activity. Both showed reduced potency compared with the natural product (18R, 19R)-iezoside (73). (18S, 19S)-Iezoside, which features a syn-configuration at C-18 and C-19 was the most active analogue, underscoring the importance of this configuration. The role of the peptide moiety was further examined by synthesising dimethylamine analogues of all diastereomers ((18R, 19R)-78, (18S, 19R)-79, (18S, 19S)-80, and (18R, 19S)-81, Fig. 20). These derivatives showed a complete loss of activity, highlighting the necessity of the peptide moiety for biological function. Finally, the aglycone derivatives (18R, 19R)-82, (18S, 19R)-83, (18S, 19S)-84, and (18R, 19S)-85 were synthesised, and (18R, 19R)-82 was the most potent among them. This derivative showed a twofold decrease in activity in the SERCA bioassay and over a 100-fold decrease against HeLa cells, potentially due to the relative instability of the aglycone in solution. In addition to these exciting results, the discovery of iezoside (73) has garnered significant interest within the scientific community, which was summarised in two recent publications.178,179
The next natural product provides another example of how cyanobacteria isolated from Japan are a valuable resource for drug discovery. Rivularia sp. was collected from Higashi-Hennazaki on Miyakojima Island, and purification of an ethanolic extract yielded two pyrrolinone-containing natural products, hennaminal (86) and hennamide (87) (Fig. 21).180 Their structures and relative configurations were determined by HRMS, and 1D and 2D NMR analyses. The absolute configuration of hennaminal (86) was established by comparing its experimental ECD spectrum with that obtained using computational chemistry. To further confirm this result, hennaminal (86) was subjected to ozonolysis followed by oxidative workup and hydrolysis (4M HCl), yielding L-leucine. The configuration of the isolated amino acid was determined using a chiral column and analytical standards. An unexpected transformation was observed when hennamide (87) was stored in CD3OD at −20 °C for one week. During this period, the compound converted completely into its dimeric form, hennamide dimer (88) (Fig. 21). Consequently, a total synthesis of hennamide (87) was undertaken to confirm its absolute configuration. The synthetic route began with the condensation of protected L-leucine 89 with Meldrum's acid, followed by cyclisation under reflux conditions. The resulting ketone was reduced with NaBH4 to give the pyrrolidone 90. The key intermediate 91 was obtained via a protection–deprotection sequence using benzyl trichloroacetimidate and piperidine. Subsequently, pyrrolidone 91 was coupled with the pentafluorophenol ester 92 to form 93, which was then converted into hennamide (87) via benzyl group deprotection, elimination, and removal of the Boc protecting group. Using the same strategy, (4S,10R)-epi-hennamide was synthesised, and a comparison of the optical rotations between the natural and synthetic compounds allowed the stereochemical assignment of C-4 and C-10 as S-configured. Furthermore, the bioactivity of hennaminal (86) and hennamide (87) was evaluated. Both compounds exhibited cytotoxicity against HeLa cells, with IC50 values exceeding 20 µM, and against the parasite T. b. rhodesiense, with IC50 values of 11 µM for hennaminal (86) and 9.7 µM for hennamide (87).
 | ||
| Fig. 21 Pyrrolinones isolated from Rivularia sp. and total synthesis of hennamide (87).180 | ||
8. Conclusions
Recent discoveries of cyanobacterial metabolites reported between 2021 and 2024 have revealed remarkable structural diversity across multiple compound classes. Many of the challenges associated with their structural elucidation have been addressed through modern analytical and computational technologies. High-performance NMR, together with artificial intelligence-assisted data analysis, has enabled structural elucidation from small sample quantities and complex spectra. Furthermore, significant advances in (meta)genomic approaches, including direct pathway cloning and heterologous expression, have facilitated the identification of numerous new compounds. Extensive efforts have been made to integrate spectrometry, spectroscopy, synthetic chemistry and (meta)genomic analyses for the structural elucidation of novel natural products. Overall, this review underscores the importance of cyanobacteria as a rich source of structurally diverse and innovative molecular scaffolds.9. Author contributions
Simon Sieber: conceptualisation, writing original draft and editing, visualisations. Elisabeth Janssen: conceptualisation, writing original draft and editing, data curation.10. Conflicts of interest
There are no conflicts to declare.11. Data availability
The review highlights the natural products isolated from cyanobacteria from 2021 to 2024. No new data were generated during this work. Furthermore, this manuscript does not include primary research results, software, or code. No new data were generated or analysed as part of this review.12. Acknowledgements
The authors thank the University of Zürich, in particular Prof. Dr Karl Gademann, and the Swiss Federal Institute of Aquatic Science and Technology (EAWAG) for their support. Dr Felix Hartrampf is thanked for his critical inputs during the revision of the manuscript.13. References
- A. H. Banday, N. ul Azha, R. Farooq, S. A. Sheikh, M. A. Ganie, M. N. Parray, H. Mushtaq, I. Hameed and M. A. Lone, Phytochem. Lett., 2024, 59, 124–135 CrossRef CAS.
- A. R. Carroll, B. R. Copp, R. A. Davis, R. A. Keyzers and M. R. Prinsep, Nat. Prod. Rep., 2019, 36, 122–173 RSC.
- A. R. Carroll, B. R. Copp, T. Grkovic, R. A. Keyzers and M. R. Prinsep, Nat. Prod. Rep., 2025, 42, 257–297 RSC.
- M. B. Weiss, R. M. Borges, P. Sullivan, J. P. B. Domingues, F. H. S. da Silva, V. G. S. Trindade, S. Luo, J. Orjala and C. M. Crnkovic, Nat. Prod. Rep., 2025, 42, 6–49 RSC.
- S. Ferrinho, H. Connaris, N. J. Mouncey and R. J. M. Goss, Water Res., 2024, 256, 121492 CrossRef CAS PubMed.
- P. N. N. de Freitas, F. R. Jacinavicius, L. S. Passos, A. O. de Souza, R. B. Dextro and E. Pinto, Algal Res., 2024, 82, 103617 CrossRef.
- M. A. Haffar, Z. Fajloun, S. Azar, J.-M. Sabatier and Z. A. Khattar, Toxins, 2024, 16, 551 CrossRef PubMed.
- L. T. Tan and N. F. Salleh, Molecules, 2024, 29, 5307 CrossRef CAS PubMed.
- H. Luesch, E. K. Ellis, Q.-Y. Chen and R. Ratnayake, Nat. Prod. Rep., 2024, 42, 208–256 RSC.
- Y. He, Y. Chen, H. Tao, X. Zhou, J. Liu, Y. Liu and B. Yang, Phytochem. Rev., 2025, 24, 483–525 CrossRef CAS.
- N. Barzkar, S. Sukhikh and O. Babich, Front. Microbiol., 2024, 14, 1285902 CrossRef PubMed.
- J. J. Huang, W. Xu, S. Lin and P. C. K. Cheung, Crit. Rev. Biotechnol., 2025, 45, 276–304 CrossRef CAS PubMed.
- R. Carpine and S. Sieber, Curr. Res. Biotechnol., 2021, 3, 65–81 CrossRef CAS.
- M. Gugger, A. Boullié and T. Laurent, Toxins, 2023, 15, 388 Search PubMed.
- M. Meunier, A. Schinkovitz and S. Derbré, Nat. Prod. Rep., 2024, 41, 1766–1786 Search PubMed.
- M. DiBello, A. R. Healy, H. Nikolayevskiy, Z. Xu and S. B. Herzon, Acc. Chem. Res., 2023, 56, 1656–1668 CrossRef CAS PubMed.
- B. B. Basnet, Z.-Y. Zhou, B. Wei and H. Wang, Crit. Rev. Biotechnol., 2025, 45, 1527–1558 Search PubMed.
- B. E. Hetzler, D. Trauner and A. L. Lawrence, Nat. Rev. Chem., 2022, 6, 170–181 Search PubMed.
- G. Hu and M. Qiu, Nat. Prod. Rep., 2023, 40, 1735–1753 RSC.
- A. Najmi, S. A. Javed, M. A. Bratty and H. A. Alhazmi, Molecules, 2022, 27, 349 CrossRef CAS PubMed.
- M. W. Mullowney, K. R. Duncan, S. S. Elsayed, N. Garg, J. J. J. van der Hooft, N. I. Martin, D. Meijer, B. R. Terlouw, F. Biermann, K. Blin, J. Durairaj, M. G. González, E. J. N. Helfrich, F. Huber, S. Leopold-Messer, K. Rajan, T. de Rond, J. A. van Santen, M. Sorokina, M. J. Balunas, M. A. Beniddir, D. A. van Bergeijk, L. M. Carroll, C. M. Clark, D.-A. Clevert, C. A. Dejong, C. Du, S. Ferrinho, F. Grisoni, A. Hofstetter, W. Jespers, O. V. Kalinina, S. A. Kautsar, H. Kim, T. F. Leao, J. Masschelein, E. R. Rees, R. Reher, D. Reker, P. Schwaller, M. Segler, M. A. Skinnider, A. S. Walker, E. L. Willighagen, B. Zdrazil, N. Ziemert, R. J. M. Goss, P. Guyomard, A. Volkamer, W. H. Gerwick, H. U. Kim, R. Müller, G. P. van Wezel, G. J. P. van Westen, A. K. H. Hirsch, R. G. Linington, S. L. Robinson and M. H. Medema, Nat. Rev. Drug Discovery, 2023, 22, 895–916 Search PubMed.
- M. Grigalunas, S. Brakmann and H. Waldmann, J. Am. Chem. Soc., 2022, 144, 3314–3329 CrossRef CAS PubMed.
- S. P. Gaudêncio, E. Bayram, L. L. Bilela, M. Cueto, A. R. Díaz-Marrero, B. Z. Haznedaroglu, C. Jimenez, M. Mandalakis, F. Pereira, F. Reyes and D. Tasdemir, Mar. Drugs, 2023, 21, 308 Search PubMed.
- Z. Huang, T. Bi, H. Jiang and H. Liu, Phytochem. Anal., 2024, 35, 5–16 CrossRef CAS PubMed.
- E. M.-L. Janssen, M. R. Jones, E. Pinto, F. Dörr, M. A. Torres, F. R. Jacinavicius, H. Mazur-Marzec, K. Szubert, R. Konkel, L. Tartaglione, C. Dell'Aversano, A. Miglione, P. McCarron, D. G. Beach, C. O. Miles, D. P. Fewer, K. Sivonen, J. Jokela, M. Wahlsten, T. H. J. Niedermeyer, F. Schanbacher, P. Leão, M. Preto, P. M. D'Agostino, M. Baunach, E. Dittmann, M. Miguel-Gordo, R. Reher, S. Sieber and E. Schymanski, Zenodo, 2024, DOI:10.5281/zenodo.13854577.
- R. D. Süssmuth and A. Mainz, Angew. Chem., Int. Ed., 2017, 56, 3770–3821 CrossRef PubMed.
- P. G. Arnison, M. J. Bibb, G. Bierbaum, A. A. Bowers, T. S. Bugni, G. Bulaj, J. A. Camarero, D. J. Campopiano, G. L. Challis, J. Clardy, P. D. Cotter, D. J. Craik, M. Dawson, E. Dittmann, S. Donadio, P. C. Dorrestein, K.-D. Entian, M. A. Fischbach, J. S. Garavelli, U. Göransson, C. W. Gruber, D. H. Haft, T. K. Hemscheidt, C. Hertweck, C. Hill, A. R. Horswill, M. Jaspars, W. L. Kelly, J. P. Klinman, O. P. Kuipers, A. J. Link, W. Liu, M. A. Marahiel, D. A. Mitchell, G. N. Moll, B. S. Moore, R. Müller, S. K. Nair, I. F. Nes, G. E. Norris, B. M. Olivera, H. Onaka, M. L. Patchett, J. Piel, M. J. T. Reaney, S. Rebuffat, R. P. Ross, H.-G. Sahl, E. W. Schmidt, M. E. Selsted, K. Severinov, B. Shen, K. Sivonen, L. Smith, T. Stein, R. D. Süssmuth, J. R. Tagg, G.-L. Tang, A. W. Truman, J. C. Vederas, C. T. Walsh, J. D. Walton, S. C. Wenzel, J. M. Willey and W. A. van der Donk, Nat. Prod. Rep., 2013, 30, 108–160 RSC.
- M. Montalbán-López, T. A. Scott, S. Ramesh, I. R. Rahman, A. J. van Heel, J. H. Viel, V. Bandarian, E. Dittmann, O. Genilloud, Y. Goto, M. J. G. Burgos, C. Hill, S. Kim, J. Koehnke, J. A. Latham, A. J. Link, B. Martínez, S. K. Nair, Y. Nicolet, S. Rebuffat, H.-G. Sahl, D. Sareen, E. W. Schmidt, L. Schmitt, K. Severinov, R. D. Süssmuth, A. W. Truman, H. Wang, J.-K. Weng, G. P. van Wezel, Q. Zhang, J. Zhong, J. Piel, D. A. Mitchell, O. P. Kuipers and W. A. van der Donk, Nat. Prod. Rep., 2020, 38, 130–239 RSC.
- M. N. Ahmed, E. Reyna-González, B. Schmid, V. Wiebach, R. D. Süssmuth, E. Dittmann and D. P. Fewer, ACS Chem. Biol., 2017, 12, 1538–1546 CrossRef CAS PubMed.
- N. M. Bösch, M. Borsa, U. Greczmiel, B. I. Morinaka, M. Gugger, A. Oxenius, A. L. Vagstad and J. Piel, Angew. Chem., Int. Ed., 2020, 59, 11763–11768 Search PubMed.
- K. Sivonen, N. Leikoski, D. P. Fewer and J. Jokela, Appl. Microbiol. Biotechnol., 2010, 86, 1213–1225 Search PubMed.
- A. H. Khan and C.-S. Phan, Front. Nat. Prod., 2025, 4, 1616031 CrossRef.
- S. Köcher, S. Resch, T. Kessenbrock, L. Schrapp, M. Ehrmann and M. Kaiser, Nat. Prod. Rep., 2019, 37, 163–174 RSC.
- R. Konkel, M. Cegłowska, K. Szubert, E. Wieczerzak, S. Iliakopoulou, T. Kaloudis and H. Mazur-Marzec, Mar. Drugs, 2023, 21, 508 CrossRef CAS PubMed.
- H. Mazur-Marzec, A. Fidor, M. Cegłowska, E. Wieczerzak, M. Kropidłowska, M. Goua, J. Macaskill and C. Edwards, Mar. Drugs, 2018, 16, 220 Search PubMed.
- M. Welker, M. Brunke, K. Preussel, I. Lippert and H. V. Döhren, Microbiology, 2004, 150, 1785–1796 Search PubMed.
- T. Okino, S. Qi, H. Matsuda, M. Murakami and K. Yamaguchi, J. Nat. Prod., 1997, 60, 158–161 CrossRef CAS.
- S. O'Brien, R. Alvariño, B. Kennedy, L. M. Botana and O. P. Thomas, Phytochemistry, 2024, 223, 114137 CrossRef PubMed.
- P. Junier, G. Cailleau, M. Fatton, P. Udriet, I. Hashmi, D. Bregnard, A. Corona-Ramirez, E. di Francesco, T. Kuhn, N. Mangia, S. Zhioua, D. Hunkeler, S. Bindschedler, S. Sieber and D. Gonzalez, Water Res.:X, 2024, 24, 100252 Search PubMed.
- M. J. Müller, A. Dorst, C. Paulus, I. Khan and S. Sieber, Chem. Commun., 2022, 58, 12560–12563 RSC.
- S. M. Brown, J. R. Blaszczak, R. K. Shriver, R. C. Jones, A. Sohrab, R. Goel, G. L. Boyer, B. Wei, K. M. Manoylov, T. R. Nelson, J. M. Zabrecky and R. Stancheva, Harmful Algae, 2025, 144, 102834 CrossRef CAS PubMed.
- C. Valadez-Cano, A. Reyes-Prieto, L. Johnston, Y. Huang, H. Morris, L. Zamlynny, A. M. Comeau, D. G. Beach, R. C. Jamieson and J. Lawrence, Toxicon, 2025, 264, 108461 CrossRef CAS PubMed.
- A. Diez-Chiappe, M. Á. Muñoz-Martín, S. Cirés, A. Quesada and E. Perona, Harmful Algae, 2025, 149, 102942 Search PubMed.
- A. Sohrab, S. Kaiser, B. Bhattarai, R. Stancheva and R. Goel, Sci. Total Environ., 2025, 997, 180194 CrossRef CAS PubMed.
- C. Valadez-Cano, N. Tromas, A. Reyes-Prieto, L. Johnston, Y. Huang, H. Morris, L. Zamlynny, D. G. Beach, R. C. Jamieson and J. Lawrence, Environ. Microbiol., 2025, 27, e70067 CrossRef CAS PubMed.
- S. A. Wood, L. Biessy and J. Puddick, Harmful Algae, 2018, 80, 88–95 CrossRef CAS PubMed.
- S. A. Wood, L. T. Kelly, K. Bouma-Gregson, J. Humbert, H. D. Laughinghouse, J. Lazorchak, T. G. McAllister, A. McQueen, K. Pokrzywinski, J. Puddick, C. Quiblier, L. A. Reitz, K. G. Ryan, Y. Vadeboncoeur, A. Zastepa and T. W. Davis, Freshw. Biol., 2020, 65, 1824–1842 CrossRef PubMed.
- R. Reher, A. T. Aron, P. Fajtová, P. Stincone, B. Wagner, A. I. Pérez-Lorente, C. Liu, I. Y. B. Shalom, W. Bittremieux, M. Wang, K. Jeong, M. L. Matos-Hernandez, K. L. Alexander, E. J. Caro-Diaz, C. B. Naman, J. H. W. Scanlan, P. M. M. Hochban, W. E. Diederich, C. Molina-Santiago, D. Romero, K. A. Selim, P. Sass, H. Brötz-Oesterhelt, C. C. Hughes, P. C. Dorrestein, A. J. O'Donoghue, W. H. Gerwick and D. Petras, Nat. Commun., 2022, 13, 4619 CrossRef CAS PubMed.
- T. Okino, H. Matsuda, M. Murakami and K. Yamaguchi, Tetrahedron Lett., 1993, 34, 501–504 CrossRef CAS.
- J. da S. P. Neto, G. M. Serra, L. P. Xavier and A. V. Santos, Int. J. Mol. Sci., 2025, 26, 6117 CrossRef PubMed.
- N. Eusébio, R. Castelo-Branco, D. Sousa, M. Preto, P. D'Agostino, T. A. M. Gulder and P. N. Leão, ACS Synth. Biol., 2022, 11, 3493–3503 CrossRef PubMed.
- C.-S. Phan, Z. Ling, J. J. Mehjabin, K. Matsuda, N. I. Prakoso, T. Umezawa, T. Wakimoto and T. Okino, J. Nat. Prod., 2024, 87, 2629–2639 CrossRef CAS PubMed.
- F. Varriale, L. Tartaglione, S.-K. Zervou, C. O. Miles, H. Mazur-Marzec, T. M. Triantis, T. Kaloudis, A. Hiskia and C. Dell'Aversano, Chemosphere, 2023, 311, 137012 CrossRef CAS PubMed.
- H. Nakamura, E. E. Schultz and E. P. Balskus, Nat. Chem. Biol., 2017, 13, 916–921 CrossRef CAS PubMed.
- N. Eusebio, A. Rego, N. R. Glasser, R. Castelo-Branco, E. P. Balskus and P. N. Leão, BMC Genom., 2021, 22, 633 CrossRef CAS PubMed.
- S. C. do Amaral, P. R. Monteiro, J. da S. P. Neto, G. M. Serra, E. C. Gonçalves, L. P. Xavier and A. V. Santos, Mar. Drugs, 2021, 19, 17 CrossRef CAS PubMed.
- T. K. Hemscheidt, Methods Enzymol., 2012, 516, 25–35 CAS.
- S. Sieber, S. M. Grendelmeier, L. A. Harris, D. A. Mitchell and K. Gademann, J. Nat. Prod., 2020, 83, 438–446 CrossRef CAS PubMed.
- T. Okino, H. Matsuda, M. Murakami and K. Yamaguchi, Tetrahedron, 1995, 51, 10679–10686 CrossRef CAS.
- S. Saha, P.-A. Bulzu, P. Urajová, J. Mareš, G. Konert, J. C. Manoel, M. Macho, D. Ewe, P. Hrouzek, J. Masojídek, R. Ghai and K. Saurav, mSphere, 2021, 6, e00562 CrossRef CAS PubMed.
- D. I. Sharif, J. Gallon, C. J. Smith and E. Dudley, ISME J., 2008, 2, 1171–1182 CrossRef CAS PubMed.
- P. R. Monteiro, S. C. do Amaral, A. S. Siqueira, L. P. Xavier and A. V. Santos, Toxins, 2021, 13, 522 CrossRef CAS PubMed.
- N. Bouaïcha, C. O. Miles, D. G. Beach, Z. Labidi, A. Djabri, N. Y. Benayache and T. Nguyen-Quang, Toxins, 2019, 11, 714 CrossRef PubMed.
- L. Rouhiainen, J. Jokela, D. P. Fewer, M. Urmann and K. Sivonen, Chem. Biol., 2010, 17, 265–273 CrossRef CAS PubMed.
- D. Tillett, E. Dittmann, M. Erhard, H. von Döhren, T. Börner and B. A. Neilan, Chem. Biol., 2000, 7, 753–764 CrossRef CAS PubMed.
- S.-K. Zervou, T. Kaloudis, S. Gkelis, A. Hiskia and H. Mazur-Marzec, Toxins, 2021, 14, 4 CrossRef PubMed.
- R. Konkel, M. Grabski, M. Cegłowska, E. Wieczerzak, G. Węgrzyn and H. Mazur-Marzec, Int. J. Environ. Res. Public Health, 2022, 19, 12346 CrossRef CAS PubMed.
- B. Bober, E. Chrapusta-Srebrny and J. Bialczyk, Eur. J. Phycol., 2020, 56, 244–254 CrossRef.
- E. Zafrir-Ilan and S. Carmeli, Tetrahedron, 2010, 66, 9194–9202 CrossRef CAS.
- D. Baliu-Rodriguez, N. J. Peraino, S. H. Premathilaka, J. A. Birbeck, T. Baliu-Rodriguez, J. A. Westrick and D. Isailovic, Environ. Sci. Technol., 2022, 56, 1652–1663 CrossRef CAS PubMed.
- K. Thomas, R. A. Perez Calderon, S. D. Giddings, I. Rajotte, C. O. Miles and P. McCarron, in Biotoxin Metrology Certificate of Analysis CRM-MCLA-20210128, National Research Council Canada, Halifax, 2022 Search PubMed.
- S. H. Premathilaka, J. A. Westrick and D. Isailovic, Anal. Chem., 2024, 96, 775–786 CrossRef CAS PubMed.
- K. A. Cottrill, C. O. Miles, L. C. Krajewski, B. R. Cunningham, W. Bragg, N. R. Boise, K. D. Victry, D. S. Wunschel, K. L. Wahl and E. I. Hamelin, Harmful Algae, 2024, 139, 102739 CrossRef CAS PubMed.
- Y. Zhang, Y. Goto and H. Suga, Trends Biochem. Sci., 2023, 48, 360–374 CrossRef CAS PubMed.
- F. Zhang, D. Zhao, Y. Wu and L. Li, Nat. Prod. Rep., 2025, 42, 1303–1343 RSC.
- F. Hubrich, Nat. Prod. Rep., 2026 10.1039/d5np00075k.
- Y. Hao, E. Pierce, D. Roe, M. Morita, J. A. McIntosh, V. Agarwal, T. E. Cheatham, E. W. Schmidt and S. K. Nair, Proc. Natl. Acad. Sci. U. S. A., 2016, 113, 14037–14042 CrossRef CAS PubMed.
- M. Purushothaman, S. Sarkar, M. Morita, M. Gugger, E. W. Schmidt and B. I. Morinaka, Angew. Chem., 2021, 133, 8541–8546 CrossRef.
- C.-S. Phan, K. Matsuda, N. Balloo, K. Fujita, T. Wakimoto and T. Okino, J. Am. Chem. Soc., 2021, 143, 10083–10087 CrossRef CAS PubMed.
- F. Hubrich, S. K. Kandy, C. Chepkirui, C. Padhi, S. Mordhorst, P. Moosmann, T. Zhu, M. Gugger, J. R. Chekan and J. Piel, Chem, 2024, 10, 3224–3242 CAS.
- K. Ishida, H. Matsuda, M. Murakami and K. Yamaguchi, Tetrahedron, 1996, 52, 9025–9030 CrossRef CAS.
- K. Ishida, H. Matsuda, M. Murakami and K. Yamaguchi, J. Nat. Prod., 1997, 60, 724–726 CrossRef CAS PubMed.
- A. Parajuli, D. H. Kwak, L. Dalponte, N. Leikoski, T. Galica, U. Umeobika, L. Trembleau, A. Bent, K. Sivonen, M. Wahlsten, H. Wang, E. Rizzi, G. D. Bellis, J. Naismith, M. Jaspars, X. Liu, W. Houssen and D. P. Fewer, Angew. Chem., Int. Ed., 2016, 55, 3596–3599 Search PubMed.
- P. Moosmann, F. Ecker, S. Leopold-Messer, J. K. B. Cahn, C. L. Dieterich, M. Groll and J. Piel, Nat. Chem., 2020, 12, 968–972 CrossRef CAS PubMed.
- P. Moosmann, R. Ueoka, L. Grauso, A. Mangoni, B. I. Morinaka, M. Gugger and J. Piel, Angew. Chem., Int. Ed., 2017, 56, 4987–4990 CrossRef CAS PubMed.
- O. Demirkiran, J. Almaliti, T. Leão, G. Navarro, T. Byrum, F. A. Valeriote, L. Gerwick and W. H. Gerwick, J. Nat. Prod., 2021, 84, 2081–2093 CrossRef CAS PubMed.
- T. Galica, N. Borbone, J. Mareš, A. Kust, A. Caso, G. Esposito, K. Saurav, J. Hájek, K. Řeháková, P. Urajová, V. Costantino and P. Hrouzek, Appl. Environ. Microbiol., 2021, 87, e03128–20 Search PubMed.
- K. Ozaki, A. Jinno, N. Natsume, S. Sumimoto, A. Iwasaki, K. Suenaga and T. Teruya, Tetrahedron, 2021, 85, 131969 CrossRef CAS.
- P. Sullivan, A. Krunic, L. J. Davis, H. S. Kim, J. E. Burdette and J. Orjala, J. Nat. Prod., 2021, 84, 2256–2264 CrossRef CAS PubMed.
- L. Ding, R. Bar-Shalom, D. Aharonovich, N. Kurisawa, G. Patial, S. Li, S. He, X. Yan, A. Iwasaki, K. Suenaga, C. Zhu, H. Luo, F. Tian, F. Fares, C. B. Naman and T. Luzzatto-Knaan, Mar. Drugs, 2021, 19, 397 Search PubMed.
- T. Li, C. Xi, Y. Yu, N. Wang, X. Wang, A. Iwasaki, F. Fang, L. Ding, S. Li, W. Zhang, Y. Yuan, T. Wang, X. Yan, S. He, Z. Cao and C. B. Naman, J. Nat. Prod., 2022, 85, 493–500 CrossRef CAS PubMed.
- A. Yamano, Y. Asato, N. Natsume, A. Iwasaki, K. Suenaga and T. Teruya, J. Nat. Prod., 2022, 85, 169–175 CrossRef CAS PubMed.
- K. Irie, G. Jeelani, T. Nozaki and A. Iwasaki, J. Nat. Prod., 2024, 87, 2292–2301 CrossRef CAS PubMed.
- N. Watanabe, G. Jeelani, T. Nozaki and A. Iwasaki, ACS Omega, 2024, 9, 36795–36801 CAS.
- B. Ryu, E. Glukhov, T. R. Teixeira, C. R. Caffrey, S. Madiyan, V. Joseph, N. E. Avalon, C. A. Leber, C. B. Naman and W. H. Gerwick, J. Nat. Prod., 2024, 87, 1601–1610 CrossRef CAS PubMed.
- B.-T. Zhang, H. Nishino, R. Kawabe, M. Kamio, R. Watanabe, H. Uchida, M. Satake and H. Nagai, Biosci., Biotechnol., Biochem., 2024, 88, 517–521 CrossRef CAS PubMed.
- G. J. Kim, S. J. Mascuch, E. Mevers, P. D. Boudreau, W. H. Gerwick and H. Choi, J. Org. Chem., 2022, 87, 1043–1055 CrossRef CAS PubMed.
- D. P. Fewer, J. Jokela, L. Heinilä, R. Aesoy, K. Sivonen, T. Galica, P. Hrouzek and L. Herfindal, Physiol. Plant., 2021, 173, 639–650 Search PubMed.
- L. M. P. Heinilä, D. P. Fewer, J. K. Jokela, M. Wahlsten, X. Ouyang, P. Permi, A. Jortikka and K. Sivonen, Org. Biomol. Chem., 2021, 19, 5577–5588 RSC.
- L. Darcel, S. Das, I. Bonnard, B. Banaigs and N. Inguimbert, Mar. Drugs, 2021, 19, 473 CrossRef CAS PubMed.
- J. J. Mehjabin, C.-S. Phan and T. Okino, J. Nat. Prod., 2024, 87, 984–993 CrossRef CAS PubMed.
- D. Overlingė, M. Cegłowska, R. Konkel and H. Mazur-Marzec, Mar. Drugs, 2024, 22, 506 CrossRef PubMed.
- S. W. Algor, A. Sukenik and S. Carmeli, Mar. Drugs, 2024, 22, 389 CrossRef PubMed.
- D. Overlingė, A. Toruńska-Sitarz, M. Cegłowska, K. Szubert and H. Mazur-Marzec, Sci. Rep., 2024, 14, 24686 CrossRef PubMed.
- R. Fernández, M. Pérez, A. Losada, S. Reboredo, A. G.-S. Juan, M. J. Martín, A. Francesch, S. Munt and C. Cuevas, Mar. Drugs, 2024, 22, 303 CrossRef PubMed.
- S. Matthew, Q.-Y. Chen, R. Ratnayake, C. S. Fermaintt, D. Lucena-Agell, F. Bonato, A. E. Prota, S. T. Lim, X. Wang, J. F. Díaz, A. L. Risinger, V. J. Paul, M. Á. Oliva and H. Luesch, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2021847118 CrossRef CAS PubMed.
- P. Marfey, Carlsberg Res. Commun., 1984, 49, 591–596 CrossRef CAS.
- T. Weinert, N. Olieric, R. Cheng, S. Brünle, D. James, D. Ozerov, D. Gashi, L. Vera, M. Marsh, K. Jaeger, F. Dworkowski, E. Panepucci, S. Basu, P. Skopintsev, A. S. Doré, T. Geng, R. M. Cooke, M. Liang, A. E. Prota, V. Panneels, P. Nogly, U. Ermler, G. Schertler, M. Hennig, M. O. Steinmetz, M. Wang and J. Standfuss, Nat. Commun., 2017, 8, 542 CrossRef PubMed.
- N. E. Avalon, M. A. Reis, C. C. Thornburg, R. T. Williamson, D. Petras, A. T. Aron, G. F. Neuhaus, M. Al-Hindy, J. Mitrevska, L. Ferreira, J. Morais, Y. E. Abiead, E. Glukhov, K. L. Alexander, F. A. Vulpanovici, M. J. Bertin, S. Whitner, H. Choi, G. Spengler, K. Blinov, A. M. Almohammadi, L. A. Shaala, W. R. Kew, L. Paša-Tolić, D. T. A. Youssef, P. C. Dorrestein, V. Vasconcelos, L. Gerwick, K. L. McPhail and W. H. Gerwick, J. Am. Chem. Soc., 2024, 146, 18626–18638 CrossRef CAS PubMed.
- T.-E. Ngo, A. Ecker, B. Ryu, A. Guild, A. Remmel, P. D. Boudreau, K. L. Alexander, C. B. Naman, E. Glukhov, N. E. Avalon, V. V. Shende, L. Thomas, S. Dahesh, V. Nizet, L. Gerwick and W. H. Gerwick, ACS Chem. Biol., 2024, 19, 619–628 CrossRef CAS PubMed.
- N. Kurisawa, G. Jeelani, T. Nozaki, A. L. Agusta, K. Suenaga and A. Iwasaki, J. Nat. Prod., 2024, 87, 1838–1843 CrossRef CAS PubMed.
- S. Kokkaliari, L. Grauso, A. Mangoni, G. Seabra, V. J. Paul and H. Luesch, Chem. – Eur. J., 2024, 30, e202401393 CrossRef CAS PubMed.
- S. Breinlinger, T. J. Phillips, B. N. Haram, J. Mareš, J. A. M. Yerena, P. Hrouzek, R. Sobotka, W. M. Henderson, P. Schmieder, S. M. Williams, J. D. Lauderdale, H. D. Wilde, W. Gerrin, A. Kust, J. W. Washington, C. Wagner, B. Geier, M. Liebeke, H. Enke, T. H. J. Niedermeyer and S. B. Wilde, Science, 2021, 371, eaax9050 CrossRef CAS PubMed.
- N. J. Thomas, C. U. Meteyer and L. Sileo, Vet. Pathol., 1998, 35, 479–487 CrossRef CAS PubMed.
- S. K. Williams, J. Kempton, S. B. Wilde and A. Lewitus, Harmful Algae, 2007, 6, 343–353 CrossRef CAS.
- S. B. Wilde, T. M. Murphy, C. P. Hope, S. K. Habrun, J. Kempton, A. Birrenkott, F. Wiley, W. W. Bowerman and A. J. Lewitus, Environ. Toxicol., 2005, 20, 348–353 Search PubMed.
- S. Adak, A. L. Lukowski, R. J. B. Schäfer and B. S. Moore, J. Am. Chem. Soc., 2022, 144, 2861–2866 Search PubMed.
- S. Gäfe and H. H. Niemann, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2023, 79, 596–609 CrossRef PubMed.
- H. Li, J.-W. Huang, L. Dai, H. Zheng, S. Dai, Q. Zhang, L. Yao, Y. Yang, Y. Yang, J. Min, R.-T. Guo and C.-C. Chen, Nat. Commun., 2023, 14, 7425 CrossRef PubMed.
- S. Adak, N. Ye, L. A. Calderone, M. Duan, W. Lubeck, R. J. B. Schäfer, A. L. Lukowski, K. N. Houk, M.-E. Pandelia, C. L. Drennan and B. S. Moore, Nat. Chem., 2024, 16, 1989–1998 CrossRef CAS PubMed.
- M. G. Ricardo, M. Schwark, D. Llanes, T. H. J. Niedermeyer and B. Westermann, Chem. – Eur. J., 2021, 27, 12032–12035 CrossRef CAS PubMed.
- L. Štenclová, S. B. Wilde, M. Schwark, J. L. Cullen, S. A. McWhorter, T. H. J. Niedermeyer, W. M. Henderson and J. Mareš, Harmful Algae, 2023, 125, 102425 CrossRef PubMed.
- M. Schwark, J. A. M. Yerena, K. Röhrborn, P. Hrouzek, P. Divoká, L. Štenclová, K. Delawská, H. Enke, C. Vorreiter, F. Wiley, W. Sippl, R. Sobotka, S. Saha, S. B. Wilde, J. Mareš and T. H. J. Niedermeyer, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2219230120 CrossRef CAS PubMed.
- N. R. Glasser, D. Cui, D. D. Risser, C. D. Okafor and E. P. Balskus, Nat. Chem., 2024, 16, 173–182 CrossRef CAS PubMed.
- H. Nakamura, H. A. Hamer, G. Sirasani and E. P. Balskus, J. Am. Chem. Soc., 2012, 134, 18518–18521 CrossRef CAS PubMed.
- E. E. Schultz, N. R. Braffman, M. U. Luescher, H. H. Hager and E. P. Balskus, Angew. Chem., Int. Ed., 2019, 58, 3151–3155 CrossRef CAS PubMed.
- P. N. Leão, H. Nakamura, M. Costa, A. R. Pereira, R. Martins, V. Vasconcelos, W. H. Gerwick and E. P. Balskus, Angew. Chem., Int. Ed., 2015, 54, 11063–11067 Search PubMed.
- T. B. Afonso, M. S. Costa, R. R. de Castro, S. Freitas, A. Silva, M. P. C. Schneider, R. Martins and P. N. Leão, J. Nat. Prod., 2016, 79, 2504–2513 Search PubMed.
- J. P. A. Reis, S. Freitas, T. Procházková and P. N. Leão, J. Nat. Prod., 2024, 87, 2709–2715 Search PubMed.
- J. P. A. Reis, S. A. C. Figueiredo, M. L. Sousa and P. N. Leão, Nat. Commun., 2020, 11, 1458 CrossRef CAS PubMed.
- D. Dehm, J. Krumbholz, M. Baunach, V. Wiebach, K. Hinrichs, A. Guljamow, T. Tabuchi, H. Jenke-Kodama, R. D. Süssmuth and E. Dittmann, ACS Chem. Biol., 2019, 14, 1271–1279 CrossRef CAS PubMed.
- J. Krumbholz, K. Ishida, M. Baunach, J. E. Teikari, M. M. Rose, S. Sasso, C. Hertweck and E. Dittmann, Angew. Chem., Int. Ed., 2022, 61, e202204545 CrossRef CAS PubMed.
- A. Guljamow, M. Kreische, K. Ishida, A. Liaimer, B. Altermark, L. Bähr, C. Hertweck, R. Ehwald and E. Dittmann, Appl. Environ. Microbiol., 2017, 83, e01510–e01517 CrossRef PubMed.
- A. G. Atanasov, S. B. Zotchev, V. M. Dirsch, I. E. Orhan, M. Banach, J. M. Rollinger, D. Barreca, W. Weckwerth, R. Bauer, E. A. Bayer, M. Majeed, A. Bishayee, V. Bochkov, G. K. Bonn, N. Braidy, F. Bucar, A. Cifuentes, G. D'Onofrio, M. Bodkin, M. Diederich, A. T. Dinkova-Kostova, T. Efferth, K. E. Bairi, N. Arkells, T.-P. Fan, B. L. Fiebich, M. Freissmuth, M. I. Georgiev, S. Gibbons, K. M. Godfrey, C. W. Gruber, J. Heer, L. A. Huber, E. Ibanez, A. Kijjoa, A. K. Kiss, A. Lu, F. A. Macias, M. J. S. Miller, A. Mocan, R. Müller, F. Nicoletti, G. Perry, V. Pittalà, L. Rastrelli, M. Ristow, G. L. Russo, A. S. Silva, D. Schuster, H. Sheridan, K. Skalicka-Woźniak, L. Skaltsounis, E. Sobarzo-Sánchez, D. S. Bredt, H. Stuppner, A. Sureda, N. T. Tzvetkov, R. A. Vacca, B. B. Aggarwal, M. Battino, F. Giampieri, M. Wink, J.-L. Wolfender, J. Xiao, A. W. K. Yeung, G. Lizard, M. A. Popp, M. Heinrich, I. Berindan-Neagoe, M. Stadler, M. Daglia, R. Verpoorte and C. T. Supuran, Nat. Rev. Drug Discovery, 2021, 20, 200–216 CrossRef CAS PubMed.
- T. F. Molinski, Nat. Prod. Rep., 2010, 27, 321–329 Search PubMed.
- D. S. Wishart, J. Magn. Reson., 2019, 306, 155–161 CrossRef CAS PubMed.
- S. Sieber, A. Carlier, M. Neuburger, G. Grabenweger, L. Eberl and K. Gademann, Angew. Chem., Int. Ed., 2015, 54, 7968–7970 CrossRef CAS PubMed.
- R. Taguchi, A. Iwasaki, A. Ebihara, G. Jeelani, T. Nozaki and K. Suenaga, Org. Lett., 2022, 24, 4710–4714 CrossRef CAS PubMed.
- N. Grimblat, M. M. Zanardi and A. M. Sarotti, J. Org. Chem., 2015, 80, 12526–12534 CrossRef CAS PubMed.
- N. Grimblat, J. A. Gavín, A. H. Daranas and A. M. Sarotti, Org. Lett., 2019, 21, 4003–4007 CrossRef CAS PubMed.
- W. Hehre, P. Klunzinger, B. Deppmeier, A. Driessen, N. Uchida, M. Hashimoto, E. Fukushi and Y. Takata, J. Nat. Prod., 2019, 82, 2299–2306 Search PubMed.
- S. G. Smith and J. M. Goodman, J. Am. Chem. Soc., 2010, 132, 12946–12959 CrossRef CAS PubMed.
- K. Umeda, A. Iwasaki, R. Taguchi, N. Kurisawa, G. Jeelani, T. Nozaki and K. Suenaga, J. Nat. Prod., 2023, 86, 2529–2538 Search PubMed.
- I. Ohtani, T. Kusumi, Y. Kashman and H. Kakisawa, J. Am. Chem. Soc., 1991, 113, 4092–4096 CrossRef CAS.
- M. Yotsu-Yamashita, K. Umeda, A. Iwasaki and K. Suenaga, Toxicon, 2024, 250, 108122 CrossRef CAS PubMed.
- F. H. Al-Awadhi, S. Kokkaliari, R. Ratnayake, V. J. Paul and H. Luesch, J. Nat. Prod., 2024, 87, 2355–2365 CrossRef CAS PubMed.
- O. Ohno, A. Iwasaki, K. Same, C. Kudo, E. Aida, K. Sugiura, S. Sumimoto, T. Teruya, E. Tashiro, S. Simizu, K. Matsuno, M. Imoto and K. Suenaga, Org. Lett., 2022, 24, 4547–4551 CrossRef CAS PubMed.
- Y. Miyamoto, A. Iwasaki, H. Fujimura, C. Kudo, N. Kurisawa, O. Ohno and K. Suenaga, J. Org. Chem., 2023, 88, 3208–3216 CrossRef CAS PubMed.
- S. A. C. Figueiredo, M. Preto, G. Moreira, T. P. Martins, K. Abt, A. Melo, V. M. Vasconcelos and P. N. Leão, Angew. Chem., Int. Ed., 2021, 60, 10064–10072 CrossRef CAS PubMed.
- K. Voráčová, J. Hájek, J. Mareš, P. Urajová, M. Kuzma, J. Cheel, A. Villunger, A. Kapuscik, M. Bally, P. Novák, M. Kabeláč, G. Krumschnabel, M. Lukeš, L. Voloshko, J. Kopecký and P. Hrouzek, PLoS One, 2017, 12, e0172850 CrossRef PubMed.
- I. Gutiérrez-del-Río, N. B. de Fraissinette, R. Castelo-Branco, F. Oliveira, J. Morais, S. Redondo-Blanco, C. J. Villar, M. J. Iglesias, R. Soengas, V. Cepas, Y. L. Cubillos, G. Sampietro, L. Rodolfi, F. Lombó, S. M. S. González, F. L. Ortiz, V. Vasconcelos and M. A. Reis, J. Nat. Prod., 2020, 83, 1885–1890 Search PubMed.
- T. J. O'Donnell, Y. Luo, W. Y. Yoshida, S. Suzuki, R. Sun and P. G. Williams, J. Nat. Prod., 2022, 85, 415–425 CrossRef PubMed.
- M. H. Medema, R. Kottmann, P. Yilmaz, M. Cummings, J. B. Biggins, K. Blin, I. de Bruijn, Y. H. Chooi, J. Claesen, R. C. Coates, P. Cruz-Morales, S. Duddela, S. Düsterhus, D. J. Edwards, D. P. Fewer, N. Garg, C. Geiger, J. P. Gomez-Escribano, A. Greule, M. Hadjithomas, A. S. Haines, E. J. N. Helfrich, M. L. Hillwig, K. Ishida, A. C. Jones, C. S. Jones, K. Jungmann, C. Kegler, H. U. Kim, P. Kötter, D. Krug, J. Masschelein, A. V. Melnik, S. M. Mantovani, E. A. Monroe, M. Moore, N. Moss, H.-W. Nützmann, G. Pan, A. Pati, D. Petras, F. J. Reen, F. Rosconi, Z. Rui, Z. Tian, N. J. Tobias, Y. Tsunematsu, P. Wiemann, E. Wyckoff, X. Yan, G. Yim, F. Yu, Y. Xie, B. Aigle, A. K. Apel, C. J. Balibar, E. P. Balskus, F. Barona-Gómez, A. Bechthold, H. B. Bode, R. Borriss, S. F. Brady, A. A. Brakhage, P. Caffrey, Y.-Q. Cheng, J. Clardy, R. J. Cox, R. D. Mot, S. Donadio, M. S. Donia, W. A. van der Donk, P. C. Dorrestein, S. Doyle, A. J. M. Driessen, M. Ehling-Schulz, K.-D. Entian, M. A. Fischbach, L. Gerwick, W. H. Gerwick, H. Gross, B. Gust, C. Hertweck, M. Höfte, S. E. Jensen, J. Ju, L. Katz, L. Kaysser, J. L. Klassen, N. P. Keller, J. Kormanec, O. P. Kuipers, T. Kuzuyama, N. C. Kyrpides, H.-J. Kwon, S. Lautru, R. Lavigne, C. Y. Lee, B. Linquan, X. Liu, W. Liu, A. Luzhetskyy, T. Mahmud, Y. Mast, C. Méndez, M. Metsä-Ketelä, J. Micklefield, D. A. Mitchell, B. S. Moore, L. M. Moreira, R. Müller, B. A. Neilan, M. Nett, J. Nielsen, F. O'Gara, H. Oikawa, A. Osbourn, M. S. Osburne, B. Ostash, S. M. Payne, J.-L. Pernodet, M. Petricek, J. Piel, O. Ploux, J. M. Raaijmakers, J. A. Salas, E. K. Schmitt, B. Scott, R. F. Seipke, B. Shen, D. H. Sherman, K. Sivonen, M. J. Smanski, M. Sosio, E. Stegmann, R. D. Süssmuth, K. Tahlan, C. M. Thomas, Y. Tang, A. W. Truman, M. Viaud, J. D. Walton, C. T. Walsh, T. Weber, G. P. van Wezel, B. Wilkinson, J. M. Willey, W. Wohlleben, G. D. Wright, N. Ziemert, C. Zhang, S. B. Zotchev, R. Breitling, E. Takano and F. O. Glöckner, Nat. Chem. Biol., 2015, 11, 625–631 CrossRef CAS PubMed.
- S. A. Kautsar, K. Blin, S. Shaw, J. C. Navarro-Muñoz, B. R. Terlouw, J. J. J. van der Hooft, J. A. van Santen, V. Tracanna, H. G. Suarez Duran, V. Pascal Andreu, N. Selem-Mojica, M. Alanjary, S. L. Robinson, G. Lund, S. C. Epstein, A. C. Sisto, L. K. Charkoudian, J. Collemare, R. G. Linington, T. Weber and M. H. Medema, Nucleic Acids Res., 2019, 48, D454–D458 Search PubMed.
- B. R. Terlouw, K. Blin, J. C. Navarro-Muñoz, N. E. Avalon, M. G. Chevrette, S. Egbert, S. Lee, D. Meijer, M. J. J. Recchia, Z. L. Reitz, J. A. van Santen, N. Selem-Mojica, T. Tørring, L. Zaroubi, M. Alanjary, G. Aleti, C. Aguilar, S. A. A. Al-Salihi, H. E. Augustijn, J. A. Avelar-Rivas, L. A. Avitia-Domínguez, F. Barona-Gómez, J. Bernaldo-Agüero, V. A. Bielinski, F. Biermann, T. J. Booth, V. J. C. Bravo, R. Castelo-Branco, F. O. Chagas, P. Cruz-Morales, C. Du, K. R. Duncan, A. Gavriilidou, D. Gayrard, K. Gutiérrez-García, K. Haslinger, E. J. N. Helfrich, J. J. J. van der Hooft, A. P. Jati, E. Kalkreuter, N. Kalyvas, K. B. Kang, S. Kautsar, W. Kim, A. M. Kunjapur, Y.-X. Li, G.-M. Lin, C. Loureiro, J. J. R. Louwen, N. L. L. Louwen, G. Lund, J. Parra, B. Philmus, B. Pourmohsenin, L. J. U. Pronk, A. Rego, D. A. B. Rex, S. Robinson, L. R. Rosas-Becerra, E. T. Roxborough, M. A. Schorn, D. J. Scobie, K. S. Singh, N. Sokolova, X. Tang, D. Udwary, A. Vigneshwari, K. Vind, S. P. J. M. Vromans, V. Waschulin, S. E. Williams, J. M. Winter, T. E. Witte, H. Xie, D. Yang, J. Yu, M. Zdouc, Z. Zhong, J. Collemare, R. G. Linington, T. Weber and M. H. Medema, Nucleic Acids Res., 2022, 51, D603–D610 CrossRef PubMed.
- M. M. Zdouc, K. Blin, N. L. L. Louwen, J. Navarro, C. Loureiro, C. D. Bader, C. B. Bailey, L. Barra, T. J. Booth, K. A. J. Bozhüyük, J. D. D. Cediel-Becerra, Z. Charlop-Powers, M. G. Chevrette, Y. H. Chooi, P. M. D'Agostino, T. de Rond, E. D. Pup, K. R. Duncan, W. Gu, N. Hanif, E. J. N. Helfrich, M. Jenner, Y. Katsuyama, A. Korenskaia, D. Krug, V. Libis, G. A. Lund, S. Mantri, K. D. Morgan, C. Owen, C.-S. Phan, B. Philmus, Z. L. Reitz, S. L. Robinson, K. S. Singh, R. Teufel, Y. Tong, F. Tugizimana, D. Ulanova, J. M. Winter, C. Aguilar, D. Y. Akiyama, S. A. A. Al-Salihi, M. Alanjary, F. Alberti, G. Aleti, S. A. Alharthi, M. Y. A. Rojo, A. A. Arishi, H. E. Augustijn, N. E. Avalon, J. A. Avelar-Rivas, K. K. Axt, H. B. Barbieri, J. C. J. Barbosa, L. G. B. Segato, S. E. Barrett, M. Baunach, C. Beemelmanns, D. Beqaj, T. Berger, J. Bernaldo-Agüero, S. M. Bettenbühl, V. A. Bielinski, F. Biermann, R. M. Borges, R. Borriss, M. Breitenbach, K. M. Bretscher, M. W. Brigham, L. Buedenbender, B. W. Bulcock, C. Cano-Prieto, J. Capela, V. J. Carrion, R. S. Carter, R. Castelo-Branco, G. Castro-Falcón, F. O. Chagas, E. Charria-Girón, A. A. Chaudhri, V. Chaudhry, H. Choi, Y. Choi, R. Choupannejad, J. Chromy, M. S. C. Donahey, J. Collemare, J. A. Connolly, K. E. Creamer, M. Crüsemann, A. A. Cruz, A. Cumsille, J.-F. Dallery, L. C. Damas-Ramos, T. Damiani, M. de Kruijff, B. D. Martín, G. D. Sala, J. Dillen, D. T. Doering, S. R. Dommaraju, S. Durusu, S. Egbert, M. Ellerhorst, B. Faussurier, A. Fetter, M. Feuermann, D. P. Fewer, J. Foldi, A. Frediansyah, E. A. Garza, A. Gavriilidou, A. Gentile, J. Gerke, H. Gerstmans, J. P. Gomez-Escribano, L. A. González-Salazar, N. E. Grayson, C. Greco, J. E. G. Gomez, S. Guerra, S. G. Flores, A. Gurevich, K. Gutiérrez-García, L. Hart, K. Haslinger, B. He, T. Hebra, J. L. Hemmann, H. Hindra, L. Höing, D. C. Holland, J. E. Holme, T. Horch, P. Hrab, J. Hu, T.-H. Huynh, J.-Y. Hwang, R. Iacovelli, D. Iftime, M. Iorio, S. Jayachandran, E. Jeong, J. Jing, J. J. Jung, Y. Kakumu, E. Kalkreuter, K. B. Kang, S. Kang, W. Kim, G. J. Kim, H. Kim, H. U. Kim, M. Klapper, R. A. Koetsier, C. Kollten, Á. T. Kovács, Y. Kriukova, N. Kubach, A. M. Kunjapur, A. K. Kushnareva, A. Kust, J. Lamber, M. Larralde, N. J. Larsen, A. P. Launay, N.-T.-H. Le, S. Lebeer, B. T. Lee, K. Lee, K. L. Lev, S.-M. Li, Y.-X. Li, C. Licona-Cassani, A. Lien, J. Liu, J. A. V. Lopez, N. V. Machushynets, M. I. Macias, T. Mahmud, M. Maleckis, A. M. Martinez-Martinez, Y. Mast, M. F. Maximo, C. M. McBride, R. M. McLellan, K. M. Bhatt, C. Melkonian, A. Merrild, M. Metsä-Ketelä, D. A. Mitchell, A. V. Müller, G.-S. Nguyen, H. T. Nguyen, T. H. J. Niedermeyer, J. H. O'Hare, A. Ossowicki, B. O. Ostash, H. Otani, L. Padva, S. Paliyal, X. Pan, M. Panghal, D. S. Parade, J. Park, J. Parra, M. P. Rubio, H. T. Pham, S. J. Pidot, J. Piel, B. Pourmohsenin, M. Rakhmanov, S. Ramesh, M. H. Rasmussen, A. Rego, R. Reher, A. J. Rice, A. Rigolet, A. Romero-Otero, L. R. Rosas-Becerra, P. Y. Rosiles, A. Rutz, B. Ryu, L.-A. Sahadeo, M. Saldanha, L. Salvi, E. Sánchez-Carvajal, C. Santos-Medellin, N. Sbaraini, S. M. Schoellhorn, C. Schumm, L. Sehnal, N. Selem, A. D. Shah, T. K. Shishido, S. Sieber, V. Silviani, G. Singh, H. Singh, N. Sokolova, E. C. Sonnenschein, M. Sosio, S. T. Sowa, K. Steffen, E. Stegmann, A. B. Streiff, A. Strüder, F. Surup, T. Svenningsen, D. Sweeney, J. Szenei, A. Tagirdzhanov, B. Tan, M. J. Tarnowski, B. R. Terlouw, T. Rey, N. U. Thome, L. R. T. Ortega, T. Tørring, M. Trindade, A. W. Truman, M. Tvilum, D. W. Udwary, C. Ulbricht, L. Vader, G. P. van Wezel, M. Walmsley, R. Warnasinghe, H. G. Weddeling, A. N. M. Weir, K. Williams, S. E. Williams, T. E. Witte, S. M. W. Rocca, K. Yamada, D. Yang, D. Yang, J. Yu, Z. Zhou, N. Ziemert, L. Zimmer, A. Zimmermann, C. Zimmermann, J. J. J. van der Hooft, R. G. Linington, T. Weber and M. H. Medema, Nucleic Acids Res., 2024, 53, D678–D690 CrossRef PubMed.
- K. Palaniappan, I.-M. A. Chen, K. Chu, A. Ratner, R. Seshadri, N. C. Kyrpides, N. N. Ivanova and N. J. Mouncey, Nucleic Acids Res., 2019, 48, D422–D430 Search PubMed.
- S. A. Kautsar, K. Blin, S. Shaw, T. Weber and M. H. Medema, Nucleic Acids Res., 2020, 49, D490–D497 CrossRef PubMed.
- K. Blin, M. H. Medema, R. Kottmann, S. Y. Lee and T. Weber, Nucleic Acids Res., 2017, 45, D555–D559 CrossRef CAS PubMed.
- K. Blin, S. Shaw, A. M. Kloosterman, Z. Charlop-Powers, G. P. van Wezel, M. H. Medema and T. Weber, Nucleic Acids Res., 2021, 49, W29–W35 CrossRef CAS PubMed.
- A. Rutz, D. Probst, C. Aguilar, D. Y. Akiyama, F. Alberti, H. E. Augustijn, N. E. Avalon, C. Beemelmanns, H. B. Barbieri, F. Biermann, A. J. Bridge, E. C. Girón, R. Cox, M. Crüsemann, P. M. D'Agostino, M. Feuermann, J. Gerke, K. G. García, J. E. Holme, J.-Y. Hwang, R. Iacovelli, J. C. J. Barbosa, N. Kaur, M. Klapper, A. M. Köhler, A. Korenskaia, N. Kubach, B. T. Lee, C. Loureiro, S. Mantri, S. Narula, D. Meijer, J. C. Navarro-Muñoz, G.-S. Nguyen, S. Paliyal, M. Panghal, L. Rao, S. Sieber, N. Sokolova, S. T. Sowa, J. Szenei, B. R. Terlouw, H. G. Weddeling, J. Yu, N. Ziemert, T. Weber, K. Blin, J. J. J. van der Hooft, M. H. Medema and M. M. Zdouc, Nucleic Acids Res., 2025, gkaf969 Search PubMed.
- J. D. Rudolf, T. A. Alsup, B. Xu and Z. Li, Nat. Prod. Rep., 2020, 38, 905–980 RSC.
- J. R. Gurr, T. J. O'Donnell, Y. Luo, W. Y. Yoshida, M. L. Hall, A. M. S. Mayer, R. Sun and P. G. Williams, J. Nat. Prod., 2020, 83, 1691–1695 CrossRef CAS PubMed.
- D. Back, T. J. O'Donnell, K. K. Axt, J. R. Gurr, J. M. Vanegas, P. G. Williams and B. Philmus, ACS Chem. Biol., 2023, 18, 1797–1807 CrossRef CAS PubMed.
- R.-A. Hughes, Y. Zhang, R. Zhang, P. G. Williams, J. S. Lindsey and E. S. Miller, Appl. Environ. Microbiol., 2017, 83, e01068–17 CAS.
- X. Jin, E. S. Miller and J. S. Lindsey, Life, 2021, 11, 356 CrossRef CAS PubMed.
- A. Ebihara, R. Taguchi, G. Jeelani, T. Nozaki, K. Suenaga and A. Iwasaki, J. Nat. Prod., 2024, 87, 1116–1123 CrossRef CAS PubMed.
- F. I. Saldívar-González, V. D. Aldas-Bulos, J. L. Medina-Franco and F. Plisson, Chem. Sci., 2021, 13, 1526–1546 RSC.
- A. Ebihara, A. Iwasaki, Y. Miura, G. Jeelani, T. Nozaki and K. Suenaga, J. Org. Chem., 2021, 86, 11763–11770 CrossRef CAS PubMed.
- R. Reher, H. W. Kim, C. Zhang, H. H. Mao, M. Wang, L.-F. Nothias, A. M. Caraballo-Rodriguez, E. Glukhov, B. Teke, T. Leao, K. L. Alexander, B. M. Duggan, E. L. V. Everbroeck, P. C. Dorrestein, G. W. Cottrell and W. H. Gerwick, J. Am. Chem. Soc., 2020, 142, 4114–4120 CrossRef CAS PubMed.
- N. L. Drake, H. C. Harris and C. B. Jaeger, J. Am. Chem. Soc., 1948, 70, 168–171 CrossRef CAS PubMed.
- N. Kurisawa, A. Iwasaki, K. Teranuma, S. Dan, C. Toyoshima, M. Hashimoto and K. Suenaga, J. Am. Chem. Soc., 2022, 144, 11019–11032 CrossRef CAS PubMed.
- D. A. Evans, D. L. Rieger, M. T. Bilodeau and F. Urpi, J. Am. Chem. Soc., 1991, 113, 1047–1049 CrossRef CAS.
- Y. K. Zhang, M. A. Sanchez-Ayala, P. W. Sternberg, J. Srinivasan and F. C. Schroeder, Org. Lett., 2017, 19, 2837–2840 CrossRef CAS PubMed.
- S. Kokkaliari, D. Luo, V. J. Paul and H. Luesch, Mar. Drugs, 2023, 21, 378 CrossRef CAS PubMed.
- N. Kurisawa, K. Teranuma, A. Noto, A. Iwasaki, Y. Kabashima, R. Nakajima, C. Toyoshima and K. Suenaga, Bull. Chem. Soc. Jpn., 2024, 97, uoae070 CrossRef CAS.
- D. Kong and T. Yamori, Bioorg. Med. Chem., 2012, 20, 1947–1951 CrossRef CAS PubMed.
- D. Trauner and K.-P. Ruehmann, Synfacts, 2022, 18, 1028 CrossRef.
- N. Kurisawa, A. Iwasaki and K. Suenaga, J. Synth. Org. Chem., Jpn., 2024, 82, 1107–1116 CrossRef CAS.
- H. Takahashi, A. Iwasaki, A. Ebihara, R. Taguchi, G. Jeelani, T. Nozaki and K. Suenaga, Org. Lett., 2023, 25, 2400–2404 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |

