 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Glucose 6-phosphate: the diversity of C-methylation in sugar moieties within natural product biosynthesis†
Ziruo Zou and
Michael Müller
and
Michael Müller *
*
Institute of Pharmaceutical Sciences, University of Freiburg, Albertstrasse 25, 79104 Freiburg, Germany. E-mail: michael.mueller@pharmazie.uni-freiburg.de
First published on 16th March 2026
Abstract
Covering: Up to 2026
Methylation is one of the most frequent and functionally significant modifications in natural product biosynthesis. This transformation generally involves the cosubstrate S-adenosyl-L-methionine (SAM) and occurs with high regio-, chemo-, and stereo-selectivity. C-Methylation serves as an effective strategy for the incorporation of C1 building blocks and plays a key role in both carbon skeleton extension and structural core decoration. A notable example is SAM-dependent C-methylation of nucleotide-activated sugar moieties catalyzed by C-methyltransferase (C-MT) during the biosynthesis of glycosylated natural products. A prerequisite in these pathways is the activation of the sugar by the attachment of a nucleotide diphosphate (NDP) tail. Sugar C-MTs exhibit remarkable substrate specificity and often act in concert with additional transformations, such as oxidation, decarboxylation, or reduction, to diversify sugar structures. Canonical SAM-dependent sugar C-MTs are capable of introducing methyl groups at C3, C4, or C5 positions of the sugar ring by an SN2-like mechanism involving enolate intermediates. In contrast, C6 methylation has been attributed to radical SAM enzymes. These distinct strategies illustrate the enzymatic versatility and sophisticated control over sugar C-methylation in nature, contributing to the structural diversity of natural products.
1. Introduction
Methylation is one of the most common enzymatic reactions in nature, especially in the specialized metabolic pathways of many organisms, where it regulates biological activity. The enzymes responsible for methylation reactions are referred to as methyltransferases (MTs). The widely studied cosubstrate S-adenosyl-L-methionine (SAM) serves as a methyl donor and enables methylation either via a classical SN2 mechanism or a radical-based process.1 SAM-dependent MTs catalyze the O-, N-, C-, and S-methylation of small molecules, such as those found in amino acids and in polymeric biosynthetic scaffolds, such as DNA and proteins, with high regio- and stereo-selectivity.2–4SAM-dependent C-methylation plays a pivotal role in the structural diversification of biomolecules by introducing the C1 building block into the carbon skeleton. This transformation usually requires activation of the substrate for SAM attack. Such reactivity is commonly found in substrates with easily enolizable structures, such as carbonyl-containing compounds or in electron-rich aromatic systems, including, but not limited to, deprotonated phenols.5 This precise addition of C1 units is essential for the synthesis of chiral compounds as an integral tool for the asymmetric formation of C–C bonds. C-Methyltransferases (C-MTs) operating in a SAM-dependent manner are primarily involved in specific metabolic pathways and methylate various compounds, including pyrroloindoles,6 aliphatic compounds,7 phenols,8,9 and keto-deoxy sugars.10,11 A key area where C-MTs have a significant impact is in the production of branched-chain methylated deoxy sugars. These sugars are often formed in connection with the biosynthesis of polyketides,12 nonribosomal peptides,13 lipopolysaccharides,14 and oligosaccharides.15
In this review, we summarize the current knowledge on keto-deoxy sugar C-MTs, with a particular focus on their substrate specificities and chemical performances in methylation. Some key examples from natural product biosynthetic pathways are discussed. An overview of the broader nucleotide-sugar methylation landscape is presented to identify general features of sugar biosynthesis and structural modifications. Ultimately, this review seeks to elucidate the unique catalytic capabilities of these enzymes for methyl transfer and their contribution to the structural and functional diversity of natural products.
2. Sugar units in glycosylated natural products
Glycosylated natural products are a diverse and biologically important class of biomolecules used in medicine, agriculture, and industry.16,17 The incorporation of unusual sugar units into natural products, which are often deoxygenated and highly functionalized, improves the water-solubility of the aglycone and facilitates transport and bioavailability, making them an important starting point for the development of new therapeutics.18From a biosynthetic point of view, monosaccharides are, in principle, activated by attachment to nucleotide monophosphate (NMP) or nucleotide diphosphate (NDP). This activation facilitates their recognition by biosynthetic tailoring enzymes and glycosyltransferases and enables their incorporation into complex biomolecules within the cell. Moreover, the NMP/NDP tail acts as a good leaving group in the glycosyl transfer reaction.19
Different metabolic pathways diverging from glycolysis serve as the starting point for the biosynthesis of sugar units, such as avilamycin A,20 a polyketide-derived antibiotic with a heptasaccharide side chain (Scheme 1). The sugar components of this oligosaccharide antibiotic are thought to originate from three primary monosaccharide units during biosynthesis: guanosine diphosphate (GDP) sugars, which are derived from fructose 6-phosphate; thymidine diphosphate (TDP) sugars and uridine diphosphate (UDP) sugars, predominantly derived from glucose 6-phosphate. Before being incorporated into the main skeleton, these monosaccharides undergo various modifications, such as C-acetylation21 and methylation,22 which contribute to increasing the structural diversity and modifying the functional properties of the final product avilamycin A.
3. Methylation in sugar biosynthesis
3.1 C3 methylation
So far, conventional methyl transfer has only been observed with TDP- and GDP-activated monosaccharides. The presence of an adjacent keto group on the substrate enables SAM-dependent C-methylation by sugar C-MTs. This process begins with deprotonation, leading to enolization, and culminates in nucleophilic attack on SAM via an SN2 mechanism.11 Notably, the site of deprotonation determines the regioselectivity of methylation, allowing for precise modification at the C3, C4, or C5 positions on the sugar ring.C3-Methyltransferases (C3-MTs) targeting NDP-activated sugars are among the most prevalent enzymes in the biosynthetic pathways of natural products in prokaryotic systems. To date, at least seven distinct groups of C3-MTs have been reported, which are primarily classified based on the structural diversity of the methylated sugar products. The strict substrate specificity and precise stereochemical outcomes reflect the remarkable diversity of bacterial metabolic pathways.
In the biosynthesis of glycosylated pyrrolo[1,4]benzodiazepine sibiromycin in Streptosporangium sibiricum and glycosylated anthracycline nogalamycin in Streptomyces nogalater, respectively, SibM7 and SnogG2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 23 catalyze C3 methylation on TDP-4-keto-L-rhamnose (Fig. 1A). Notably, the sibiromycin biosynthetic gene cluster (BGC) lacks a gene encoding TDP-glucose 4,6-dehydratase necessary for substrate formation; this deficiency may be compensated for by an endogenous TDP-glucose dehydratase present in Streptosporangium sibiricum.9
23 catalyze C3 methylation on TDP-4-keto-L-rhamnose (Fig. 1A). Notably, the sibiromycin biosynthetic gene cluster (BGC) lacks a gene encoding TDP-glucose 4,6-dehydratase necessary for substrate formation; this deficiency may be compensated for by an endogenous TDP-glucose dehydratase present in Streptosporangium sibiricum.9
Maduropeptin, a glycosylated natural product derived from Actinomadura madurae ATCC 39144, is known for its potent DNA-damaging activity. The enediyne core is highly reactive and responsible for its biological activity; the glycosyl moieties improve solubility and stability.24,25 The biosynthesis of the aminosugar in maduropeptin starts with glucose 1-phosphate; it undergoes unique C6-dehydrogenation and oxidative decarboxylation before methylation. This ring-contraction pathway is common for the modification of UDP sugars,26–28 but rare for TDP sugars (Scheme 2A). It is assumed that the following methylation mediated by MdpA4 (Fig. 1B) leads to a remarkable stereochemical inversion at C3 of TDP-4-keto-pentose, where the C3 hydroxyl group is converted from an axial to an equatorial position.29 In chrysomycin biosynthesis, ChryCMT (Fig. 1C) is thought to catalyze a similar C3 methylation of TDP-4-keto-6-deoxy-D-glucose,30 because of its 50–60% sequence identity with other C3-MTs. Although its enzymatic activity has not yet been experimentally confirmed, it is presumed to follow a similar stereochemical pattern to MdpA4, including the rearrangement of the C3 hydroxyl group from an equatorial to an axial orientation during catalysis.
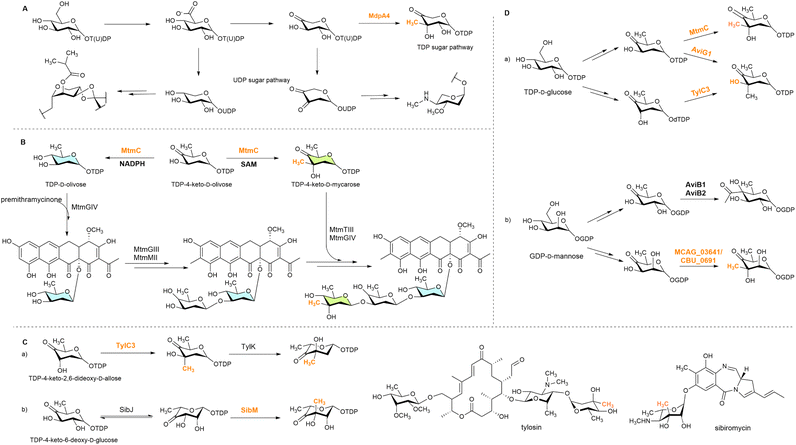 | ||
| Scheme 2 Representative C3 methylation during natural product biosynthesis: (A) UDP sugar pathway of UDP-4-keto-6-deoxy-D-glucose and MdpA4-joined TDP sugar C-methylation; (B) bifunctional MT MtmC involved in two branches of the biosynthetic step;32 (C) two consecutive steps of the C3 methylation and epimerization in the biosynthesis of tylosin (a) and sibiromycin (b). (D) C3 methylation in the TDP and GDP sugar pathways: (a) AviG1 and TylC3 produce TDP-4-keto-2,6-dideoxy-3-methyl-D-allose starting from different substrates and (b) GDP-D-mannose is used for the synthesis of two keto-deoxy sugars for oxidative decarboxylation and methylation, respectively. | ||
MtmC in mithramycin trisaccharide biosynthesis methylates TDP-4-keto-D-olivose in the same stereoselective manner to generate the equatorial-methylated product (Fig. 1D).31,32 The same methylation pattern can be observed in the biosynthesis of chromomycin, a member of the aureolic acid family produced by Streptomyces griseus.33 The crystal structure of MtmC, in complex with the byproduct SAH and the substrate TDP-4-keto-D-olivose, was solved by Jürgen Rohr and colleagues.34 In vitro enzyme assays and molecular docking results show that NADPH can effectively bind within the active site, indicating that MtmC allows different biosynthetic pathways to produce two building blocks, either by reduction to form TDP-D-olivose or by methylation to generate TDP-4-keto-D-mycarose. Both products are essential components of the trisaccharide assembly line (Scheme 2B).
TylC3—the first streptomycete C-MT characterized in vitro in the biosynthesis of tylosin—utilizes TDP-4-keto-2,6-dideoxy-D-allose as its substrate.35,36 Interestingly, methylation occurs at a TDP-deoxy sugar with an axial C3 hydroxyl substituent, a feature that distinguishes this pathway from those catalyzed by other C3-MTs. This axial orientation is established prior to methylation through the combined action of TylX3 and TylC1, along with a metal-assisted dehydroxylation step.36 In contrast to C3-MT pathways involving SibM and SnogG2, where 3,5-epimerization of the sugar scaffold precedes methylation, the epimerization in the tylosin biosynthetic pathway occurs after methylation at C3 (Scheme 2C). This sequential transformation leads to a similar skeletal structure, but results in a distinct stereocenter at C3. A comparable methylation pattern is also proposed in the biosynthesis of lankamycin37 and erythromycin A.38 The C-methylated sugar units are further decorated with O-methylation and/or O-acetylation before incorporation into the iconic polyketide core.
For the biosynthesis of avilamycin in Streptomyces viridochromogenes Tü57, the C3-MT AviG1 shows a contrasting stereochemical behavior compared to MtmC (Scheme 2D).20,22 Oxidoreductase AviT is located in the same BGC as AviG1 and is responsible for the formation of the methylation substrate. It exhibits a significant sequence similarity with NDP-3-ketoreductase LanT from Streptomyces cyanogenus39 (52% identical aa) and CmmW from Streptomyces griseus33 (46.8% identical aa). The similarity between AviT and LanT/CmmW suggests that AviG1 favors the same keto-deoxy sugar substrate, TDP-4-keto-D-olivose, as CmmC, indicating that AviG1 and TylC3 follow different stereocatalytic mechanisms but ultimately generate the same products (Scheme 2D).
Another potential substrate featuring a structurally similar sugar moiety for methylation in the avilamycin biosynthesis pathway is GDP-4-keto-6-deoxy-D-glucose. Thiamine diphosphate (ThDP)-dependent enzymes AviB1 and AviB2 are hypothesized to catalyze the acetylation of this GDP-sugar (Scheme 2D).20
Derived from the same precursor as GDP-4-keto-6-deoxy-D-glucose, GDP-4-keto-6-deoxy-D-mannose stands out as the only known keto-deoxy sugar with a GDP tail that undergoes keto-activated C-methylation. The putative MT MCAG_03641 performs C3 methyl transfer to GDP-4-keto-6-deoxy-D-mannose (Fig. 1G), preserving the stereochemistry of the C3 hydroxyl group during the biosynthesis of everninomicin 13-384-1 (everninomicin A, ziracin) in Micromonospora carbonacea var. africana ATCC39149.40,41 A similar process occurs in the biosynthesis of everninomicin B41 and flambamycin.42 Additionally, an MT homologous to MCAG_03641, CBU_0691, was identified by in silico analysis in Coxiella burnetii. This enzyme appears to methylate GDP-4-keto-6-deoxy-D-mannose in the same way as MCAG_03641 and may contribute to lipopolysaccharide biosynthesis.14 Interestingly, everninomicin D, a structurally related natural product from Micromonospora sp. SCSIO 07395, lacks this axial hydroxyl group on ring D compared to everninomicin 13-384-1 and may instead undergo a TDP sugar methylation pathway involving C-MT EvdM3 (Scheme 3A).43
 | ||
| Scheme 3 (A) Structures of everninomicin 13-384-1 and everninomicin D. (B)(a) Methylation of TDP-3-amino-2,3,6-trideoxy-4-keto-D-glucose and the variations of the nitrosugars in natural products. (b) Proposed catalytic mechanism of TcaB9 (PDB ID: 4E32).10 | ||
The family of nitrosugar-containing natural products includes a variety of compounds, such as tetrocarcin,44 lobophorin,45 arisostatin A,46 and rubradirin.47 Although multiple C3-methylated nitrosugar variants are observed among these natural products, methylation uniformly occurs at the amino sugar stage (Scheme 3B). In the biosynthesis of tetrocarcin, TcaB9 catalyzes the transfer of a methyl group from SAM to TDP-3-amino-2,3,6-trideoxy-4-keto-D-glucose at C3 (Fig. 1F).11,48 An X-ray crystallographic model of TcaB9 in complex with its substrate reveals that the reaction proceeds with retention of the axial configuration at C3–OH. In this process, His 225 potentially plays a critical role in facilitating methyl transfer (Scheme 3B). However, the identity of the active site base responsible for deprotonation remains unclear. Compared to other C3-MT crystal structures, such as KijD1 (PDB ID: 5T67) and MtmC (PDB ID: 4RVH), the amino acid residues distribution in the active site is highly conserved. Moreover, there is no definitive evidence to date as to whether other C3-MTs, such as KijD1,49,50 EvaC,51 DvaC (also named as OrfX),13 and EvdM8,43 also methylate the same substrate as TcaB9, as their specific substrates have not yet been identified.
3.2 C4 methylation
CyaA3, identified in the biosynthetic pathway of cyanosporaside A in Salinispora pacifica CNS-143, stands out as a canonical SAM-dependent C-MT known to catalyze C4 methylation of TDP-4-keto-6-deoxy-D-glucose.52 Sequence analysis reveals that CyaA3 exhibits less than 40% identity with other sugar C-MTs, distinguishing it from previously characterized enzymes. This divergence suggests an independent evolutionary trajectory of CyaA3 and implies a potentially unique catalytic mode for C4 methyl transfer. It is proposed to facilitate the H-3′ deprotonation at TDP-4-keto-6-deoxy-D-glucose, which is followed by enolization and hydrogen transfer, ultimately leading to nucleophilic methylation and ensuring regioselective methylation at C4 (Scheme 4A). | ||
| Scheme 4 (A) CyaA3-catalyzed C4 methylation with a proposed H+-transfer mechanism. (B) C4 methylation in the biosynthesis of gentamicin by GenD1. | ||
Radical SAM-dependent MTs have also been implicated in C4 methylation of keto-deoxy sugars. GenD1—a cobalamin (Cbl)-dependent class B radical MT is involved in the biosynthesis of the aminoglycoside antibiotic gentamicin from Micromonospora echinospora (Scheme 4B).53,54 Its role was confirmed by gene disruption and in vitro enzyme assay, as a C4-MT that transforms gentamicin A into gentamicin X2. Notably, GenD1 is characterized by high substrate flexibility, as it can utilize multiple compounds, including 3″-dehydro-3″-oxo-gentamicin A2 (DAA2), gentamicin A, gentamicin A-2, and gentamicin Ae to generate gentamicin A-1, gentamicin X2, gentamicin X2-1, and G418 (Scheme 5).53 This versatility highlights its critical role in the diverse and flexible biosynthetic pathways leading to gentamicin production.
 | ||
| Scheme 5 Network of methyl transfer reactions in the gentamicin biosynthetic pathway and the proposed methyl transfer mechanism of GenK.53,60 | ||
3.3 C5 methylation
In addition to the remarkable diversity of naturally occurring C3-MTs, several C5-MTs have also been identified by bioinformatic analysis and in vitro activity assays. The function of a C5-MT CloU was identified via in-frame gene inactivation within the BGC of clorobiocin, demonstrating that its gene product performs C5 methylation on the keto-deoxy sugar after 3,5-double epimerization of TDP-4-keto-6-deoxy-D-glucose by CloW (Scheme 6A).10,55 In 2005, NovU, a C-MT from Streptomyces spheroides involved in novobiocin biosynthesis, was the first sugar C5-MT to be functionally characterized in vitro.56 Its homologs SgcA3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 57,58 and CouU,55 are deduced to transfer the methyl group similarly and show 58.5% and 86.0% sequence identity to NovU, respectively.
57,58 and CouU,55 are deduced to transfer the methyl group similarly and show 58.5% and 86.0% sequence identity to NovU, respectively.
Furthermore, it is possible to incorporate a methyl group into GDP sugars. The BGC of tiacumicin contains the putative GDP-specific C5-MT TiaS2,59 which potentially methylates the same substrate as C3-MT MCAG_03641 and CBU_0691 (Scheme 6B), namely GDP-4-keto-6-deoxy-D-mannose, but the stereochemistry of the methylation remains elusive.
3.4 C6 methylation
To date, only Cbl-dependent MTs have been shown to catalyze C6 methylation of sugar moieties. This type of methylation proceeds via a radical mechanism and does not require activation through the attachment of an NDP moiety. GenK was identified to methylate a nonactivated sp3 carbon in the late-stage modification of gentamicin biosynthesis.60 The iron-sulfur cluster facilitates the cleavage of SAM, generating a gentamicin substrate radical that subsequently accepts a methyl group from methylcobalamin (Scheme 5). Similar to the C4-MT GenD1, GenK exhibits substrate flexibility and accepts multiple sugar substrates. Furthermore, these methylation events can occur in combination with the N-MT GenN in an interchangeable sequence, forming a flexible and interconnected methylation network (Scheme 5).534. Biosynthetic network of keto-deoxy sugars in canonical C-methylation
Although the substrates of SN2-mediated C-methylation of keto-deoxy sugars are structurally different, they share a common backbone: a six-membered sugar ring with a keto group in the C3 or C4 position linked to either a TDP or GDP tail. The biosynthesis of these sugar moieties begins with the predominant precursor, D-glucose. Once transported as glucose 6-phosphate, it is converted into glucose 1-phosphate via hexose phosphate mutase. It is then transformed in a two-step process into TDP-4-keto-6-deoxy-D-glucose, which is a key intermediate in the biosynthesis of all methylatable TDP sugars (1 → 4; Scheme 7). Similarly, GDP-activated sugars originate from fructose 6-phosphate (8), which is obtained by phosphoglucose isomerisation of glucose 6-phosphate. Phosphomannose isomerase and hexose phosphate synthase then convert fructose 6-phosphate into mannose 1-phosphate (10). Subsequently, the GDP tail is tailored into the sugar core via the corresponding nucleotidyltransferases, followed by the assistance of 4,6-dehydrogenase, generating GDP-4-keto-6-deoxy-D-mannose (12), ready for C3 and C5 methylation. Structural variations among TDP-linked substrates depend primarily on the extent of deoxygenation. TDP-4-keto-6-deoxy-D-glucose is first modified by sugar 2,3-dehydrogenase, followed by either a sugar 3-aminotransferase (e.g., TcaB8,44 EvaB,38 DvaB13,61) or a sugar 3-hydroxytransferase (e.g., TylC1,35 EryBII,38 AviT20). These resulting intermediates then serve as substrates for MTs such as TcaB9, EvaC, AviG1, TylC3, EryBIII, and MtmC, each of which has distinct preferences for the axial or equatorial hydroxyl/amino group at C3. This variability contributes to the stereochemical diversity of the final products. TDP-4-keto pentose (22) represents a rare substrate for methylation, generated by oxidative decarboxylation at C6, and has only been reported in the biosynthesis of maduropeptin. Additionally, 3-epimerization or 3,5-epimerization of keto-deoxy sugars is also essential for the natural assembly of monosaccharide biosynthetic pathways. Such transformations may occur either before or after dehydroxylation, as exemplified by the actions of NovW,56 SibJ,9 and TylK.62 In the biosynthesis of chromomycin A3, the epimerization is proposed to take place even after C-methylation by CmmF, as suggested by Nuria Menéndez et al.635. Preferences of sugar C-MTs: a bioinformatics perspective
Phylogenetic analysis provides valuable insights into the methylation preferences of the classical sugar C-MTs based on existing knowledge. The evolutionary trajectory of sugar C-MTs is primarily determined by the stereochemistry of the resulting methylated products. In particular, TylC3-like enzymes diverge into a distinct phylogenetic branch from the other C3-, C4-, and C5-MTs, possibly due to the axial-to-equatorial shift of the C3 hydroxyl group in the methylation process, which might be the most obvious discrepancy in the course of evolution (Fig. 2). The apparent phylogenetic distribution of TylC3 and SibM in separate branches indicates that the two-step modification of the TDP sugar units by methylation and epimerization may have different priorities in the biosynthesis of tylosin and sibiromycin, as shown in Scheme 2C. MdpA4 remains independent of other C3-MTs, especially ChryCMT, suggesting that the absence of C6 influences the anchoring of substrates in the active site during evolution. C4-MT CyaA3 forms a distinct clade, separate from the major group of C3- and C5-MTs, suggesting that it may employ a novel catalytic mechanism for C4-methylation. Notably, the rest of the canonical sugar C3-, C4-, and C5-MTs do not exhibit strict sequence-based boundaries in their substrate preferences, underscoring the uniformity and versatility of substituted keto-deoxy sugars, such as KijD1, TcaB9, and EvdM8, which appear to have co-evolved, suggesting a shared evolutionary trajectory. | ||
| Fig. 2 Phylogenetic analysis of C-MTs in the biosynthesis of sugar moieties. Yellow: clade of C3-MTs; green: C4-MTs; and blue: C5-MTs. The evolutionary history was inferred using the maximum likelihood method and the Whelan and Goldman + freq. model.64 The tree with the highest log likelihood (−20133.94) is shown. The initial tree(s) for the heuristic search were obtained automatically by applying the neighbor-join and BioNJ algorithms to a matrix of pairwise distances estimated using the JTT model, and then selecting the topology with a superior log likelihood value. A discrete gamma distribution was used to model the differences in evolutionary rate between the sites [5 categories (+G, parameter = 1.4597)]. The rate variation model allowed for some sites to be evolutionarily invariable ([+I], 2.89% sites). The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. This analysis involved 40 amino acid sequences. The final dataset contained a total of 462 positions. The evolutionary analyses were conducted in MEGA11.65 | ||
As there is a lack of detailed biosynthetic studies on these C3-MTs, their catalytic mechanism and stereochemistry remain unclear. The 3,5-epimerization of the substrate could be an important factor in the evolution of C3-MTs, as evidenced by the clustering of SnogG2 and SibM with C5-MTs such as NovU, CloU, and SgcA3. In addition, the distinction between GDP- and TDP-specific C3-MTs is not clearly defined, despite the difference in the stereochemistry at the C2 position. Meanwhile, TiaS2 diverges from other C5-MTs, further enhancing the complexity of the evolutionary landscape. The phylogenetic clustering of sugar C-MTs allows a tentative assignment of methylation substrates by comparison with characterized systems. For instance, KedS4 clusters with TylC3 and may act in an analogous manner on TDP-4-keto-2,6-dideoxy-D-allose. In this scenario, C-methylation would be followed by 3,5-epimerization and subsequent tailoring reactions to obtain TDP-L-mycarose as the sugar unit for incorporation. A similar proposal has been made for NocS3,66 which suggests a comparable methylation mechanism within the biosynthesis of apoptolidin and chromomycin, due to its phylogenetic proximity to ApoM5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 67 and CmmC.33
67 and CmmC.33
Certain exceptions could be found in the phylogenetic tree, such as the comparison of MtmC and MmyC with CmmC, which belong to the same aureolic acid family.68 One possible explanation is that MtmC functions as a bifunctional enzyme capable of accommodating both SAM and NADPH, which may have driven its independent evolutionary path.
6. Parallels and differences between sugar C-MTs within tailoring enzyme networks
Modification of sugar units is part of an intricate universe that encompasses the biosynthesis of glycosylated natural products. A central feature of this process is C–C bond formation, which drives the expansion of carbon skeletons. Chain elongation and branching can be achieved through various enzymatic strategies, including aldolases,69,70 transketolases71 and other ThDP-dependent enzymes,72 and both canonical and radical SAM-dependent MTs. Among these, C-methylation of NDP-activated keto-deoxy sugars by sugar MTs plays a pivotal role in C1 building-block integration. It often occurs in tandem with oxidation, decarboxylation, or reduction and contributes to the construction of structurally diverse sugar units.It is worthy of note that ThDP-dependent enzymes are capable of modifying analogous nucleoside substrates, albeit via wholly disparate mechanisms in sugar-containing natural product biosynthesis, thus providing divergent pathways for sugar skeleton modification.20,73,74 The ability of two distinct enzyme classes to operate on a common substrate presents intriguing opportunities, particularly in the design of artificial sugar analogs through the strategic combination of these enzymes for late-stage sugar modification. It also emphasizes the high reactivity of NDP-linked deoxy keto sugars, as shown in the bifunctional C-MT MtmC. Efficient methylation, therefore, likely requires strong pathway commitment of sugar C-MTs within the cellular environment. In contrast to C-MTs that modify α-keto acids75,76 or those associated with polyketide synthase (PKS) pathways,77,78 which need metal ions to facilitate methyl transfer, methylation with sugar C-MTs function without metal cofactors. This observation suggests that the active site in sugar MTs stabilizes the enolate intermediate primarily through electrostatic interactions.
In addition to the biosynthetic pathway in which natural TDP- or GDP-linked keto-deoxy sugars are derived from glucose 6-phosphate/fructose 6-phosphate, an alternative route has also been reported. In this case, the potential methylation precursor TDP-3-keto-6-deoxy-D-glucose is generated from TDP-D-quinovose through a radical-initiated deamination reaction,79,80 highlighting the remarkable diversity of sugar biosynthetic strategies in nature. This is just one of many examples mentioned in this review that are reminiscent of the screening hypothesis described by Firn and Jones.81
7. Conclusion
In this review, we survey sugar C-MTs involved in early- and late-stage biosynthetic modifications, including enzymes acting via SN2-type and radical SAM-associated mechanisms. Methylation of deoxy keto sugar exhibits remarkable diversity across biosynthetic pathways and enables highly stereoselective methylation at C3, C4, C5, or C6 positions, ultimately yielding at least ten distinct sugar units. Among these enzymes, C3-MTs are most frequently identified (based on bioinformatic analysis or experimental studies). Here, we not only emphasize the methylation itself but also highlight that it is typically integrated with additional tailoring reactions, such as epimerization, dehydroxylation, and transamination. These enzymes work together to expand the accessible chemical space of sugar cores, and the process accounts for much of the structural variation observed in glycosylated natural products or polysaccharides.Limited phylogenetic divergence among sugar C-MTs, coupled with the high conservation of active-site residues observed in structural studies, complicates predictions of substrate preference and stereochemical outcome from sequence information alone. For most sugar C-MTs, we argue that what is really constrained by evolution is not the “C3, C4, or C5” position for methylation, but maybe rather the relative orientation of the substrate and SAM within the active site.82,83 Elucidation of sugar C-MTs mechanisms from both structural and chemical perspectives can shed light on how regio- and stereo-chemical behaviors are controlled during monosaccharide biosynthesis. Reported examples show that methylated sugar units display marked structural variation, consistent with precise control over methyl group installation by sugar C-MTs.
As exemplified by the so-called “magic methyl effect” in drug discovery,84 methyl group installation has the potential to exert a disproportionate influence on molecular bioactivity and physicochemical properties. Within this context, knowledge of the NDP-activated keto-deoxy sugar methylation network can provide valuable insights into glycosylated natural product biosynthesis and is expected to lead to significant progress in the study and discovery of previously unknown glycosylated natural products and specialized metabolites, such as lipopolysaccharides. In addition, the methylation network delineates the substrate tolerance of sugar C-MTs toward non-native substrates and provides a basis for their application in artificial or new-to-nature modification of sugars. Future studies on sugar C-methylation, particularly within pathway-committed biosynthetic contexts, have the potential to clarify how targeted methylation of sugar units contributes to the functional diversification of natural products.
8. Conflicts of interest
There are no conflicts to declare.9. Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: Table S1. See DOI: https://doi.org/10.1039/d5np00071h.10. Acknowledgements
We thank Juliane Breiltgens for fruitful discussions. The Deutsche Forschungsgemeinschaft (DFG project 510974120) is acknowledged for funding. Z. Z. thanks the Chinese Scholarship Council (CSC) for a PhD fellowship.11. References
- D. K. Liscombe, G. V. Louie and J. P. Noel, Nat. Prod. Rep., 2012, 29, 1238–1250 RSC.
- J. C. Wu and D. V. Santi, J. Biol. Chem., 1987, 262, 4778–4786 CrossRef CAS PubMed.
- H. Ogawa, T. Gomi, F. Takusagawa and M. Fujioka, Int. J. Biochem. Cell Biol., 1998, 30, 13–26 CrossRef CAS PubMed.
- M. Jansson, S. T. Durant, E.-C. Cho, S. Sheahan, M. Edelmann, B. Kessler and N. B. La Thangue, Nat. Cell Biol., 2008, 10, 1431–1439 CrossRef CAS PubMed.
- J. Breiltgens and M. Müller, Nat. Prod. Rep., 2026 10.1039/d5np00070j.
- J. Liu, T. Ng, Z. Rui, O. Ad and W. Zhang, Angew. Chem., Int. Ed., 2014, 53, 136–139 CrossRef CAS PubMed.
- M. Komatsu, M. Tsuda, S. Ōmura, H. Oikawa and H. Ikeda, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 7422–7427 CrossRef CAS.
- K. Ishida, K. Fritzsche and C. Hertweck, J. Am. Chem. Soc., 2007, 129, 12648–12649 CrossRef CAS PubMed.
- W. Li, A. Khullar, S. Chou, A. Sacramo and B. Gerratana, Appl. Environ. Microbiol., 2009, 75, 2869–2878 CrossRef CAS PubMed.
- A. Freitag, S.-M. Li and L. Heide, Microbiology, 2006, 152, 2433–2442 CrossRef CAS PubMed.
- N. A. Bruender and H. M. Holden, Protein Sci., 2012, 21, 876–886 CrossRef CAS PubMed.
- P. Wang, W. Kim, L. B. Pickens, X. Gao and Y. Tang, Angew. Chem., Int. Ed., 2012, 51, 11136–11140 CrossRef CAS PubMed.
- S. Pelzer, R. Süßmuth, D. Heckmann, J. Recktenwald, P. Huber, G. Jung and W. Wohlleben, Antimicrob. Agents Chemother., 1999, 43, 1565–1573 CrossRef CAS PubMed.
- G. Flores-Ramirez, S. Janecek, J. A. Miernyk and L. Skultety, Proteome Sci., 2012, 10, 67 CrossRef CAS.
- S. Gaisser, A. Trefzer, S. Stockert, A. Kirschning and A. Bechthold, J. Bacteriol., 1997, 179, 6271–6278 CrossRef CAS PubMed.
- S. Mizrahy and D. Peer, Chem. Soc. Rev., 2012, 41, 2623–2640 RSC.
- J. Muthukumar, R. Chidambaram and S. Sukumaran, J. Food Sci. Technol., 2021, 58, 2453–2466 CrossRef CAS PubMed.
- B. La Ferla, C. Airoldi, C. Zona, A. Orsato, F. Cardona, S. Merlo, E. Sironi, G. D'Orazio and F. Nicotra, Nat. Prod. Rep., 2011, 28, 630–648 RSC.
- C. J. Thibodeaux, C. E. Melançon and H.-w. Liu, Angew. Chem., Int. Ed., 2008, 47, 9814–9859 CrossRef CAS PubMed.
- G. Weitnauer, A. Mühlenweg, A. Trefzer, D. Hoffmeister, R. D. Süßmuth, G. Jung, K. Welzel, A. Vente, U. Girreser and A. Bechthold, Chem. Biol., 2001, 8, 569–581 CrossRef CAS PubMed.
- C. Hofmann, R. Boll, B. Heitmann, G. Hauser, C. Dürr, A. Frerich, G. Weitnauer, S. J. Glaser and A. Bechthold, Chem. Biol., 2005, 12, 1137–1143 CrossRef CAS PubMed.
- G. Weitnauer, S. Gaisser, L. Kellenberger, P. F. Leadlay and A. Bechthold, Microbiology, 2002, 148, 373–379 CrossRef CAS PubMed.
- S. Torkkell, T. Kunnari, K. Palmu, P. Mäntsälä, J. Hakala and K. Ylihonko, Mol. Genet. Genomics, 2001, 266, 276–288 CrossRef CAS PubMed.
- N. Zein, W. Solomon, K. L. Colson and D. R. Schroeder, Biochemistry, 1995, 34, 11591–11597 CrossRef CAS PubMed.
- D. R. Schroeder, K. L. Colson, S. E. Klohr, N. Zein, D. R. Langley, M. S. Lee, J. A. Matson and T. W. Doyle, J. Am. Chem. Soc., 1994, 116, 9351–9352 CrossRef CAS.
- R. E. Campbell, S. C. Mosimann, I. Van De Rijn, M. E. Tanner and N. C. J. Strynadka, Biochemistry, 2000, 39, 7012–7023 CrossRef CAS PubMed.
- M. Bar-Peled, C. L. Griffith and T. L. Doering, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 12003–12008 CrossRef CAS PubMed.
- S. D. Breazeale, A. A. Ribeiro and C. R. H. Raetz, J. Biol. Chem., 2002, 277, 2886–2896 CrossRef CAS.
- S. G. Van Lanen, T. Oh, W. Liu, E. Wendt-Pienkowski and B. Shen, J. Am. Chem. Soc., 2007, 129, 13082–13094 CrossRef CAS PubMed.
- M. K. Kharel, S. E. Nybo, M. D. Shepherd and J. Rohr, ChemBioChem, 2010, 11, 523–532 CrossRef CAS PubMed.
- G. Wang, P. Pahari, M. K. Kharel, J. Chen, H. Zhu, S. G. Van Lanen and J. Rohr, Angew. Chem., Int. Ed., 2012, 51, 10638–10642 CrossRef CAS.
- L. L. Remsing, J. Garcia-Bernardo, A. Gonzalez, E. Künzel, U. Rix, A. F. Braña, D. W. Bearden, C. Méndez, J. A. Salas and J. Rohr, J. Am. Chem. Soc., 2002, 124, 1606–1614 CrossRef CAS PubMed.
- N. Menendez, M. Nurealam, A. Brana, J. Rohr, J. Salas and C. Mendez, Chem. Biol., 2004, 11, 21–32 CAS.
- J. M. Chen, C. Hou, G. Wang, O. V. Tsodikov and J. Rohr, Biochemistry, 2015, 54, 2481–2489 CrossRef CAS PubMed.
- H. Chen, G. Agnihotri, Z. Guo, N. L. S. Que, X. H. Chen and H.-w. Liu, J. Am. Chem. Soc., 1999, 121, 8124–8125 CrossRef CAS.
- H. Chen, Z. Zhao, T. M. Hallis, Z. Guo and H.-w. Liu, Angew. Chem., Int. Ed., 2001, 40, 607–610 CrossRef CAS.
- M. Zhang, B. Shuang and K. Arakawa, Bioorg. Med. Chem. Lett., 2023, 80, 129125 CrossRef CAS PubMed.
- V. A. Mironov, O. V. Sergienko, I. N. Nastasyak and V. N. Danilenko, Appl. Biochem. Microbiol., 2004, 40, 531–541 CrossRef CAS.
- H. Zhang, Y. Wang, J. Wu, K. Skalina and B. A. Pfeifer, Chem. Biol., 2010, 17, 1232–1240 CrossRef CAS PubMed.
- T. J. Hosted, T. X. Wang, D. C. Alexander and A. C. Horan, J. Ind. Microbiol. Biotechnol., 2001, 27, 386–392 CrossRef CAS PubMed.
- M. Zhu, L. Wang, H. Zhang, L. Zhang, B. Tan, Q. Huang, Y. Zhu and C. Zhang, ACS Synth. Biol., 2023, 12, 1520–1532 CrossRef CAS PubMed.
- L. Ninet, F. Benazet, Y. Charpentie, M. Dubost, J. Florent, J. Lunel, D. Mancy and J. Preud'Homme, Experientia, 1974, 30, 1270–1272 CrossRef CAS PubMed.
- E. M. Limbrick, A. E. Yñigez-Gutierrez, C. C. Dulin, D. K. Derewacz, J. M. Spraggins, K. M. McCulloch, T. M. Iverson and B. O. Bachmann, ChemBioChem, 2020, 21, 3349–3358 CrossRef CAS PubMed.
- J. Fang, Y. Zhang, L. Huang, X. Jia, Q. Zhang, X. Zhang, G. Tang and W. Liu, J. Bacteriol., 2008, 190, 6014–6025 CrossRef CAS PubMed.
- C. Yue, J. Niu, N. Liu, Y. Lü, M. Liu, Y. Li, M. Wang, M. Shao, S. Qian and Y. Huang, Pak. J. Pharm. Sci., 2016, 29, 287–293 CAS.
- H. Komaki, N. Ichikawa, A. Hosoyama, M. Hamada and Y. Igarashi, Antibiotics, 2021, 10, 1447 CrossRef CAS PubMed.
- C.-G. Kim, J. Lamichhane, K.-I. Song, V. D. Nguyen, D.-H. Kim, T.-S. Jeong, S.-H. Kang, K.-W. Kim, J. Maharjan, Y.-S. Hong, J. S. Kang, J.-C. Yoo, J.-J. Lee, T.-J. Oh, K. Liou and J. K. Sohng, Arch. Microbiol., 2008, 189, 463–473 CrossRef CAS PubMed.
- N. A. Bruender, J. B. Thoden, M. Kaur, M. K. Avey and H. M. Holden, Biochemistry, 2010, 49, 5891–5898 CrossRef CAS PubMed.
- H. Zhang, J. A. White-Phillip, C. E. Melançon, H. Kwon, W. Yu and H.-w. Liu, J. Am. Chem. Soc., 2007, 129, 14670–14683 CrossRef CAS PubMed.
- G. T. Dow, J. B. Thoden and H. M. Holden, Protein Sci., 2016, 25, 2282–2289 CrossRef CAS PubMed.
- H. Chen, M. G. Thomas, B. K. Hubbard, H. C. Losey, C. T. Walsh and M. D. Burkart, Proc. Natl. Acad. Sci. U. S. A., 2000, 97, 11942–11947 CrossRef CAS PubMed.
- A. L. Lane, S.-J. Nam, T. Fukuda, K. Yamanaka, C. A. Kau, P. R. Jensen, W. Fenical and B. S. Moore, J. Am. Chem. Soc., 2013, 135, 4171–4174 CrossRef CAS PubMed.
- S. Li, J. Guo, A. Reva, F. Huang, B. Xiong, Y. Liu, Z. Deng, P. F. Leadlay and Y. Sun, Proc. Natl. Acad. Sci. U. S. A., 2018, 115, 1340–1345 CrossRef CAS PubMed.
- J. Guo, F. Huang, C. Huang, X. Duan, X. Jian, F. Leeper, Z. Deng, P. F. Leadlay and Y. Sun, Chem. Biol., 2014, 21, 608–618 CrossRef CAS PubMed.
- L. Heide, Nat. Prod. Rep., 2009, 26, 1241 RSC.
- T. T. Thu Thuy, H. C. Lee, C.-G. Kim, L. Heide and J. K. Sohng, Arch. Biochem. Biophys., 2005, 436, 161–167 CrossRef PubMed.
- W. Liu, S. D. Christenson, S. Standage and B. Shen, Science, 2002, 297, 1170–1173 CrossRef CAS PubMed.
- S. Van Lanen and B. Shen, Curr. Top. Med. Chem., 2008, 8, 448–459 CrossRef CAS PubMed.
- Y. Xiao, S. Li, S. Niu, L. Ma, G. Zhang, H. Zhang, G. Zhang, J. Ju and C. Zhang, J. Am. Chem. Soc., 2011, 133, 1092–1105 CrossRef CAS PubMed.
- H. J. Kim, R. M. McCarty, Y. Ogasawara, Y. Liu, S. O. Mansoorabadi, J. LeVieux and H.-w. Liu, J. Am. Chem. Soc., 2013, 135, 8093–8096 CrossRef CAS PubMed.
- S. Donadio, M. Sosio, E. Stegmann, T. Weber and W. Wohlleben, Mol. Genet. Genomics, 2005, 274, 40–50 CrossRef CAS PubMed.
- H. Takahashi, Y. Liu and H.-w. Liu, J. Am. Chem. Soc., 2006, 128, 1432–1433 CrossRef CAS PubMed.
- N. Menéndez, M. Nurealam, A. Brana, J. Rohr, J. Salas and C. Méndez, Chem. Biol., 2004, 11, 21–32 Search PubMed.
- S. Whelan and N. Goldman, Mol. Biol. Evol., 2001, 18, 691–699 CrossRef CAS PubMed.
- K. Tamura, G. Stecher and S. Kumar, Mol. Biol. Evol., 2021, 38, 3022–3027 CrossRef CAS PubMed.
- Y. Ding, Y. Yu, H. Pan, H. Guo, Y. Li and W. Liu, Mol. BioSyst., 2010, 6, 1180–1185 CrossRef CAS PubMed.
- Y. Du, D. K. Derewacz, S. M. Deguire, J. Teske, J. Ravel, G. A. Sulikowski and B. O. Bachmann, Tetrahedron, 2011, 67, 6568–6575 CrossRef CAS PubMed.
- L. J. Stevenson, J. Bracegirdle, L. Liu, A. V. Sharrock, D. F. Ackerley, R. A. Keyzers and J. G. Owen, RSC Chem. Biol., 2021, 2, 556–567 RSC.
- J. Wagner, R. A. Lerner and C. F. Barbas, Science, 1995, 270, 1797–1800 CrossRef CAS PubMed.
- X. Garrabou, J. A. Castillo, C. Guérard-Hélaine, T. Parella, J. Joglar, M. Lemaire and P. Clapés, Angew. Chem., Int. Ed., 2009, 48, 5521–5525 CrossRef CAS PubMed.
- N. J. Turner, Curr. Opin. Biotechnol., 2000, 11, 527–531 CrossRef CAS PubMed.
- C. Dresen, M. Richter, M. Pohl, S. Lüdeke and M. Müller, Angew. Chem., Int. Ed., 2010, 49, 6600–6603 CrossRef CAS PubMed.
- J. Steitz, L. Krug, L. Walter, K. Hernández, C. Röhr, P. Clapés and M. Müller, Angew. Chem., Int. Ed., 2022, 61, e202113405 CrossRef CAS PubMed.
- L. Krug, D. Bjarnesen, L. Lanza, L. Lindemann, N. D. Fessner and M. Müller, Angew. Chem., Int. Ed., 2024, 63, e202403535 CAS.
- C. Sommer-Kamann, A. Fries, S. Mordhorst, J. N. Andexer and M. Müller, Angew. Chem., Int. Ed., 2017, 56, 4033–4036 CrossRef CAS PubMed.
- C. Sommer-Kamann, J. Breiltgens, Z. Zou, S. Gerhardt, R. Saleem-Batcha, F. Kemper, O. Einsle, J. N. Andexer and M. Müller, ChemBioChem, 2024, 25, e202400258 CrossRef CAS PubMed.
- M. A. Skiba, A. P. Sikkema, N. A. Moss, C. L. Tran, R. M. Sturgis, L. Gerwick, W. H. Gerwick, D. H. Sherman and J. L. Smith, ACS Chem. Biol., 2017, 12, 3039–3048 CrossRef CAS PubMed.
- Y. Lao, M. A. Skiba, S. W. Chun, A. R. H. Narayan and J. L. Smith, ACS Chem. Biol., 2022, 17, 2088–2098 CrossRef CAS PubMed.
- Y. Xue, L. Zhao, H.-w. Liu and D. H. Sherman, Proc. Natl. Acad. Sci. U. S. A., 1998, 95, 12111–12116 CrossRef CAS PubMed.
- G. Lin, S.-H. Choi, M. W. Ruszczycky and H.-w. Liu, J. Am. Chem. Soc., 2015, 137, 4964–4967 CrossRef CAS PubMed.
- R. D. Firn and C. G. Jones, Nat. Prod. Rep., 2003, 20, 382 RSC.
- J. Breiltgens, A. Paul, Z. Zou, J. N. Andexer and M. Müller, J. Nat. Prod., 2025, 88, 2750–2756 CrossRef CAS PubMed.
- J. Breiltgens, Z. Zou, J. N. Andexer and M. Müller, ChemBioChem, 2026 DOI:10.1002/cbic.70294.
- H. Schönherr and T. Cernak, Angew. Chem., Int. Ed., 2013, 52, 12256–12267 CrossRef PubMed.
Footnote |
| † Dedicated to Roman Kaiser on the occasion of his 80th birthday. |
| This journal is © The Royal Society of Chemistry 2026 |




