Progress, challenges, and opportunities in the field of biosynthetic reactions involving ambimodal transition states
Hidayat
Hussain
 *a,
K. N.
Houk
*a,
K. N.
Houk
 b,
Ching Ching
Lam
b,
Ching Ching
Lam
 b,
Satyajit D.
Sarker
b,
Satyajit D.
Sarker
 c and
Lutfun
Nahar
c and
Lutfun
Nahar
 d
d
aInternational Joint Laboratory of Medicinal Food Development and Health Products Creation, Biological Engineering Technology Innovation Center of Shandong Province, Heze Branch of Qilu University of Technology (Shandong Academy of Sciences), Heze, China. E-mail: hussainchem3@gmail.com
bDepartment of Chemistry and Biochemistry, University of California Los Angeles, Los Angeles, CA 90095-1569, USA
cCentre for Natural Products Discovery, School of Pharmacy and Biomolecular Sciences, Liverpool John Moores University, James Parsons Building, Byrom Street, Liverpool L3 3AF, UK
dLaboratory of Growth Regulators, Palacký University and Institute of Experimental Botany, The Czech Academy of Sciences, Šlechtitelů 27, Olomouc, Czech Republic
First published on 18th November 2025
Abstract
Covering: up to June 2025
High selectivity is generally observed in the biosynthesis of complex natural molecules. Evolution usually leads to enzymes that favor the formation of a particular isomer rather than one of the many other potential molecules. Recent discoveries of enzymes with multiple sequential post-transition state bifurcations (PTSB) after ambimodal transition states demonstrate the impact of dynamics on selectivity. PTSB cause a single ambimodal transition state (TS) to form multiple products. This is different from conventional energetically-controlled mechanisms, where two discrete transition states have different energy barriers. Selectivity arising from ambimodal TSs cannot be fully explained by transition state theory. The presence of PTSB on enzyme catalyzed reaction surfaces has been discovered recently at a significantly higher rate. For both uncatalyzed and catalyzed reactions, computational chemists are devising techniques to comprehend which elements of molecular structure and vibrations govern the product selectivity in systems that contain bifurcations. This review describes enzyme-catalyzed reactions involving ambimodal transition states, and recent advances in understanding how enzymes control selectivity in such reactions.
1 Introduction
Enzymes are highly efficient and specific catalysts that play a crucial role in facilitating chemical reactions essential for metabolic and regulatory processes in living organisms.1–3 A deeper understanding of how enzymes control selectivity and accelerate reactions is important for advancing biochemical and bioengineering research aimed at designing and improving enzyme functionality. A key area of research is how enzyme conformational changes, ranging from nanoseconds to sub-picoseconds, influence the reactivity and selectivity of enzymes. Although nanosecond-scale motions are well-recorded in numerous enzymes, the function of faster, sub-picosecond movements in regulating enzyme activity is not yet fully comprehended. The outcome of an enzymatic reaction is linked to the transition state and its stabilization, which is affected by the dynamics of the enzyme active site residues.1–3Enzymologists use a variety of structural and kinetic data to differentiate between mechanistic possibilities. It has been suggested that a number of ‘textbook’ mechanisms are inaccurate in significant respects. For example, quantum mechanics/molecular mechanics (QM/MM) molecular dynamics simulations suggest that the reaction of native hen egg white lysozyme (HEWL) with its natural substrate forms a covalent intermediate. This is different from the ‘textbook’ mechanism involving an oxocarbenium ion intermediate for this important enzyme.4,5 There have been many long and heated debates about reaction mechanisms and catalytic principles within the field of enzymology, but these are difficult to resolve through experimentation alone. Scientists have also questioned the applicability of transition state theory to enzyme reactions, and the role of dynamics.6 The field of computational enzymology is undergoing rapid development, and computational modelling is playing a key role in aiding experimental studies of enzyme-catalysed reactions.
No detailed or comprehensive review or highlights have been published on biosynthetic reactions with post-transition state bifurcations on the potential energy surface. A very short ‘News & Views’ article was published by Hornsby and Paton in 2014,7 in which only one example of a bifurcating biosynthetic reaction was discussed. Moreover, Hare and Tantillo8 published a review on the dynamic behaviour of carbocations and its impact on terpene biosynthesis in 2016. Nonetheless, the scope of their coverage is limited to just two primary illustrations of bifurcation reactions. Additionally, in 2017, Hare and Tantillo9 published a review of post-transition state bifurcations in organic reactions, focusing primarily on synthetic reactions. However, they only examined three enzymatic reactions. In 2019, Tantillo10 published a review on the use of quantum chemical calculations to interrogate mechanisms of natural product biosynthesis. The review covered only one example of a biosynthetic bifurcation reaction. Here, we review examples of enzymatic reactions with ambimodal transition states and PTSBs and highlight how both experimental and computational approaches can aid in identifying and elucidating the underlying reaction mechanisms.
2 Bifurcating reactions
A transition state (TS) is the point of highest energy along the minimum-energy route connecting reactants and products. Typically, a TS connects one set of reactants to one set of products. When analyzing reaction selectivity, most chemists will instinctively consider kinetic effects or transition state theory (TST). In accordance with the findings of TST, the reaction pathway that exhibits the lowest activation barrier is the most favourable, and consequently leads to the major product (Fig. 1A).11,12 In TST, the product ratio from competing pathways is affected by the difference in Gibbs free energy of their respective transition states.13,14 However, the number of reactions exhibiting non-statistical (non-TST) behaviour reported is steadily growing, where the traditional TST breaks down.14 Many of these examples are organic reactions, in which reaction dynamic effects can also play a pivotal role. Such reactions characteristically exhibit a bifurcating potential energy surface (PES) with a valley-ridge inflection (VRI) point (Fig. 1B) or a shallow entropic intermediate (Fig. 1C) consequent to an ambimodal TS that can lead to multiple product pathways.11,12,15,16 This process is known as post-transition state bifurcation (PTSB). A single TS gives rise to multiple products via a VRI point15,16 (Fig. 1B and C). | ||
| Fig. 1 Typical energy profiles in chemical and biosynthetic reactions.11,12 (A) the reaction pathway that exhibits the lowest activation barrier is the most favourable, and consequently leads to the major product (P1); (B) a reaction with an ambimodal TS (TS1) and a valley-ridge inflection point (VRI) on the potential energy surface;11,12 (C) a reaction with a shallow intermediate (INT) where the sequential TSs (TS2A and TS2A) are low in energy compared to the first TS.11,12 In this case, selectivity may also be influenced by reaction dynamic effects. Reproduced from ref. 11 with permission from the IUPAC. | ||
This type of PES is associated with a reaction mechanism that deviates from the usual stepwise or concerted designations. This type has been classified as a two-step, no-intermediate mechanism.13 The existence of a PTSB makes it more difficult to predict product selectivity because a single TS structure enables direct access to two or more products.17,18 Such reactions may have multiple transition states after the rate determining ambimodal TS. In addition to the established thermodynamic, kinetic and tunnelling factors that influence reaction selectivity, the PTSB introduces a new mechanistic factor controlling selectivity.19
3 Isoprenoids
3.1 Bifurcation in monoterpenes
Major et al. and Tantillo et al., respectively, investigated the detailed mechanistic pathway for the biosynthesis of bornyl diphosphate (BPP) from geranyl diphosphate through computations.20–22 Based on gas phase calculations, they predicted that PTSB might be involved in the biosynthesis of BPP, catalysed by bornyl diphosphate synthase (BPPS) (Scheme 1). From the bornyl cation, the reaction pathways bifurcate and lead to bornyl diphosphate (2) and camphene (4), which is a byproduct produced via a camphyl cation (3).21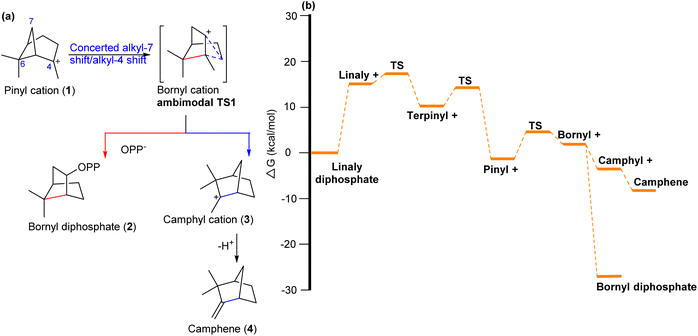 | ||
| Scheme 1 (a) Carbocation rearrangement reaction involving PTSB in the biosynthesis of bornyl diphosphate. (b) The X-ray crystallography structure of dimeric (+)-bornyl diphosphate synthase from Salvia officinalis, reported by Whittington et al. (PDB ID: 1N23),23 was used as a starting point for the simulations. Based on the free energy profile of carbocations in Salvia officinalis (+)-bornyl diphosphate synthase (So-BPPS), the bornyl cation was identified as an enzyme-induced bifurcation point.21 Reproduced from ref. 21 with permission from the Royal Society of Chemistry. | ||
Major and Weitman21 predicted from computational studies that achieving high reaction specificity is difficult due to enzyme dynamics. The electrostatic interactions between the enzyme and the bornyl cation are also found to be insufficiently strong, leading to the formation of by-products. Experimental evidence supporting this conclusion comes in the form of the crystal structure of So-BPPS with (1R,4S)-2-azabornane, which is an analogue of the bornyl cation.23 The results showed that product ratios in systems with PTSBs can be significantly influenced by non-covalent and electrostatic interactions.
3.2 Bifurcation in sesquiterpene biosynthesis
A possible biosynthetic route to neopupukeanane (8), pupukeanane (9), and abeopupukeanane (10), molecules under the class of sesquiterpene, was proposed by various research groups respectively (Scheme 2a).24,25 The cation 7 formed from the cyclization of cation 5 was proposed to undergo a 1,2-alkyl shift to form either a neopupukeanane cation 8 or a pupukeanane cation 9. The latter cation (9) would be a direct precursor of pupukeane sesquiterpenes and an indirect precursor of abeopupukeane sesquiterpenes (via cation 10).24 Tantillo and colleagues investigated the mechanism of the biosynthesis using DFT calculations.24 Two sequential 1,2-alkyl shifts from cation 7 lead to the tertiary carbene 10 or 11 (Scheme 2a). On the reaction pathway from cationic 7 to cation 10 or cation 11, structures similar to secondary carbocation 9 persisted temporarily. Carbocation 9, which was thought to be formed by the migration of the C3–C4 bond in cation 7, is not a minimum at all levels of theory tested. Tantillo and colleagues26 further demonstrated that the part of the PES near the cation 9 was somewhat flat and had at least three exit pathways, towards the cations 7, 10, and 11. Garson and colleagues27 also show that the cation 9 is non-classical and occupies a comparatively flat region on the potential energy surface, while being higher in energy than cation 7. Earlier work on similar but simpler organic systems also highlights features of PTSBs on the PES, with this supported by trajectories from quasiclassical dynamics simulations.28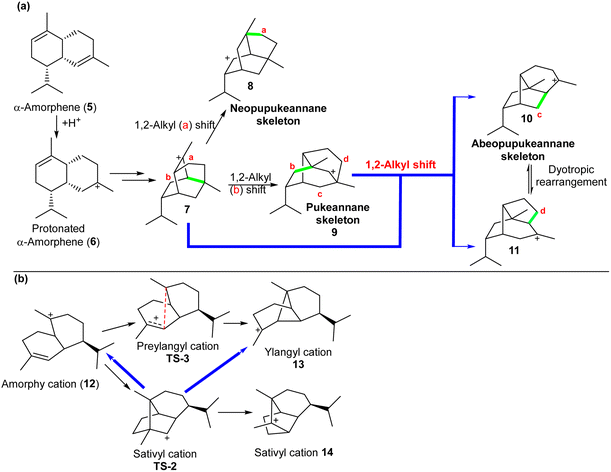 | ||
| Scheme 2 (a) Carbocations involved in proposed mechanisms for the formation of pupukeanane and related sesquiterpene natural products. Newly formed σ-bonds in structures are coloured in green. (b) Biosynthetic carbocation rearrangements leading to ylangene and sativene. The paths in blue are “dynamically-controlled”. Reproduced from ref. 26 and 29 with permission from the American Chemical Society and Royal Society of Chemistry respectively. | ||
Ylangene and sativene also belong to the class of sesquiterpene compounds. The lifetimes of classical secondary carbocations that lead to ylangene and sativene were estimated and compared by Pemberton and Tantillo29 using direct dynamics calculations. It was found that these carbocations exist near transition state structures (TSS). TS-2 is a TS that can lead to either the terpinyl-like amorphyl cation 12 or the camphyl-like sativyl cation 14 (Scheme 2b). The cation 14 was observed to be present in all 110 trajectories that were initiated from TS-2 in the product direction. In the reactant direction, the formation of 12 occurred in 42 of the trajectories, but the formation of pinyl-like ylangyl cation 13 occurred in 68 of the trajectories. This indicates that a bifurcation may exist along the pathway from TS-2 back towards 12, highlighting another case where dynamic effects need to be considered. It should be noted that structures resembling 12 are not generally involved in trajectories leading to 13 and the secondary carbocation TS-2 is not preceded by any non-classical region. Pemberton and Tantillo29 hypothesised, based on these computational studies, that a bifurcation is likely to be encountered along the pathway from TS-2 to cations 12/13. The authors' claim needs to be verified through further computational and experimental biosynthetic studies.
3.3 Bifurcation in diterpene biosynthesis
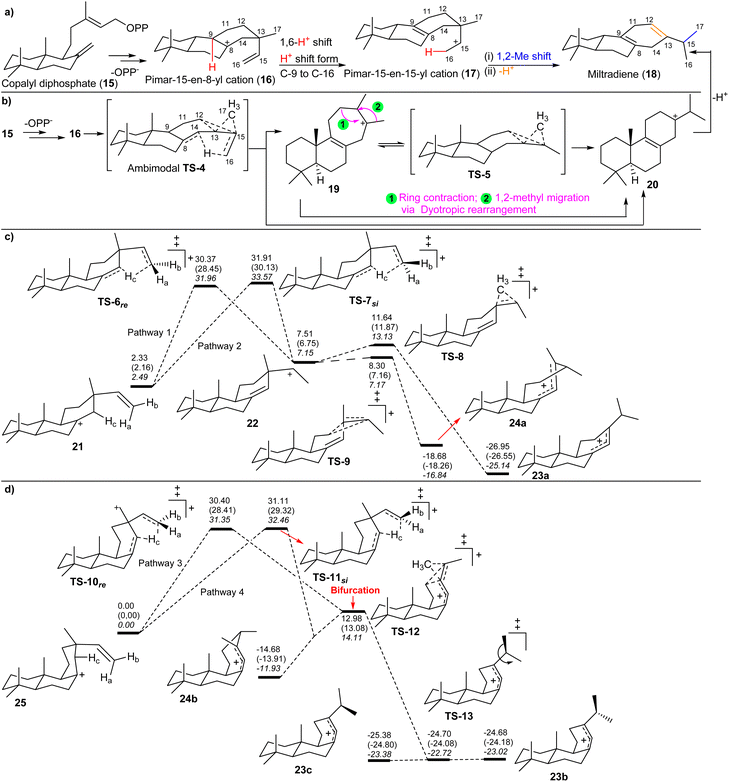 | ||
| Scheme 3 (a) Previous proposed mechanism for miltiradiene (18) formation.30 (b) A comprehensive proposed mechanism from Hong and Tantillo31,32 based on computations. (c) Energy profile of the reaction pathways from 21 that involve the secondary cation 22.31 (d) Energy profile of the reaction pathways from 25.31 Energies are reported relative to 25 in kcal mol−1. Three sets of values are shown: the values at the top of each set were computed using B3LYP/6-31+G(d,p); the values in parentheses were computed with mPW1PW91/6-31+G(d,p)//B3LYP/6-31 p G(d,p); and the italicized values were computed with MPWB1K/6-31+G(d,p)//B3LYP/6-31+G(d,p).31 Reproduced from ref. 31 and 32 with permission from the Springer Nature. | ||
Hong and Tantillo31,32 investigated the details of the diterpene miltiradiene (18) through computational analysis, as shown in Scheme 3c and d. The rearrangement of conformer 21 proceeds via intramolecular proton transfer to the π-face (pathways 1 and 2, Scheme 3c), resulting in the secondary cation 22. Subsequently, a low-barrier [1,2] methyl shift led to the formation of the isopropyl group in abietadiene (22 to 23a). Conformer 25 reacts differently. While TS structures for intramolecular proton transfer to the π-face were located as a saddle point on the PES (pathways 3 and 4, Scheme 3d), IRC calculations revealed that neither was directly linked to a secondary cation minimum. The TS-10re, the proton transfer TS to the re-face of the π-bond (pathway 3) appeared to be linked to another TS structure (TS-12), which is a characteristic of a bifurcating surface. IRC calculations on TS-12 revealed that it links particular conformers of cations 23a–c and 24a,bvia an unusual dyotropic rearrangement. These results therefore suggest that the pathway from TS-10re to the products is dynamically controlled. The resemblance of TS-12 to a hyper-conjugated secondary carbocation suggests this rearrangement involves concerted but highly asynchronous alkyl-shifting events. Calculations further indicate that proton transfer to the si face of the π-bond in 25 could also potentially result in the formation of either of the cations 23a–c and 24a,b. This is possible by passing through the region of the PES where a bifurcation occurs.
The exact nature of control by the enzyme has yet to be determined. Nevertheless, the very existence of multiple branches, together with the relatively flat energy surface of biosynthesis, is intriguing in terms of how nature may be able to control molecular diversity from a few starting materials. The findings of Hong and Tantillo32 demonstrated that the selectivity of organic reaction mechanisms can be influenced by dynamical tendencies together with complex potential energy surfaces. Thus, it is necessary to look beyond transition-state theory.
3.4 Bifurcation in saccharides
3-Deoxy-D-manno-octulosonate 8-phosphate (KDO8P, 31) is a component of the lipopolysaccharide in most Gram-negative bacteria. KDO8P synthase speeds up the condensation of PEP (26) and A5P (27) to create KDO8P (31) (Scheme 4a), a vital building block in the enzymatic synthesis of the endotoxin. Earlier studies have shown that there are two possible pathways for the condensation of PEP (26) and A5P (27). Pathway A entails the occurrence of a carbanion intermediate 28, which is formed through the attack of a hydroxide ion on C-2 of the PEP (26). Conversely, the path B involves a transient oxocarbenium zwitterionic intermediate 30, which is formed when the C-3 of the PEP (26) attacks the C-1 (aldehyde) of the A5P (27). This is subsequently followed by a reaction with water at C-2.33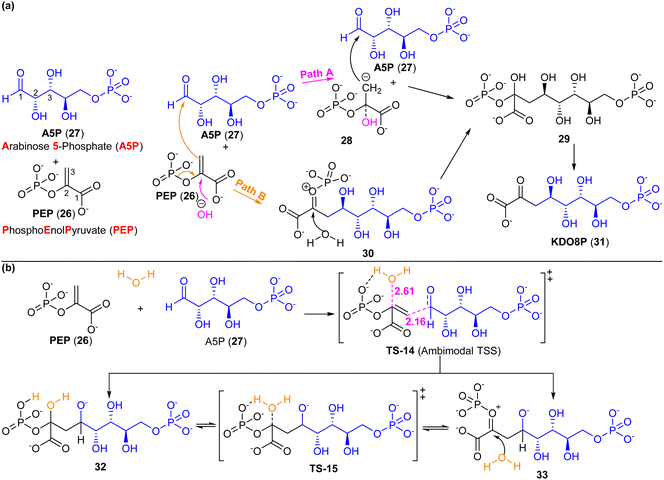 | ||
| Scheme 4 (a) Two stepwise mechanisms have been proposed for the condensation of PEP (26) and A5P (27) prior to computational studies. (b) Proposed mechanism from Tao et al. with a PTSB based on computations.33 Reproduced from ref. 33 with permission from the American Chemical Society. | ||
Tao et al.33 investigated the synthesis of KDO8P (formed by the condensation of PEP and A5P) in the C11N form of the Aquifex aeolicus KDO8PS enzyme, which serves as a non-metal analogue of the native enzyme. They used QM/MM calculations with ONIOM to construct the PESs that include all possible reaction pathways systematically. A number of potential mechanisms were examined. The lowest energy pathway corresponded to the syn addition of water to the si side of PEP (26) and the PEP (26) to the re side of A5P (27). This pathway had an ambimodal TS (TS-14, Scheme 4b), which could result in two products, 32 and 33.9,33 Product 32 has a lower energy than 33 and the system is not predicted to exist predominantly in the pure product 33 state. 32 and 33 can be interconverted via TS-15. Their findings revealed that the lowest energy reaction path does not align with any of the previously proposed stepwise mechanisms (Scheme 4a; pathway A and B). Instead, they concluded that the reaction proceeds through a partially concerted reaction mechanism involving PEP (26), A5P (27), and water. This highlights the involvement of PTSB (Scheme 4b) in the reaction between PEP (26) and A5P (27).9,33
Analyses were performed on the TS structures and reaction trajectories from computations. The C–O and C–C distances are 2.61 and 2.16 Å at the TS-14, which corresponds to a barrier of ∼14 kcal mol−1. The two hydrogen atoms in the water molecule are ∼1.8 Å from the carbonyl oxygen atom of A5P (27) and a phosphate oxygen atom of PEP (26), respectively. On trajectories from TS-14 towards the direction of TS-7, the C–O bond maintains a distance of 2.6 Å and the C–C bond is fully formed at 1.6 Å. The author suggested that this region of the PES corresponds to the location of the valley inflection point, from which the reaction can proceed to product 32 or 33.9,33
4 Ambimodal bispercyclic reactions
4.1 Ambimodal [4 + 2] vs. [6 + 4] cycloadditions with a stable [4 + 2] product
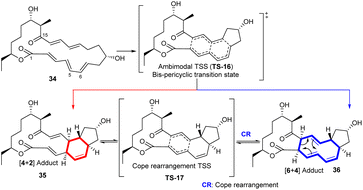 | ||
| Scheme 5 The ambimodal [4 + 2]/[6 + 4] cycloaddition involved in the biosynthesis of spinosyn A; The red arrow represents the path to the [4 + 2] adduct and the blue arrow represents the path to the [6 + 4] adduct. Reproduced from ref. 35 with permission from the American Chemical Society. | ||
Three independent research groups investigated the mechanistic details of the SpnF-catalysed [4 + 2] and [6 + 4] cycloaddition bifurcation computationally.36,40,41 Houk et al. studied the reaction in the enzyme using the environment-perturbed transition-state sampling (EPTSS) method, which involves QM/MM treatments to calculate the kinetic barriers.36 SpnF controls the outcome of the ambimodal reaction via TS-16 and disfavors the pathway to the [6 + 4] adduct 36 over Diels–Alder adduct 35. This fact, combined with the thermodynamic instability of the [6 + 4] adduct, justifies why scientists have never observed this product in experiments.34 In the enzyme environment, TS-16 is stablised by both intramolecular H-bonding and hydrophobic binding interactions. TS-16 also has a lower barrier compared to an alternative TS that goes directly to the Diels–Alder product.36 Quasiclassical dynamics simulations were performed and the outcomes of the trajectories were analyzed. In the gas phase, the ratio of 35 to 36 is 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1.1, whereas in water, the ratio is 1.6
1.1, whereas in water, the ratio is 1.6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. In the enzyme SpnF, 11
1. In the enzyme SpnF, 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 35 and 36 were obtained. In implicit hexane, the ratio of 35 to 36 is 1.5
1 35 and 36 were obtained. In implicit hexane, the ratio of 35 to 36 is 1.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, which indicates that the [4 + 2]-adduct is promoted over the [6 + 4]-adduct by an increase in solvent polarity.
1, which indicates that the [4 + 2]-adduct is promoted over the [6 + 4]-adduct by an increase in solvent polarity.
The enzyme SpnF influences the ambimodal TS geometry and the post-TS bifurcation dynamics through the active site. The enzymatic perturbation of the energy surface results in the favouring of the [4 + 2]-adduct formation. This illustration exemplifies how an enzyme operates on a vibrational timescale to direct post-transition state trajectories towards the Diels–Alder adduct.36
Zheng and Thiel40 suggest that enzymatic acceleration is largely caused by an H-bond between Thr196 and the C-15 carbonyl. This results in the withdrawal of electron density and enhanced reactivity of the dienophile. Furthermore, it is proposed that the enzyme facilitates the adoption of the s-cis C5–C6 substrate diene conformation. This conformation is a prerequisite for the cycloaddition, which may be stabilized by H-bonding between Trp256 and the C1 carbonyl. Further evidence demonstrated that, compared with the gas phase, the enzyme significantly lowered the barrier of TS-16, which was consistent with experimental outcomes. Examination of the optimised geometries at a QM/MM level suggests that the enzyme may reorganise the substrate within the active site. This reorganization accelerates the [4 + 2] cycloaddition and hinders the [6 + 4] cycloaddition via interactions with active-site residues.
Moreover, Chen et al.41 posit a series of non-covalent interactions (π–π stacking) between the substrate and the residues Trp256 and Tyr23, which facilitate the positioning of the substrate into a reactive conformation. In their model, the Thr196-C15 carbonyl H-bond is absent. Instead, the presence of intramolecular interactions within the TS, along with H-bonds between the TS and residues (Glu152, His42, and Trp256) contributes to a reduced barrier. The authors also propose that the trajectory from the ambimodal TS is influenced by the transfer of kinetic energy from hydrophobic residues to the substrate through femtosecond-scale vibrational collisions.
4.2 Ambimodal [4 + 2] vs. [6 + 4] cycloadditions with stable [6 + 4] products
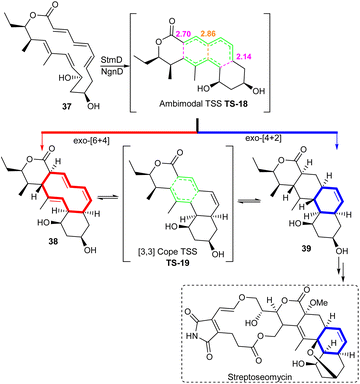 | ||
| Scheme 6 Streptoseomycin biosynthesis involves an ambimodal [6 + 4] cycloaddition/[4 + 2] Diels–Alder reaction catalyzed by StmD/NgnD. The red arrow represents the pathways to the [6 + 4] adduct and the blue arrow represents the pathway to the [4 + 2] adduct. Reproduced from ref. 42 with permission from the Springer Nature. | ||
DFT calculations demonstrated that the exo-[6 + 4] and exo-[4 + 2] adducts, 38 and 39, are generated from the reaction via a single transition state TS-18. Quasiclassical dynamics simulations show that the bifurcation predominantly leads to the [6 + 4] adduct 38 and the [4 + 2] adduct 39 is the minor product (3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio of 38/39). In TS-18, the partial C–C bond (2.70 Å) formed in the [6 + 4] adduct 38 is shorter compared to the partial C–C bond formed in the [4 + 2] product 39 (2.86 Å). Thermodynamically, 38 is also more stable than 39.42 Wang and his research team3 looked into the NgnD enzyme-catalysed PTSB reaction of [6 + 4] and [4 + 2] cycloaddition of compound 37 with the theozyme models to understand the role of key residues at the binding site. Specifically, the residues Y55, W67, M69 and Y13 influence the outcome of the ambimodal reaction, changing the product distribution compared to when the enzyme is absent.
1 ratio of 38/39). In TS-18, the partial C–C bond (2.70 Å) formed in the [6 + 4] adduct 38 is shorter compared to the partial C–C bond formed in the [4 + 2] product 39 (2.86 Å). Thermodynamically, 38 is also more stable than 39.42 Wang and his research team3 looked into the NgnD enzyme-catalysed PTSB reaction of [6 + 4] and [4 + 2] cycloaddition of compound 37 with the theozyme models to understand the role of key residues at the binding site. Specifically, the residues Y55, W67, M69 and Y13 influence the outcome of the ambimodal reaction, changing the product distribution compared to when the enzyme is absent.
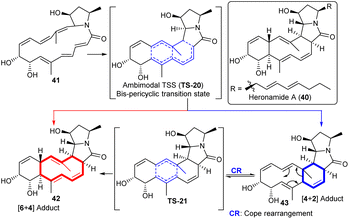 | ||
| Scheme 7 Heronamides biosynthesis involves an ambimodal [6 + 4] cycloaddition/[4 + 2] Diels–Alder reaction. The red arrow represents the pathway to the [6 + 4] adduct and the blue arrow represents the pathway to the [4 + 2] adduct. Reproduced from ref. 45 with permission from the American Chemical Society. | ||
The [6 + 4]/[4 + 2] ambimodal TS, TS-20, is bis-pericyclic in nature. In addition, the [6 + 4] adduct 42 is found to be more thermodynamically stable than the [4 + 2] adduct 43, which could convert into the [6 + 4] adduct 42via a Cope rearrangement TS, TS-21. The [4 + 2] product 43 is less stable due to steric repulsion between the two methyl groups and the combined effect of the tether. In a separate study, Zhang et al.46 performed quasiclassical dynamics simulations in both the water and gas phase environments. The trajectories suggest the ratio of [6 + 4] 42 to [4 + 2] adducts 43 is 56![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 in water compared to 16
1 in water compared to 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 in the gas phase. The [6 + 4] adduct 42 has a much greater dipole moment compared to [4 + 2] adducts 43. The molecule with a greater dipole moment is expected to be more stable in the polar water phase.
1 in the gas phase. The [6 + 4] adduct 42 has a much greater dipole moment compared to [4 + 2] adducts 43. The molecule with a greater dipole moment is expected to be more stable in the polar water phase.
4.3 Ambimodal intramolecular vs. hetero-Diels–Alder cycloaddition
Leporins are alkaloid compounds characterized by a tricyclic structure. The biosynthesis of leporin C (46) involves an ambimodal [4 + 2] hetero-Diels–Alder (HDA)/[4 + 2] intramolecular Diels–Alder (IMDA) cycloadditions (Scheme 8). The most direct pathway from (E)-QM (44) to leporin C (46) is via the HDA reaction, which is catalyzed by LepI, a pericyclase enzyme. Alternatively, the (E)-1 enzyme-bound compound can undergo competing IMDA cyclisation, producing the spirobicyclic product 45, which can then be released from the enzyme. The PTSB of the bis-pericyclic ambimodal TS (TS-22) controls the periselectivity. In the case of the IMDA route, LepI can then recapture the spirobicyclic intermediate 45 and convert it to the leporin C (46) by a retro-Claisen rearrangement (TS-23).47,48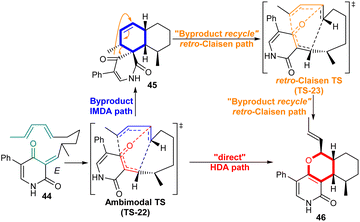 | ||
| Scheme 8 LepI-catalysed reactions to leporin C (46); (1) ‘direct’ path (HDA reaction), (2) ‘by-product recycle’ path (IMDA reaction/retro-Claisen rearrangement). The blue arrow represents the path to the IMDA adduct and the red arrow represents the path to the HDA adduct. Reproduced from ref. 47 with permission from the Springer Nature. | ||
Quasiclassical dynamic simulations confirmed that TS-22 is an ambimodal TS. TS-22 is a standard saddle point on the PES with temporarily formed bonds that can result in two distinct pericyclic products, the IMDA adduct 45 and the HDA adduct 46. (E)-44 is the reactant of the two products.47 The IMDA adduct 45 is preferred under non-enzymatic conditions because it has a shorter C–C bond (2.91 Å), which is formed in 45, compared to the longer C–O bond (3.34 Å), which is formed in 46, in TS-22. The experimental and computational findings demonstrated that the presence of water as the solvent increased the product ratio of HAD adduct 46. LepI also shifted the selectivity outcome of the post-ambimodal TS bifurcation towards the HDA adduct, much more significantly than in water.47
4.4 Ambimodal [4 + 2] vs. [2 + 4] cycloaddition
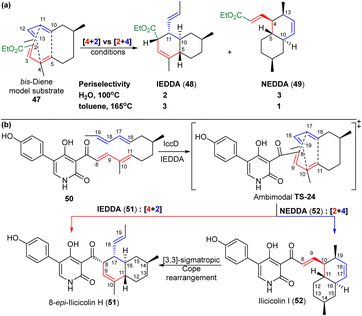 | ||
| Scheme 9 (a) Williams et al.49 model study using an ethyl ester of the acyclic precursor. (b) Ilicicolins biosynthesis involves an ambimodal [4 + 2] cycloaddition/[2 + 4] Diels–Alder reaction. Reproduced from ref. 49 and 50 with permission from the Elsevier and American Chemical Society respectively. | ||
Houk and Tang50 investigated the biosynthesis of ilicicolin H via a 2-pyridone-4-based Diels–Alder reaction involving a trans-bisdiene chain 50 (Scheme 9b). They first examined the periselectivity in the absence of pericyclase IccD (Scheme 9b). The reaction of 50 in toluene yielded 8-epi-ilicicolin H (IEDDA product; 51) and ilicicolin I (NEDDA product; 52) in a ratio of 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. Conversely, the reaction with IccD produced 99% of the IEDDA product 51 and less than 1% of the NEDDA product 52. The authors considered the possibility a conversion from product 52 to product 51via a [3,3]-sigmatropic Cope rearrangement, given that products 51 and 52 originates from the same facial- and endo-selective IEDDA and NEDDA reactions, respectively. However, product 52 did not convert to 51, either in the presence of IccD or under heating in water at 100 °C.
1. Conversely, the reaction with IccD produced 99% of the IEDDA product 51 and less than 1% of the NEDDA product 52. The authors considered the possibility a conversion from product 52 to product 51via a [3,3]-sigmatropic Cope rearrangement, given that products 51 and 52 originates from the same facial- and endo-selective IEDDA and NEDDA reactions, respectively. However, product 52 did not convert to 51, either in the presence of IccD or under heating in water at 100 °C.
The findings with the model substrate using computations are consistent with results from synthetic studies. The energy profile of the reaction from 50 to 51/52 in the gas phase was derived at the DFT level.50 The computations demonstrated that the pathway via ambimodal TS-24 is the most energetically favourable compared to other possibilities. TS-24 generates both the IEDDA adduct 51 (creating C11–C16 and C8–C17 bonds) and the NEDDA adduct 52 (creating C11–C16 and C10–C19 bonds) via a PTSB. The reaction from bis-diene 50 to cycloadducts 51 and 52 is extremely exothermic. Conversion from 51 to 52 or vice versa via a Cope rearrangement is unlikely to occur even with enzyme catalysis due to the large kinetic barrier (more than 40 kcal mol−1), although this Cope rearrangement TS saddle point is lower in energy on the PES compared to TS-24.
Studies of quasiclassical trajectories in the gas phase revealed that the products 51 and 52 were present in a ratio of 98![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, whereas the experimental ratio with the IccD enzyme was 99
2, whereas the experimental ratio with the IccD enzyme was 99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. Based on these results, the authors suggest that the IccD-catalysed reaction occurs in a hydrophobic pocket, directing the predominant formation of product 51via ambimodal TS-24.50
1. Based on these results, the authors suggest that the IccD-catalysed reaction occurs in a hydrophobic pocket, directing the predominant formation of product 51via ambimodal TS-24.50
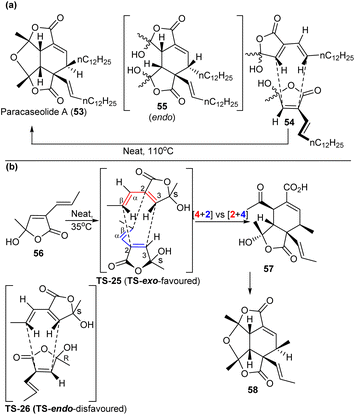 | ||
| Scheme 10 (a) Syntheses of paracaseolide A (53) from previous works, where thermal dimerization of alkenylbutenolide 54 is the key step. This cycloaddition has been presumed to proceed through the endo pathway, leading to 55. (b) The stereochemical outcome of spontaneous butenolide 56 dimerization cleanly produces the exo adduct 57 and then to 58. Reproduced from ref. 52 with permission from the Springer Nature. | ||
Hoye et al.52 investigated the biosynthesis of 53, with a particular focus on the dimerisation of 54via a Diels–Alder cycloaddition. They utilised a simpler methyl-bearing analogue 56 to gain valuable insights into the intricate mechanism. 56 behaved similarly to 54 in terms of reactivity in an enzyme-free [4 + 2] cycloaddition-based dimerisation reaction, which produced dimer 58 as the major product. X-ray analysis of the intermediate, 57, revealed that dimerisation is both regio- and stereoselective. The [4 + 2] cycloaddition occurs via a route in which the diene interacted with the dienophile in an exo orientation relative to the carbonyl group on the dienophile (Scheme 10b).
To understand the stereochemical outcome, Hoye et al.52 conducted a DFT study to calculate the relative energies of TS structures that correspond to the exo versus the endo modes of 56 dimerisation. In line with experimental observations, the lowest energy TS structure for the exo dimerisation is favoured by 3.9 kcal mol−1 compared to the endo pathway. The exo TS (TS-25) is preferred energetically over the endo TS (TS-26) due to its C2-symmetric geometry, which allows for enhanced stabilization through the delocalization of the unsaturated centre. The C2-symmetric characteristics also implies that the reaction proceeds through an exo TS that is bispericyclic in nature, where the competing [4 + 2] and [2 + 4] cycloaddition modes fully converged.
4.5 Ambimodal [4 + 2] vs. [4 + 3] cycloadditions
Delitschiapyrones A (59) and B (60) are products of the reaction between naphthoquinone 61 and 2-pyrone 62. Delitschiapyrone A (59) has an exceptional 6/6/7/5/6-fused pentacyclic ring scaffold. The postulated biosynthetic pathway (Scheme 11) showed that precursors 61 and 62 could react in an exo [4 + 2] fashion. The cycloadduct is delitschiapyrone B (60), which has not previously been isolated from cultures. Subsequently, an α-ketol rearrangement of 60 leads to ring expansion to form the seven-membered ring of molecule 63. The enzymes that catalyse the [4 + 2] cycloaddition/α-ketol rearrangement cascade remain unknown. The cyclisation of compound 63 finally resulted in the completion of the postulated biosynthesis of delitschiapyrone A (59).58 The intermolecular Diels–Alder reaction might be spontaneous, as suggested by the results of a bioinspired total synthesis of natural product 59, which involved just stirring a mixture of two Diels–Alder reactants.59 | ||
| Scheme 11 Ambimodal [4 + 2] and [4 + 3] cycloadditions are involved in the biosynthesis of delitschiapyrones. Reproduced from ref. 60 with permission from the American Chemical Society respectively. | ||
DFT studies by Houk et al.60 have shown that a 42.2 kcal mol−1 energy barrier must be overcome to generate compound 63 from compound 60. However, compound 63 is 5.9 kcal mol−1 less thermodynamically stable than compound 60. This energetics suggests that 63 cannot be formed. Nevertheless, a considerable quantity of 59, the spontaneous cyclization product of 63, was isolated from both the reaction mixture and the culture.58,59 Based on the disagreement between theory and experiment, Houk et al.60 predicted that the reaction does not follow the proposed cascade reaction pathway.
The mechanisms and reaction dynamics of the biosynthetic formation of compound 59 were then investigated60 using DFT calculations and quasiclassical dynamics simulations. Houk et al. propose that the reaction occurs through a Diels–Alder/α-ketol rearrangement or [4 + 2]/[4 + 3] ambimodal TS (TS-27). Structural analysis of the TS-27 revealed that the competing bond-forming distances of bonds 1 (C3–C13) and 2 (C4–C13) are similar. The post-transition state bifurcation from TS-27 results in the formation of either delitschiapyrone B (60) or a precursor 63 of delitschiapyrone A (59). This finding is highly significant for subsequent studies on the biosynthetic pathway of 59 involving gene deletion and heterologous expression.
5 Conclusion and future perspective
What determines the selectivity of a chemical reaction? This question has been the focus of extensive research. It is often assumed that the product ratio is influenced by either the relative free energies of the product (thermodynamic control) or by the activation barriers along the pathways to product formation (kinetic control). Recent studies have highlighted that features beyond thermodynamics and kinetics, namely dynamic effects, can significantly influence selectivity control.The likelihood of a single TS leading to multiple products expands the scope of potential reaction pathways, blurring the conventional differentiation between stepwise and concerted mechanisms. Such features on the PES shift selectivity control to after the TS, challenging the traditional view of kinetic control as being determined at the TS crossing. Evidence for post-TS bifurcations is increasingly observed in a growing number and complexity of synthetic and biosynthetic processes.12
This review summarizes examples of biosynthetic reactions featuring ambimodal transition states with bifurcating pathways beyond the TS. These cases demonstrate how selectivity in bioorganic reactions can arise from inherent dynamical tendencies together with complex PESs, and underscore the need to consider more than just the transition state when proposing the mechanism. These prior examples highlight that post-TS bifurcations are not a rare phenomenon. Caution is warranted when encountering similar scenarios in future studies—particularly in reactions involving carbocationic rearrangements, pericyclic reactions, and, perhaps more broadly, other biosynthesis reactions.
This shift in perspective also raises a central mechanistic question: to what extent can enzymes actively shape or bias ambimodal outcomes? For example, could enzymes reshape the PTSB energy surface to favour one pathway over another? Tantillo et al.32 have suggested that such dynamic selection plays a role in the biosynthesis of terpenes. While this proposal differs from the traditional concept of catalysis through lowering reaction barriers, it recognizes the inherent dynamic complexity of real reactions.
When selectivity forecasts based on transition state theory do not correspond with experimental findings, even after extensive benchmarking and conformational exploration, it will always be beneficial to investigate the role of dynamic effects. Several correlational models11 have been developed to provide a rapid estimation of selectivity based on the ambimodal TS. Quasiclassical dynamics simulations are more expensive but offer accurate predictions and more in-depth insights into the reaction mechanism. Combining these approaches enables a more comprehensive understanding of how reaction dynamics influence selectivity. However, this may not be sufficient for biosynthetic systems. Current computational studies mainly focus on the reactive substrate within the femtosecond timescale. Further investigations are needed to capture detailed interactions with the enzyme active site and/or pyrophosphate counterion across broader timescales. Given the increasing quantity and significance of such reactions in the domains of biosynthetic chemistry, there is demand for more sophisticated computational tools to predict reaction outcomes accurately.
6 Author contributions
H. H.: conceptualized, supervised, and edited the review process. K. N. H, C. C. L, S. S. and L. N. wrote, edited and proofread the manuscript.7 Conflicts of interest
There are no conflicts to declare.8 Data availability
There is no new data were created or analyzed in the study and that, therefore, data sharing is not applicable.9 Acknowledgments
H. Hussain is thankful to the Alexander von Humboldt Foundation for its generous support in providing the opportunity to do work in Germany which facilitated the writing of this review. K. N. H. thanks the National Science Foundation (CHE-2153972) for financial support. C. C. L. thanks Croucher Foundation and the Croucher Postdoctoral Fellowship program for the financial support of this project. L.N. gratefully acknowledges the support from the European Regional Development Project ENOCH #CZ.02.1.01/0.0/0.0/16_019/0000868, and the Czech Science Foundation Project #23-05474S.10 References
- S. J. Benkovic and S. Hammes-Schiffer, Science, 2003, 301, 1196–1202 CrossRef CAS.
- S. Marti, M. Roca, J. Andres, V. Moliner, E. Silla, I. Tunon and J. Bertran, Chem. Soc. Rev., 2004, 33, 98–107 RSC.
- Y. Hou, J. Chen, W. Liu, G. Zhu, Q. Yang and X. Wang, Molecules, 2024, 29, 5518 CrossRef CAS PubMed.
- D. J. Vocadlo, G. J. Davies, R. Laine and S. G. Withers, Nature, 2001, 412, 835–838 CrossRef CAS.
- A. L. Bowman, I. M. Grant and A. J. Mulholland, Chem. Commun., 2008, 4425–4427 RSC.
- R. Lonsdale, K. E. Ranaghan and A. J. Mulholland, Chem. Commun., 2010, 46, 2354–2372 RSC.
- C. E. Hornsby and R. S. Paton, Nat. Chem., 2014, 6, 88–89 CrossRef CAS.
- S. R. Hare and D. J. Tantillo, Beilstein J. Org. Chem., 2016, 12, 377–390 CrossRef CAS.
- S. R. Hare and D. J. Tantillo, Pure Appl. Chem., 2017, 89, 679–698 CrossRef CAS.
- D. J. Tantillo, WIREs Comput. Mol. Sci., 2020, 10, e1453 CrossRef CAS.
- C. C. Lam and K. N. Houk, Pure Appl. Chem., 2025, 97, 1135–1148 CrossRef CAS.
- C. C. Lam and J. M. Goodman, Chem. Sci., 2023, 14, 12355–12365 RSC.
- J. L. Bao and D. G. Truhlar, Chem. Soc. Rev., 2017, 46, 7548–7596 RSC.
- P. Bharadwaz, M. Maldonado-Domınguez and M. Srnec, Chem. Sci., 2021, 12, 12682–12694 RSC.
- D. H. Ess, S. E. Wheeler, R. G. Iafe, L. Xu, N. Celebi-Ölcüm and K. N. Houk, Angew. Chem., Int. Ed., 2008, 47, 7592–7601 CrossRef CAS PubMed.
- W. Shin, Y. Hou, X. Wang and Z. J. Yang, J. Chem. Theory Comput., 2024, 20, 10942–10951 CrossRef CAS PubMed.
- X. S. Xue, C. S. Jamieson, M. Garcia-Borràs, X. Dong, Z. Yang and K. N. Houk, J. Am. Chem. Soc., 2019, 141, 1217–1221 CrossRef CAS.
- A. Martin-Somer, X. S. Xue, C. S. Jamieson, Y. Zou and K. N. Houk, J. Am. Chem. Soc., 2023, 145, 4221–4230 CrossRef CAS.
- X. Wang, C. Zhang, Y. Jiang, W. Wang, Y. Zhou, Y. Chen, B. Zhang, R. X. Tan, H. M. Ge, Z. J. Yang and Y. Liang, J. Am. Chem. Soc., 2021, 143, 21003–21009 CrossRef CAS PubMed.
- M. Weitman and D. T. Major, J. Am. Chem. Soc., 2010, 132, 6349–6360 CrossRef CAS PubMed.
- D. T. Major and M. Weitman, J. Am. Chem. Soc., 2012, 134, 19454–19462 CrossRef CAS.
- Y. J. Hong and D. J. Tantillo, Org. Biomol. Chem., 2010, 8, 4589–4600 RSC.
- D. A. Whittington, M. L. Wise, M. Urbansky, R. M. Coates, R. B. Croteau and D. W. Christianson, Proc. Natl. Acad. Sci. U. S. A., 2002, 99, 15375–15380 CrossRef CAS.
- S. Jaisamut, S. Prabpai, C. Tancharoen, S. Yuenyongsawad, S. Hannonbua, P. Kongsaeree and A. Plubrukarn, J. Nat. Prod., 2013, 76, 2158–2161 CrossRef CAS.
- F. Le Bideau, M. Kousara, L. Chen, L. Wie and F. Dumas, Chem. Rev., 2017, 117, 6110 CrossRef CAS PubMed.
- C. H. McCulley and D. J. Tantillo, J. Phys. Chem. A, 2018, 122, 8058–8061 CrossRef CAS PubMed.
- D. C.-M. Sim, N. L. Hungerford, E. H. Krenske, G. K. Pierens, K. T. Andrews, T. S. Skinner-Adams and M. J. Garson, Aust. J. Chem., 2020, 73, 129–136 CrossRef CAS.
- S. R. Hare, R. P. Pemberton and D. J. Tantillo, J. Am. Chem. Soc., 2017, 139, 7485–7493 CrossRef CAS.
- R. P. Pemberton and D. J. Tantillo, Chem. Sci., 2014, 5, 3301–3308 RSC.
- W. Gao, M. L. Hillwig, L. Huang, G. Cui, X. Wang, J. Kong, B. Yang and R. J. Peters, Org. Lett., 2009, 11, 5170–5173 CrossRef CAS PubMed.
- Y. J. Hong and D. J. Tantillo, Nat. Chem., 2009, 1, 384–389 CrossRef CAS.
- Y. J. Hong and D. J. Tantillo, Nat. Chem., 2014, 6, 104–111 CrossRef CAS.
- P. Tao, D. L. Gatti and H. B. Schlegel, Biochemistry, 2009, 48, 11706–11714 CrossRef CAS.
- H. J. Kim, M. W. Ruszczycky, S. Choi, Y. Liu and H. Liu, Nature, 2011, 473, 109–112 CrossRef CAS PubMed.
- A. Patel, Z. Chen, Z. Yang, O. Gutierrez, H. W. Liu, K. N. Houk and D. A. Singleton, J. Am. Chem. Soc., 2016, 138, 3631–3634 CrossRef CAS PubMed.
- Z. Yang, S. Yang, P. Yu, Y. Li, C. Doubleday, J. Park, A. Patel, B. Jeon, W. K. Russell, H. Liu, D. H. Russell and K. N. Houk, Proc. Natl. Acad. Sci., 2018, 115, E848–E855 CAS.
- M. G. Medvedev, A. A. Zeifman, F. N. Novikov, I. S. Bushmarinov, O. V. Stroganov, I. Y. Titov, G. G. Chilov and I. V. Svitanko, J. Am. Chem. Soc., 2017, 139, 3942–3945 CrossRef CAS.
- C. D. Fage, E. A. Isiorho, Y. Liu, D. T. Wagner, H. Liu and A. T. Keatinge-Clay, Nat. Chem. Biol., 2015, 11, 256–258 CrossRef CAS.
- B. A. Hess and L. Smentek, Org. Biomol. Chem., 2012, 10, 7503–7509 RSC.
- Y. Zheng and W. Thiel, J. Org. Chem., 2017, 82, 13563–13571 CrossRef CAS PubMed.
- N. Chen, F. Zhang, R. Wu and B. A. Hess, ACS Catal., 2018, 8, 2353–2358 CrossRef CAS.
- B. Zhang, K. B. Wang, W. Wang, X. Wang, F. Liu, J. Zhu, J. Shi, L. Y. Li, H. Han, K. Xu, H. Y. Qiao, X. Zhang, R. H. Jiao, K. N. Houk, Y. Liang, R. S. Tan and H. M. Ge, Nature, 2019, 568, 122–126 CrossRef CAS PubMed.
- R. Raju, A. M. Piggott, M. M. Conte and R. J. Capon, Org. Biomol. Chem., 2010, 8, 4682–4689 RSC.
- R. Sugiyama, S. Nishimura, N. Matsumori, Y. Tsunematsu, A. Hattori and H. Kakeya, J. Am. Chem. Soc., 2014, 136, 5209–5212 CrossRef CAS PubMed.
- P. Yu, A. Patel and K. N. Houk, J. Am. Chem. Soc., 2015, 137, 13518–13523 CrossRef CAS PubMed.
- C. Zhang, X. Wang, Y. Chen, Z. He, P. Yu and Y. Liang, J. Org. Chem., 2020, 85(14), 9440–9445 CrossRef CAS PubMed.
- M. Ohashi, F. Liu, Y. Hai, M. Chen, M. Tang, Z. Yang, M. Sato, K. Watanabe, K. N. Houk and Y. Tang, Nature, 2017, 549, 502–506 CrossRef.
- Y. Cai, Y. Hai, M. Ohashi, C. S. Jamieson, M. Garcia-Borras, K. N. Houk, J. Zhou and Y. Tang, Nat. Chem., 2019, 11, 812–820 CrossRef CAS PubMed.
- D. R. Williams, R. D Gaston and I. B. Horton, Tetrahedron Lett., 1985, 26, 1391–1394 CrossRef CAS.
- Z. Zhang, C. S. Jamieson, Y. L. Zhao, D. Li, M. Ohashi, K. N. Houk and Yi Tang, J. Am. Chem. Soc., 2019, 141, 5659–5663 CrossRef CAS.
- X. L. Chen, H. L. Liu, J. Li, G. R. Xin and Y. W. Guo, Org. Lett., 2011, 13, 5032–5035 CrossRef CAS PubMed.
- T. Wang and T. R. Hoye, Nat. Chem., 2015, 7, 641–645 CrossRef CAS PubMed.
- J. P. Yin, C. L. Tang, L. X. Gao, W. P. Ma, J. Y. Li, Y. Li, J. Li and F. J. Nan, Org. Biomol. Chem., 2014, 12, 3441–3445 RSC.
- D. Noutsias and G. Vassilikogiannakis, Org. Lett., 2012, 14, 3565–3567 CrossRef CAS.
- L. Vasamsetty, F. A. Khan and G. Mehta, Tetrahedron Lett., 2013, 54, 3522–3525 CrossRef CAS.
- D. S. Giera and C. B. W. Stark, RSC Adv., 2013, 3, 21280–21284 RSC.
- J. Boukouvalas and M. A. Jean, Tetrahedron Lett., 2014, 55, 4248–4250 CrossRef CAS.
- J. G. Luo, X. B. Wang, Y. M. Xu, J. M. U'Ren, A. E. Arnold, L. Y. Kong and A. A. Gunatilaka, Org. Lett., 2014, 16, 5944–5947 CrossRef CAS.
- K. Kurasawa, E. Kwon, S. Kuwahara and M. Enomoto, Org. Lett., 2018, 20(15), 4645–4648 CrossRef CAS PubMed.
- Y. Zou and K. N. Houk, J. Am. Chem. Soc., 2021, 143, 11734–11740 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |




