Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Nanoparticle ultrastructure allows reversible pH sensitivity using 19F NMR and in vivo19F MRI
Alvja
Mali
a,
Mariah
Daal
a,
Natalia
Jirát-Ziółkowska
 bc,
Nicolas
Stumpe
d,
Naiara
Larreina Vicente
a,
N. Koen
van Riessen
a,
Visakh V. S.
Pillai
e,
Francesco Simone
Ruggeri
e,
Cyril
Cadiou
f,
Françoise
Chuburu
bc,
Nicolas
Stumpe
d,
Naiara
Larreina Vicente
a,
N. Koen
van Riessen
a,
Visakh V. S.
Pillai
e,
Francesco Simone
Ruggeri
e,
Cyril
Cadiou
f,
Françoise
Chuburu
 f,
Daniel
Jirak
f,
Daniel
Jirak
 bg,
Paul B.
White
h and
Mangala
Srinivas
bg,
Paul B.
White
h and
Mangala
Srinivas
 *ai
*ai
aDepartment of Cell Biology and Immunology, Wageningen University and Research, P. O. Box 338, 6700 AH Wageningen, The Netherlands. E-mail: mangala.srinivas@wur.nl
bInstitute for Clinical and Experimental Medicine, Prague, Czech Republic
cInstitute of Biophysics and Informatics, First Faculty of Medicine, Charles University, Prague, Czech Republic
dInstitute for Molecular Cardiology, Heinrich Heine University, Düsseldorf, Germany
ePhysical Chemistry and Soft Matter, Wageningen University and Research, Wageningen, The Netherlands
fUniversity of Reims Champagne Ardenne, CNRS, ICMR UMR 7312, Reims, France
gFaculty of Health Studies, Technical University of Liberec, Liberec, Czech Republic
hInstitute for Molecules and Materials, Radboud University, Nijmegen, The Netherlands
iCenya Imaging B.V., Amsterdam, The Netherlands
First published on 28th January 2026
Abstract
Fluorine-19 magnetic resonance imaging (19F MRI) is a powerful imaging modality that provides background-free imaging. Its sensitivity can be modulated through environmentally responsive probe designs exploiting paramagnetic relaxation enhancement (PRE) and related mechanisms. Here, we present a novel pH-sensitive 19F MRI nanosystem in which signal activation is governed by nanoparticle (NP) ultrastructure. Poly(lactic-co-glycolic acid) (PLGA) NPs were synthesized co-encapsulating perfluoro-15-crown-5-ether (PFCE) and lipophilic gadolinium (Gd) chelates, yielding either single-core or multi-core structures. While single-core NPs, the conventional structure for polymer-encapsulated perfluorocarbons, remained pH-insensitive, multi-core NPs exhibited a pronounced and reversible pH-dependent 19F signal modulation. The 19F T1 decreased slightly (from ∼700 to ∼600 ms) in both NP types due to the presence of Gd, whereas T2 shortened markedly at neutral pH (∼20 ms, “OFF” state) and increased substantially under acidic conditions (“ON” state); this variation was absent in single-core NPs. The pH-dependent 19F T2 behaviour was consistent across both high-field (14 T) and low-field (3 T) MRI. Such behaviour of multi-core NPs was validated in vitro and in vivo, as the 19F signal reappeared only after lysosomal internalization in RAW 264.7 cells and was selectively activated within acidic tumour regions, but not at the neutral subcutaneous injection site. By exploiting nanoparticle ultrastructure to achieve reversible signal switching, this work introduces a general strategy for activatable 19F MRI probes for imaging applications in tumours and other pH-sensitive disease environments. It demonstrates, for the first time, PRE-driven signal enhancement and suppression governed by structural rearrangement rather than chemical modification in response to pH variation.
1 Introduction
Non-invasive imaging techniques are crucial for early disease detection, driving the need for more sensitive and specific molecular imaging probes. Among emerging strategies, nanoparticle (NP)-based contrast agents have gained particular attention for their ability to enhance imaging sensitivity, improve targeting, and support multimodal imaging. Their tunable physicochemical properties, such as size, surface charge, and composition, enable control over circulation time, tissue accumulation, and contrast agent encapsulation, making them adaptable tools for a wide range of imaging modalities.1,2 These advantages have made NPs increasingly relevant in biomedical applications aimed at visualizing molecular and cellular processes in vivo.3 Clinical imaging enables real-time, non-invasive monitoring of physiological and pathological conditions, with magnetic resonance imaging (MRI) offering superior soft-tissue contrast, deep tissue penetration, and high spatial resolution without ionizing radiation.4However, conventional proton (1H) MRI suffers from strong background signals caused by the high-water content in biological tissues (65–90%), reducing contrast and making it difficult to distinguish specific targets, particularly early-stage tumours.4–6 This has led to growing interest in alternative MRI-active nuclei, such as 23Na,719F,8,9 and 31P,10,11 which offer improved specificity and signal clarity. Among them, fluorine-based MRI (19F MRI) has gained significant attention due to its high sensitivity, comparable to that of protons, and its exceptional specificity, as fluorine is nearly absent in biological tissues (≤10−6 M).12,13 These characteristics have made 19F MRI a powerful tool for diverse applications, including tracking cellular therapies and quantitative imaging.14–17 Among the different fluorine labels, perfluorocarbons (PFCs) are the most commonly used in 19F MRI, owing to their high fluorine content, excellent molecular mobility, and favourable safety profile.18 PFC-based agents have also been used clinically.19,20
Beyond direct fluorine imaging, activatable 19F MRI probes enhance sensitivity by exploiting changes in relaxivity6,21 or chemical shift22,23 to detect biological processes or trigger drug release.24,25 Stimulus-induced relaxivity alterations can create on/off signal switches, significantly improving 19F MRI sensitivity. These effects are primarily regulated by their structural properties, particularly their influence on transverse relaxation (T2).26 Two key strategies regulate 19F MR signal modulation: (1) fluorine nuclei tethered to paramagnetic centres via stimuli-responsive linkers, where proximity modulates the paramagnetic relaxation enhancement (PRE) effect,27–29 and (2) reversible fluorine agent aggregation, where restricted motion quenches the signal and disassembly restores it.21,30,31 PRE is highly distance-dependent, following an inverse sixth power relationship (r−6) with fluorine–chelate separation.32–34 Thus, precise nanoscale control over molecular reconfigurations is essential for responsive probe design.26,35 Nanomaterials provide ideal platforms for designing such activatable probes, as their tuneable physicochemical properties allow optimization of PRE effects and fluorine relaxation behaviour for enhanced responsiveness.36,37
Among external stimuli, pH is particularly relevant, as it is tightly regulated in normal tissues (7.2–7.4) but disrupted under pathological conditions such as inflammation and cancer.38 Tumours exhibit a characteristic acidic extracellular microenvironment due to metabolic reprogramming.39,40 As a hallmark of disease, these variations make pH-sensitive imaging valuable for detection and monitoring.41
Most pH-responsive polymer NPs used for 19F MRI contain ionizable groups, such as tertiary amines, with varying acid dissociation constants (pKa values).42–46 Fluctuations in pH induce protonation or deprotonation of these groups, leading to polymer swelling or shrinking, which indirectly modulates47 the fluorine signal.48,49 However, a key challenge for these agents remains increasing the usable fluorine signal while maintaining sufficient fluorine mobility.45 Higher fluorine content often increases hydrophobicity and fluorophilicity, leading to aggregation in aqueous environments, which reduces molecular mobility and signal intensity, and may cause toxicity risks in vivo.18,24
Our previous research demonstrated the successful co-encapsulation of lipophilic gadolinium (Gd) chelates and perfluoro-15-crown-5-ether (PFCE) within a polymer matrix (poly(D,L-lactide-co-glycolide), PLGA), yielding NPs with either multi-core or single-core structures, as confirmed by Small-Angle Neutron Scattering (SANS).50–52
While PRE in PFCs typically requires fluorophilic chelators,53,54 in our earlier methodological study,51 we showed that NP ultrastructure alone significantly influences 19F MR relaxation, with multi-core NPs exhibiting stronger paramagnetic effects than single-core NPs. That same work also revealed only preliminary indications of pH-sensitive 19F MR relaxation, uniquely associated with the multi-core configuration, without systematic mechanistic analysis or validation under physiologically relevant conditions. Moreover, the internal structure was previously shown to influence in vivo clearance52 and in vitro cellular uptake.55
Building on these findings, the present study systematically investigates the mechanism, reversibility, and physiological relevance of this pH responsiveness. Using both high- and low-field MRI (14 T and 3 T), we observed pH-dependent 19F MRI signal changes, with multi-core NPs exhibiting pronounced pH responsiveness even under physiologically relevant conditions. In proof-of-concept in vivo experiments (7 T MR), a 19F signal was observed only in the tumour and not subcutaneously. Overall, we have built on earlier structural observations towards a translational imaging context, with further mechanistic understanding of the reversibility of the signal generation.
2 Results and discussion
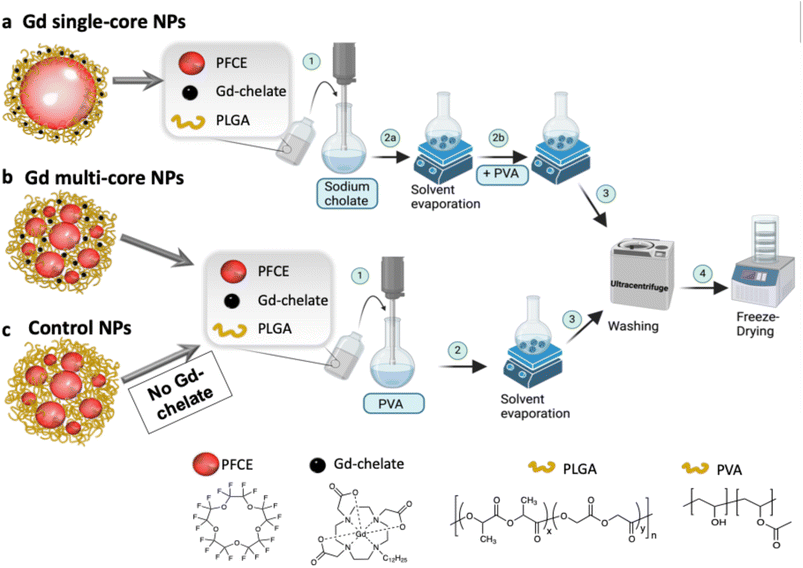
PFCs are advantageous for 19F MRI due to their high fluorine content but present challenges such as fluorophilicity, low surface tension, and slow clearance.18 Customizable nanocarriers, such as lipid- or polymer-based NPs, address these limitations.18,56–58 Our group has focused on optimizing and characterizing PLGA-loaded PFCE NPs, demonstrating that slight procedural modifications yield NPs with either multi-core or single-core ultrastructure (Fig. 1a and b).50,59 The ultrastructure plays a crucial role in PFCE clearance, with multi-core NPs exhibiting 15-fold faster clearance in vivo compared to single-core NPs.52 These structural differences also influenced MR properties, particularly when a lipophilic Gd-chelate was co-encapsulated with PFCE in the PLGA matrix. Gd entrapment increased with the amount of Gd-chelate added, with higher encapsulation observed for chelates containing longer hydrophobic chains.51 However, the consistent pH sensitivity across different Gd chelates suggests that the NP ultrastructure, rather than the chelator itself, determines the response. Based on these findings, only the chelator with the longer hydrophobic tail was used because of its slightly higher encapsulation efficiency.51 In this study, three types of NPs were investigated: (1) Gd single-core, (2) Gd multi-core, and (3) control NPs (Gd-free) (Fig. 1a–c). Gd single-core and multi-core NPs were used to investigate the role of NP ultrastructure in pH sensitivity, while control NPs served to determine whether the observed effects were intrinsic to the PLGA matrix or specifically due to the co-encapsulation of Gd-chelate and PFCE.To ensure comparability across pH sensitivity experiments, NPs were standardized based on (1) Gd entrapment measured by ICP-OES, (2) PFCE entrapment (% wt), and (3) T1 and T2 relaxation times measured by 19F NMR (Table 1). Due to inherent batch-to-batch variation, Table 1 reports average values from multiple batches used in this study. While Gd entrapment remained consistent across NP types, PFCE entrapment varied slightly, with Gd multi-core NPs exhibiting the lowest encapsulation efficiency. The co-encapsulation of Gd-chelate did not significantly alter the NP parameters, such as diameter and PDI (Table S1), with multi-core NPs (∼200 nm, PDI 0.2) and single-core NPs (∼160 nm, PDI 0.15) showing similar size distributions.
| NP type | Batches | Gd entrapped µg (Gd) per mg (NP) | 19F T1 (ms) | 19F T2 (ms) | PFCE% wt |
|---|---|---|---|---|---|
| Gd multi-core | 9 | 2.3 ± 0.7 | 550 ± 40 | 21 ± 9 | 14 ± 2 |
| Gd single-core | 4 | 2.0 ± 1.0 | 640 ± 15 | 55 ± 10 | 18 ± 4 |
| Control | 5 | N/A | 750 ± 35 | 390 ± 50 | 22 ± 4 |
The encapsulation of paramagnetic chelates significantly affected 19F MR properties, particularly in multi-core NPs, where it induced line broadening, potentially affecting quantification accuracy. The paramagnetic effect was more pronounced in T2 than in T1, with greater T2 shortening observed in multi-core NPs due to T2's higher sensitivity to local magnetic field variations, which are amplified by paramagnetic agents such as Gd.32,60 The effect of paramagnetic ions on the relaxation times of 19F nuclei depends on their distance from the fluorinated core.54,61,62 Previous SANS measurements indicated that each PFCE core in multi-core NPs has a radius of 9–12 nm with a 4 nm shell thickness, while single-core NPs have a core radius of approximately 20 nm with a similar shell thickness.50 This configuration results in a maximum Gd to PFCE distance of approximately 15 nm in multi-core NPs, compared to about 24 nm in single-core NPs.51 Given that PRE decreases steeply with increasing distance,35 the robust PRE effect observed in multi-core NPs suggests that additional structural factors, such as PFCE confinement or restricted mobility, may enhance fluorine relaxation beyond what would be expected based on spatial averages alone. In contrast, the larger, less confined single-core NPs allow for greater PFCE mobility, reducing these interactions and weakening PRE. These structural differences likely account for the stronger PRE effect observed in multi-core NPs.
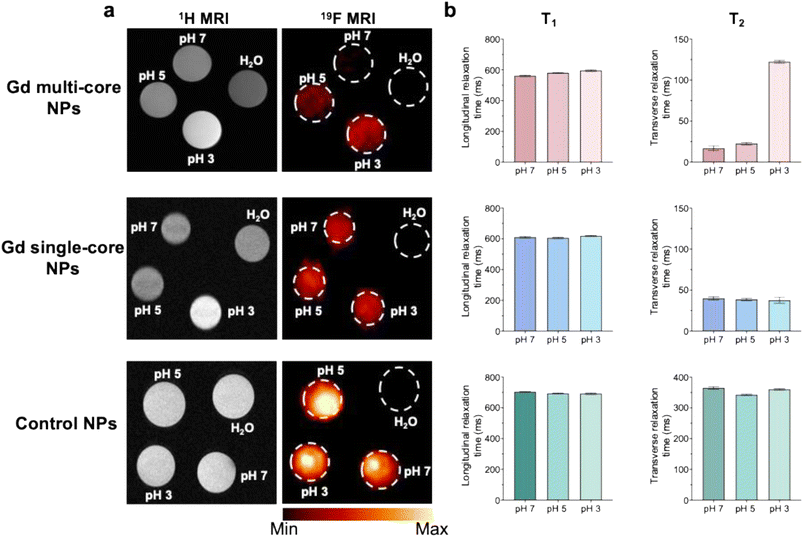
19F PRE has long been used to improve 19F MR signal sensitivity and to develop stimuli-responsive systems by incorporating specific functional groups sensitive to environmental changes, which modulate the 19F signal.18,24,26,28 Beyond chemical modifications, the internal structure of PLGA NPs also influences PRE upon pH response. Indeed, Gd single-core NPs showed minimal changes in 19F relaxation times across different pH conditions. In contrast, multi-core NPs exhibited a pronounced response to pH variations, with T2 relaxation times increasing 15-fold from neutral to acidic pH.5119F MRI at 14 T (Fig. 2a) confirmed these observations, showing a marked 19F signal increase for multi-core NPs under acidic conditions, whereas no changes were observed in single-core and control NPs. The T2 relaxation time of multi-core NPs increased with decreasing pH, while the T1 and T2 values for control NPs and Gd single-core remained stable (Fig. 2b and S1). Notably, even at acidic pH, the T2 of Gd multi-core NPs does not fully recover to the level observed for pristine PFCE NPs. This likely reflects residual PRE effects from the encapsulated Gd chelate, which continues to influence nearby 19F nuclei.
In addition, control NPs exhibited higher 19F signal intensity compared to Gd-loaded NPs, attributed to their higher PFCE content (% wt) (Table 1). Additionally, the defined area of higher intensity (Fig. 2a) likely reflects NP sedimentation due to the shape of the sample tube.
While 19F PRE effects are primarily dictated by the proximity and interaction of Gd chelate with the fluorinated core, the enhanced 1H MRI contrast observed at pH 3 for Gd multi-core NPs (Fig. 2a) suggests that acidic environments induce structural changes in the polymer matrix, increasing water permeability and Gd–water interactions. The 1H T1 relaxation time (Table S2 and Fig. S2) further supports this, showing a significant reduction in T1 at pH 3, while T1 values at pH 5 and 7 remain comparable. These findings suggest that acidic pH induces polymer swelling, likely due to carboxylic group protonation, which reduces rigidity and enhances water permeability, ultimately facilitating water access to Gd and amplifying its paramagnetic effects. Previous NMR spectroscopy studies corroborate this, confirming water's presence within the polymer network and its proximity to the PFCE cores.50
To confirm the pH sensitivity observed at 14 T, multi-core NPs were further investigated using a low-field (3 T) MRI scanner (Fig. 3). The results at 3 T were consistent with those at 14 T, as T1 and T2 of control NPs remained constant across pH conditions (Fig. 3b and d). In contrast, multi-core NPs exhibited an increase in T2 under acidic conditions (Fig. 3c), corresponding to a brighter 19F signal and reinforcing their pH responsiveness.
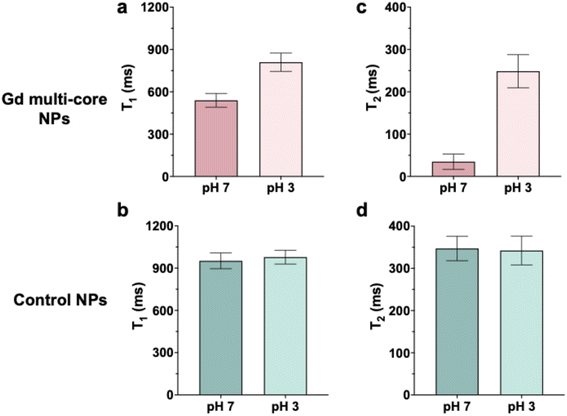 | ||
| Fig. 3 19F T1 (a and b) and T2 (c and d) relaxation times of Gd multi-core and control NPs. All samples were measured at the same concentration 6.5 mg mL−1 on a 3 T MR scanner. | ||
Given the lack of pH sensitivity in single-core NPs, subsequent investigations focused on multi-core NPs with Gd-free PLGA–PFCE NPs used as controls to confirm the effect of Gd-chelate interactions rather than intrinsic polymer properties.
To rule out the possibility that the observed effects were due to PLGA hydrolysis or Gd-chelate release under acidic conditions, the stability of encapsulated Gd was previously confirmed using 1H NMR.51 Furthermore, Gd release was mimicked by supplementing GdCl3 (gadolinium chloride) to Gd-free NPs; while proton relaxation was similarly affected, the 19F signal remained unchanged, confirming that the lipophilic Gd-chelates remained confined within the hydrophobic PLGA matrix even in acidic environments.51
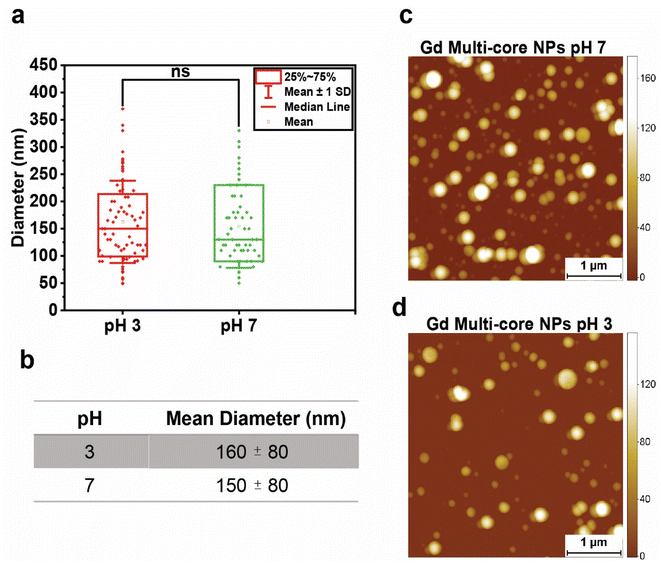
With hydrolysis excluded, the pH-dependent effect was next examined in terms of possible NP swelling, which remained undetectable by DLS (Fig. S3 and Tables S3, S4). Therefore, the size of Gd multi-core NPs was measured using AFM at both acidic (pH 3) and neutral (pH 7) pH. While statistical analysis showed no significant size difference (Fig. 4a), the mean diameter at pH 3 was slightly larger (∼10 nm, Fig. 4b). Although this increase was not statistically significant, even minor changes in nanoscale dimensions could, in principle, influence the PRE effect due to its high sensitivity to minor distance variations in effective Gd–19F distance.34
Importantly, the absence of a statistically significant change in particle size indicates that geometric expansion alone is unlikely to account for the pronounced pH-dependent changes observed in 19F relaxation. Instead, these results suggest that additional mechanisms contribute, including pH-induced alterations in internal nanoparticle dynamics, such as increased PFCE mobility or changes in the effective correlation time within the polymer matrix, which can strongly modulate PRE without requiring large changes in average Gd–19F separation.
AFM imaging (Fig. 4c and d) further supported the structural stability of Gd multi-core NPs, showing that the NPs retained a spherical shape across different pH levels.
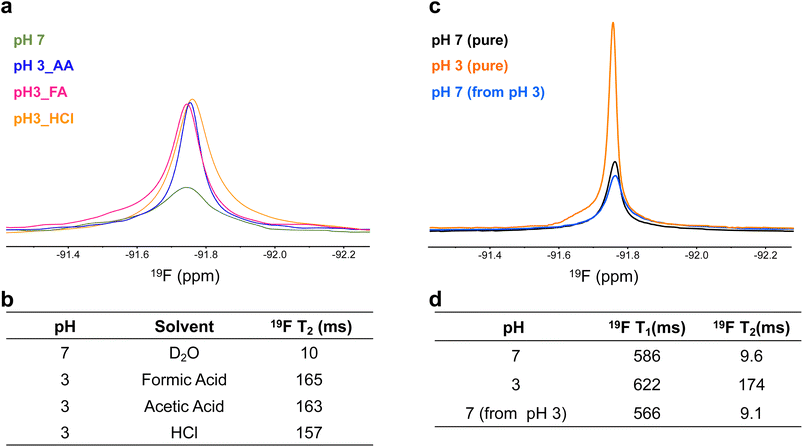
To determine whether this effect was driven solely by increased permeability and swelling or also influenced by the surrounding environment, 19F signal intensity and T2 values were examined in different acidic media. Gd multi-core NPs were resuspended in formic acid, acetic acid, HCl, and D2O. The consistent 19F signal intensities and T2 values (Fig. 5a and b) across these conditions suggest that the pH sensitivity is inherent to the NP structure rather than specific ion interactions. Furthermore, DLS measurements confirmed that the particle size remained stable, indicating that the NPs were not structurally degraded by the acid exposure (Table S3 and Fig. S3).
A key feature of the Gd multi-core NPs is their reversible pH-dependent 19F signal modulation, as observed by 19F NMR. To investigate this reversibility, samples from the same batch were compared: one resuspended at pH 7, one at pH 3, and the latter subsequently neutralized to pH 7 using 5 N NaOH. Initially, the 19F signal at pH 3 was higher, but upon neutralization, it decreased to match that of the sample originally resuspended at pH 7 (Fig. 5c). This behaviour aligns with the increase in T2 values under acidic conditions (Fig. 5d). Reproducibility of this reversible behaviour across independently prepared nanoparticle batches is shown in the SI (Table S5).
DLS measurements confirmed that the NPs remained structurally stable throughout the pH transition, with no effective changes in size or PDI (Table S4). A similar reversible effect was observed in NPs resuspended in acetic acid (Table S6), reinforcing that the pH responsiveness arises from the NP ultrastructure rather than the acidic medium itself.
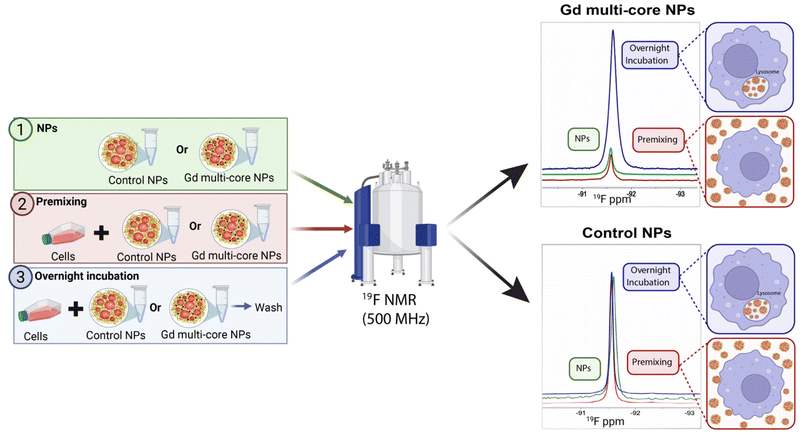
To further investigate the pH responsiveness of Gd multi-core NPs under biologically relevant conditions, in-cell 19F NMR spectroscopy was performed using RAW 264.7 macrophages. Previous phantom experiments were conducted under non-physiological pH conditions, which do not fully reflect the moderate acidity of cellular compartments. Thus, assessing whether the NPs exhibit similar pH sensitivity under physiological conditions was necessary. For this purpose, cells were labelled with Gd multi-core and control NPs and analysed via19F NMR in a Shigemi tube containing culture medium with 5% D2O. Cell viability remained high before and after measurements.
Three experimental conditions were analysed: (1) NPs in medium, to assess the influence of the medium's biochemical composition on pH responsiveness, (2) premixing, representing extracellular NP presence at pH 7, and (3) overnight incubation, mimicking NP uptake and intracellular processing (Fig. 6).
PLGA-based particles of this size are primarily internalized through fluid-phase pinocytosis and clathrin-mediated endocytosis.63,64 Due to their terminal carboxylic acid groups, negatively charged PLGA NPs tend to accumulate in lysosomes, unlike positively charged NPs, which can escape lysosomes and exhibit perinuclear localization.65 Therefore, we expected that the acidic lysosomal environment would influence the 19F relaxivity of the Gd multi-core NPs.
Under premixing conditions (Fig. 6), the 19F signal of Gd multi-core NPs was initially low, similar to the baseline signal of the NPs in medium. However, following overnight incubation, the 19F signal increased substantially, indicating responsiveness to the mildly acidic intracellular environment. In contrast, the control NPs exhibited consistently high signal intensity both before and after incubation, with no significant increase over time, indicating no pH responsiveness (Fig. 6). T2 values (Table S7) further support these observations, showing an increase for Gd multi-core NPs from ∼30 ms to ∼200 ms after incubation, whereas control NPs remained stable (∼750 ms). The fact that the T2 of Gd multi-core NPs remains at 208 ms, significantly lower than that of control NPs (Table S7), suggests that the 19F nuclei still experience the PRE effect from Gd. This indicates that Gd has not been entirely released, and the particles remain structurally intact, rather than undergoing complete degradation.
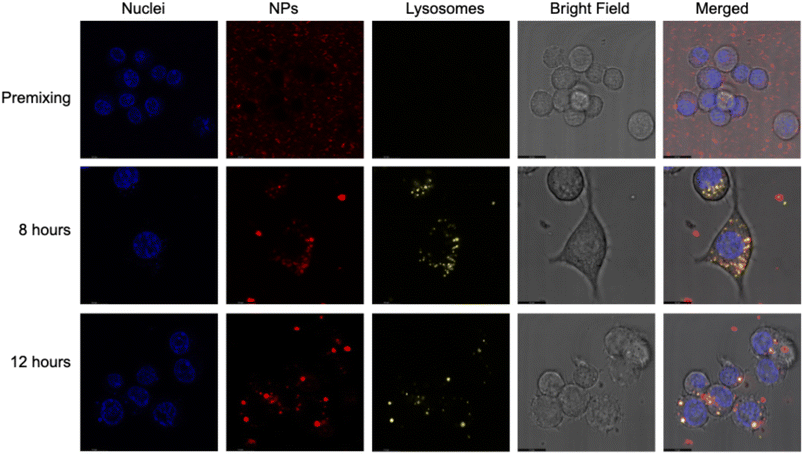
Confocal microscopy confirmed intracellular localization, showing that Gd multi-core NPs remained outside the cells under premixing conditions but colocalized with lysosomes after 8 hours and overnight incubation (12 hours) (Fig. 7). The increase in the 19F NMR signal after overnight incubation aligns with this lysosomal localization, suggesting that the mildly acidic environment enhances the pH responsiveness of Gd multi-core NPs under physiological conditions.
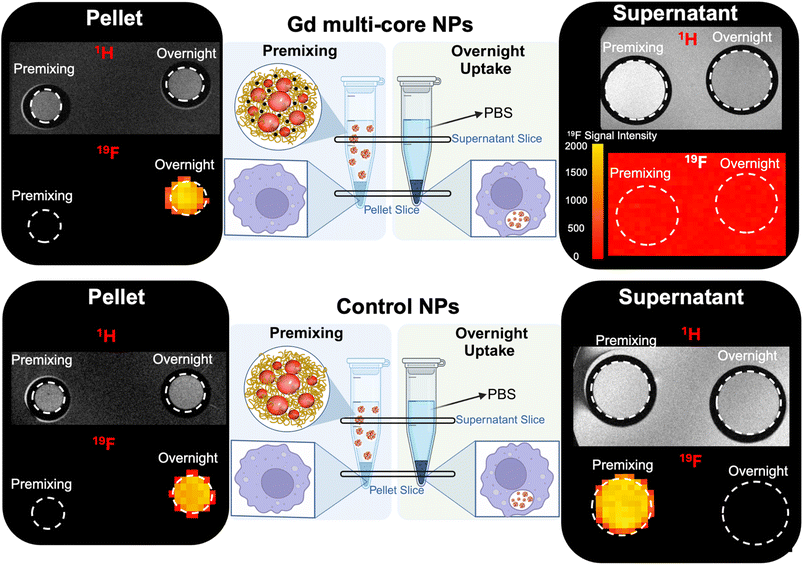
The trends observed in the NMR studies were further corroborated by in vitro19F and 1H MRI (14 T) of RAW 264.7 macrophages incubated with Gd multi-core or control NPs (Fig. 8). Cells were incubated under the same conditions described above for the in-cell NMR, namely, “premixing”, which mimics extracellular NP presence at neutral pH, and “overnight incubation”, which allows for intracellular processing in mildly acidic compartments. After incubation, cells were fixed, resuspended in PBS, and imaged. Both the pellet and the supernatant were imaged to distinguish signal contributions from internalized versus extracellular particles (Fig. 8).
In the supernatant of the cells incubated with Gd multi-core NPs, no 19F MR signal was detected after overnight incubation, as uninternalized NPs were washed away. Similarly, under the premixing conditions, the 19F signal remained undetectable, consistent with the previously observed “off” state due to the PRE effect of Gd (Fig. 8 and S4). However, under premixing condition, an increased 1H signal confirmed the presence of Gd multi-core NPs in the supernatant, indicating Gd-driven contrast enhancement (Fig. 8). This behaviour differs from that observed in aqueous suspension (Fig. 2), where 1H signal enhancement occurred only at pH 3, likely due to polymer swelling induced by carboxyl group protonation. In contrast, the enhanced 1H signal in PBS at pH 7 suggests that ionic interactions with salts in the buffer may also contribute to mild NP swelling. We hypothesize that these interactions disrupt polymer packing and increase hydration, allowing greater water permeability around the Gd chelates and enhancing relaxivity. However, this effect alone was insufficient to mitigate the PRE effect and restore the 19F MR signal. This indicates that while some structural rearrangement may occur in PBS, it does not significantly alter the Gd–PFCE proximity required to relieve PRE suppression.
In the cell pellet, no 19F MR signal was observed under the premixing conditions, consistent with the NPs remaining extracellular in PBS (pH 7), where the PRE effect continues to suppress the 19F signal. After overnight incubation, however, the 19F signal became clearly detectable in the pellet. This signal recovery suggests that intracellular uptake and subsequent trafficking into mildly acidic compartments, such as late endosomes or lysosomes, alleviated the PRE effect, likely through ultrastructural rearrangement that increased the average Gd–PFCE distance or altered PFCE mobility.
For the control NPs, which lack Gd, a clear 19F MR signal was observed in the supernatant under premixing condition and remained detectable in the pellet after overnight incubation (Fig. 8), with no evidence of signal modulation. This confirms that these NPs are not pH-responsive and that their signal remains stable over time regardless of location.
Interestingly, these findings reveal an additional advantage of Gd multi-core NPs for 19F MRI-based cell tracking.66 In ex vivo cell labelling workflows, it is often difficult to distinguish between truly internalized NPs and extracellular surface-bound particles that persist despite washing, potentially confounding in vivo imaging readouts. In our system, only particles that are internalized and processed within the mildly acidic intracellular environment undergo structural changes sufficient to restore the 19F signal. In contrast, extracellular particles remain undetectable by 19F MRI due to the ongoing PRE effect. This selective signal activation improves labelling specificity and offers a promising strategy to enhance the reliability of 19F MRI for tracking labelled cells in vivo.
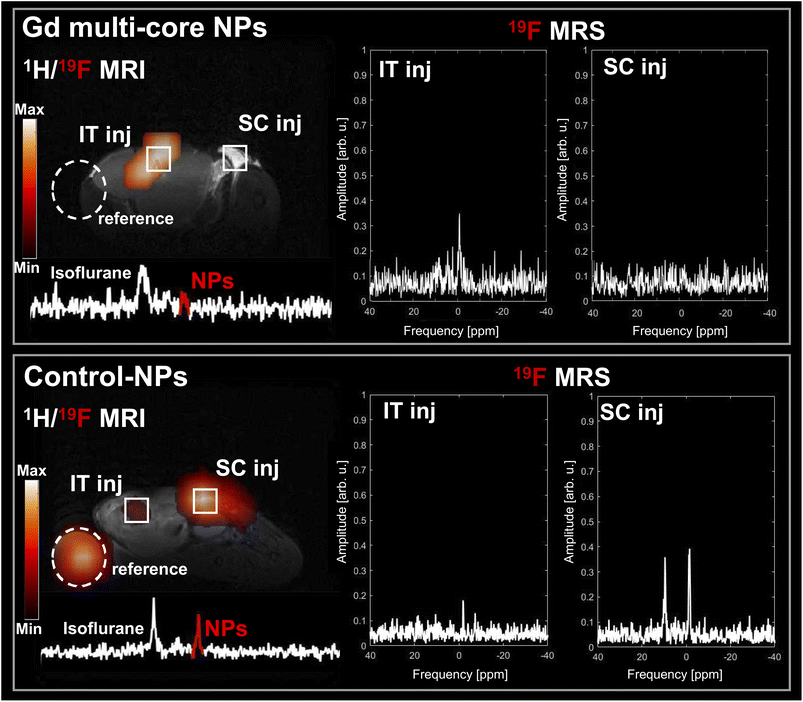
Building on the strong in vitro pH response of Gd multi-core NPs, their behaviour was further evaluated in an in vivo tumour model as a proof of concept. As shown in Fig. 9, the Gd multi-core NPs produced a strong 19F MR signal within the tumour, whereas no detectable signal was observed in the reference tube or at the subcutaneous injection site, both mimicking the pH 7 conditions previously studied in vitro. The presence of the 19F MR signal within the tumour was further confirmed by ex vivo imaging after tumour extraction (Fig. S5), demonstrating the responsiveness of Gd multi-core NPs to the acidic tumour microenvironment. In contrast, control NPs remained clearly visible by 19F MR in both the subcutaneous injection and reference tube, while the 19F MR signal in the tumour was notably lower. This may be due to differences in NP distribution or local microenvironmental factors affecting MR signal decay. These findings highlight the potential of Gd multi-core NPs as a smart, pH-responsive probe capable of selectively activating 19F MRI contrast in acidic environments, making them advantageous for precision imaging and diagnosis in tumour microenvironments and other conditions where pH variations are critical.
Collectively, our results indicate that the pH responsiveness is not driven by Gd release, PLGA degradation, or specific ionic effects. At pH 3, reduced 1H T1 values (Table S2) suggest matrix swelling and increased permeability. However, at more physiological acidity (pH 5), 1H T1 remains comparable to pH 7 despite clear 19F signal enhancement, indicating that matrix softening alone is insufficient to explain the effect. Minor factors, such as localized PFCE mobility and other physicochemical changes, may contribute synergistically. Still, among all tested variables, internal ultrastructural rearrangement remains the most plausible and dominant mechanism driving the pH-dependent 19F MRI signal modulation.
Additionally, multi-core PFCE–PLGA NPs have previously demonstrated excellent biocompatibility, with no toxicity observed even at high doses, further supporting their suitability for clinical translation.52 Interestingly, previous studies with PFCE–PLGA NPs incorporating hydrophilic gadoteridol showed enhancement of the 1H MR signal, but no significant modulation of the 19F signal.50
Although our multi-core design leads to a lower absolute 19F signal and shorter T2 values (Fig. 2a, b and 3), it represents a rational design compromise, prioritizing stimulus-responsive behavior over maximal signal intensity. This approach is particularly well-suited for applications requiring conditional signal activation, such as imaging of pH-sensitive pathological environments.
This distinction is important: if the primary goal is to enhance the 19F signal under all conditions, other paramagnetic centers such as Fe3+ or Mn2+, which preferentially shorten T1 with minimal impact on T2, may be more appropriate.35,54 In contrast, our Gd-based multi-core system is designed to enable 19F ON/OFF switching. Here, effective modulation of the 19F signal depends not only on the presence of Gd chelates but also critically on maintaining a short Gd–19F distance, as achieved within the multi-core nanoparticle structure.
While the present in vivo experiments demonstrate proof-of-concept pH-responsive 19F MRI signal activation, intratumoral and subcutaneous administration do not fully reflect clinically relevant diagnostic scenarios, where intravenous delivery is currently the predominant route. Following systemic administration, dilution, clearance, and reduced local nanoparticle concentration are expected to further limit signal-to-noise ratio and detection sensitivity. In this context, the relatively low absolute 19F signal and short T2 values represent a limitation for whole-body diagnostic imaging. Nevertheless, by prioritizing conditional signal activation over maximal signal intensity, this platform may be particularly suited for applications involving localized accumulation, targeted delivery, or image-guided administration.
3 Conclusion
This study presents a novel pH-sensitive 19F MRI nanosystem in which NP ultrastructure modulates signal activation via the PRE effect, without requiring fluorophilic chelators or molecular linkers. By co-encapsulating PFCE and Gd-chelates within a PLGA matrix, we showed that multi-core NPs exhibit strong pH-responsive behaviour, enhancing 19F MR signal intensity under acidic conditions. In contrast, single-core NPs, representing the conventional structure for polymer-encapsulated PFCs, remained unaffected by pH changes. These findings were validated across high-field (14 T) and low-field (3 T) MRI platforms, as well as in vitro and in vivo, where tumour acidity selectively activated the 19F MR signal.Notably, multi-core NPs exhibited reversible pH-dependent 19F signal enhancement under controlled in situ NMR conditions, a phenomenon not previously described in the literature. This reversibility supports the hypothesis that the observed effect is primarily driven by internal ultrastructural rearrangement, rather than by polymer degradation or Gd release, thereby preserving nanoparticle integrity. In contrast, single-core NPs remained pH-insensitive, emphasizing the role of nanoscale organization in modulating PRE-based contrast.
Given their high fluorine content, biocompatibility, and tunable response, Gd multi-core NPs offer a promising platform for stimulus-responsive MRI probes, with potential applications in tumour imaging and other pH-sensitive disease conditions. Their improved safety profile is further supported by the use of a biodegradable polymer matrix and chelated Gd, helping to mitigate concerns regarding free Gd toxicity. Moreover, the ability of these NPs to selectively activate the 19F signal only after cellular internalization provides an additional advantage for accurate and reliable 19F MRI-based cell tracking. More broadly, this work demonstrates that the nanoparticle architecture can be leveraged as a switchable contrast mechanism, offering a promising strategy for adaptive diagnostic imaging and stimulus-responsive theranostic applications.
4 Materials and methods
4.1 Materials
All chemicals were used as received. Poly(D,L-lactide-co-glycolide) (PLGA) Resomer RG 502H, with a lactide![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) glycolide molar ratio of 50
glycolide molar ratio of 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50, was obtained from Evonik Industries AG, Essen, Germany. Polyvinyl alcohol (PVA, 9000–10
50, was obtained from Evonik Industries AG, Essen, Germany. Polyvinyl alcohol (PVA, 9000–10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 Mw, 80% hydrolyzed), dichloromethane (DCM), deuterium oxide (D2O), trifluoroacetic acid (TFA), gadolinium chloride (GdCl3), and 3-aminopropyl-triethoxysilane (APTES) were purchased from Sigma-Aldrich (St. Louis, MO, USA). Perfluoro-15-crown-5 ether (PFCE) was obtained from Exfluor (Texas, USA). Water was purified using a Purelab Chorus water purification system from Elga (Milli-Q water). Lipophilic gadolinium (Gd) chelate was synthesized as described previously.67 Phosphate-buffered saline (PBS), Dulbecco's Modified Eagle Medium (DMEM), fetal bovine serum (FBS), and 1% penicillin/streptomycin were purchased from Gibco, Thermo Fisher Scientific, USA.
000 Mw, 80% hydrolyzed), dichloromethane (DCM), deuterium oxide (D2O), trifluoroacetic acid (TFA), gadolinium chloride (GdCl3), and 3-aminopropyl-triethoxysilane (APTES) were purchased from Sigma-Aldrich (St. Louis, MO, USA). Perfluoro-15-crown-5 ether (PFCE) was obtained from Exfluor (Texas, USA). Water was purified using a Purelab Chorus water purification system from Elga (Milli-Q water). Lipophilic gadolinium (Gd) chelate was synthesized as described previously.67 Phosphate-buffered saline (PBS), Dulbecco's Modified Eagle Medium (DMEM), fetal bovine serum (FBS), and 1% penicillin/streptomycin were purchased from Gibco, Thermo Fisher Scientific, USA.
4.2 Methods
After sonication, the organic solvent was allowed to evaporate overnight, solidifying the particles. The NPs were washed four times with Milli-Q water by centrifugation at 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g (4 °C, 35 minutes) with supernatant replacement after each wash. The NPs were then resuspended in 5 mL of water, frozen using liquid nitrogen, and freeze-dried. The resulting powder was stored at −20 °C. Control NPs were prepared using the same procedure, excluding the addition of the Gd-chelate solution to the organic phase. For confocal microscopy experiments, NPs were prepared using the same procedure, with the addition of a fluorescent dye (AttoOxa12, ATTO-TEC GmbH, Germany) to enable imaging. In this manuscript, NPs containing Gd-chelate will be referred to as Gd multi-core NPs, while those without Gd-chelate will be referred to as control NPs (Fig. 1).
000 × g (4 °C, 35 minutes) with supernatant replacement after each wash. The NPs were then resuspended in 5 mL of water, frozen using liquid nitrogen, and freeze-dried. The resulting powder was stored at −20 °C. Control NPs were prepared using the same procedure, excluding the addition of the Gd-chelate solution to the organic phase. For confocal microscopy experiments, NPs were prepared using the same procedure, with the addition of a fluorescent dye (AttoOxa12, ATTO-TEC GmbH, Germany) to enable imaging. In this manuscript, NPs containing Gd-chelate will be referred to as Gd multi-core NPs, while those without Gd-chelate will be referred to as control NPs (Fig. 1).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 625 g, 1.5 wt%) and sonicated under the same conditions described above (40% amplitude, 3 minutes). Sodium cholate served as a surfactant during the sonication step. The desired amount of Gd-chelate (c = 0.0166 mg µL−1) was added to the organic phase prior to sonication. Following sonication, DCM was evaporated overnight under stirring to solidify the particles. To replace the surfactant, PVA solution (10 g, 2 wt%) was added to the emulsion, which was continuously stirred at 4 °C for 4 days. The single-core NPs were then washed four times with Milli-Q water through centrifugation (16
625 g, 1.5 wt%) and sonicated under the same conditions described above (40% amplitude, 3 minutes). Sodium cholate served as a surfactant during the sonication step. The desired amount of Gd-chelate (c = 0.0166 mg µL−1) was added to the organic phase prior to sonication. Following sonication, DCM was evaporated overnight under stirring to solidify the particles. To replace the surfactant, PVA solution (10 g, 2 wt%) was added to the emulsion, which was continuously stirred at 4 °C for 4 days. The single-core NPs were then washed four times with Milli-Q water through centrifugation (16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 × g, 35 minutes, 4 °C), resuspended in 5 mL of water, frozen with liquid nitrogen, and freeze-dried. The resulting powder was stored at −20 °C. In this manuscript, single-core NPs containing Gd-chelate will be referred to as Gd single-core NPs (Fig. 1).
000 × g, 35 minutes, 4 °C), resuspended in 5 mL of water, frozen with liquid nitrogen, and freeze-dried. The resulting powder was stored at −20 °C. In this manuscript, single-core NPs containing Gd-chelate will be referred to as Gd single-core NPs (Fig. 1).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 volume ratio. The mixture was incubated at 40 °C for two days to ensure complete degradation of the NPs. After degradation, the samples were diluted to a total volume of 10 mL prior to measurement.
1 volume ratio. The mixture was incubated at 40 °C for two days to ensure complete degradation of the NPs. After degradation, the samples were diluted to a total volume of 10 mL prior to measurement.
In the overnight incubation condition, cells were seeded in 6-well plates and incubated for ∼12 hours with Gd multi-core or control NPs at a concentration of 2 mg per 1.0 × 106 cells to allow uptake. After incubation, excess NPs were removed by washing the cells three times with PBS, followed by scraping and resuspension in fresh medium.
In the premixing condition, cells were mixed with the desired NPs immediately before measurement, without allowing time for uptake.
For MRI measurements, the same conditions were used, except that after washing and collecting the cells, cells were fixed with 4% paraformaldehyde (PFA) at room temperature for 30 minutes. Following fixation, cells were washed three additional times, resuspended in PBS, and centrifuged to form a pellet before scanning.
For quantitative 19F NMR spectroscopy, 5 mg of NPs was resuspended in 500 µL of D2O and mixed with 1 vol% TFA (100 µL) as an internal reference (δ = 0.00 MHz). The solution was transferred into a 5 mm NMR tube, and measurements were performed using 8 scans with a 25-second interscan relaxation delay.
For relaxation time measurements, 5 mg of NPs were resuspended in D2O or a D2O-based solvent with the desired pH at the same concentration used for quantitative analysis right prior to the measurement and transferred to an NMR tube. The 19F 90° pulse was calibrated prior to measurement and T1 and T219F relaxation times were measured as described below.
T 1 measurements were conducted using the inversion recovery method, which involves a 180° pulse followed by variable delay times to allow recovery of the longitudinal magnetization, followed by a 90° detection pulse. T1 was initially estimated using a simple 1D sequence, followed by a full pseudo-2D acquisition with delays and an interscan delay set to 5 × T1 (longest). Recovery curves were fit to an exponential growth function to calculate T1. The 19F T1 experiments were performed without 1H decoupling.
T 2 measurements were performed using the Carr–Purcell–Meiboom–Gill (CPMG) sequence, which involves a 90° pulse followed by a train of tau-180-tau sequences, where tau is a variable delay. For rapidly T2-relaxing nuclei (T2 < 30 ms), a tau of 0.5 ms was used and for slower T2-relaxing nuclei, a tau of 1.2 ms was used. T2 was first estimated using a simple 1D sequence, followed by a full pseudo-2D acquisition with a list of refocusing pulse repetitions ranging from 2 to 1750 and an interscan delay set to 5 × T1 (longest). The recovery was fit to a mono-exponential decay function to determine T2 relaxation time.
For in-cell 19F NMR experiments cells were prepared as described above for the premixing and overnight incubation conditions. Following cell labelling, washing, and harvesting, cells were resuspended in 0.5 mL of full medium supplemented with 10% D2O and transferred to a 5 mm Shigemi tube. The inner Shigemi tube was omitted to prevent compression of the cells. Cell viability was assessed using trypan blue staining and quantified with an Invitrogen™ Countess™ 3 (Thermo Fisher Scientific, USA) automated cell counter. All NMR data analysis was performed using Mestrenova 14.3.0 (Mestrelab Research, Barcelona, Spain).
4.2.8.1. 14 T MRI. MR experiments with phantoms were conducted on a Bruker wide-bore 600 MHz Avance NEO spectrometer (Bruker, Ettlingen, Germany) equipped with a Micro 2.5 microimaging gradient system and Paravision 360 imaging software. A MicWB40 microMRI probehead, fitted with a 30 mm diameter dual-tuned 1H/19F coil, was used for all experiments. All 19F sequences were run with a chemical shift offset appropriate for the NP fluorine signal (approximately −91.6 ppm).
Fluorine and hydrogen MRI were performed on (1) Gd single-core NPs, (2) Gd multi-core NPs, and (3) control NPs at a concentration of 6.5 mg mL−1. Phantoms were prepared using 200 µL Eppendorf tubes, each containing NPs at pH 3, 5, or 7, along with a control tube containing water. All four tubes were placed in a 25 mm NMR tube for simultaneous scanning. To evaluate relaxation time changes following cell uptake, cells were treated as described in Section 4.2.6, using the overnight incubation and premixing conditions. T1 and T2 relaxation times were then measured for both conditions in the cell pellets and the supernatant.
T 1 relaxation times were measured using a Flow-Sensitive Alternating Inversion Recovery with Rapid Acquisition with Relaxation Enhancement (FAIRRARE) sequence with the following parameters: echo time (TE) of 4.36 ms, repetition time (TR) of 5000 ms, and inversion times (TI) of 23, 100, 250, 500, 800, 2000, and 4000 ms, yielding seven MR images. The number of averages (NA) was set to 60 for the control NPs and 240 for the Gd NPs. The matrix size was 32 × 32, with a field of view (FOV) of 20 × 20 mm, a slice thickness (ST) of 10 mm, and a RARE factor of 32. Total scan time was 35 minutes for control NPs and 140 minutes for Gd NPs.
T 2 relaxation times were measured using a multi-scan multi-echo (MSME) sequence with the following parameters: TE of 20 ms, echo spacing of 20 ms, and a total of 60 echo images. TR was 5000 ms, with NA set to 30. The matrix size was 32 × 32, with a FOV of 20 × 20 mm and a slice thickness of 10 mm. The total scan time was 80 minutes.
Phantom localization was established using a 1H RARE sequence with the following parameters: TE of 20 ms, TR of 1000 ms, NA of 4, matrix size of 256 × 256, FOV of 20 × 20 mm, slice thickness of 0.4 mm, and a RARE factor of 8. The scan time was approximately 2 minutes. A subsequent 19F RARE sequence was performed using the same orientation as 1H (axial). The parameters for the 19F MR sequence were: TE of 18 ms, TR of 2075 ms, NA of 512, matrix size of 40 × 34, FOV of 20 × 20 mm, slice thickness of 10 mm, and a RARE factor of 34, with a scan time of approximately 18 minutes.
4.2.8.2 3 T MRI. To assess the pH sensitivity at clinically relevant magnetic field strengths, NPs were examined using a 3 T Bruker BioSpec system with ParaVision 360 v.3.3 software (Bruker, Ettlingen, Germany). The same conditions described in Section 4.2.8.1 were used, including the types of NPs, concentration, and phantom preparation.
T 1 relaxation times were measured using an inversion recovery RARE sequence with a TE of 6.5 ms, a TR of 5000 ms, and TIs of 23, 100, 250, 500, 800, 2000, and 4000 ms, producing seven MR images. The NA was set to 60, with a matrix size of 32 × 32, a spatial resolution of 1.5 × 1.5 mm2 and a slice thickness of 5 mm. The RARE factor was set to 32, and the total scan time was 35 minutes.
T 2 relaxation times were measured using a MSME sequence. Parameters for the control samples and the Gd-loaded sample at pH 3 were as follows: TE of 20 ms, echo spacing of 20 ms, and a total of 60 echo images, with a TR of 5000 ms and NA set to 60. The matrix size was 32 × 32, with a spatial resolution of 1.5 × 1.5 mm2 and a slice thickness of 5 mm. For the Gd samples at pH values of 5 and 7, the parameters were adjusted as follows: TE of 6.3 ms, echo spacing of 6.3 ms, and a total of 60 echo images, with a TR of 1800 ms and NA set to 60. The matrix size, spatial resolution and slice thickness remained the same.
Fluorescently labelled multi-core NPs (AttoOxa12) were incubated at 2 mg per 1 × 106 for either 12 or 8 hours prior to imaging. At each endpoint, cells were washed three times with PBS to remove excess NPs, and fresh medium with (or without) LysoTracker™ Green DND-26 (Thermo Fisher Scientific, USA) was added at a concentration of 100 nM for 2 hours. Fifteen minutes before washing, a drop of Hoechst EasyProbe (ABP Biosciences, Beltsville, USA) was added to each well for nuclear staining.
To image the premixing conditions, the particles were added to live cells in situ and immediately imaged. The acquisition settings were consistent between samples, and the same post-processing workflow was applied for the analysis.
4.2.10.1 4T1 cell incubation and tumor induction. The murine mammary carcinoma 4T1 cell line was cultured in Roswell Park Memorial Institute (RPMI) 1640 media (Gibco, Thermo Fisher Scientific) supplemented with 2 mmol per L L-glutamine, 10% FBS, and 1% penicillin–streptomycin. All cells were incubated under standard conditions (37 °C; 5% CO2). To induce tumour growth in vivo, a cell suspension (2 × 106; V = 0.1 mL in PBS) was subcutaneously injected into the right mammary fat pad of healthy Balb/c mice. Tumour growth was monitored daily.
4.2.10.2 NP phantom measurement. The NPs (control and Gd multi-core) were tested prior to the in vivo experiments using a custom-made dual 1H/19F surface radiofrequency (RF) coil on a 7 T MR experimental scanner (Bruker, Ettlingen, Germany). The half-saddle RF coil (30 mm diameter) was tuned and matched to the Larmor frequencies of both 1H and 19F nuclei for optimal performance. Phantom experiments of aqueous control NPs and Gd multi-core NPs (10 mg/100 µL) were performed to establish sensitivity. Phantom localization was achieved using 1H MRI with a T2-weighted turbo spin-echo RARE sequence with the following parameters: TR of 1000 ms, TE of 36 ms, 2 NA, RARE factor of 8, FOV of 20 × 20 mm2, matrix size of 256 × 256 and 0.078 × 0.078 mm2 resolution. 19F MR non-localized spectroscopy (19F MRS) was assessed using a single-pulse sequence with a TR of 1000 ms, a bandwidth (BW) of 41 ppm, and a NA of 1–600 to observe the MR sensitivity at different scan times (1 second to 10 minutes). The chemical offset (−91 ppm) was determined from the phantom measurements. 19F MR images were acquired by MR spectroscopic imaging (MRSI), which enables spectrally resolved signal acquisition from a 2D voxel slab, providing targeted signal separation at a specific resonance frequency. Following parameters were used: TR of 30 ms, BW of 80 ppm, FOV of 20 × 20 mm2, total scan time of 30 seconds to 1 hour (NA of 1–175), slice thickness of 5 mm, matrix size of 64 × 64 and 0.313 × 0.313 mm2 resolution. All phantom images were measured in the coronal plane.
4.2.10.3 Tumour imaging. In vivo 19F MRS measurements were performed using the same RF dual coil and MR sequence parameters on a 7 T MR scanner as in phantom experiments, with a modified BW (50 ppm). The 19F MRSI sequence (TR of 100 ms) was optimized for the animal measurements with a FOV of 35 × 35 mm2, a slice thickness of 25 mm, and a scan time of 10 minutes (NA of 17). Measurements were conducted directly after NP injection (5 mg/100 µL), administered subcutaneously (SC) and intratumorally (IT), at 24 hours post-injection (day 1) and on day 5 (only control NPs). Control NPs and Gd multi-core NPs (2 mg/100 µL) were mixed in 4% gelatine and used as signal references placed next to the animal. Tumour containing Gd-multi-core NPs was extracted at day 1 and measured ex vivo using the same sequences.
Conflicts of interest
There are no conflicts of interest to declare.Data availability
All data supporting the findings of this study are available within the article and its supplementary information (SI). Supplementary information: Fig. S1–S5 and Tables S1–S7 providing additional data supporting the main findings of the manuscript. These include detailed nanoparticle size and polydispersity measurements by DLS, 19F and 1H MR relaxometry data, reversibility experiments confirming pH sensitivity, and MR images of in vitro and ex vivo samples. See DOI: https://doi.org/10.1039/d5na01005e.Acknowledgements
All experiments related to in vivo tumour visualization were funded by the Ministry of Health CR-DRO (Institute for Clinical and Experimental Medicine IKEM, project no. IN00023001). This project was also supported by the ZonMW PSIDER consortium – TRACER (Grant No. 0250022110001) and the EU Horizon 2020 Programme – Phoenix (Grant No. 953110). We sincerely thank Niels Wever from the ANU Department of Animal Nutrition for performing the ICP-OES analysis. We acknowledge the uNMR-NL Grid, a distributed state-of-the-art Magnetic Resonance facility for the Netherlands (NWO grant 184.035.002), and the MAGNEtic resonance research FacilitY (MAGNEFY) at Wageningen University for providing access to the NMR instruments. We are also grateful to Dominik Havlicek and Andrea Galisova from IKEM for their invaluable assistance with the in vivo MRI experiments. We would also like to thank Marcia Arini Ardyanti for her contribution to the preparation of the graphical abstract.References
- B. R. Smith and S. S. Gambhir, Nanomaterials for In Vivo Imaging, Chem. Rev., 2017, 117(3), 901–986 Search PubMed.
- O. Maxouri, Z. Bodalal, M. Daal, S. Rostami, I. Rodriguez, L. Akkari, M. Srinivas, R. Bernards and R. Beets-Tan, How to 19F MRI: applications, technique, and getting started, BJR|Open, 2023, 5(1), 20230019 CrossRef PubMed.
- N. MacRitchie, M. Frleta-Gilchrist, A. Sugiyama, T. Lawton, I. B. McInnes and P. Maffia, Molecular imaging of inflammation - Current and emerging technologies for diagnosis and treatment, Pharmacol. Ther., 2020, 211, 107550 Search PubMed.
- T.-H. Shin, Y. Choi, S. Kim and J. Cheon, Recent advances in magnetic nanoparticle-based multi-modal imaging, Chem. Soc. Rev., 2015, 44(14), 4501–4516 RSC.
- E. Di Gregorio, G. Ferrauto, S. Lanzardo, E. Gianolio and S. Aime, Use of FCC-NMRD relaxometry for early detection and characterization of ex-vivo murine breast cancer, Sci. Rep., 2019, 9(1), 4624 CrossRef PubMed.
- X. Zhu, X. Tang, H. Lin, S. Shi, H. Xiong, Q. Zhou, A. Li, Q. Wang, X. Chen and J. Gao, A Fluorinated Ionic Liquid-Based Activatable 19F MRI Platform Detects Biological Targets, Chem, 2020, 6(5), 1134–1148 CAS.
- L. O. Poku, M. Phil, Y. Cheng, K. Wang and X. Sun, 23Na-MRI as a Noninvasive Biomarker for Cancer Diagnosis and Prognosis, J. Magn. Reson. Imaging, 2021, 53(4), 995–1014 CrossRef PubMed.
- J. Ruiz-Cabello, B. P. Barnett, P. A. Bottomley and J. W. M. Bulte, Fluorine (19F) MRS and MRI in biomedicine, NMR Biomed., 2011, 24(2), 114–129 CrossRef CAS PubMed.
- J. Chen, P. Pal and T. Ahrens Eric, Systems Engineering Approach Towards Sensitive Cellular Fluorine-19 MRI, NMR Biomed., 2025, 38(1), e5298 Search PubMed.
- O. Koshkina, T. Rheinberger, V. Flocke, A. Windfelder, P. Bouvain, N. M. Hamelmann, J. M. J. Paulusse, H. Gojzewski, U. Flögel and F. R. Wurm, Biodegradable polyphosphoester micelles act as both background-free 31P magnetic resonance imaging agents and drug nanocarriers, Nat. Commun., 2023, 14(1), 4351 CrossRef CAS PubMed.
- N. Jirát-Ziółkowska, K. Sulková, L. Kracíková, L. Androvič, D. Havliček, R. Laga and D. Jirák, Bio-responsive polymers for dual 31P/19F-magnetic resonance to detect reactive oxygen species in vivo, Commun. Mater., 2024, 5(1), 178 CrossRef.
- J. Chen, G. M. Lanza and S. A. Wickline, Quantitative magnetic resonance fluorine imaging: today and tomorrow, Wiley Interdiscip. Rev.:Nanomed. Nanobiotechnol., 2010, 2(4), 431–440 CAS.
- J. Wahsner, E. M. Gale, A. Rodríguez-Rodríguez and P. Caravan, Chemistry of MRI Contrast Agents: Current Challenges and New Frontiers, Chem. Rev., 2019, 119(2), 957–1057 CrossRef CAS PubMed.
- M. Srinivas, E. H. J. G. Aarntzen, J. W. M. Bulte, W. J. Oyen, A. Heerschap, I. J. M. de Vries and C. G. Figdor, Imaging of cellular therapies, Adv. Drug Delivery Rev., 2010, 62(11), 1080–1093 CrossRef CAS PubMed.
- M. Srinivas, A. Heerschap, E. T. Ahrens, C. G. Figdor and I. J. de Vries, (19)F MRI for quantitative in vivo cell tracking, Trends Biotechnol., 2010, 28(7), 363–370 Search PubMed.
- T. Boissenot, E. Fattal, A. Bordat, S. Houvenagel, J. Valette, H. Chacun, C. Gueutin and N. Tsapis, Paclitaxel-loaded PEGylated nanocapsules of perfluorooctyl bromide as theranostic agents, Eur. J. Pharm. Biopharm., 2016, 108, 136–144 Search PubMed.
- T. A. Tunca Arın, D. Havlíček, D. F. Dorado Daza, N. Jirát-Ziółkowska, O. Pop-Georgievski, D. Jirák and O. Sedlacek, Water-soluble fluorinated copolymers as highly sensitive 19F MRI tracers: From structure optimization to multimodal tumor imaging, Mater. Today Bio, 2025, 31, 101462 CrossRef PubMed.
- A. Mali, E. L. Kaijzel, H. J. Lamb and L. J. Cruz, 19F-nanoparticles: Platform for in vivo delivery of fluorinated biomaterials for 19F-MRI, J. Controlled Release, 2021, 338, 870–889 Search PubMed.
- E. T. Ahrens, B. M. Helfer, C. F. O'Hanlon and C. Schirda, Clinical cell therapy imaging using a perfluorocarbon tracer and fluorine-19 MRI, Magn. Reson. Med., 2014, 72(6), 1696–1701 CrossRef CAS PubMed.
- E. T. Ahrens, B. M. Helfer, C. F. O’Hanlon, D. R. Lister, J. L. Bykowski, K. Messer, B. I. Leach, J. Chen, H. Xu, G. A. Daniels and E. E. W. Cohen, Method for estimation of apoptotic cell fraction of cytotherapy using in vivo fluorine-19 magnetic resonance: pilot study in a patient with head and neck carcinoma receiving tumor-infiltrating lymphocytes labeled with perfluorocarbon nanoemulsion, J. Immunother. Cancer, 2023, 11(6), e007015 CrossRef PubMed.
- Y. Takaoka, T. Sakamoto, S. Tsukiji, M. Narazaki, T. Matsuda, H. Tochio, M. Shirakawa and I. Hamachi, Self-assembling nanoprobes that display off/on 19F nuclear magnetic resonance signals for protein detection and imaging, Nat. Chem., 2009, 1(7), 557–561 Search PubMed.
- D. Janasik, K. Jasiński, W. P. Węglarz, I. Nemec, P. Jewula and T. Krawczyk, Ratiometric pH-Responsive 19F Magnetic Resonance Imaging Contrast Agents Based on Hydrazone Switches, Anal. Chem., 2022, 94(8), 3427–3431 CrossRef CAS PubMed.
- J.-X. Yu, V. D. Kodibagkar, R. R. Hallac, L. Liu and R. P. Mason, Dual 19F/1H MR Gene Reporter Molecules for in Vivo Detection of β-Galactosidase, Bioconjugate Chem., 2012, 23(3), 596–603 CrossRef CAS PubMed.
- X. Zhu, P. Zhang, D. Liu, L. Tao, J. Du and X. Gao, Stimuli-responsive 19F MRI probes: From materials design to in vitro detection and in vivo diagnosis, TrAC, Trends Anal. Chem., 2024, 172, 117607 CrossRef CAS.
- F. Liu, X. Li, Y. Li, S. Xu, C. Guo and L. Wang, Visualization of drug release in a chemo-immunotherapy nanoplatform via ratiometric 19F magnetic resonance imaging, Chem. Sci., 2024, 15(42), 17397–17406 RSC.
- T. Nakamura, H. Matsushita, F. Sugihara, Y. Yoshioka, S. Mizukami and K. Kikuchi, Activatable 19F MRI Nanoparticle Probes for the Detection of Reducing Environments, Angew. Chem., Int. Ed., 2015, 54(3), 1007–1010 CrossRef CAS PubMed.
- S. Mizukami, R. Takikawa, F. Sugihara, Y. Hori, H. Tochio, M. Wälchli, M. Shirakawa and K. Kikuchi, Paramagnetic Relaxation-Based 19F MRI Probe To Detect Protease Activity, J. Am. Chem. Soc., 2008, 130(3), 794–795 CrossRef CAS PubMed.
- K. Akazawa, F. Sugihara, T. Nakamura, S. Mizukami and K. Kikuchi, Highly Sensitive Detection of Caspase-3/7 Activity in Living Mice Using Enzyme-Responsive 19F MRI Nanoprobes, Bioconjugate Chem., 2018, 29(5), 1720–1728 CrossRef CAS PubMed.
- X. Yue, Z. Wang, L. Zhu, Y. Wang, C. Qian, Y. Ma, D. O. Kiesewetter, G. Niu and X. Chen, Novel 19F Activatable Probe for the Detection of Matrix Metalloprotease-2 Activity by MRI/MRS, Mol. Pharm., 2014, 11(11), 4208–4217 CrossRef CAS PubMed.
- Y. Yuan, H. Sun, S. Ge, M. Wang, H. Zhao, L. Wang, L. An, J. Zhang, H. Zhang and B. Hu, et al., Controlled Intracellular Self-Assembly and Disassembly of 19F Nanoparticles for MR Imaging of Caspase 3/7 in Zebrafish, ACS Nano, 2015, 9(1), 761–768 CrossRef CAS PubMed.
- P. Huang, W. Guo, G. Yang, H. Song, Y. Wang, C. Wang, D. Kong and W. Wang, Fluorine Meets Amine: Reducing Microenvironment-Induced Amino-Activatable Nanoprobes for 19F-Magnetic Resonance Imaging of Biothiols, ACS Appl. Mater. Interfaces, 2018, 10(22), 18532–18542 CrossRef CAS PubMed.
- G. M. Clore and J. Iwahara, Theory, Practice, and Applications of Paramagnetic Relaxation Enhancement for the Characterization of Transient Low-Population States of Biological Macromolecules and Their Complexes, Chem. Rev., 2009, 109(9), 4108–4139 CrossRef CAS PubMed.
- J. Iwahara and G. M. Clore, Detecting transient intermediates in macromolecular binding by paramagnetic NMR, Nature, 2006, 440(7088), 1227–1230 CrossRef CAS PubMed.
- D. Xie, M. Yu, R. T. Kadakia and E. L. Que, 19F Magnetic Resonance Activity-Based Sensing Using Paramagnetic Metals, Acc. Chem. Res., 2020, 53(1), 2–10 CrossRef CAS PubMed.
- Y. Mo, C. Huang, C. Liu, Z. Duan, J. Liu and D. Wu, Recent Research Progress of 19F Magnetic Resonance Imaging Probes: Principle, Design, and Their Application, Macromol. Rapid Commun., 2023, 44(16), 2200744 CrossRef CAS PubMed.
- J. Zheng, X. Fang, L. Li, R. Zhang and C. Li, Biomolecule-responsive nanoprobes for living cell analysis, TrAC, Trends Anal. Chem., 2023, 169, 117387 CrossRef CAS.
- D. J. Cooke, E. Y. Maier, T. L. King, H. Lin, S. Hendrichs, S. Lee, N. N. Mafy, K. M. Scott, Y. Lu and E. L. Que, Dual Nanoparticle Conjugates for Highly Sensitive and Versatile Sensing Using 19F Magnetic Resonance Imaging, Angew. Chem., Int. Ed., 2024, 63(4), e202312322 CrossRef CAS PubMed.
- S. Hajjar and X. Zhou, pH sensing at the intersection of tissue homeostasis and inflammation, Trends Immunol., 2023, 44(10), 807–825 Search PubMed.
- K. Rajamäki, T. Nordström, K. Nurmi, K. E. O. Åkerman, P. T. Kovanen, K. Öörni and K. K. Eklund, Extracellular Acidosis Is a Novel Danger Signal Alerting Innate Immunity via the NLRP3 Inflammasome, J. Biol. Chem., 2013, 288(19), 13410–13419 Search PubMed.
- X. Zhang, Y. Lin and R. J. Gillies, Tumor pH and its measurement, J. Nucl. Med., 2010, 51(8), 1167–1170 Search PubMed.
- M. H. Cho, S. H. Shin, S. H. Park, D. K. Kadayakkara, D. Kim and Y. Choi, Targeted, Stimuli-Responsive, and Theranostic 19F Magnetic Resonance Imaging Probes, Bioconjugate Chem., 2019, 30(10), 2502–2518 Search PubMed.
- M. Oishi, S. Sumitani and Y. Nagasaki, On−Off Regulation of 19F Magnetic Resonance Signals Based on pH-Sensitive PEGylated Nanogels for Potential Tumor-Specific Smart 19F MRI Probes, Bioconjugate Chem., 2007, 18(5), 1379–1382 Search PubMed.
- X. Huang, G. Huang, S. Zhang, K. Sagiyama, O. Togao, X. Ma, Y. Wang, Y. Li, T. C. Soesbe and B. D. Sumer, et al., Multi-Chromatic pH-Activatable 19F-MRI Nanoprobes with Binary ON/OFF pH Transitions and Chemical-Shift Barcodes, Angew. Chem., Int. Ed., 2013, 52(31), 8074–8078 CrossRef CAS PubMed.
- M. Zalewski, D. Janasik, A. Kapała, M. Minoshima, F. Sugihara, W. Raj, J. Pietrasik, K. Kikuchi and T. Krawczyk, pH-Sensitive Polymethacrylates as Potential Contrast Agents in 19F MRI, Macromol. Chem. Phys., 2022, 223(14), 2200027 CrossRef CAS.
- C. Guo, S. Xu, A. Arshad and L. Wang, A pH-responsive nanoprobe for turn-on 19F-magnetic resonance imaging, Chem. Commun., 2018, 54(70), 9853–9856 Search PubMed.
- Q. Wang, Y. Yu, Y. Chang, X. Xu, M. Wu, G. R. Ediriweera, H. Peng, X. Zhen, X. Jiang and D. J. Searles, et al., Fluoropolymer-MOF Hybrids with Switchable Hydrophilicity for 19F MRI-Monitored Cancer Therapy, ACS Nano, 2023, 17(9), 8483–8498 CrossRef CAS PubMed.
- N. Jirát-Ziółkowska, M. Vít, O. Groborz, K. Kolouchová, D. Červený, O. Sedláček and D. Jirák, Long-term in vivo dissolution of thermo- and pH-responsive, 19F magnetic resonance-traceable and injectable polymer implants, Nanoscale Adv., 2024, 6(12), 3041–3051 Search PubMed.
- K. Wang, H. Peng, K. J. Thurecht, S. Puttick and A. K. Whittaker, Biodegradable core crosslinked star polymer nanoparticles as 19F MRI contrast agents for selective imaging, Polym. Chem., 2014, 5(5), 1760–1771 RSC.
- T. A. Tunca Arın and O. Sedlacek, Stimuli-Responsive Polymers for Advanced 19F Magnetic Resonance Imaging: From Chemical Design to Biomedical Applications, Biomacromolecules, 2024, 25(9), 5630–5649 Search PubMed.
- O. Koshkina, G. Lajoinie, F. B. Bombelli, E. Swider, L. J. Cruz, P. B. White, R. Schweins, Y. Dolen, E. A. W. van Dinther and N. K. van Riessen, et al., Multicore Liquid Perfluorocarbon-Loaded Multimodal Nanoparticles for Stable Ultrasound and 19F MRI Applied to In Vivo Cell Tracking, Adv. Funct. Mater., 2019, 29(19), 1806485 CrossRef PubMed.
- A. Mali, M. Verbeelen, P. B. White, A. H. J. Staal, N. K. van Riessen, C. Cadiou, F. Chuburu, O. Koshkina and M. Srinivas, The internal structure of gadolinium and perfluorocarbon-loaded polymer nanoparticles affects (19)F MRI relaxation times, Nanoscale, 2023, 15(44), 18068–18079 RSC.
- A. H. J. Staal, K. Becker, O. Tagit, N. Koen van Riessen, O. Koshkina, A. Veltien, P. Bouvain, K. R. G. Cortenbach, T. Scheenen and U. Flögel, et al., In vivo clearance of 19F MRI imaging nanocarriers is strongly influenced by nanoparticle ultrastructure, Biomaterials, 2020, 261, 120307 Search PubMed.
- C. Wang, S. R. Adams, H. Xu, W. Zhu and E. T. Ahrens, β-Diketonate-Iron(III) Complex: A Versatile Fluorine-19 MRI Signal Enhancement Agent, ACS Appl. Bio Mater., 2019, 2(9), 3836–3842 Search PubMed.
- A. A. Kislukhin, H. Xu, S. R. Adams, K. H. Narsinh, R. Y. Tsien and E. T. Ahrens, Paramagnetic fluorinated nanoemulsions for sensitive cellular fluorine-19 magnetic resonance imaging, Nat. Mater., 2016, 15(6), 662–668 Search PubMed.
- N. Larreina Vicente, M. Srinivas and O. Tagit, Perfluorocarbon-Loaded Poly(lactide-co-glycolide) Nanoparticles from Core to Crust: Multifaceted Impact of Surfactant on Particle Ultrastructure, Stiffness, and Cell Uptake, ACS Appl. Polym. Mater., 2025, 7(5), 2864–2878 Search PubMed.
- I. Tirotta, V. Dichiarante, C. Pigliacelli, G. Cavallo, G. Terraneo, F. B. Bombelli, P. Metrangolo and G. Resnati, 19F Magnetic Resonance Imaging (MRI): From Design of Materials to Clinical Applications, Chem. Rev., 2015, 115(2), 1106–1129 Search PubMed.
- M. Cametti, B. Crousse, P. Metrangolo, R. Milani and G. Resnati, The fluorous effect in biomolecular applications, Chem. Soc. Rev., 2012, 41(1), 31–42 Search PubMed.
- P. Bouvain, S. Temme and U. Flögel, Hot spot 19F magnetic resonance imaging of inflammation, WIREs Nanomedicine and Nanobiotechnology, 2020, 12(6), e1639 CrossRef CAS PubMed.
- M. Srinivas, L. J. Cruz, F. Bonetto, A. Heerschap, C. G. Figdor and I. J. de Vries, Customizable, multi-functional fluorocarbon nanoparticles for quantitative in vivo imaging using 19F MRI and optical imaging, Biomaterials, 2010, 31(27), 7070–7077 CrossRef CAS PubMed.
- G. M. Clore, Practical Aspects of Paramagnetic Relaxation Enhancement in Biological Macromolecules, Methods Enzymol., 2015, 564, 485–497 CAS.
- A. de Vries, R. Moonen, M. Yildirim, S. Langereis, R. Lamerichs, J. A. Pikkemaat, S. Baroni, E. Terreno, K. Nicolay, G. J. Strijkers and H. Grüll, Relaxometric studies of gadolinium-functionalized perfluorocarbon nanoparticles for MR imaging, Contrast Media Mol. Imaging, 2014, 9(1), 83–91 Search PubMed.
- A. M. Neubauer, J. Myerson, S. D. Caruthers, F. D. Hockett, P. M. Winter, J. Chen, P. J. Gaffney, J. D. Robertson, G. M. Lanza and S. A. Wickline, Gadolinium-modulated 19F signals from perfluorocarbon nanoparticles as a new strategy for molecular imaging, Magn. Reson. Med., 2008, 60(5), 1066–1072 CrossRef CAS PubMed.
- F. Danhier, E. Ansorena, J. M. Silva, R. Coco, A. Le Breton and V. Préat, PLGA-based nanoparticles: an overview of biomedical applications, J. Controlled Release, 2012, 161(2), 505–522 CrossRef CAS PubMed.
- B. Yameen, W. I. Choi, C. Vilos, A. Swami, J. Shi and O. C. Farokhzad, Insight into nanoparticle cellular uptake and intracellular targeting, J. Controlled Release, 2014, 190, 485–499 CrossRef CAS PubMed.
- Z.-G. Yue, W. Wei, P.-P. Lv, H. Yue, L.-Y. Wang, Z.-G. Su and G.-H. Ma, Surface Charge Affects Cellular Uptake and Intracellular Trafficking of Chitosan-Based Nanoparticles, Biomacromolecules, 2011, 12(7), 2440–2446 CrossRef CAS PubMed.
- E. T. Ahrens, B. M. Helfer, C. F. O'Hanlon, D. R. Lister, J. L. Bykowski, K. Messer, B. I. Leach, J. Chen, H. Xu, G. A. Daniels and E. E. W. Cohen, Method for estimation of apoptotic cell fraction of cytotherapy using in vivo fluorine-19 magnetic resonance: pilot study in a patient with head and neck carcinoma receiving tumor-infiltrating lymphocytes labeled with perfluorocarbon nanoemulsion, J. Immunother. Cancer, 2023, 11(6), e007015 CrossRef PubMed.
- G. Rigaux, V. G. Roullin, C. Cadiou, C. Portefaix, L. Van Gulick, G. Bœuf, M. C. Andry, C. Hoeffel, L. Vander Elst and S. Laurent, et al., A new magnetic resonance imaging contrast agent loaded into poly(lacide-co-glycolide) nanoparticles for long-term detection of tumors, Nanotechnology, 2014, 25(44), 445103 CrossRef CAS PubMed.
- F. S. Ruggeri, S. Vieweg, U. Cendrowska, G. Longo, A. Chiki, H. A. Lashuel and G. Dietler, Nanoscale studies link amyloid maturity with polyglutamine diseases onset, Sci. Rep., 2016, 6(1), 31155 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |