Open Access Article
Open Access ArticleAn enzymatic microfluidic sensor for β-lactam antibiotics based on cross-linked ancestral β-lactamase crystals
Isaac
Rodríguez-Ruiz
 a,
Pablo
Moya-Garrido
b,
Valeria A.
Risso
cd,
Sergio
Martínez-Rodríguez
a,
Pablo
Moya-Garrido
b,
Valeria A.
Risso
cd,
Sergio
Martínez-Rodríguez
 def,
Sébastien
Teychené
a,
José Manuel
Sanchez-Ruiz
cd and
José A.
Gavira
def,
Sébastien
Teychené
a,
José Manuel
Sanchez-Ruiz
cd and
José A.
Gavira
 *bdf
*bdf
aLaboratoire de Génie Chimique, CNRS, INP, UPS, Toulouse, France
bLaboratory of Crystallographic Studies, IACT-CSIC, Avd. de las Palmeras, 4, 18100-Armilla, Granada, Spain. E-mail: j.gavira _ad_ csic.es
cDepartamento de Quimica Fisica, Facultad de Ciencias, Universidad de Granada, 18071 Granada, Spain
dUnidad de Excelencia de Química Aplicada a Biomedicina y Medioambiente (UEQ), Universidad de Granada, 18071 Granada, Spain
eDepartment of Biochemistry and Molecular Biology III and Immunology, University of Granada, 18071 Granada, Spain
fRaw Materials, Human and Environmental Health, UGR, Associated Unit of the CSIC by the IACT-CSIC, Avd. de las Palmeras, 4, 18100-Armilla, Granada, Spain
First published on 22nd January 2026
Abstract
A microfluidic enzymatic sensor based on cross-linked crystals of an ancestral β-lactamase (βLa-CLECs) was developed for the detection of β-lactam antibiotics under continuous flow conditions. The sensor is presented in a modular configuration, consisting of a separated microprobe for enzymatic catalysis, allowing for precise enzyme loading, and a photonic lab-on-a-chip (PhLoC) platform for detection by in-line spectrophotometric measurements. The exceptional thermal stability of the reconstructed Precambrian β-lactamase made it suitable for the steps required in device fabrication and carrier-free immobilization treatment, while its inherent promiscuity, i.e., its capability to degrade a variety of lactam antibiotics, including man-made third-generation antibiotics, broadens its potential application. Full catalytic activity of the ancestral enzyme was retained after immobilization, and inhibition by antibiotics such as ampicillin and sulbactam was detected at concentrations as low as parts per billion. These results support the use of ancient enzymes as stable and responsive biorecognition elements in cost-effective, high-throughput analytical systems targeting environmental pollutants and pharmaceutical compounds.
Introduction
Microfluidic systems have emerged as a revolutionary technology with growing importance across a range of scientific disciplines1 and industrial applications.2,3 These miniaturized devices enable the precise manipulation of small liquid volumes within microscopic channels, unlocking a wide array of possibilities in fields as diverse as biotechnology,3 pharmacology, medicine, and analytical chemistry.1 Notable applications include enzyme catalysis assays,4 protein crystallization,5 and the synthesis of molecules and nano- or microparticles.2 The integration of enzyme catalytic reactions into microfluidic systems working under continuous flow conditions represents a promising technology with several strategic benefits.6,7A key advantage of microfluidics is compartmentalization, which facilitates precise control over reactions at the microscale. This greatly enhances surface-to-volume ratios, leading to improved efficiency, reduced energy consumption, lower operational costs and minimal waste generation. Furthermore, it also shortens analysis times. In biotechnology, microfluidic systems have been employed for cell culture, DNA sequencing, protein separation, and high-throughput assays.3 In pharmacology, they have contributed significantly to the early-stage drug screening and the development of more precise drug delivery methods. Microfluidic technology is especially relevant in biosensing applications, where enzymes can serve as pollutant detectors or biorecognition elements. These uses span from immunoassays and clinical diagnoses to the monitoring of cell and microbial cultures, offering higher sensitivity than traditional bulk analysis techniques.
Nevertheless, bridging the gap between microfluidic technology and real-world sensing applications remains a challenge. Analyte detection necessitates the integration of a transducer. Photonic Lab-on-a-Chip platforms (PhLoCs) offer a cost-effective and well-established solution for non-invasive on-chip spectrophotometric analysis8 and have already demonstrated substantial utility in various applications. Beyond biosensing, enzyme catalysis plays a pivotal and dynamic role in industry. Continuous processing is a key goal for reducing costs and maximizing yields. Packed-bed reactors are commonly used for continuous-flow biocatalytic processes, where the substrates pass through a column packed with a heterogeneous catalyst, allowing complete transformation due to controlled residence times.9,10
However, the development of cost-effective and efficient immobilization strategies that stabilize, separate, and enable enzyme reuse (without compromising the catalytic function) remains a thriving area of research. Carrier-free immobilization methods, such as Cross-Linked Enzyme Aggregates (CLEAs), have shown progress in enhancing enzyme stability and robustness, particularly in large-scale operations.11 Similarly, Cross-Linked Enzyme Crystals (CLECs) have already demonstrated exceptional efficiency, especially in chiral ester resolution.12 A further development involves reinforced CLECs (rCLECs), where protein crystals are grown within a gel matrix that becomes incorporated into the crystals. The gel medium permits controlled diffusion of the cross-linker and gives new properties to the composite crystals, such as enhanced stability and tunable dissolution rates.9,13
Several approaches are typically used to enhance/modulate enzyme activity and/or strength: (i) the rational design, including site-directed mutagenesis or (ii) the search for the desired functionality in genome libraries by combinatorial methods i.e. direct evolution.14 A different approach to search for enhanced properties is based on sequence analysis, following the “Consensus Approach”15 and the “Ancestral Protein Reconstruction”.16 The first methodology is one of the simplest computational strategies, which uses statistical information of sequence alignments to generate the so-called “consensus sequence”. The consensus sequence search has occasionally led to variants with enhanced stability, and sometimes, with modulated biological function, while “Ancestral Protein Reconstruction” has allowed the production of proteins with enhanced properties (hyperstability and substrate promiscuity).17 Several ancestral β-lactamases have already been crystallized and their 3D structures determined, demonstrating that enzyme fold conservation extends up to approximately four billion years ago. A punctuated-equilibrium model of structural evolution was observed, in which new folds emerge over comparatively short periods, followed by long phases of structural stasis.18 Resurrected ancient enzymes were found to be hyperstable compared to extant enzymes and exhibited enhanced promiscuity,17 supported by a greater degree of active-site deformability.19 Both features, high stability and promiscuity, are advantageous for the integration of enzymes as rCLEC materials. Lysozyme and formamidase were previously crystallized, cross-linked, and enzymatically characterized in the so-called McCLEC (microfluidic chip for cross-linked enzyme crystal) device.20 The system was further improved by integrating an in situ spectrophotometric detection system based on photonic lab-on-a-chip technology. Successful operation under continuous flow was demonstrated using lipase.21 The use of enzymes as biocatalysts in microfluidic reactors has been previously assayed, and a number of different approaches have been reported.22,23
Lactamase has also been immobilized and employed as an active material in a capillary column microreactor. In that system, γ-lactamase retained 100% of its initial activity at 80 °C, despite a 70% reduction in catalytic activity due to immobilization compared to the free enzyme.22
In recent years, various analytical methods have been reported for the determination of β-lactam antibiotics to complement conventional instrumental techniques such as liquid chromatography–mass spectrometry, spectrophotometric, capillary electrophoresis or fluorescence methods.24 Among these, electrochemical sensors based on molecularly imprinted polymers (MIPs) have demonstrated nanomolar detection limits and good recovery rates in environmental and food-related matrices, highlighting their potential for sensitive and cost-effective on-site analysis.25–27
Other approaches, such as surface plasmon resonance (SPR) sensors combined with MIPs, have shown excellent selectivity for β-lactam antibiotics, although these systems typically rely on less portable and more sophisticated instrumentation.24 Rapid tests based on lateral flow immunoassays (LFIA) offer outstanding portability and short response times on the order of minutes, making them particularly suitable for in situ screening, although less sensitive when compared to electrochemical or SPR-based methods.28,29 Other technologies, such as a sensor for kanamycin to be used in environmental water samples, integrate portable plastic gold electrodes within a microfluidic chip, with good stability and a low detection limit.30
In this work, a novel microfluidic probe based on cross-linked crystals of an ancestral β-lactamase (βLa-CLECs) integrated into OSTEMER microfluidic chips provides an alternative sensing strategy that leverages enzymatic specificity, robustness under immobilization conditions, and compatibility with polymer-based microfluidic devices. The system operates under continuous flow with integrated detection, enabling the sensing of two common antibiotics, ampicillin and sulbactam, and is also able to detect CTX and CRO.
Our system compares well with other gold-standard technologies for antibiotic detection systems,26 with the LoD in the order of ppb and an analysis time lower than CA30 minutes. It presents a greater versatility, allowing for ex situ production of the crystalline active material and the precise quantification of loaded material. This new configuration offers distinct advantages for practical sensing applications, particularly where operational stability, reproducibility, and device integration are critical.
Experimental
Ancestral lactamase production
His-tagged ancestral β-lactamase was purified in sufficient quantity using protocols previously established in our laboratory.17,31 Briefly, Escherichia coli BL21 (DE3) cells were transformed with pET24-b vectors carrying the selected ancestral lactamase gene. Transformed colonies were grown on solid LB agar supplemented with kanamycin (50 µg mL−1) and then cultured in liquid LB medium (supplemented with 50 µg per mL kanamycin) at 37 °C. Once the culture reached an optical density (Abs600) of 0.6–0.8, protein expression was induced with 1 mM IPTG for 3 hours. Cells were harvested by centrifugation and stored at −80 °C until further use. Purification was carried out via immobilized metal affinity chromatography (IMAC) using a 1 mL Ni2+-charged column (GE Healthcare), followed by size-exclusion chromatography (Sephadex 16/60 S75) on an FPLC system. The resulting protein samples were extensively dialyzed in 20 mM Tris pH 8.0, aliquoted, flash-frozen in liquid nitrogen, and stored at −80 °C until further use.Crystallization and cross-linked crystal production
Optimal conditions for the crystallization and cross-linking of ancestral lactamase were determined using traditional crystallization techniques, along with SDS-PAGE analysis to confirm complete fixation of CLECs.Crystallization was carried out using the vapor diffusion method set up in a mushroom device, which allowed up to 100 drops to be run simultaneously, enabling medium-scale crystal production (Fig. S1). Each reservoir contained 5 mL of precipitant solution (5.0 M sodium formate and 0.1 M sodium acetate pH 4.0), and drops were prepared by mixing protein and precipitant solutions in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio (2 µL each). Freshly purified ancestral lactamase at 20 mg mL−1 was combined with the precipitant cocktail and allowed to equilibrate for one month at 293 K.
1 ratio (2 µL each). Freshly purified ancestral lactamase at 20 mg mL−1 was combined with the precipitant cocktail and allowed to equilibrate for one month at 293 K.
The crystals were subjected to a mild cross-linking treatment by supplementing the reservoir solution with 5% (v/v) glutaraldehyde, followed by equilibration for one week. For rapid βLa-CLEC production, glutaraldehyde was directly added to the drops at a final concentration of 2.5% (v/v). Cross-linked crystals were recovered, transferred to Eppendorf tubes, washed with Milli-Q water, and gently centrifuged. After excess water was removed, the βLa-CLECs were lyophilized and stored at room temperature.
For quantitative experiments, lyophilized βLa-CLECs were weighed using a Mettler Toledo XP26 high precision balance.
Enzyme assays
The catalytic activity of lactamase crystals was monitored spectrophotometrically by tracking the degradation of nitrocefin. Nitrocefin exhibits absorbance at 217 nm, corresponding to the 7-acyl group, and at 386 nm. Upon hydrolysis of the β-lactam ring by the enzyme, the 386 nm peak disappears and a new absorbance peak emerges at 482 nm. Both wavelengths were used to evaluate the extent of the reaction over time.According to commercial recommendations, a concentrated stock solution of nitrocefin was prepared in DMSO at 10.0 mg mL−1 concentration. Working solutions were obtained by diluting the stock solution in 100 mM HEPES buffer (pH 7.0) and stored at −80 °C, protected from light using aluminum foil (DMSO concentration was always kept below 1% in the working solution). After thawing to room temperature, substrate samples were kept in the dark to prevent light-induced degradation.
For inhibition assays, cefotaxime (CTX), ceftriaxone (CRO), ampicillin (AMP), and sulbactam (SUL) were prepared in 100 mM HEPES buffer (pH 7.0).
Microprobe conception and operation
The biocatalytic microdevices were fabricated using an OSTEMER 322 (Mercene Labs, Sweden) through injection molding, employing PDMS molds produced via standard soft lithography techniques based on a cost-effective dry film method.32 OSTEMER, a thiol–ene–epoxy composite, was specifically chosen for this application due to its unique two-step curing process, which enables complete reticulation. A detailed fabrication protocol is provided elsewhere.33The final microfluidic device consists of two distinct parts (Fig. 1A and B), which are aligned and assembled to form a complete system. Each structure is first molded and exposed to UV light, resulting in a solid yet flexible and tacky OSTEMER component. Structure A includes a square-shaped well where βLa-CLECs are manually deposited (Fig. 1C), with the amount precisely measured using a high-precision balance. In addition, structure A incorporates pillar-like features distributed along and across the 3 cm wide microchannel to ensure a uniform fluid distribution throughout the flow path. The loading and subsequent operation of the devices are facilitated by the strong adhesion of the βLa-CLECs to the OSTEMER surface, enabled by the presence of free epoxy groups.
Next, structure B (containing the inlet/outlet ports for fluid injection and defining the main microfluidic channel geometry) is carefully aligned and brought into contact with structure A, thereby sealing the microfluidic network. A final thermal curing step is then carried out at a mild temperature (∼50 °C) to complete the epoxy reticulation without compromising the stability of the ancestral biological material, which has been shown to exhibit greater thermal resistance than standard enzymes.17 The depth of different features comprising both structures A and B can be customized based on the average size of the βLa-CLECs, allowing for optimization of the volume-to-surface ratio by minimizing the cross-sectional area of the fluidic channels (Fig. 1D).
To operate the microdevices, various solutions (either containing only the substrate or a mixture of substrate and inhibitor) are loaded into glass syringes (Fig. 1E, syringes 1 and 2, respectively) and injected into the microdevices (highlighted in red in Fig. 1E) through a T-junction, at well-controlled flow rates using a standard syringe pump system (Nemesys, Cetoni, Germany). Given that the reagents are introduced at constant flow rates, these can be directly translated into residence times within the microdevices, representing the contact time between the solutions and the βLa-CLECs. Under steady-state conditions, this contact time dictates a consistent level of substrate conversion.
By adjusting the flow rate, the residence time is inversely modified: higher flow rates lead to shorter contact times and thus lower conversion, whereas lower flow rates result in longer contact times and higher conversion rates. Similarly, at a constant total flow rate, varying the substrate-to-inhibitor ratio allows modulation of the conversion efficiency, depending on the inhibitor's presence and concentration (Fig. S2).
To monitor substrate conversion in real time, a Photonic Lab-on-a-Chip (PhLoC8) device (highlighted in blue in Fig. 1E) is coupled in-line at the outlet of the biocatalytic reactor. The PhLoC system, described in detail elsewhere,34 functions as a microfluidic UV-vis cuvette featuring three optical paths of varying lengths, which extend the linear detection range while minimizing the required sample volume (∼4 µL). On either side of each optical path—defined by the length and width of a microfluidic channel—self-aligning microchannels are incorporated to ensure precise and easy positioning of fiber optics relative to 2D beam-collimating microlenses. These microlenses are designed to match the numerical aperture of the fiber optics.
For this study, only the longest optical path (1 cm, equivalent to a standard cuvette spectrometer) was utilized for real-time, on-line measurements. Light was coupled into and out of the PhLoC using 230 µm multimode pigtailed fiber optics (Thorlabs, Dachau, Germany). Illumination was provided by a DH-2000 Mikropack Deuterium-Tungsten Halogen Light Source (Ocean Optics, Dunedin, FL, USA), covering the 215–2000 nm wavelength range. Spectral data were acquired using an Ocean Optics MAYA 2000 spectrophotometer (Ocean Optics, Dunedin, FL, USA), enabling continuous monitoring of UV-vis absorption profiles for any solution passing through the detection region of the PhLoC.
Results and discussion
TEM-1 β-lactamases are enzymes that allow bacteria to resist antibiotics by hydrolyzing β-lactam compounds. Sequence reconstruction studies have led to the development of several putative Precambrian β-lactamases mimicking the modern TEM-1 enzyme. Among these, the enzyme reconstructed as the last common ancestor of Gram-negative bacteria (GNCA), estimated to be two billion years old, was selected for use in the antibiotic detector system based on cross-linked crystals. The crystallization behaviour and 3D structure of GNCA had been previously characterized by our group,17 revealing that it retains the typical lactamase fold despite significant sequence divergence (122 amino acid differences relative to TEM-1 β-lactamase). Two main properties justified the selection of this ancestral enzyme. First, it exhibited the highest thermal stability among the reconstructed nodes, with a melting temperature (Tm) increase of approximately 35 °C. Second, it demonstrated the greatest substrate promiscuity, showing substantially enhanced activity against third-generation antibiotics, whereas TEM-1 β-lactamase remains specialized for penicillins.17The initial activity of the histidine-tagged ancestral lactamase under standard assay conditions was confirmed using nitrocefin. Upon addition of the diluted enzyme, the slight yellow color of the nitrocefin solution turned pale pink. The same qualitative test was used to assess the activity in non-cross-linked ancestral lactamase crystals, which dissolved rapidly upon exposure to nitrocefin and immediately catalyzed the reaction (Fig. S3A). When stabilized βLa-CLECs were tested, a single crystal was sufficient to induce the color change from clear to pale pink (Fig. S3B).
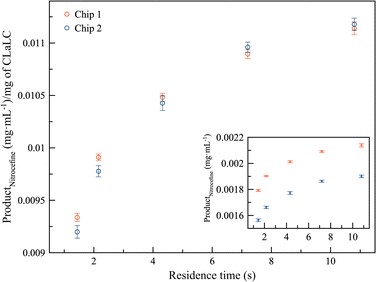
Previous work demonstrated that crystallization, cross-linking, and enzymatic characterization of lipase crystals could be successfully conducted in a microfluidic device operating under continuous flow conditions.21 In the present study, the device design was modified and adapted to allow precise loading of enzyme crystals, regardless of individual crystal size. Two microprobes were prepared with different amounts of βLa-CLECs and tested with a fixed nitrocefin concentration of 0.016 mg mL−1 at five different flow rates (i.e., residence times). Fig. 2 shows the high reproducibility between both microprobes after normalization by crystal weight. The independent measurements for both devices are also shown as an inset in the same figure.
The enzymatic activity of the βLa-CLEC microprobes was fully characterized using five different substrate concentrations, ranging from 5 to 40 µg mL−1, and ten distinct flow rates (i.e., residence times) for each concentration. Each experimental cycle was performed in triplicate to assess reproducibility, represented as standard deviation in Fig. 3A. Measurement reproducibility was further verified by accounting for uncertainties in substrate solution preparation and potential degradation. To evaluate this, two consecutive cycles were conducted using substrate solutions prepared at approximately 12 µg mL−1. As shown in Fig. 3B, the experimental outcomes aligned with expected results, with only slight variation attributed to differences in the initial substrate concentration.
Finally, the utility of the βLa-CLEC microfluidic probes for detecting marginal amounts of antibiotics in solution was evaluated by assessing the inhibitory effect on a robust β-lactamase. Four antibiotics—cefotaxime (CTX), ceftriaxone (CRO), ampicillin (AMP), and sulbactam (SUL) were used. The amount of nitrocefin was set at 10.0 µg mL−1 at a constant residence time, allowing for the competitive inhibitors to effectively reduce reaction rates. Under these conditions, increasing concentrations of sulbactam or ampicillin resulted in a proportional decrease in enzymatic activity, confirming competition for the active site. CTX and CRO did not exhibit inhibition levels greater than 10% at concentrations near 10.0 ppm and were therefore not analyzed further. Since the molecular weight of CTX and CRO are 455.46 and 554.57 g mol−1, respectively, compared to 349.41 and 233.24 g mol−1, for ampicillin and sulbactam, respectively, we can think of diffusion constraint within crystal channels.
Ampicillin (AMP), one of the most widely used antibiotics globally, was tested over a concentration range of 4 to 200 ppm (Fig. 4A). The minimum detectable concentration was determined through two rounds of experiments conducted at maximum concentrations of 10.0 ppm and 1.0 ppm, each repeated for reproducibility. A concentration of 1.0 ppm resulted in 10% loss of activity, while 250 ppb caused nearly a 3% reduction (inset in Fig. 4A). In the case of sulbactam (SUL), a single experiment was enough to reveal that the minimum detectable concentration was 200 ppb, which corresponded to a loss of enzymatic activity exceeding 10%, and confirming the robustness of the method.
Conclusions
In this work, an enzymatic microfluidic probe was developed using cross-linked crystals of a reconstructed ancestral β-lactamase (βLa-CLECs). The enzyme, corresponding to a variant of the last common ancestor of Gram-negative bacteria, was selected for its high thermal stability (Tm enhancement of ∼35 °C) and broad substrate promiscuity toward third-generation β-lactam antibiotics. Cross-linked enzyme crystals were produced, precisely weighed, and manually loaded into a modular OSTEMER-based microdevice, thanks to a specific 2-structure open design, allowing for consistent distribution of the βLa-CLECs and reliable adhesion within the microfluidic structure. The modular microprobes were connected in series with a photonic detection module for in-line on-chip UV-vis detection.The catalytic activity of the immobilized enzyme was retained after crystallization and cross-linking, as confirmed by the complete characterization of nitrocefin hydrolysis over a wide concentration range and under tunable residence times, controlled by varying the substrate injection flow rates. The inhibitory effect of selected antibiotics was evaluated. Ampicillin and sulbactam could be detected with high sensitivity and reproducibility. The lower limit of detection was achieved with sulbactam, 200 ppb, with activity losses exceeding 10% at those levels.
These findings confirm that cross-linked crystals of ancestral enzymes provide a stable and responsive biocatalytic material suitable for integration into lab-on-a-chip sensing platforms. The demonstrated sensitivity, stability, and reproducibility highlight the potential of this system for detecting antibiotics in aqueous samples at environmentally relevant concentrations under continuous flow.
Author contributions
The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Other data are contained in a mpzz file type for MagicPlot 3.0.1 software. Data from the four chips considered in this work are included. Supplementary information: (1) cross-linking & (2) experimental setup. See DOI: https://doi.org/10.1039/d5na00906e.Abbreviations
| (r)CLECs | (reinforced) Cross-linked enzyme crystals |
| CLEAs | Cross-linked enzyme aggregates |
| βLa-CLECs | Cross-linked ancestral β-lactamase crystals |
| McCLECs | Microfluidic chip for cross-linked enzyme crystals |
| PhLoC | Photonic lab-on-a-chip |
| SDS-PAGE | Sodium dodecyl sulfate polyacrylamide gel electrophoresis |
| AMP | Ampicillin |
| SUL | Sulbactam |
| CTX | Cefotaxime |
| CRO | Ceftriaxone |
| LoD | Limit of detection |
Acknowledgements
This work was partly funded by the Spanish Ministry of Science, Innovation and Agencia Estatal de Investigación http://dx.doi.org/10.13039/501100011033 grant PID2020-116261GB-I00 (JAG), the Institut National Politechnique de Toulouse ETI 2025 program (project InCySe) (IRR), and Junta de Andalucia-Consejeria de Universidad, Investigación e Innovación (PAIDI 2020) P21.00112 (VAR). PMG acknowledges the support of the CSIC through the JAE program (2025).Notes and references
- J. Liu, Q. Fu, Q. Li, Y. Yang, Y. Zhang, K. Yang, G. Sun, J. Luo, W. Lu and J. He, Pharm. Fronts, 2024, 6, e69–e100 Search PubMed.
- A. Zafar, C. Takeda, A. Manzoor, D. Tanaka, M. Kobayashi, Y. Wadayama, D. Nakane, A. Majeed, M. A. Iqbal and T. Akitsu, Molecules, 2024, 29(2), 398 CrossRef CAS PubMed.
- J. Bahnemann and A. Grünberger, Microfluidics in Biotechnology, 2022, vol. 179 Search PubMed.
- M. P. Cardoso Marques, A. Lorente-Arevalo and J. M. Bolivar, Adv. Biochem. Eng./Biotechnol., 2022, 179, 211–246 CrossRef CAS PubMed.
- J. Jang, W. S. Kim, T. S. Seo and B. J. Park, Chem. Eng. J., 2024, 495, 153657 CrossRef CAS.
- J. Britton, S. Majumdar and G. A. Weiss, Chem. Soc. Rev., 2018, 47, 5891–5918 RSC.
- J. M. Woodley, React. Chem. Eng., 2024, 9, 2028–2033 RSC.
- I. Rodríguez-Ruiz, T. N. Ackermann, X. Muñoz-Berbel and A. Llobera, Anal. Chem., 2016, 88, 6630–6637 CrossRef.
- R. Fernández-Penas, C. Verdugo-Escamilla, S. Martínez-Rodríguez and J. A. Gavira, Cryst. Growth Des., 2021, 21, 1698–1707 Search PubMed.
- A. Lambarska, K. Szymanska and U. Hanefeld, Green Chem., 2024, 26, 10718–10738 RSC.
- R. B. R. Valério, I. G. de Sousa, A. L. G. Cavalcante, J. E. da Silva Souza, T. G. Rocha, F. S. Neto, M. C. M. de Souza and J. C. S. dos Santos, in Biocatalyst Immobilization, Elsevier, 2023, pp. 87–114 Search PubMed.
- J. Jegan Roy and T. Emilia Abraham, Chem. Rev., 2004, 104, 3705–3722 CrossRef CAS PubMed.
- R. Contreras-Montoya, M. Arredondo-Amador, G. Escolano-Casado, M. C. Mañas-Torres, M. González, M. Conejero-Muriel, V. Bhatia, J. J. Díaz-Mochón, O. Martínez-Augustin, F. S. de Medina, M. T. Lopez-Lopez, F. Conejero-Lara, J. A. Gavira and L. Á. de Cienfuegos, ACS Appl. Mater. Interfaces, 2021, 13, 11672–11682 CrossRef CAS PubMed.
- S. Kara and F. Rudroff, Enzyme Cascade Design and Modelling, Springer International Publishing, Cham, 2021 Search PubMed.
- B. T. Porebski and A. M. Buckle, Protein Eng., Des. Sel., 2016, 29, 245–251 CrossRef CAS PubMed.
- V. A. Risso, J. M. Sanchez-Ruiz and S. B. Ozkan, Curr. Opin. Struct. Biol., 2018, 51, 106–115 Search PubMed.
- V. A. Risso, J. A. Gavira, D. F. Mejia-Carmona, E. A. Gaucher and J. M. Sanchez-Ruiz, J. Am. Chem. Soc., 2013, 135, 2899–2902 CrossRef CAS PubMed.
- A. Ingles-Prieto, B. Ibarra-Molero, A. Delgado-Delgado, R. Perez-Jimenez, J. M. Fernandez, E. A. Gaucher, J. M. Sanchez-Ruiz and J. A. Gavira, Structure, 2013, 21, 1690–1697 Search PubMed.
- T. Zou, V. A. Risso, J. A. Gavira, J. M. Sanchez-Ruiz and S. B. Ozkan, Mol. Biol. Evol., 2015, 32, 132–143 CrossRef CAS PubMed.
- M. Conejero-Muriel, I. Rodríguez-Ruiz, S. Martínez-Rodríguez, A. Llobera and J. A. Gavira, Lab Chip, 2015, 15, 4083–4089 Search PubMed.
- M. Conejero-Muriel, I. Rodríguez-Ruiz, C. Verdugo-Escamilla, A. Llobera and J. A. Gavira, Anal. Chem., 2016, 88, 11919–11923 Search PubMed.
- A. M. Hickey, B. Ngamsom, C. Wiles, G. M. Greenway, P. Watts and J. A. Littlechild, Biotechnol. J., 2009, 4, 510–516 Search PubMed.
- I. Rodríguez-Ruiz, M. Conejero-Muriel, T. N. Ackermann, J. A. Gavira and A. Llobera, Lab Chip, 2015, 15, 1133–1139 Search PubMed.
- G. Liang, L. Song, Y. Gao, K. Wu, R. Guo, R. Chen, J. Zhen and L. Pan, Toxics, 2023, 11, 513 CrossRef CAS PubMed.
- R. López, S. Khan, A. Wong, M. del P. T. Sotomayor and G. Picasso, Front. Chem., 2021, 9, 1–9 Search PubMed.
- M. Frigoli, M. P. Krupa, G. Hooyberghs, J. W. Lowdon, T. J. Cleij, H. Diliën, K. Eersels and B. van Grinsven, Sensors, 2024, 24, 5576 Search PubMed.
- V. B. Chau Nguyen, J. Reut, A. G. Ayankojo and V. Syritski, Talanta, 2025, 287, 127580 Search PubMed.
- A. Pollap and J. Kochana, Biosensors, 2019, 9(2), 61 CrossRef CAS PubMed.
- L. Kaiser, J. Weisser, M. Kohl and H.-P. Deigner, Sci. Rep., 2018, 8, 5628 CrossRef PubMed.
- Y. Bao, K. Cai, Y. Liu and B. Li, Chin. J. Chem., 2024, 42, 171–176 CrossRef CAS.
- V. A. Risso, S. Martinez-Rodriguez, A. M. Candel, D. M. Krüger, D. Pantoja-Uceda, M. Ortega-Muñoz, F. Santoyo-Gonzalez, E. A. Gaucher, S. C. L. Kamerlin, M. Bruix, J. A. Gavira and J. M. Sanchez-Ruiz, Nat. Commun., 2017, 8, 1–13 Search PubMed.
- I. Rodríguez-Ruiz, S. Teychené, N. Van Pham, D. Radajewski, F. Lamadie, A. Llobera and S. Charton, Talanta, 2017, 170, 180–184 CrossRef PubMed.
- R. K. Ramamoorthy, E. Yildirim, I. Rodriguez-Ruiz, P. Roblin, L.-M. Lacroix, A. Diaz, R. Parmar, S. Teychené and G. Viau, Lab Chip, 2024, 24, 327–338 Search PubMed.
- I. Rodríguez-Ruiz, F. Lamadie and S. Charton, Anal. Chem., 2018, 90, 2456–2460 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |