Open Access Article
Open Access ArticleOne system, three functions: an electroactive reconfigurable organic receptor for adaptive binding, information encoding, and metal-free oxidation
Logeshwari Seethapathy a,
Rohith Muthusamy
a,
Rohith Muthusamy a,
Chinmoy K. Hazra
a,
Chinmoy K. Hazra b,
Jayanta Samanta
b,
Jayanta Samanta a,
Susnata Pramanik
a,
Susnata Pramanik a and
Rajorshi Das
a and
Rajorshi Das *a
*a
aDepartment of Chemistry, College of Engineering and Technology, SRM Institute of Science and Technology, Kattankulathur, Chennai, 603203, India. E-mail: rajorshd@srmist.edu.in; rajorika@gmail.com
bDepartment of Chemistry, Indian Institute of Technology Delhi, Hauz Khas, South Delhi, 110016, India
First published on 7th April 2026
Abstract
Living systems rely on biomolecular receptors capable of selectively responding to multiple chemical cues, enabling precise information processing and adaptive functionality. Inspired by this paradigm, we report a multifunctional synthetic receptor that functions in stimuli-responsive conformational switching, programmable information encoding, and metal-free oxidation within a single molecular system. The receptor features a rigid, planar core that undergoes an unexpected and extensive deformation upon chemical alkylation, adopting a bowl-shaped geometry that can be fully reverted to its original flat structure upon guest recognition. This reversible transformation enables adaptive, induced-fit binding of diverse guest molecules, allowing the host to modulate its geometry to optimize donor–acceptor interactions and maximize complex stability. Electrochemical studies show that the resulting cationic species 1(PF6)3 is highly electron-deficient and serves as an efficient oxidant for metal-free oxidative transformations. Furthermore, the programmable conformational and redox states of the receptor provide a basis for constructing tunable donor–acceptor systems capable of molecular-level information encoding and decoding.
New conceptsThis study introduces a new conceptual framework in supramolecular chemistry by demonstrating that a traditionally rigid, planar ligand, 2,4,6-tri(4-pyridyl)-1,3,5-triazine (TPT), can undergo pronounced and reversible conformational deformation without metal coordination. Simple N-alkylation induces an unprecedented planar-to-bowl transformation of the free ligand, revealing chemically triggered flexibility in an otherwise static scaffold. Importantly, this deformation is reversible: selective guest recognition restores the planar geometry, enabling size-adaptable induced-fit host–guest interactions reminiscent of biological receptors. The conformational switching directly modulates the electronic structure of the system, leading to controlled repositioning of the HOMO–LUMO levels. This feature allows chemical information to be encoded and decoded through donor–acceptor charge-transfer interactions, establishing a platform for molecular-level encryption and decryption. Furthermore, the enhanced electron deficiency of trialkylated TPT enables efficient metal-free oxidation chemistry. Overall, this work establishes chemical stimulus-induced flexibility in a rigid free ligand as a powerful new concept, integrating adaptive recognition, information processing, and reactivity within a single organic system. |
Introduction
Biomolecular receptors are essential components of living systems, processing information through selective responses to chemical signals.1 They can recognize multiple stimuli and exhibit induced-fit binding, where the presence of one molecule changes the shape and size of the parent biological host influencing the cooperative binding.1–4 The working principles of these natural systems provide a foundation for the synthesis of programmable molecular architectures, which can be designed according to the requirements. Such systems may generate distinct outputs based on specific input(s), enabling applications in molecular logic,5,6 sensing,7,8 and catalysis.9One promising approach to achieving such functionality is the use of self-assembled architecture with size-adaptable guest binding. Raymond,10 Fujita,11–14 Clever,15–17 Nitschke,18–21 Mukherjee22,23 and some other groups24–27 have reported several supramolecular systems that can alter their architecture in the presence of external guest molecules. For example, Raymond's M2L3 helicate cage undergoes a structural transformation to a tetrahedron via guest encapsulation.10 An M18L6 molecular capsule reported by Fujita can expand its inner core by reassembling its components to a larger capsule (M24L8) to accommodate guest molecules inside its cavity.13 In a similar fashion, an M2L4 cage reported by Clever converts its structure to an interlocked [3]3-catenane triggered by an excess amount of halide ions.15 Sun's M4L2 metallohelix can also potentially transform its secondary structure to a tertiary protein-like structure after anion binding.27
All the examples described above elaborate on the reassembly of supramolecular structures in the presence of guest molecules, which follows a self-correction pathway, a routine procedure of metal–ligand dissociation and reassociation to generate thermodynamically more stable product(s). In these cases, the rigid ligands used for the construction of supramolecular structures remain rigid, and only the reconstitution of metal–ligand self-assembly with a greater number of metal ions and ligands occurs, leading to the formation of larger supramolecular structures. The induction of structural flexibility in a rigid ‘free’ ligand (where the ligand is not bound to any metal ion forming a constrained supramolecular assembly) through external stimuli is extremely difficult, and therefore it has been unprecedented. However, if possible, it would lead to the generation of novel materials that can not only show induced fit guest-recognition ability, but may also exhibit other functions including chemical transformations and information security.
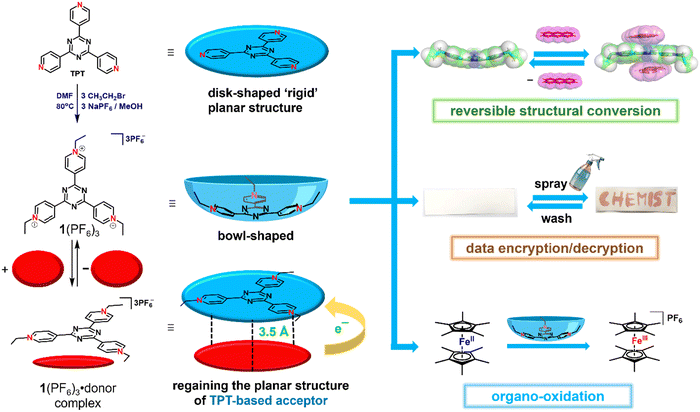
Herein, in an unprecedented reaction, we demonstrate that the rigidity of a ligand can significantly be altered by chemical stimuli without changing the core chemical structure of the rigid ligand. We found that the alkylation of a planar 2,4,6-tri(4-pyridyl)-1,3,5-triazine (TPT) yielded an unusual ‘bowl-shaped’ trialkylated tripyridiniumtriazine derivative, 1(PF6)3, which can regain its planarity upon charge-complementary molecular recognition (Scheme 1). This trialkylation not only imparts remarkable structural flexibility to the TPT molecule but also generates a highly electron-deficient, high-valent cationic system. Notably, the strong electron-accepting nature of triethylated TPT, 1(PF6)3, makes it a promising candidate for its applications in data encryption/decryption and metal-free oxidation reactions.
Results and discussion
Induction of chemical flexibility in a rigid TPT motif
TPT is a well-known tripodal ligand widely used in supramolecular chemistry for the construction of a variety of self-assembled structures.28,29 According to our literature survey, over 300 crystal structures of TPT–metal complexes have been reported to date. Notably, the majority of these complexes exhibit TPT in a planar conformation, consistent with its inherent rigid molecular geometry. However, there is only an isolated example in which the TPT unit adopts a non-planar configuration. In this rare case, the triazine core displays a measurable deviation from planarity, typically attributed to the severe strain induced by the formation of a rigid supramolecular cage.30In our approach, structural flexibility was introduced into the otherwise rigid TPT molecule by modulating its electronic environment through chemical and charge-transfer processes, as illustrated in Scheme 1. The tricationic species 1(PF6)3 was synthesized following a straightforward two-step procedure. Initially, TPT was reacted with an excess of ethyl bromide at 50 °C for 48 hours in a sealed tube, resulting in the formation of the tri-N-ethylpyridinium tribromide salt 1(Br)3. Subsequent anion exchange with hexafluorophosphate gave the corresponding hexafluorophosphate salt 1(PF6)3 in 65% yield as a colorless solid. The product 1(PF6)3 exhibits good solubility in polar solvents such as acetonitrile, dimethylformamide, and dimethyl sulfoxide, whereas it remains insoluble in diethyl ether and other non-polar solvents.
The formation of 1(PF6)3 was confirmed by multinuclear NMR spectroscopy, high-resolution electrospray ionization mass spectrometry (HR-ESI MS), single-crystal X-ray diffraction (SCXRD) and cyclic voltammetry analysis. In the 1H NMR spectrum, two peaks were obtained in the aromatic region at δ = 9.52 and 9.48 ppm in DMSO-d6 for pyridinium protons (Fig. S1 in the SI). The other CH3 and CH2 protons appeared at δ = 1.68 and 4.84 ppm, respectively. The 13C {1H} NMR spectrum shows the appearance of the aromatic carbon atoms at δ = 169.0, 148.5, 146.2 and 127.1 ppm and the alkyl carbon atoms at 57.1 and 16.5 ppm, as expected (Fig. S2). The molecular composition of 1(PF6)3 was further verified by HR-ESI MS (Fig. S12). The spectrum showed the highest intensity peak for the molecular ion [1(PF6)2]+ with the expected isotopic distribution patterns (the signal was obtained at m/z = 689.1559 for [1(PF6)2]+; calcd m/z = 689.1580 for C24H27F12P2N6).
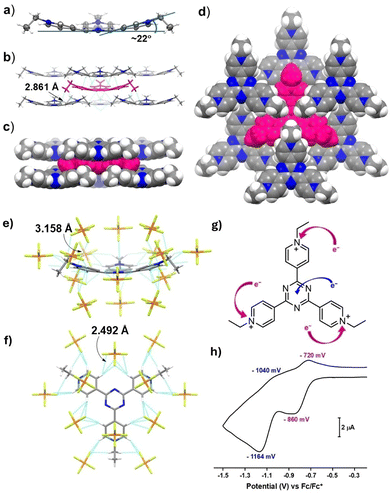
The formation of the trication 1(PF6)3 is unequivocally confirmed by the SCXRD analysis. Suitable crystals were grown by slow evaporation of the solvent from a concentrated acetonitrile solution of 1(PF6)3. The compound crystallized in the trigonal space group R3m with a three-fold symmetry. The structural analysis reveals an unusual bowl-shaped structure of [1]3+, unlike the conventional planar form of metal-coordinated TPT. The molecular solid-state structure of [1]3+ along with its packing motifs is displayed in Fig. 1. The angle between the ethylated pyridinium ring and the central triazine plane measures 22° as depicted in Fig. 1a. The packing structure shows that one tricationic [1]3+ is in short contact with six nearby [1]3+ (Fig. 1b–d) and fifteen hexafluorophosphate (PF6−) ions via C–H⋯π (2.861 Å), F⋯π (3.158 Å, Fig. 1e and f), and non-classical hydrogen bonds (C–H⋯F, 2.463–2.625 Å).
In order to realize the solution-state structure of 1(PF6)3 and examine the possibility of conformational interconversion in solution, we performed two complementary 1H NMR experiments. First, concentration-dependent 1H NMR spectra of 1(PF6)3 were recorded in DMSO-d6 (2.5, 10, and 20 mg in 0.5 mL). At lower concentrations, two well-resolved pyridinium proton signals are observed. However, at higher concentrations, these peaks progressively merge into two broad resonances (Fig. S3). This behaviour suggests intermolecular interactions at elevated concentrations, likely arising from the proximity between pyridinium units of neighbouring molecules, which may induce partial distortion of the bowl-shaped conformation and dynamic averaging of the signals.
Second, a donor–acceptor titration experiment was conducted by adding increasing amounts (0.2–1.0 eq.) of pyrene as a donor to a constant concentration of 1(PF6)3. Upon gradual addition (notably around 0.6 eq.), the initially resolved pyridinium signals broaden and eventually coalesce into a single broad peak, consistent with strong π–π/charge-transfer interactions and guest-induced structural reorganization from a bowl-shaped to a more planar geometry (Fig. S11).
Next, we investigated the electrochemical behaviour of 1(PF6)3 to realize the electron deficiency caused by the trialkylation of tripyridyltriazine. It is worth mentioning that the triazine is an inherent electron-deficient moiety frequently used as a hydrophobic unit to impose electron deficiency in a system. After trialkylation, the electron-deficient TPT becomes even more electron-deficient due to the loss of electrons from the three pyridyl units. As expected, the cyclic voltammogram of 1(PF6)3 shows two redox waves in the reduction region. The first wave corresponds to the one-electron reduction of the pyridyl cation that is observed at E1/2 = −790 mV, and the second wave found at E1/2 = −1102 mV can be attributed to the one-electron reduction of the triazine unit. The cyclic voltammetry data along with the square-wave voltammogram are depicted in Fig. 1g and h and Fig. S16. The observed potentials match well with those of the previously reported alkylated pyridinium compounds.31
Charge-complementary molecular recognition and bowl-to-disc conversion
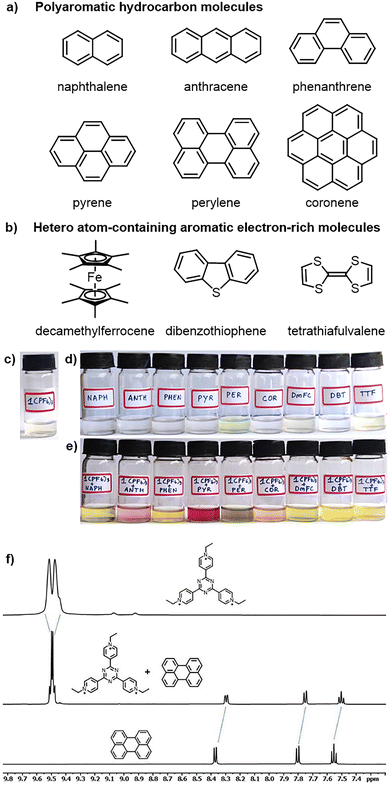
Considering the cationic nature and the strong electron affinity of 1(PF6)3, we next investigated its guest-recognition ability toward electron-rich guest molecules. The initial screening of guests covers a range of differently sized molecules ranging from the two-ring system naphthalene to the seven-ring system coronene depicted in Fig. 2a and b.In all cases, 1.5 equivalents of guests were added to the acetonitrile solution of 1(PF6)3. While the individual solutions of 1(PF6)3 and naphthalene, anthracene, phenanthrene, pyrene or coronene are colourless or light yellow, their mixtures gave yellow, green or deep red coloured solutions as shown in Fig. 2c–e. The change of colours from their parent solutions indicates charge transfer from the electron-rich polyaromatic hydrocarbon (PAH) guests to the electron-poor tricationic 1(PF6)3.
The formation of host–guest complexes was confirmed by 1H NMR, UV-vis, and infrared (IR) spectroscopy and X-ray crystallographic analysis. Suitable quality single crystals were obtained by slow evaporation of a concentrated acetonitrile solution from their host–guest mixtures. The co-crystals were dissolved in DMSO-d6, and the resulting solutions were subjected to 1H NMR studies, which show the presence of both donor (neutral guest) units and the acceptor (cationic host) unit, confirming the formation of charge-complementary adducts (Fig. 2f and Fig. S4–S9 in the SI). The NMR spectral analysis also indicated the upfield shifts of the chemical resonances of all the guest molecules (donors) compared to the strong acceptor 1(PF6)3. For example, while the aromatic proton resonances of perylene appear at δ = 7.55, 7.80 and 8.37 ppm and the pyridinium protons appear at 9.49 and 9.52 ppm, their donor–acceptor adduct shows resonances at δ = 7.50, 7.75 and 8.29 (for perylene) as well as at 9.48 and 9.50 ppm (for pyridinium protons) in DMSO-d6. Similar observations have been made for pyrene and anthracene molecules (Fig. S4–S7).
The observed upfield shifts of the perylene proton signals upon complexation with 1(PF6)3 arise from the shielding effects generated by the host molecules, for example, π–π stacking, ring-current effects, and recognition between the two [1]3+. These effects primarily influence the electronic environment of the ‘guest’ perylene protons, leading to detectable chemical shift changes. In contrast, the proton resonances of the host 1(PF6)3 remain essentially unchanged due to the fast exchange on the NMR time scale. This mainly happens in the case of dynamic host–guest equilibrium. The rapid exchange between free and bound states can lead to averaged signals for the host. However, in the case of 1(PF6)3·pyrene, the 1H NMR spectrum shows a significant shift of the pyridinium protons to the upfield region (Fig. S10 and S11).
The 1H NMR spectra also indicate that all the donor molecules exhibit fast exchange kinetics with the acceptor 1(PF6)3 on the NMR timescale. To further elucidate the binding stoichiometry and determine the corresponding binding constants, photoluminescence titration experiments were carried out.32,33 The binding studies revealed that the binding isotherms of the guests, such as naphthalene, anthracene, and pyrene, were best fitted to a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 stoichiometric model, indicating that one planar donor can recognize up to two molecules of the acceptor or vice versa (Table 1 and Tables S6–S8, Fig. S17–S19). For perylene, the binding isotherm shows a 1
2 stoichiometric model, indicating that one planar donor can recognize up to two molecules of the acceptor or vice versa (Table 1 and Tables S6–S8, Fig. S17–S19). For perylene, the binding isotherm shows a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 binding stoichiometry. Owing to the larger and more extended π-conjugated surface of perylene, it can form a stronger and more extensive π–π interaction with a single acceptor unit (Table S9 and Fig. S20).
1 binding stoichiometry. Owing to the larger and more extended π-conjugated surface of perylene, it can form a stronger and more extensive π–π interaction with a single acceptor unit (Table S9 and Fig. S20).
| Entry | Guests | K11 (M−1) | K12 (M−1) | NMR kinetics |
|---|---|---|---|---|
| a K11 and K12 values are calculated by the global fit V0.5 analysis from fluorescence titrations using a constant guest concentration of 0.01 mM and gradual addition of 5 mM 1(PF6)3 in ACN at 298 K. | ||||
| 1 | Naphthalene | 4476 ± 121 | 70![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 234 ± 152 234 ± 152 |
Fast |
| 2 | Anthracene | 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 774 ± 28 774 ± 28 |
699 ± 11 | Fast |
| 3 | Pyrene | 299 ± 32 | 2792 ± 51 | Fast |
| 4 | Perylene | 451 ± 4 | Nil | Fast |
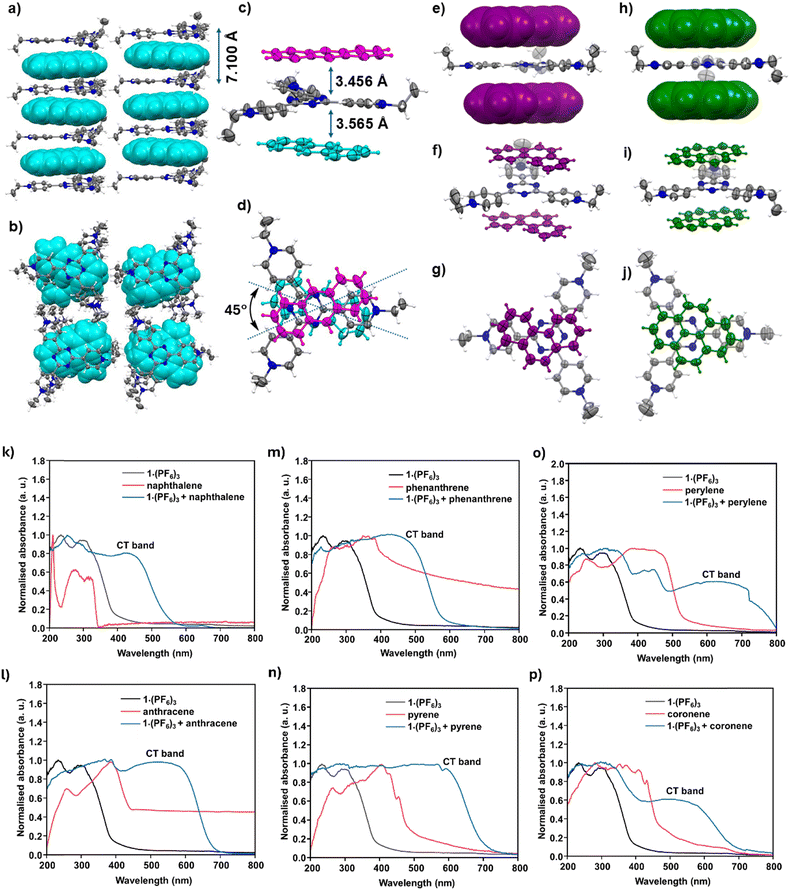
The recognition of electron-rich molecules by the acceptor 1(PF6)3 was unequivocally confirmed by X-ray crystallographic studies. The diffraction analysis of 1(PF6)3·anthracene shows the presence of one anthracene and one 1(PF6)3 in the asymmetric unit. The anthracene molecule is trapped between the two molecules of triethyltripyridiniumtriazine units by the charge-complementary electrostatic as well as π–π stacking interactions (Fig. 3a and b and Fig. S13). The distance between the donor and the acceptor is around 3.5 Å (Fig. 3c). The two anthracene molecules separated by one [1]3+ are lying in two different orientations with an angle of 45° (Fig. 3d). The distance between the two [1]3+ units measures 7.1 Å, as displayed in Fig. 3a.
Interestingly, the ‘bowl’-shaped structure of [1]3+ became flattened after the recognition of the electron-rich polyaromatic hydrocarbon anthracene molecule. The molecular recognition was driven by the charge-complementary interactions allowing charge separation from the donor (D) to acceptor (A) components to form an infinite ⋯Dd+–Ad−–Dd+–Ad−–Dd+–Ad−⋯ chain via the electrostatic interactions between the anthracene and the tricationic [1]3+. In principle, the π–π interactions between the donor and acceptor components become strongest when both are planar giving the maximum overlap between them rather than the interaction between a planar and a bowl-shaped species, which only gives a point interaction. Thus, the overall stability of the charge-transfer complex drives the flattening of the ‘bowl’ shaped 1(PF6)3 to a planar structure in the presence of planar guest molecules.
To extend the concept of flattening of [1]3+ in the presence of planar donors, we also attempted co-crystallization of 1(PF6)3 with other polyaromatic hydrocarbon donor molecules (Fig. 3). Gratifyingly, the single crystal X-ray diffraction studies with the crystals of 1(PF6)3·phenanthrene and 1(PF6)3·pyrene unequivocally confirm the presence of a planar conformation of [1]3+ trapped between the two phenanthrene or two pyrene molecules (Fig. 3e and h). Unlike the anthracene in 1(PF6)3·anthracene, the orientation of phenanthrene and pyrene molecules is found to be perfectly aligned on top of each other, as depicted in Fig. 3f, g and i, j. Intriguingly, all the hexafluorophosphate (PF6−) counter-anions in these donor–acceptor adducts are pushed away from the central triazine core due to the strong π–π stacking of PAH molecules, further elaborating the fact that the noncovalent electrostatic interactions in these systems are much stronger than the halogen (F)⋯π interactions (Fig. S13–S15).
The formation of charge-complementary interaction-driven recognition of electron-rich donors by 1(PF6)3 is also evident from UV-vis spectroscopy. The characteristic colour change associated with CT interaction becomes prominent only at relatively high concentrations. Under these conditions, the absorbance values rapidly approach 5–6 a.u., exceeding the reliable linear detection range of the spectrophotometer and compromising quantitative analysis. At lower concentrations suitable for UV-vis measurements, the CT band is too weak to be distinctly resolved in solution. To address this limitation, we performed solid-state UV-vis measurements, where the donor–acceptor interaction could be clearly detected through the emergence of a CT band in the visible region. Thus, while solution-phase detection is constrained by instrumental limits, the solid-state data provide clear evidence of charge-transfer interaction.
The solid-state UV-vis spectra of all neutral donors and 1(PF6)3 and of the D–A adducts from 1(PF6)3·naphthalene to 1(PF6)3·coronene were recorded in an integrating sphere to reduce intensity losses due to scattering and reflection. As shown in Fig. 3k–p, each of the D–A complexes displays a new band in their UV-vis spectra in addition to their individual bands of the neutral donor and acceptor units. Such bands, often found in the visible-NIR region, are characteristic of the charge-transfer (CT) transition in their D–A adducts, confirming the ability of the newly synthesized 1(PF6)3 acceptor to recognize electron-rich donors.
For example, while 1(PF6)3 and anthracene34 display intense bands at 220 and 300 nm, and 250 and 390 nm, respectively, their charge-complementary adduct 1(PF6)3·anthracene shows discernible broad bands at 380 and ca. 560 nm. Similarly, phenanthrene35 to coronene molecules can also be successfully recognized by the tripyridiniumtriazine cation [1]3+. Notably, the CT bands for more electron-rich anthracene, pyrene, perylene and coronene are observed at the lower energy region from 500 to 700 nm compared to the less electron-rich naphthalene showing its CT band at the relatively higher energy region at λmax = 440 nm.
Interestingly, the acceptor 1(PF6)3 can be separated easily from its CT adducts by dissolving the D–A crystals in acetonitrile followed by column chromatography (Scheme 1). The single crystals obtained from the purified 1(PF6)3 were further mounted on an X-ray diffractometer, which showed identical cell parameters, confirming the reversal of the bowl structure of 1(PF6)3.
Data encryption/decryption with 1(PF6)3
The increasing demand for secure information storage and transfer has driven chemists to explore a new frontier at the interface of chemistry, materials science and cryptography.36–38 Unlike traditional digital encryption, molecular-level approaches harness chemical reactivity, conformational switching, supramolecular recognition, and stimuli-responsive behavior to encode, store, and conceal information in physical matter.38,39 These strategies provide unique advantages in non-digital operation, low detectability, and environmental responsiveness, offering fundamentally new platforms for secure communication and anti-counterfeiting technologies.Since the electron-deficient receptor 1(PF6)3 undergoes predictable transformations and exhibits distinct color changes in response to chemical stimuli, we explored its potential for molecular-level data encryption and decryption using donor–acceptor-based charge-transfer interactions (Fig. 4). To demonstrate this concept, a piece of paper was encoded with the word ‘CHEMIST’ written using an acetonitrile solution of 1(PF6)3 (Fig. 4b). Because 1(PF6)3 is colorless, the message became completely invisible after drying for a few minutes, resulting in an unrecognizable white sheet (Fig. 4c). Spraying the surface with an acetonitrile solution of pyrene, a suitable electron donor that rapidly forms a visible charge-transfer complex with 1(PF6)3, revealed the hidden message (Fig. 4d and e). The decoded information could then be erased by gently washing the sheet with acetonitrile, which removed the donor and restored the blank appearance (Fig. 4f and g). Notably, this gentle washing step leaves the acceptor embedded within the paper, enabling regeneration of the message upon reapplication of the pyrene spray (Fig. 4h and i). Because the donor–acceptor binding is relatively weak, gentle washing with acetonitrile preferentially removes the externally applied donor, which is more loosely associated and more readily solvated. In contrast, the pre-deposited and dried 1(PF6)3 remains largely retained on the paper substrate due to its stronger interactions with the paper material (cellulose matrix), enabling the acceptor to persist under mild washing conditions. For complete removal of the encrypted data, the sheet can be washed two to three times to remove the residual acceptor from the sheet of paper.
This reversible process elaborated above highlights the potential of 1(PF6)3 for recyclable, stimuli-responsive, molecular-level information security. Interestingly, this method can also be applied in rewritable printing technology. Fabricating paper containing similar 1(PF6)3 acceptor-like molecules may enable its use as a rewritable medium, where pyrene solution can be used as the ink.
1(PF6)3 as an organo-oxidant
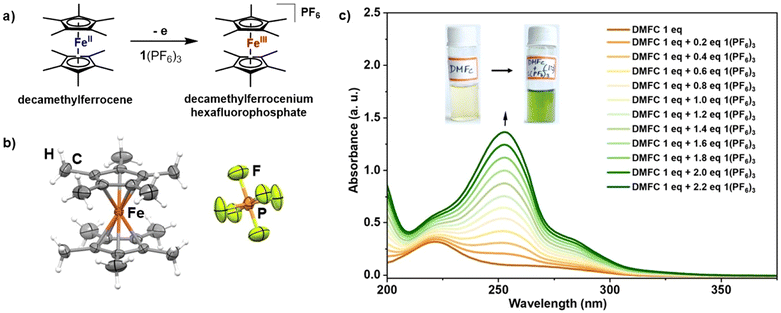
Inspired by the results of charge-complementary molecular recognition of planar electron-rich PAH molecules by the electron-deficient 1(PF6)3, we next investigated its ability to fully oxidize electron-rich metal complexes like decamethylferrocene (DMFC) as shown in Fig. 5a. In principle, for successful oxidation, the oxidation potential of a compound must be equal to, less than or at least close to the reduction potential of an oxidant. In the case of DMFC, the Fe2+/3+ oxidation process is found to occur at −550 mV against the ferrocene/ferrocenium standard in dichloromethane,40–42 which is lower than the first reduction potential of 1(PF6)3, observed at E1/23+/0 = −290 mV in acetonitrile. Since the redox potential strictly depends on the solvents used for measurement, it can be assumed that the tricationic 1(PF6)3 can potentially oxidize DMFC in acetonitrile. As expected, while adding the colourless 1(PF6)3 into the orange acetonitrile solution of DMFC, an immediate formation of a green solution was observed (Fig. 2d and e). The color change confirms the formation of a decamethylferrocenium cation (DMFC+).43 The in situ oxidation of DMFC can also be monitored by UV-vis spectroscopy. The pristine DMFC shows a band at λ = 222 nm. Upon gradual addition of the acceptor 1(PF6)3, a new band appears around 252 nm along with an increase in the intensity of the 222 nm band (Fig. 5c). The oxidation of DMFC is also verified with the addition of silver hexafluorophosphate (EAg/Ag+ = 40 mV in acetonitrile vs. Fc/Fc+ redox couple), which shows identical results to those observed with 1(PF6)3 addition (Fig. S22 and S23).After slow evaporation at ambient temperature, the resulting solution yielded green rod-shaped crystals, which, through the X-ray crystallographic study, were confirmed to be the decamethylferrocenium cation [Me5C5FeIIIC5Me5]+, the one-electron oxidized product of the parent DMFC. The molecular structure of decamethylferrocenium hexafluorophosphate is displayed in Fig. 5b.
It is noteworthy that such oxidation has previously been performed only with strong oxidants, for example, cerium(IV) ammonium nitrate (CAN), ferric chloride (FeCl3) or silver cations (Ag+).40 However, in our case, the oxidation can be done purely under metal-free conditions.
Theoretical calculations
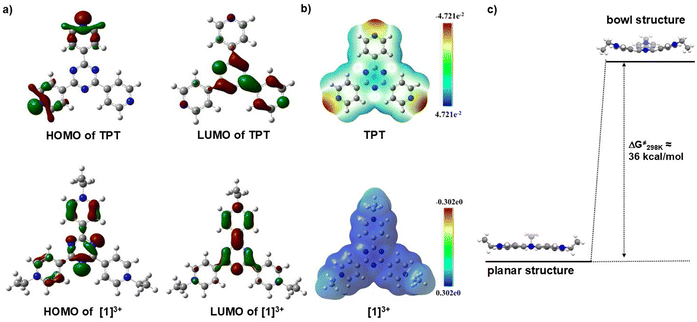
The structural deformation of the rigid planar TPT induced by trialkylation was analyzed using density functional theory (DFT) calculations. We first calculated the effect of alkylation on the TPT moiety. The highest occupied molecular orbitals (HOMOs) and the lowest unoccupied molecular orbitals (LUMOs) of TPT and triethylated TPT 1(PF6)3 are illustrated in Fig. 6a. The calculations show a significant change of HOMOs on the TPT moiety after alkylation, while the LUMO remains almost unchanged. This is obvious due to the fact that the electron-rich pyridine ring with one nonbonding lone pair of electrons becomes electron-poor after alkylation. Therefore, while the HOMOs of TPT reside on the pyridine ring, the alkylation yielded a total shift from the pyridinium ring to a comparatively electron-rich triazine core. This theoretical HOMO–LUMO observation is also supported by the Hirshfeld plot (Fig. 6b).Notably, the geometry-optimized structure of triethyltripyridiniumtriazine [1]3+ provided a perfectly planar conformation, which contrasts with the isolated ‘bowl’-shaped structure of [1]3+ (Fig. 1, vide supra), obtained from the single-crystal XRD analysis. The reason behind the stabilization of the bowl structure is not very clear. Therefore, we performed single-point energy calculations using the same functional and basis set to observe the difference in energy between the two conformations. The calculations revealed that the ground state energy of the bowl structure is 0.05743124 HF units (∼36 kcal mol−1, HF = Hartree–Fock) higher than the planar structure (Fig. 6c). This energy difference is perfectly in line with the bending vibration (elastic molecules) for such a moderately sized cationic species with four rigid and planar six-membered rings.44–46 However, we assume that the uneven hydrophobic interactions between the cationic [1]3+ and fifteen counter-anions (PF6−) play a significant role in its stabilization in the bowl form (Fig. 1e and f).
Conclusion
We have described the design and synthesis of a stimuli-responsive molecular system that exhibits unique chemically induced structural flexibility on a rigid ligand scaffold. Unlike traditional supramolecular systems where guest-induced transformation occurs through metal–ligand reassembly, this work showcases a rare example of conformational switching in a ‘free’ ligand, 2,4,6-tri(4-pyridyl)-1,3,5-triazine (TPT), via N-alkylation. The resultant trialkylated tricationic ligand 1(PF6)3 adopts a bowl-shaped conformation, which flattens upon binding planar electron-rich guests, enabled by charge-transfer and π–π interactions. This bowl-to-disc conversion opens new possibilities for dynamic host–guest systems and redox-tunable molecular devices. This structural adaptability is particularly important in supramolecular systems, leading to emergent properties such as all-or-nothing templating effects and varying degrees of cooperativity.1,47–50 The binding of different guest molecules to the host can either enhance or diminish the effective interactions between them, allowing for precise control over molecular recognition.Furthermore, the pronounced electron deficiency of 1(PF6)3 enables its use as a selective organo-oxidant, offering an efficient metal-free platform for oxidative transformations. Its performance in encryption–decryption experiments demonstrates the receptor's capability for recyclable, stimuli-responsive, molecular-level information security. The rapid and reversible generation of charge-transfer signals, combined with the intrinsic chemical reconfigurability of the system, positions 1(PF6)3 as a highly versatile molecular device for secure data storage, visual encoding, and emerging stimuli-gated information technologies.
Author contributions
Rajorshi Das: conceptualization, supervision and original draft preparation; Logeshwari Seethapathy and Rohith M: investigation and data curation; Chinmoy K. Hazra: experimental support; Jayanta Samanta and Rajorshi Das: crystallographic analysis; Susnata Pramanik and Logeshwari Seethapathy: theoretical studies; all authors: writing – review and editing.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5mh02406d.CCDC 2476293–2476297 contain the supplementary crystallographic data for this paper.51a–e
Acknowledgements
The authors gratefully acknowledge the funding support from SRM Institute of Science and Technology (SRMIST) under the Selective Excellence Research Initiative (SERI) (grant no. SRMIST/R/AR(A/SERI2024/174/17-342)). The authors thank SRM-Central Instrumentation Facility (SCIF) and SRM Nanotechnology Research Centre (NRC) for providing essential instrumentation facilities. L. S. and R. M. are grateful to SRMIST for their doctoral fellowship. The authors also thank Dr Anandhakumar S. and Mr Abilash S. for the measurement of electrochemical data.References
- D. D. Boehr, R. Nussinov and P. E. Wright, Nat. Chem. Biol., 2009, 5, 789–796 CrossRef CAS PubMed.
- H. R. Bosshard, Physiology, 2001, 16, 171–173 CrossRef CAS PubMed.
- M. Nocker and P. Cozzini, Curr. Top. Med. Chem., 2011, 11, 133–147 CrossRef CAS PubMed.
- K. A. Wreggett and J. W. Wells, J. Biol. Chem., 1995, 270, 22488–22499 Search PubMed.
- S. Erbas-Cakmak, S. Kolemen, A. C. Sedgwick, T. Gunnlaugsson, T. D. James, J. Yoon and E. U. Akkaya, Chem. Soc. Rev., 2018, 47, 2228–2248 RSC.
- C. Emanuelson, A. Bardhan and A. Deiters, ACS Synth. Biol., 2024, 13, 538–545 CrossRef CAS PubMed.
- J. Krämer, R. Kang, L. M. Grimm, L. De Cola, P. Picchetti and F. Biedermann, Chem. Rev., 2022, 122, 3459–3636 CrossRef PubMed.
- A. Patel and P. Peralta-Yahya, Biochemistry, 2023, 62, 187–195 CrossRef CAS PubMed.
- K.-Y. Wang, J. Zhang, Y.-C. Hsu, H. Lin, Z. Han, J. Pang, Z. Yang, R.-R. Liang, W. Shi and H.-C. Zhou, Chem. Rev., 2023, 123, 5347–5420 Search PubMed.
- M. Scherer, D. L. Caulder, D. W. Johnson and K. N. Raymond, Angew. Chem. Int. Ed., 1999, 38, 1587–1592 Search PubMed.
- K. Kumazawa, Y. Yamanoi, M. Yoshizawa, T. Kusukawa and M. Fujita, Angew. Chem. Int. Ed., 2004, 43, 5936–5940 CrossRef CAS PubMed.
- T. Sawada, H. Hisada and M. Fujita, J. Am. Chem. Soc., 2014, 136, 4449–4451 Search PubMed.
- S. Wang, T. Sawada, K. Ohara, K. Yamaguchi and M. Fujita, Angew. Chem. Int. Ed., 2016, 55, 2063–2066 Search PubMed.
- K. Iizuka, H. Takezawa and M. Fujita, J. Am. Chem. Soc., 2024, 146, 32311–32316 CrossRef CAS PubMed.
- R. Zhu, J. Lübben, B. Dittrich and G. H. Clever, Angew. Chem. Int. Ed., 2015, 54, 2796–2800 Search PubMed.
- A. Walther, I. Regeni, J. J. Holstein and G. H. Clever, J. Am. Chem. Soc., 2023, 145, 25365–25371 CrossRef CAS PubMed.
- S. Sudan, F. Fadaei-Tirani, R. Scopelliti, K. E. Ebbert, G. H. Clever and K. Severin, Angew. Chem. Int. Ed., 2022, 61, e202201823 Search PubMed.
- I. A. Riddell, M. M. J. Smulders, J. K. Clegg, Y. R. Hristova, B. Breiner, J. D. Thoburn and J. R. Nitschke, Nat. Chem., 2012, 4, 751–756 CrossRef CAS PubMed.
- D. M. Wood, W. Meng, T. K. Ronson, A. R. Stefankiewicz, J. K. M. Sanders and J. R. Nitschke, Angew. Chem. Int. Ed., 2015, 54, 3988–3992 CrossRef CAS PubMed.
- C. T. McTernan, J. A. Davies and J. R. Nitschke, Chem. Rev., 2022, 122, 10393–10437 Search PubMed.
- F. J. Rizzuto and J. R. Nitschke, Nat. Chem., 2017, 9, 903–908 Search PubMed.
- R. Banerjee, P. Bhandari, N. Hickey and P. S. Mukherjee, J. Am. Chem. Soc., 2025, 147, 23049–23059 Search PubMed.
- M. Aggarwal, R. Banerjee, N. Hickey and P. S. Mukherjee, Angew. Chem. Int. Ed., 2024, 63, e202411513 CrossRef CAS PubMed.
- H. Lee, P. Elumalai, N. Singh, H. Kim, S. U. Lee and K.-W. Chi, J. Am. Chem. Soc., 2015, 137, 4674–4677 CrossRef CAS PubMed.
- L.-X. Cai, Y.-H. Hu, L.-P. Zhou, P.-M. Cheng, X.-Q. Guo, Y.-T. Chan and Q.-F. Sun, Chem. Sci., 2025, 16, 7956–7962 RSC.
- A. V. Virovets, E. Peresypkina and M. Scheer, Chem. Rev., 2021, 121, 14485–14554 Search PubMed.
- X.-Q. Guo, L.-P. Zhou, S.-J. Hu, L.-X. Cai, P.-M. Cheng and Q.-F. Sun, J. Am. Chem. Soc., 2021, 143, 6202–6210 CrossRef CAS PubMed.
- M. Fujita, M. Tominaga, A. Hori and B. Therrien, Acc. Chem. Res., 2005, 38, 369–378 Search PubMed.
- R. Chakrabarty, P. S. Mukherjee and P. J. Stang, Chem. Rev., 2011, 111, 6810–6918 CrossRef CAS PubMed.
- N. Hafezi, J. M. Holcroft, K. J. Hartlieb, E. J. Dale, N. A. Vermeulen, C. L. Stern, A. A. Sarjeant and J. F. Stoddart, Angew. Chem. Int. Ed., 2015, 54, 456–461 Search PubMed.
- L.-X. Cai, S.-C. Li, D.-N. Yan, L.-P. Zhou, F. Guo and Q.-F. Sun, J. Am. Chem. Soc., 2018, 140, 4869–4876 CrossRef CAS PubMed.
- J. Samanta and R. Natarajan, Org. Lett., 2016, 18, 3394–3397 CrossRef CAS PubMed.
- S. Ibáñez, P. Salvà, L. N. Dawe and E. Peris, Angew. Chem. Int. Ed., 2024, 63, e202318829 CrossRef PubMed.
- R. Taheri-Ledari, J. Rahimi and A. Maleki, Mater. Res. Express, 2020, 7, 015067 CrossRef CAS.
- M. R. M. Izawa, D. M. Applin, L. Norman and E. A. Cloutis, Icarus, 2014, 237, 159–181 CrossRef CAS.
- H. Zhang, Q. Li, Y. Yang, X. Ji and J. L. Sessler, J. Am. Chem. Soc., 2021, 143, 18635–18642 CrossRef CAS PubMed.
- S. D. Dahlhauser, C. D. Wight, S. R. Moor, R. A. Scanga, P. Ngo, J. T. York, M. S. Vera, K. J. Blake, I. M. Riddington, J. F. Reuther and E. V. Anslyn, ACS Cent. Sci., 2022, 8, 1125–1133 CrossRef CAS.
- Y. Shen, X. Le, Y. Wu and T. Chen, Chem. Soc. Rev., 2024, 53, 606–623 RSC.
- W. Zhong, J. Zhang, Y. Lin, S. Li, Y. Yang, W.-J. Wang, C. Si, F. E. Kühn, Z. Zhao, X.-M. Cai and B. Z. Tang, Chem. Sci., 2024, 15, 3920–3927 RSC.
- N. G. Connelly and W. E. Geiger, Chem. Rev., 1996, 96, 877–910 CrossRef CAS PubMed.
- R. Das, M. Linseis, S. M. Schupp, L. Schmidt-Mende and R. F. Winter, Chem. – Eur. J., 2022, 28, e202104403 CrossRef CAS PubMed.
- R. Das, M. Linseis, S. Scheerer, K. Zoller, L. Senft, I. Ivanović-Burmazović and R. F. Winter, Inorg. Chem., 2022, 61, 12662–12677 CrossRef CAS PubMed.
- B. Su, R. P. Nia, F. Li, M. Hojeij, M. Prudent, C. Corminboeuf, Z. Samec and H. H. Girault, Angew. Chem. Int. Ed., 2008, 47, 4675–4678 CrossRef CAS PubMed.
- B. Zhu, B. Cheng, L. Zhang and J. Yu, Carbon Energy, 2019, 1, 32–56 CrossRef CAS.
- V. Mehmeti and M. Sadiku, Computation, 2022, 10, 68 CrossRef CAS.
- S. Ehrlich, J. Moellmann and S. Grimme, Acc. Chem. Res., 2013, 46, 916–926 CrossRef CAS PubMed.
- U. B. Choi, H. Sanabria, T. Smirnova, M. E. Bowen and K. R. Weninger, Biomolecules, 2019, 9, 114 CrossRef CAS PubMed.
- E. Lopez-Fontal, L. Milanesi and S. Tomas, Chem. Sci., 2016, 7, 4468–4475 RSC.
- S. Whitelam, C. Rogers, A. Pasqua, C. Paavola, J. Trent and P. L. Geissler, Nano Lett., 2009, 9, 292–297 CrossRef CAS PubMed.
- T. M. Nguyen and T. N. Truong, npj Comput. Mater., 2025, 11, 372 CrossRef CAS.
- (a) CCDC 2476293: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2p3sf1; (b) CCDC 2476294: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2p3sg2; (c) CCDC 2476295: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2p3sh3; (d) CCDC 2476296: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2p3sj4; (e) CCDC 2476297: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2p3sk5.
| This journal is © The Royal Society of Chemistry 2026 |