 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Self-assembly of phenylalanine oligopeptides: development of transferable bottom-up coarse-grained potentials
Mason Hooten a and
Meenakshi Dutt
a and
Meenakshi Dutt *b
*b
aDepartment of Biomedical Engineering, Rutgers The State University of New Jersey, Piscataway, NJ 08854, USA
bDepartment of Chemical and Biochemical Engineering, Rutgers The State University of New Jersey, Piscataway, NJ 08854, USA. E-mail: meenakshi.dutt@rutgers.edu
First published on 9th March 2026
Abstract
Self-assembled nanostructures comprising Phenylalanine-based oligopeptides are widely utilized in various applications across the domains of electronics, personal care, and biomedicine. Numerous factors, including sequence, determine the morphology of these nanostructures. Computational approaches such as bottom-up coarse-grained (CG) models coupled with the molecular dynamics simulation technique can be used to predict morphologies as a function of the sequence. These approaches efficiently resolve the multiple scales relevant to the self-assembly process while generating CG representations that are consistent with the all-atom representations. However, bottom-up CG models are computationally demanding to develop. This study proposes an approach to develop bottom-up CG models of Phenylalanine oligopeptides that are transferable between sequences to resolve their assembly. The approach builds upon a previously reported bottom-up CG model for the Phenylalanine tripeptide to develop parameters for new degrees of freedom associated with other oligopeptides. The performance of the new CG model is analyzed by comparing simulation results with corresponding results obtained using different versions of the Martini force field. The predictions from the new force field are also compared with available experimental results.
Design, System, ApplicationSelf-assembled nanostructures composed of phenylalanine-based oligopeptides are utilized in various domains, including electronics, biomedicine, pharmaceuticals, and personal care. The extensive use of these nanostructures stems from their multifunctionality and morphological polymorphism, which is influenced by numerous factors, including the oligopeptide sequence. However, the ability to predict the morphology of these self-assembled nanostructures as a function of the sequence remains limited. The development of this predictive capability will significantly reduce experimental effort and costs, thereby advancing numerous domains. This capability can be developed through computational approaches that efficiently bridge scales pertinent to the assembly, packing, and organization of oligopeptides into nanostructures, while simultaneously preserving the chemical details of their molecular conformations. In this study, we propose a bottom-up coarse-graining approach to develop a transferable, multiresolution, coarse-grained model for various phenylalanine oligopeptides to resolve their self-assembly. The hybrid approach employs a structural coarse-graining method for bonded interactions and a variational approach for nonbonded interactions. The approach is employed to enable bridging the scales relevant to the process of self-assembly while simultaneously resolving consistent all-atom and coarse-grained representations of the molecules. The approach proposed and used in this study can be applied to investigate self-assembly in pure and mixed molecular systems across diverse classes of biomolecules and polymers. |
I. Introduction
Self-assembled nanostructures comprising phenylalanine-based peptides are ubiquitous and are utilized in diverse areas, including electronics, biomedicine, pharmaceuticals, and personal care. Experimental studies have reported a variety of phenylalanine-containing oligopeptides self-assembling into ordered nanostructures. Yet, the ability to predict the morphology of the self-assembled nanostructures as a function of the phenylalanine oligopeptide sequence remains limited. The development of this predictive capability will significantly reduce experimental effort and costs. This capability can be developed through computational approaches that efficiently bridge scales pertinent to the assembly, packing, and organization of oligopeptides into nanostructures, while simultaneously preserving the chemical details of their molecular conformations. One such approach is a suitable coarse-grained (CG) molecular model1–42 used in conjunction with the molecular dynamics (MD) simulation technique.There are two overarching philosophies underlying the coarse-graining of molecular representations: top-down and bottom-up.43–54 The top-down coarse-graining approaches will use macroscopic, thermodynamic, and experimental results to parameterize the CG model. Meanwhile, bottom-up CG approaches use atomistic structure and properties to parameterize the CG model. In this study, a bottom-up coarse-graining approach is used to develop a transferable CG force field for di- and tetraphenylalanine (FF and FFFF, respectively) to resolve their self-assembly in an aqueous solution. A hybrid approach, utilizing a structural coarse-graining method for the bonded interactions and a variational approach for the nonbonded interactions, is employed to enable scale bridging while simultaneously resolving consistent all-atom (AA) and CG representations30 of the molecules.
There has been considerable interest in the development of CG models with reduced degrees of freedom that can accurately simulate the self-assembly process across a range of system compositions.55 Coarse-grained models have resolved the process in systems that span a range of chemical variables, including the composition of thermodynamic states,56 local milieu,57 chain composition,58 and peptide sequence lengths.11 The tunable chemical details captured by bottom-up CG models make them attractive for simulating this process in various systems. However, the computational cost of developing such models increases exponentially with the complexity of their representation, force field parameters, and composition of the system.
More recently, Mohammadi et al. reported the development of a bottom-up nonbonded force field representing interactions between the 20 proteinogenic amino acids and a one-site water model.27 Their model optimizes Lennard–Jones parameters using artificial neural network-assisted particle swarm optimization, targeting the atomistic free energy of solvation for each amino acid. The resulting nonbonded force field is used in concert with earlier-derived bonded parameters58 to simulate self-assembly of peptide amphiphiles. Simulations of two different amphiphile systems revealed distinct nanostructure morphologies — fibers versus micelles — that matched their respective experimental observations. This ability of a single parameter set to distinguish between systems represents an improvement over an earlier version of their model. It highlights the value of optimal solvation potentials in studies of solvent-mediated self-assembly.
Extensions to the relative entropy minimization (REM)59 technique have also demonstrated the capacity to reproduce both monomer structure and supramolecular arrangement over a range of system compositions. In the regularized REM (reg-REM) method, Liebl and Voth added a regularization term to capture the binding affinity between protein monomers.60 One of their reg-REM models, designed to capture the binding affinity of CA/SP1 monomers in an 18-monomer lattice, also faithfully simulates the structure of a large virus-like particle comprising 3180 monomers, demonstrating its transferability to a larger system size. In another study, Pretti and Shell developed a procedure that includes a term for tuning the model to recover specific secondary structures.61 Training the model on an ensemble of simulations of short peptides, they found that the model recovered secondary structure characteristics of amyloidogenic sequences not included in the training set.
An earlier study34 developed a bottom-up CG model of FFF with potential energy functions derived from AA reference data. The model was used to predict the formation of self-assembled nanospheres, which agreed with corresponding experimental results. Whereas such models can potentially predict self-assembly pathways, they suffer from high computational costs arising from the generation of the AA reference sample and the routines used for extracting CG potential energy functions (namely, force-matching (FM), Boltzmann inversion (BI), and iterative Boltzmann inversion (IBI)). The rationale underlying the selection of FM to determine CG nonbonded interactions is due to its ability to capture the potential energy surface associated with multibody interactions. The BI approach was chosen to compute the CG bonded interactions because it is well-suited for determining localized interactions and creating potentials that reproduce structural properties.62,63 The adoption of the IBI approach enables the iterative refinement of the CG bonded potentials initially determined using the BI approach. The refined bonded potentials allow better reproduction of the structural properties. Furthermore, bottom-up models are typically designed to reproduce precise structural representations, such that a fully parameterized model cannot be reused for different chemical species.
To mitigate the high costs associated with developing bottom-up CG models, it is desirable that the fully parameterized models can be substantially reused in developing structurally similar models. The ability to reuse model parameters from one structure to create a model of a new structure is typically referred to as transferability. A transferred model, thus, incorporates some of the structure and parameterization details of an earlier model but applies them to a different species. This study proposes a new approach for developing transferable, multiresolution, bottom-up CG models of biomolecules to resolve the dynamics underlying their self-assembly. The approach (1) preserves the nonbonded CG potentials,34 capturing multibody interactions, while (2) refining the CG bonded potentials to accurately resolve the local structure of the molecules. The approach is sufficiently modular that the specific coarse-graining methods used to develop the nonbonded and bonded potentials can be substituted with analogous methods.
In this study, CG models of two peptides – diphenylalanine (FF) and tetraphenylalanine (FFFF) – are developed, validated, and compared with corresponding results from the Martini CG force field. Simulations using the developed models demonstrate self-assembled nanostructures that are distinct from those reported using the FFF model34 and from one another. The results suggest that a judicious process of transfer modeling can reproduce atomistic structural targets for a family of short aromatic peptides. Some portions of the FFF model34 are transferred to the new models, demonstrating a significant reduction in the cost of developing bottom-up CG models. The approach proposed and used in this study can potentially be applied to investigate the process of self-assembly in pure and mixed molecular systems, encompassing biomolecules and polymers of different chemistries. The CG approach can also be adopted to conceive designs and iteratively optimize molecular sequences with target functionalities. For example, folding, binding or assembly. This can be implemented by integrating the approach into a high-throughput workflow that identifies molecular sequences, starting from a candidate pool of sequences or blocks, with specific functionalities.
II. Methods
This section describes the methods used to define and parameterize the new models, followed by the details of the CG simulations conducted using the parameterized bottom-up models. Descriptions of the Martini CG simulations and backmapping efforts used to analyze the simulations' results are also provided. Codes and example input files for running simulations, developing potentials, and performing the analyses presented here are available in a public GitHub repository.64 All molecular visualizations are prepared using VMD.65A. Mapping schemes and CG model
Phenylalanine di- and tetra-peptides are modeled at CG resolution, using new CG models of FF and FFFF (new models are F2 and F4, respectively). Coarse-grained pseudo atoms or CG beads are defined in terms of atomistic chemical structure, as shown in Fig. 1. The peptide backbone is an alternating chain of alpha carbons and amide groups. The aromatic side chains are represented by three-bead rings to approximate the planar nature and orientability that are observed in the atomistic representation of phenyl rings. The peptides are capped with N- and C-terminal beads. | ||
| Fig. 1 Representations of (a) FF, (b) FFF, and (c) FFFF. CG representations are shown as transparent spheres overlayed on the AA representation in licorice. The FFF model, AR3, is inherited from an earlier study34 and transferred to the representations of FF and FFFF to create the F2 and F4 models, respectively. Terminal beads NH3 (blue) and COO (red) appear once in each model. Backbone beads CA (yellow) appear once for each side chain, with interspersed AMD (orange) beads representing amide groups. The PHE representation comprises three beads per side chain, PHA, PHB, and PHC (purple). | ||
The CG force field includes parameters for bonded interactions required to maintain the peptide's conformation, as well as nonbonded interactions between the peptide and solvent beads. Potential energy functions are applied to the bonds, angles, and dihedrals in the bonded structures and the interpeptide, peptide–water, and water–water nonbonded interactions. The potential energy functions are nonparametric and derived from the atomistic reference simulations using the methods described in subsequent sections. Although the atomistic reference force field separates the potential energy's electrostatic and van der Waals components, the CG force field does not explicitly represent electrostatic charge. Thus, pairwise nonbonded interactions are incorporated in a single tabulated energy function that reflects the cumulative effects of electrostatics and van der Waals interactions observed in the AA reference simulations.
The CG models for FF and FFFF utilize the same set of 7 pseudo atom types, as defined initially in the AR3 model,34 as shown in Fig. 1. The F2 model contains 11 pseudo atoms, compared to 43 atoms in the atomistic reference, resulting in a nearly 4-to-1 mapping ratio. The F2 pseudo atoms are connected with a network of 12 bonds, implying 17 angles and 16 proper dihedrals. All bonds, angles, and dihedrals are parameterized with potential energy functions. The F4 model also achieves a nearly 4-to-1 mapping ratio with 21 pseudo atoms compared with 83 atoms in the atomistic reference. The model encompasses parameters for all 24 bonds, 35 angles, and 36 dihedrals.
B. All atom molecular dynamics simulations
For each peptide species, FF and FFFF, MD simulations of AA reference systems at two different peptide concentrations are performed. The low-concentration systems comprise a single peptide, while the high-concentration systems comprise 25 peptides, all dissolved in an aqueous solution. All systems are solvated in explicit water in a 4 nm cubic simulation box with three-dimensional periodic boundaries. Systems are initialized with peptides randomly distributed in the simulation box. The peptides are configured with a net positive charge at the N-terminus and a net negative charge at the C-terminus, consistent with the expected zwitterionic presentation at physiological pH. Because the peptides are charge-neutral, counterions are not required.The AA MD simulations use the Amber99 force field and SPC/E water model. Steepest descent energy minimization and an equilibration MD simulation run in the NVT ensemble are performed before the production MD simulations. The production simulations sample the NPT ensemble and use a leapfrog integrator with a 2 fs time step. The temperature is maintained at 300 K using the Nose–Hoover thermostat, and the pressure is maintained at 1 bar using the Parrinello–Rahman barostat. The Lennard–Jones cutoff is set to 1.2 nm, with long-range dispersion correction applied for energy and pressure. The particle mesh Ewald (PME) method with a cutoff of 1.2 nm is used to calculate the electrostatic interactions. All bonds are constrained with the LINCS algorithm. All simulations in this study — AA and CG — are performed using Gromacs 2018.
C. AR3 - previously developed model of FFF-bonded structure and intermolecular interactions
In an earlier study,34 two CG models of the phenylalanine tripeptide (FFF) were developed and used to investigate its self-assembly using CG MD simulations. Parameterization of the models employed a hybrid strategy, focusing on equilibrium structures and intermolecular forces. First, peptide nonbonded interactions were calculated using FM with exclusions,66,67 targeting forces in a reference atomistic system with a high molar concentration of peptides. Water interactions were derived using IBI.68,69 Parameters for the bonded interactions were then iteratively refined to reproduce structural characteristics associated with conformations of a single peptide molecule dissolved in water. Simulations using both iteratively refined models predicted the formation of nanostructures, which compared favorably with experimental results.The second model in that earlier study represented the aromatic ring of the side chain with three pseudo atoms and was thus designated AR3 (for aromatic ring 3 beads). This study investigates the transferability of the AR3 model for FFF to phenylalanine peptides with altered chain lengths. The F2 and F4 models inherit potentials from AR3, including the entire nonbonded force field and most of the bond, angle, and dihedral potentials. The nonbonded force field, including peptide–peptide, peptide–water, and water–water interactions, remains unchanged from the AR3 model. Only the bonded parameters are updated or iterated, as discussed below.
D. Boltzmann inversion and iterative Boltzmann inversion
The bonded potentials of the F2 and F4 models, which have no pre-existing analog in the AR3 CG model, are initially derived using BI. In BI, a target bonded degree of freedom (DOF) distribution is first obtained by analyzing reference AA trajectories in terms of the CG representation. Distributions from a canonical ensemble follow a Boltzmann distribution, which relates the probability of observing a particular configuration to the system's energy, where lower-energy configurations are observed more frequently. Inverting the Boltzmann distribution equation thus yields a potential of mean force—eqn (1) through (3) describe this relation for bonds, angles, and dihedrals.
 | (1) |
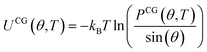 | (2) |
UCG(ϕ, T) = −kBT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ln(PCG(ϕ, T)) ln(PCG(ϕ, T))
| (3) |
Once initial guesses are available for all potential energy functions for the bonded DOFs, the angles and dihedrals are corrected using the IBI scheme. In IBI, trial MD simulations are performed using an initial set of potentials, and DOF distributions are subsequently calculated. The potentials are refined proportionately to the difference between the trial and reference distributions, using the iterative procedure defined in eqn (4).
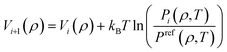 | (4) |
E. AA sampling and potential development
The versatile object-oriented toolkit for coarse-graining applications (VOTCA)66 software contains tools for mapping CG representations, calculating distributions, and developing CG potentials using BI and IBI. VOTCA version 1.6 samples distributions and calculates potentials from atomistic reference MD simulations. Initial parameterizations of new bonded potentials are calculated with BI performed on smooth DOF distributions from an AA MD simulation of a single peptide dissolved in water. IBI is applied successively to correct the resulting bonded potentials, with trial CG MD simulations performed at the same low concentration to produce the initial bonded DOF distributions in AA MD simulations. The IBI test MD simulations are run using the complete transferred force field. The full version of the bonded force field encompassing the lower-order DOFs is typically applied in the simulations testing IBI refinements. Thus, in the case of angle refinements, the inherited bond potentials are also utilized, whereas for dihedral refinement, both the inherited bond and angle potentials are applied.F. Evaluating IBI improvements with earth mover's distance
At each step of IBI, the distributions of the target parameterized DOFs are calculated from the test simulation trajectory, and the results are used to correct the potentials. A scheme targeting site–site pair correlation functions has been proposed to enable an iteratively corrected pair potential to converge, along with associated convergence in the implied conformational ensemble.70 The practical considerations of simulations complicate perfect convergence, for example, in the cost of sampling enough frames to produce smooth DOF distributions. In IBI of angle and dihedral potentials in the peptide models, the quality of the test distributions improves considerably with a small number of iterations before visibly declining. The surmise is that this is due to the accumulation of numerical errors arising from the rough CG potential energy surface. Therefore, since convergence cannot be guaranteed, a method is chosen to quantify the similarity of the target AA distributions and the test CG distributions at each IBI step.The Earth Mover's Distance (EMD), also known as the Wasserstein metric, measures the distance between two distributions. The EMD characterizes how far the masses of two distributions would need to move to resemble one another. A comprehensive analytical description of the EMD can be found in ref. 71 and 72. This study calculates the EMD between each test distribution and its respective atomistic reference after each IBI step. The EMD values for the complete set of IBI DOFs — angles or dihedrals — are summed to yield an error index in the conformational ensemble, with a lower value indicating a lower error. The IBI process is stopped the first time this value increases, and the iterated potentials corresponding to the previous step, i.e., the IBI step, are selected, which results in the first minimum of the conformational error.
G. Bottom-up CG simulations
CG MD simulations of systems encompassing two concentrations of peptides are performed, matching the high and low concentrations of the peptides in the AA reference simulations. The low-concentration (single-peptide) simulations enable the determination of the conformational DOFs for comparison with corresponding results for the atomistic reference. High-concentration systems with 25, 32, 64, and 128 peptides are simulated in CG representation to observe differences in their self-assembly as a function of the number of peptides. The cubic simulation box is scaled to maintain a constant molarity among the high-concentration systems.The F2 and F4 models are parameterized using tabulated potentials transferred from the AR3 force field, modified as discussed in the results and discussion section, with a potential energy cutoff of 1.2 nm. Initial configurations in each CG simulation are produced by mapping a frame from an equilibrated AA reference trajectory to its CG representation. The CG MD simulations are run in a cubic box of constant volume, sized to match the equilibrated box volume in a corresponding AA reference simulation. The CG MD calculations use a leapfrog stochastic dynamics integrator with a 1 fs time step. Temperature is maintained at 300 K using an inverse friction coefficient of 0.2 ps.
H. Martini 2 and 3 coarse-grained molecular dynamics simulations
The Martini framework is a versatile CG force field, comprising a library of pre-specified bead types and prototypes that represent the bonded structures of biomolecular building blocks for which the system is parameterized. While Martini began as a force field for lipid bilayer simulations two decades ago, major revisions since then have expanded the set of pseudo atoms and parameters to target a growing number of applications in biomolecular simulations. Martini 2 introduced a protein force field to govern amino acid interactions, allowing simulations of peptide aggregation. The most recent version, Martini 3, was released in 2021 and provides toolkits for constructing biomolecules based on the structures of amino acids, phospholipids, and nucleic acids.The development of Martini as a general tool for biomolecular simulation has, of course, come with challenges and tradeoffs. One such tradeoff is that the current version of Martini, the most flexible to date, has been refined to reduce spurious aggregation (the famous “stickiness” problem). Still, at the same time, it has become less capable of accurately simulating the self-assembly of short peptides. Earlier studies using Martini 2 reported a variety of nanostructures in homogeneous systems of FF in aqueous solution, including tubes, vesicles, and bilayers.5 A recent systematic investigation into the evolution of Martini as a framework for the self-assembly of short peptides also found that Martini 2 simulations using FF predicted the formation of spherical or elongated vesicles. In contrast, FF simulations with Martini 3 did not yield identifiable nanostructures.73
For this study, simulations of FF and FFFF were conducted using Martini 2 and Martini 3 to compare the results with those from the bottom-up models. Initially, simulations were performed using Martini 3 to examine the formation of peptide nanostructures in the most recent iteration of the force field. The results demonstrated that the behavior of the Martini 3 models was not consistent with earlier versions of Martini, such as Martini 2.2.
Systems of 32 peptides FF or FFFF were prepared in a cubic simulation box with the exact initial dimensions as the AA simulation. The Martini 2.2 simulations are solvated with a 90/10 split of water and antifreeze particles. The Martini 3.0 simulations use the initial release water model. Simulations use the Martini 2.2 and 3.0.0 protein parameter sets, respectively.
The cubic simulation boxes, with three-dimensional periodic boundary conditions, are sized to match the peptide concentration of the high-concentration bottom-up CG MD simulations. Peptides are mapped to the Martini CG representation using the martinize2 program. The peptide molecules are randomly dispersed in the simulation box along with the water beads for the initial configuration. A short equilibration simulation is performed before production MD simulations.
The Martini 2 and Martini 3 production simulations are parameterized following the guidance of reference.74 The simulations run for 40 to 80 million iterations using a 25 fs time step (corresponding to a nominal simulation time of 1 to 2 microseconds) and a leap-frog integrator. The temperature is set to 300 K using the velocity rescale thermostat with a stochastic term,75,76 and the pressure is set to 1 bar using the Berendsen barostat.77 The Lennard-Jones potential uses a switch tapering to a cutoff of 1.2 nm. Reaction-field electrostatic interactions with a potential shift are calculated with a dielectric permittivity of 15 and a cutoff of 1.2 nm.
I. Backmapping and backmapped-atomistic (BA) simulations
The backward method of Wassenaar et al. is adapted to project AA coordinates onto CG coordinates, creating a backmapped atomistic configuration. The backmapped configuration undergoes energy minimization, excluding nonbonded interactions, followed by another minimization including nonbonded interactions. These steps are followed by a series of position-restrained NVT simulations with time steps progressing from 0.2 fs to 2 fs. A short equilibration MD simulation produces a stable backmapped AA configuration. This is followed by the production MD simulation encompassing 10 million iterations using a time step of 2 fs, in the NPT ensemble.III. Results and discussion
The new F2 and F4 models inherit their nonbonded force field from the previously developed AR3 model. The AR3 model includes potentials for the bonds, angles, and dihedrals related to the peptide-bonded structure. Thus, the new models required only new bonded potentials to be developed, where the F2 and F4 bonded structures deviated from that of AR3.A. Bonded potentials
Initial potentials are calculated via BI using a distribution function corresponding to each new DOF. An AA MD simulation of a single peptide in water is run for 500 ns with a time step of 2 fs and a sampling rate of 2 ps. The resulting sample of 250![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 frames calculates the CG-mapped distributions of each new bonded interaction for each CG model. These distributions are inverted via BI, completing the intrapeptide potentials needed to maintain the conformation of a single peptide in CG MD simulation.
000 frames calculates the CG-mapped distributions of each new bonded interaction for each CG model. These distributions are inverted via BI, completing the intrapeptide potentials needed to maintain the conformation of a single peptide in CG MD simulation.
The CG test simulations for IBI are run for 10 million steps with a 2 fs time step and sampled at 2 ps intervals, resulting in a sample of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 frames per iteration. At each iteration, the distributions of the DOFs corresponding to the complete set of target potentials -- angles or dihedrals -- are evaluated and compared with distributions from reference AA trajectories. While the first few steps of IBI generally improve the similarity of the CG and AA distributions, there is a point at which subsequent IBI steps increase the error. This failure to converge is attributed to computational limitations and the relatively large number of concurrently parameterized DOFs in the model. To select a set of iteratively refined potentials, after each IBI step, the EMD is calculated between the CG test and AA reference distributions for each conformational DOF (angle or dihedral), and the EMD values are summed over the set of DOF potentials being resolved (all angles or all dihedrals).
000 frames per iteration. At each iteration, the distributions of the DOFs corresponding to the complete set of target potentials -- angles or dihedrals -- are evaluated and compared with distributions from reference AA trajectories. While the first few steps of IBI generally improve the similarity of the CG and AA distributions, there is a point at which subsequent IBI steps increase the error. This failure to converge is attributed to computational limitations and the relatively large number of concurrently parameterized DOFs in the model. To select a set of iteratively refined potentials, after each IBI step, the EMD is calculated between the CG test and AA reference distributions for each conformational DOF (angle or dihedral), and the EMD values are summed over the set of DOF potentials being resolved (all angles or all dihedrals).
The set of potentials associated with the first minimum of the summed EMD is selected for each IBI run. In the F2 model, total angle EMD reaches its minimum after initial BI plus four rounds of iteration. Applying these angle potentials, total dihedral EMD reaches its first minimum after BI plus one iteration. In the F4 model, angle potentials are chosen after BI plus five iterations, and dihedrals are selected after BI plus one iteration.
B. Individual peptide structure
The fully parameterized F2 and F4 models are each used to simulate a single peptide solvated in water. Distributions are calculated for the parameterized DOFs as well as for two unparameterized measures of overall conformation, the end-to-end distance and radius of gyration (Rg). In the case of F4, the rotameric arrangement of consecutive side chains is also evaluated. The conformational results for the F4 model are discussed in the following sections, with an equivalent treatment of the results for the F2 model available in the SI.Angles in F4 also typically match the sampled ranges of the atomistic reference, with some deviations. As with F2, the F4 angles describing the extension of the side chains away from the peptide backbone are somewhat biased but take on values generally within the range sampled by the AA reference. However, the backbone angle bridging the first and second amide groups across the second backbone alpha carbon (AMD1-CA2-AMD2) samples configurations from 1.5 to 2.0 rad, suggesting a slightly collapsed conformation in the CG simulations compared to the AA reference (see SI Fig. S7). Although IBI refines the potential in this angle, the initial estimate is inherited from the AR3 model in an earlier study.34 The reference AA distribution was used to develop the original samples in the 1.5 to 2.0 rad range; thus, the AR3 potential function achieves a local minimum. Despite several iterations of IBI, the propagation of these configurations to the F4 model highlights the importance of the initial parameterizations on the eventual shape of refined potentials.
As end-to-end distance and Rg will be influenced by nonbonded interactions, comparing these quantities between highly concentrated AA and CG systems, where nonbonded interactions are fully present, is examined. The end-to-end distance in the AA and CG representations of the 25 peptide system samples falls within the same range and shares some peak locations. Although the atomistic curve exhibits four distinct modes in the short-range sampling, the CG curve displays only two modes. This is expected, as the nonbonded interactions are resolved at the same high concentration as the 25 peptide systems. The bonded DOFs are refined using these nonbonded interactions; thus, the single peptide conformation may be interpreted as biased in their absence.
The rotameric character of individual peptides and those in the aggregate for the four models of FFFF simulated – AA, F4, Martini2, and Martini3 – was calculated. The simulations of monomers using the bottom-up F4 force field show rotamer distributions vary along the chain, indicating the incorporation of specific conformational details gleaned from the AA reference simulations. In comparison, the Martini simulations typically show a random relationship, indicating nonspecific behavior of the side chains. The F4 simulations of aggregates also show a comparative spreading of the rotamer distributions. This flexibility is reminiscent of that seen in the AA reference and is key to the formation of the branched aggregates shown in Fig. 4. In comparison, the distributions from the Martini aggregates become sharper, reflecting their apparent tendency to snap sharply into place in the bilayer-like aggregates shown in Fig. 5.
To examine the tendency of the side chains, their relative orientation (namely, isomerization or rotamerization) in a single peptide is measured. The coordination between adjacent side chains in the tetramers is calculated as the dihedral angle measured from the proximal bead in each side chain across their respective backbone beads. A value of 0 in this calculation corresponds to side chains projecting in the same direction as the backbone, i.e., a cisoid conformation. In contrast, a value of pi describes side chains which are diametrically opposite, i.e., transoid.
The parity of the CG representations in the F4 and Martini models naturally allows for direct comparison, where the differences between the two models (namely, Martini backbones do not specify explicit amide groups) will not impact measurements of the relative rotations. This feature is also calculated in simulated systems of multiple peptides, including representative aggregates from CG simulations.
Cisoid rotamerization is observed among the second, third, and fourth side chains in simulations of a single peptide using the AA, the bottom-up CG, and the Martini models, as seen in Fig. 2. However, the models differ in their results for coordination between the first and second side chains. The parity of the CG representations in the F4 and Martini models allows for direct comparison. The differences between the two models (namely, amide groups are not explicitly specified in Martini backbones) do not impact measurements of the relative rotations. This feature is also calculated in simulated systems of multiple peptides, including representative aggregates from CG simulations.
In the AA representation, an isolated peptide in water exhibits a transoid arrangement of the first two side chains (SC1-SC2) and a cisoid arrangement of the remaining three side chains (SC2-SC3 and SC3-SC4). The three rotameric measurements are more flexible in simulations with multiple peptides, where clear transoid arrangements also appear among the last three side chains —a feature that does not appear in any of the multi-peptide CG simulations.
An individual peptide simulated with the F4 model adopts a different SC1-SC2 arrangement, with the most probable orientation centered around pi/2. Similar to the more straightforward transoid arrangement of the AA model, this allows for the proximity of the side chain to the backbone, resulting in hydrophobic shielding. This level of shielding may be unavailable in a cisoid arrangement, where the side chains would become crowded and sterically interfere with each other. This effect may modulate the excluded volume of the CG beads involved. In aggregates resulting from the F4 model, all rotamers are cisoid. This is consistent with an intuitive picture of the backbone maximizing the shielding of the hydrophobic side chains by burying them in the aggregate's interior.
For the Martini models, all rotamer distributions have a single mode at 0. For single-peptide simulations using Martini 2, a distinct central peak is observed. In contrast, the single-peptide distributions for Martini 3 appear essentially uniform, albeit with a slight increase in frequency over the range of [−1, 1]. For the multiple-peptide simulations, the distributions for Martini 2 are centered around zero much more sharply with no transoid probability. The multiple peptide systems using Martini 3 also demonstrate a more pronounced cisoid mode, with some likelihood of transoid arrangements. Furthermore, the Martini 3 multi-peptide distributions are noticeably smoother than those of others and appear almost Gaussian in shape. Given the predominance of cisoid rotamerization in the Martini models of the peptides, it is essential to note that these models do not apply a potential energy function to the backbone dihedrals. Although the Martini models include zwitterionic potentials, charges are assigned as either +1 or −1 on the N- and C-terminal backbone beads, respectively. Thus, the observed configurations primarily arise from intermolecular interactions with water, with polar interactions modulating the intermolecular arrangement and reinforcing the cisoid tendency.
C. Systems of multiple peptides
Systems of multiple peptides simulated with the F2 and F4 models aggregate into ordered nanostructures. Systems of 25, 32, 64, and 128 peptides were simulated with both bottom-up models, all with an equal concentration of peptides. Ten independent trajectories (using different seeds) were run for systems of 32 and 64 peptides, along with a single trajectory for the 25- and 125-peptide reference systems.Structures formed in the F4 simulations are considerably more diverse, including prolate droplets, cylindrical rods, wavy rods, and branched structures. As with simulations using F2 and the earlier AR3 models, side chain beads are typically shielded at the interior of the aggregates. The F4 nanostructures are solid, allowing no water to intrude among the packed side chain beads. The minimum (transverse) diameters of the various structures are 2 to 3 nm. The structure of the cylindrical rods differs from that observed in the earlier AR3 simulations. The AR3 rods consisted of planar layers comprising three peptides each, stacked in a longitudinal direction. The rods formed using the F4 model do not create such distinct layers. Instead, the peptides are randomly oriented, shielding the side chains in the aggregate and exposing the main chain to the aqueous solvent. The 128 peptide system resolves a single branched network with continuous connections across the periodic boundaries in all directions.
The ten Martini 3 simulations comprising 32 FF peptides in aqueous solution yielded no ordered nanostructures. A representative final state of the FF systems is visualized in Fig. 5(a). In the simulations of 32 FFFF peptides, final states typically showed a stable lamella with side chains on the interior. The charged terminal beads are alternating on the upper and lower surfaces. A representative final state of the FFFF systems is visualized in Fig. 5(b) and (c).
All ten simulations using the Martini 2 model for FF resulted in rodlike morphologies, with the charged terminal backbone beads clustering on the surface. These charged beads are sometimes bunched up, visually suggesting an upper and lower layer to the aggregate, with side chain beads in the middle layer. The rod appears twisted in others, and the charged beads are more evenly distributed over the whole surface. All ten simulations using the Martini 2 model for FFFF uniformly produced layered morphologies in which consecutive peptide backbones are arranged antiparallel with side chain beads fanned out toward the interior of the layer. The consistent and close arrangement of the charged terminal beads yields a layer that appears striped, with backbone beads exposed to the solvent. Typical structures are shown in Fig. 6.
The bottom-up CG simulations for FF demonstrate the formation of spherical micelles, or nanospheres, which represent a potential transient state in the self-assembly pathway leading to the formation of nanorods, as captured by simulations using the Martini 2 model for FF. These results deviate significantly from corresponding results obtained using the Martini 3 model for FF. The formation of branched fibrils, as captured by bottom-up CG simulations for FFFF, differs from the self-assembled nanostructures observed using the two Martini models for FFFF.
As an aside, the bilayers formed in the Martini FFFF simulations can be presumed to be precursors to the formation of layered nanoplates. However, the fact that bilayers are the only morphology represented may indicate that assembly dynamics are driven by the hydrophobicity of the side chains and the length of the peptides, which allow for a preferential arrangement of the charged backbone terminals. In the Martini model, the peptides also lack explicit representation of the amino groups, such that the polar interactions of those groups and their associated effects on the formation of secondary structures cannot be determined. It is noted that these Martini simulations utilized amino acid representations generated using the defaults of the martinize2 program. Modifications to the model, including manual charge distribution, modulation of water self-interaction, and the addition of a dihedral potential, may improve the results.
The system sizes in this study are relatively small, raising the possibility that finite-size effects influence the final nanostructures and their dynamics of assembly. This may be the case despite the mitigating strategy of using potential cutoffs less than half the length of the simulation box (a common approach, in observance of the minimum image convention in MD simulations79). Regarding the hypothesis that the observed dynamics of assembly represent essential characteristics of the peptide chemistry, a favorable comparison of the particle RDFs between the CG and AA systems (Fig. S5 and S11) is observed. This comparison suggests that the CG force field accurately captures nonbonded interactions. Also, the dynamics of assembly are observed to be stable across system sizes (Fig. 9). And while the number of peptides in the simulations – both the raw numbers and the concentrations – suggests the need for discretion when interpreting the results in the context of macroscopic bench experiments, the stability of the dynamics and structures is found to be a positive indicator of the applicability of the CG models.
FF systems. The AA MD simulation of 32 FF peptides shows many contacts between the side chains of the first residue and between those of the first and second residue. As expected, the N-terminal is frequently in contact with C-termini, but there are also many contacts between the N-terminal and the backbone adjacent to the second side chain. The interpeptide contact of the C-terminal is relatively uncommon. The simulation of 64 peptides reveals similar patterns in the proportions of contacts, except for a significantly larger proportion of contacts between the side chains of the second residues.
In the systems using the F2 model, the 32 and 64-peptide maps are nearly identical. Most contacts occur between beads of the first side chains with other first and second side chains, with fewer contacts between beads of the second side chains, a pattern similar to that of the 32-peptide AA system. The CG systems demonstrate practically no contacts involving the terminal beads. The terminal beads are pushed to the micelle surface and excluded from close contact by the backbone curvature induced by packing the side chains in the micelle interior.
In the BA maps, as in the AA maps, the N-terminal makes a moderate number of contacts with the C-terminal and backbone at the second side chain. This constitutes a rebound in these contacts after they disappeared in the CG simulation, but their numbers are in a smaller proportion than in the reference AA system. The BA system also shows contacts highly concentrated on the side chains, but now more evenly distributed among the possible configurations (i.e., intermolecular contacts between the first side chain on each monomer, the first and second side chains, and the second side chains).
The contact maps for systems of 32 and 64 peptides in the AA representation differ. For example, the 32 peptide map shows higher contact counts on the first side chain, while the 64 peptide map shows higher contact counts on the second side chain. The contact maps calculated from CG trajectories for the two system sizes are substantially similar. This is expected as the CG MD simulation for both system sizes predicted identical nanostructure morphology. However, the dissimilarity between the two system sizes in AA representation suggests an underlying behavior that the parameterization of the CG model has captured, which is not visible in either of these long AA simulations. It should be noted that the CG model converges to these stable structures for both system sizes in only tens of nanoseconds of simulated time.
The BA simulations allow the backmapped CG morphology (i.e., a set of micelles) to be relaxed by the atomistic force field. This relaxation occurs due to a combination of detailed peptide reconfiguration and increased spatial resolution, allowing the atoms in the peptides to interact more smoothly over the length of the chain. The 32- and 64-peptide BA contact maps show some differences, but they appear substantially similar nonetheless. This is despite notable dissimilarity between the respective reference AA systems. It is therefore surmised that the equilibrium configurations predicted by the CG model provide plausible conformations and arrangements of molecules within the predicted nanostructures.
FFFF systems. Comparisons between the contact maps calculated from the simulations of FFFF peptides are similar to those seen in the FF peptide systems. The results for the AA trajectories for systems of 32 and 64 peptides are appreciably dissimilar, while those computed from the CG trajectories are highly similar. The BA systems show similarity in regions where the CG simulations show dominant contacts. The BA systems appear more similar to one another than the AA systems. However, the degree of this apparent similarity is difficult to gauge from visual inspection of the contact map. Indeed, even for these relatively short peptides, the bead-scale resolution makes for a contact map that resists straightforward visual interpretation.
To overcome the difficulty of direct interpretation among contact maps with different gross sample counts, the virtual two-dimensional free energy surfaces (FES) implied by the contact maps are compared. First, the contact maps are Boltzmann inverted to yield a two-dimensional FES. Then, using a two-dimensional implementation of the EMD algorithm, the EMD between pairs of FES is calculated at each simulated resolution and across two system sizes. This approach is adapted from an earlier study by Hunkler et al.,80 who employed a similar method to evaluate backmapping-based sampling by comparing FESs from AA, CG, and backmapped AA simulations of ASP and GLU oligomers. A lower EMD indicates greater similarity between the two sampled FESs. Comparisons of FES similarity among systems as measured by two-dimensional EMD is shown in Fig. 8.
 | ||
| Fig. 8 EMD values are calculated to compare the two-dimensional FES obtained by Boltzmann inversion of the contact maps shown in Fig. 7. Lower values indicate closer concordance between FES, higher values indicate divergence between FES. (a) FES are compared from AA, CG, and BA simulations in systems of 32 and 64 peptides, for peptide sequences FF and FFFF. The FES implied by AA and BA simulations are typically more similar than those implied by CG simulation. (b) FES are compared between the systems of 32 and 64 peptides at each simulated resolution, for peptide sequences FF and FFFF. FES implied by CG simulations at different system sizes are more divergent in simulations using the F4 force field than those using the F2 force field. | ||
Fig. 8(a) shows that for both peptide chain lengths, the EMD values between CG and AA and those between CG and BA are much larger than those calculated between AA and BA. This is unsurprising since the AA and BA simulations proceed under the same force field, and thus their molecules should sample similar regions. However, given that the nanostructures represented by the BA contact maps inherit some contact propensity from the evolution of the CG simulations, the fact that the AA and BA FES match well demonstrates that the molecular arrangements resolved by the CG MD simulations are, at a high level, consistent with the equilibrium conditions captured using the atomistic force field. Fig. 8(b) compares FES from the nanostructures in the 32 and 64 peptide systems for each peptide sequence, thereby gauging the consistency of the FES associated with the force fields used at each resolution across system sizes.
With that caveat, interaction count time series typically look linear at the early stages before transitioning to exponential behavior. Comparing the evolving structure during aggregation will help identify the underlying dominant processes. The dynamics of the evolving structure can be approximated as linear to understand the relative rates of change of interaction counts. This approximation enables the calculation of an intercept (minimum count, available from single-peptide simulations) and an appropriate scaling factor. In this study, the counts are scaled by the maximum possible counts, i.e., where all available particles overlap (a forbidden state under a typical MD model).
By observing the relationships between peptide clustering and counts of mutual interpeptide interactions, one can develop an understanding of the dynamics underlying the formation of nanostructures. Fig. 9 illustrates the evolution of bead-wise contacts calculated for ensembles of 32 and 64 peptides, using FF and FFFF with the F2 and F4 models, respectively. Fig. S15 shows that the relative proportions of contacts established in the first 2.5 million simulation steps remain stable through to 100 million steps. Fig. 10 depicts the average aggregate size in each ensemble, scaled by the number of peptides in the simulation, to illustrate the proportion of system peptides represented by the average aggregate size over time. This framing clearly shows in the F2 systems a tendency to form aggregates comprising only a fraction of the total peptides in simulation, compared to the F4 systems, which tend to form a single aggregate involving all peptides. Alternative framings of this tendency are shown in Fig. S16.
The FF simulations typically resolve the formation of multiple micelles within 5 ns of simulated time. These micelles do not assemble into a single nanostructure when the simulations are run for an extended duration. Fig. 9 shows that early aggregation in both systems is characterized by a rapid increase in interactions between side chains, indicative of the shielding process of the side chains due to the hydrophobic effect. Interactions between the complementary terminal groups gradually increase during the simulations, reflecting their progressive organization on the surfaces of the individual micelles. Interactions involving main chain beads remain at their minimum values throughout the FF simulations, since the tightly packed side chains bar the main chain beads from interacting with the interiors of the micelle, and most micelles do not approach each other. Interactions between amide groups are essentially zero throughout both simulations. Hence, the process of micelle formation in these simulations is driven almost entirely by the hydrophobic effect, with nominal influence of the polar groups on the evolution of the micelle surfaces.
In the simulation with 32 FFFF peptides, the system forms a single straight nanorod within 1 ns of simulated time. As in the FF simulations, early aggregation exhibits a rapid increase in interactions between side chains, which occurs much more quickly than other bead interactions. Interactions between amide groups increase gradually, followed by interactions involving main-chain beads. Complementary terminal beads have essentially no interactions. Therefore, the nanorod formation in this simulation is initially driven by the hydrophobic effect, with subsequent long-range organization into a nanostructure.
The simulation of 64 FFFF peptides revealed the formation of a highly networked three-dimensional nanostructure, where the branches are typically 2 nm in diameter, consistent with the results for the 128-peptide system shown in Fig. 4(c). The rapid increase in side chain interactions mirrors that of the 32-peptide system. However, in the 64 peptide system, the amide group interactions also rapidly increase at the same pace as the side chains, with a slight lag in main chain interactions. This relatively rapid spatial organization of the backbones in the 64 peptide system, driven by interactions between the amide groups, signals the simultaneous formation of multiple proto-rods similar to those observed in the 32 peptide system, which form throughout the simulation box and coalesce into a network-like nanostructure.
Mayans and Alemán83 have compiled data on self-assembly in di-, tri, and tetraphenylalanine with and without protective capping groups, and in a variety of solvents ranging from apolar to polar, summarized in the reprinted Fig. 11. In the highest-concentration aqueous solutions, the authors report that uncapped FF and FFFF peptides assemble into nanotubes, microtubules, and fibrils. Mayans and Alemán also note the influence of solvent polarity and terminal capping on the aggregate morphologies. Under various conditions, the morphological landscape of FF solutions grows to include plates, ribbons, and toroids, while FFFF is also seen to produce branching, flower-like, and braid-like structures.
 | ||
| Fig. 11 Summary of assemblies formed by phenylalanine di-, tri, and tetra-peptides as a function of solvent polarity and peptide concentration: (a) uncapped and (b) capped at both the N- and C-terminus with fluorenyl groups. Reprinted from ref. 83. | ||
There is appreciable similarity in the morphological diversity of the simulated FFFF nanostructures with the corresponding experimental results. Specifically, the networked or branched fibrils in the FFFF simulations are consistent with experimental observation under equivalent solution conditions. It is surmised that the appearance of such structures in the simulations reflects a decreased influence of the charged terminal groups in analogy to the charge shielding provided by capping groups in experiments. This decrease in influence may be due to various factors, including charge distribution over the atoms of the reference AA peptides and within the CG representation.
In the case of FF, the early phase of aggregation of the FF peptides typically begins with the formation of large incoherent aggregates before forming distinct octamers, as demonstrated by the time evolution of the average number of molecules per cluster. The formation of these predicted nano-micelles is a plausible transient state in the pathways to the formation of larger equilibrium morphologies reported in experiments.85 However, the simulations do not demonstrate the formation of larger nanostructures and therefore appear to deviate from the experimental results. This discrepancy is likely due to a combination of physical effects and limitations of the CG methodology. It is noted that the simulated concentrations are set at a high value to exceed the (unknown) critical micelle concentration and ensure aggregation, a common strategy in simulations investigating aggregation.1,5,11,20,34,86–90 The simulated peptide concentrations in this study (200–400 mg mL−1) are two to four orders of magnitude higher than the range of values reported by Mayans and Alemán. The solvent content of the simulations is therefore much lower, reducing the entropic contributions of water to the formation of the aggregates. This mitigation of entropic forces may partially explain why the micelles do not collapse into more densely packed aggregates.
It may also be that the CG model is too coarse34 to represent the chemical details required for close packing. Positing once again that micelles are genuinely on the continuum of concentration-dependent morphologies, the CG force field may be too smooth to allow relaxation into closely packed nanostructures. Experiments show that the supramolecular organization of phenylalanine dipeptides is an Ostwald process, that is, a process in which self-assembled aggregates partially dissociate, with the solution progressively “ripening” into a state where lower energy aggregate morphologies dominate.91 In this case, the micelles may represent a relatively high-energy early aggregation state, but with model conditions that do not permit the required partial dissociation. These arguments suggest improvements to the model, such as increasing the resolution of the representation. Furthermore, the detailed dynamics of the self-assembly process may reflect the physicochemical properties of high-concentration solutions, whose behavior may differ fundamentally from that of much lower in vitro concentrations used in experiments.92
IV. Conclusions
A new approach is proposed for developing transferable, multiresolution CG force fields that resolve the dynamics underlying the self-assembly of short phenylalanine oligopeptides. As part of the study, two new bottom-up CG models representing di- and tetra-phenylalanine oligopeptides have been developed. Each model was partially transferred from an earlier CG model of triphenylalanine, including the bonded structure and nonbonded interactions. The F2 model faithfully recapitulated all structural targets of the atomistic reference simulation sets. The F4 model accurately recovered bonds and angles, but exhibited notable deviations from the atomistic reference, particularly in dihedral angles. However, the F4 model partially resolved side rotamerization characteristics, which are highly relevant to molecular packing.Predicted self-assembled nanostructures encompassing FFFF reflect polymorphism consistent with prior experimental studies. The F2 model's predictions of the early phase of aggregation are in agreement with corresponding experimental results. The nanostructures predicted in the F2 model beyond this phase appear to deviate from those reported in experiments. However, it is surmised that concentration effects and a highly coarse representation of FF may result in structures associated with transient states in the aggregation process.
Comparable peptide systems were also simulated using two versions of the popular Martini model, allowing a performance comparison on structural and computational metrics. While Martini has an advantage over bottom-up models in computational performance, the latter can rapidly converge on equilibrium structures with reasonable accuracy, and future performance upgrades can be used to resolve larger systems.
These results are by no means an indication that Martini is not a suitable framework for investigating short peptide aggregation. For one, the total counts of peptides in the simulations presented here are quite small for Martini, which is routinely used to simulate systems hundreds or thousands of times larger. The Martini ecosystem also supports a variety of structural constraints, and several targeted refinements of the force field have been described. Rather, the comparison simply shows that a bottom-up model trained on modest reference data can resolve fine-scale system dynamics that are fundamentally distinct from those seen in a very popular top-down model.
This study demonstrates that chemically specific models may be transferred and judiciously refined to simulate various systems within a family of aromatic peptides. Notably, these CG models are parameterized solely with nonbonded interactions, as well as 2-, 3-, and 4-body bonded interactions, and the parameterizations are readily interpreted in terms of reference structures. These CG models are computationally efficient in scale bridging while providing a representation that is consistent with corresponding AA representations. These attributes enable the CG models to capture multiscale processes such as self-assembly while simultaneously generating chemically accurate molecular conformations on demand. Currently, efforts are underway to automate the demonstrated hybrid CG strategy, thereby increasing efficiency in the development process to a level comparable to that of machine learning-enabled CG strategies. The CG approach discussed in this study could be potentially adopted to investigate multiscale processes involving multiple molecules in pure or mixed molecular systems, encompassing biomolecules or polymers of different chemistries. Furthermore, the CG approach can be adopted to propose designs and iteratively optimize molecular sequences with specific functionalities, such as folding, binding, or assembly. To this end, the approach can be integrated into a high-throughput workflow93 that identifies specific molecular sequences, starting with candidates of sequences or blocks, with target functionalities.
Author contributions
Mason Hooten: data curation (lead), formal analysis (lead), investigation (lead), methodology (lead), software (lead), validation (lead), writing—original draft preparation (lead), visualization (lead), writing—review and editing (equal). Meenakshi Dutt: conceptualization (lead), supervision (lead), project administration (lead), funding acquisition (lead), resources (lead), writing—review and editing (equal).Conflicts of interest
There are no conflicts of interest to declare.Data availability
Codes and example input files for running simulations, developing potentials, and performing the analyses presented here are available in a public GitHub repository [https://github.com/duttm/Hybrid_Bottom-Up_Coarse-Grained_Model_for_Aromatic_Peptides (accessed 2026-02-22)].Supplementary information (SI): containing structural distribution plots, alternative clustering profiles, performance evaluation of the CG systems, and supplementary model description. See DOI: https://doi.org/10.1039/d5me00207a.
Acknowledgements
The authors acknowledge financial support from the NSF CAREER DMR-1654325, DMREF DMR-2118860, and OAC-1835449 awards. Portions of the studies presented have used computational resources via NSF ACCESS (allocation DMR-140125).References
- F. Aydin and M. Dutt, Bioinspired Vesicles Encompassing Two-Tail Phospholipids: Self-Assembly and Phase Segregation via Implicit Solvent Coarse-Grained Molecular Dynamics, J. Phys. Chem. B, 2014, 118(29), 8614–8623, DOI:10.1021/jp503376r.
- L. Chong and M. Dutt, Design of PAMAM-COO Dendron-Grafted Surfaces to Promote Pb(II) Ion Adsorption, Phys. Chem. Chem. Phys., 2015, 17(16), 10615–10623, 10.1039/C5CP00309A.
- F. Aydin and M. Dutt, Surface Reconfiguration of Binary Lipid Vesicles via Electrostatically Induced Nanoparticle Adsorption, J. Phys. Chem. B, 2016, 120(27), 6646–6656, DOI:10.1021/acs.jpcb.6b02334.
- L. Chong, F. Aydin and M. Dutt, Implicit solvent coarse-grained model of polyamidoamine dendrimers: Role of generation and pH, J. Comput. Chem., 2016, 37(10), 920–926 CrossRef CAS PubMed.
- S. Mushnoori, K. Schmidt, V. Nanda and M. Dutt, Designing Phenylalanine-Based Hybrid Biological Materials: Controlling Morphology via Molecular Composition, Org. Biomol. Chem., 2018, 16(14), 2499–2507, 10.1039/C8OB00130H.
- A. Banerjee, A. Tam and M. Dutt, Dendronized Vesicles: Formation, Self-Organization of Dendron-Grafted Amphiphiles and Stability, Nanoscale Adv., 2021, 3(3), 725–737, 10.1039/D0NA00773K.
- A. Garaizar and J. R. Espinosa, Salt dependent phase behavior of intrinsically disordered proteins from a coarse-grained model with explicit water and ions, J. Chem. Phys., 2021, 155, 12503 CrossRef PubMed.
- J. A. Joseph, A. Reinhardt, A. Aguirre, P. Y. Chew, K. O. Russell, J. R. Espinosa, A. Garaizar and R. Collepardo-Guevara, Physics-driven coarse-grained model for biomolecular phase separation with near-quantitative accuracy, Nat. Comput. Sci., 2021, 1(11), 732–743 CrossRef PubMed.
- K. H. Shen, M. Fan and L. M. Hall, Molecular dynamics simulations of ion-containing polymers using generic coarse-grained models, Macromolecules, 2021, 54(5), 2031–2052 CrossRef CAS.
- T. Sun, V. Minhas, N. Korolev, A. Mirzoev, A. P. Lyubartsev and L. Nordenskiöld, Bottom-up coarse-grained modeling of DNA, Front. Mol. Biosci., 2021, 8, 645527 CrossRef CAS PubMed.
- A. Banerjee, C. Y. Lu and M. Dutt, A Hybrid Coarse-Grained Model for Structure, Solvation and Assembly of Lipid-like Peptides, Phys. Chem. Chem. Phys., 2022, 24(3), 1553–1568, 10.1039/D1CP04205J.
- I. Biskupek, C. Czaplewski, J. Sawicka, E. Iłowska, M. Dzierżyńska, S. Rodziewicz-Motowidło and A. Liwo, Prediction of aggregation of biologically-active peptides with the UNRES coarse-grained model, Biomolecules, 2022, 12(8), 1140 CrossRef CAS PubMed.
- P. N. Depta, M. Dosta, W. Wenzel, M. Kozlowska and S. Heinrich, Hierarchical coarse-grained strategy for macromolecular self-assembly: application to hepatitis B virus-like particles, Int. J. Mol. Sci., 2022, 23(23), 14699 CrossRef CAS PubMed.
- P. Khan, R. Kaushik and A. Jayaraj, Approaches and perspective of coarse-grained modeling and simulation for polymer–nanoparticle hybrid systems, ACS Omega, 2022, 7(51), 47567–47586 CrossRef CAS PubMed.
- A. P. Latham and B. Zhang, Unifying coarse-grained force fields for folded and disordered proteins, Curr. Opin. Struct. Biol., 2022, 72, 63–70 CrossRef CAS PubMed.
- P. Miszta, P. Pasznik, S. Niewieczerzał, K. Młynarczyk and S. Filipek, COGRIMEN: Coarse-Grained Method for Modeling of Membrane Proteins in Implicit Environments, J. Chem. Theory Comput., 2022, 18(9), 5145–5156 CrossRef CAS PubMed.
- A. Sahoo, P. Y. Lee and S. Matysiak, Transferable and polarizable coarse grained model for proteins–ProMPT, J. Chem. Theory Comput., 2022, 18(8), 5046–5055 CrossRef CAS.
- J. C. Shillcock, C. Lagisquet, J. Alexandre, L. Vuillon and J. H. Ipsen, Model biomolecular condensates have heterogeneous structure quantitatively dependent on the interaction profile of their constituent macromolecules, Soft Matter, 2022, 18(35), 6674–6693 RSC.
- C. Tan, J. Jung, C. Kobayashi, D. U. L. Torre, S. Takada and Y. Sugita, Implementation of residue-level coarse-grained models in GENESIS for large-scale molecular dynamics simulations, PLoS Comput. Biol., 2022, 18(4), e1009578 CrossRef CAS PubMed.
- A. Banerjee and M. Dutt, A Hybrid Approach for Coarse-Graining Helical Peptoids: Solvation, Secondary Structure, and Assembly, J. Chem. Phys., 2023, 158(11), 114105, DOI:10.1063/5.0138510.
- A. Banerjee and M. Dutt, Self-Organization of Mobile, Polyelectrolytic Dendrons on Stable, Amphiphile-Based Spherical Surfaces, Langmuir, 2023, 39(9), 3439–3449, DOI:10.1021/acs.langmuir.2c03386.
- A. Drajkowska and A. Molski, Aggregation and partitioning of amyloid peptide fragments in the presence of a lipid bilayer: a coarse grained molecular dynamics study, Biophys. Chem., 2023, 300, 107051 CrossRef CAS PubMed.
- Y. Ge, X. Wang, Q. Zhu, Y. Yang, H. Dong and J. Ma, Machine Learning-Guided Adaptive Parametrization for Coupling Terms in a Mixed United-Atom/Coarse-Grained Model for Diphenylalanine Self-Assembly in Aqueous Ionic Liquids, J. Chem. Theory Comput., 2023, 19(19), 6718–6732, DOI:10.1021/acs.jctc.3c00809.
- U. Kapoor, Y. C. Kim and J. Mittal, Coarse-grained models to study protein–DNA interactions and liquid–liquid phase separation, J. Chem. Theory Comput., 2023, 20(4), 1717–1731 CrossRef PubMed.
- F. Klein, M. Soñora, L. H. Santos, E. N. Frigini, A. Ballesteros-Casallas, M. R. Machado and S. Pantano, The SIRAH force field: A suite for simulations of complex biological systems at the coarse-grained and multiscale levels, J. Struct. Biol., 2023, 215(3), 107985 CrossRef CAS PubMed.
- D. Liang, Z. Zhang, M. Rafailovich, M. Simon, Y. Deng and P. Zhang, Coarse-grained modeling of the sars-cov-2 spike glycoprotein by physics-informed machine learning, Computation, 2023, 11(2), 24 CrossRef CAS.
- E. Mohammadi, S. Y. Joshi and S. A. Deshmukh, Development, Validation, and Applications of Nonbonded Interaction Parameters between Coarse-Grained Amino Acid and Water Models, Biomacromolecules, 2023, 24(9), 4078–4092, DOI:10.1021/acs.biomac.3c00441.
- S. Mondal and Q. Cui, Membrane remodeling and vesicle formation by biomolecular condensates: A coarse grained simulation study, Biophys. J., 2023, 122(3), 65a–66a CrossRef.
- S. Mushnoori, C. Y. Lu, K. Schmidt and M. Dutt, A Coarse-Grained Molecular Dynamics Study of Phase Behavior in Co-Assembled Lipomimetic Oligopeptides, J. Mol. Graphics Modell., 2023, 125, 108624, DOI:10.1016/j.jmgm.2023.108624.
- Y. Peng, A. J. Pak, A. E. P. Durumeric, P. G. Sahrmann, S. Mani, J. Jin, T. D. Loose, J. Beiter and G. A. Voth, OpenMSCG: A Software Tool for Bottom-Up Coarse-Graining, J. Phys. Chem. B, 2023, 127(40), 8537–8550, DOI:10.1021/acs.jpcb.3c04473.
- J. Wu, W. Xue and G. A. Voth, K-means clustering coarse-graining (KMC-CG): A next generation methodology for determining optimal coarse-grained mappings of large biomolecules, J. Chem. Theory Comput., 2023, 19(23), 8987–8997 CrossRef CAS PubMed.
- J. Zha and F. Xia, Developing hybrid all-atom and ultra-coarse-grained models to investigate taxol-binding and dynein interactions on microtubules, J. Chem. Theory Comput., 2023, 19(16), 5621–5632 CrossRef CAS PubMed.
- A. E. Durumeric, Y. Chen, F. Noé and C. Clementi, Learning data efficient coarse-grained molecular dynamics from forces and noise, arXiv, 2024, preprint, DOI:10.48550/arXiv.2407.01286.
- M. Hooten, A. Banerjee and M. Dutt, Multiscale, Multiresolution Coarse-Grained Model via a Hybrid Approach: Solvation, Structure, and Self-Assembly of Aromatic Tripeptides, J. Chem. Theory Comput., 2024, 20(4), 1689–1703, DOI:10.1021/acs.jctc.3c00458.
- A. T. Shivgan, J. K. Marzinek, A. Krah, P. Matsudaira, C. S. Verma and P. J. Bond, Coarse-grained model of glycosaminoglycans for biomolecular simulations, J. Chem. Theory Comput., 2024, 20(8), 3308–3321 CrossRef CAS PubMed.
- R. Zhang, L. Yang, X. Xiao and H. Liu, Dissipative Particle Dynamics Simulation of Protein Folding in Explicit and Implicit Solvents: Coarse-Grained Model for Atomic Resolution, J. Chem. Theory Comput., 2024, 20(15), 6904–6916 CrossRef CAS PubMed.
- J. Gao, R. Dong, S. Yang, Z. Zhao and Y. W. Zhang, Coarse-grained molecular dynamics investigation on the interaction between κ-and β-casein aggregates and curcumin, PLoS One, 2025, 20(7), e0328010 CrossRef CAS PubMed.
- S. Y. Joshi, T. Kumarage, R. Ashkar and S. A. Deshmukh, Development of Transferable Coarse-Grained Lipid Models with Optimized Structural and Elastic Membrane Properties, J. Chem. Theory Comput., 2025, 21, 9890–9908 CrossRef CAS PubMed.
- A. Jussupow, D. Bartley, L. J. Lapidus and M. Feig, COCOMO2: A coarse-grained model for interacting folded and disordered proteins, J. Chem. Theory Comput., 2025, 21(4), 2095–2107 CrossRef CAS PubMed.
- D. U. La Torre and Y. Sugita, CGBack: Diffusion Model for Backmapping Large-Scale and Complex Coarse-Grained Molecular Systems, J. Chem. Inf. Model., 2025, 65, 9974–9986 CrossRef.
- A. S. Rauh, G. Tesei and K. Lindorff-Larsen, A coarse-grained model for disordered proteins under crowded conditions, Protein Sci., 2025, 34(8), e70232 CrossRef CAS PubMed.
- I. Yasuda, S. von Bülow, G. Tesei, E. Yamamoto, K. Yasuoka and K. Lindorff-Larsen, Coarse-grained model of disordered RNA for simulations of biomolecular condensates, J. Chem. Theory Comput., 2025, 21(5), 2766–2779 CrossRef CAS PubMed.
- S. Y. Joshi and S. A. Deshmukh, A Review of Advancements in Coarse-Grained Molecular Dynamics Simulations, Mol. Simul., 2021, 47(10–11), 786–803, DOI:10.1080/08927022.2020.1828583.
- A. Liwo, C. Czaplewski, A. K. Sieradzan, A. G. Lipska, S. A. Samsonov and R. K. Murarka, Theory and practice of coarse-grained molecular dynamics of biologically important systems, Biomolecules, 2021, 11(9), 1347 CrossRef CAS PubMed.
- J. Jin, A. J. Pak, A. E. P. Durumeric, T. D. Loose and G. A. Voth, Bottom-up Coarse-Graining: Principles and Perspectives, J. Chem. Theory Comput., 2022, 18(10), 5759–5791, DOI:10.1021/acs.jctc.2c00643.
- L. Borges-Araújo, I. Patmanidis, A. P. Singh, L. H. Santos, A. K. Sieradzan, S. Vanni, C. Czaplewski, S. Pantano, W. Shinoda, L. Monticelli and A. Liwo, Pragmatic coarse-graining of proteins: models and applications, J. Chem. Theory Comput., 2023, 19(20), 7112–7135 CrossRef PubMed.
- W. G. Noid, Perspective: Advances, challenges, and insight for predictive coarse-grained models, J. Phys. Chem. B, 2023, 127(19), 4174–4207 CrossRef CAS PubMed.
- K. L. Saar, D. Qian, L. L. Good, A. S. Morgunov, R. Collepardo-Guevara, R. B. Best and T. P. Knowles, Theoretical and data-driven approaches for biomolecular condensates, Chem. Rev., 2023, 123(14), 8988–9009 CrossRef CAS PubMed.
- I. Zaporozhets and C. Clementi, Multibody terms in protein coarse-grained models: A top-down perspective, J. Phys. Chem. B, 2023, 127(31), 6920–6927 CrossRef CAS PubMed.
- A. Banerjee, M. Hooten, N. Srouji, R. Welch, J. Shovlin and M. Dutt, A Perspective on Coarse-Graining Methodologies for Biomolecules: Resolving Self-Assembly over Extended Spatiotemporal Scales, Front. Soft Matter, 2024, 4, 1361066, DOI:10.3389/frsfm.2024.1361066.
- W. G. Noid, R. J. Szukalo, K. M. Kidder and M. C. Lesniewski, Rigorous Progress in Coarse-Graining, Annu. Rev. Phys. Chem., 2024, 75, 21–45, DOI:10.1146/annurev-physchem-062123-010821.
- S. Liu, C. Wang and B. Zhang, Toward Predictive Coarse-Grained Simulations of Biomolecular Condensates, Biochemistry, 2025, 64(8), 1750–1761 CrossRef CAS PubMed.
- A. B. Poma, A. H. Caldas, L. F. Cofas-Vargas, M. S. Jones, A. L. Ferguson and L. M. Sandonas, Recent Advances in Machine Learning and Coarse-Grained Potentials for Biomolecular Simulations and Their Applications, Biophys. J., 2025, 125, 327–343, DOI:10.1016/j.bpj.2025.06.019.
- P. C. Whitford and J. N. Onuchic, Simulating biomolecules for physiological timescales, Curr. Opin. Struct. Biol., 2025, 92, 103039 CrossRef CAS PubMed.
- O. Engin, A. Villa, C. Peter and M. Sayar, A Challenge for Peptide Coarse Graining: Transferability of Fragment-Based Models, Macromol. Theory Simul., 2011, 20(7), 451–465, DOI:10.1002/mats.201100005.
- S. P. Carmichael and M. S. Shell, A New Multiscale Algorithm and Its Application to Coarse-Grained Peptide Models for Self-Assembly, J. Phys. Chem. B, 2012, 116(29), 8383–8393, DOI:10.1021/jp2114994.
- B. Ozgur and M. Sayar, Assembly of Triblock Amphiphilic Peptides into One-Dimensional Aggregates and Network Formation, J. Phys. Chem. B, 2016, 120(39), 10243–10257, DOI:10.1021/acs.jpcb.6b07545.
- O. Conway, Y. An, K. K. Bejagam and S. A. Deshmukh, Development of Transferable Coarse-Grained Models of Amino Acids, Mol. Syst. Des. Eng., 2020, 5(3), 675–685, 10.1039/C9ME00173E.
- M. S. Shell, The Relative Entropy Is Fundamental to Multiscale and Inverse Thermodynamic Problems, J. Chem. Phys., 2008, 129(14), 144108, DOI:10.1063/1.2992060.
- K. Liebl and G. A. Voth, A Hybrid Bottom-Up and Data-Driven Machine Learning Approach for Accurate Coarse-Graining of Large Molecular Complexes, J. Chem. Theory Comput., 2025, 21(9), 4846–4854, DOI:10.1021/acs.jctc.5c00063.
- E. Pretti and M. S. Shell, Characterizing the Sequence Landscape of Peptide Fibrillization with a Bottom-Up Coarse-Grained Model, J. Phys. Chem. B, 2025, 129(14), 3559–3570, DOI:10.1021/acs.jpcb.4c07248.
- T. C. Moore, C. R. Iacovella and C. McCabe, Derivation of Coarse-Grained Potentials via Multistate Iterative Boltzmann Inversion, J. Chem. Phys., 2014, 140(22), 224104, DOI:10.1063/1.4880555.
- H. Wang, C. Hartmann, C. Schütte and L. Delle Site, Grand-Canonical-like Molecular-Dynamics Simulations by Using an Adaptive-Resolution Technique, Phys. Rev. X, 2013, 3(1), 011018, DOI:10.1103/PhysRevX.3.011018.
- M. Hooten and M. Dutt Hybrid Bottom-Up Coarse-Grained Model for Aromatic Peptides, 2023, GitHub repository, https://github.com/duttm/Hybrid_Bottom-Up_Coarse-Grained_Model_for_Aromatic_Peptides, (accessed 2026-02-22).
- W. Humphrey, A. Dalke and K. Schulten, VMD: Visual Molecular Dynamics, J. Mol. Graphics, 1996, 14(1), 33–38, DOI:10.1016/0263-7855(96)00018-5.
- V. Rühle, C. Junghans, A. Lukyanov, K. Kremer and D. Andrienko, Versatile Object-Oriented Toolkit for Coarse-Graining Applications, J. Chem. Theory Comput., 2009, 5(12), 3211–3223, DOI:10.1021/ct900369w.
- S. Izvekov and G. A. Voth, A Multiscale Coarse-Graining Method for Biomolecular Systems, J. Phys. Chem. B, 2005, 109(7), 2469–2473, DOI:10.1021/jp044629q.
- F. Müller-Plathe, Coarse-Graining in Polymer Simulation: From the Atomistic to the Mesoscopic Scale and Back, ChemPhysChem, 2002, 3(9), 754–769, DOI:10.1002/1439-7641(20020916)3:9<754::AID-CPHC754>3.0.CO;2-U.
- D. Reith, M. Pütz and F. Müller-Plathe, Deriving Effective Mesoscale Potentials from Atomistic Simulations, J. Comput. Chem., 2003, 24(13), 1624–1636, DOI:10.1002/jcc.10307.
- A. K. Soper, Empirical Potential Monte Carlo Simulation of Fluid Structure, Chem. Phys., 1996, 202(2), 295–306, DOI:10.1016/0301-0104(95)00357-6.
- Y. Rubner, C. Tomasi and L. J. Guibas, A Metric for Distributions with Applications to Image Databases, In Sixth International Conference on Computer Vision, 1998, pp. 59–66, DOI:10.1109/ICCV.1998.710701.
- Y. Rubner, C. Tomasi and L. J. Guibas, The Earth Mover's Distance as a Metric for Image Retrieval, Int. J. Comput. Vis., 2000, 40(2), 99–121, DOI:10.1023/A:1026543900054.
- A. Van Teijlingen, M. C. Smith and T. Tuttle, Short Peptide Self-Assembly in the Martini Coarse-Grain Force Field Family, Acc. Chem. Res., 2023, 56(6), 644–654, DOI:10.1021/acs.accounts.2c00810.
- L. Monticelli, S. K. Kandasamy, X. Periole, R. G. Larson, D. P. Tieleman and S.-J. Marrink, The MARTINI Coarse-Grained Force Field: Extension to Proteins, J. Chem. Theory Comput., 2008, 4(5), 819–834, DOI:10.1021/ct700324x.
- G. Bussi, D. Donadio and M. Parrinello, Canonical Sampling through Velocity Rescaling, J. Chem. Phys., 2007, 126(1), 014101, DOI:10.1063/1.2408420.
- G. Bussi, T. Zykova-Timan and M. Parrinello, Isothermal-Isobaric Molecular Dynamics Using Stochastic Velocity Rescaling, J. Chem. Phys., 2009, 130(7), 074101, DOI:10.1063/1.3073889.
- H. J. C. Berendsen, J. P. M. Postma, W. F. Van Gunsteren, A. DiNola and J. R. Haak, Molecular Dynamics with Coupling to an External Bath, J. Chem. Phys., 1984, 81(8), 3684–3690, DOI:10.1063/1.448118.
- W. Zhang, M. J. o. Anteunis and F. A. m. Borremans, Aromatic Side Chain Rotational Preference, Int. J. Pept. Protein Res., 1986, 28(6), 593–602, DOI:10.1111/j.1399-3011.1986.tb03297.x.
- M. P. Allen and D. J. Tildesley, Computer Simulation of Liquids, Oxford university press, 2017 Search PubMed.
- S. Hunkler, T. Lemke, C. Peter and O. Kukharenko, Back-Mapping Based Sampling: Coarse Grained Free Energy Landscapes as a Guideline for Atomistic Exploration, J. Chem. Phys., 2019, 151(15), 154102, DOI:10.1063/1.5115398.
- S. Catalini, F. Bagni, S. Cicchi, M. Di Donato, A. Iagatti, A. Lapini, P. Foggi, C. Petrillo, A. Di Michele, M. Paolantoni, G. Schirò, L. Comez and A. Paciaroni, Multiple Length-Scale Control of Boc-Protected Diphenylalanine Aggregates through Solvent Composition, Mater. Adv., 2024, 5(9), 3802–3811, 10.1039/D4MA00018H.
- C. Diaferia, G. Morelli and A. Accardo, Fmoc-Diphenylalanine as a Suitable Building Block for the Preparation of Hybrid Materials and Their Potential Applications, J. Mater. Chem. B, 2019, 7(34), 5142–5155, 10.1039/C9TB01043B.
- E. Mayans and C. Alemán, Revisiting the Self-Assembly of Highly Aromatic Phenylalanine Homopeptides, Molecules, 2020, 25(24), 6037, DOI:10.3390/molecules25246037.
- D. Zaguri, M. R. Zimmermann, G. Meisl, A. Levin, S. Rencus-Lazar, T. P. J. Knowles and E. Gazit, Kinetic and Thermodynamic Driving Factors in the Assembly of Phenylalanine-Based Modules, ACS Nano, 2021, 15(11), 18305–18311, DOI:10.1021/acsnano.1c07537.
- C. Liu, Y. Dan, J. Yun, L. Adler-Abramovich and J. Luo, Unveiling the Assembly Transition of Diphenylalanine and Its Analogs: From Oligomer Equilibrium to Nanocluster Formation, ACS Nano, 2025, 19(13), 13250–13263, DOI:10.1021/acsnano.5c00433.
- F. Aydin, P. Ludford and M. Dutt, Phase Segregation in Bio-Inspired Multi-Component Vesicles Encompassing Double Tail Phospholipid Species, Soft Matter, 2014, 10(32), 6096–6108, 10.1039/C4SM00998C.
- F. Aydin, G. Uppaladadium and M. Dutt, Harnessing Nanoscale Confinement to Design Sterically Stable Vesicles of Specific Shapes via Self-Assembly, J. Phys. Chem. B, 2015, 119(32), 10207–10215, DOI:10.1021/acs.jpcb.5b02239.
- F. Aydin, G. Uppaladadium and M. Dutt, The Design of Shape-Tunable Hairy Vesicles, Colloids Surf., B, 2015, 128, 268–275, DOI:10.1016/j.colsurfb.2015.01.049.
- S. Mushnoori, C. Y. Lu, K. Schmidt, E. Zang and M. Dutt, Peptide-Based Vesicles and Droplets: A Review, J. Phys.: Condens. Matter, 2020, 33(5), 053002, DOI:10.1088/1361-648X/abb995.
- X. Yu and M. Dutt, Implementation of Dynamic Coupling in Hybrid Molecular Dynamics–Lattice Boltzmann Approach: Modeling Aggregation of Amphiphiles, Comput. Phys. Commun., 2020, 257, 107287, DOI:10.1016/j.cpc.2020.107287.
- A. Levin, T. O. Mason, L. Adler-Abramovich, A. K. Buell, G. Meisl, C. Galvagnion, Y. Bram, S. A. Stratford, C. M. Dobson and T. P. Knowles, Ostwald's Rule of Stages Governs Structural Transitions and Morphology of Dipeptide Supramolecular Polymers, Nat. Commun., 2014, 5(1), 5219 CrossRef CAS PubMed.
- B. Strodel, Amyloid Aggregation Simulations: Challenges, Advances and Perspectives, Curr. Opin. Struct. Biol., 2021, 67, 145–152, DOI:10.1016/j.sbi.2020.10.019.
- S. C. Mushnoori, E. Zang, A. Banerjee, M. Hooten, A. Merzky, M. Turilli, S. Jha and M. Dutt, Pipelines for Automating Compliance-Based Elimination and Extension (PACE2): A Systematic Framework for High-Throughput Biomolecular Materials Simulation Workflows, JPhys Mater., 2023, 7(1), 015006, DOI:10.1088/2515-7639/ad08d0.
| This journal is © The Royal Society of Chemistry 2026 |








