Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Investigating the role of cytochrome bd oxidases in the antibacterial activity of madecassic acid and derivatives thereof
Samantha A.
Henry
a,
Geraud N.
Sansom
 a,
Thao Thi Phuong
Tran
a,
Thao Thi Phuong
Tran
 b,
Ryan A.
Boughton
a,
Guy
Joiner
a,
Calum M.
Webster
a,
H. Ireshika C.
de Silva
b,
Ryan A.
Boughton
a,
Guy
Joiner
a,
Calum M.
Webster
a,
H. Ireshika C.
de Silva
 c,
Michelle D.
Garrett
c,
Michelle D.
Garrett
 a,
Christopher J.
Serpell
a,
Christopher J.
Serpell
 *d,
Gary K.
Robinson
*d,
Gary K.
Robinson
 a and
Mark
Shepherd
a and
Mark
Shepherd
 *a
*a
aSchool of Natural Sciences, University of Kent, Canterbury, CT2 7NH, UK. E-mail: M.Shepherd@kent.ac.uk
bInstitute of Chemistry, Vietnam Academy of Science and Technology, 18 Hoang Quoc Viet Road, Nghia Do, Hanoi, Vietnam
cDepartment of Chemistry, University of Colombo, Colombo 03, Sri Lanka
dSchool of Pharmacy, University College London, 29-39 Brunswick Square, London, WC1N 1AX, UK. E-mail: chris.serpell@ucl.ac.uk
First published on 13th January 2026
Abstract
Natural products are valuable starting points for drug discovery, although individual modes of action are often difficult to pin down. Ursanes such as madecassic acid have been shown to have antibacterial properties, but a variety of mechanisms have been proposed. In this paper, we report previously uninvestigated activity against cytochrome bd oxidases which are only found in prokaryotes and are therefore promising new targets, using madecassic acid and a set of synthetically modified derivatives. Our work shows that madecassic acid and its derivatives can block activity of these enzymes, while phenotypic effects in membrane and whole organism assays are more complex, consistent with modulation of multiple pathways depending upon molecular structure. This provides a new route to ursane-based antibacterial action while highlighting the importance of chemical modifications in fine-tuning biological activity of natural products.
Introduction
Ursanes are pentacyclic terpenoids which have been shown to have antibacterial properties.1–4 Madecassic acid, an ursane produced by the common Asian herb Centella asiatica, has been shown to have good activity against S. aureus and moderate activity against other bacterial species, with evidence consistent with a mechanism including increasing membrane and wall permeability, interaction with DNA, and inhibition of protein synthesis.5 This is similar to evidence for ursolic acid, a related compound also found to be active against Gram-positive bacteria.6 Unpurified extracts of C. asiatica have shown to be active against Gram-positive and -negative bacteria.7 These trends have been demonstrated across a range of modified ursanes,8 and the potential of asiatic acid to inhibit bacterial growth had led to its incorporation into gel materials to aid in vivo wound healing.9Cytochrome bd oxidases are attractive drug targets for discovery of new antibacterials, since they are found only in prokaryotic organisms,10 and therefore permit specific killing of bacteria. The proteins are found on the inner membrane, where their role is to couple reduction of molecular oxygen with oxidation of quinols, resulting in a proton motive force used in the generation of ATP. Blocking cytochrome bd oxidase activity would therefore impede the capacity of bacteria to access energy required for growth.11Escherichia coli, a Gram-negative bacterium responsible for many urinary tract and bloodstream infections, has two bd oxidases: bd-I which is expressed in microaerobic environments, and bd-II which is predominant in anaerobic conditions.12
We have discovered effective steroidal ligands for bd-I which inhibit growth and provide lethality against bacteria, with the strongest measured affinities in E. coli.13 Since ursane-type structures resemble sterols, it is possible that binding of bacterial cytochrome oxidases is a key component in the antibacterial mechanism of action of the compound class. Since literature reports that a number of mechanisms are at play, it would also be useful to know if the balance of these processes can be modulated through structural refinement of the compounds. In this paper we test madecassic acid (MA) and three synthetic analogues for cytochrome oxidase inhibition through molecular docking, measurement of oxygen consumption in isolated membranes, and testing against live E. coli models, each engineered to have access to different oxidases.
Experimental procedures
Chemical synthesis
Madecassic acid (MA) was obtained by extraction of leaves grown in Quang Tho II Commune, Hue Province, VietNam, following our reported precedure.14 Deuterated solvents were purchased from Cambridge Isotope Laboratories Inc. Solvents and reagents were purchased from Thermo Fisher Scientific, Fluorochem, or Alfa Aesar.A Bruker AVII 400 MHz spectrometer was used to record NMR spectra, and each spectrum was calibrated to the known chemical shift of the residual solvent peak of the deuterated solvent used. Chemical shifts were reported in part per million (ppm) and J coupling values were reported in Hz. Proton NMR spectra were obtained at 400 MHz and 13C spectra were obtained at 101 MHz. Spectral data was processed using MestReNova software. Electrospray mass spectrometry data was obtained using a Thermo MSQPlus instrument fitted with a Zorbax SB-C18 5 μm 3.0 × 150 mm column using H2O + 0.1% formic acid and MeOH + 0.1% formic acid or H2O + 0.1% TFA and MeCN + 0.1% TFA mobile phases. Data was analysed using Chromeleon™ Chromatography Data System (CDS) Software. A Bruker micrOTOF-Q LCMS system was used to obtain high-resolution mass spectrometry data, samples were dissolved in HPLC-grade methanol and injected using direct injection mode with a mobile phase system of 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 MeOH and H2O. Data was processed using Bruker Compass Hystar software.
50 MeOH and H2O. Data was processed using Bruker Compass Hystar software.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH2Cl2) giving MA-2 as a white needle crystalline solid (183 mg 72.6% yield). 1H NMR (400 MHz, CDCl3) δ 5.27 (d, J = 3.6 Hz, 1H), 5.22 (td, J = 11.3, 4.8 Hz, 2H), 5.00 (d, J = 10.3 Hz, 1H), 4.33 (d, J = 3.8 Hz, 1H), 3.93 (d, J = 11.9 Hz, 1H), 3.70 (d, J = 12.0 Hz, 1H), 2.19 (d, J = 11.2 Hz, 1H), 2.15–0.80 (m, 46H). 13C NMR (101 MHz, CDCl3) δ 183.9, 171.0, 170.6, 170.6, 137.3, 125.7, 75.0, 70.0, 68.0, 65.5, 52.5, 48.3, 48.0, 47.9, 45.9, 42.5, 40.8, 39.2, 38.9, 38.7, 37.4, 36.7, 30.7, 28.0, 24.2, 23.6, 23.4, 21.3, 21.2, 21.2, 21.0, 20.9, 18.8, 18.6, 17.1, 15.5, 14.3. HREI-MS: m/z calculated for C36H54O9Na [M + Na+] 653.3660; observed 653.3672.
CH2Cl2) giving MA-2 as a white needle crystalline solid (183 mg 72.6% yield). 1H NMR (400 MHz, CDCl3) δ 5.27 (d, J = 3.6 Hz, 1H), 5.22 (td, J = 11.3, 4.8 Hz, 2H), 5.00 (d, J = 10.3 Hz, 1H), 4.33 (d, J = 3.8 Hz, 1H), 3.93 (d, J = 11.9 Hz, 1H), 3.70 (d, J = 12.0 Hz, 1H), 2.19 (d, J = 11.2 Hz, 1H), 2.15–0.80 (m, 46H). 13C NMR (101 MHz, CDCl3) δ 183.9, 171.0, 170.6, 170.6, 137.3, 125.7, 75.0, 70.0, 68.0, 65.5, 52.5, 48.3, 48.0, 47.9, 45.9, 42.5, 40.8, 39.2, 38.9, 38.7, 37.4, 36.7, 30.7, 28.0, 24.2, 23.6, 23.4, 21.3, 21.2, 21.2, 21.0, 20.9, 18.8, 18.6, 17.1, 15.5, 14.3. HREI-MS: m/z calculated for C36H54O9Na [M + Na+] 653.3660; observed 653.3672.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 75%, EtOAc
75%, EtOAc![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CHCl3) to yield MA-3Boc as a white solid (236 mg, 95.8%). 1H NMR (400 MHz, CDCl3) δ 5.85 (t, J = 5.4 Hz, 1H), 5.34 (t, J = 3.5 Hz, 1H), 5.22 (td, J = 11.0, 4.7 Hz, 1H), 5.01 (d, J = 10.3 Hz, 1H), 4.57 (s, 1H), 4.34 (t, J = 3.8 Hz, 1H), 3.94 (d, J = 11.9 Hz, 1H), 3.71 (d, J = 12.0 Hz, 1H), 3.23 (dq, J = 13.3, 6.8 Hz, 1H), 3.14–2.93 (m, 3H), 2.17–0.77 (m, 72H). 13C NMR (101 MHz, CDCl3) δ 177.9, 170.9, 170.5, 170.5, 139.4, 125.5, 79.1, 75.0, 70.1, 67.4, 65.4, 54.2, 48.1, 47.9, 47.8, 46.0, 43.1, 42.5, 40.6, 39.9, 39.6, 39.2, 38.8, 38.8, 37.4, 37.3, 31.0, 30.2, 29.5, 29.5, 29.3, 29.3, 29.3, 29.2, 28.5, 27.9, 27.2, 26.9, 24.9, 23.5, 23.3, 21.3, 21.2, 21.0, 20.9, 18.7, 18.3, 17.3, 15.4. HREI-MS: m/z calculated for C51H84N2O10Na [M + Na+] 907.6018; observed 907.6052.
CHCl3) to yield MA-3Boc as a white solid (236 mg, 95.8%). 1H NMR (400 MHz, CDCl3) δ 5.85 (t, J = 5.4 Hz, 1H), 5.34 (t, J = 3.5 Hz, 1H), 5.22 (td, J = 11.0, 4.7 Hz, 1H), 5.01 (d, J = 10.3 Hz, 1H), 4.57 (s, 1H), 4.34 (t, J = 3.8 Hz, 1H), 3.94 (d, J = 11.9 Hz, 1H), 3.71 (d, J = 12.0 Hz, 1H), 3.23 (dq, J = 13.3, 6.8 Hz, 1H), 3.14–2.93 (m, 3H), 2.17–0.77 (m, 72H). 13C NMR (101 MHz, CDCl3) δ 177.9, 170.9, 170.5, 170.5, 139.4, 125.5, 79.1, 75.0, 70.1, 67.4, 65.4, 54.2, 48.1, 47.9, 47.8, 46.0, 43.1, 42.5, 40.6, 39.9, 39.6, 39.2, 38.8, 38.8, 37.4, 37.3, 31.0, 30.2, 29.5, 29.5, 29.3, 29.3, 29.3, 29.2, 28.5, 27.9, 27.2, 26.9, 24.9, 23.5, 23.3, 21.3, 21.2, 21.0, 20.9, 18.7, 18.3, 17.3, 15.4. HREI-MS: m/z calculated for C51H84N2O10Na [M + Na+] 907.6018; observed 907.6052.
MA-3Boc (131.2 mg, 0.148 mmol) was then dissolved in CH2Cl2 (3.7 mL), 4 M HCl in dioxane (3.7 mL, 14.8 mmol) was added to the solution at 0 °C. The reaction was stirred and left to warm to room temperature overnight. After 21 hours, solvent was removed under reduced pressure. The residue was washed into a separating funnel with a mixture of CH2Cl2 (50 mL) and sat. NaHCO3(aq) (50 mL) and aqueous layer removed. It was then extracted with a further sat. NaHCO3(aq) (2 × 50 mL) and brine (1 × 50 mL). Dried over anhydrous MgSO4, filtered and reduced under vacuum. Extracted product was clean by NMR and required no further purification yielding MA-3 (105.8 mg, 90.9%). 1H NMR (400 MHz, CDCl3) δ 5.92 (t, J = 5.4 Hz, 1H), 5.34 (d, J = 3.7 Hz, 1H), 5.21 (td, J = 10.9, 4.5 Hz, 1H), 5.00 (d, J = 10.3 Hz, 1H), 4.83 (s, 2H), 4.31 (d, J = 5.1 Hz, 1H), 3.94 (d, J = 11.7 Hz, 1H), 3.70 (dd, J = 12.5, 2.6 Hz, 1H), 3.28–2.94 (m, 3H), 2.85 (s, 2H), 2.25–0.73 (m, 62H). 13C NMR (101 MHz, CDCl3) δ 178.1, 171.1, 170.7, 170.6, 139.4, 125.5, 75.0, 70.2, 67.2, 65.4, 54.1, 48.1, 47.9, 47.8, 46.0, 43.8, 43.1, 42.5, 40.4, 39.9, 39.7, 39.2, 38.8, 37.4, 37.3, 31.0, 29.5, 29.4, 29.3, 29.3, 29.2, 29.1, 27.8, 27.1, 26.7, 24.9, 23.5, 23.3, 21.3, 21.3, 21.0, 20.9, 18.7, 18.3, 17.3, 15.4.
Computation and molecular docking
Ligand docking to an AlphaFold2 model of E. coli cytochrome bd-I was performed essentially as described in detail previously.13 Briefly, preparation of in silico ligand and protein files was performed using AutoDockTools and PyMOL.15 The equation Kd = exp(ΔG/(R × T)) with ΔG = binding energy (kcal mol−1), R = gas constant = (1.986 cal mol−1 K−1) and T = temperature (298 K) was used to estimate the dissociation constants.Clog![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values were calculated using the Chemical Properties tool in Signals ChemDraw v23.1.2.7.
P values were calculated using the Chemical Properties tool in Signals ChemDraw v23.1.2.7.
Bacterial strains
EC958 is a multidrug-resistant E. coli O25: H4-ST131 isolate.16,17 Generation of the EC958 bd-I only strain (genotype ΔcyoA appCB::Cm) is described elsewhere.13Growth assays
Starter cultures were grown in 10 mL LB (E. coli) in sterile 50 mL conical flasks at 180 rpm and 37 °C until stationary phase was reached and were used to inoculate 50 mL of fresh growth medium in 250 mL conical flasks. M9 minimal medium was used for E. coli (16 g L−1 Na2HPO4·2H2O, 3 g L−1 KH2PO4, 0.5 g L−1 NaCl, 1 g L−1 NH4Cl, 0.24 g L−1 MgSO4, 0.01 g L−1 CaCl2, 0.1% casamino acids and 2% glycerol). Drug stocks were prepared in DMSO so that their final concentrations were 40× higher than the working concentrations. Greiner F-bottom sterile 96-well plates were prepared by adding 100 μL of a 2× concentrated growth medium, 61.7 μL sterile milliQ H2O, 5 μL of the drug and 33.3 μL of cell culture (final OD600 of ∼0.1). Cells were grown in a FLUOstar Omega plate reader at double orbital pattern setting.Oxygen consumption assays
Membrane preparation and oxygen consumption were performed as described previously.13 Briefly, cells were grown to exponential phase then harvested via centrifugation for 20 min at 4000 rpm and 4 °C. The cell pellet was resuspended in ice-cold sonication buffer (20 mM Tris/HCl at pH 7.4, 2 mM MgCl2, and 1 mM EGTA). The resuspended cells were sonicated (6 × 30s on ice at 15 μM) before centrifugation at 44![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 rpm for 1 h and 4 °C to isolate membranes. The membrane pellet was resuspended in 20 mM Tris/HCl (pH 7.4) at a final concentration of 100 mg mL−1 and stored at −20 °C. For oxygen measurements a Rank Brothers oxygen electrode with a 4 mL closed chamber was used at 37 °C, which contained 50 mM HEPES pH 7.4, 0.5 mg mL−1 membranes (based on wet membranes) and DMSO-solubilised drug (added from 40× final concentration). A final concentration of 8 mM succinate (pH 7.4) was added (from 160 mM stock) to initiate the reaction with a single run lasting 15–20 min.
000 rpm for 1 h and 4 °C to isolate membranes. The membrane pellet was resuspended in 20 mM Tris/HCl (pH 7.4) at a final concentration of 100 mg mL−1 and stored at −20 °C. For oxygen measurements a Rank Brothers oxygen electrode with a 4 mL closed chamber was used at 37 °C, which contained 50 mM HEPES pH 7.4, 0.5 mg mL−1 membranes (based on wet membranes) and DMSO-solubilised drug (added from 40× final concentration). A final concentration of 8 mM succinate (pH 7.4) was added (from 160 mM stock) to initiate the reaction with a single run lasting 15–20 min.
Viability assays
Viability assays were performed essentially as described previously.13 Briefly, E. coli was grown overnight in 10 mL LB medium and was used to inoculate M9 medium. 20 μL of each drug was added to the wells of row A of a 96 well plate followed by 180 μL of cells (OD600 of 0.1). Cells were exposed to drug for 3 hours at 37 °C. Following drug exposure, serial dilutions were performed in 1 × phosphate buffered saline (pH 7.4) before being spotted onto LB agar plates overnight to determine changes in cell survival. Six repeats were performed for each concentration of drug which included two biological repeats and three technical repeats of each.Results
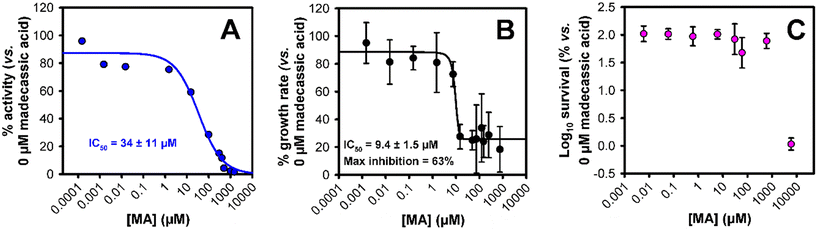
Both MA and ubiquinol-8 (UQ-8) the natural substrate of cytochrome bd-I were docked to the quinol site of the E. coli AlphaFold2 cytochrome bd-I structural model.13 The binding affinity values predicted by AutoDock Vina (Fig. 1) indicate that MA is substantially more tightly bound by cytochrome bd-I (1.14 μM) than UQ-8 (108.9 μM). The carboxylic acid of MA was found to reside in the hydrophobic cleft (red, Fig. 1) close to where the aromatic unit of UQ-8 sits, while the pentacyclic hydrocarbon occupies a similar space to that of the oligoisoprene tail of UQ-8. The polar units on MA point out of the protein cavity towards the aqueous environment.These predictions suggest that MA would be able to competitively inhibit cytochrome bd-I. To test this prediction, we performed oxygen consumption assays to measure oxidoreductase activity using isolated E. coli EC958 cytochrome bd-I only membranes (Fig. 2A),13 and MA obtained by extraction of C. asiatica,14 resulting in an IC50 of 34 ± 11 μM. This activity is consistent with the docking results in that MA would be expected to outcompete UQ-8. On this basis, we advanced to assessing the ability of MA to inhibit the growth of cytochrome bd-I only E. coli cells (Fig. 2B), giving an IC50 of 9.4 ± 1.5 μM, with a maximum inhibition of 63%. We further conducted a viability assay (Fig. 2C) which showed that MA did not kill bd-I only E. coli cells. The observed lack of lethality may relate to permeability of the cell membrane and/or wall to MA, but is consistent with other tests on Gram-negative bacteria.5
To investigate the contribution of cytochrome bd-I to MA-mediated respiratory inhibition, growth experiments were also performed on the wild type EC958 strain, which encodes an additional respiratory oxidase cytochrome bo′ that is expressed under the aerobic conditions tested (Fig. S1A). These data revealed an IC50 of ∼10 μM and maximal inhibition of 48%, which is consistent with the wild type responding to respiratory inhibition in a similar way to the bd-I only strain. In addition, the wild type strain was not susceptible to MA-mediated killing at concentrations below 1 mM (Fig. S1B), confirming that MA is bacteriostatic but not bactericidal towards both strains.
We then decided to explore variations upon the MA skeleton which might alter access to proteins on the bacterial inner membrane. For this reason, we examined modified versions. These were based on transformations prominent in the literature for such compounds,18,19 enabling easy development in future, and operated on opposite ends of the molecule to provide initial independent evaluation of modification sites. Acetylating the alcohols at positions 2, 3, and 23 gave MA-2, lowering polarity from Clog![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P of MA at 3.97 to 6.64. Additionally, conversion of the carboxylic acid into an amide using 1,10-diaminodecane to add a hydrophobic chain and introduce a cationic group (likely to interact with bacterial lipids) gave MA-3 (Clog
P of MA at 3.97 to 6.64. Additionally, conversion of the carboxylic acid into an amide using 1,10-diaminodecane to add a hydrophobic chain and introduce a cationic group (likely to interact with bacterial lipids) gave MA-3 (Clog![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P = 8.63). We also produced the 1,10-diaminodecane amide without acetylation (MA-4, Clog
P = 8.63). We also produced the 1,10-diaminodecane amide without acetylation (MA-4, Clog![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P = 5.96) to create a double-mutant cycle. These compounds have been previously studied for anticancer activity, and were synthesised in the same manner, starting from MA.18 As before, these were docked into the quinol binding site of cytochrome bd-I (Fig. 3). Compared with the predicted Kd of 1.14 μM for MA, acetylation resulted in a slightly stronger affinity (Kd = 0.96 μM) for MA-2. However, when the long chain amine was also added to give MA-3, the affinity was substantially weakened (Kd = 14.4 μM). The chain appears to add too much bulk, and impedes the binding geometry. Removal of some of the molecule's steric bulk in the unacetylated MA-4 resulted in a moderate improvement in Kd (8.64 μM). Importantly, all of these are modelled to bind more strongly than UQ-8, and therefore we could expect inhibitory activity.
P = 5.96) to create a double-mutant cycle. These compounds have been previously studied for anticancer activity, and were synthesised in the same manner, starting from MA.18 As before, these were docked into the quinol binding site of cytochrome bd-I (Fig. 3). Compared with the predicted Kd of 1.14 μM for MA, acetylation resulted in a slightly stronger affinity (Kd = 0.96 μM) for MA-2. However, when the long chain amine was also added to give MA-3, the affinity was substantially weakened (Kd = 14.4 μM). The chain appears to add too much bulk, and impedes the binding geometry. Removal of some of the molecule's steric bulk in the unacetylated MA-4 resulted in a moderate improvement in Kd (8.64 μM). Importantly, all of these are modelled to bind more strongly than UQ-8, and therefore we could expect inhibitory activity.
Oxygen consumption assays were then performed with bd-I-only membranes (Fig. 4a). The tighter predicted binding of MA-2 compared with MA correlated with elevated inhibition (IC50 = 15 μM, Fig. 4A), as did the slightly weaker binding of MA-4 correspond to a lowered inhibition (IC50 = 79 μM, Fig. 4G). However, MA-3 was more active than would be expected from the predicted binding (IC50 = 10 μM, Fig. 4D). This is potentially due to the more lipophilic nature of that compound which may result in it being drawn into the membranes. The same compound performed well in the growth inhibition of bd-I only E. coli cells (IC50 = 16 μM, Fig. 4E), although was not as potent as MA (IC50 = 9.4 μM, Fig. 2B). MA-2 was much less active (IC50 = 116 μM, Fig. 4B) despite having good activity against membranes, while MA-4′s weak performance against membranes was echoed in its limited (IC50 > 250 μM, Fig. 4H) growth inhibition. In light of these results, it was therefore a surprise when MA-4 revealed itself as the only compound tested which had any capacity (albeit weak) to kill E. coli bd-I only cells (Fig. 4I), with an LC50 of 304 μM.
Discussion
These results show that inhibitory activity of bacterial cytochromes is involved in the antibacterial activity of MA, with binding to the ubiquinol site of cytochrome bd-I being of particular interest, to which MA and all our derivatives are modelled to bind more strongly than the natural substrate. In line with these results, MA itself shows inhibition of enzymatic activity and bacterial growth, although without any observed killing of E. coli – this is consistent with other studies which found that Gram-positive bacteria were more vulnerable.5Having applied chemical modifications to MA, we were able to improve enzymatic inhibition activity in membrane assays for two out of three variants (MA-2 and MA-3). Unmodified MA proved best for growth inhibition, although MA-3, the most highly modified version was close behind, despite being expected to bind the protein the least strongly. However, it was only MA-4, ranked third in predicting binding and last in enzymatic effects, which was found to kill E. coli. It must be said therefore that the links between predicted binding, and the different levels of biological activity (protein activity in isolated membranes, growth inhibition, cell killing) are mixed. This supports previous work by other groups which have identified a variety of mechanisms through which MA has antibacterial effects including membrane disruption, inhibition of protein synthesis, and topoisomerase inhibition.6 In particular, we note that our modifications make the compound significantly more hydrophobic as reflected in a shift of calculated partition coefficient of four log10 units between MA and MA-3, as well as swapping an anionic (at biological pH) carboxylate for a cationic ammonium; both of these changes are likely to increase interactions with lipids. The proteins of interest reside in the inner membrane of E. coli, and their natural substrate is primarily found in the lipid bilayer. This increased lipidic interaction could therefore both increase the local concentration of MA derivatives, or conversely result in competitive sequestration within the lipid bilayer, away from the protein. Convolution of these effects with the other mechanisms at play have led to the overall result that MA-4 has the greatest bactericidal activity. The myriad ways in which natural products such as MA can interact with biological macromolecules leads to complications which are hard to avoid – in our case, the possibility that MA might interact with succinate dehydrogenase5 was a potential limitation for the isolated membrane components of this study, since the oxygen consumption assay relies upon this complex to maintain the supply of ubiquinol for cytochrome bd-mediated oxygen consumption. It was therefore necessary to perform additional control experiments to exclude MA-mediated SDH inhibition and further demonstrate the inhibition of cytochrome bd-I activity where electron delivery is independent of SDH. To introduce an additional level of rigour to these investigations, an alternative bd-I only strain was engineered where an oxidase null mutant strain was complemented with a plasmid encoding the cytochrome bd operon (SI). Membranes from this E. coli EcoM4 pSU2718G-cydABX-his6 strain were prepared and SDH activity measurements confirmed that 1 mM MA did not inhibit SDH in isolated membranes (Fig. S2). To verify that MA directly targets cytochrome bd-I directly, oxygen consumption experiments were undertaken using the same membranes and duroquinol as the electron donor (delivers electrons directly to bd-I). These data (Fig. S3) confirmed that 1 mM MA completely abolishes bd-I activity, further supporting the direct binding of MA to bd-I in EC958 membranes.
Plants have evolved a certain set of natural products which have a particular role for that plant. Because they interact with the biological machinery in one organism, there is a good chance that they will do something in other organisms, and this has historically led to many successful medicines.20 However, there is no reason to expect natural products to behave exactly as we would wish, and off-target effects are common, since the compounds were selected for a different role. Chemical modification can make a big difference,21 and we have shown here that modifications can alter the behaviour of MA in bacterial models, and this provides a route to focus the activity of a natural product into a particular mechanism. Through strategic chemical modification, it should therefore be possible to both better understand the different mechanisms of triterpenoid antibacterial activity, and create molecules which maximise their potency against specific targets. Cytochrome bd oxidases are particularly attractive in this regard since they are only found in prokaryotic organisms. Given the versatile modification chemistry open to madecassic acid,18,19,22,23 further work to maximise its activity against these proteins would be well justified.
Conclusion
Madecassic acid and three derivatives thereof have been modelled to, and shown to have, a high affinity for cytochrome bd-I oxidase, and experimental observations of enzymatic activity, bacterial growth, and bactericidal assays are consistent with this modelling. Alterations of the chemical structure have been shown to influence the biological activity, although the relationships between the different assay outcomes are non-linear. These findings show that inhibition of cytochrome bd oxidases is a further component of the antibacterial activity of madecassic acid, in addition to mechanisms already investigated by others. Further strategic chemical modification could result in new antimicrobial compounds based upon madecassic acid or other similar plant-derived compounds, which are specifically targeted towards the unique aspects of bacterial biochemistry.Author contributions
All authors contributed to writing the manuscript. TTPT extracted and purified the MA from C. asiatica. GNS and HICDS performed organic transformations. SAH was involved in planning and performing the microbiological experiments and data analysis. RAB developed the duroquinol assay, CMW engineered the complementation vector, GJ performed the SDH assays and duroquinol experiments, GKR contributed to the planning of the microbiological experiments and data analysis. CJS and MDG obtained funding and supervised the chemistry. MS also obtained funding and was responsible for all microbiological experimental design and data analysis.Conflicts of interest
There are no conflicts of interest.Data availability
Processed data for this paper is included in the main manuscript. Raw data for this paper (molecular characterisation, assay readout) is available from the authors upon request. Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5md01116g.Acknowledgements
We thank the UK's Engineering and Physical Science Research Council and Global Challenges Research Fund (EP/T020164/1), the UK's Biotechnology and Biological Sciences Research Council (BB/T008741/1; BB/Z517343/1), the University of Kent, and Vietnam Academy of Science and Technology (VAST) for financial support.References
- D. Wojnicz, D. Tichaczek-Goska, K. Korzekwa, M. Kicia and A. Hendrich, Anti-Enterococcal Activities of pentacyclic Triterpenes, Adv. Clin. Exp. Med., 2017, 26(3), 483–490, DOI:10.17219/acem/62245.
- Z. Sycz, D. Tichaczek-Goska, A. Jezierska-Domaradzka and D. Wojnicz, Are Uropathogenic Bacteria Living in Multispecies Biofilm Susceptible to Active Plant Ingredient—Asiatic Acid?, Biomolecules, 2021, 11(12), 1754, DOI:10.3390/biom11121754.
- W.-H. Liu, T.-C. Liu and M.-C. Mong, Antibacterial Effects and Action Modes of Asiatic Acid, Biomedicine, 2015, 5(3), 16, DOI:10.7603/s40681-015-0016-7.
- D. Wojnicz, D. Tichaczek-Goska and M. Kicia, pentacyclic Triterpenes Combined with Ciprofloxacin Help to Eradicate the Biofilm Formed in Vitro by Escherichia coli, Indian J. Med. Res., 2015, 141(3), 343–353, DOI:10.4103/0971-5916.156631.
- C. Wei, P. Cui and X. Liu, Antibacterial Activity and Mechanism of Madecassic Acid against Staphylococcus aureus, Molecules, 2023, 28(4), 1895, DOI:10.3390/molecules28041895.
- G. Liu, P. Qin, X. Cheng, L. Wu, W. Zhao and W. Gao, Evaluation of the Mechanistic Basis for the Antibacterial Activity of Ursolic Acid against Staphylococcus aureus, Front. Microbiol., 2024, 15 DOI:10.3389/fmicb.2024.1389242.
- P. Chonsut, C. Romyasamit, A. Konyanee, N. Niyomtham, L. Goodla and A. Mordmuang, Potential Activities of Centella Asiatica Leaf Extract against Pathogenic Bacteria-Associated Biofilms and Its Anti-Inflammatory Effects, Adv. Pharmacol. Pharm. Sci., 2024, 2024(1), 5959077, DOI:10.1155/2024/5959077.
- Z. Xie, X. Dai and L. Wang, Novel 23-Hydroxybetulinic Acid Derivatives: Semi-Synthesis and Antibacterial Activity, Russ. J. Gen. Chem., 2024, 94(10), 2770–2788, DOI:10.1134/S1070363224100219.
- K. Kandaswamy, S. P. Panda, M. R. Shaik, S. A. Hussain, P. Deepak, N. Thiyagarajulu, D. Jain, A. P. M. Antonyraj, R. Subramanian, A. Guru and J. Arockiaraj, Formulation of Asiatic Acid-Loaded Polymeric Chitosan-Based Hydrogel for Effective MRSA Infection Control and Enhanced Wound Healing in Zebrafish Models, Int. J. Biol. Macromol., 2025, 293, 137425, DOI:10.1016/j.ijbiomac.2024.137425.
- V. B. Borisov, R. B. Gennis, J. Hemp and M. I. Verkhovsky, The Cytochrome bd Respiratory Oxygen Reductases, Biochim. Biophys. Acta, Bioenerg., 2011, 1807(11), 1398–1413, DOI:10.1016/j.bbabio.2011.06.016.
- L. N. Jeffreys, A. Ardrey, T. A. Hafiz, L.-A. Dyer, A. J. Warman, N. Mosallam, G. L. Nixon, N. E. Fisher, W. D. Hong, S. C. Leung, G. Aljayyoussi, J. Bibby, D. V. Almeida, P. J. Converse, N. Fotouhi, N. G. Berry, E. L. Nuermberger, A. M. Upton, P. M. O'Neill, S. A. Ward and G. A. Biagini, Identification of 2-Aryl-Quinolone Inhibitors of Cytochrome Bd and Chemical Validation of Combination Strategies for Respiratory Inhibitors against Mycobacterium tuberculosis, ACS Infect. Dis., 2023, 9(2), 221–238, DOI:10.1021/acsinfecdis.2c00283.
- A. Giuffrè, V. B. Borisov, M. Arese, P. Sarti and E. Forte, Cytochrome bd Oxidase and Bacterial Tolerance to Oxidative and Nitrosative Stress, Biochim. Biophys. Acta, Bioenerg., 2014, 1837(7), 1178–1187, DOI:10.1016/j.bbabio.2014.01.016.
- S. A. Henry, C. M. Webster, L. N. Shaw, N. J. Torres, M.-E. Jobson, B. C. Totzke, J. K. Jackson, J. E. McGreig, M. N. Wass, G. K. Robinson and M. Shepherd, Steroid Drugs Inhibit Bacterial Respiratory Oxidases and Are Lethal Toward Methicillin-Resistant Staphylococcus aureus, J. Infect. Dis., 2024, 230(1), e149–e158, DOI:10.1093/infdis/jiad540.
- T. T. P. Tran, N. T. Pham, N. X. Nguyen, C. V. Tran, L. V. Tran, L. T. Nguyen, D. T. Nguyen, N. T. N. Huynh, T. L. Nguyen, Y. T. H. Be, M. D. Garrett, C. J. Serpell and T. Van Sung, A Comparative Study of Organic and Conventional Production Methods on Biomass Accumulation and Bioactive Compounds Content in Centella asiatica (L.) Urban in Vietnam, Ind. Crops Prod., 2025, 237, 122251, DOI:10.1016/j.indcrop.2025.122251.
- D. Seeliger and B. L. de Groot, Ligand Docking and Binding Site Analysis with PyMOL and Autodock/Vina, J. Comput.-Aided Mol. Des., 2010, 24(5), 417–422, DOI:10.1007/s10822-010-9352-6.
- M.-H. Nicolas-Chanoine, X. Bertrand and J.-Y. Madec, Escherichia coli ST131, an Intriguing Clonal Group, Clin. Microbiol. Rev., 2014, 27(3), 543–574, DOI:10.1128/cmr.00125-13.
- B. M. Forde, N. L. B. Zakour, M. Stanton-Cook, M.-D. Phan, M. Totsika, K. M. Peters, K. G. Chan, M. A. Schembri, M. Upton and S. A. Beatson, The Complete Genome Sequence of Escherichia coli EC958: A High Quality Reference Sequence for the Globally Disseminated Multidrug Resistant E. Coli O25b:H4-ST131 Clone, PLoS One, 2014, 9(8), e104400, DOI:10.1371/journal.pone.0104400.
- T. Van Loc, V. T. Q. Nhu, T. Van Chien, L. T. T. Ha, T. T. P. Thao and T. Van Sung, Synthesis of Madecassic Acid Derivatives and Their Cytotoxic Activity, Z. Naturforsch., B: J. Chem. Sci., 2018, 73(2), 91–98, DOI:10.1515/znb-2017-0172.
- C. V. Tran, T. T. P. Tran, A. T. Nguyen, L. V. Tran, N. T. Pham, L. T. Nguyen, D. T. Nguyen, M. D. Garrett, N. T. Nguyen, T. T. Do, C. J. Serpell and S. V. Tran, Synthesis and Cytotoxic Activity of Madecassic Acid–Silybin Conjugate Compounds in Liver Cancer Cells, RSC Med. Chem., 2024, 15, 3418–3432, 10.1039/D4MD00170B.
- D. J. Newman and G. M. Cragg, Natural Products as Sources of New Drugs from 1981 to 2014, J. Nat. Prod., 2016, 79(3), 629–661, DOI:10.1021/acs.jnatprod.5b01055.
- M. Decker, Hybrid Molecules Incorporating Natural Products: Applications in Cancer Therapy, Neurodegenerative Disorders and Beyond, Curr. Med. Chem., 2011, 18(10), 1464–1475, DOI:10.2174/092986711795328355.
- A. S. C. Valdeira, D. A. Ritt, D. K. Morrison, J. B. McMahon, K. R. Gustafson and J. A. R. Salvador, Synthesis and Biological Evaluation of New Madecassic Acid Derivatives Targeting ERK Cascade Signaling, Front. Chem., 2018, 6, 434, DOI:10.3389/fchem.2018.00434.
- A. S. C. Valdeira, E. Darvishi, G. M. Woldemichael, J. A. Beutler, K. R. Gustafson and J. A. R. Salvador, Madecassic Acid Derivatives as Potential Anticancer Agents: Synthesis and Cytotoxic Evaluation, J. Nat. Prod., 2019, 82(8), 2094–2105, DOI:10.1021/acs.jnatprod.8b00864.
| This journal is © The Royal Society of Chemistry 2026 |