Open Access Article
Open Access ArticleExploration of a benzothiophene scaffold for use as adjuvants with β-lactam antibiotics against methicillin-resistant Staphylococcus aureus
Monica A. Stefaniaka,
Vijay S. Gondilb,
Emily P. Gillisa,
Ansley M. Nemetha,
Allen G. Oliver a,
Roberta J. Melandera,
Paul M. Dunman*b and
Christian Melander
a,
Roberta J. Melandera,
Paul M. Dunman*b and
Christian Melander *a
*a
aDepartment of Chemistry and Biochemistry, University of Notre Dame, Notre Dame, IN 46556, USA. E-mail: cmelande@nd.edu
bDepartment of Microbiology and Immunology, University of Rochester Medical Center, Rochester, New York 14642, USA. E-mail: paul_dunman@urmc.rochester.edu
First published on 5th March 2026
Abstract
Rising antibiotic resistance coupled with diminishing investment in antibiotic development has led to limited treatment options for multi drug-resistant bacterial infections. Penicillin binding protein 2a (PBP2a) is a key determinant of resistance to many β-lactam antibiotics in methicillin-resistant Staphylococcus aureus (MRSA), while over-expression of PBP4 mediates resistance to fifth-generation cephalosporins. We previously identified benzothiophene NDM-335 as an inhibitor of PBP4 that overcomes PBP4 mediated resistance to ceftobiprole, and have shown that other PBP4 inhibitor scaffolds can be tuned to overcome PBP2a-mediated resistance. Based upon this precedent, we conducted a structure activity relationship (SAR) study on the NDM-335 scaffold to uncover compounds that overcome PBP2a-mediated oxacillin resistance in MRSA, with the lead compound lowering the minimum inhibitory concentration 32-fold at 5 μM. Further development of this new adjuvant lead could potentially deliver a new combination therapy for treating MRSA infections.
1. Introduction
Antibiotic resistance is a well-established global health threat that was exacerbated during the coronavirus disease pandemic in 2019 (COVID-19).1 The Centers for Disease Control and Prevention (CDC) reported that almost 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 people died from antimicrobial resistant bacterial infections during the pandemic's first year, with 40% of these deaths caused by nosocomial infections.1 Staphylococcus aureus is one of the six ESKAPE pathogens (Enterococcus faecium, S. aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and the Enterobacter species), a group of bacterial pathogens that are frequently associated with hospital associated infections, and are notorious for their multidrug resistance.2 The CDC reported in July 2024 that infections caused by ESKAPE pathogens increased by a combined 20% during the pandemic compared to the pre-pandemic period.1,3 Although the rate of methicillin resistant S. aureus (MRSA) infections returned to pre-pandemic levels in 2022, continued stewardship and antibiotic development are required to control MRSA infections.3
000 people died from antimicrobial resistant bacterial infections during the pandemic's first year, with 40% of these deaths caused by nosocomial infections.1 Staphylococcus aureus is one of the six ESKAPE pathogens (Enterococcus faecium, S. aureus, Klebsiella pneumoniae, Acinetobacter baumannii, Pseudomonas aeruginosa, and the Enterobacter species), a group of bacterial pathogens that are frequently associated with hospital associated infections, and are notorious for their multidrug resistance.2 The CDC reported in July 2024 that infections caused by ESKAPE pathogens increased by a combined 20% during the pandemic compared to the pre-pandemic period.1,3 Although the rate of methicillin resistant S. aureus (MRSA) infections returned to pre-pandemic levels in 2022, continued stewardship and antibiotic development are required to control MRSA infections.3
The introduction of new antibiotics has historically been employed to combat multidrug resistant (MDR) bacterial threats; however, the rapid emergence of resistance, coupled with large pharmaceutical companies' exit from antibiotic development has prompted orthogonal approaches to address this problem.4 One solution is the use of adjuvants, which are compounds that do not possess standalone antimicrobial properties, but potentiate the activity of partner antibiotics by circumventing bacterial resistance mechanisms.5 The most well-known, and only clinically approved adjuvants to date, are serine β-lactamase inhibitors, such as clavulanic acid which is paired with amoxicillin and marketed as Augmentin.6 In addition to Augmentin, avibactam is a non-β-lactam serine β-lactamase inhibitor that was clinically approved in 2015 and is administered in combination with ceftazidime.7 Outside of β-lactamase inhibitors, many other resistance mechanisms represent potential targets for the adjuvant approach.8
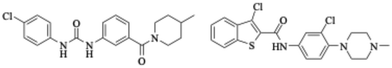
Penicillin binding proteins (PBPs) are transpeptidases/transglycosylases that play a role in the formation of the peptidoglycan component of the bacterial cell wall. S. aureus encodes four native PBPs; two essential (PBP1 and PBP2) and two non-essential (PBP3 and PBP4).9–11 While resistance to β-lactams in MRSA predominantly occurs through the non-native PBP2a (encoded by mecA), which has a lower affinity for most β-lactams than do native PBPs, PBP2a-independent β-lactam resistance has also been reported.12–14 This latter resistance is derived from the overexpression of PBP4, which confers resistance to the fifth-generation cephalosporins, ceftobiprole (BPR) and ceftaroline, which are otherwise active against MRSA.15 This discovery prompted us to screen for potentiators of BPR in a PBP4 overexpressing strain of S. aureus to identify potential inhibitors of PBP4.16 One hit from this screen was the phenyl urea ChemBridge 9314848 (Fig. 1), which inhibits binding of a fluorescently labeled penicillin molecule (Bocillin-FL) to PBP4 and, interestingly, PBP2a.17 Compound 9314848 also potentiates β-lactams against MRSA strains with PBP2a-derived resistance, and analog synthesis led to the identification of compounds with increased activity with select penicillins and cephalosporins.18
Another compound identified from the screen for PBP4 inhibitors was NDM-335 (ChemBridge 7974147) (Fig. 1).16 At 25 μM, NDM-335 lowers the minimum inhibitory concentration (MIC) of BPR to 8 μg mL−1 against the BPR-resistant, PBP4 over-expressor CRB without affecting expression of pbp4.16 CRB is a derivative of the MRSA strain COL that lacks the mecA gene coding for PBP2a, and over-expresses PBP4 leading to broad resistance to β-lactams, including fifth generation cephalosporins.13 Given that ChemBridge 9314848 could be tuned to reduce β-lactam resistance in PBP2a-expressing strains with normal PBP4 expression levels, we elected to further explore the potential of NDM-335 and analogs as antibiotic adjuvants targeting PBP2a-dependent resistance.
2. Results and discussion
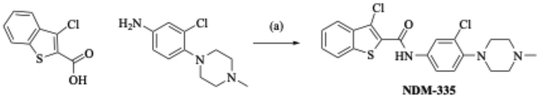
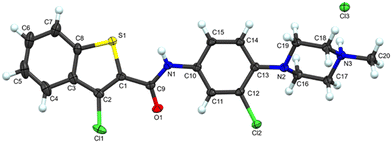
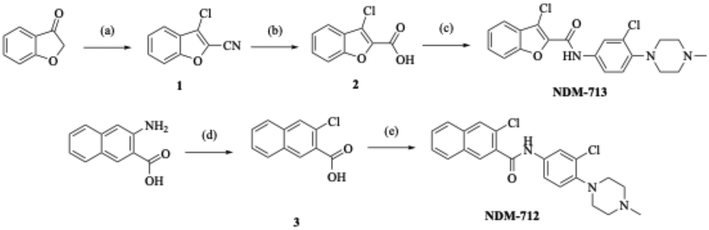
We first resynthesized NDM-335 via a 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) hydrochloride mediated coupling of commercially available 3-chlorobenzothiophene-2-carboxylic acid and 3-chloro-4-(4-methylpiperazin-1-yl)aniline (Scheme 1).19 We further verified the structure of NDM-335 by solving its crystal structure (Fig. 2).We first determined the standalone MIC of NDM-335 against the MRSA USA300 strain BAA-1556 (PBP2a/normal PBP4 expression)20 under standard microdilution assay conditions to be >200 μM. We began a series of oxacillin potentiation assays, which are commonly used to identify other inhibitors of PBP2a related pathways.21,22 At 60 μM (<30% standalone MIC), NDM-335 lowers the oxacillin MIC against this strain from 32 μg mL−1 to 0.125 μg mL−1 (256-fold). Against MRSA, the Clinical Laboratory Standards Institute (CLSI) breakpoint for oxacillin is 2 μg mL−1.23 We sequentially performed a dose–response study to determine the lowest concentration of NDM-335 that lowers the oxacillin MIC to or below the clinical breakpoint, and term this the adjuvant's minimum effective concentration (MEC). Against BAA-1556, at 7.5 μM, NDM-335 lowers the oxacillin MIC to 0.25 μg mL−1, while at 3.75 μM, it suppresses the MIC to 8 μg mL−1 (Table 1), indicating the MEC lies between 7.5 and 3.75 μM. We then evaluated NDM-335 against two additional MRSA strains: ATCC 43300 and AH-1263 (standalone oxacillin MICs of 32 μg mL−1 for each strain), both of which also harbor the mecA gene and show normal PBP4 expression levels. NDM-335 lowers the oxacillin MIC to 0.5 μg mL−1 at 5 μM in ATCC 43300, and to 0.25 μg mL−1 at 7.5 μM in AH-1263. As a control, we measured potentiation of BPR by NDM-335 against BAA-1556. In this strain, the BPR MIC is 1 μg mL−1, and this is only reduced four-fold in the presence of 12.5 μM (and concentrations up to 200 μM) NDM-335, indicating strong specificity for PBP2a-dependent resistance.
| NDM-335 standalone MIC (μM) | NDM-335 concentration (μM) | Oxacillin MIC (μg mL−1) [fold reduction] |
|---|---|---|
| MRSA ATCC BAA-1556 | ||
| >200 | — | 32 |
| 60 | 0.125 [256] | |
| 30 | 0.125 [256] | |
| 15 | 0.125 [256] | |
| 7.5 | 0.25 [128] | |
| 3.75 | 8 [4] | |
| MRSA ATCC 43300 | ||
| >200 | — | 32 |
| 60 | 0.125 [256] | |
| 30 | 0.125 [256] | |
| 15 | 0.125 [256] | |
| 7.5 | 0.125 [256] | |
| 5 | 0.5 [64] | |
| 4 | 4 [8] | |
| MRSA AH-1263 | ||
| >200 | — | 32 |
| 60 | 0.125 [256] | |
| 30 | 0.25 [128] | |
| 15 | 0.25 [128] | |
| 7.5 | 0.25 [128] | |
| 5 | 8 [4] | |
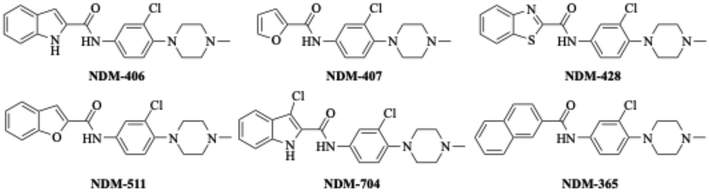
With these promising results, we initiated a structure activity relationship (SAR) study of the benzothiophene region of NDM-335 to determine whether compounds with enhanced adjuvant activity could be identified. To this end, we synthesized a library of compounds in which the benzothiophene was replaced with alternative heterocycles (or a naphthyl ring), while conserving the 3-chloro-4-(4-methylpiperazine) moiety. Following the synthetic route outlined in Scheme 1, the indole, 3-chloroindole, benzothiazole, furan, benzofuran, and naphthalene derivatives (Fig. 3) were synthesized from the respective commercially available carboxylic acids. Indole, 3-chloroindole, benzothiazole, benzofuran, and naphthalene were selected for their structural similarities to benzothiophene, and to analyze the impact of altering the heteroatom of the benzothiophene head group. The furan was chosen to assess the necessity of the benzyl ring and the sulfur atom.
The 3-chlorobenzofuran (NDM-713) analog required a more complex synthesis (Scheme 2), starting with subjecting benzofuran-3(2H)-one to Vilsmeier–Haack conditions24 to provide 3-chlorobenzofuran-2-carbonitrile, 1. The nitrile was then treated with sodium hydroxide to provide key intermediate 2, which was then reacted with 3-chloro-4-(4-methylpiperazin-1-yl)aniline via an EDC-mediated coupling to yield NDM-713. To access the 2-chloronapthalene (NDM-712) derivative (Scheme 2), 3-amino-2-naphthoic acid was subjected to Sandmeyer conditions to provide 3-chloro-2-naphthoic acid, 3.25 3-Chloro-2-naphthoic acid was then reacted with 3-chloro-4-(4-methylpiperazin-1-yl)aniline via an EDC coupling to provide amide NDM-712.
The stand-alone MIC of each compound was first determined against the same three MRSA strains described above (Table 2), with all compounds, except the 3-chloroindole derivative NDM-704, returning MICs of 100 to >200 μM. Due to low solubility of NDM-704 in stock solution of DMSO, it was tested at ten-fold lower starting concentrations and returned a stand-alone MIC greater than 20 μM in BAA-1556, 43300, and AH-1263. Oxacillin MICs in the presence of each analog are summarized in Table 2. The naphthyl (NDM-365), indole (NDM-406), and furan (NDM-407) derivatives fail to potentiate oxacillin against BAA-1556 and AH-1263, returning unchanged oxacillin MICs of 32 μg mL−1 at 60 μM. Against 43300, NDM-365 and NDM-407 exhibit minimal activity, lowering the oxacillin MIC only four-fold to 8 μg mL−1 at 60 μM. Against this same strain, NDM-406 lowers the oxacillin MIC 32-fold to 1 μg mL−1 at 60 μM, and 64-fold to 0.5 μg mL−1 at 30 μM, but lost significant activity at 15 μM with an oxacillin MIC of 4 μg mL−1 (eight-fold reduction). Against BAA-1556, the benzofuran derivative (NDM-511) lowers the oxacillin MIC 32-fold to 1 μg mL−1 at 60 μM, and four-fold (8 μg mL−1) at 30 μM. Against 43300, NDM-511 returns an oxacillin MIC of 0.25 μg mL−1 at 60 μM, which increases to 4 μg mL−1 at 30 μM. Against AH-1263, NDM-511 suppresses the oxacillin MIC to 0.5 μg mL−1 at 60 μM, which increases to 8 μg mL−1 at 30 μM. Against BAA-1556 and 43300, NDM-704 is inactive at 6 μM, returning oxacillin MICs of 16 μg mL−1, an insignificant two-fold drop from the standalone oxacillin MIC. In AH-1263, NDM-704 delivers an eight-fold reduction in oxacillin MIC to 4 μg mL−1 at 6 μM. The 2-chloronaphthyl derivative (NDM-712) lowers the oxacillin MIC to 16 μg mL−1 and 8 μg mL−1 at 60 μM, in BAA-1556 and 43300, respectively. NDM-712 reduces the oxacillin MIC to the breakpoint of 2 μg mL−1 at 60 μM in AH-1263. and to 4 μg mL−1 at 30 μM. The 3-chlorobenzofuran derivative (NDM-713) suppresses the oxacillin MIC to 0.125 μg mL−1, 0.125 μg mL−1, and 0.25 μg mL−1 against BAA-1556, 43300, and AH-1263, respectively, at 30 μM. At 15 μM, the oxacillin MIC increases to 32 μg mL−1, 16 μg mL−1, and 16 μg mL−1 against these three strains. Finally, the benzothiazole derivative (NDM-428) lowers the MIC of oxacillin below the breakpoint at 5 μM in BAA-1556 (to 2 μg mL−1) and at 7.5 μM in AH-1263 (to 0.5 μg mL−1) but interestingly is not active in 43300 (returning an oxacillin MIC of 16 μg mL−1). From this, it appears that the presence of a sulfur atom at the one-position of the heterocycle is important for oxacillin adjuvant activity.
| Compound | Standalone MIC (μM) | Concentration (μM) | Oxacillin MIC (μg mL−1) [fold reduction] |
|---|---|---|---|
| MRSA ATCC BAA-1556 | |||
| — | — | — | 32 |
| NDM-365 | >200 | 60 | 32 [−] |
| NDM-406 | >200 | 60 | 32 [−] |
| NDM-407 | >200 | 60 | 32 [−] |
| NDM-428 | >200 | 60 | 2 [16] |
| 30 | 1 [32] | ||
| 15 | 1 [32] | ||
| 7.5 | 1 [32] | ||
| 5 | 2 [16] | ||
| 4 | 4 [8] | ||
| NDM-511 | >200 | 60 | 1 [32] |
| 30 | 8 [4] | ||
| NDM-704 | >20 | 6 | 16 [2] |
| NDM-712 | >200 | 60 | 16 [2] |
| NDM-713 | 100 | 30 | 0.125 [256] |
| 15 | 32 [−] | ||
| MRSA ATCC 43300 | |||
| — | — | — | 32 |
| NDM-365 | >200 | 60 | 8 [4] |
| NDM-406 | >200 | 60 | 1 [32] |
| 30 | 0.5 [64] | ||
| 15 | 4 [8] | ||
| NDM-407 | >200 | 60 | 8 [4] |
| NDM-428 | >200 | 60 | 16 [2] |
| NDM-511 | 200 | 60 | 0.25 [128] |
| 30 | 4 [8] | ||
| NDM-712 | >200 | 60 | 8 [4] |
| NDM-713 | 100 | 30 | 0.125 [256] |
| 15 | 16 [2] | ||
| NDM-704 | >20 | 6 | 16 [2] |
| MRSA AH-1263 | |||
| — | — | — | 32 |
| NDM-365 | >200 | 60 | 32 [−] |
| NDM-406 | >200 | 60 | 32 [−] |
| NDM-407 | >200 | 60 | 32 [−] |
| NDM-428 | >200 | 60 | 0.5 [64] |
| 30 | 0.5 [64] | ||
| 15 | 0.25 [128] | ||
| 7.5 | 0.5 [64] | ||
| 5 | 8 [4] | ||
| NDM-511 | >200 | 60 | 0.5 [64] |
| 30 | 8 [4] | ||
| NDM-712 | >200 | 60 | 2 [16] |
| 30 | 4 [8] | ||
| NDM-713 | 100 | 30 | 0.25 [128] |
| 15 | 16 [2] | ||
| NDM-704 | >20 | 6 | 4 [8] |
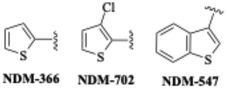
Based upon the importance of the sulfur atom, we next investigated compounds with various sulfur containing heterocycles (Fig. 4). Each derivative was synthesized from the respective carboxylic acid using the approach outlined in Scheme 1. Analogs synthesized include: NDM-366 that contains a thiophene, NDM-702 with a 3-chlorothiophene, and NDM-547 that harbors a benzothiophene group attached at the 3-position, rather than the 2-position as seen in NDM-335.
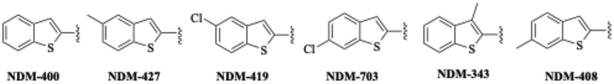
The thiophene derivatives NDM-366 and NDM-702 have standalone MICs greater than 200 μM against all three strains, while NDM-547 exhibits a standalone MIC of 100 μM against each strain (Table 3). NDM-702 is inactive at 60 μM, returning oxacillin MICs of 16 μg mL−1 against all three strains. NDM-366 is inactive against BAA-1556, and AH-1263, and only lowers the oxacillin MIC four-fold to 8 μg mL−1 against 43300 at 60 μM. NDM-547 returns an oxacillin MIC of 0.5 μg mL−1 at 30 μM in BAA-1556, but loses activity at lower concentrations, only reducing the oxacillin MIC to 4 μg mL−1 at 15 μM. Against 43300, NDM-547 lowers the oxacillin MIC to 1 μg mL−1 at 15 μM, but fails to reach breakpoint levels at 10 μM, returning an MIC of 4 μg mL−1. In AH-1263, NDM-547 lowers the MIC of oxacillin to 0.5 μg mL−1 at 30 μM, and to 16 μg mL−1 at 15 μM. In aggregate, these data underscore the importance of a benzothiophene motif for activity, with connection at the C-2 position delivering the highest activity. To follow up on this observation, additional C-2 substituted benzothiophenes were synthesized (Fig. 5) again via the route shown in Scheme 1. These include: the unsubstituted analog (NDM-400), 5-methylbenzothiophene (NDM-427), 5-chlorobenzothiophene (NDM-419), 6-chlorobenzothiophene (NDM-703), 3-methylbenzothiophene (NDM-343), and 6-methylbenzothiophene (NDM-408).
| Compound | Standalone MIC (μM) | Concentration (μM) | Oxacillin MIC (μg mL−1) [fold reduction] |
|---|---|---|---|
| MRSA ATCC BAA-1556 | |||
| — | — | — | 32 |
| NDM-366 | >200 | 60 | 32 [−] |
| NDM-547 | 100 | 30 | 0.5 [64] |
| 15 | 4 [8] | ||
| NDM-702 | >200 | 60 | 16 [2] |
| MRSA ATCC 43300 | |||
| — | — | — | 32 |
| NDM-366 | >200 | 60 | 8 [4] |
| NDM-547 | 100 | 30 | 0.5 [64] |
| 15 | 1 [32] | ||
| 10 | 4 [8] | ||
| 7.5 | 4 [8] | ||
| NDM-702 | >200 | 60 | 16 [2] |
| MRSA AH-1263 | |||
| — | — | — | 32 |
| NDM-366 | >200 | 60 | 32 [−] |
| NDM-547 | 100 | 30 | 0.5 [64] |
| 15 | 16 [2] | ||
| NDM-702 | >200 | 60 | 16 [2] |
Standalone MIC and adjuvant activity for these analogs is summarized in Table 4. The 5-chlorobenzothiophene analog NDM-419 is inactive at 60 μM against BAA-1556 and AH-1263, while it is only moderately active against 43300, returning oxacillin MICs of 32 μg mL−1, 16 μg mL−1, and 8 μg mL−1, respectively. The 6-chloro derivative NDM-703 exhibits a similar lack of activity, returning oxacillin MICs of 16 μg mL−1, 8 μg mL−1, and 16 μg mL−1 against BAA-1556, 43300, and AH-1263, respectively at 60 μM. The 6-methyl derivative NDM-408 is moderately active against BAA-1556 and AH-1263, lowering the oxacillin MIC four-fold to 8 μg mL−1 at 60 μM; however, it exhibits greater activity against 43300, lowering the oxacillin MIC to 2 μg mL−1 at 5 μM. The unsubstituted derivative NDM-400 lowers the oxacillin MIC to 1 μg mL−1 at 7.5 μM and 2 μg mL−1 at 5 μM in BAA-1556 and 43300, respectively, and to 0.5 μg mL−1 at 15 μM against AH-1263. The 5-methyl derivative NDM-427 retains activity at 7.5 μM against all three strains, lowering the oxacillin MIC to 2 μg mL−1. As a result of the initial SAR study, the parent NDM-335, the unsubstituted benzothiophene NDM-400, the 6-methylbenzothiophene NDM-408, the 5-methylbenzothiophene NDM-427, and the benzothiazole NDM-428 were subject to further investigation. NDM-335, NDM-408, NDM-427, and NDM-428 were chosen as lead compounds because they potentiate oxacillin at concentrations of 10 μM or lower in at least two out of the three strains tested. While NDM-400 does not achieve this same standard, we chose it as a lead compound due to its activity at 5 μM in MRSA 43300 and 15 μM in MRSA AH-1263.
| Compound | Standalone MIC (μM) | Concentration (μM) | Oxacillin MIC (μg mL−1) [fold reduction] |
|---|---|---|---|
| MRSA ATCC BAA-1556 | |||
| — | — | — | 32 |
| NDM-408 | >200 | 60 | 8 [4] |
| NDM-343 | >200 | 60 | 2 [16] |
| 30 | 2 [16] | ||
| 15 | 4 [8] | ||
| NDM-400 | >200 | 60 | 0.125 [256] |
| 30 | 0.125 [256] | ||
| 15 | 0.25 [128] | ||
| 7.5 | 1 [32] | ||
| 5 | 16 [2] | ||
| NDM-419 | >200 | 60 | 32 [−] |
| NDM-427 | >200 | 60 | 0.25 [128] |
| 30 | 0.5 [64] | ||
| 15 | 1 [32] | ||
| 7.5 | 2 [16] | ||
| 5 | 8 [4] | ||
| NDM-703 | >200 | 60 | 16 [2] |
| MRSA ATCC 43300 | |||
| — | — | — | 32 |
| NDM-343 | >200 | 60 | 0.25 [128] |
| 30 | 2 [16] | ||
| 15 | 4 [8] | ||
| NDM-400 | >200 | 60 | 0.125 [256] |
| 30 | 0.125 [256] | ||
| 15 | 0.5 [64] | ||
| 7.5 | 2 [16] | ||
| 5 | 2 [16] | ||
| 4 | 4 [8] | ||
| NDM-408 | >200 | 60 | 0.25 [256] |
| 30 | 0.25 [256] | ||
| 15 | 0.25 [256] | ||
| 7.5 | 0.5 [64] | ||
| 5 | 2 [16] | ||
| 4 | 4 [8] | ||
| NDM-419 | >200 | 60 | 8 [4] |
| NDM-427 | >200 | 60 | 0.125 [256] |
| 30 | 0.125 [256] | ||
| 15 | 0.125 [256] | ||
| 7.5 | 2 [16] | ||
| 5 | 4 [8] | ||
| NDM-703 | >200 | 60 | 8 [4] |
| MRSA AH-1263 | |||
| — | — | — | 32 |
| NDM-343 | >200 | 60 | 16 [2] |
| NDM-400 | >200 | 60 | 0.125 [256] |
| 30 | 0.25 [128] | ||
| 15 | 0.5 [64] | ||
| 10 | 16 [2] | ||
| NDM-408 | >200 | 60 | 8 [4] |
| NDM-419 | >200 | 60 | 16 [2] |
| NDM-427 | >200 | 60 | 0.125 [256] |
| 30 | 0.5 [64] | ||
| 15 | 0.5 [64] | ||
| 7.5 | 2 [16] | ||
| 5 | 8 [4] | ||
| NDM-703 | >200 | 60 | 16 [2] |
Lead compounds were tested against the methicillin sensitive S. aureus (MSSA) strain ATCC 29213 as a control (Table 5). MSSA 29213 lacks PBP2a, leading us to predict that potentiation activity would be lost. NDM-400, NDM-408, NDM-427, and NDM-428 lower the MIC of oxacillin to 0.125 μg mL−1 at 60 μM, which is two-fold lower than the standalone oxacillin MIC of 0.25 μg mL−1 in MSSA 29213. MIC changes of two fold are considered statistically insignificant, thus strongly indicating that the potentiation activity is related to PBP2a, or other regulatory networks involved in PBP2a-mediated resistance.
| Compound | Standalone MIC (μM) | Concentration (μM) | Oxacillin MIC (μg mL−1) [fold reduction] |
|---|---|---|---|
| MSSA ATCC 29213 | |||
| — | — | — | 0.25 |
| NDM-335 | >200 | 60 | 0.125 [2] |
| NDM-400 | >200 | 60 | 0.125 [2] |
| NDM-408 | >200 | 60 | 0.125 [2] |
| NDM-427 | >200 | 60 | 0.125 [2] |
| NDM-428 | >200 | 60 | 0.125 [2] |
Following the study of benzothiophene derivatives in combination with oxacillin, activity of lead compounds with two additional β-lactams: penicillin G (Table 6), and the second-generation cephalosporin, cefoxitin (Table 7) was explored. In BAA-1556 and MRSA 43300, three of the five compounds (NDM-408, NDM-427, and NDM-428) did not reduce the MIC of penicillin G to its clinical breakpoint of 0.125 μg mL−1 at 60 μM (Table 7).23 NDM-335 lowers the penicillin G MIC to 0.0625 at 60 μM, 30 μM, and 15 μM, but fails to achieve breakpoint levels at 10 μM. NDM-400 returns a penicillin G MIC of 0.015625 μg mL−1 at 60 μM and 0.125 μg mL−1 at 30 μM but fails to achieve breakpoint levels at 15 μM. In MRSA 43300, NDM-335, NDM-408, NDM-427, and NDM-428 do not achieve breakpoint level MICs when dosed at 60 μM. NDM-400 lowers the penicillin G MIC to 0.03125 μg mL−1 at 60 μM, but the MIC increases to 0.5 μg mL−1 when dosed at 30 μM. Against MRSA AH-1263, NDM-408, NDM-427, and NDM-428 return MICs greater than the penicillin G breakpoint of 0.125 μg mL−1 at 60 μM. NDM-335 and NDM-400 return penicillin G MICs of 0.125 μg mL−1 and 0.015625 μg mL−1 respectively at 60 μM. Both compounds return penicillin MICs of 0.125 μg mL−1 at 30 μM, but activity is lost at 15 μM. At 15 μM, NDM-335 and NDM-400 return penicillin G MICs of 4 μg mL−1.
| Compound | Standalone MIC (μM) | Concentration (μM) | Penicillin G MIC (μg mL−1) [fold reduction] |
|---|---|---|---|
| MRSA ATCC BAA-1556 | |||
| — | — | — | 4 |
| NDM-335 | >200 | 60 | 0.0625 [64] |
| 30 | 0.0625 [64] | ||
| 15 | 0.0625 [64] | ||
| 10 | 0.5 [8] | ||
| NDM-400 | >200 | 60 | 0.015625 [256] |
| 30 | 0.125 [32] | ||
| 15 | 0.25 [16] | ||
| NDM-408 | >200 | 60 | 2 [2] |
| NDM-427 | >200 | 60 | 0.25 [16] |
| NDM-428 | >200 | 60 | 2 [2] |
| MRSA ATCC 43300 | |||
| — | — | — | 4 |
| NDM-335 | >200 | 60 | 0.25 [16] |
| NDM-400 | >200 | 60 | 0.03125 [128] |
| 30 | 0.5 [8] | ||
| NDM-408 | >200 | 60 | 8 [−] |
| NDM-427 | >200 | 60 | 1 [4] |
| NDM-428 | >200 | 60 | 8 [−] |
| MRSA AH-1263 | |||
| — | — | — | 4 |
| NDM-335 | >200 | 60 | 0.125 [32] |
| 30 | 0.125 [32] | ||
| 15 | 4 [−] | ||
| NDM-400 | >200 | 60 | 0.015625 [256] |
| 30 | 0.125 [32] | ||
| 15 | 4 [−] | ||
| NDM-408 | >200 | 60 | 2 [2] |
| NDM-427 | >200 | 60 | 0.25 [16] |
| NDM-428 | >200 | 60 | 2 [2] |
| Compound | Standalone MIC (μM) | Concentration (μM) | Cefoxitin MIC (μg mL−1) [fold reduction] |
|---|---|---|---|
| MRSA ATCC BAA-1556 | |||
| — | — | — | 32 |
| NDM-335 | >200 | 60 | 16 [2] |
| NDM-400 | >200 | 60 | 0.125 [256] |
| >200 | 30 | 16 [2] | |
| NDM-408 | >200 | 60 | 16 [2] |
| NDM-427 | >200 | 60 | 16 [2] |
| NDM-428 | >200 | 60 | 32 [−] |
| MRSA ATCC 43300 | |||
| — | — | — | 32 |
| NDM-335 | >200 | 60 | 32 [−] |
| NDM-400 | >200 | 60 | 0.5 [64] |
| 30 | 16 [2] | ||
| NDM-408 | >200 | 60 | 32 [−] |
| NDM-427 | >200 | 60 | 16 [2] |
| NDM-428 | >200 | 60 | 32 [−] |
| MRSA AH-1263 | |||
| — | — | — | 32 |
| NDM-335 | >200 | 60 | 16 [2] |
| NDM-400 | >200 | 60 | 16 [2] |
| NDM-408 | >200 | 60 | 16 [2] |
| NDM-427 | >200 | 60 | 8 [4] |
| NDM-428 | >200 | 60 | 32 [−] |
Lead compounds were tested with cefoxitin. Against MRSA BAA-1556, NDM-335, NDM-408, NDM-427, and NDM-428 fail to lower the cefoxitin MIC to its clinical breakpoint level of 4 μg mL−1.23 NDM-335 lowers the cefoxitin MIC to 0.125 μg mL−1 at 60 μM, and 16 μg mL−1 at 30 μM. Against MRSA 43300, NDM-335, NDM-408, NDM-427, and NDM-428 fail to lower the cefoxitin MIC to breakpoint levels. NDM-400 lowers the cefoxitin MIC to 0.5 μg mL−1 at 60 μM and 16 μg mL−1 at 30 μM, showing a loss of activity at 30 μM. In MRSA AH-1263, NDM-400, NDM-408, NDM-427, and NDM-428 fail to lower the cefoxitin MIC to breakpoint levels. Taken together, these results are similar to other adjuvants we have developed, where there appears to be an optimal antibiotic partner within the general antibiotic class.18,26,27
Growth curves were then constructed for NDM-335, NDM-400, and NDM-427 using MRSA ATCC 43300 as the test strain (Fig. 6–8). Each compound at its MEC in combination with oxacillin at the breakpoint concentration (2 μg mL−1) was examined, along with 2× and 4× this dosing. Based on the changes in colony forming units (CFUs), the combination of NDM-335, NDM-400, and NDM-427 with oxacillin was determined to be bacteriostatic.28
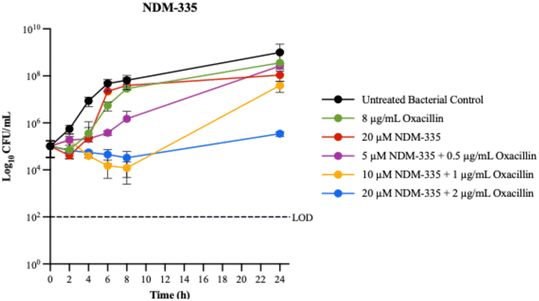 | ||
| Fig. 6 MRSA 43300 growth curves for NDM-335 and oxacillin. Data ± SD from at least three replicates. | ||
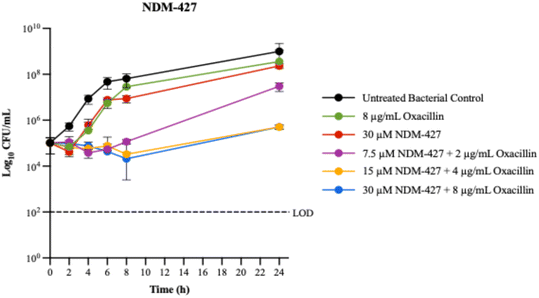 | ||
| Fig. 7 MRSA 43300 growth curves for NDM-427 and oxacillin. Data ± SD from at least three replicates. | ||
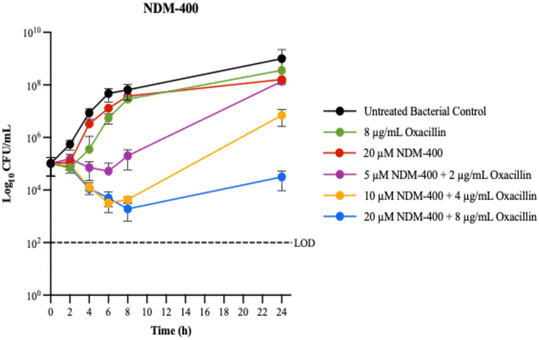 | ||
| Fig. 8 MRSA 43300 growth curves for NDM-400 and oxacillin. Data ± SD from at least three replicates. | ||
S. aureus codes for a repertoire of two component regulatory systems (TCS), including VraRS, ArlRS, WalKR, BlaIR, GraRS, SaeRS, that have been shown to directly or indirectly modulate resistance to cell wall active antibiotics.29 Thus, to begin a preliminary investigation into the mechanism of action, lead compounds were examined against a panel of deletion strains that each have one of the 15 nonessential S. aureus two component systems deleted, as well as a mecA transposon mutant, and activity was compared to that against the parent (MRSA AH-1263).30 First, the oxacillin MIC was determined for each strain (Table 8). Four strains have significantly lower oxacillin MICs than the parent strain, indicating involvement of these gene products in oxacillin resistance. AH 2360 (ΔgraRS),30 AH 2087 (ΔvraRS),30 AH 2090 (ΔkdpDE),30 and AH 5929 (mecA:Tn mutant)31 have oxacillin MICs of 0.5 μg mL−1 (64-fold), 4 μg mL−1 (eight-fold), 8 μg mL−1 (four-fold), and 0.25 μg mL−1 (128-fold), respectively. In all four strains, each compound has a standalone MIC >200 μM. Against the graRS mutant, no compounds potentiate the activity of oxacillin (MIC lowered no more than two-fold), suggesting that GraRS is required for adjuvant activity. Against the vraRS deletion strain AH 2087, all compounds further potentiate the activity of oxacillin, implying that the VraRS regulatory system is not the target of this series of compounds. Against the kdpDE deletion strain AH 2090, only NDM-428 does not further potentiate oxacillin (MIC lowered only two-fold), indicating that KdpDE is not the general target for this series of compounds, but further investigation is required to determine if it plays a role in the activity of NDM-428. Finally, against the mecA transposon mutant AH 5929, no compound further potentiates the activity of oxacillin, suggesting that PBP2a is also required for adjuvant activity. In aggregate, this indicates that the potential targets for these compounds are involved in the GraRS two component signaling system and/or PBP2a (in both cases either direct binding or downregulation of the genes).
| Compound | Standalone MIC (μM) | Concentration (μM) | Oxacillin MIC (μg mL−1) [fold reduction] |
|---|---|---|---|
| AH 1263 | |||
| — | — | — | 32 |
| NDM-335 | >200 | 7.5 | 0.25 [128] |
| NDM-400 | >200 | 15 | 0.5 [64] |
| NDM-408 | >200 | 60 | 8 [4] |
| NDM-427 | >200 | 7.5 | 2 [16] |
| NDM-428 | >200 | 7.5 | 0.5 [64] |
| AH 5929 (mecA:Tn) | |||
| — | — | — | 0.25 |
| NDM-335 | >200 | 7.5 | 0.125 [2] |
| NDM-400 | >200 | 15 | 0.125 [2] |
| NDM-408 | >200 | 60 | 0.125 [2] |
| NDM-427 | >200 | 7.5 | 0.125 [2] |
| NDM-428 | >200 | 7.5 | 0.25 [−] |
| AH 2087 ΔvraRS | |||
| — | — | — | 4 |
| NDM-335 | >200 | 7.5 | 0.5 [8] |
| NDM-400 | >200 | 15 | 0.125 [32] |
| NDM-408 | >200 | 60 | 1 [4] |
| NDM-427 | >200 | 7.5 | 0.25 [16] |
| NDM-428 | >200 | 7.5 | 1 [4] |
| AH 2090 ΔkdpDE | |||
| — | — | — | 8 |
| NDM-335 | >200 | 7.5 | 0.25 [32] |
| NDM-400 | >200 | 15 | 0.25 [32] |
| NDM-408 | >200 | 60 | 0.5 [16] |
| NDM-427 | >200 | 7.5 | 0.25 [32] |
| NDM-428 | >200 | 7.5 | 4 [2] |
| AH 2360 ΔgraRS | |||
| — | — | — | 0.5 |
| NDM-335 | >200 | 7.5 | 0.5 [−] |
| NDM-400 | >200 | 15 | 0.5 [−] |
| NDM-408 | >200 | 60 | 0.5 [−] |
| NDM-427 | >200 | 7.5 | 0.25 [2] |
| NDM-428 | >200 | 7.5 | 0.5 [−] |
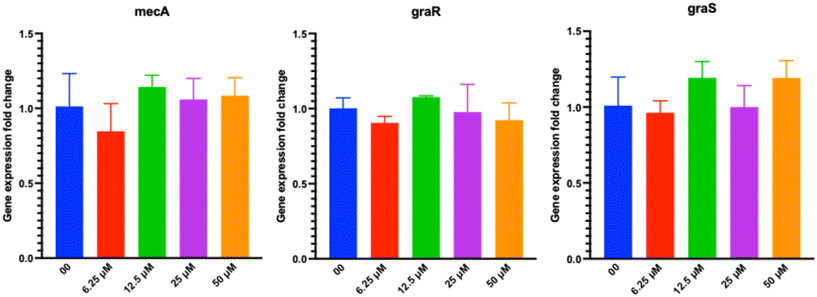
To follow up on these studies, we performed qRT-PCR to determine if the mechanism of the compounds was underpinned by downregulation of mecA, graR, or graS using NDM-335 and BAA-1556 as our representative adjuvant/strain combination (Fig. 9). At all concentrations studied (6.25–50 μM), no statistically significant downregulation of any of the genes was observed, indicating that neither the regulatory genes driving transcription of mecA, nor the genes encoding the GraRS TCS are the targets of this compound.
To establish the therapeutic index (TI) of lead compounds, their effect on viability of the hepatocellular carcinoma cell line, HepG2, was determined. The TI for conventional antibiotic development is defined as (mammalian cell CC50)/(antibiotic MIC), with a TI ≥ 50 desirable for further development. Because adjuvants, by definition, are typically non-toxic to bacteria, we define adjuvant TI as (mammalian cell CC50)/MEC. The CC50 was determined for the five lead compounds (Table 9). The parent compound, NDM-335, has a CC50 of 96.8 μM, an active concentration of 7.5 μM, and a TI of 12.9. NDM-400 returns a CC50 of 145.9 μM, an active concentration ranging from 5 μM to 7.5 μM across the three strains, resulting in a TI ranging from 19.5 to 29.2. NDM-408 has a CC50 of 81.4 μM, an active concentration of 5 μM, and a TI of 16.3. NDM-427 returns a CC50 of 81.4 μM, an active concentration of 7.5 μM, and a TI of 10.9. Finally, NDM-428 has a CC50 of 179.6 μM, an active concentration ranging from 5 μM to 7.5 μM, and a TI ranging from 24.0 to 35.9. While none of the compounds return a TI ≥ 50, all but one compound (NDM-427) has an increased TI compared to the parent compound.
| NDM-335 | NDM-400 | NDM-408 | NDM-427 | NDM-428 | |
|---|---|---|---|---|---|
| CC50 (μM) | 96.8 ± 12.0 | 145.9 ± 13.5 | 89.6 ± 1.9 | 81.4 ± 10.4 | 179.6 ± 29.98 |
| TI | 12.9 | 19.5–29.2 | 16.3 | 10.9 | 24.0–35.9 |
Conclusion
Through an initial SAR study on benzothiophene NDM-335, we identified several novel compounds that lower the MIC of oxacillin to or below the CLSI breakpoint of 2 μg mL−1 against at least one MRSA strain at 10 μM or lower. Structural trends revealed a strong correlation between activity and sulfur-containing heterocycles, with benzothiophene and benzothiazole motifs providing the most robust adjuvant activity. Substitution patterns within the benzothiophene ring significantly influence potency and strain-based activity, underscoring the sensitivity of this scaffold to modifications. Activity was largely restricted to oxacillin and penicillin G, with limited potentiation of cefoxitin, suggesting antibiotic-class selectivity consistent with PBP resistance mechanisms. Preliminary mechanistic studies further support a role for PBP2a in the observed adjuvant activity. Loss of potentiation in a mecA transposon mutant and in a graRS deletion strain indicates that PBP2a and GraRS-mediated regulation are required for activity. Follow up qRT-PCR analysis has indicated neither controlling mecA expression, nor the genes encoding the GraRS TCS are the targets of these compounds. We are currently purifying PBP2a from BAA-1556 to perform both binding and inhibition assays to further establish the mechanism of action of these compounds and will be reported in due course. This work validates the benzothiophene scaffold as a promising foundation for β-lactam adjuvant development and provides a framework for future efforts aimed at enhancing potency, maintaining appropriate safety metrics, and defining mechanistic activity. Compared to our previously identified PBP2a-dependent oxacillin adjuvants, the benzothiophenes appear more promising than the phenyl urea scaffold for further development as they remain active in potentiation assays at lower concentrations (10 μM or lower), while phenyl urea derivatives are active at high concentrations (60 μM).18Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article has been provided either within the manuscript or in the supplementary information (SI).Supplementary information: the SI includes details on the bacterial strains, media and culture conditions used. Methods for determination of compound minimum inhibitory concentrations as well as antibiotic potentiation. General synthetic procedures are included as well as 1H NMR data for known compounds. Full characterization is provided for all novel compounds which includes 1H NMR/13C NMR/HRMS/UV/IR data. A 1H NMR spectra is provided for each novel compound. Compound crystallographic data has also been included. See DOI: https://doi.org/10.1039/d5md01109d.
CCDC 2514565 contains the supplementary crystallographic data for this paper.32
Acknowledgements
The authors would like to thank Professor Alex Horswill for providing bacterial strains, as well as the University of Notre Dame and the National Institutes of Health (RO1AI167284 to C. M.; P50 AR072000 to V. G. and P. M. D.) for financial support.References
- CDC, Antibiotic Resistance Threats in the United States, 2019 Search PubMed.
- M. S. Mulani, E. E. Kamble, S. N. Kumkar, M. S. Tawre and K. R. Pardesi, Emerging Strategies to Combat ESKAPE Pathogens in the Era of Antimicrobial Resistance: A Review, Front. Microbiol., 2019, 10, 539 CrossRef PubMed.
- CDC, Antibiotic resistance threats in the United States, 2013, 2013 Search PubMed.
- J. Conly and B. Johnston, Where are all the new antibiotics? The new antibiotic paradox, Can. J. Infect. Dis. Med. Microbiol., 2005, 16(3), 159–160 CrossRef PubMed.
- E. E. Gill, O. L. Franco and R. E. Hancock, Antibiotic adjuvants: diverse strategies for controlling drug-resistant pathogens, Chem. Biol. Drug Des., 2015, 85(1), 56–78 CrossRef CAS PubMed.
- U. Imtiaz, E. Billings, J. R. Knox, E. K. Manavathu, S. A. Lerner and S. Mobashery, Inactivation of Class A ß-Lactamases by Clavulanic Acid: The Role of Arginine-244 in a Proposed Nonconcerted Sequence of Events, J. Am. Chem. Soc., 1993, 115, 4435–4442 CrossRef CAS.
- D. E. Ehmann, H. Jahic, P. L. Ross, R. F. Gu, J. Hu and G. Kern, et al., Avibactam is a covalent, reversible, non-beta-lactam beta-lactamase inhibitor, Proc. Natl. Acad. Sci. U. S. A., 2012, 109(29), 11663–11668 CrossRef CAS PubMed.
- R. J. Melander and C. Melander, The Challenge of Overcoming Antibiotic Resistance: An Adjuvant Approach?, ACS Infect. Dis., 2017, 3(8), 559–563 CrossRef CAS PubMed.
- C. G. Bon, J. C. Grigg, J. Lee, C. S. Robb, N. A. Caveney and L. D. Eltis, et al., Structural and kinetic analysis of the monofunctional Staphylococcus aureus PBP1, J. Struct. Biol., 2024, 216(2), 108086 CrossRef CAS PubMed.
- R. Kylväjä, T. Ojalehto, V. Kainulainen, R. Virkola and B. Westerlund-Wikström, Penicillin binding protein 3 of Staphylococcus aureus NCTC 8325-4 binds and activates human plasminogen, BMC Res. Notes, 2016, 9, 389 CrossRef PubMed.
- E. A. Masters, K. L. de Mesy Bentley, A. L. Gill, S. P. Hao, C. A. Galloway and A. T. Salminen, et al., Identification of Penicillin Binding Protein 4 (PBP4) as a critical factor for Staphylococcus aureus bone invasion during osteomyelitis in mice, PLoS Pathog., 2020, 16(10), e1008988 CrossRef CAS PubMed.
- L. C. Chan, A. Gilbert, L. Basuino, T. M. da Costa, S. M. Hamilton and K. R. Dos Santos, et al., PBP 4 Mediates High-Level Resistance to New-Generation Cephalosporins in Staphylococcus aureus, Antimicrob. Agents Chemother., 2016, 60(7), 3934–3941 CrossRef CAS PubMed.
- R. Banerjee, M. Gretes, C. Harlem, L. Basuino and H. F. Chambers, A mecA-negative strain of methicillin-resistant Staphylococcus aureus with high-level β-lactam resistance contains mutations in three genes, Antimicrob. Agents Chemother., 2010, 54(11), 4900–4902 CrossRef CAS PubMed.
- L. Basuino, A. Jousselin, J. A. N. Alexander, N. C. J. Strynadka, M. G. Pinho and H. F. Chambers, et al., PBP4 activity and its overexpression are necessary for PBP4-mediated high-level β-lactam resistance, J. Antimicrob. Chemother., 2018, 73(5), 1177–1180 CrossRef CAS PubMed.
- G. Memmi, S. R. Filipe, M. G. Pinho, Z. Fu and A. Cheung, Staphylococcus aureus PBP4 is essential for beta-lactam resistance in community-acquired methicillin-resistant strains, Antimicrob. Agents Chemother., 2008, 52(11), 3955–3966 CrossRef CAS PubMed.
- M. Young, D. J. Walsh, E. Masters, V. S. Gondil, E. Laskey and M. Klaczko, et al., Identification of Identification of Staphylococcus aureus Penicillin Binding Protein 4 (PBP4) inhibitors, Antibiotics, 2022, 11(10), 1351 CrossRef CAS PubMed.
- V. S. Gondil, H. S. Butman, M. Young, D. J. Walsh, Y. Narkhede and M. J. Zeiler, et al., Development of phenyl-urea-based small molecules that target penicillin-binding protein 4, Chem. Biol. Drug Des., 2024, 103(6), e14569 CrossRef CAS PubMed.
- H. S. Butman, M. A. Stefaniak, D. J. Walsh, V. S. Gondil, M. Young and A. H. Crow, et al., Phenyl urea based adjuvants for β-lactam antibiotics against methicillin resistant Staphylococcus aureus, Bioorg. Med. Chem. Lett., 2025, 121, 130164 CrossRef CAS PubMed.
- K. Ghosh, T. Sarkar and A. P. Chattopadhyay, Anthracene appended pyridinium amide-urea conjugate in selective fluorometric sensing of L-N-acetylvaline salt, Beilstein J. Org. Chem., 2010, 6, 1211–1218 CrossRef CAS PubMed.
- B. A. Diep, S. R. Gill, R. F. Chang, T. H. Phan, J. H. Chen and M. G. Davidson, et al., Complete genome sequence of USA300, an epidemic clone of community-acquired meticillin-resistant Staphylococcus aureus, Lancet, 2006, 367(9512), 731–739 CrossRef CAS PubMed.
- V. T. Nguyen, B. T. Birhanu, V. Miguel-Ruano, C. Kim, M. Batuecas and J. Yang, et al., Restoring susceptibility to β-lactam antibiotics in methicillin-resistant Staphylococcus aureus, Nat. Chem. Biol., 2025, 21(4), 482–489 Search PubMed.
- C. Santiago, E. L. Pang, K. H. Lim, H. S. Loh and K. N. Ting, Inhibition of penicillin-binding protein 2a (PBP2a) in methicillin resistant Staphylococcus aureus (MRSA) by combination of ampicillin and a bioactive fraction from Duabanga grandiflora, BMC Complementary Altern. Med., 2015, 15, 178 CrossRef PubMed.
- CLSI, Performance Standards for Antimicrobial Susceptibility Testing; Nineteenth Informational Supplement, Clinical and Laboratory Standards Institute, Wayne, PA, 2009, p. 152 Search PubMed.
- R. A. Irgashev, A. S. Steparuk and G. L. Rusinov, A new convenient synthetic route towards 2-(hetero)aryl-substituted thieno[3,2-b]indoles using Fischer indolization, Org. Biomol. Chem., 2018, 16(26), 4821–4832 RSC.
- H. L. van de Wouw, J. Y. Lee, M. A. Siegler and R. S. Klausen, Innocent BN bond substitution in anthracene derivatives, Org. Biomol. Chem., 2016, 14(12), 3256–3263 Search PubMed.
- M. J. Zeiler, G. M. Connors, G. M. Durling, A. G. Oliver, L. Marquez and R. J. Melander, et al., Synthesis, Stereochemical Confirmation, and Derivatization of 12(S),16ε-Dihydroxycleroda-3,13-dien-15,16-olide, a Clerodane Diterpene That Sensitizes Methicillin-Resistant Staphylococcus aureus to β-Lactam Antibiotics, Angew. Chem., Int. Ed., 2022, 61(17), e202117458 Search PubMed.
- M. Dettweiler, R. J. Melander, G. Porras, C. Risener, L. Marquez and T. Samarakoon, et al., A Clerodane Diterpene from, ACS Infect. Dis., 2020, 6(7), 1667–1673 Search PubMed.
- A. Ishak, N. Mazonakis, N. Spernovasilis, K. Akinosoglou and C. Tsioutis, Bactericidal versus bacteriostatic antibacterials: clinical significance, differences and synergistic potential in clinical practice, J. Antimicrob. Chemother., 2025, 80(1), 1–17 CrossRef CAS PubMed.
- L. Bleul, P. Francois and C. Wolz, Two-Component Systems of S. aureus: Signaling and Sensing Mechanisms, Genes, 2021, 13(1), 34 CrossRef PubMed.
- M. J. White, J. M. Boyd, A. R. Horswill and W. M. Nauseef, Phosphatidylinositol-specific phospholipase C contributes to survival of Staphylococcus aureus USA300 in human blood and neutrophils, Infect. Immun., 2014, 82(4), 1559–1571 CrossRef PubMed.
- P. D. Fey, J. L. Endres, V. K. Yajjala, T. J. Widhelm, R. J. Boissy and J. L. Bose, et al., A genetic resource for rapid and comprehensive phenotype screening of nonessential Staphylococcus aureus genes, MBio, 2013, 4(1), e00537-12 Search PubMed.
- CCDC 2514565: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc2qdm0s.
| This journal is © The Royal Society of Chemistry 2026 |