 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Design, synthesis, and anticancer evaluation of novel pyrrole–pyrazoline/chalcone hybrids: in vitro and computational insights into EGFR inhibition
Mansour S. Alturki *a,
Marwa F. Ahmed*b,
Abdulaziz H. Al Khzema,
Mohamed S. Gomaa
*a,
Marwa F. Ahmed*b,
Abdulaziz H. Al Khzema,
Mohamed S. Gomaa a,
Mohammad Sarafroza,
Nada Tawfeeq
a,
Mohammad Sarafroza,
Nada Tawfeeq a,
Mashael M. Alharbid,
Abdulaziz K. Al Mousleme,
Mohammed F. Aldawsarif,
Wajin R. Alruwilig,
Shah Alam Khanh,
Radwan El-Haggar
a,
Mashael M. Alharbid,
Abdulaziz K. Al Mousleme,
Mohammed F. Aldawsarif,
Wajin R. Alruwilig,
Shah Alam Khanh,
Radwan El-Haggar i and
Atiah H. Almalki*bc
i and
Atiah H. Almalki*bc
aDepartment of Pharmaceutical Chemistry, College of Pharmacy, Imam Abdulrahman Bin Faisal University, P. O. Box 1982, Dammam 31441, Eastern Province, Kingdom of Saudi Arabia. E-mail: msalturki@iau.edu.sa
bDepartment of Pharmaceutical Chemistry, College of Pharmacy, Taif University, P.O. Box 11099, Taif 21944, Saudi Arabia. E-mail: marwa.farg@tu.edu.sa; marwafarag80@yahoo.com; ahalmalki@tu.edu.sa
cAddiction and Neuroscience Research Unit, Taif University, Taif 21944, Kingdom of Saudi Arabia
dDepartment of Chemistry, College of Science, King Faisal University, Al-Ahsa 31982, Saudi Arabia
eDepartment of Pharmaceutical Sciences, College of Clinical Pharmacy, King Faisal University, Al-Ahsa 31982, Saudi Arabia
fDepartment of Pharmaceutics, College of Pharmacy, Prince Sattam Bin Abdulaziz University, Al-kharj-11942, Saudi Arabia
gCollege of Pharmacy, Imam Abdulrahman Bin Faisal University, P.O. Box 1982, Dammam 31441, Saudi Arabia
hDepartment of Pharmaceutical Chemistry, College of Pharmacy, National University of Science and Technology, PO Box 620, PC 130, Muscat, Oman
iPharmaceutical Chemistry Department, Faculty of Pharmacy, Capital (Formerly Helwan) University, 11795, Ain Helwan, Cairo, Egypt
First published on 27th April 2026
Abstract
A novel series of pyrrole–pyrazoline/chalcone hybrids were designed, synthesized and evaluated for antiproliferative activity to target epidermal growth factor receptor (EGFR) inhibition. All the synthesized compounds were evaluated in the NCI panel of 59 human cancer cell lines, where compound 6b emerged as the most active analogue. Enzyme-binding assays confirmed its potent EGFR inhibitory activity (IC50 = 0.225 μM), comparable to the reference inhibitor erlotinib (IC50 = 0.198 μM). Flow cytometry analysis of human breast cancer cells (MCF-7) showed that 6b induces significant cell arrest in the G2/M phase. Real-time polymerase chain reaction (RT-PCR) experiments further confirmed the molecular mechanisms, revealing that 6b modulated key apoptotic regulators, significantly increasing the Bax/Bcl-2 ratio and upregulating p53, BAX, and caspase-7, while concurrently suppressing Bcl-2 expression. Molecular simulation studies provide evidence for the preferential binding of 6b to the active state of EGFR, consistent with the experimental results. The synthetic strategy used to prepare the pyrrole–pyrazoline/chalcone scaffold is simple, hence providing efficient access to the title compounds whose potential can be further explored as an EGFR-targeted anticancer chemotype.
1. Introduction
Cancer is one of the most challenging diseases worldwide; the World Health Organization (WHO) estimated 20 million new cases and 9.7 million deaths in 2022. Data covering 185 countries and 36 cancers indicated lung, breast and colorectal cancers to be the three major cancer types in 2022.1 Carcinogenesis is a multifactorial process comprising several metabolic changes within a single cell. Conventional cancer treatment approaches include surgery, chemotherapy, radiation therapy, and hormonal therapy but each of these strategies poses physical, financial and psychological challenges.2 Amongst them, chemotherapy remains a primary approach for cancer management; however, it often fails to fully eradicate tumors and is frequently associated with significant toxicity and the development of resistance.3 Although, in the recent past, tremendous progress with oncological therapies has been made, severe side effects (safety and toxicity) and the development of drug resistance due to chemotherapy remain a dilemma.3 Given the increasing burden of cancer, there is a pressing need to develop newer chemotherapeutic agents with novel mechanisms of action for the prevention and treatment of this deadly disease.2In addition, the low specificity of most chemotherapeutic agents often leads to modest efficacy or poor safety profiles. One of the most promising strategies to address these issues is the use of targeted cancer therapies.4,5
The epidermal growth factor receptor (EGFR), a transmembrane receptor tyrosine kinase (TK), is essential for appropriate epidermal cell development and differentiation. EGFR is a ligand-binding receptor composed of a single transmembrane domain, an extracellular domain, and a cytoplasmic region that contains a conserved protein tyrosine kinase (PTK) surrounding a series of regulatory sequences.6,7 The EGFR's crucial function in cell signaling pathways makes it a prominent target in cancer treatment. EGFR is a well-established target implicated in the development and prognosis of various types of cancer. EGFR overexpression is associated with poor treatment outcomes for patients with a number of human malignancies, including breast, lung, colorectal, prostate, and brain cancers,8 which is due to the development of resistance to hormone therapy, chemotherapy, and radiation.9 Therefore, this important protein has been and is still the target of various drug discovery projects in the ongoing battle with cancer. Several EGFR targeting ligands have shown clinical success and decent numbers have acquired FDA approval for the treatment of various types of cancer including among others, erlotinib and lapatinib.10 Accordingly, there have been a growing number of recommendations for the use of EGFR inhibitors as the preferred first-line treatment due to their superior efficacy and safety compared to conventional chemotherapy.11,12
The chemistry of heterocyclic compounds is one of the most prominent branches of organic chemistry. These molecules have gained a great deal of interest in recent years, both for their biological applications and their utility in human society. Pyrrole derivatives are a class of biologically active heterocyclic compounds that can be used as scaffolds for the development of antimicrobial,13–15 antimalarial,16–18 antiviral,19–21 and anti-inflammatory drugs.22 Furthermore, studies have reported that the pyrrole nucleus is an essential core capable of exhibiting potent anticancer activity23–25 (Fig. 1a). Chalcone chemistry has attracted considerable interest due to its simple chemistry and broad spectrum of potential biological activities.23,25,26 There have been several compounds (Fig. 1a) containing chalcone motifs identified as tyrosine kinases inhibitors (TKIs) of EGFR that have been demonstrated to be potent antitumor agents.27,28 Additionally, pyrazoline derivatives have gained popularity over the past several years due to their diverse range of biological properties.29–37 Potent pyrazolines and acetylpyrazoline-based derivatives against the TK of EGFR were reported as novel chemotherapeutic agents (Fig. 1a).20,21,38–44 The majority of US Food and Drug Administration (FDA) approved small molecule anticancer drugs contain heterocyclic rings, with nearly 60% of them in general possessing a nitrogen heterocycle.45 Some of the examples of US FDA approved small molecule EGFR-TKIs (Fig. 1b) containing at least one more nitrogen atom as part of their pharmacophore motif include gefitinib, erlotinib, icotinib, lapatinib, vandetanib afatinib, dacomitinib, canertinib (aminoquinazoline-based derivatives), brigatinib (aminopyrimidine derivative), neratinib, pelitinib (2nd generation 4-aminoquinoline derivatives), osimertinib (3rd generation irreversible EGFR-TKI, pyrrolopyrimidine derivative), olmutinib (thieno [3,2-d] pyrimidine-based drug), rociletinib (2,4-diaminopyrimidine-based compound), naquotinib, avitinib (pyrrolopyrimidine-based irreversible EGFR inhibitor), and nazartinib (aminobenzimidazole-based derivative).46–49
The epidermal growth factor receptor (EGFR) is a well-validated oncogenic target characterized by a conserved ATP-binding pocket within its tyrosine kinase domain (TKD). Effective EGFR inhibitors typically share key pharmacophoric features essential for high-affinity binding and kinase inhibition. Hydrogen bonding with the hinge region is a critical pharmacophoric element with the ability to form hydrogen bonds with backbone amide groups in the hinge region of EGFR, particularly involving residues such as MET769. This interaction anchors the inhibitor within the ATP-binding site and is central to inhibitory potency. Hydrophobic/aromatic interactions through aromatic rings and hydrophobic moieties enable π–π stacking and van der Waals contacts with hydrophobic pockets adjacent to the hinge region, enhancing binding affinity and specificity. Key residues involved include LEU694, VAL702, LEU764, and LEU768. Ionic and polar interactions through charged or polar groups capable of ionic–π or hydrogen bonding interactions with residues such as LYS721, GLU738, and ASP831 (part of the DFG motif) contribute to stabilizing the ligand conformation and improving selectivity.
The proposed hybrid compounds strategically integrate these essential pharmacophoric elements through a rational mix-and-match approach informed by prior antitumor scaffolds. The pyrrole ring serves as an aromatic scaffold capable of engaging in π–π stacking interactions with aromatic residues lining the EGFR binding site, contributing hydrophobic contacts and enhancing binding affinity. The chalcone α,β-unsaturated ketone and pyrazoline ring systems provide hydrogen bond acceptors/donors capable of forming key interactions with the hinge region, particularly mimicking the hydrogen bonding pattern with MET769 as observed in clinically approved inhibitors like erlotinib. Acetyl groups on the pyrazoline scaffold modulate electronic properties and steric bulk, optimizing the fit within hydrophobic pockets and potentially improving metabolic stability and pharmacokinetics. Substitution on the aromatic rings with electron-withdrawing fluoro substituents may enhance metabolic stability and modulate electronic density to strengthen hydrogen bonding and van der Waals interactions within the ATP-binding pocket or that with electron-donating methyl groups may contribute hydrophobic contacts and influence the binding orientation, providing complementary activity profiles (Fig. 2). These compounds offer the distinct benefit of being easily synthesized through simple and efficient chemical procedures, facilitating effective molecular optimization and in-depth mechanistic investigations in molecular pharmacology.
These newly synthesized compounds were submitted to the National Cancer Institute (NCI) for evaluation of their antiproliferative activities. A follow-up study was conducted to determine which of the drugs is most effective as an EGFR TKI. This work also introduced the Annexin V-FITC apoptosis test. The cell cycle activity of the most effective compound was also determined to determine the possible stage at which the novel derivatives may inhibit cancer cell proliferation. Moreover, in addition to the levels of Bax and Bcl2, this investigation evaluated the levels of caspase-7. Finally, a molecular modeling approach was employed to identify binding interactions against the WT EGFR TK therapeutic target. ADMET studies and molecular dynamics simulations were conducted to evaluate the pharmacokinetic and pharmacodynamic properties of the potential EFGR inhibitors. This study aimed to develop novel EGFR inhibitors that are highly and efficiently accessible through simple, rapid and clean synthetic chemistry. This would eventually speed up the drug optimization phase and facilitate the chemical development process in the clinical trial phase.
2. Experimental
2.1. Chemistry
All reagents and solvents used in the following experiments were obtained from commercial sources. A digital melting point apparatus Electrothermal Stuart 5MP3 was utilized to measure melting points. Elemental analysis was performed on a Vario EL III elemental analyzer. The IR spectrograms of the compounds were validated using an infrared spectrophotometer (Perkin Elmer 9712). NMR spectra were obtained using Joel-Ex and Varian-Gemini NMR spectrometers for the investigation of 1H NMR and 13C NMR (500 MHz for 1H-NMR and 101 MHz for 13C-NMR). For the mass spectra, a Shimadzu GC mass spectrometer MSQp1000EX was used. High-resolution mass spectrometry (HRMS) measurements were carried out on an ACQUITY UPLC–QTOF Premier system (Waters) using ESI in positive mode. Data were acquired over m/z 100–1300 with lock-mass correction (HP-1221) to ensure mass accuracy.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1590 (C
O), 1590 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1HNMR (DMSO-d6): δ 6.43 (d, 1H, J = 15.5 Hz, CH
C). 1HNMR (DMSO-d6): δ 6.43 (d, 1H, J = 15.5 Hz, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 6.62 (d, 2H, J = 8.4 Hz, H-pyrrole), 6.97 (d, 2H, J = 8.2 Hz, H-pyrrole), 7.34–7.82 (m, 9H, Ar–H, CH
), 6.62 (d, 2H, J = 8.4 Hz, H-pyrrole), 6.97 (d, 2H, J = 8.2 Hz, H-pyrrole), 7.34–7.82 (m, 9H, Ar–H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ) and 8.18 (s, 1H, Ar–H). 13C NMR (DMSO-d6): 189.4, 145.3, 142.4, 138.5, 135.3, 131.2, 129.1, 128.4, 127.6, 126.5, 124.1, 122.2, 120.6, 109.4, 107.3. MS: m/z = 273.34.
) and 8.18 (s, 1H, Ar–H). 13C NMR (DMSO-d6): 189.4, 145.3, 142.4, 138.5, 135.3, 131.2, 129.1, 128.4, 127.6, 126.5, 124.1, 122.2, 120.6, 109.4, 107.3. MS: m/z = 273.34.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1594 (C
O), 1594 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1HNMR (DMSO-d6): δ 6.38 (d, 1H, J = 15.4 Hz, CH
C). 1HNMR (DMSO-d6): δ 6.38 (d, 1H, J = 15.4 Hz, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 6.59 (d, 2H, J = 8.3 Hz, H-pyrrole), 6.91 (d, 2H, J = 8.0 Hz, H-pyrrole), 7.48–7.83 (m, 8H, Ar–H, CH
), 6.59 (d, 2H, J = 8.3 Hz, H-pyrrole), 6.91 (d, 2H, J = 8.0 Hz, H-pyrrole), 7.48–7.83 (m, 8H, Ar–H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ) and 8.16 (s, 1H, Ar–H). 13C NMR (DMSO-d6): 189.6, 160.7, 145.2, 142.5, 138.4, 136.7, 130.8, 129.6, 128.4, 127.4, 125.3, 124.1, 121.2, 120.3, 115.7, 112.8, 110.6, 107.2. MS: m/z = 291.33.
) and 8.16 (s, 1H, Ar–H). 13C NMR (DMSO-d6): 189.6, 160.7, 145.2, 142.5, 138.4, 136.7, 130.8, 129.6, 128.4, 127.4, 125.3, 124.1, 121.2, 120.3, 115.7, 112.8, 110.6, 107.2. MS: m/z = 291.33.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1594 (C
O), 1594 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1H NMR (DMSO-d6): δ 2.47 (s, 3H, CH3), 6.33 (d, 1H, J = 15.0 Hz, CH
C). 1H NMR (DMSO-d6): δ 2.47 (s, 3H, CH3), 6.33 (d, 1H, J = 15.0 Hz, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 6.68 (d, 2H, J = 8.3 Hz, H-pyrrole), 6.95 (d, 2H, J = 8.0 Hz, H-pyrrole), 7.23–7.62 (m, 8H, Ar–H, CH
), 6.68 (d, 2H, J = 8.3 Hz, H-pyrrole), 6.95 (d, 2H, J = 8.0 Hz, H-pyrrole), 7.23–7.62 (m, 8H, Ar–H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ) and 8.21 (s, 1H, Ar–H). 13C NMR (DMSO-d6): 189.8, 145.3, 142.4, 138.5, 136.2, 132.7, 129.6, 128.6, 128.3, 127.2, 126.6, 125.4, 122.3, 119.4, 111.7, 107.1, 21.7. MS: m/z = 287.36.
) and 8.21 (s, 1H, Ar–H). 13C NMR (DMSO-d6): 189.8, 145.3, 142.4, 138.5, 136.2, 132.7, 129.6, 128.6, 128.3, 127.2, 126.6, 125.4, 122.3, 119.4, 111.7, 107.1, 21.7. MS: m/z = 287.36.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O), 1593 (C
O), 1593 (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1H NMR (DMSO-d6): δ 6.25 (d, 1H, J = 15.5 Hz, CH
C). 1H NMR (DMSO-d6): δ 6.25 (d, 1H, J = 15.5 Hz, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ), 6.44 (d, 2H, J = 8.6 Hz, H-pyrrole), 6.95 (d, 2H, J = 8.2 Hz, H-pyrrole), 7.12–7.66 (m, 8H, Ar–H, CH
), 6.44 (d, 2H, J = 8.6 Hz, H-pyrrole), 6.95 (d, 2H, J = 8.2 Hz, H-pyrrole), 7.12–7.66 (m, 8H, Ar–H, CH![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) ) and 8.23 (s, 1H, Ar–H). 13C NMR (DMSO-d6): 190.2, 160.3, 145.4, 141.1, 137.6, 135.8, 130.7, 129.5, 126.4, 125.3, 121.2, 118.7, 116.6, 111.8, 107.4. MS: m/z = 291.33.
) and 8.23 (s, 1H, Ar–H). 13C NMR (DMSO-d6): 190.2, 160.3, 145.4, 141.1, 137.6, 135.8, 130.7, 129.5, 126.4, 125.3, 121.2, 118.7, 116.6, 111.8, 107.4. MS: m/z = 291.33.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1H NMR (DMSO-d6): δ 3.04 (dd, 1H, J = 11.2, 5.0 Hz, H5 pyrazoline), 3.23 (dd, 1H, J = 11.6, 5.9 Hz, Hb4 pyrazoline), 4.97 (dd, 1H, J = 12.0, 5.7 Hz, Ha4 pyrazoline), 6.36 (d, 2H, J = 8.6 Hz, H-pyrrole), 6.91 (d, 2H, J = 8.2 Hz, H-pyrrole), 7.24–7.58 (m, 9H, Ar–H) and 8.60 (s, NH, D2O exchangeable). 13C NMR (DMSO-d6): 151.5, 143.4, 142.0, 134.5, 129.3, 128.5, 126.9, 126.7, 125.2, 123.4, 120.3, 110.9, 106.2, 51.3, 44.9. MS: m/z = 287.37.
C). 1H NMR (DMSO-d6): δ 3.04 (dd, 1H, J = 11.2, 5.0 Hz, H5 pyrazoline), 3.23 (dd, 1H, J = 11.6, 5.9 Hz, Hb4 pyrazoline), 4.97 (dd, 1H, J = 12.0, 5.7 Hz, Ha4 pyrazoline), 6.36 (d, 2H, J = 8.6 Hz, H-pyrrole), 6.91 (d, 2H, J = 8.2 Hz, H-pyrrole), 7.24–7.58 (m, 9H, Ar–H) and 8.60 (s, NH, D2O exchangeable). 13C NMR (DMSO-d6): 151.5, 143.4, 142.0, 134.5, 129.3, 128.5, 126.9, 126.7, 125.2, 123.4, 120.3, 110.9, 106.2, 51.3, 44.9. MS: m/z = 287.37.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1H NMR (DMSO-d6): δ 3.08 (dd, 1H, J = 11.0, 5.1 Hz, H5 pyrazoline), 3.28 (dd, 1H, J = 11.4, 5.7 Hz, Hb4 pyrazoline), 4.89 (dd, 1H, J = 12.1, 5.7 Hz, Ha4 pyrazoline), 6.21 (d, 2H, J = 8.4 Hz, H-pyrrole), 6.97 (d, 2H, J = 8.0 Hz, H-pyrrole), 7.13–7.46 (m, 8H, Ar–H) and 8.52 (s, NH, D2O exchangeable). 13C NMR (DMSO-d6): 162.8, 151.9, 145.4, 142.4, 136.7, 130.1, 128.5, 126.0, 123.4, 122.4, 120.1, 115.8, 115.9, 109.9, 106.5, 51.5, 43.8. MS: m/z = 305.36. LC-HRMS: calculated for C19H17N3F (M + H): 306.1407, found: 306.1480.
C). 1H NMR (DMSO-d6): δ 3.08 (dd, 1H, J = 11.0, 5.1 Hz, H5 pyrazoline), 3.28 (dd, 1H, J = 11.4, 5.7 Hz, Hb4 pyrazoline), 4.89 (dd, 1H, J = 12.1, 5.7 Hz, Ha4 pyrazoline), 6.21 (d, 2H, J = 8.4 Hz, H-pyrrole), 6.97 (d, 2H, J = 8.0 Hz, H-pyrrole), 7.13–7.46 (m, 8H, Ar–H) and 8.52 (s, NH, D2O exchangeable). 13C NMR (DMSO-d6): 162.8, 151.9, 145.4, 142.4, 136.7, 130.1, 128.5, 126.0, 123.4, 122.4, 120.1, 115.8, 115.9, 109.9, 106.5, 51.5, 43.8. MS: m/z = 305.36. LC-HRMS: calculated for C19H17N3F (M + H): 306.1407, found: 306.1480.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C). 1H NMR (DMSO-d6): δ 2.30 (s, 3H, CH3), 3.11 (dd, 1H, J = 11.0, 5.2 Hz, H5 pyrazoline), 3.23 (dd, 1H, J = 11.4, 5.7 Hz, Hb4 pyrazoline), 4.99 (dd, 1H, J = 12.1, 5.6 Hz, Ha4 pyrazoline), 6.29 (d, 2H, J = 8.7 Hz, H-pyrrole), 6.96 (d, 2H, J = 8.5 Hz, H-pyrrole), 7.15–7.47 (m, 8H, Ar–H) and 8.59 (s, NH, D2O exchangeable). 13C NMR (DMSO-d6): 151.6, 143.4, 142.2, 138.2, 134.5, 129.8, 128.4, 127.1, 125.0, 123.6, 123.4, 122.3, 120.4, 110.8, 106.7, 51.5. 42.6, 21.8. MS: m/z = 301.39.
C). 1H NMR (DMSO-d6): δ 2.30 (s, 3H, CH3), 3.11 (dd, 1H, J = 11.0, 5.2 Hz, H5 pyrazoline), 3.23 (dd, 1H, J = 11.4, 5.7 Hz, Hb4 pyrazoline), 4.99 (dd, 1H, J = 12.1, 5.6 Hz, Ha4 pyrazoline), 6.29 (d, 2H, J = 8.7 Hz, H-pyrrole), 6.96 (d, 2H, J = 8.5 Hz, H-pyrrole), 7.15–7.47 (m, 8H, Ar–H) and 8.59 (s, NH, D2O exchangeable). 13C NMR (DMSO-d6): 151.6, 143.4, 142.2, 138.2, 134.5, 129.8, 128.4, 127.1, 125.0, 123.6, 123.4, 122.3, 120.4, 110.8, 106.7, 51.5. 42.6, 21.8. MS: m/z = 301.39.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). 1H NMR (DMSO-d6): δ 2.41 (s, 3H, COCH3), 3.72 (m, 1H, H5 pyrazoline), 3.89 (m, 1H, Hb4 pyrazoline), 5.54 (m, 1H, Ha4 pyrazoline), 6.21 (d, 2H, J = 8.6 Hz, H-pyrrole), 6.97 (d, 2H, J = 8.4 Hz, H-pyrrole), 7.28–7.63 (m, 9H, Ar–H). 13C NMR (DMSO-d6): 168.2, 151.3, 142.4, 141.6, 135.5, 129.5, 128.4, 127.0, 126.7, 125.1, 123.6, 120.3, 110.9, 106.5, 62.9, 42.9, 23.2. MS: m/z = 329.40.
O). 1H NMR (DMSO-d6): δ 2.41 (s, 3H, COCH3), 3.72 (m, 1H, H5 pyrazoline), 3.89 (m, 1H, Hb4 pyrazoline), 5.54 (m, 1H, Ha4 pyrazoline), 6.21 (d, 2H, J = 8.6 Hz, H-pyrrole), 6.97 (d, 2H, J = 8.4 Hz, H-pyrrole), 7.28–7.63 (m, 9H, Ar–H). 13C NMR (DMSO-d6): 168.2, 151.3, 142.4, 141.6, 135.5, 129.5, 128.4, 127.0, 126.7, 125.1, 123.6, 120.3, 110.9, 106.5, 62.9, 42.9, 23.2. MS: m/z = 329.40.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). 1H NMR (DMSO-d6): δ 2.45 (s, 3H, COCH3), 3.67 (m, 1H, H5 pyrazoline), 3.85 (m, 1H, Hb4 pyrazoline), 5.58 (m, 1H, Ha4 pyrazoline), 6.25 (d, 2H, J = 8.5 Hz, H-pyrrole), 6.92 (d, 2H, J = 8.4 Hz, H-pyrrole), 7.06–7.50 (m, 8H, Ar–H). 13C NMR (DMSO-d6): 168.3, 162.7, 151.6, 143.6, 142.3, 130.5, 129.4, 126.2, 123.6, 122.7, 118.5, 114.9, 113.5, 111.9, 106.4, 62.5, 41.8, 23.2. MS: m/z = 347.39.
O). 1H NMR (DMSO-d6): δ 2.45 (s, 3H, COCH3), 3.67 (m, 1H, H5 pyrazoline), 3.85 (m, 1H, Hb4 pyrazoline), 5.58 (m, 1H, Ha4 pyrazoline), 6.25 (d, 2H, J = 8.5 Hz, H-pyrrole), 6.92 (d, 2H, J = 8.4 Hz, H-pyrrole), 7.06–7.50 (m, 8H, Ar–H). 13C NMR (DMSO-d6): 168.3, 162.7, 151.6, 143.6, 142.3, 130.5, 129.4, 126.2, 123.6, 122.7, 118.5, 114.9, 113.5, 111.9, 106.4, 62.5, 41.8, 23.2. MS: m/z = 347.39.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O). 1H NMR (DMSO-d6): δ 2.24 (s, 3H, CH3), 2.40 (s, 3H, COCH3), 3.64 (m, 1H, H5 pyrazoline), 3.88 (m, 1H, Hb4 pyrazoline), 5.51 (m, 1H, Ha4 pyrazoline), 6.30 (d, 2H, J = 8.6 Hz, H-pyrrole), 6.86 (d, 2H, J = 8.2 Hz, H-pyrrole), 7.13–7.51 (m, 8H, Ar–H). 13C NMR (DMSO-d6): 168.1, 151.5, 143.4, 141.9, 138.3, 134.7, 129.1, 128.4, 127.2, 126.7, 125.0, 123.8, 123.6, 120.3, 110.8, 106.4, 66.2, 41.8, 23.4, 21.6. MS: m/z = 343.43.
O). 1H NMR (DMSO-d6): δ 2.24 (s, 3H, CH3), 2.40 (s, 3H, COCH3), 3.64 (m, 1H, H5 pyrazoline), 3.88 (m, 1H, Hb4 pyrazoline), 5.51 (m, 1H, Ha4 pyrazoline), 6.30 (d, 2H, J = 8.6 Hz, H-pyrrole), 6.86 (d, 2H, J = 8.2 Hz, H-pyrrole), 7.13–7.51 (m, 8H, Ar–H). 13C NMR (DMSO-d6): 168.1, 151.5, 143.4, 141.9, 138.3, 134.7, 129.1, 128.4, 127.2, 126.7, 125.0, 123.8, 123.6, 120.3, 110.8, 106.4, 66.2, 41.8, 23.4, 21.6. MS: m/z = 343.43.2.2. In vitro cytotoxicity
Antiproliferative activity was assessed using the NCI-60 human tumor cell line panel maintained and tested by the National Cancer Institute Developmental Therapeutics Program. The panel consists of established human cancer cell lines purchased from the American Type Culture Collection (ATCC, Manassas, VA, USA). Only established commercial cell lines were used.The NCI-60 human cancer cell lines used in this study were obtained from repositories of the National Cancer Institute Development Therapeutics Program (NCI DTP). In vitro tests were performed on a total of 59 human cancer cell lines using the method described by Monks et al.60 to assess the cytotoxicity of all synthesized compounds by the National Cancer Institute (NCI, USA). The antiproliferative activities of all the synthesized compounds were tested at the NCI, USA, against a panel of 59 human cancer cell lines representing nine cancer types, including leukemia, lung, colon, CNS, melanoma, ovarian, renal, prostate, and breast cancers. The assay was performed following a modified sulforhodamine B (SRB) protocol according to the method of Monks et al.; some major steps are mentioned here for the ease of reproducibility.60
Cancer cells were cultured using RPMI-1640 medium supplemented with 5% fetal bovine serum (FBS) and antibiotics as appropriate. Cells were seeded in 96-well plates, 5000–10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well depending upon the cell line, and allowed to settle for around 24 h in a 5% CO2 incubator at 37 °C. The test compounds dissolved in DMSO were serially diluted in concentrations ranging from 10−4 to 10−8 M. The cells were treated in triplicate and further incubated for 48 h.
000 cells per well depending upon the cell line, and allowed to settle for around 24 h in a 5% CO2 incubator at 37 °C. The test compounds dissolved in DMSO were serially diluted in concentrations ranging from 10−4 to 10−8 M. The cells were treated in triplicate and further incubated for 48 h.
After the treatment regime, 10% cold trichloroacetic acid was added for fixing cells at 4 °C for approximately 1 h; the plates were then washed, dried, and stained with 0.4% SRB dissolved in 1% acetic acid for 30 min at room temperature. Following this, excess dye was washed off with 1% acetic acid. The dye that remained was dissolved using 10 mM Tris base (pH 10.5), and absorbance was measured at 564 nm.
2.3. EGFR inhibition assay
A HTRF test51 was used to assess the inhibitory activity of the most promising compound 6b against EGFR. Compound 6b/erlotinib was incubated with the EGFR kinase and its substrate for five minutes before adding ATP to start the enzymatic reaction. After that, the reaction was completed within 30 minutes at room temperature. An EDTA detection reagent was added after 1 hour to stop the reaction. Each concentration was tested in triplicate within an experiment, and the assay was repeated in three independent experiments (n = 3). IC50 values are expressed as the mean ± standard deviation (SD).512.4. Cell cycle analysis
Human breast cancer cells (MCF-7) were seeded into six-well plates at 1 × 105 cells per well and allowed to attach for 24 hours at 37 °C in a humid atmosphere with 5% CO2 in complete culture medium supplemented with 10% FBS. After attachment, treatment with DMSO (0.1% v/v) or compound 6b (10 μM) was started. The cells were treated for 48 hours while incubation conditions remained unchanged. At the end of treatment, the cells were washed off the medium twice with PBS, followed by 70% cold ethanol addition dropwise with gentle vortexing for fixing. Fixed cells could be stored at 4 °C for at least 1 hour to maximize fixing efficacy. Before analysis, the fixed cells were washed with PBS to remove ethanol traces and stained with a PI solution in the presence of RNase A to stain the DNA specifically. The cells were incubated at 37 °C for 15 minutes to allow proper staining. Cell cycle analysis was conducted on a BD FACS Caliber flow cytometer. Data acquisition and analysis were then performed to calculate percentages of G0/G1, S, and G2/M cells. This method enabled the assessment of cell cycle arrest induced by compound 6b in MCF-7 cells. Experiments were performed in triplicate (n = 3).522.5. Apoptosis assay
To determine apoptosis induction by compound 6b, Annexin V-FITC/PI double staining was carried out using an Annexin V-FITC/PI apoptosis detection kit (BD Biosciences, San Diego, CA, USA) according to slight modifications of the manufacturer's instructions for MCF-7 cells.MCF-7 cells were plated into T-75 flasks at a cell density of 4 × 106 cells per flask and allowed to attach after overnight incubation. The following day, the cells were treated for 24 hours with compound 6b at its IC50 concentration of 0.225 μM in a humidified incubator with 5% CO2 at 37 °C. After completion of the treatment, the cells were harvested by trypsinization and pelleted by centrifugation. Around 0.5 × 106 cells of all samples were then washed twice, with cold phosphate-buffered saline, and resuspended in 100 μL of 1× binding buffer. The cells were stained by adding 5 μL of Annexin V-FITC and 5 μL of propidium iodide (PI). The samples were incubated for 15 minutes at room temperature in the dark.
Following incubation, 400 μL of 1× binding buffer was added to each tube, and the samples were shortly analyzed using the BD FACS Caliber flow cytometer (BD Biosciences, San Jose, CA, USA). The data were processed to quantify the percentage of cells in the early apoptotic phase and experiments were performed in triplicate (n = 3).52
2.6. Caspase-7 activity
Caspase-7 activity was measured as per a previously reported method.53 MCF-7 cells (1 × 106) at half maximal inhibitory concentrations of compound 6b for 24 h were harvested and washed with ice-cold phosphate-buffered saline (PBS). Cell pellets were lysed with 50 mM Tris–HCl (pH 7.5), 10 mM EDTA, 0.2 M NaCl, 1.5 mM PMSF, and 1% SDS. Lysates were then boiled for 10 min and centrifuged; the supernatants were stored at −20 °C. About 10 to 20 μg of protein was separated onto SDS gels and subjected to nitrocellulose membranes. Membranes were subjected to anti-caspase-7 antibodies and secondary antibodies, and chemiluminescence detection was done. The caspase-7 was compared against untreated controls.2.7. Measurement of p53, BAX, and Bcl-2 expression
Levels of tumor suppressor gene p53, anti-apoptotic marker Bcl-2, and apoptotic marker BAX were evaluated, as previously reported.40 MCF-7 cells that were treated with compound 6b were determined by using an iScriptTM One-Step RT-PCR kit with SYBR® Green (Bio-Rad, USA). After the RNA was extracted and quantified, 1 μg of total RNA was used in each 20 μL reaction. The RT-PCR conditions were as follows: reverse transcription at 50 °C (10 min), initial denaturation at 95 °C (5 min), and 40 cycles warming up at 95 °C (10 s) and cooling down at 60 °C (30 s).2.8. DNA fragmentation analysis
Using salting-out DNA isolates, agarose gel electrophoresis was performed to examine fragmentation of genomic DNA;40 MCF-7 cells were therefore grown with and without compound 6b. After the cells had been washed and counted, they were subjected to DNA extraction. To prepare the TE buffer, DNA samples were first meticulously resuspended. The DNA laddering was examined using an ultraviolet transilluminator and ethidium bromide staining after the material was run over a 1.5 percent TAE agarose gel.2.9. Molecular modeling
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 atoms.70,71
000 atoms.70,71The final system was retrieved upon executing the MD simulation. This system was relaxed under the default relaxation protocol and the simulation period was set to 100 ns. Using the Nose–Hover chain thermostat and Martyna–Tobias–Klein barostat, simulation was conducted under an isothermal–isobaric NPT entity at a temperature of 310 K and a pressure of 1.103 bar.70,72 The reversible reference system propagator algorithm (RESPA) integrator was used to accurately predict the molecular motions for near and far atoms using time step sizes of 2.0 fs and 6.0 fs, respectively. Furthermore, the Coulombic cutoff was selected to be 9.0 Å as per the default program. Following the simulation run, the results were loaded onto Maestro workspace through the Desmond simulation interaction diagram and analyzed.72,73
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P), hydrogen bond donors, and hydrogen bond acceptors).75
P), hydrogen bond donors, and hydrogen bond acceptors).753. Results and discussion
3.1. Chemistry
All compounds were synthesized using the protocol specified in Schemes 1 and 2. The Paal–Knorr reaction produced (3-pyrrol-1-yl)acetophenone 3 by condensing 3-amino acetophenone 1 with 2,5-dimethoxytetrahydrofuran 2. Using a sodium hydroxide catalyst, chalcones 5a–d were synthesized in ethanol by Claisen Schmidt condensation of 3 with substituted aromatic aldehydes (4a–d; Scheme 1).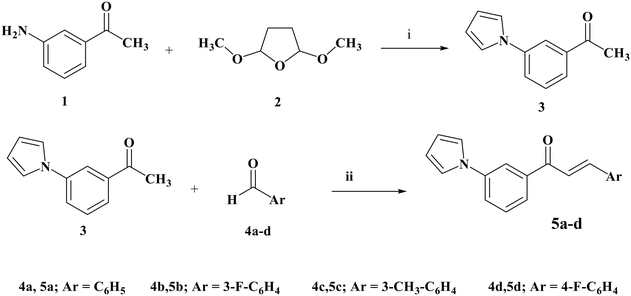 | ||
| Scheme 1 Synthesis of compounds 5a–d. Reagents and conditions: (i) glacial acetic acid, reflux 3 h. (ii) NaOH, absolute ethanol, stirring 24 h. | ||
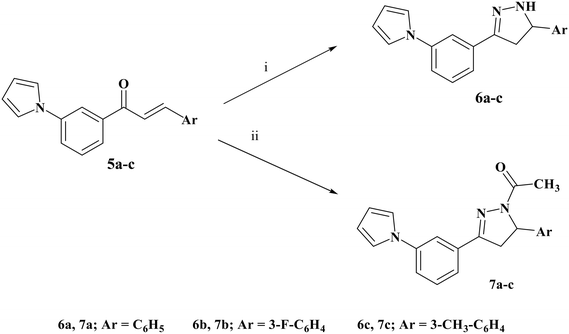 | ||
| Scheme 2 Synthesis of compounds 6a–c and 7a–c. Reagents and conditions: (i) hydrazine hydrate, absolute ethanol, reflux 16 h. (ii) Hydrazine hydrate, glacial acetic acid, reflux 20 h. | ||
The proposed structures of chalcones 5a–d were thoroughly confirmed by elemental analysis and comprehensive spectral characterization techniques. Infrared (IR) spectroscopy exhibited characteristic absorption bands for the carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) group ranging from 1720 to 1725 cm−1, indicative of the α,β-unsaturated ketone functionality typical of chalcones. Aromatic C–H stretching vibrations appeared near 3000 cm−1, consistent with the aromatic rings present in the molecules. The 1H NMR spectra further validated the structures, showing a distinctive doublet for the olefinic proton at δ 6.25–6.43 ppm with a coupling constant (J) of approximately 15 Hz, confirming the trans-configuration of the double bond. The corresponding olefinic proton appeared as multiplets overlapping with aromatic proton signals, reflecting the expected spin coupling. All carbon resonances appeared at anticipated chemical shifts in the 13C NMR spectra, confirming the chalcone backbone and substitution pattern.
O) group ranging from 1720 to 1725 cm−1, indicative of the α,β-unsaturated ketone functionality typical of chalcones. Aromatic C–H stretching vibrations appeared near 3000 cm−1, consistent with the aromatic rings present in the molecules. The 1H NMR spectra further validated the structures, showing a distinctive doublet for the olefinic proton at δ 6.25–6.43 ppm with a coupling constant (J) of approximately 15 Hz, confirming the trans-configuration of the double bond. The corresponding olefinic proton appeared as multiplets overlapping with aromatic proton signals, reflecting the expected spin coupling. All carbon resonances appeared at anticipated chemical shifts in the 13C NMR spectra, confirming the chalcone backbone and substitution pattern.
Chalcone derivatives 5a–c were reacted with hydrazine hydrate in ethanol to yield pyrazoline derivatives 6a–c (Scheme 2). Elemental and spectral analyses confirmed the formation of these products. IR spectra showed the disappearance of the carbonyl absorption bands and the emergence of broad NH stretching peaks at 3385–3390 cm−1, characteristic of pyrazoline ring formation. The 1H NMR spectra revealed the loss of olefinic doublet signals and the appearance of three characteristic doublets of doublets at δ 3.04–3.11 ppm, 3.23–3.28 ppm, and 4.89–4.99 ppm, corresponding to the protons on the pyrazoline ring, confirming successful cyclization.
In a parallel synthetic route, chalcones 5a–c were treated with hydrazine hydrate in the presence of glacial acetic acid to afford acetyl pyrazoline derivatives 7a–c (Scheme 2). Structural confirmation was achieved through elemental and spectral analyses. IR spectra revealed the disappearance of amino group absorption bands and the appearance of a carbonyl band at 1662–1670 cm−1, indicative of acetyl functionality. The 1H NMR spectra displayed a singlet at δ 2.24–2.45 ppm, attributable to the acetyl methyl protons. All carbon resonances in the 13C NMR spectra appeared at expected chemical shifts, consistent with the proposed acetyl pyrazoline structures.
Together, the combined elemental and spectral data provide comprehensive validation of each synthetic step—from chalcone formation to pyrazoline ring closure and acetylation. The IR, 1H NMR, and 13C NMR spectral features consistently support the structural assignments and confirm the expected chemical transformations. These detailed characterization results ensure the reliability of the synthetic methodology and the purity of the final products.
The synthetic pathway utilized simple, time-consuming, and cost-efficient methods. The starting materials are readily available and cheap materials, the reaction time is short, and the reaction conditions are regular. The products are easily collected by filtration in pure and high yield.
3.2. Biological activity
| Panel | Subpanel | 5a | 5b | 5c | 5d | 6a |
|---|---|---|---|---|---|---|
| Leukemia | CCRF-CEM | 99.55 | 103.36 | 109.74 | 106.60 | 96.14 |
| HL-60(TB) | 101.62 | 84.8 | 86.3 | 80.60 | 98.01 | |
| K-562 | 95.95 | 73.96 | 78.16 | 84.45 | 108.06 | |
| MOLT-4 | 106.04 | 96.16 | 98.24 | 99.71 | 100.94 | |
| RPMI-8226 | 93.26 | 98.23 | 98.38 | 100.89 | 97.32 | |
| SR | 100.51 | 84.3 | 91.71 | 101.36 | 112.65 | |
| Non-small cell lung cancer | A549/ATCC | 99.88 | 103.58 | 101.05 | 103.38 | 101.42 |
| EKVX | 100.65 | 102 | 99.36 | 97.17 | 86.87 | |
| HOP-62 | 116.18 | 123.03 | 120.42 | 114.41 | 108.14 | |
| NCI-H226 | 100.97 | 95.49 | 94.43 | 96.60 | 95.14 | |
| NCI-H23 | 86.53 | 91.54 | 95.61 | 91.65 | 92.05 | |
| NCI-H322M | 97.56 | 94.21 | 97.88 | 99.28 | 102.59 | |
| NCI-H460 | 107.98 | 100.61 | 111.92 | 108.03 | 97.69 | |
| NCI-H522 | 96.97 | 98.49 | 97.49 | 99.04 | 97.86 | |
| Colon cancer | COLO 205 | 121.6 | 123.97 | 121.73 | 118.42 | 116.68 |
| HCC-2998 | 109.01 | 111.44 | 108.47 | 107.96 | 111.99 | |
| HCT-116 | 91.57 | 83.81 | 92.77 | 89.12 | 103.48 | |
| HCT-15 | 91.88 | 92.49 | 90.33 | 91.46 | 97.4 | |
| HT29 | 109.21 | 117.46 | 114.39 | 110.73 | 117.43 | |
| KM12 | 101.43 | 99.19 | 103.91 | 96.45 | 99.24 | |
| SW-620 | 93.04 | 92.16 | 102.47 | 102.69 | 102.3 | |
| CNS cancer | SF-268 | 115.55 | 113.92 | 112.82 | 108.28 | 107.41 |
| SF-295 | 114.8 | 102.28 | 100.95 | 106.01 | 104.08 | |
| SF-539 | 105.3 | 106.43 | 107.03 | 107.83 | 109.4 | |
| SNB-19 | 94.57 | 93.27 | 97.37 | 97.17 | 99.02 | |
| SNB-75 | 103.57 | 106.03 | 97.88 | 117.07 | 125.19 | |
| U251 | 98.05 | 98 | 102.94 | 100.91 | 103.17 | |
| Melanoma | LOX IMVI | 95.85 | 91.91 | 99.27 | 90.87 | 100.85 |
| MALME-3M | 96.49 | 100.25 | 93.8 | 96.59 | 86.91 | |
| M14 | 99.72 | 104.21 | 105.98 | 107.42 | 100.91 | |
| MDA-MB-435 | 103.55 | 102.65 | 105.02 | 99.65 | 106.71 | |
| SK-MEL-2 | 99.58 | 110.11 | 108.63 | 108.22 | 114.24 | |
| SK-MEL-28 | 108.37 | 105 | 106.34 | 105.85 | 108.55 | |
| SK-MEL-5 | 99.35 | 98.4 | 99.24 | 99.24 | 99.03 | |
| UACC-257 | 114.66 | 111.75 | 107.37 | 109.33 | 100.64 | |
| UACC-62 | 81.33 | 80.71 | 84.8 | 81.2 | 92.81 | |
| Ovarian cancer | IGROV1 | 101.47 | 100.15 | 101.83 | 103.23 | 100.68 |
| OVCAR-4 | 97.43 | 101.73 | 100.04 | 99.98 | 99.83 | |
| OVCAR-5 | 106.86 | 105.28 | 107.36 | 109.29 | 108.52 | |
| OVCAR-8 | 104.89 | 102.44 | 102.04 | 103.1 | 100.16 | |
| NCI/ADR-RES | 103.59 | 105.69 | 104.29 | 109.28 | 102.62 | |
| SK-OV-3 | 99.71 | 117.26 | 104.31 | 99.41 | 96.9 | |
| Renal cancer | 786-0 | 103.09 | 95.91 | 96.7 | 99.02 | 108.87 |
| A498 | 85.8 | 82.33 | 96.98 | 88.88 | 97.29 | |
| ACHN | 104.54 | 108.64 | 106.13 | 107.44 | 101.84 | |
| CAKI-1 | 91.57 | 92.07 | 93.33 | 90.39 | 92.32 | |
| RXF 393 | 104.79 | 95.74 | 97.24 | 90.04 | 108.78 | |
| SN12C | 101.37 | 101.44 | 108.61 | 104.67 | 109.12 | |
| TK-10 | 139.02 | 106.25 | 115.09 | 137.51 | 166.31 | |
| UO-31 | 84.7 | 89.07 | 99.42 | 87.04 | 90.13 | |
| Prostate cancer | PC-3 | 100.79 | 97.2 | 106.48 | 106.18 | 97.29 |
| DU-145 | 100.94 | 97.37 | 104.22 | 104.56 | 108.1 | |
| Breast cancer | MCF7 | 72.19 | 73.18 | 70.69 | 73.68 | 66.8 |
| MDA-MB-231/ATCC | 101.42 | 107.66 | 110.69 | 100.79 | 98.02 | |
| HS 578T | 98.86 | 107.36 | 108.02 | 98.34 | 85.53 | |
| BT-549 | 135.75 | 153.28 | 139.59 | 137.37 | 138.14 | |
| T-47D | 89.48 | 94.42 | 100.39 | 94.79 | 92.59 | |
| MDA-MB-468 | 111.12 | 108.91 | 108.55 | 106.92 | 115.34 |
| Panel | Subpanel | 6b | 6c | 7a | 7b | 7c |
|---|---|---|---|---|---|---|
| Leukemia | CCRF-CEM | 101.59 | 102.71 | 102.64 | 94.39 | 101.01 |
| HL-60(TB) | 108.02 | 115.55 | 96.19 | 107.89 | 93.51 | |
| K-562 | 98.1 | 95.31 | 89.05 | 99.87 | 92.74 | |
| MOLT-4 | 130.77 | 107.25 | 82.81 | 90.29 | 83.73 | |
| RPMI-8226 | 97.17 | 95.39 | 90.08 | 93.66 | 99.15 | |
| SR | NT | 104.75 | 104.95 | 96.21 | 94.84 | |
| Non-small cell lung cancer | A549/ATCC | 101.42 | 103.57 | 100.35 | 95.19 | 99.23 |
| EKVX | 88.07 | 95.51 | 88.5 | 95.66 | 96.85 | |
| HOP-62 | 106.95 | 127.09 | 108.62 | 109.06 | 113.73 | |
| HOP-92 | NT | 128.05 | 100.77 | NT | NT | |
| NCI-H226 | 90.52 | 99.81 | 92.31 | 94.54 | 89.98 | |
| NCI-H23 | 90.14 | 93.54 | 89.89 | 82.07 | 87.71 | |
| NCI-H322M | 96.92 | 97.37 | 88.83 | 90.33 | 92.11 | |
| NCI-H460 | 95.37 | 101.79 | 98.19 | 107.05 | 104.48 | |
| NCI-H522 | 97.93 | 96.57 | 94.51 | 94.8 | 94.88 | |
| Colon cancer | COLO 205 | 111.59 | 117.7 | 98.56 | 103.62 | 110.17 |
| HCC-2998 | 108.54 | 110.44 | 112.06 | 105.99 | 107.85 | |
| HCT-116 | 102.68 | 107.78 | 95.79 | 92.49 | 93.2 | |
| HCT-15 | 99.9 | 101.07 | 98.35 | 92.8 | 97.74 | |
| HT29 | 109.22 | 107.81 | 108.02 | 100.23 | 114.33 | |
| KM12 | 101.61 | 102.17 | 99.46 | 103.04 | 101.25 | |
| SW-620 | 98.43 | 95.33 | 96.83 | 95.97 | 93.58 | |
| CNS cancer | SF-268 | 109.07 | 112.22 | 102.76 | 112.81 | 107.64 |
| SF-295 | 111.41 | 103.52 | 94.8 | 106.74 | 99.31 | |
| SF-539 | 104 | 106.8 | 98.35 | 101.44 | 100.42 | |
| SNB-19 | 98.66 | 91.94 | 89.69 | 88.51 | 89.71 | |
| SNB-75 | 113.78 | 109.48 | 99.16 | 104.44 | 130.8 | |
| U251 | 99.84 | 109.34 | 97.89 | 93.41 | 94.98 | |
| Melanoma | LOX IMVI | 94.08 | 108.55 | 91.18 | 95.53 | 95.21 |
| MALME-3M | 89.96 | 97.08 | 92.4 | 88.26 | 95.69 | |
| M14 | 96.45 | 108.88 | 101.87 | 97.61 | 103.59 | |
| MDA-MB-435 | 107.8 | 106.7 | 102.07 | 102.21 | 107.15 | |
| SK-MEL-2 | 104.61 | 102.93 | 104.26 | 102.79 | 98.43 | |
| SK-MEL-28 | 110.85 | 105.01 | 105.23 | 103.19 | 105.46 | |
| SK-MEL-5 | 99.5 | 99.03 | 97.02 | 98.73 | 98.62 | |
| UACC-257 | 106.57 | 111.27 | 106.65 | 103.65 | 105.57 | |
| UACC-62 | 92.57 | 91.32 | 78.01 | 80.96 | 82.72 | |
| Ovarian cancer | IGROV1 | 94.42 | 100.88 | 90.69 | 96.92 | 93.63 |
| OVCAR-4 | 105 | 106.73 | 93.58 | 100.8 | 107.56 | |
| OVCAR-5 | 114.43 | 106.86 | 98.14 | 95.78 | 95.26 | |
| OVCAR-8 | 99.8 | 103.1 | 95.46 | 98.19 | 95.0 | |
| NCI/ADR-RES | 106.04 | 108.51 | 93.57 | 95.89 | 95.12 | |
| SK-OV-3 | 95.59 | 106.37 | 95.51 | 97.26 | 109.88 | |
| Renal cancer | 786-0 | 106.41 | 104.05 | 105.86 | 102.06 | 99.85 |
| A498 | 100.85 | 95.54 | 92.64 | 101.26 | 90.98 | |
| ACHN | 100.3 | 106.9 | 93.05 | 101.28 | 93.8 | |
| CAKI-1 | 88.84 | 97.44 | 81.1 | 86.46 | 88.64 | |
| RXF 393 | 95.8 | 104.97 | 95.21 | 86.15 | 97.03 | |
| SN12C | 103.56 | 94.55 | 98.94 | 97.12 | 94.91 | |
| TK-10 | 192.89 | 143.84 | 139.21 | 123.1 | 128.19 | |
| UO-31 | 77.59 | 89.45 | 71.06 | 75.37 | 81.23 | |
| Prostate cancer | PC-3 | 95.14 | 96.72 | 83.32 | 84.92 | 85.07 |
| DU-145 | 109.19 | 107.36 | 105.52 | 113.63 | 105.64 | |
| Breast cancer | MCF7 | 59.13 | 82.22 | 75.46 | 75.09 | 71.3 |
| MDA-MB-231/ATCC | 95.66 | 94.38 | 89.88 | 94.8 | 99.92 | |
| HS 578T | 92.58 | 94.29 | 89.37 | 91.01 | 92.8 | |
| BT-549 | 144.51 | 128.5 | 131.3 | 125.51 | 129.01 | |
| T-47D | 87.95 | 97.86 | 85.39 | 89.02 | 89.59 | |
| MDA-MB-468 | 110.50 | 118.74 | 79.09 | 90.36 | 93.21 |
In series 5, compound 5a with an unsubstituted phenyl moiety exhibited moderate growth inhibition in non-small cell lung cancer NCI-H23 (13.47%), melanoma UACC-62 (18.67%), renal cancer A498 (14.20%), renal cancer UO-31 (15.30%), and breast cancer MCF7 (27.81%). Introduction of a fluoro substituent at the 3-position in 5b enhanced antiproliferative effects across leukemia K-562 (26.04%), leukemia SR (15.70%), colon cancer HCT-116 (16.19%), melanoma UACC-62 (19.29%), renal cancer A498 (17.67%), and breast cancer MCF7 (26.82%). The 3-methylphenyl analog 5c showed activity against leukemia K-562 (21.84%), melanoma UACC-62 (15.20%), and breast cancer MCF7 (29.31%). Compound 5d, bearing a 4-fluorophenyl group, inhibited leukemia HL-60 (TB) (19.40%), leukemia K-562 (15.55%), melanoma UACC-62 (18.80%), renal cancer UO-31 (12.96%), and breast cancer MCF7 (26.32%).
In series 6, compound 6a (phenyl moiety) reduced cell growth in NSCLC EKVX (13.13%), melanoma MALME-3M (13.09%), breast cancer MCF7 (33.20%), and breast cancer HS 578T (14.47%). The 3-fluorophenyl derivative 6b exhibited enhanced inhibition in renal cancer CAKI-1 (11.16%), breast cancer MCF7 (40.87%), and breast cancer T-47D (12.05%). The 3-methylphenyl compound 6c was active against renal cancer UO-31 (10.55%) and breast cancer MCF7 (17.78%).
In series 7, compound 7a (phenyl moiety) showed growth inhibition in melanoma UACC-62 (21.99%), renal cancer CAKI-1 (18.90%), renal cancer UO-31 (28.94%), prostate cancer PC-3 (16.68%), breast cancer MCF7 (24.54%), breast cancer T-47D (14.61%), and breast cancer MDA-MB-468 (20.91%). The 3-fluorophenyl derivative 7b inhibited NSCLC NCI-H23 (17.93%), melanoma UACC-62 (19.04%), renal cancer UO-31 (24.63%), prostate cancer PC-3 (15.08%), and breast cancer MCF7 (24.91%). The 3-methylphenyl analog 7c was active against leukemia MOLT-4 (16.27%), melanoma UACC-62 (17.28%), prostate cancer PC-3 (14.93%), and breast cancer MCF7 (28.70%).
The structure–activity relationship (SAR) analysis indicates that phenyl ring substitutions significantly influence antiproliferative activity, likely via modulation of EGFR affinity and downstream signaling. Fluoro substituents, as in 5b, 6b, and 7b, enhance activity potentially through increased lipophilicity, metabolic stability, and electronic effects that strengthen binding interactions within the EGFR ATP-binding pocket. The electron-withdrawing nature of fluorine can optimize hydrogen bonding and van der Waals contacts with key residues, improving binding affinity and potency. Positioning of the fluoro group (meta or para) may also favor steric complementarity within the receptor site, consistent with observed higher growth inhibition, such as 40.87% inhibition of MCF7 cells by 6b.
Methyl substituents in 5c, 6c, and 7c exert distinct effects: as electron-donating groups via hyperconjugation, they increase electron density on the aromatic ring and promote hydrophobic interactions within EGFR's binding pockets. However, steric bulk introduced by methyl groups may influence binding orientation, accounting for varied activity profiles compared to fluoro-substituted analogs. This is reflected in significant antiproliferative effects, including 29.31% inhibition of MCF7 cells by 5c.
Overall, fluoro and methyl substituents modulate the electronic and steric environment of chalcone, pyrazoline, and acetyl pyrazoline scaffolds, impacting EGFR affinity and antiproliferative efficacy. Fluoro substituents predominantly enhance potency via electron-withdrawing effects and metabolic stability, while methyl groups contribute through electron donation and hydrophobic interactions, yielding complementary activity profiles across diverse cancer cell lines. A significant finding is that all tested compounds consistently exhibited their highest antiproliferative activity against MCF-7 cells, which are characterized by relatively elevated EGFR expression. Intriguingly, 7c containing a 3-methylphenyl moiety inhibited the growth of leukemia MOLT-4 (16.27%), melanoma UACC-62 (17.28%), prostate cancer PC-3 (14.93%), and breast cancer MCF7 (28.70%).
| Compound | IC50 μM |
|---|---|
| 6b | 0.225 ± 0.03 |
| Erlotinib | 0.198 ± 0.04 |
| Compound | Caspase-7 conc. | FLD |
|---|---|---|
| 6b/MCF7 | 396.13 | 7.38 |
| Cont. MCF7 | 52.97 | 1 |
Compared with the control group, compound 6b markedly increased the total apoptotic population (18.22 ± 1.9% vs. 1.57 ± 0.6%). This effect was primarily attributed to a pronounced elevation in late apoptosis (12.61 ± 1.4% vs. 0.52 ± 0.4%), while early apoptosis was also increased (4.34 ± 0.8% vs. 0.64 ± 0.3%). In contrast, necrotic cell death remained low and showed only a slight increase following 6b treatment (1.27 ± 0.2% vs. 0.41 ± 0.1%) (Fig. 5). These findings indicate that 6b predominantly induces apoptotic cell death in MCF-7 cells rather than necrosis.
| Cod. | Bax conc. | FLD | P53 conc. | FLD | Bcl-2 conc. | FLD |
|---|---|---|---|---|---|---|
| 6b/MCF7 | 408.35 | 6.8 | 425.64 | 7.58 | 2.17 | 0.37 |
| Cont. MCF7 | 60.05 | 1 | 56.19 | 1 | 5.86 | 1 |
These apoptotic changes can be mechanistically linked to the inhibition of EGFR signaling by compound 6b, as demonstrated by docking, molecular dynamics, and cytotoxicity data. EGFR inhibition disrupts downstream pro-survival pathways such as PI3K/AKT and RAS/RAF/MEK/ERK, which normally promote cell proliferation and suppress apoptosis. The blockade of these pathways reduces the transcription and activity of anti-apoptotic proteins like Bcl-2, while enhancing pro-apoptotic factors including Bax and the tumor suppressor p53. The elevated Bax/Bcl-2 ratio increases mitochondrial outer membrane permeability, leading to cytochrome c release, caspase activation, and programmed cell death.
Therefore, the observed modulation of Bax, Bcl-2, and p53 gene expression upon 6b treatment aligns with its demonstrated EGFR inhibitory activity and provides a cohesive mechanistic link between receptor blockade and activation of the mitochondrial apoptotic pathway in MCF-7 cells (Table 5).
3.3. In silico analysis
6b was then docked in the wild-type EGFR TK (active state) using the extra precision mode in Glide. The results showed that 6b adapted a relatively similar docking pattern in the active site that is close to the reference drug erlotinib pose, which suggests the potential affinity of 6b to the EGFR active site (Fig. 9). The calculated RMSD for the binding pose in the reference to erlotinib was 0.0217 Å.
To examine the strength of the interaction between the ligand and EGFR TK protein, as well as to understand the nature of the interaction and determine the energy contribution of each ligand residue, molecular mechanics, generalized Born, and surface area solvation (MM/GBSA) free binding energy calculations were conducted.81 These are known for their accuracy and cost-effectiveness compared to other methods. The results showed 6b having lower binding affinity than the reference antagonist, erlotinib (Table 6).
| Compound | Glide score docking (rigid)a kcal mol−1 | Induced-fit docking (IFD) (flexible)a kcal mol−1 | Free binding energy (MM/GBSA)a kcal mol−1 | Ionic interaction | H-bond interaction | Hydrogen bond length (Å) | Hydrophobic interactions |
|---|---|---|---|---|---|---|---|
| a More negative values indicate higher binding interactions within the binding pocket. | |||||||
| 6b–EGFR (active conformation) | −6.488 | −8.774 | −58.42 | LYS721 | ASP831, THR766 | ASP831 = 1.87 | LEU694, VAL702 |
| GLU738 | MET769 | THR766 = 1.83 | |||||
| ASP831 | H2O (10) | MET769 = 2.96 | |||||
| H2O (10) = 1.86 and 2.23 | |||||||
| Erlotinib–EGFR (active conformation) | −9.668 | −10.706 | −94.21 | LYS704 | LYS704 | LYS704 = 3.06 | LEU694 |
| LYS721 | MET769 | MET769 = 1.95 | VAL702 | ||||
| ASP831 | CYS773 | CYS773 = 2.07 | ALA719 | ||||
| H2O (10) | H2O (10) = 1.92 | LEU764 | |||||
| LEU768 | |||||||
| MET769 | |||||||
| CYS773 | |||||||
| LEU820 | |||||||
The docking analysis of compound 6b within the active site of the wild-type EGFR tyrosine kinase domain (TKD) revealed a binding mode closely resembling that of erlotinib, with a low RMSD of 0.0217 Å indicating high pose accuracy. This similarity suggests that 6b can effectively occupy the ATP-binding pocket, a critical determinant of EGFR inhibition.
Key molecular interactions underpinning 6b's binding affinity include a robust hydrogen bonding network formed by the pyrazoline nitrogen atom with essential residues ASP831, THR766, and MET769, as well as coordination with a conserved water molecule. These hydrogen bonds stabilize the ligand within the binding pocket, mimicking crucial interactions observed in potent EGFR inhibitors.
In addition to hydrogen bonds, ionic–π interactions with residues LYS721 and GLU738 contribute electrostatic complementarity, facilitating proper ligand orientation and enhancing binding strength. The aromatic pyrazoline and pyrrolo–phenyl moieties engage in π–π stacking interactions with aromatic side chains lining the pocket, further stabilizing the ligand through favorable van der Waals forces and aromatic overlap.
Hydrophobic interactions with residues LEU694 and VAL702 anchor hydrophobic regions of 6b, reducing desolvation penalties and improving binding thermodynamics. These contacts are essential for maintaining the ligand's conformation and enhancing affinity.
Although rigid docking Glide scores for 6b were approximately 50% lower than those for erlotinib, induced fit docking (IFD), which accounts for receptor flexibility, improved binding scores to about 20% lower than the reference, reflecting a more accurate representation of the binding event.
Complementary MM/GBSA free energy calculations quantified the energetic contributions of these interactions, confirming the ligand's favorable binding profile despite somewhat lower affinity than erlotinib. The combined hydrogen bonding, π–π stacking, ionic–π, and hydrophobic interactions collectively rationalize 6b's observed antiproliferative activity, particularly its potent inhibition of MCF-7 cells known for elevated EGFR expression.
 | ||
| Fig. 11 RMSF of EGFR TKD (active state) when simulated in complex with 6b (A) or erlotinib (B) as well as the individual RMSDs of 6b and erlotinib (C). | ||
Furthermore, a closer observation of the histogram of interactions revealed MET769 as the major contributor of the established binding poses in both 6b and erlotinib. This amino acid residue participated in both H-bonds and water bridges in the case of 6b while it exclusively established H-bonds in the erlotinib–EGFR TKD (active state) complex (Fig. 12A and B). 6b established various hydrophobic interactions while erlotinib had about an equal number of hydrophobic interactions and those bridged by water molecules (Fig. 12B).
| Compound | Glide score docking (rigid)a kcal mol−1 | Induced-fit docking (IFD) (flexible)a kcal mol−1 | Free binding energy (MM/GBSA)a kcal mol−1 | Ionic interaction | H-bond interaction | Hydrogen bond length (Å) | Hydrophobic interactions |
|---|---|---|---|---|---|---|---|
| a More negative values indicate higher binding interactions within the binding pocket. | |||||||
| 6b–EGFR(inactive conformation) | −5.987 | −8.223 | −19.31 | LYS721 | THR830 | THR830 = 2.80 | VAL702 |
| ASP831 | ASP831 | ASP831 = 2.46 | LEU820 | ||||
| LEU834 | |||||||
| Erlotinib–EGFR(inactive conformation) | −7.779 | −10.409 | −45.64 | LYS721 | MET769 | MET769 = 2.15 | VAL702 |
| ASP776 | CYS773 | CYS773 = 1.96 | ALA719 | ||||
| ASP831 | ASP831 = 3.17 | MET742 | |||||
| LEU764 | |||||||
| LEU768 | |||||||
| MET769 | |||||||
| CYS773 | |||||||
| LEU820 | |||||||
| LEU834 | |||||||
Hydrophobic interactions with LEU694 predominate in the 6b inactive complex (Fig. 15A), while erlotinib maintains critical hydrogen bonding with MET769 (Fig. 15B), a key residue for EGFR inhibition.
These findings align with the known behavior of established EGFR inhibitors such as erlotinib, which preferentially bind to and stabilize the active conformation of EGFR TKD through persistent hydrogen bonds with residues like MET769. The relative instability and weaker interactions of 6b in the inactive state suggest reduced inhibitory potential in this conformation, consistent with the mechanism of many clinically validated EGFR inhibitors that target the active kinase state for effective inhibition.
Therefore, the MD results position 6b as a compound with binding characteristics and conformational preferences similar to known EGFR inhibitors, favoring the active TKD conformation for stable engagement. This supports the docking and binding energy data indicating 6b's potential as a competitive EGFR inhibitor with a binding profile characteristic of effective ATP-competitive inhibitors. The enhanced activity of compound 6b is attributed to the 3-fluoro substituent, which improves binding within the EGFR ATP pocket by optimizing hydrophobic contacts, maintaining the correct orientation toward the hinge residue Met793, and increasing ligand lipophilicity without steric penalty. These effects result in stronger ligand–receptor stabilization and superior biological activity relative to the phenyl and methyl analogues.
| ADMET parameters | Erlotinib (reference) | 6b |
|---|---|---|
| Absorption | ||
| Water solubility (log mol L−1) | −4.403 | −4.977 |
Caco-2 permeability (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Papp in 10−6 cm s−1) Papp in 10−6 cm s−1) |
1.238 | 1.649 |
| HIA (% absorbed) | 95.549 | 96.803 |
| Distribution | ||
| BBB permeability (log BB) | −0.67 | 0.767 |
| CNS permeability (log PS) | −3.384 | −1.545 |
| Metabolism | ||
| CYP450 2D6 substrate (yes/no) | No | No |
| CYP3A4 substrate (yes/no) | Yes | Yes |
| CYP450 1A2 inhibitor (yes/no) | Yes | Yes |
| CYP450 2C19 inhibitor (yes/no) | Yes | Yes |
| CYP450 2C9 inhibitor | Yes | No |
| CYP450 2D6 inhibitor (yes/no) | No | No |
| CYP450 3A4 inhibitor (yes/no) | Yes | Yes |
| Excretion | ||
| Total clearance (log ml min−1 kg−1) | 0.591 | 0.385 |
| Renal OCT2 substrate (yes/no) | No | Yes |
| Toxicity | ||
| Ames toxicity (yes/no) | No | Yes |
| Max. tolerated dose (log mg kg−1 per day) | 0.002 | 0.226 |
| hERG1 inhibition (yes/no) | No | No |
| Acute oral toxicity (LD50) (mol kg−1) | 2.368 | 2.17 |
| Hepatotoxicity (yes/no) | Yes | No |
Toxicological descriptors for AMES toxicity, acute oral toxicity, maximum tolerated dose, hepatotoxicity, and hERG inhibition were also computed to evaluate the pharmacodynamic properties of 6b. Interestingly, it showed favorable features to hERG inhibition and hepatotoxicity molecular descriptors, except for the AMES descriptor. Additionally, it displayed good predictive values below the acute oral toxicity test threshold. On the other hand, erlotinib was found to have a low estimated value for the toxic dose threshold of chemicals, which is consistent with other reports.83,84
The drug-likeness properties of 6b were assessed. Lipinski's rule of five was used in this study to determine the necessary descriptors. According to this rule, a molecule is considered a drug-like or lead contender if it has a molecular weight of less than 500 g mol−1. The hydrogen bond acceptor and donor number are less than 10 and 5, respectively, and the octanol–water partition coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P) is not greater than 5.75 If more than one of these properties is violated, the compound's absorption and penetration profile may not be good. 6b's drug-like profile is excellent, as shown in Table 9.
P) is not greater than 5.75 If more than one of these properties is violated, the compound's absorption and penetration profile may not be good. 6b's drug-like profile is excellent, as shown in Table 9.
| Compound | M. Wt. (K per Dalton) | Log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
No. H-bond donor | No. H-bond acceptor | No. of violation | Lipinski alert |
|---|---|---|---|---|---|---|
| 6b | 305.354 | 5.13 | 1 | 2 | 1 | Accepted |
| Erlotinib (reference) | 393.441 | 4.243 | 1.5 | 7.4 | 0 | Accepted |
4. Conclusion
This study reports the successful design and synthesis of a novel series of pyrrole–pyrazoline hybrids as potent EGFR inhibitors with promising antiproliferative activity across diverse human cancer cell lines. Compound 6b emerged as a lead candidate, demonstrating comparable EGFR enzymatic inhibition to erlotinib and significant cytotoxicity, particularly against MCF-7 breast cancer cells characterized by elevated EGFR expression (Fig. 16). Comprehensive SAR analysis revealed that fluoro substituents enhance potency through electronic and steric modulation of EGFR binding, whereas methyl groups contribute distinct hydrophobic interactions, collectively informing scaffold optimization.Molecular docking and dynamics simulations corroborated the preferential and stable binding of 6b to the active conformation of EGFR TKD, highlighting critical hydrogen bonding, ionic–π, π–π stacking, and hydrophobic interactions that underpin its inhibitory mechanism. The observed modulation of apoptotic markers, including increased Bax/Bcl-2 ratio and p53 upregulation, establishes a mechanistic link between EGFR blockade and mitochondrial apoptosis induction.
These findings underscore the therapeutic potential of this scaffold as a basis for developing selective EGFR-targeted anticancer agents. Future work will focus on structural modifications to enhance selectivity, metabolic stability, and pharmacokinetic profiles, guided by the established SAR and computational insights, aiming to advance these hybrids toward clinical evaluation.
Not applicable. This study utilized only established cancer cell lines (NCI-58 panel), which were obtained from publicly available repositories. No human subjects or identifiable personal data were involved; therefore, Institutional Review Board (IRB) approval was not required.
Conflicts of interest
The authors declare no conflicts of interest.Data availability
EGFR (active state) (PDB ID: 1M17) and EGFR (inactive state) (PDB ID: 2GS7) data for in silico experiments using molecular docking were obtained from the Research Collaboratory for Structural Bioinformatics (RCSB) Protein Data Bank (PDB). PDB DOI: “https://doi.org/10.2210/pdb1M17/pdb and https://doi.org/10.2210/pdb2GS7/pdb (accessed on 27 January 2024)”. The data presented in this study are available in the article and the supplementary information (SI).Supplementary information is available. See DOI: https://doi.org/10.1039/d5md00800j.
Acknowledgements
The authors would like to acknowledge the Deanship of Graduate Studies and Scientific Research, Taif University for funding this work. The authors are thankful to the Developmental Therapeutics Program of the United States National Cancer Institute for performing the anticancer testing.References
- World Health Organization, Global cancer burden growing amidst mounting need for services [Internet]. 2024 Feb 1 [cited 2025 Jan 31], Available from: https://www.who.int/news/item/01-02-2024-global-cancer-burden-growing--amidst-mounting-need-for-services Search PubMed.
- A. Zafar, S. Khatoon, M. J. Khan, J. Abu and A. Naeem, Advancements and limitations in traditional anticancer therapies: a comprehensive review of surgery, chemotherapy, radiation therapy, and hormonal therapy, Discover Oncol., 2025, 16(1), 607, DOI:10.1007/s12672-025-02198-8.
- Z. Hu and H. Bousbaa, Recent advances in anticancer strategies, Cancers, 2025, 17(2), 173, DOI:10.3390/cancers17020173.
- B. Liu, H. Zhou and L. Tan, et al. Exploring treatment options in cancer: tumor treatment strategies, Signal Transduction Targeted Ther., 2024, 9, 175, DOI:10.1038/s41392-024-01856-7.
- Z. Schrank, G. Chhabra and L. Lin, et al. Current molecular-targeted therapies in NSCLC and their mechanism of resistance, Cancers, 2018, 10(7), 224, DOI:10.3390/cancers10070224.
- M. A. Lemmon and J. Schlessinger, Cell signaling by receptor tyrosine kinases, Cell, 2010, 141(7), 1117–1134, DOI:10.1016/j.cell.2010.06.011.
- R. Roskoski Jr., The ErbB/HER family of protein-tyrosine kinases and cancer, Pharmacol. Res., 2014, 79, 34–74, DOI:10.1016/j.phrs.2013.11.002.
- R. Rosell, T. Moran and C. Queralt, et al. Screening for EGFR mutations in lung cancer, N. Engl. J. Med., 2009, 361(10), 958–967, DOI:10.1056/NEJMoa0904554.
- F. Ciardiello and G. Tortora, A novel approach in the treatment of cancer: targeting the epidermal growth factor receptor, Clin. Cancer Res., 2001, 7(10), 2958–2970 CAS.
- T. Zubair and D. Bandyopadhyay, Small molecule EGFR inhibitors as anti-cancer agents: discovery, mechanisms of action, and opportunities, Int. J. Mol. Sci., 2023, 24(3), 2651, DOI:10.3390/ijms24032651.
- M. Maemondo, A. Inoue and K. Kobayashi, et al. Gefitinib or chemotherapy for non–small-cell lung cancer with mutated EGFR, N. Engl. J. Med., 2010, 362(25), 2380–2388, DOI:10.1056/NEJMoa0909530.
- G. A. Masters, S. Temin and C. G. Azzoli, et al. Systemic therapy for stage IV non–small-cell lung cancer: American Society of Clinical Oncology clinical practice guideline update, J. Clin. Oncol., 2015, 33(30), 3488–3515, DOI:10.1200/JCO.2015.62.1342.
- G. S. Basarab, P. J. Hill and C. E. Garner, et al. Optimization of pyrrolamide topoisomerase II inhibitors toward identification of an antibacterial clinical candidate (AZD5099), J. Med. Chem., 2014, 57(14), 6060–6082, DOI:10.1021/jm500462x.
- M. S. Mohamed, R. A. El-Domany and R. H. Abdel-Hameed, Synthesis of certain pyrrole derivatives as antimicrobial agents, Acta Pharm., 2009, 59(2), 145–158, DOI:10.2478/v10007-009-0016-9.
- M. S. Mohamed, A. E. Rashad, M. E. A. Zaki and S. S. Fatahala, Synthesis and antimicrobial screening of some fused heterocyclic pyrroles, Acta Pharm., 2005, 55(3), 237–249 CAS , Available from: https://hrcak.srce.hr/16761.
- D. T. Mahajan, V. H. Masand, K. N. Patil, T. B. Hadda and V. Rastija, Integrating GUSAR and QSAR analyses for antimalarial activity of synthetic prodiginines against multi drug resistant strain, Med. Chem. Res., 2013, 22, 2284–2292, DOI:10.1007/s00044-012-0223-7.
- P. W. Okanya, K. I. Mohr, K. Gerth, R. Jansen and R. Muller, Marinoquinolines A–F, pyrroloquinolines from Ohtaekwangia kribbensis (Bacteroidetes), J. Nat. Prod., 2011, 74(4), 603–608, DOI:10.1021/np100625a.
- B. Pacorel, S. C. Leung and A. V. Stachulski, et al. Modular synthesis, and in vitro and in vivo antimalarial assessment of C-10 pyrrole Mannich base derivatives of artemisinin, J. Med. Chem., 2010, 53(2), 633–640, DOI:10.1021/jm901216v.
- W. Peng, D. C. Peltier and M. J. Larsen, et al. Identification of thieno[3,2-b]pyrrole derivatives as novel small molecule inhibitors of neurotropic alphaviruses, J. Infect. Dis., 2009, 199(7), 950–957, DOI:10.1086/597275.
- H. H. Wang, K. M. Qiu and H. E. Cui, et al. Synthesis, molecular docking, and evaluation of thiazolyl-pyrazoline derivatives containing benzodioxole as potential anticancer agents, Bioorg. Med. Chem., 2013, 21(2), 448–455, DOI:10.1016/j.bmc.2012.11.020.
- W. M. Zhang, M. Xing and T. T. Zhao, et al. Synthesis, molecular modeling and biological evaluation of cinnamic acid derivatives with pyrazole moieties as novel anticancer agents, RSC Adv., 2014, 4, 37197–37207, 10.1039/C4RA05257A.
- M. Biava, C. Battilocchio and G. Poce, et al. Improving the solubility of a new class of anti-inflammatory pharmacodynamic hybrids that release nitric oxide and inhibit cyclooxygenase-2 isoenzyme, Eur. J. Med. Chem., 2012, 58, 287–298, DOI:10.1016/j.ejmech.2012.10.014.
- A. Detsi, M. Majdalani, C. A. Kontogiorgis, D. Hadjipavlou-Litina and P. Kefalas, Natural and synthetic 2′-hydroxy-chalcones and aurones: synthesis, characterization and evaluation of the antioxidant and soybean lipoxygenase inhibitory activity, Bioorg. Med. Chem., 2009, 17(23), 8073–8085, DOI:10.1016/j.bmc.2009.10.002.
- L. Mishra, H. Itokawa and K. F. Bastow, et al. Anti-HIV and cytotoxic activities of Ru(II)/Ru(III) polypyridyl complexes containing 2,6-(2′-benzimidazolyl)-pyridine/chalcone as co-ligand, Bioorg. Med. Chem., 2001, 9(7), 1667–1671, DOI:10.1016/S0968-0896(01)00074-8.
- D. F. Romagnolo and O. I. Selmin, Flavonoids and cancer prevention: a review of the evidence, J. Nutr. Gerontol. Geriatr., 2012, 31(3), 206–238, DOI:10.1080/21551197.2012.702534.
- D. A. Israf, T. A. Khaizurin, A. Syahida, N. H. Lajis and S. Khozirah, Cardamonin inhibits COX and iNOS expression via inhibition of p65 NF-κB nuclear translocation and Iκ-B phosphorylation in RAW 264.7 macrophage cells, Mol. Immunol., 2007, 44(5), 673–679, DOI:10.1016/j.molimm.2006.04.025.
- K. Sangpheak, L. Tabtimmai and S. Seetaha, et al. Biological evaluation and molecular dynamics simulation of chalcone derivatives as epidermal growth factor-tyrosine kinase inhibitors, Molecules, 2019, 24(6), 1092, DOI:10.3390/molecules24061092.
- E. B. Yang, Y. J. Guo, K. Zhang, Y. Z. Chen and P. Mack, Inhibition of epidermal growth factor receptor tyrosine kinase by chalcone derivatives, Biochim. Biophys. Acta, 2001, 1550(2), 144–152, DOI:10.1016/S0167-4838(01)00276-X.
- H. A. Abdel-Aziz, H. S. A. El-Zahabi and K. M. Dawood, Microwave-assisted synthesis and in-vitro anti-tumor activity of 1,3,4-triaryl-5-N-arylpyrazole-carboxamides, Eur. J. Med. Chem., 2010, 45(6), 2427–2432, DOI:10.1016/j.ejmech.2010.02.026.
- H. A. Abdel-Aziz, A. A. I. Mekawey and K. M. Dawood, Convenient synthesis and antimicrobial evaluation of some novel 2-substituted-3-methylbenzofuran derivatives, Eur. J. Med. Chem., 2009, 44(9), 3637–3644, DOI:10.1016/j.ejmech.2009.02.020.
- F. M. Awadallah, G. A. Piazza and B. D. Gary, et al. Synthesis of some dihydro-pyrimidine based compounds bearing pyrazoline moiety and evaluation of their antiproliferative activity, Eur. J. Med. Chem., 2013, 70, 273–279, DOI:10.1016/j.ejmech.2013.10.003.
- C. N. Cavasotto, M. A. Ortiz, R. A. Abagyan and F. J. Piedrafita, In silico identification of novel EGFR inhibitors with antiproliferative activity against cancer cells, Bioorg. Med. Chem. Lett., 2006, 16(7), 1969–1974, DOI:10.1016/j.bmcl.2005.12.067.
- S. Jiang, H. Lu, S. Liu, Q. Zhao, Y. He and A. K. Debnath, N-substituted pyrrole derivatives as novel HIV-1 entry inhibitors that interfere with gp41 six-helix bundle formation and block virus fusion, Antimicrob. Agents Chemother., 2004, 48(11), 4349–4359, DOI:10.1128/AAC.48.11.4349-4359.2004.
- P. C. Lv, D. D. Li, Q. S. Li, X. Lu, Z. P. Xiao and H. L. Zhu, Synthesis, molecular docking and evaluation of thiazolyl-pyrazoline derivatives as EGFR TK inhibitors and potential anticancer agents, Bioorg. Med. Chem. Lett., 2011, 21(18), 5374–5377, DOI:10.1016/j.bmcl.2011.07.010.
- G. Menozzi, L. Mosti, P. Fossa, F. Mattioli and M. Ghia, ω-Dialkylaminoalkyl ethers of phenyl-(5-substituted 1-phenyl-1H-pyrazol-4-yl)methanols with analgesic and anti-inflammatory activity, J. Heterocyclic Chem., 1997, 34(3), 963–968, DOI:10.1002/jhet.5570340339.
- R. Sridhar, P. T. Perumal and S. Etti, et al. Design, synthesis and anti-microbial activity of 1H-pyrazole carboxylates, Bioorg. Med. Chem. Lett., 2004, 14(24), 6035–6040, DOI:10.1016/j.bmcl.2004.09.066.
- I. Vujasinovic, A. P. Radicevic, K. Brajsa and B. Bertosa, Synthesis and biological validation of novel pyrazole derivatives with anticancer activity guided by 3D-QSAR analysis, Bioorg. Med. Chem., 2012, 20(6), 2101–2110, DOI:10.1016/j.bmc.2012.01.032.
- H. A. S. Abbas and S. S. Abd El-Karim, Design, synthesis and anticervical cancer activity of new benzofuran–pyrazol-hydrazono-thiazolidin-4-one hybrids as potential EGFR inhibitors and apoptosis inducing agents, Bioorg. Chem., 2019, 89, 103035, DOI:10.1016/j.bioorg.2019.103035.
- S. S. Abd El-Karim, Y. M. Syam, A. M. El Kerdawy and T. M. Abdelghany, New thiazol-hydrazono-coumarin hybrids targeting human cervical cancer cells: synthesis, CDK2 inhibition, QSAR and molecular docking studies, Bioorg. Chem., 2019, 86, 80–96, DOI:10.1016/j.bioorg.2019.01.026.
- M. F. Ahmed and A. H. Almalki, Design, synthesis, antiproliferative activity, and cell cycle analysis of new thiosemicarbazone derivatives targeting ribonucleotide reductase, Arabian J. Chem., 2021, 14, 102989, DOI:10.1016/j.arabjc.2021.102989.
- K. M. Qiu, H. H. Wang and L. M. Wang, et al. Design, synthesis and biological evaluation of pyrazolyl-thiazolinone derivatives as potential EGFR and HER-2 kinase inhibitors, Bioorg. Med. Chem., 2012, 20(6), 2010–2018, DOI:10.1016/j.bmc.2012.01.051.
- B. Sharma, V. J. Singh and P. A. Chawla, Epidermal growth factor receptor inhibitors as potential anticancer agents: an update of recent progress, Bioorg. Chem., 2021, 116, 105393, DOI:10.1016/j.bioorg.2021.105393.
- X. X. Tao, Y. T. Duan, L. W. Chen, D. J. Tang, M. R. Yang and P. F. Wang, et al. Design, synthesis and biological evaluation of pyrazolyl-nitroimidazole derivatives as potential EGFR/HER-2 kinase inhibitors, Bioorg. Med. Chem. Lett., 2016, 26(2), 677–683, DOI:10.1016/j.bmcl.2015.11.040.
- N. H. Nasab, F. Azimian, Y. S. Eom, F. H. Shah and S. J. Kim, Synthesis, anticancer evaluation, and molecular docking studies of thiazolyl-pyrazoline derivatives, Bioorg. Med. Chem. Lett., 2023, 80, 129105, DOI:10.1016/j.bmcl.2022.129105.
- S. G. Nayak, V. B. Das, V. Kamat and V. K. D., Advances in the development of potent heterocyclic anticancer agents: a critical review, Synth. Commun., 2025, 55(19), 1437–1463, DOI:10.1080/00397911.2025.2529464.
- G. Iancu, D. Şerban, C. D. Badiu, C. Tănăsescu, M. S. Tudosie, C. Tudor, D. O. Costea, A. Zgura, R. Iancu and D. Vasile, Exp. Ther. Med., 2022, 23, 114, DOI:10.3892/etm.2021.11037.
- H. Ciftci, M. Otsuka, M. Fujita and B. Sever, New naphthalene-linked pyrazoline-thiazole hybrids as prominent antilung and antibreast cancer inhibitors, Turk. J. Chem., 2024, 48(6), 856–866, DOI:10.55730/1300-0527.3704.
- P. Shi, Y. Oh and L. Deng, et al. Overcoming acquired resistance to AZD9291, a third-generation EGFR inhibitor, through modulation of MEK/ERK-dependent Bim and Mcl-1 degradation, Clin. Cancer Res., 2017, 23(21), 6567–6579, DOI:10.1158/1078-0432.CCR-17-1574.
- V. R. Solomon, C. Hu and H. Lee, Design and synthesis of anti-breast cancer agents from 4-piperazinylquinoline: a hybrid pharmacophore approach, Bioorg. Med. Chem., 2010, 18(4), 1563–1572, DOI:10.1016/j.bmc.2010.01.001.
- H. Mahmoudi and A. A. Jafari, Facile preparation of sulfonic acid-functionalized magnetite-coated maghemite as a magnetically separable catalyst for pyrrole synthesis, ChemCatChem, 2013, 5, 3743–3749, DOI:10.1002/cctc.201300623.
- A. Monks, D. Scudiero and P. Skehan, et al. Feasibility of a high-flux anticancer drug screen using a diverse panel of cultured human tumor cell lines, J. Natl. Cancer Inst., 1991, 83(11), 757–766, DOI:10.1093/jnci/83.11.757.
- Y. Jia, C. M. Quinn, A. I. Gagnon and R. Talanian, Homogeneous time-resolved fluorescence and its applications for kinase assays in drug discovery, Anal. Biochem., 2006, 356(2), 273–281, DOI:10.1016/j.ab.2006.05.006.
- M. F. Ahmed, E. Y. Santali, E. M. M. El-Deen, I. A. Naguib and R. El-Haggar, Development of pyridazine derivatives as potential EGFR inhibitors and apoptosis inducers: design, synthesis, anticancer evaluation, and molecular modeling studies, Bioorg. Chem., 2021, 106, 104473, DOI:10.1016/j.bioorg.2020.104473.
- W. T. Huang, J. Liu and J. F. Liu, et al. Synthesis and biological evaluation of conjugates of deoxypodophyllotoxin and 5-FU as inducer of caspase-3 and -7, Eur. J. Med. Chem., 2012, 49, 48–54, DOI:10.1016/j.ejmech.2011.12.005.
- R. A. Friesner, J. L. Banks and R. B. Murphy, et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy, J. Med. Chem., 2004, 47(7), 1739–1749, DOI:10.1021/jm0306430.
- J. H. Park, Y. Liu, M. A. Lemmon and R. Radhakrishnan, Erlotinib binds both inactive and active conformations of the EGFR tyrosine kinase domain, Biochem. J., 2012, 448, 417, DOI:10.1042/BJ20121513.
- X. Zhang, J. Gureasko, K. Shen, P. A. Cole and J. Kuriyan, An allosteric mechanism for activation of the kinase domain of epidermal growth factor receptor, Cell, 2006, 125(6), 1137–1149, DOI:10.1016/j.cell.2006.05.013.
- C. Lu, C. Wu and D. Ghoreishi, et al. OPLS4: improving force field accuracy on challenging regimes of chemical space, J. Chem. Theory Comput., 2021, 17(7), 4291–4300, DOI:10.1021/acs.jctc.1c00302.
- J. C. Shelley, A. Cholleti, L. L. Frye, J. R. Greenwood, M. R. Timlin and M. Uchimaya, Epik: a software program for pKa prediction and protonation state generation for drug-like molecules, J. Comput.-Aided Mol. Des., 2007, 21(12), 681–691, DOI:10.1007/s10822-007-9133-z.
- A. Monks, D. Scudiero, P. Skehan, R. Shoemaker, K. Paull, D. Vistica, C. Hose, J. Langley, P. Cronise and A. Vaigro-Wolff, et al., J. Natl. Cancer Inst., 1991, 83, 757–766, DOI:10.1093/jnci/83.11.757.
- R. Jaundoo, J. Bohmann and G. E. Gutierrez, et al. Using a consensus docking approach to predict adverse drug reactions in combination drug therapies for Gulf War illness, Int. J. Mol. Sci., 2018, 19(11), 3355, DOI:10.3390/ijms19113355.
- S. K. Kwofie, B. Dankwa and E. A. Odame, et al. In silico screening of isocitrate lyase for novel anti-buruli ulcer natural products originating from Africa, Molecules, 2018, 23(7), 1550, DOI:10.3390/molecules23071550.
- M. S. Alturki, Exploring marine-derived compounds: in silico discovery of selective ketohexokinase (KHK) inhibitors for metabolic disease therapy, Mar. Drugs, 2024, 22(10), 455, DOI:10.3390/md22100455.
- W. Sherman, T. Day, M. P. Jacobson, R. A. Friesner and R. Farid, Novel procedure for modeling ligand/receptor induced fit effects, J. Med. Chem., 2006, 49(2), 534–553, DOI:10.1021/jm050540c.
- M. Tutone, I. Pibiri, L. Lentini, A. Pace and A. M. Almerico, Deciphering the nonsense readthrough mechanism of action of ataluren: an in silico compared study, ACS Med. Chem. Lett., 2019, 10(4), 522–527, DOI:10.1021/acsmedchemlett.8b00558.
- I. Massova and P. A. Kollman, Combined molecular mechanical and continuum solvent approach (MM-PBSA/GBSA) to predict ligand binding, Perspect. Drug Discovery Des., 2000, 18, 113–135, DOI:10.1023/A:1008763014207.
- P. Tuffery and P. Derreumaux, Flexibility and binding affinity in protein–ligand, protein–protein and multi-component protein interactions: limitations of current computational approaches, J. R. Soc., Interface, 2012, 9(66), 20–33, DOI:10.1098/rsif.2011.0584.
- T. A. Rants'o, C. J. van Vuuren and R. L. van Zyl, Optimization of covalent docking for organophosphates interaction with Anopheles acetylcholinesterase, J. Mol. Graphics Modell., 2022, 110, 108054, DOI:10.1016/j.jmgm.2021.108054.
- H. A. L. Filipe and L. M. S. Loura, Molecular dynamics simulations: advances and applications, Molecules, 2022, 27(7), 2105, DOI:10.3390/molecules27072105.
- A. Hospital, J. R. Goñi, M. Orozco and J. L. Gelpí, Molecular dynamics simulations: advances and applications, Adv. Appl. Bioinf. Chem., 2015, 8, 37–47, DOI:10.2147/AABC.S70333.
- M. S. Alturki, R. A. Alkhodier and M. S. Gomaa, et al. Structure-based discovery of orthosteric non-peptide GLP-1R agonists via integrated virtual screening and molecular dynamics, Int. J. Mol. Sci., 2025, 26(13), 6131, DOI:10.3390/ijms26136131.
- K. M. Callahan and B. Roux, Molecular dynamics of ion conduction through the selectivity filter of the NaVAb sodium channel, J. Phys. Chem. B, 2018, 122(43), 10126–10142, DOI:10.1021/acs.jpcb.8b09678.
- M. S. Alturki, A. H. Al Khzem and M. S. Gomaa, et al. Gallic acid: a potent metabolite targeting shikimate kinase in Acinetobacter baumannii, Metabolites, 2024, 14(12), 727, DOI:10.3390/metabo14120727.
- T. Pantsar, KRAS (G12C)–AMG 510 interaction dynamics revealed by all-atom molecular dynamics simulations, Sci. Rep., 2020, 10, 11992, DOI:10.1038/s41598-020-68950-y.
- D. E. V. Pires, T. L. Blundell and D. B. Ascher, pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures, J. Med. Chem., 2015, 58(9), 4066–4072, DOI:10.1021/acs.jmedchem.5b00104.
- C. A. Lipinski, Lead- and drug-like compounds: the rule-of-five revolution, Drug Discovery Today: Technol., 2004, 1(4), 337–341, DOI:10.1016/j.ddtec.2004.11.007.
- K. Cao and S. W. G. Tait, Apoptosis and cancer: force awakens, phantom menace, or both?, Int. Rev. Cell Mol. Biol., 2018, 337, 135–152, DOI:10.1016/bs.ircmb.2017.12.003.
- C. M. Pfeffer and A. T. K. Singh, Apoptosis: a target for anticancer therapy, Int. J. Mol. Sci., 2018, 19(2), 448, DOI:10.3390/ijms19020448.
- B. Antonsson, Mitochondria and the Bcl-2 family proteins in apoptosis signaling pathways, Mol. Cell. Biochem., 2004, 256(1–2), 141–155, DOI:10.1023/B:MCBI.0000009865.70898.36.
- Y. R. Saadat, N. Saeidi, S. Z. Vahed, A. Barzegari and J. Barar, An update to DNA ladder assay for apoptosis detection, BioImpacts, 2015, 5(1), 25, DOI:10.15171/bi.2015.01.
- C. N. Cavasotto, Binding free energy calculation using quantum mechanics aimed for drug lead optimization, in Quantum Mechanics in Drug Discovery, 2020, pp. 257–268, DOI:10.1007/978-1-0716-0282-9_16.
- U. Dasmahapatra, C. K. Kumar and S. Das, et al. In-silico molecular modelling, MM/GBSA binding free energy and molecular dynamics simulation study of novel pyrido fused imidazo[4,5-c]quinolines as potential anti-tumor agents, Front. Chem., 2022, 10, 991369, DOI:10.3389/fchem.2022.991369.
- L. Guan, H. Yang and Y. Cai, et al. ADMET-score–a comprehensive scoring function for evaluation of chemical drug-likeness, MedChemComm, 2019, 10, 148–157, 10.1039/C8MD00472B.
- M. Vigano, M. L. Milia and M. V. Grassini, et al. Hepatotoxicity of small molecule protein kinase inhibitors for cancer, Cancers, 2023, 15, 1766, DOI:10.3390/cancers15061766.
| This journal is © The Royal Society of Chemistry 2026 |















