Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
The influence of polymer topology and side chain functionality on the Schiff-base reactivity of biocompatible polypeptide hydrogels
Robert D.
Murphy
 *a,
Franz
Lobianco
b,
Viviane
Chiaradia
a,
Josué M.
Galindo
a,
Tom
Hodgkinson
*a,
Franz
Lobianco
b,
Viviane
Chiaradia
a,
Josué M.
Galindo
a,
Tom
Hodgkinson
 b and
Andreas
Heise
b and
Andreas
Heise
 acd
acd
aDepartment of Chemistry, RCSI University of Medicine and Health Sciences, 123 St. Stephen's Green, Dublin 2, Ireland. E-mail: robertdmurphy@rcsi.com
bTissue Engineering Research Group, Department of Anatomy and Regenerative Medicine, RCSI University of Medicine and Health Sciences, 123, St Stephen's Green, Dublin 2, Ireland
cThe SFI Centre for Advanced Materials and BioEngineering Research, RCSI University of Medicine and Health Science, Dublin 2, Ireland
dCURAM the SFI Research Centre for Medical Devices, Department of Chemistry, RCSI University of Medicine and Health Sciences, Dublin 2, Ireland
First published on 6th March 2026
Abstract
Presented are bio-inspired, synthetic polypeptide-based hydrogels derived from Schiff-base chemistry. Through the N-carboxyanhydride ring opening polymerisation (NCA ROP), polypeptides with linear and star topologies were readily synthesised and modified by post-polymerisation modification to form aldehyde and acyl hydrazide rich complimentary crosslinkers. Through mixing pure polymer topologies (e.g., linear and linear crosslinkers) or topological blends (e.g., star and linear crosslinkers), a wide range of mechanical properties could be accessed, forming robust, biocompatible hydrogels.
1. Introduction
Hydrogels are an important class of soft viscoelastic materials used in a wide variety of applications which include tissue engineering,1 and flexible electronics.2 The chemistry of hydrogels is highly versatile,3 though conventionally, chain-growth polymerisation reactions are utilised to induce covalent network crosslinking.4 As an alternative approach, covalent crosslinking of hydrogels can also be achieved using step-growth reactions of complimentary reactive functional groups.5 A more recent adaptation of step-growth reactivity comes in the form of dynamic covalent chemistry (DCC), which has been used to great effect to enhance embedded properties, converting hydrogel networks from static to dynamic.6,7 Hydrogels with truly modular properties and on-demand reversibility have been fabricated through reactions which form boronate esters,8,9 disulphides,10,11 Diels–Alder adducts,12,13 and Schiff-base adducts.14,15The Schiff-base reaction in particular has garnered much attention to this end, whereby the dynamic, reversible nature of bonds formed through condensation of nucleophilic (e.g., amino) and aldehyde groups, can endow hydrogels with unique properties such as pH response, self-healing capabilities (reversible bonding), and tuneable gelation kinetics.16 Since the Schiff-base formation is in equilibrium with hydrolysis,17 the pH response and reversibility is contingent on the pKa of the nucleophilic species used, which can be amine, hydrazide, acyl hydrazide, or aminooxy groups.18 Schiff-base governed crosslinking is widely adapted for hydrogels, using natural biopolymers,19 or synthetic polymers.15 It has also emerged as a key tool for crosslinking of proteins or (poly)peptides, an underexplored class of materials ideally suitable for hydrogel fabrication.20 Bioinspired synthetic polypeptides can be readily prepared via the N-carboxyanhydride ring opening polymerisation (NCA ROP),21,22 with a range of architectures,23,24 enabling rapid access to versatile hydrogel materials based on the chemistry of amino acid residue.25–31 To this end, Schiff-base reactivity can be leveraged from these modular synthetic polypeptides, via straightforward side chain modification.32 The Yin group have successfully demonstrated a range of side chain modified polypeptides, such as oxidation of diols to aldehydes,33 and amidation with acyl hydrazides,34 enabling hydrogel fabrication from Schiff-base reactivity. The studies also highlight the hydrogels’ adaptability as viable cartilage tissue engineering scaffolds, demonstrating the remarkable potential for polypeptides within biomedical applications.35
In terms of design principles for Schiff-base chemistry, some studies have elucidated key differences in reactivity profiles and stability of aldehyde reactions with oximes, amines and acyl hydrazides.36 To this end, the nature of aldehyde chemistry has also been examined recently.37 In addition to nucleophile type, polymer architecture may also influence Schiff-base reactivity, with star or branched polymer topologies differing significantly from traditional linear polymers.38 While star or branched polymers have been used for Schiff-base hydrogel crosslinking,14,39 there are only a few studies which loosely demonstrate how polymer architectures can change reactivity and hydrogel properties,40,41 while no studies have utilised polypeptides for this purpose. Therefore, a more systematic understanding of how polymer topological blends (linear or star) may impact Schiff-base reactivity would be beneficial for the field.
Herein, we disclose the synthesis and characterisation of a library of linear and star-shaped polypeptides, exploring their use as a versatile Schiff-base hydrogel platform based on polymer topology. Using NMR spectroscopy, circular dichroism and rheological analysis, the structure–property relationship could be elucidated to identify gelation times and gel strength, which was found to be directly governed by polymer architecture. Moreover, these remarkably tuneable polypeptide hydrogels also served as highly biocompatible matrices for encapsulated human mesenchymal stem cells (hMSCs).
2. Experimental
2.1. Linear poly(γ-benzyl-L-glutamine) (l-PBLG)
γ-benzyl-L-glutamate NCA (42.00 g, 1.60 × 103 mmol) was dissolved in 1000 mL of CHCl3. Hexylamine (161.00 mg, 1.60 mmol) and 18-C-6 (423.00 mg, 1.60 mmol) in 5 mL CHCl3 were then quickly added to the solution. The reaction was continued at room temperature until full conversion of NCA was confirmed by FTIR spectroscopy. The solution was then reduced in vacuo to 1/3 its volume, followed by precipitation into excess diethyl ether (1.1 L) and drying in vacuo (yield: 31.40 g, 89%).2.2. Star poly(γ-benzyl-L-glutamate) (s-PBLG)
γ-benzyl-L-glutamate NCA (5.00 g, 1.90 × 101 mmol) was dissolved in 80 mL of CHCl3. G1 PPI dendrimer (DAB-Am-4) (50.00 mg, 1.58 × 10−1 mmol) in 1 mL CHCl3 was then quickly added to the solution. The reaction was continued at room temperature until full conversion of NCA was confirmed by FTIR spectroscopy. The solution was then precipitated into excess diethyl ether (300 mL) and dried in vacuo (yield: 3.95 g, 95%).2.3. Linear poly(hydroxyethyl-L-glutamine) (l-PHELG)
The l-PBLG (6.00 g, 2.53 × 10−1 mmol) was dissolved in 70 mL DMF, before addition of ethanolamine (16.70 g, 2.73 × 102 mmol) and 2-hydroxypyridine (5.20 g, 5.46 × 101 mmol). The reaction solution was stirred for 3 days at 50 °C, before cooling and precipitation into 350 mL diethyl ether. The residue was then dissolved in 30 mL deionised water and dialysed for 3 days before lyophilisation to yield l-PHELG (yield: 3.75 g, 80%). *s-PHELG was prepared in a similar manner.2.4. Linear poly(hydroxyethyl-L-glutamine-co-formylethylester-L-glutamine) (PHELG-co-PFEELG) (l-PBzAld)
The l-PHELG (210.00 mg, 1.05 × 10−2 mmol) was dissolved in 5 mL DMSO, before addition of 4-dimethylaminopyridine (4.30 mg, 3.53 × 10−2 mmol), 4-formyl-benzoic acid (53.00 mg, 3.53 × −1 mmol), and 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide hydrochloride (96.80 mg, 5.05 × 10−1 mmol). The reaction was stirred for 3 days at room temperature, before precipitation into 80 mL diethyl ether and drying. The residue was then dissolved in 5 mL deionised water and dialysed for 3 days before lyophilisation to yield l-PBzAld (yield: 165.00 mg, 76%). *s-PBzAld was prepared in a similar manner.2.5. Linear poly(acyl hydrazide-L-glutamine) (PAHLG) (l-PAHz1)
The l-PBLG (4.00 g, 1.57 × 10−1 mmol) was dissolved in 160 mL DMF, before dropwise addition of hydrazine hydrate (11.70 g, 3.64 × 102 mmol). The reaction solution was stirred for 1 day at room temperature, before precipitation into 800 mL diethyl ether. The residue was then suspended in 50 mL deionised water, and adjusted to pH 4 with 1M HCl. It was dialysed for 3 days before lyophilisation to yield l-PAHz1 (yield: 1.60 g, 49%). *s-PAHz1 was prepared in a similar manner.2.6. Linear poly(acyl hydrazide-L-glutamine-co-hydroxyethyl-L-glutamine) (PAHLG-co-PHELG) (l-PAHz2)
The l-PBLG (1.00 g, 3.92 × 10−2 mmol) was dissolved in 10 mL DMF, before addition of hydrazine hydrate (200.00 mg, 6.27 mmol) and ethanolamine (1.72 g, 2.82 × 101 mmol). The reaction solution was stirred for 3 days at 50 °C, before cooling and precipitation into 120 mL diethyl ether and drying. The residue was then suspended in 20 mL deionised water, and adjusted to pH 4 with 1M HCl. It was dialysed for 3 days before lyophilisation to yield l-PAHz2 (yield: 505.00 mg, 63%). *s-PAHz1 was prepared in a similar manner.2.7. CD spectroscopy
CD spectroscopy was performed on an Applied Photophysics Chirascan Plus Circular Dichroism Spectrometer at 20 °C. Polymer solutions were prepared a concentration of 0.01 mg mL−1 and analysed in a quartz cuvette with a path length of 1 cm. Secondary structural features of recorded CD data were determined using online software BestSel.2.8. Hydrogel formulation
Hydrogels were prepared by separately dissolving the PBzAld and PAHz polypeptides in phosphate buffered saline (PBS) solution at 5.0 wt% respectively, to form viscous solutions. The two solutions were then blended together to form hydrogels at room temperature. See Table S3 for more details.2.9. Rheology
Rheological measurements of hydrogels were completed on an MCR 301 digital rheometer (Anton Paar). Hydrogels were formulated as described above (Section S2.8, SI) and then immediately analysed on the rheometer. All experiments were conducted at room temperature (20 °C) using a parallel plate (PP25, Anton Paar) consisting of a 25 mm diameter geometry and a gap length of 0.095 mm. The use of a protective hood was employed to prevent evaporation.3. Results and discussion
3.1. Synthesis and characterisation of polypeptide topologies
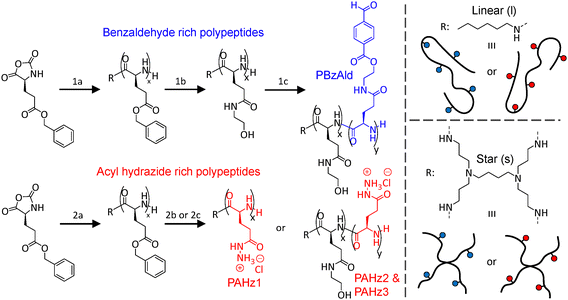
For the synthesis of the polymer library, the NCA monomer of benzyl-L-glutamate (BLG) was used to create polypeptides with readily amenable side chains (Fig. 1). BLG NCA was initially synthesised and its purity confirmed by 1H NMR spectroscopy (Fig. S1). To create linear and star polymer topologies, ring opening polymerisation (ROP) of BLG NCA was conducted using monofunctional hexylamine in the presence of a crown ether (18-C-6) catalyst, or multifunctional 4-armed poly(propylene imine) (PPI) dendrimer initiators (Fig. 1, step 1a/2a).42 A monomer/initiator ([M]0/[I]0) ratio of 100/1 was maintained to afford linear and star poly(BLG) (l-PBLG) and (s-PBLG) with an overall degree of polymerisation (DP) of 116 and 110 respectively, as confirmed by 1H NMR spectroscopy (Fig. S2 and S3). Size exclusion chromatography (SEC) confirmed the formation of l-PBLG116 and s-PBLG110 with controllable molecular weights (Mn) and low dispersities (Đ) (Fig. S4). Linear and star polypeptides with aldehyde and acyl hydrazide side chains were then formed by post-polymerisation modification of PBLGs. Aldehyde decorated polypeptides were accessed through aminolysis of the PBLGs, subjecting the benzyl esters to ethanolamine and 2-hydroxypyridine (Fig. 1, step 1b). Non-ionic, hydrophilic functionalities were endowed to linear and star topologies, with 1H NMR spectroscopy confirming resultant poly(hydroxyethyl-L-glutamine) (PHELG) structures (Fig. S5 and S6). The hydroxy side chains were then condensed with 4-formylbenzoic acid, targeting 30% of PHELG of linear (35/116) and star (33/110) polypeptides for benzaldehyde (BzAld) functionalisation, using 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide.hydrochloride (EDC·HCl) and 4-dimethylaminopyridine (DMAP) (Fig. 1, step 1c). The degree of functionalisation (DF) of the newly formed linear and star poly(hydroxyethyl-L-glutamine-co-formylethylester-L-glutamine) (PHELG-co-PFEELG), termed l-PBzAld and s-PBzAld herein, was found to be 9 units (8%) respectively, as determined by 1H NMR spectroscopy (Fig. S7 and S8). Acyl hydrazide (AHz) functionalised polypeptides were formed in a similar manner, whereby increasing amounts of AHz groups were introduced via aminolysing with either hydrazine monohydrate alone (Fig. 1, step 2b), or a mixture of hydrazine monohydrate with ethanolamine (Fig. 1, step 2c).43 This targeted homopolypeptides of poly(acyl hydrazide-L-glutamine) (PAHLG), and copolypeptides of poly(acyl hydrazide-L-glutamine-co-hydroxyethyl-L-glutamine) (PAHLG-co-PHELG). Amount of AHz for DF% was targeted at 100%, 20% and 10% for polypeptides, specifically termed l-PAHz1-3 for linear, and s-PAHz1-3 for star herein. Successful synthesis was confirmed by 1H NMR spectroscopy for linear polypeptides (Fig. S9–S11). AHz functionalisation was 100% (116 units) for l-PAHz1, while higher amounts of AHz were noted for l-PAHz2 (57 units, 49%) and l-PAHz3 (29 units, 25%), likely due to the higher reactivity of hydrazine monohydrate in contrast to ethanolamine, both used at 8-fold excess per BLG group. Similar characteristics were observed for s-PAHz1-3, with DF of 100% (110 units), 50% (55 units), and 25% (28 units) confirmed by 1H NMR analysis (Fig. S12–S14). A summary of data for the modifications with BzAld and AHz groups can be found in Table S1.3.2. Determination of polypeptide secondary structure
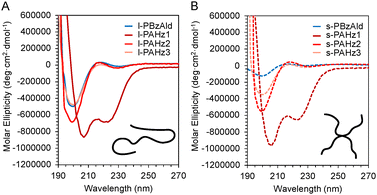
Circular dichroism (CD) spectroscopy was used to understand impact of side chain residues and topology on polypeptide secondary structure (Fig. 2 and Table S2). The online database program BestSel was used to calculate secondary structure content from experimental CD spectra.44 Trends were observed for side chain modifications (BzAld and AHz) of the linear and star polypeptides. For l- and s-PBzAld copolypeptides, 0% α-helical content was observed for both, while 34% and 28% β-sheets, 23% and 21% turns, and 43% and 51% random conformations were observed respectively. In contrast, significant α-helical content of 83% and 76% was noted for the l- and s-PAHz1 homopolypeptides, with 6% β-sheet motifs observed for s-PAHz, along with 17% and 18% random conformations respectively. Reducing acyl hydrazide groups and increasing hydroxyl ethyl for the linear copolypeptides resulted in only 1% α-helical character for l-PAHz2 and l-PAHz3, which also featured ∼38% and 30% β-sheets, 20% and 22% turns, with both having 41% random motifs respectively. Similar profiles were noted for star copolypeptides s-PAHz2 and s-PAHz3, with 0% α-helices observed, 37% and 27% β-sheet character, 22% and 24% turns, and 41% and 49% random conformations. In summary, secondary structures were under more influence of side group chemistry rather than polymer topology.3.3. Evaluation of hydrogel crosslinking kinetics
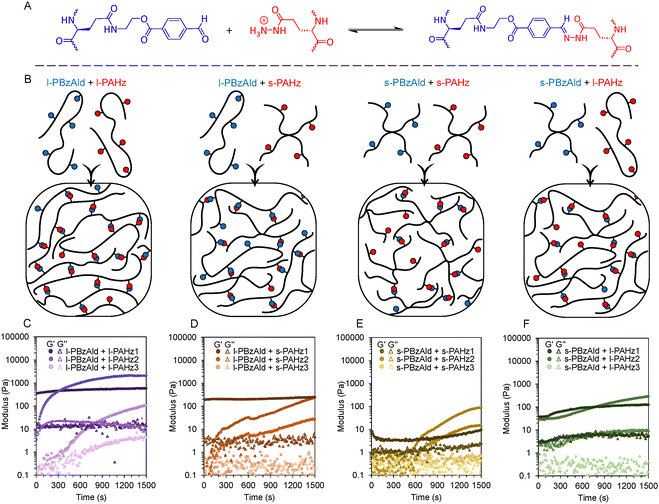
Polypeptides with pendant benzaldehyde (BzAld) and acyl hydrazide (AHz) groups were then mixed at room temperature to induce a Schiff-base reaction (Fig. 3A and Video S1), crosslinking them into hydrogels through a dynamic covalent acyl hydrazone bond. Complimentary PBzAld and PAHz polypeptide crosslinkers, fixed at 5.0 wt% respectively using phosphate buffered saline (PBS) were screened for Schiff-base reactivity based on polymer topology, to identify the influence of architecture.The library was screened through mixing homo-topological blends of linear PBzAld and PAHz, star PBzAld and PAHz, and hetero-topological blends of linear PBzAld and star PAHz or linear PAHz and star PBzAld (Fig. 3B). Rheological analysis was conducted in order to identify crosslinking kinetics for hydrogel formation through the Schiff-base reaction at room temperature with time sweeps presented over a 1500 s interval (25 min) to show key differences (Fig. 3C-F). Polypeptides were readily dissolved in phosphate buffer saline (PBS) solution, facilitating crosslinking to form ionically stable hydrogels. For linear homo-topological blends (Fig. 3C), regimes varied according to amount of AHz groups, while BzAld content remained constant. For example, l-PBzAld crosslinked almost instantaneously with l-PAHz1, signified by an immediate gel point due to a non-detectable crossover of storage modulus (G′) and loss modulus (G″), whereby G′ plateaued to 530 Pa, 260 s after mixing. Slower Schiff-base kinetics were observed with l-PBzAld and l-PAHz2, whereby a gel point was observed at 30 s, with G′ plateauing to 2030 Pa after 1200 s. The reaction of l-PBzAld with l-PAHz3 (the least AHz groups) was even more sluggish, having a gel point at 360 s, and taking 1700 s to plateau to a G′ of 110 Pa.
Mixing l-PBzAld as a hetero-topological blend with star polypeptides with AHz groups resulted in weaker gels, and slower crosslinking kinetics (Fig. 3D). For example, a similar non-detectable gel point was observed when blending with s-PAHz1, with G′ plateauing to 240 Pa after 200 s. After mixing with s-PAHz2, a gel point was noted after 100 s, while a G′ of 460 Pa was determined after a plateau at 1800 s. For s-PAHz3, it required 2980 s after mixing for a gel point, almost plateauing at 5500 s providing a G′ of 260 Pa (Fig. S15).
Similar to the linear analogues, rheological regimes of star homo-topological polypeptide blends varied (Fig. 3E). Instantaneous crosslinking of s-PBzAld and s-PAHz1 was noted, with displaying a 10 Pa G′ plateau after 100 s. When reacting s-PBzAld with s-PAHz2 with less AHz groups, it took 550 s for a gel point, forming gels with G′ of 90 Pa after 1490 s. Interestingly for s-PAHz3, no gelation was observed. Using s-PBzAld as a star homo-topological polypeptide blend with l-PAHz (1-3), improvements on purely star-star blends were observed (Fig. 3F). Rapid gelation and plateauing at 620 s for a G′ of 120 Pa was observable for s-PBzAld and l-PAHz1. Similarly for s-PBzAld and l-PAHz2, an instant gel point was noted with a plateau at 1730 s providing a hydrogel with 410 Pa strength. Very slow reactivity was noted for s-PBzAld and l-PAHz3, whereby the gel point took 1740 s, stabilising to 60 Pa after 2340 s. Ultimately, purely linear homo-topological blends provided fastest Schiff-base reactivity and best gel strengths (G′) for hydrogel formation, while decreasing the amount of AHz reduced reaction kinetics, as summarised for the screened library of polypeptide blends in Table S3.
3.4. Hydrogel dynamic properties and biocompatibility
The blend of l-PBzAld and l-PAHz2 was identified as a lead candidate to further analyse its rheological behaviour and biocompatibility. The molar ratio of BzAld/AHz was varied between 0.33, 0.25, and 0.16, in order to identify the influence of stoichiometry on dynamic properties. A molar ratio of 1 was avoided in order to demonstrate dynamic behaviour, as benzaldehyde-based Schiff base bonds have been shown to form static matrices at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
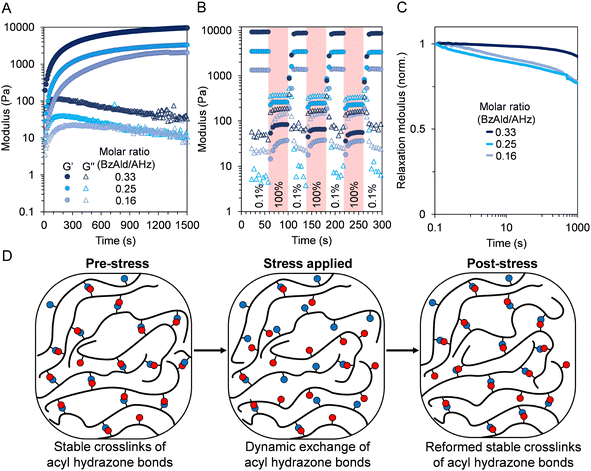
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 benzaldehyde/acyl hydrazide ratio.37,45 Maintaining excess of acyl hydrazide was therefore anticipated to improve dynamic properties such as self-recovery and stress relaxation. For all tested ratios, similar timeframes were noted for full crosslinking (∼1200 s), with a decrease in G′ moduli (9240, 3200, and 2300 Pa) observed with decreasing molar ratio (Fig. 4A). As a control experiment, the hydrogel media was changed from PBS to Dulbecco's modified eagle medium (DMEM), with gelation time and G′ both impacted (Fig. S16). Full crosslinking (plateaued G′) was only observed after 6700 s, while G′ reduced to 1170 Pa, likely due to the presence of glucose (a reducing sugar) in the media, which could form Schiff-base bonds with pendant acyl hydrazides.46 Aside from this, all hydrogels could also demonstrate remarkable self-recovery behaviour, first deforming and then almost instantaneously reforming their network upon cycling between high (100%) and low (0.1%) strain (Fig. 4B). The dynamic nature of the formed acyl hydrazone crosslinks was also probed by stress relaxation experiments (Fig. 4C). For hydrogels with a higher molar ratio of BzAld to AHz (0.33), a longer stress relaxation halftime was observed, in line with the higher density of acyl hydrazone crosslinks. Decreasing the molar ratio to 0.25 and 0.16 resulted in a faster relaxation accordingly, consistent with less dynamic bonds (Fig. 4D).
1 benzaldehyde/acyl hydrazide ratio.37,45 Maintaining excess of acyl hydrazide was therefore anticipated to improve dynamic properties such as self-recovery and stress relaxation. For all tested ratios, similar timeframes were noted for full crosslinking (∼1200 s), with a decrease in G′ moduli (9240, 3200, and 2300 Pa) observed with decreasing molar ratio (Fig. 4A). As a control experiment, the hydrogel media was changed from PBS to Dulbecco's modified eagle medium (DMEM), with gelation time and G′ both impacted (Fig. S16). Full crosslinking (plateaued G′) was only observed after 6700 s, while G′ reduced to 1170 Pa, likely due to the presence of glucose (a reducing sugar) in the media, which could form Schiff-base bonds with pendant acyl hydrazides.46 Aside from this, all hydrogels could also demonstrate remarkable self-recovery behaviour, first deforming and then almost instantaneously reforming their network upon cycling between high (100%) and low (0.1%) strain (Fig. 4B). The dynamic nature of the formed acyl hydrazone crosslinks was also probed by stress relaxation experiments (Fig. 4C). For hydrogels with a higher molar ratio of BzAld to AHz (0.33), a longer stress relaxation halftime was observed, in line with the higher density of acyl hydrazone crosslinks. Decreasing the molar ratio to 0.25 and 0.16 resulted in a faster relaxation accordingly, consistent with less dynamic bonds (Fig. 4D).
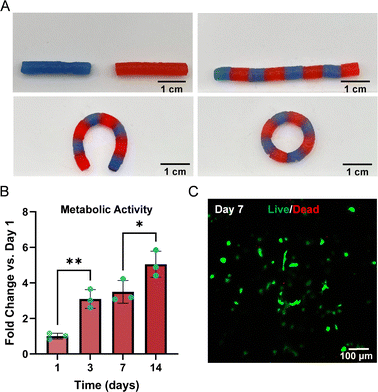
Further analysis of the l-PBzAld + l-PAHz2 (0.33 molar ratio) gel properties included a high gel fraction (G) of 87.6 ± 4.2%, and a relatively low swelling ratio (Q) of 7.9 ± 0.6 (Table S4). The dynamic bond relaxation and self-healing behaviour was also captured macroscopically (Fig. 5A). Remarkable self-healing was observed in rapid timeframes (30 min) after cutting tubular hydrogel structures, and placing them in direct contact after to reform the Schiff-base bonds, without the need for temperature or additives. The reformed hydrogels could also be readily manipulated into ring conformations, denoting the true dynamic yet robust nature of these Schiff-base polypeptide hydrogels. Subsequently, hydrogel biocompatibility was assessed through resazurin-based metabolic activity and live/dead viability assays, whereby hMSCs were encapsulated in l-PBzAld/l-PAHz2 hydrogel, with a BzAld/AHz molar ratio of 0.33. The metabolic activity (Fig. 5B) of encapsulated cells increased significantly at day 3 compared to day 1 (3.1-fold the initial activity), remaining constant from day 3 to day 7 (no significant change) and seeing yet another significant increase on day 14 (5.05-fold the initial activity). Cells viability was also observed in live/dead assay (Fig. 5C) at day 7, where 90.97% of imaged cells were flagged as alive.
4. Conclusions
In summary, a series of benzaldehyde modified and acyl hydrazide modified polypeptides were used to form robust dynamic hydrogels, with highly tuneable properties. Side chain modification of polypeptides had direct influence on secondary structure, while changes in polymer topology had no significant effect, as evidenced by CD spectroscopy. Hydrogel properties were readily modulated by blending homo- (e.g., linear/linear) and hetero- (e.g., linear/star) topological mixtures, providing insight into how architectural design can be leveraged to modulate hydrogel mechanical regimes. Biocompatibility results indicate a promising capability of the presented material to not only support, but also stimulate cellular bioactivity, as encapsulated cells remained viable and with increasing metabolic activity even after 14 days in culture.Author contributions
The project idea was conceived by R. D. M. The original manuscript and figures were drafted by R. D. M. Experiments were designed by R. D. M., F. L., V. C., J. M. G., T. H., and A. H., and performed by R. D. M., F. L., V. C., and J. M. G. All authors have provided input and given approval to the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
The data generated from the current study is not publicly available but can be provided by the authors upon reasonable request.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ma01468a.
Acknowledgements
J. M. G. acknowledges financial support from the European Union's Horizon Europe programme under grant agreement No 101203568. V. C. acknowledges financial support of Taighde Éireann-Research Ireland under the EPSRC-Research Ireland Joint Funding of Research (grant number 24/EPSRC/4005).Notes and references
- C. D. Spicer, Polym. Chem., 2020, 11, 184–219 RSC.
- G. Zhu, N. Javanmardia, L. Qian, F. Jin, T. Li, S. Zhang, Y. He, Y. Wang, X. Xu, T. Wang and Z. Q. Feng, Int. J. Biol. Macromol., 2024, 281, 136115 CrossRef CAS PubMed.
- B. G. Soliman, A. K. Nguyen, J. J. Gooding and K. A. Kilian, Adv. Mater., 2024, 36, 1–24 CrossRef PubMed.
- P. Sikdar, M. M. Uddin, T. M. Dip, S. Islam, M. S. Hoque, A. K. Dhar and S. Wu, Mater. Adv., 2021, 2, 4532–4573 RSC.
- Y. Gao, K. Peng and S. Mitragotri, Adv. Mater., 2021, 33, 2006362 CrossRef CAS PubMed.
- M. R. Arkenberg, H. D. Nguyen and C. C. Lin, J. Mater. Chem. B, 2020, 8, 7835–7855 RSC.
- M. M. Perera and N. Ayres, Polym. Chem., 2020, 11, 1410–1423 RSC.
- Y. Chen, W. Qian, R. Chen, H. Zhang, X. Li, D. Shi, W. Dong, M. Chen and Y. Zhao, ACS Macro Lett., 2017, 6, 1129–1133 CrossRef CAS PubMed.
- H. Kang, W. Wei, L. Sun, R. Yu, E. Yang, X. Wu and H. Dai, Chem. Mater., 2023, 35, 2408–2420 CrossRef CAS.
- G. M. Scheutz, J. L. Rowell, S. T. Ellison, J. B. Garrison, T. E. Angelini and B. S. Sumerlin, Macromolecules, 2020, 53, 4038–4046 CrossRef CAS.
- B. R. Nelson, B. E. Kirkpatrick, C. E. Miksch, M. D. Davidson, N. P. Skillin, G. K. Hach, A. Khang, S. N. Hummel, B. D. Fairbanks, J. A. Burdick, C. N. Bowman and K. S. Anseth, Adv. Mater., 2024, 36, 1–15 CrossRef PubMed.
- S. Kirchhof, A. Strasser, H. J. Wittmann, V. Messmann, N. Hammer, A. M. Goepferich and F. P. Brandl, J. Mater. Chem. B, 2015, 3, 449–457 RSC.
- C. M. Madl and S. C. Heilshorn, Chem. Mater., 2019, 31, 8035–8043 CrossRef CAS PubMed.
- J. Liu, X. Zhang, X. Chen, L. Qu, L. Zhang, W. Li and A. Zhang, Polym. Chem., 2018, 9, 378–387 RSC.
- F. Y. Lin, N. H. Dimmitt, M. Moraes de Lima Perini, J. Li and C. C. Lin, Adv. Healthcare Mater., 2022, 11, 1–11 Search PubMed.
- J. Xu, Y. Liu and S. Hsu, Molecules, 2019, 24, 1–21 Search PubMed.
- F. L. C. Morgan, I. A. O. Beeren, L. Moroni and M. B. Baker, Chem. Mater., 2025, 37, 2709–2719 CrossRef CAS PubMed.
- Z. Zhang, C. He and X. Chen, Mater. Chem. Front., 2018, 2, 1765–1778 RSC.
- S. Li, M. Pei, T. Wan, H. Yang, S. Gu, Y. Tao, X. Liu, Y. Zhou, W. Xu and P. Xiao, Carbohydr. Polym., 2020, 250, 116922 CrossRef CAS PubMed.
- K. Sahajpal, S. Shekhar, A. Kumar, B. Sharma, M. K. Meena, A. K. Bhagi and S. Sharma, J. Mater. Chem. B, 2022, 10, 3173–3198 RSC.
- Q. Li, Y. Lan, W. Wang, G. Ji, X. Li and Z. Song, Macromolecules, 2023, 56, 7023–7031 CrossRef CAS.
- Y. Wu, K. Chen, J. Wang, M. Chen, W. Dai and R. Liu, J. Am. Chem. Soc., 2024, 146, 24189–24208 CrossRef CAS PubMed.
- S. Stefanovic, K. McCormick, S. Fattah, R. Brannigan, S. A. Cryan and A. Heise, Polym. Chem., 2023, 14, 3151–3159 RSC.
- E. Tinajero-Díaz, N. Judge, B. Li, T. Leigh, R. D. Murphy, P. D. Topham, M. J. Derry and A. Heise, ACS Macro Lett., 2024, 13, 1031–1036 CrossRef PubMed.
- A. P. Nowak, V. Breedveld, L. Pakstis, B. Ozbas, D. J. Pine and T. J. Deming, Nature, 2002, 417, 424–428 CrossRef CAS PubMed.
- R. Murphy, D. P. Walsh, C. A. Hamilton, S. A. Cryan, M. I. Het Panhuis and A. Heise, Biomacromolecules, 2018, 19, 2691–2699 CrossRef CAS PubMed.
- R. D. Murphy, S. Kimmins, A. J. Hibbitts and A. Heise, Polym. Chem., 2019, 10, 4675–4682 RSC.
- R. D. Murphy, R. V. Garcia, S. J. Oh, T. J. Wood, K. D. Jo, J. Read de Alaniz, E. Perkins and C. J. Hawker, Adv. Mater., 2023, 35, 2207542 CrossRef CAS PubMed.
- R. D. Murphy, C. Delaney, S. Kolagatla, L. Florea, C. J. Hawker and A. Heise, Adv. Funct. Mater., 2023, 33, 1–9 CrossRef.
- R. V. Garcia, E. A. Murphy, N. J. Sinha, Y. Okayama, J. M. Urueña, M. E. Helgeson, C. M. Bates, C. J. Hawker, R. D. Murphy and J. Read de Alaniz, Small, 2023, 19, 1–8 CrossRef PubMed.
- A. F. Roca-Arroyo, J. A. Gutierrez-Rivera, L. D. Morton and D. A. Castilla-Casadiego, Gels, 2025, 11, 1–52 CrossRef PubMed.
- T. J. Deming, Chem. Rev., 2016, 116, 786–808 CrossRef CAS PubMed.
- S. Yan, X. Zhang, K. Zhang, H. Di, L. Feng, G. Li, J. Fang, L. Cui, X. Chen and J. Yin, J. Mater. Chem. B, 2016, 4, 947–961 RSC.
- S. Li, D. Niu, T. Shi, W. Yun, S. Yan, G. Xu and J. Yin, ACS Biomater. Sci. Eng., 2023, 9, 2625–2635 CrossRef CAS PubMed.
- A. Rasines Mazo, S. Allison-Logan, F. Karimi, N. J.-A. Chan, W. Qiu, W. Duan, N. M. O’Brien-Simpson and G. G. Qiao, Chem. Soc. Rev., 2020, 49, 4737–4834 RSC.
- T. Behroozi Kohlan, Y. Wen, C. Mini and A. Finne-Wistrand, Carbohydr. Polym., 2024, 338, 122173 CrossRef CAS PubMed.
- S. M. Hull, J. Lou, C. D. Lindsay, R. S. Navarro, B. Cai, L. G. Brunel, A. D. Westerfield, Y. Xia and S. C. Heilshorn, Sci. Adv., 2023, 9, 1–14 Search PubMed.
- J. M. Ren, T. G. McKenzie, Q. Fu, E. H. H. Wong, J. Xu, Z. An, S. Shanmugam, T. P. Davis, C. Boyer and G. G. Qiao, Chem. Rev., 2016, 116, 6743–6836 CrossRef CAS PubMed.
- D. E. Apostolides, T. Sakai and C. S. Patrickios, Macromolecules, 2017, 50, 2155–2164 CrossRef CAS.
- F. Jia, J. M. Kubiak, M. Onoda, Y. Wang and R. J. Macfarlane, Adv. Sci., 2021, 8, 1–8 Search PubMed.
- Y. H. Lin, J. Lou, Y. Xia and O. Chaudhuri, Adv. Healthcare Mater., 2024, 13, 1–12 Search PubMed.
- M. Byrne, P. D. Thornton, S. A. Cryan and A. Heise, Polym. Chem., 2012, 3, 2825–2831 RSC.
- D. Hlavatá, H. Pivcová, J. Spěváček, D. Nosková and F. Rypáček, Collect. Czech. Chem. Commun., 1993, 58, 2415–2427 CrossRef.
- A. Micsonai, F. Wien, L. Kernya, Y. H. Lee, Y. Goto, M. Réfrégiers and J. Kardos, Proc. Natl. Acad. Sci. U. S. A., 2015, 112, E3095–E3103 CrossRef CAS PubMed.
- N. de Paiva Narciso, R. S. Navarro, A. E. Gilchrist, M. L. M. Trigo, G. Aviles Rodriguez and S. C. Heilshorn, Adv. Healthcare Mater., 2023, 12, 1–14 Search PubMed.
- A. B. Uceda, L. Mariño, R. Casasnovas and M. Adrover, Biophys. Rev., 2024, 16, 189–218 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |