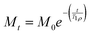Elucidating molecular level interfacial interactions between a de novo protein and nucleated calcite with solid-state NMR
Emily G. S.
Close
 *a,
Andrew S.
Lipton
*a,
Andrew S.
Lipton
 a,
Marlo
Zorman
a,
Marlo
Zorman
 bg,
Harley
Pyles
bg,
Harley
Pyles
 cf,
Thi Kim
Hoang Trinh
cf,
Thi Kim
Hoang Trinh
 a,
Garry W.
Buchko
a,
Garry W.
Buchko
 ad,
Ying
Chen
ad,
Ying
Chen
 a,
Chun-Long
Chen
a,
Chun-Long
Chen
 ab,
Christopher J.
Mundy
ab,
Christopher J.
Mundy
 ab,
David
Baker
ab,
David
Baker
 cef and
Wendy J.
Shaw
cef and
Wendy J.
Shaw
 *a
*a
aPacific Northwest National Laboratory, Richland, WA, USA. E-mail: emily.close@pnnl.gov; Wendy.shaw@pnnl.gov
bDepartment of Chemical Engineering, University of Washington, Seattle, WA, USA
cInstitute for Protein Design, University of Washington, Seattle, WA, USA
dSchool of Molecular Biosciences, Washington State University, Pullman, WA, USA
eHoward Hughes Medical Institute, University of Washington, Seattle, WA, USA
fDepartment of Biochemistry, University of Washington, Seattle, WA, USA
gMolecular Engineering & Sciences Institute, University of Washington, Seattle, WA, USA
First published on 12th March 2026
Abstract
Biomineralization is the process by which organisms use biomolecules to produce hierarchically structured organic–inorganic composites. Using biology as inspiration, a protein construct (FD31) was previously designed to accelerate formation of nano-calcite with an unconventional {110} face. To understand the molecular interactions essential for protein aided calcite nucleation, solid-state nuclear magnetic resonance (ssNMR) spectroscopy was used in this work to characterize the FD31–calcite interface at the atomic level. Glutamic acid side chains designed to interact directly with calcium ions on the surface were found to have dynamics on the sub-millisecond timescale, indicating possible interactions between the protein and surface waters that were not included in the original model. Dipolar ssNMR recoupling techniques also showed that the protein backbone is ∼2 Å closer to the surface than in the original docking model. Refined molecular simulations were done in the presence of explicit waters, which resulted in the protein backbone closer to the surface than in the original docking structure, providing better agreement with experiment and highlighting the important role played by water in FD31–calcite interactions. While this work provides the first experimental confirmation that FD31 interactions with calcite are localized to the surface of the protein designed to serve as a template, these studies do indicate a more dynamic binding and closer binding mode between FD31 and the nucleated surface than originally proposed. In all, this enhanced molecular insight into the FD31–calcite interface has advanced our fundamental understanding of the atomic interactions at the organic–inorganic interface and will aid in the design of biological templates for the nucleation of inorganic crystals.
Introduction
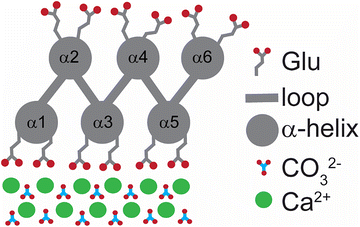
Biominerals are composite materials containing biomolecules and inorganic crystals, which are produced and used by organisms to help stiffen and strengthen tissue.1,2 The most prevalent family of biominerals contain carbonates,3,4 which are found in the shells, skeletons, and exoskeletons of freshwater, terrestrial, and marine organisms.3,5,6 While calcium carbonate exists in the three primary crystalline polymorphs of vaterite, aragonite, or calcite, the latter is the most thermodynamically stable and thus the most prevalent.7 To control the nucleation, growth, and assembly of crystalline calcium carbonate, organisms rely on biomacromolecules such as proteins.8,9 While it is known that the acidic functional groups of aspartate and glutamate play an important role in the formation of calcium carbonate biominerals,10–14 the structural details at the organic–inorganic interfaces involved in these processes are largely unknown. This incomplete molecular understanding has made it difficult to efficiently design proteins and other polymers with similar capabilities of templating biomimetic materials.In theory, a protein designed with carboxylate residues positioned to electrostatically and structurally match the geometric lattice of calcium carbonate crystals should be able to control the nucleation and growth of calcium carbonate mineralization.15 This design strategy is inspired by the mechanism employed by ice-nucleating proteins, which contain surfaces capable of epitaxial-like lattice matching of the charged residues to the ice lattice, aiding in the formation of ice via preorganization of ice-like water molecules.16 To generate a protein capable of templating calcium carbonate crystals, Baker and coworkers designed a series of de novo helical repeat proteins with flat surfaces containing carboxylic acid functional groups lattice matched to calcium carbonate nuclei. This study resulted in the identification of a 19 kDa protein, FD31 (Fig. 1), which was capable of lowering the free energy of nano-calcite formation by bypassing formation of an intermediate vaterite phase.17
While a protein construct capable of templating nano-calcite was identified, their process required screening of numerous potential protein designs.17 An experimentally validated molecular understanding of what made this protein an effective template is still lacking. Solid-state NMR is well suited for characterizing organic–inorganic interfaces due to its ability to provide residue-specific structural18–22 and dynamic18,21,23 information, as well as measure distances between NMR active nuclei.24–26 To date, a number of ssNMR studies have been used to characterize interactions between biomolecules and inorganic crystals,27,28 revealing insights into the protein residues involved in surface interactions25,29,30 and the structure and orientation of proteins on the surface.30 In this study, we utilized ssNMR dynamics and distance-dependent experiments, complemented by molecular dynamics simulations, to explore the molecular intricacies of the FD31–calcite interface. (Fig. 1).
Experimental
Unless otherwise noted, all chemicals were purchased from Sigma and used without any additional modification.FD31 and FD31* expression
Genes encoding the original FD31 sequence and a variant with 25 glutamic acid residues substituted with aspartic acid residues (FD31*) were ordered from Genscript (Piscataway, NJ). Both constructs containing an N-terminal His6-tag followed by a Tobacco Etch Virus (TEV) cleavage site were cloned into pET-28b+ vectors between Ndel and Xhol restriction enzyme sites. A flexible linker, SWSGGSGG, containing one additional tryptophan residue was added to each gene construct to increase the UV absorbance of the resulting protein at 280 nm. Both cloned genes were transformed into BLR(DE3) Escherichia coli (Novagen, Rahway, NJ) on Listeria Broth (LB) agar plates (Research Products International (RPI), Mount Prospect, Il) supplemented with 50 µg mL−1 kanamycin (RPI). To make glycerol stocks, a colony from each plate was added to 5 mL sterile LB supplemented with 50 µg mL−1 of kanamycin and grown at 37 °C/180 rpm to an OD600 between 0.8 and 1. The cells were then mixed with 5 mL of filtered 50% glycerol, shaken at 180 rpm for 2 minutes, divided into 1 mL aliquots in sterile 1.7 mL microcentrifuge tubes (Eppendorf, Hamburg, Germany), flash frozen with liquid nitrogen, and stored at −80 °C.Protein expression in rich media was done in 500 mL sterile ZY media (50 mM tryptone (RPI) and 12 mM yeast extract (RPI)). To prepare a starter culture, a 1 mL glycerol stock was thawed and added to 25 mL sterile LB supplemented with 50 µg mL−1 of kanamycin and grown at 37 °C/180 rpm to an OD600 ∼0.8. To express FD31 or FD31* via autoinduction, 500 mL of pre-warmed ZY media was inoculated with 10 mL sterile 50× 5052 (0.5% (v/v) glycerol, 128.8 mM glucose, 292.2 mM α-lactose), 25 mL starter culture, 0.5 mL sterile 1000× FeCl3, 25 mL sterile 20× NPS (1 M anhydrous sodium phosphate dibasic, 1 M anhydrous potassium phosphate monobasic, 0.5 M ammonium sulfate) and 1 mL of sterile 1 M MgSO4. The inoculated media was left at 37 °C/180 rpm for ∼20 h, and the next morning the cells were pelleted at 3500×g for 15 minutes and stored at −80 °C until needed.
For FD31, two samples were prepared with residue-specific 13C- and 15N-labels; uniformly labeled threonine (FD31-U-T) and uniformly labeled glutamic acid (FD31-U-E), in which all carbon and nitrogen residues in threonine or glutamic acid were 13C and 15N labeled (Table S1). For FD31*, two samples were prepared with residue-specific 13C-labeled amino acids; backbone carbonyl labeled glutamic acid (FD31*-E’) and side chain carbonyl labeled glutamic acid (FD31*-Eδ) and one sample was prepared with residue specific 15N-labeled glutamic acid (FD31*-EN) (Table S1). For selectively labeled protein expression, a modified Redfield-medium was used containing 3.7 mM adenine, 2.3 mM guanosine, 1.6 mM thymine, 4.5 mM uracil, 1.8 mM cytosine, 18.3 mM anhydrous sodium acetate, 12.7 mM succinic acid, 14.0 mM ammonium chloride, 21.3 mM NaOH, 60.3 mM anhydrous K2HPO4, 14 µM CaCl2·2H2O, 7 µM ZnSO4·7H2O, 12 µM MnSO4·H2O, 170 µM thiamine, 41 µM niacin, 2 µM biotin, 10 µM kanamycin, 111 mM glucose, 4.0 mM MgSO4, and 10 µM FeCl3. The medium also contained 19 of the 20 common amino acids except asparagine, as follows: 5.6 mM Ala, 2.3 mM Arg, 3.0 mM Asp, 0.4 mM Cys, 2.7 mM Gln, 4.4 mM Glu, 29.2 mM Gly, 0.6 mM His, 1.7 mM Ile, 1.7 mM Leu, 2.9 mM Lys, 1.7 mM Met, 0.8 mM Phe, 0.9 mM Pro, 20.0 mM Ser, 1.9 mM Thr, 0.9 mM Tyr, 2.0 mM Val, and 0.2 mM Trp.31–34 To minimize “scrambling” of the targeted amino acid, only one-quarter the amount of unlabeled target amino acid was added at the start of culture growth. A 25 mL starter culture was prepared as described above and used to inoculate 750 mL of pre-warmed modified Redfield-medium. The cells were grown to OD600 1.0 at 37 °C/180 rpm and then the 13C- or 15N-labeled target amino acids were added. Only one half the amount from the original recipe was added of labeled Glu (14.6 mM) or Thr (0.4 mM). Ten minutes after addition of the labeled amino acid, protein expression was induced via addition of 1 mM isopropyl-β-D-1-thiogalactopyranoside. After 30 min (Glu) or 60 min (Thr) of induction, the cells were pelleted at 3500×g for 15 minutes and stored at −80 °C until needed.
FD31 and FD31* purification
A frozen FD31 or FD31* cell pellet was thawed in ∼25 mL of lysis buffer (20 mM Tris, 500 mM NaCl, 30 mM imidazole, 5% (v/v) glycerol, 1 mM PMSF, pH 8) and lysed via sonication at 40% amplitude on a Q500 Sonicator (Fisher Scientific, Hampton, NH) for 10 minutes (30 sec on, 30 sec off). The lysate was clarified via centrifugation at 19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800×g for 1 h at 4 °C, and the supernatant run over a 5 mL prepacked HisTrap column (Cytiva, Marlborough, MA) loaded with wash buffer (20 mM Tris, 500 mM NaCl, 30 mM imidazole, 5% (v/v) glycerol, pH 8). His-tagged protein was eluted from the column using elution buffer (20 mM Tris, 500 mM NaCl, 500 mM imidazole, 5% (v/v) glycerol, pH 8) and then buffer exchanged into TEV cleavage buffer (50 mM Tris, 50 mM NaCl, pH 8) using 10 kDa molecular weight cutoff (MWCO) Amicon Ultra Centrifugal Filters (Sigma, St. Louis, MO). To exchange buffer, the protein was concentrated down to ∼1 mL in the Amicon, diluted with ∼15 mL of cleavage buffer, and then spun down at 3500×g until the final volume remaining in the filter was ∼1 mL. This was repeated four times. The protein concentration was estimated using its calculated molar extinction coefficient at 280 nm (ProtParam (expasy.org/protparam/) (εpre-cleavage = 6990 M−1 cm−1, εpost-cleavage = 5500 M−1 cm−1)) using 2 µL aliquots on a Thermo Scientific NanoDrop One (Waltham, MA). Cleavage was performed at 25 °C overnight using 1 mg of TEV protease (prepared in-house) for every 25 mg of FD31/FD31*. A second HisTrap chromatography step was run to isolate cleaved FD31 from His6-tagged TEV protease and unreacted His-tagged protein. The flowthrough containing cleaved FD31 was concentrated using 10 kDa MWCO Amicon Ultra Centrifugation devices and further purified via size exclusion chromatography on a HiLoad 26/600 Superdex 75 column (Cytiva, Marlborough, MA) in SEC buffer (20 mM Tris, 100 mM NaCl, pH 8). The purified proteins were then exchanged into MOPS buffer (10
800×g for 1 h at 4 °C, and the supernatant run over a 5 mL prepacked HisTrap column (Cytiva, Marlborough, MA) loaded with wash buffer (20 mM Tris, 500 mM NaCl, 30 mM imidazole, 5% (v/v) glycerol, pH 8). His-tagged protein was eluted from the column using elution buffer (20 mM Tris, 500 mM NaCl, 500 mM imidazole, 5% (v/v) glycerol, pH 8) and then buffer exchanged into TEV cleavage buffer (50 mM Tris, 50 mM NaCl, pH 8) using 10 kDa molecular weight cutoff (MWCO) Amicon Ultra Centrifugal Filters (Sigma, St. Louis, MO). To exchange buffer, the protein was concentrated down to ∼1 mL in the Amicon, diluted with ∼15 mL of cleavage buffer, and then spun down at 3500×g until the final volume remaining in the filter was ∼1 mL. This was repeated four times. The protein concentration was estimated using its calculated molar extinction coefficient at 280 nm (ProtParam (expasy.org/protparam/) (εpre-cleavage = 6990 M−1 cm−1, εpost-cleavage = 5500 M−1 cm−1)) using 2 µL aliquots on a Thermo Scientific NanoDrop One (Waltham, MA). Cleavage was performed at 25 °C overnight using 1 mg of TEV protease (prepared in-house) for every 25 mg of FD31/FD31*. A second HisTrap chromatography step was run to isolate cleaved FD31 from His6-tagged TEV protease and unreacted His-tagged protein. The flowthrough containing cleaved FD31 was concentrated using 10 kDa MWCO Amicon Ultra Centrifugation devices and further purified via size exclusion chromatography on a HiLoad 26/600 Superdex 75 column (Cytiva, Marlborough, MA) in SEC buffer (20 mM Tris, 100 mM NaCl, pH 8). The purified proteins were then exchanged into MOPS buffer (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) mM, pH 7) via four rounds of buffer exchange in 10 kDa MWCO Amicon filters, lyophilized, and stored at −20 °C until needed.
mM, pH 7) via four rounds of buffer exchange in 10 kDa MWCO Amicon filters, lyophilized, and stored at −20 °C until needed.
FD31 and FD31* nucleated CaCO3 crystallization
All nucleation conditions are based on those used in the original work where FD31 was designed and validated as a template of calcium carbonate, but have been scaled up 10-fold to yield sufficient sample for ssNMR analysis. A 20 mL solution of 100 mM CaCl2 and 2 µM FD31 or FD31* was prepared by adding 0.294 g calcium chloride dihydrate and 7.6 mg protein to 20 mL MilliQ water in a 50 mL falcon tube. A second 20 mL solution of 100 mM NaHCO3 was prepared by adding 0.168 g sodium bicarbonate to a 50 mL falcon tube with 20 mL MilliQ water. The sodium bicarbonate solution was added to the calcium chloride and protein solution and stirred on a magnetic stir plate overnight at 25 °C. As the pH of this solution is always between 7–8, the glutamic acid residues in FD31 are expected to be fully deprotonated during nucleation. The next morning the FD31 nucleated calcite solution was spun down at 3500×g for 45 minutes and the supernatant removed. For nucleated calcite samples prepared for NMR, the samples were then resuspended in ∼20 µL of supernatant and centrifuged into 5 mm zirconia rotors. For nucleated calcite samples prepared for SEM, the samples were dried at 80 °C for ∼4 h and stored at room temperature until needed.Solid-state NMR (ssNMR) experiments
Cross Polarization Magic Angle Spinning (CP-MAS), T1ρ, and Rotational Echo DOuble Resonance (REDOR) ssNMR experiments were done on a spectrometer operating at a 1H resonance frequency of 300 MHz. A three-channel Agilent VNMRS console was used with either a two-channel (T1ρ, CP-MAS) or three-channel (REDOR) 5 mm VT Chemagnetics (Otzuka Electronics, Osaka, Japan) probe with ceramic housing. Unless otherwise noted, all ssNMR data was collected using a 3.5 µs 90°-pulse for proton with a 1 msec contact time for CP,35,36 a recycle delay of 5–6 sec, and 72 kHz of SPINAL-64 proton decoupling.37 For direct pulse (DP) experiments on 13C, a recycle delay of 10 sec was used. To account for transverse (T2) relaxation, the final REDOR data were plotted as the ratio of the rephased signal (S) over the reference signal (S0) as a function of the rephasing period. The rephased signal (S) was collected with a recoupling period for both the 15N and 13C channels, while the reference signal (S0) was collected with a recoupling period only for 15N. Dividing S by S0 effectively cancels out the exponential decay factor attributed to T2 relaxation, thus isolating contributions from only heteronuclear dipolar interactions. As these heteronuclear dipolar interactions are directly correlated to the distance between the two nuclei, this information can be extracted to provide a distance between the two nuclei at Å level resolution.38 For the frozen experiments, the spectrometer was equipped with an FTS AirJet (FTS Systems, Stone Ridge, NY). The temperature was verified utilizing ethylene glycol39 or 207Pb NMR of PbNO3,40 with identical experimental conditions (MAS and VT gas flow), and by collecting a one-dimensional 1H spectrum before and after freezing to verify increased linewidths under frozen conditions (Fig. S1).Solution-state NMR experiments
Solution-state NMR data was collected at 20 °C on a Bruker Avance Neo spectrometer operating at a 1H resonance frequency of 600 MHz and equipped with a 5 mm TCI H&F/C/N-D cryogenically cooled probe with Z-axis gradient. Direct pulse 13C experiments were recorded using the standard Bruker pulse sequence zgpg30 with a 5 second recycle delay, 11.1 µs 90° pulse, and 64 scans.Scanning electron microscopy (SEM)
SEM/ESD images were collected on samples placed on silicon wafers using a Thermofisher Apreo Low Vac 2S Scanning Electron Microscope coupled to an Energy Dispersive Spectroscope (EDS) from Oxford Instruments (Abingdon, UK). To prepare the samples, the FD31 nucleated calcite was deposited onto a metal specimen coated by conductive carbon discs. Non-binding calcite was removed using a duster. For composition analysis, the instrument was operated at 20 kV with a step size of 0.2 µm.Circular dichroism (CD)
Circular dichroism data was collected on Aviv Model 410 spectropolarimeter (Lakewood, NJ). Quartz cuvettes with a 0.1 cm (mountable cuvette) path length were used with protein concentrations of ∼0.1 mg mL−1. Steady-state wavelength spectra were collected at pH 7 in 10 mM phosphate buffer. Averaged spectra were converted to mean residue ellipticity θ = (Mres × θobs)/(10 × d × c) where Mres = 112.9, θobs is the observed ellipticity (°) at wavelength λ, d is the pathlength (cm) and c is the protein concentration (g mL−1).Molecular dynamics (MD)
All atom molecular dynamics (MD) simulations were run with GROMACS 2024.3.46 The protein FD31 was docked on the {110} facet of calcite with Rosetta,47 then solvated, neutralized with sodium ions, and minimized. The crystal structure was periodic across the x–y boundaries and had dimensions of ∼10.3 nm × ∼10.4 nm × 3 nm, with a buffer of ∼10 nm of solvent placed above the slab. The system was equilibrated in the NPT ensemble for 500 ps, using a 1 fs timestep, the Berendsen barostat, and semi-isotropic pressure coupling. A production stage of 100 ns was run in the NVT ensemble, and the last 80 ns were used for analysis. A total of three replicas were run. Simulations used the CHARMM36 forcefield in conjunction with SPC/E water models and the calcite forcefield implemented by Shen et al.48–50Results and discussion
Dynamics of FD31 on nucleated calcite
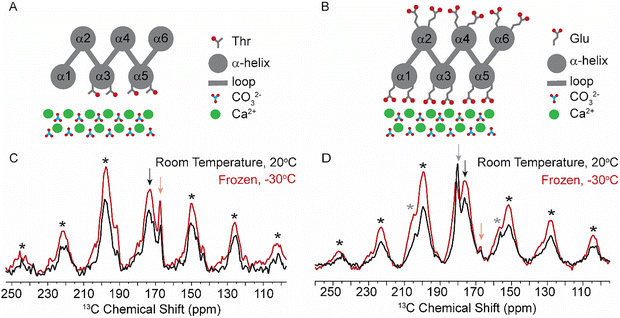
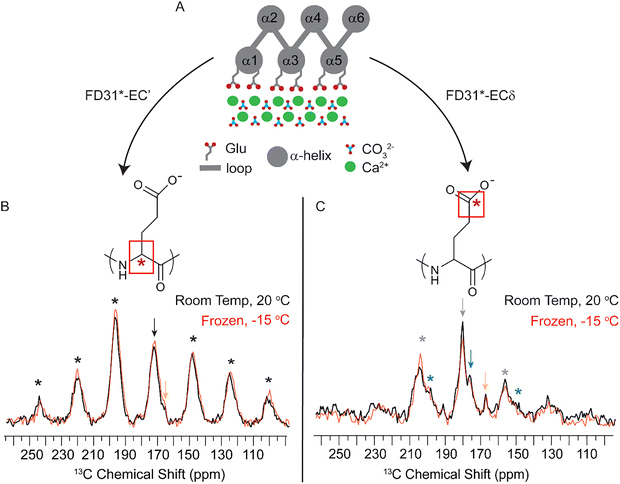
In the de novo structure, three of the six helices in FD31 form a surface of precisely spaced glutamic acid residues designed to lattice match with calcium carbonate nuclei growing in solution, while the opposite surface is composed of three other helices containing both positively and negatively charged surface residues at random spacings that are not designed to bind CaCO3 (Fig. 1 and Fig. S3). To determine if FD31 interacts with the CaCO3 surface as designed, NMR active nuclei (13C, 15N) were incorporated into FD31 at selective sites to enable dynamics measurements of the surface immobilized protein using ssNMR experiments (Table S1). Since protein dynamics are directly correlated to mobility, isotopic labels were incorporated in regions of the protein predicted to interact with the surface, specifically at all threonine (Fig. 2A) or glutamic acid residues (Fig. 2B). Glutamic acid was an obvious choice for labeling as 18 of the 43 glutamic acid side chains were predicted to directly interact with calcium ions on the crystal surfaces (Fig. 2B). However, there is also a second population of 25 glutamic acid residues predicted to be exposed to the aqueous solution (Fig. 2B). If solvent exposed, these glutamic acid residues are likely highly mobile when the protein interacts with calcite and could potentially complicate dynamic measurements. Additionally, glutamic acid is used by E. coli in the biosynthesis of multiple other amino acids, so scrambling of the desired isotopic label could occur.51 For these reasons, threonine was also chosen for selective labeling, as this residue is only present in regions of the protein predicted to be involved in surface interactions (Fig. 2A), and this amino acid is used in the biosynthesis of fewer amino acids in E. coli compared to glutamic acid.52The FD31-U-T and FD31-U-E proteins were each used to produce nucleated calcite samples for 13C ssNMR dynamics experiments. Highly dynamic threonine and/or glutamic acid residues would suggest limited interactions with the surface, whereas minimal to no dynamics would suggest a tight binding interaction with CaCO3. To probe mobility on the sub-millisecond timescale, cross polarization efficiency was assessed by collecting a 13C{1H} Cross Polarization Magic Angle Spinning (CP-MAS) spectrum at the slow spinning speed of 1800 Hz to produce a chemical shift anisotropy (CSA) pattern. At this spinning speed, the rotor period is 555 µs, making the experiment sensitive to motions on the ∼0.5 ms timescale. Spectra were collected under frozen conditions, a control for immobility of all residues, and at room temperature, where mobility would be expected if the protein was not interacting tightly with the surface. In proteins, carbonyls are a good indicator of mobilities on this timescale as they have distinct CSA patterns in mobile versus immobile environments.53 The room temperature and frozen CSA spectra were subjected to a Herzfeld-Berger Analysis (HBA) to obtain asymmetry (η) and span (Ω) parameters. Differences in these values as a function of temperature would signal mobility changes at these residue positions. For a highly mobile carbonyl, η will be close to 0 and Ω will be relatively small. For a mobile carbonyl that is spun at the magic angle the isotropic peak will be the highest in intensity, and there will be small to no spinning sidebands present (Fig. S4, left). On the other hand, for an immobile carbonyl, η is typically ≥ 0.8 and the Ω will be relatively large. For an immobile carbonyl that is spun at the magic angle, the isotropic peak will likely be lower in intensity than at least one corresponding side band, and there are likely to be multiple spinning side bands present due to the large CSA values typical of immobile carbonyl systems (Fig. S4, right).
For FD31-U-T bound to nucleated calcite, the backbone carbonyl (13C′) CSA pattern suggests limited mobility of this residue at room temperature, as the η value was ≥0.8 at both temperatures and the Ω value decreased modestly between −30 and 20 °C (Fig. 2C, Table 1 and Fig. S5). For FD31-U-E bound to nucleated calcite, the CSA pattern contained two isotropic peaks at ∼176 ppm and ∼180 ppm (Fig. 2D), which were assigned to the backbone (13C′) and side chain (13Cδ) carbonyls, respectively, based on average chemical shift assignments for glutamic acid deposited in the Biological Magnetic Resonance Data Bank (BMRB).55 Similar to calcite bound FD31-U-T, the calcite bound FD31-U-E 13C′ CSA pattern also suggested limited mobility at room temperature as the isotropic peak at ∼176 ppm had an η ≥ 0.8 at both temperatures, and the Ω changed only slightly between −30 and 20 °C (Fig. 2D, Table 1 and Fig. S5). The decrease in intensity for the CSA pattern of calcite bound FD31-U-T and FD31-U-E 13C’ does suggest a slight increase in mobility from −30 to 20 °C, as cross polarization efficiency decreases as a function of increased mobility, resulting in decreased signal to noise as mobility increases. For the calcite bound FD31-U-E 13Cδ CSA pattern, the isotropic peak at ∼180 ppm became sharper at room temperature and the number of side bands decreased from 2 sets to 1 set, consistent with an increase in mobility at 20 °C (Fig. 2D, grey asterisks). The η value also suggests increased mobility as a function of temperature, as it went from 1.0 in the frozen control to 0.3 at 20 °C. The Ω increased from −30 to 20 °C, which would suggest a decrease in mobility, however, this is likely due to the challenges fitting this spectrum due to overlap between the 13C′ and 13Cδ chemical shifts. While the CSA parameters all indicate increased mobility of the side-chain carbonyl as a function of increased temperature, the presence of FD31-U-E 13Cδ signal in the CP experiment rules out a highly mobile state at room temperature, as highly mobile systems typically exhibit poor cross-polarization efficiency and thus would not be detectable in CP experiments. Therefore, FD31-U-E 13Cδ likely occupies an intermediate mobility regime at room temperature, as the CSA pattern is inconsistent with a highly immobilized system and the presence of signal in a CP experiment is inconsistent with a highly mobile system.
| Temperature | Hertzfeld-Berger analysis | T 1ρ | ||||||
|---|---|---|---|---|---|---|---|---|
| Ω (ppm) | η | σ iso (ppm) | Linewidth (ppm) | Result | nRMSD | T 1ρ (msec) | Result | |
| a Overlap between the 13C′ and 13Cδ chemical shifts made fitting the individual Ω parameter for these resonances difficult. | ||||||||
| Threonine 13C′ | ||||||||
| 20 °C | 111.6 | 1.0 | 173.0 | 14.8 | Immobile | 0.15 | 37.3 | Immobile |
| −30 °C | 133.5 | 0.8 | 173.0 | 14.4 | Immobile | 0.12 | 21.1 | Immobile |
| Glutamic Acid 13C′ | ||||||||
| 20 °C | 117.5a | 0.8 | 175.6 | 11.0 | Immobile | 0.08 | 25.7 | Immobile |
| −30 °C | 108.4a | 1.0 | 176.0 | 13.4 | Immobile | 0.14 | 16.6 | Immobile |
| Glutamic Acid 13Cδ | ||||||||
| 20 °C | 83.2a | 0.3 | 180.2 | 6.3 | Mobile | 0.08 | 26.8 | Immobile |
| −30 °C | 52.5a | 1.0 | 181.0 | 6.7 | Immobile | 0.14 | 14.8 | Immobile |
To assess mobility on the millisecond timescale, 13C T1ρ relaxation times were measured56 for all carbonyls in the calcite bound FD31-U-T and FD31-U-E samples. For both samples, all carbonyl 13C T1ρ relaxation times were > 25 ms at 20 °C, values characteristic of immobile sites (Table 1 and Fig. S6). While the measured T1ρ values were smaller for each carbonyl under the frozen condition, which is indicative of higher mobility at −30 °C, this is likely just due to noise in the spectra, as the raw temperature curves show there is minimal decrease in signal intensity as a function of time under both temperature conditions (Fig. S6).
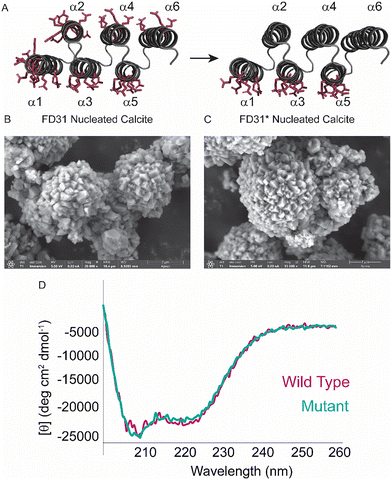
Limited mobilities on both tested timescales for the threonine and glutamic acid 13C′ in calcite nucleated FD31 could be from constrained mobility in the α-helical conformation, interactions with the surface, or a combination of both. Different mobilities for the glutamic acid side chain carbonyl on the 0.5 millisecond (CSA, mobile) and ∼10 millisecond (T1ρ, immobile) timescales could be explained by two distinct possibilities. The first is that the CSA is being dominated by the population of glutamic acid residues located in the three helices exposed to the bulk solution (Fig. 2B), as these residues would be expected to be highly mobile at room temperature. To address this possibility and prevent the mobile glutamic acid side chain population from potentially skewing the average dynamics measurement, an FD31 construct was designed with the 25 glutamic acid residues facing the bulk solution substituted to aspartic acid residues (FD31*) (Fig. 3A). Aspartic acid, the only other negatively charged amino acid, was chosen to maintain the protein's overall charge distribution and preserve the ability of FD31* to fold. The second explanation is that overlap between the 13C′ and 13Cδ peaks made fitting these individual peaks difficult, which lead to incorrect interpretations in the mobility for each individual carbonyl resonance. To address this in later experiments, only one carbonyl was selectively 13C enriched per sample.
Characterization of FD31*
To confirm substituting 25 glutamic acid residues for aspartic acid did not significantly alter the structure and calcite nucleating properties of FD31, calcite samples nucleated with either FD31 or FD31* were screened with scanning electron microscopy (SEM). Both complexes were observed to be morphologically similar with a diameter of ∼5 µm and have equally rough surfaces (Fig. 3B and C). Calcite prepared using the same protocol but without FD31 or FD31* protein present displays the classical rhombohedron morphology characteristic expected for calcite {104} in SEM (Fig. S7). The two proteins were further characterized by comparing the profiles of steady-state circular dichroism (CD) wavelength spectra in the absence of calcite. Both proteins had identical CD profiles, confirming that the 25 Glu to Asp substitutions did not alter the helical content of the protein in solution (Fig. 3D). In summary, the data suggest that the 25 glutamic acid to aspartic acid substitutions do not alter the structure of the protein prior to calcite nucleation, or the ability of the protein to nucleate calcite. This enabled us to use FD31* as a mimetic system for FD31 to probe the dynamics of only glutamic acid residues designed to interact with the inorganic surface.Dynamics of FD31* on nucleated calcite
In the original FD31-U-E nucleated calcite CSA spectra (Fig. 2D) the 13Cδ (grey arrow) and 13C′ (black arrow) resonances overlapped, making it difficult to analyze the contribution of each resonance to the spectrum. To eliminate this challenge and simplify the spectrum, two FD31* samples prepared with either the backbone (FD31*-EC′) or the side chain (FD31*-ECδ) carbonyls selectively 13C-labeled (Table S1), and these proteins were used to produce two separate nucleated calcite samples (Fig. 4). For the 13C{1H} CP-MAS spectrum of FD31*-EC′ calcite, the CSA was characteristic of an immobile system with η = 0.7 at −15 and 20 °C and little change in the Ω values as a function of temperature (Table 2, Fig. 4B and Fig. S8). There was also no intensity change observed in the glutamic acid 13C′ resonance at −15 and 20 °C, further suggesting that the mobility of the backbone did not change as a function of temperature at these glutamic acid sites. For the 13C{1H} CP-MAS spectrum of FD31*-ECδ, the CSA was characteristic of a mobile system at room temperature with η close to 0 and smaller Ω values at 20 °C than at −15 °C (Table 2, Fig. 4C and Fig. S8), and the presence of only one set of spinning side bands. While the main isotropic peak at ∼180 ppm also had narrow linewidths, a spectral feature indicative of a mobile system, overlap between the main resonance and an unexpected shoulder peak made accurate fitting of this peak difficult, so limited conclusions can be drawn from this feature alone. To rule out the possibility of a highly mobile population present for FD31*-ECδ that was not visible in the CSA profile due to poor cross polarization efficiency for highly mobile systems, a direct excitation spectrum was acquired for FD31*-ECδ nucelated calcite. The overall protein line shape and CSA pattern appears consistent between the CP and DP spectra, confirming that the only resonance for FD31*-ECδ corresponds to a semi-mobile protein population that is visible with both direct excitation and cross polarization pulses (Fig. S9). The unexpected shoulder peak observed in the FD31*-ECδ CSA spectrum (Fig. 4C, blue arrow) is discussed in more detail below.| Herzfeld-Berger analysis | T 1ρ analysis | |||||||
|---|---|---|---|---|---|---|---|---|
| Ω (ppm) | η | σ iso (ppm) | Linewidth (ppm) | Results | nRMSD | T 1ρ (ms) | Results | |
| FD31*-EC′ nucleated calcite | ||||||||
| 20 °C | 140.0 | 0.7 | 173.8 | 13.93 | Immobile | 0.10 | 93.2 | Immobile |
| −15 °C | 137.3 | 0.7 | 173.8 | 14.36 | Immobile | 0.09 | ||
| FD31*-ECd nucleated calcite, main peak | ||||||||
| 20 °C | 110.2 | 0.0 | 180.4 | 8.89 | Mobile | 0.15 | 90.6 | Immobile |
| −15 °C | 126.0 | 0.0 | 180.2 | 7.99 | Mobile | 0.10 | ||
| FD31*-ECd nucleated calcite, shoulder | ||||||||
| 20 °C | 107.0 | 0.0 | 176.1 | 6.19 | Mobile | 0.15 | 61.5 | Immobile |
| −15 °C | 98.14 | 0.0 | 176.4 | 5.03 | Mobile | 0.10 | ||
While CSA parameters provide only a qualitative assessment of mobility, examples from literature consistently show that a mobile carbonyl produces a CSA pattern characterized by both an isotropic peak with higher intensity relative to neighboring sidebands and an overall spectrum with poor signal-to-noise compared to a more static carbonyl (Fig. S4, bottom left). Examples from literature include the backbone carbonyls in leucine 42 of LRAP on hydroxyapatite (HAP),30 isoleucine 11 and glycine 12 in salivary peptide on HAP57 and phenylalanine 7 and leucine 8 in statherin on HAP,58 which all display this mobile profile in their hydrated, surface-bound state at room temperature. This same mobile pattern is observed in this work for FD31*-ECδ. In contrast, an immobile carbonyl will produce a CSA pattern in which a neighboring sideband is likely higher in intensity than the main isotropic peak, there will be multiple sets of spinning sidebands present, and the overall signal-to-noise of the spectrum will be high (Fig. S4, bottom right. Examples from literature include the backbone carbonyls in phosphorylated serine 2 and 3 in LRAP on HAP,57 in phosphorylated serine 2 and 3 in statherin bound to HAP58 and in N-terminal region of a peptoid occluded in calcium carbonate,59 which all display an immobile profile in their hydrated, surface-bound state at room temperature. This same immobile pattern is observed in this work for FD31*-EC′. In summary, based off the CSA patterns and subsequent analysis, the glutamic side chains carbonyls had more mobility on the sub-millisecond timescale compared to the backbone carbonyls.
To assess mobility on the millisecond timescale, 13C T1ρ relaxation times were measured. Both the FD31*-EC′ and FD31*-ECδ nucleated calcite samples had 13C′ T1ρ values indictive of highly immobile, rigid systems (Table 2 and Fig. S10A, B). While the rigidity of the FD31*-EC′ could be either from interactions with the surface or just limited mobility within an α-helical backbone, immobility at the FD31*-ECδ site does confirm an interaction between FD31* and the surface, as the glutamic acid sidechains would not be expected to be immobile in this system if they were not interacting with calcite. Due to the very poor S/N for FD31*-ECδ nucleated calcite sample that necessitated very long acquisition times, and the expectation that they would also be rigid, T1ρ values were not measured for either sample at −15 °C.
Interestingly, the CSA (mobile) and 13C T1ρ relaxation (immobile) data for calcite bound FD31*-ECδ appear to contradict each other (Table 2). The CSA features identify that the glutamic acid 13Cδ has mobility on the sub-millisecond timescale (Fig. 4C) while the T1ρ decay shows the glutamic acid 13Cδ has limited to no mobility in the millisecond timescale (Fig. S10B), suggesting that the side chain experiences different mobilities on different timescales. The most probable explanation for the observed side chain dynamics is that direct interactions with calcium ions on the surface immobilize the residue on the millisecond timescale, but interactions with water molecules on the surface cause minor fluctuations on the sub-millisecond timescale. While the original model for FD31 nucleated calcite assumed that the glutamic acid side chains displace surface bound water to interact with calcium directly, these dynamics results suggest that residual water molecules are likely still present on the surface.
Investigation of shoulder peak in FD31*-ECδ nucleated calcite
An unexpected shoulder (Fig. 4C, blue arrow) was also observed in the FD31*-ECδ calcite CSA spectrum. Superimposition of the FD31 and FD31*-ECd nucleated calcite CSA spectra confirm that the main resonance (∼180 ppm) originally assigned to the Glu side chain carbonyl (13Cδ) superimposes on the main resonance in the FD31*-ECδ, and the shoulder has a chemical shift slightly downfield from the resonance assigned to backbone carbonyl in the FD31*-E′ spectrum (Fig. 5A). Possible explanations for this unexpected peak are contamination of C′ labeled glutamic acid in the starting material used to grow the FD31*-ECδ cell paste, the adoption of two glutamic acid populations caused by interactions with the calcite surface or scrambling of the isotopic label during protein synthesis in E. coli. The contamination explanation was ruled out by collecting a one-dimension 13C solution-state NMR spectrum for the labeled amino acid used in protein synthesis, which contained a single isotopically enriched resonance with a chemical shift expected for the Glu side chain carbonyl carbon (Fig. S11). The explanation that interactions with the surface promoted adoption of two distinct FD31*-ECδ populations was also ruled out by collecting a 13C{1H} CP-MAS spectrum on lyophilized FD31*-ECδ which showed that the main resonance and the shoulder were both present in the absence of calcite, and therefore, the shoulder did not arise as a result of surface interactions (Fig. 5B). The biosynthetic scrambling explanation is possible as E. coli uses glutamic acid as a starting material to synthesize arginine, glutamine, and proline (Fig. S12)52 to incorporate the glutamic acid 13Cδ into the respective 13Cδ positions in these amino acids. Of these three scrambling possibilities, only the 13Cδ for glutamine has a chemical shift (180.4 ppm) in the range that corresponds to the chemical shift of the shoulder in the FD31*-Eδ calcite spectrum (Fig. 5A and Fig. S12: Arg Cδ ∼43.2 ppm; Pro Cδ ∼ 48.8 ppm).55 Because FD31 contains four glutamine and 18 glutamic acid residues, the maximum signal intensity for glutamine is expected to be ∼22% of that for glutamic acid. To get an accurate measure of signal intensity for the two residues, lyophilized FD31*-Eδ was dissolved in D2O and a one-dimension 13C solution-state NMR spectrum collected (Fig. 5C). Peak integrals were calculated using MestReNova for the Cδ (180–182 ppm) and shoulder (176.5–178 ppm) regions. The integral for the shoulder region is about ∼14% of that for the Cδ region, confirming that biosynthetic scrambling of glutamic acid into glutamine is a viable explanation for the shoulder peak.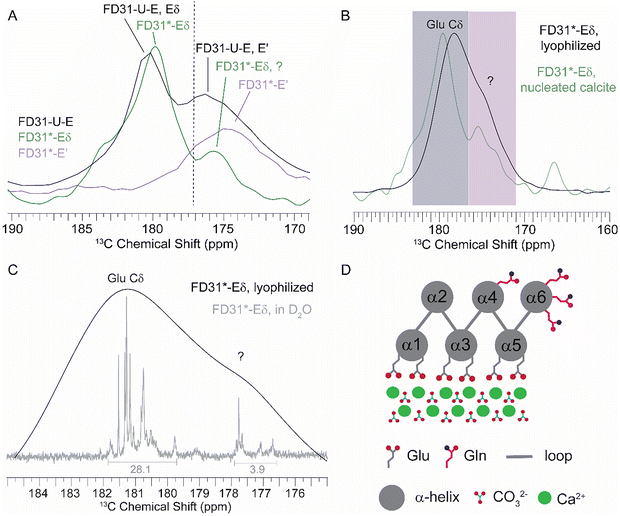 | ||
| Fig. 5 Characterization of shoulder peak from FD31*-ECδ nucleated calcite CSA spectrum. (A) Superimposed spectra of the isotropic resonance for FD31*-EC′ (purple), FD31*-ECδ (green) and FD31-U-E (black). (B) Superimposed 13C{1H} CP-MAS for FD31*-ECδ in the lyophilized state (black) and on the surface of nucleated calcite (green), showing the main resonance (Glu Cδ; grey shaded area) and shoulder resonance (?; purple shaded area) are both present when FD31* is on and off the calcite surface. Note the resonance for Glu Cδ is perturbed ∼ 3 ppm when on the surface, consistent with a change in the chemical environment due to interaction with the calcite surface. On the other hand, the shoulder resonance does not experience a change in chemical shift, consistent with the shoulder corresponding to a region of the protein that does not interact with the surface. (C) Superimposed CP-MAS of FD31*-ECδ in the lyophilized state (black) and one-dimension 13C solution NMR of FD31*-ECδ dissolved in D2O (gray) showing that the integral of the shoulder (?) (176.5–178ppm) is about ∼14% of the Cδ (180–182ppm) region. The multiple peaks observed in the solution state spectrum are from the 18 different glutamic acid residues in FD31*. (D) The modified model of FD31 templated CaCO3 formation from Fig. 1 highlighting the location of all four glutamine residues in the complex. | ||
With the knowledge that the shoulder peak in the FD31*-Eδ CSA spectrum must belong to the Cδ of glutamine due to scrambling, the superimposed spectra comparing the lyophilized and nucleated calcite FD31*-Eδ spectra was re-analyzed. Interestingly, only the peak assigned to the Cδ of glutamic acid shows a chemical shift change, whereas there is no significant chemical shift perturbation of this shoulder resonance between the lyophilized and calcite bound state (Fig. 5B). This is the first direct molecular evidence that the negatively charged carboxylic acid groups in glutamic acid are experiencing electrostatic interactions with the positively charged calcium ions in the calcite, as this would modify the electronic environment at the 13Cδ carbon and result in the observed chemical shift perturbation. This conclusion also supports the original docking model, as the four glutamine residues in FD31* are predicted to sit opposite of the FD31-calicite interface in the original model and would therefore not be expected to undergo a change in chemical environment that results in a change in chemical shift as a function of being on and off the surface (Fig. 5D).17
Orientation of Glu side chain carbonyl relative to the calcite surface
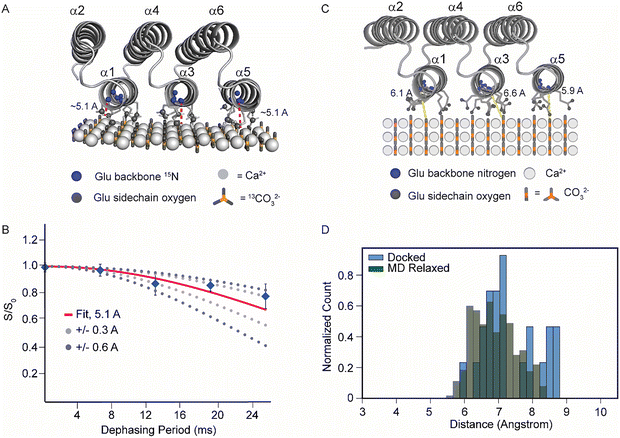
To establish the molecular orientation of FD31* on the calcite surface, ssNMR distance-dependent measurements were conducted to determine the distance between the FD31* backbone to the calcite surface using a Rotational Echo DOuble Resonance (REDOR) experiment. In a REDOR experiment, a series of pulses are applied to re-introduce dipolar coupling between two spin systems. If the two spins are tightly coupled, i.e. 7 Å or less apart for a 15N-13C spin system, the distance between them can be accurately measured by comparing experimental data to simulated dephasing curves.60 To facilitate this experiment for our system, 15N-labeled glutamic acid residues were incorporated into FD31* (FD31*-EN) and 13C nuclei were incorporated onto the calcite surface (Fig. 6A). The dephasing observed in the FD31*-E15N-Ca13CO3 spin system over 128 rotor cycles showed the best alignment with a modeled distance of approximately 5.1 Å (χ2 = 0.0139). However, the data also exhibited relatively low error when compared to greater distances of 5.4 Å (χ2 = 0.0174) and 5.7 Å (χ2 = 0.0236) (Fig. 6B and Table S3). In the original docking model with fully extended glutamic acid side chains,17 the backbone was an average of ∼7–9 Å from the calcite surface (Fig. S13), which is about ∼2–4 Å farther than the experimentally measured distances in our REDOR experiment. Previous computational and experimental studies have highlighted the importance of water molecules in stabilizing protein binding to highly charged interfaces.61–64 Therefore, the absence of explicit solvent interactions in the initial docking calculations could account for the larger protein-to-surface distances observed in silico.In order to investigate the role of solvent interactions at the protein-calcite interface, a 100 ns molecular dynamics (MD) simulation was run in explicit water. Upon relaxation, protein secondary structure contracted but remained stable, and the protein moved closer to the surface than in the original docking model (Fig. 6C, D and Fig. S14). On average, the carbon at position 5 in the surface-bound glutamic acid sidechains were ∼0.4 Å closer to the surface in the 100 ns MD simulation compared to the original docking model (Fig. S14). As a result, the protein backbone was pulled towards the surface, shifting the distribution of distances between glutamic acid backbone nitrogen atoms and the calcite surface from ∼7–9 Å originally (Fig. S13) to ∼6–8 Å (Fig. 6D), a range more consistent to the experimental measurements. This improved agreement between computation and experiment in the refined MD simulations reinforces that omitting explicit waters in the original docking model likely resulted in artificially long protein-surface distances.
However, a slight discrepancy remains as the experimental results show a FD31*-E15N-Ca13CO3 distance of ∼5.1–5.7 Å, which is ∼1–2 Å closer than the water-refined MD simulations. Notably, the newly refined docking model predicted that some glutamic acid sidechains would splay away from the surface (Fig. 6C). During the simulations, these sidechains did not reorient to bind, likely due to the high energetic cost associated with dehydrating calcite-bound waters.63–66 Such energetic barriers would prevent these splayed glutamic acid residues from binding closer to the surface in the simulations, even though the resulting interactions would be more favorable. In contrast, under experimental conditions the sidechains may overcome these barriers through motions on extended timescales or alternative nucleation pathways, which bypass the need for water displacement. Such mechanisms could explain the slightly closer protein-calcite distances observed in our ssNMR experiments compared to the computational model.
To determine if a ∼5.1-5.7 Å FD31*-E15N-Ca13CO3 distance was energetically favorable if the FD31* sidechains were fully extended at the surface, an additional simulation was run in which the protein was positioned 4 Å from the surface. This initial positioning created a conformation in which the protein sidechains sterically clashed with the surface, so a subsequent energy minimization was run to allow the sidechains to relax into a stable conformation. This resulted in an average protein backbone-surface distance distribution that closely matched the experimental results, confirming that the distances measured with ssNMR are achievable if the protein sidechains are fully elongated at the surface as long as there is no water later separating the protein and calcite surface (Fig. S15). In all, this provides possible insight into the molecular arrangement of FD31 at the surface interface.
Conclusion
In this study, we sought to uncover the molecular mechanisms underlying FD31's ability to lower the free energy of calcite nucleation and subsequent ability to template calcite {110}. Our current study sheds light on the complex nucleation pathway associated with the formation of calcite. Our MD simulation data support results from the literature that sidechains struggle to displace tightly bound interfacial water molecules. However, ssNMR experimental results suggest an inconsistency as the FD31 glutamic acid sidechains must be displacing water to achieve a ∼5.1 Å distance measured between the protein and the calcite surface. In our experimental system, calcite is nucleated by combining a solution containing protein and calcium ions with a solution containing bicarbonate ions. As we currently hypothesize that the protein plays a role in templating calcium carbonate, the most likely explanation for the distances observed in our experiments is that the protein templates amorphous calcium carbonate (ACC), which is then dehydrated during the transition to calcite (Fig. S16). However, the presence of dynamics for the glutamic acid sidechain on the microsecond timescale does suggest the presence of some residual water molecules, which can be explained by the residual water molecules interspersed between sidechain residues at the interface as observed in the 100 ns MD simulation (Fig. S17). This newly proposed mechanism would allow for a direct protein-calcite interaction without the displacement of tightly bound interfacial waters. The results gleaned from this study will motivate future investigation into the potential mechanism of dehydration of the calcium carbonate surface during templated nucleation.These findings also highlight a key limitation of simple geometry-based lattice-matching models, and suggest the omission of explicit solute interactions and bound surface waters likely accounts for discrepancies between predictions based on these models and experimental measurements. Incorporating explicit hydration effects in future models will be critical to designing more accurate constructs for protein-templated biomineralization. Ultimately, this study underscores the importance of integrating experimental and computational approaches to refine de novo protein designs for controlling crystallization processes, paving the way for improved biomolecular engineering strategies.
Conflicts of interest
There are no conflicts of interest to declare.Data availability
All experimental data files for this article, including all Nuclear Magnetic Resonance and Circular Dichroism spectra, are available on DataHub at https://doi.org/10.25584/3001598. All simulation input files and analysis scripts can be accessed at https://github.com/zormanmarlo/DHR_rsc.The supplementary information consists of supplementary figures and tables that compliment all NMR and MD experiments found in the main text. See DOI: https://doi.org/10.1039/d5ma01322d.
Acknowledgements
This work was supported by the US Department of Energy (DOE), Office of Science, Office of Basic Energy Sciences as part of the Energy Frontier Research Centers program through CSSAS – The Center for the Science of Synthesis Across Scales under Award Number DE-SC0019288 at the University of Washington and FWP-77248 at Pacific Northwest National Laboratory (PNNL). Garry Buchko, who provided training and aided in manuscript preparation, was supported by the National Institutes of Health, National Institute for Dental and Craniofacial Research under Grant number DE-015347. Circular dichroism was performed at the Environmental and Molecular Sciences Laboratory (EMSL), a national scientific user facility at PNNL sponsored by the U.S. DOE Office of Biological and Environmental Research under award number 61587. PNNL is a multi-program national laboratory operated by Battelle Memorial Institute under contract no. DE-AC05-76RL01830 for the DOE.References
- L. Addadi, S. Raz and S. Weiner, Taking Advantage of Disorder: Amorphous Calcium Carbonate and Its Roles in Biomineralization, Adv. Mater., 2003, 15(12), 959–970, DOI:10.1002/adma.200300381.
- E. Beniash, Biominerals-hierarchical nanocomposites: the example of bone, Wiley Interdiscip. Rev.: Nanomed. Nanobiotechnol., 2011, 3(1), 47–69, DOI:10.1002/wnan.105.
- L. A. Estroff, Introduction: Biomineralization, Chem. Rev., 2008, 108(11), 4329–4331, DOI:10.1021/cr8004789.
- C. A. Schmidt, et al., Myriad Mapping of nanoscale minerals reveals calcium carbonate hemihydrate in forming nacre and coral biominerals, Nat. Commun., 2024, 15(1), 1812, DOI:10.1038/s41467-024-46117-x.
- F. C. Meldrum, Calcium carbonate in biomineralisation and biomimetic chemistry, Int. Mater. Rev., 2013, 48(3), 187–224, DOI:10.1179/095066003225005836.
- N. K. Dhami, M. S. Reddy and A. Mukherjee, Biomineralization of calcium carbonates and their engineered applications: a review, Front. Microbiol., 2013, 4, 314, DOI:10.3389/fmicb.2013.00314.
- M. M. H. Al Omari, I. S. Rashid, N. A. Qinna, A. M. Jaber and A. A. Badwan, Calcium Carbonate, Profiles of Drug Substances, Excipients and Related Methodology, Academic Press, 2016, pp. 31–132 Search PubMed.
- M. Rose-Martel, S. Smiley and M. T. Hincke, Novel identification of matrix proteins involved in calcitic biomineralization, J. Proteomics, 2015, 116, 81–96, DOI:10.1016/j.jprot.2015.01.002.
- S. Mann, et al., Crystallization at Inorganic-organic Interfaces: Biominerals and Biomimetic Synthesis, Science, 1993, 261(5126), 1286–1292, DOI:10.1126/science.261.5126.1286.
- B. A. Gotliv, L. Addadi and S. Weiner, Mollusk shell acidic proteins: in search of individual functions, ChemBioChem, 2003, 4(6), 522–529, DOI:10.1002/cbic.200200548.
- G. Fu, S. Valiyaveettil, B. Wopenka and D. E. Morse, CaCO3 biomineralization: acidic 8-kDa proteins isolated from aragonitic abalone shell nacre can specifically modify calcite crystal morphology, Biomacromolecules, 2005, 6(3), 1289–1298, DOI:10.1021/bm049314v.
- B. A. Gotliv, et al., Asprich: A novel aspartic acid-rich protein family from the prismatic shell matrix of the bivalve Atrina rigida, ChemBioChem, 2005, 6(2), 304–314, DOI:10.1002/cbic.200400221.
- B. Pokroy, M. Kapon, F. Marin, N. Adir and E. Zolotoyabko, Protein-induced, previously unidentified twin form of calcite, Proc. Natl. Acad. Sci. U. S. A., 2007, 104(18), 7337–7341, DOI:10.1073/pnas.0608584104.
- J. Aizenberg, G. Lambert, S. Weiner and L. Addadi, Factors involved in the formation of amorphous and crystalline calcium carbonate: a study of an ascidian skeleton, J. Am. Chem. Soc., 2002, 124(1), 32–39, DOI:10.1021/ja016990l.
- G. K. Hunter, Interfacial aspects of biomineralization, Curr. Opin. Solid State Mater. Sci., 1996, 1(3), 430–435, DOI:10.1016/s1359-0286(96)80036-2.
- I. K. Voets, From ice-binding proteins to bio-inspired antifreeze materials, Soft Matter, 2017, 13(28), 4808–4823, 10.1039/c6sm02867e.
- F. A. Davila-Hernandez, et al., Directing polymorph specific calcium carbonate formation with de novo protein templates, Nat. Commun., 2023, 14(1), 8191, DOI:10.1038/s41467-023-43608-1.
- J. R. Long, W. J. Shaw, P. S. Stayton and G. P. Drobny, Structure and dynamics of hydrated statherin on hydroxyapatite as determined by solid-state NMR, Biochemistry, 2001, 40(51), 15451–15455, DOI:10.1021/bi010864c.
- M. Gilbert, et al., Chimeric peptides of statherin and osteopontin that bind hydroxyapatite and mediate cell adhesion, J. Biol. Chem., 2000, 275(21), 16213–16218, DOI:10.1074/jbc.M001773200.
- E. L. Buckle, A. Prakash, M. Bonomi, J. Sampath, J. Pfaendtner and G. P. Drobny, Solid-State NMR and MD Study of the Structure of the Statherin Mutant SNa15 on Mineral Surfaces, J. Am. Chem. Soc., 2019, 141(5), 1998–2011, DOI:10.1021/jacs.8b10990.
- C. Zerfass, G. W. Buchko, W. J. Shaw, S. Hobe and H. Paulsen, Secondary structure and dynamics study of the intrinsically disordered silica-mineralizing peptide P(5) S(3) during silicic acid condensation and silica decondensation, Proteins, 2017, 85(11), 2111–2126, DOI:10.1002/prot.25366.
- A. Porebska, M. Rozycka, R. Holubowicz, Z. Szewczuk, A. Ozyhar and P. Dobryszycki, Functional derivatives of human dentin matrix protein 1 modulate morphology of calcium carbonate crystals, FASEB J., 2020, 34(5), 6147–6165, DOI:10.1096/fj.201901999R.
- J. M. Gibson, J. M. Popham, V. Raghunathan, P. S. Stayton and G. P. Drobny, A solid-state NMR study of the dynamics and interactions of phenylalanine rings in a statherin fragment bound to hydroxyapatite crystals, J. Am. Chem. Soc., 2006, 128(16), 5364–5370, DOI:10.1021/ja056731m.
- R. J. Arachchige, et al., Solid-State NMR Identification of Intermolecular Interactions in Amelogenin Bound to Hydroxyapatite, Biophys. J., 2018, 115(9), 1666–1672, DOI:10.1016/j.bpj.2018.08.027.
- M. Ndao, J. T. Ash, P. S. Stayton and G. P. Drobny, The Role of Basic Amino Acids in the Molecular Recognition of Hydroxyapatite by Statherin using Solid State NMR, Surf. Sci., 2010, 604(15–16), L39–L42, DOI:10.1016/j.susc.2010.02.026.
- J. M. Gibson, V. Raghunathan, J. M. Popham, P. S. Stayton and G. P. Drobny, A REDOR NMR study of a phosphorylated statherin fragment bound to hydroxyapatite crystals, J. Am. Chem. Soc., 2005, 127(26), 9350–9351, DOI:10.1021/ja050910m.
- A. Roehrich and G. Drobny, Solid-state NMR studies of biomineralization peptides and proteins, Acc. Chem. Res., 2013, 46(9), 2136–2144, DOI:10.1021/ar300321e.
- W. J. Shaw, Solid-state NMR studies of proteins immobilized on inorganic surfaces, Solid State Nucl. Magn. Reson., 2015, 70, 1–14, DOI:10.1016/j.ssnmr.2014.10.003.
- Y. P. Du, et al., Study of Binding Interaction between Pif80 Protein Fragment and Aragonite, Sci. Rep., 2016, 6, 30883, DOI:10.1038/srep30883.
- W. J. Shaw, K. Ferris, B. Tarasevich and J. L. Larson, The structure and orientation of the C-terminus of LRAP, Biophys. J., 2008, 94(8), 3247–3257, DOI:10.1529/biophysj.107.119636.
- R. J. Arachchige, et al., Solid-State NMR Identification of Intermolecular Interactions in Amelogenin Bound to Hydroxyapatite, Biophys. J., 2018, 115(9), 1666–1672, DOI:10.1016/j.bpj.2018.08.027.
- G. W. Buchko, J. Bekhazi, J. R. Cort, N. B. Valentine, M. L. Snead and W. J. Shaw, 1H, 13C, and 15N resonance assignments of murine amelogenin, an enamel biomineralization protein, Biomol NMR Assign, 2008, 2(1), 89–91, DOI:10.1007/s12104-008-9092-x.
- G. W. Buchko, G. Lin, B. J. Tarasevich and W. J. Shaw, A solution NMR investigation into the impaired self-assembly properties of two murine amelogenins containing the point mutations T21 > I or P41 > T, Arch. Biochem. Biophys., 2013, 537(2), 217–224, DOI:10.1016/j.abb.2013.07.015.
- R. H. Griffey, A. G. Redfield, R. E. Loomis and F. W. Dahlquist, Nuclear magnetic resonance observation and dynamics of specific amide protons in T4 lysozyme, Biochemistry, 1985, 24(4), 817–822, DOI:10.1021/bi00325a001.
- A. Pines, J. S. Waugh and M. G. Gibby, Proton-Enhanced Nuclear Induction Spectroscopy - Method for High-Resolution Nmr of Dilute Spins in Solids, J. Chem. Phys., 1972, 56(4), 1776, DOI:10.1063/1.1677439.
- A. Pines, M. G. Gibby and J. S. Waugh, Proton-Enhanced Nmr of Dilute Spins in Solids, J. Chem. Phys., 1973, 59(2), 569–590, DOI:10.1063/1.1680061.
- N. Sinha, C. V. Grant, C. H. Wu, A. A. De Angelis, S. C. Howell and S. J. Opella, SPINAL modulated decoupling in high field double- and triple-resonance solid-state NMR experiments on stationary samples, J. Magn. Reson., 2005, 177(2), 197–202, DOI:10.1016/j.jmr.2005.07.008.
- O. Toke, Three Decades of REDOR in Protein Science: A Solid-State NMR Technique for Distance Measurement and Spectral Editing, Int. J. Mol. Sci., 2023, 24(17), 13637, DOI:10.3390/ijms241713637.
- A. L. Van Geet, Calibration of the methanol and glycol nuclear magnetic resonance thermometers with a static thermistor probe, Anal. Chem., 1968, 40(14), 2227–2229 CrossRef CAS.
- A. Bielecki and D. P. Burum, Temperature-Dependence of Pb-207 Mas Spectra of Solid Lead Nitrate - an Accurate, Sensitive Thermometer for Variable-Temperature, J. Magn. Reson., Ser. A, 1995, 116(2), 215–220, DOI:10.1006/jmra.1995.0010.
- S. G. J. van Meerten, W. M. J. Franssen and A. P. M. Kentgens, ssNake: A cross-platform open-source NMR data processing and fitting application, J. Magn. Reson., 2019, 301, 56–66, DOI:10.1016/j.jmr.2019.02.006.
- U. Duvvuri, et al., Water magnetic relaxation dispersion in biological systems: the contribution of proton exchange and implications for the noninvasive detection of cartilage degradation, Proc. Natl. Acad. Sci. U. S. A., 2001, 98(22), 12479–12484, DOI:10.1073/pnas.221471898.
- J. Choi, et al., Depolarization Dynamics in a Strongly Interacting Solid-State Spin Ensemble, Phys. Rev. Lett., 2017, 118(9), 093601, DOI:10.1103/PhysRevLett.118.093601.
- T. Gullion and J. Schaefer, Rotational-echo double-resonance NMR, J. Magn. Reson., 2011, 213(2), 413–417, DOI:10.1016/j.jmr.2011.09.003.
- M. Bak, J. T. Rasmussen and N. C. Nielsen, SIMPSON: a general simulation program for solid-state NMR spectroscopy, J. Magn. Reson., 2011, 213(2), 366–400, DOI:10.1016/j.jmr.2011.09.008.
- M. J. Abraham, et al., GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers, SoftwareX, 2015, 1–2, 19–25, DOI:10.1016/j.softx.2015.06.001.
- R. Das and D. Baker, Macromolecular modeling with rosetta, Annu. Rev. Biochem., 2008, 77, 363–382, DOI:10.1146/annurev.biochem.77.062906.171838.
- H. J. C. Berendsen, J. R. Grigera and T. P. Straatsma, The missing term in effective pair potentials, J. Phys. Chem., 2002, 91(24), 6269–6271, DOI:10.1021/j100308a038.
- J. Huang, et al., CHARMM36m: an improved force field for folded and intrinsically disordered proteins, Nat. Methods, 2016, 14(1), 71–73, DOI:10.1038/nmeth.4067.
- J.-W. Shen, C. Li, N. F. A. van der Vegt and C. Peter, Understanding the Control of Mineralization by Polyelectrolyte Additives: Simulation of Preferential Binding to Calcite Surfaces, J. Phys. Chem. C, 2013, 117(13), 6904–6913, DOI:10.1021/jp402341w.
- D. Lacabanne, B. H. Meier and A. Bockmann, Selective labeling and unlabeling strategies in protein solid-state NMR spectroscopy, J. Biomol. NMR, 2018, 71(3), 141–150, DOI:10.1007/s10858-017-0156-z.
- S. Mondal, D. Shet, C. Prasanna and H. S. Atreya, High yield expression of proteins in E. coli for NMR studies, Adv. Biosci. Biotechnol., 2013, 04(06), 751–767, DOI:10.4236/abb.2013.46099.
- J. W. ApSimon, et al., NMR—the chemical shift—IV, Tetrahedron, 1970, 26(1), 119–146, DOI:10.1016/0040-4020(70)85014-1.
- J. Herzfeld and A. E. Berger, Sideband Intensities in Nmr-Spectra of Samples Spinning at the Magic Angle, J. Chem. Phys., 1980, 73(12), 6021–6030, DOI:10.1063/1.440136.
- E. L. Ulrich, et al., BioMagResBank, Nucleic Acids Res., 2007, 36, D402–D408, DOI:10.1093/nar/gkm957.
- E. Fukushima and S. B. Roeder, Experimental pulse NMR. Perseus Books Publishing, 1981 Search PubMed.
- W. J. Shaw, J. R. Long, A. A. Campbell, P. S. Stayton and G. P. Drobny, A Solid State NMR Study of Dynamics in a Hydrated Salivary Peptide Adsorbed to Hydroxyapatite, J. Am. Chem. Soc., 2000, 122(29), 7118–7119, DOI:10.1021/ja000878q.
- J. R. Long, W. J. Shaw, P. S. Stayton and G. P. Drobny, Structure and Dynamics of Hydrated Statherin on Hydroxyapatite As Determined by Solid-State NMR, Biochemistry, 2001, 40(51), 15451–15455, DOI:10.1021/bi010864c.
- E. G. S. Close, et al., Molecular Interactions between a Biomimetic Polymer and CaCO3 Characterized by Solid-State Nuclear Magnetic Resonance, J. Phys. Chem. C, 2025, 129(28), 13010–13022, DOI:10.1021/acs.jpcc.5c02180.
- T. Gullion and J. Schaefer, Rotational-echo double-resonance NMR, J. Mag. Res., 1989, 81(1), 196–200, DOI:10.1016/0022-2364(89)90280-1.
- J. L. Prelesnik, et al., Ion-dependent protein–surface interactions from intrinsic solvent response, Proc. Natl. Acad. Sci. U. S. A., 2021, 118(26), e2025121118, DOI:10.1073/pnas.2025121118.
- R. G. Alberstein, et al., Discrete Orientations of Interfacial Waters Direct Crystallization of Mica-Binding Proteins, J. Phys. Chem. Lett., 2022, 14(1), 80–87, DOI:10.1021/acs.jpclett.2c02948.
- S.-H. Chong and S. Ham, Dynamics of Hydration Water Plays a Key Role in Determining the Binding Thermodynamics of Protein Complexes, Sci. Rep., 2017, 7(1), 8744, DOI:10.1038/s41598-017-09466-w.
- B. Wallner, R. E. Pavlovicz, H. Park and F. DiMaio, Efficient consideration of coordinated water molecules improves computational protein-protein and protein-ligand docking discrimination, PLoS Comput. Biol., 2020, 16(9), e1008103, DOI:10.1371/journal.pcbi.1008103.
- C. L. Freeman, J. H. Harding, D. Quigley and P. M. Rodger, Simulations of Ovocleidin-17 Binding to Calcite Surfaces and Its Implications for Eggshell Formation, J. Phys. Chem. C, 2011, 115(16), 8175–8183, DOI:10.1021/jp200145m.
- A. Valiya Parambathu, L. Wang, D. Asthagiri and W. G. Chapman, Apolar Behavior of Hydrated Calcite (10
![[1 with combining macron]](https://www.rsc.org/images/entities/char_0031_0304.gif) 4) Surface Assists in Naphthenic Acid Adsorption, Energy Fuels, 2019, 33(7), 6119–6125, DOI:10.1021/acs.energyfuels.9b00877.
4) Surface Assists in Naphthenic Acid Adsorption, Energy Fuels, 2019, 33(7), 6119–6125, DOI:10.1021/acs.energyfuels.9b00877.
| This journal is © The Royal Society of Chemistry 2026 |