 Open Access Article
Open Access ArticleAngiogenic properties of poly(NAG-co-NAPA) nanoparticles: assessment via gray-level co-occurrence matrix-based image processing
Sukanya
Patra
a,
Aniket Dayanand
Lokhande
a,
Gurmeet
Singh
a,
Divya
Pareek
a,
Saumya
Jaiswal
b,
Prem Shankar
Gupta
a,
Abhijit
Majumder
 b,
Jac Fredo Agastinose
Ronickom
a and
Pradip
Paik
b,
Jac Fredo Agastinose
Ronickom
a and
Pradip
Paik
 *a
*a
aSchool of Biomedical Engineering, Indian Institute of Technology (Banaras Hindu University), Varanasi, Uttar Pradesh 221005, India. E-mail: paik.bme@iitbhu.ac.in
bDepartment of Chemical Engineering, Indian Institute of Technology Bombay, Mumbai, Maharashtra 400076, India
First published on 27th January 2026
Abstract
Angiogenesis is a complex physiological process that is involved in the formation of new blood vessels. Despite advancements in assessing angiogenesis, there is still a substantial opportunity for the development of more accessible, reliable, and automated processes for estimating this complex phenomenon. Although in vitro and in vivo studies are instrumental for identifying the molecular players in angiogenesis, computational and mathematical models are likewise crucial to understand and explain the formation of endothelial cell networks. Quantifying angiogenesis is difficult due to the non-availability of suitable methods. Manual analysis and quantification of such results with existing approaches is typically labour-intensive and affected by inter-experimental variability. With this in mind, the main aim of the present work is to develop a new method for assessing angiogenesis using a gray-level co-occurrence matrix (GLCM)-based textural-feature image processing tool, which could be co-related with more reliable parameters. To establish this method, amino acid-based model copolymer nanoparticles (NPs), i.e. poly[(N-acryloyl glycine)-co-(N-acryloyl-(L-phenylalanine methyl ester))], or p(NAG-co-NAPA), have been synthesized and tested for cell viability. These p(NAG-co-NAPA) NPs exhibited enhanced metabolic activity up to ∼115–120% with L929 (mouse fibroblast) cells, HUVECs (human umbilical endothelial cells) and RAW 264.7 macrophages. In the 2nd step, the pro-angiogenic properties of p(NAG-co-NAPA) NPs were investigated through an ‘in ovo’ model using the CAM assay. Approximately 1000 microscopic images of newly formed blood vessels were segmented using adaptive thresholding, and their angiogenic properties were analyzed using seven GLCM-based textural features. Furthermore, to predict their regeneration efficiency, the tube formation assay was performed using HUVEC cells and the results were cross-verified with the Angiotool method to establish the GLCM-based textural-feature method as a preliminary quantitative framework for the assessment of angiogenesis. In all, we have established a new approach for angiogenesis analysis, with the conclusion that p(NAG-co-NAPA) NPs could be paramount for various therapeutic applications.
1. Introduction
In biological systems, each organ is supplied by an artery that delivers oxygen and nutrients throughout the body.1,2 Endothelial cells, the major constituent of arteries, are responsible for forming tubular vessels, which are lined by a basement membrane, mural cells, and extracellular matrix1 that can be modified by suitable biomaterials and medicines.3 This complex three-dimensional structure of blood vessels can be remodeled to respond to stimulation by chemokines or cytokines that are released and/or by synthetic biomaterials, including polymers,4 during embryonic development and various diseases and disorders.5,6 The process of angiogenesis is a unique phenomenon that is involved in the formation of new blood vessels from pre-existing ones, and it can be controlled using amino acid-based biopolymers as therapeutics.7–10 In healthy tissues, it is tightly regulated to maintain vascular homeostasis and respond to changes in metabolic demand. However, under pathological conditions, it is dysregulated due to varying factors, e.g. due to tumors and cancers, and this subsequently leads to the formation of abnormal, leaky blood vessels.11 Thus, an accurate evaluation and quantification of angiogenesis is crucial for qualifying a novel therapeutic, such as a polymeric material, for use in clinical applications.In recent times, computational tools have gained popularity in predicting therapeutic doses to provide clinical decision support for multiple biomedical applications of designed NPs.12–15 In regular practice, microscopic images acquired through angiogenesis assays, such as in ovo or in vivo models, serve as a foundational approach for analyzing tissue and cellular architectures in both physiological and pathological states. These imaging techniques allow for the quantitative assessment of vascular parameters, including vessel length density, branch points, and morphological complexity, thus providing valuable insights into the mechanisms underlying angiogenesis and disease progression. Numerous CAM assay studies have employed diverse vasculature quantification methods, leading to outcome heterogeneity in the results.16–19 Image analysis is carried out through the available tools, such as ImageJ, AngioTool and Angioquant, which each have their limitations, including computational demand, manual steps, long processing times, and costly software.20 Although MATLAB-based tools like RAVE21 and REAVER22 have emerged, RAVE lacks 3D image analysis capabilities, and REAVER remains under development. AI-based platforms (e.g., the IKOSA CAM assay application and ROI selection) have been introduced to quantify branching points and mean vessel thickness with higher precision than traditional thresholding methods, though these approaches depend on adequate training datasets and may misidentify non-vascular structures.23,24 Machine learning and deep learning frameworks for vascular image analysis have been applied to 3D microfluidic and retinal angiogenesis models, enabling morphological tracking across time and the extraction of multiple quantitative features, yet their generalizability across modalities and sensitivity to imaging conditions are remaining challenges.25–27 The absence of standardized CAM vasculature quantification methodology limits inter-study data analysis and comparison. Thus, a new method for the accurate analysis of angiogenesis is essential. To address the said limitations, our present work utilizes a Gaussian adaptive threshold method followed by gray-level co-occurrence matrix (GLCM)-based textural feature analysis for the segmentation and angiogenesis assessment of a model amino-acid-based biopolymer. This thresholding method precisely segments blood vessels from the background,28 outperforming contour29 and clustering30 algorithms. The texture-based features provide a more comprehensive description of the image and incorporate information about the spatial distribution of changes in gray intensity levels.31 For instance, GLCM textural features have been applied to assess treatment response in glioblastoma multiform patients using MR images, demonstrating significant differences in second-order statistics between responsive and progressive disease states.32 Similarly, GLCM parameters have been employed to differentiate tissue characteristics in thyroiditis using ultrasound data, underscoring its utility in quantitative image interpretation beyond conventional intensity-based methods.33 More recent work has also explored advanced computational and machine-learning frameworks that integrate GLCM descriptors for enhanced feature extraction in medical image analysis.34 By accounting for both spatial and intensity information, this method analyzes angiogenesis and predicts therapeutic doses of anti/pro-angiogenic biomolecules for multiple therapeutic applications.
Further, a wide variety of pro-angiogenic biomolecules and angiogenesis inhibitors have been designed using inorganic materials, polymers, their composites, etc.35 It is reported that the angiogenic response could be controlled by tailoring the biomaterial structure and properties.4 Many peptide-, gene-, and growth-factor-based therapies have also been reported, which were used in a multifaceted approach to enhance angiogenesis and tissue regeneration.36–38 However, the future potential of these approaches is constrained by multiple factors, such as variable responses, specificity to certain cell types, short duration of action, differences between preclinical and clinical outcomes, regulatory challenges, and issues related to cost-effectiveness.39–41 Furthermore, amino acids play a significant role in regulating and maintaining vascular function, including coagulation, vascular tone, cell growth and differentiation, redox homeostasis, immune and inflammatory responses, fibrinolysis, etc.42,43 The essential amino acid L-phenylalanine promotes tetrahydrobiopterin synthesis, increases nitrite levels by activating the guanosine triphosphate cyclase hydrolase pathway, and leads to improved endothelial function;44–46 however, in some reports it was found that phenylalanine was associated with macrovascular diseases.47 Similarly, glycine participates in many physiological pathways and is found to be both angiogenic and antiangiogenic. Amin et al. have reported the effect of glycine on angiogenesis during embryonic development via the CAM assay, showing that exogenous glycine inhibited blood vessel growth by >50%.48 Dietary glycine has been reported as a potent anti-angiogenic agent which reduces wound healing and tumor growth by reducing iNOS expression.49 Contrary to the above discussion, Guo et al. have reported that glycine shows angiogenic behavior both in vitro and in vivo.50 Similarly, in our previous studies, glycine-based polymeric nanoparticles and hydrogels were found to promote angiogenesis, leading to enhanced tissue and neuron regeneration.7,10 By taking account of the above studies, we speculated that a combination of glycine and phenylalanine may show pro-angiogenic behavior by complementing each other and this motivated us to consider p(NAG-co-NAPA) NPs as a model copolymer, consisting of N-acryloyl glycine (NAG) and N-acryloyl-L-phenylalanine methyl ester (NAPA) as two monomers in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio. Their respective homopolymers p(NAG) and p(NAPA) were considered as internal control groups in this study.7,51,52 Further, the therapeutic applications are optimized by analysing the angiogenic properties through standard in vitro and in ovo models (chorioallantoic membrane (CAM) assay) across a range of doses and time points.
1 ratio. Their respective homopolymers p(NAG) and p(NAPA) were considered as internal control groups in this study.7,51,52 Further, the therapeutic applications are optimized by analysing the angiogenic properties through standard in vitro and in ovo models (chorioallantoic membrane (CAM) assay) across a range of doses and time points.
Accordingly, this work synthesized the model copolymer, p(NAG-co-NAPA) NPs, through mini-emulsion free radical polymerization followed by structural confirmation through standard characterization techniques. Hemocompatibility and biocompatibility studies were performed against RBCs, L929 (mouse fibroblast), RAW 264.7 (macrophages), and HUVEC cells. Based on the observations, the dose- and time-dependent angiogenic properties of p(NAG-co-NAPA) copolymer NPs were studied through an in ovo model. The microscopic images obtained from the CAM assay were segmented using an image-processing-based Gaussian adaptive threshold followed by gray-level co-occurrence matrix (GLCM) code to compute 7 different gray textural features (number of pixels of blood vessels, entropy, energy, mean, contrast, variance, and dissimilarity) related to angiogenesis analysis. Furthermore, to predict the regeneration efficiency, the tube formation assay was performed using HUVEC cells and the results were cross-verified with the Angiotool method to establish the GLCM-based textural feature method as a preliminary framework for the quantitative estimation of angiogenesis. Finally, by combining the experimental and computational results, the potential of the new image processing approach for analyzing angiogenesis has been developed, and the future potential of p(NAG-co-NAPA) NPs for wound care and tissue regeneration has been established.
2. Experimental
2.1. Materials
All chemicals were used as received and without further purification, unless otherwise stated. Glycine (98%, Qualigens), L-phenylalanine methyl ester (Sigma), triethylamine (≥99.5%, Merck), 1,4-dioxan (99%) extra-pure (Merck), magnesium sulphate anhydrous (SRL), potassium Hydroxide (KOH) (Sigma), hexadecane anhydrous (HD) (99%, Sigma), SDS (sodium dodecyl sulfate) (90%, Merck), divinyl benzene (DVB) (Alfa Aesar), acryloyl chloride stab. with 400 ppm phenothiazine (96%, Alfa Aesar), ethyl acetate (Qualligen), diethyl ether (Qualligen), hydrochloric acid, 2,2′-azo-bis-isobutyronitrile (AIBN, 98%, SRL), dichloromethane (Merck), sodium bisulphate (SRL), toluene (Merck), sodium chloride (Sigma), DMSO-d6 (Sigma), CDCl3 (Sigma), phosphate-buffered saline (pH ∼7.4) (Himedia), isopropanol (Qualligen), phosphotungstic acid (Sigma), Triton-X (Loba Chemie), DNS (sodium chloride and dextrose injection IP (0.9% and 5% w/v)) (Jedux), calcium chloride (Merck), methylthiazol tetrazolium (MTT, 99.9%, Himedia), Dulbecco's modified Eagle medium (DMEM, Cell Clone), penicillin streptomycin cocktail (Himedia), Geltrex (Gibco), calcein AM (Thermo Fisher Scientific) trypsin-EDTA (Himedia), 4% paraformaldehyde solution (Himedia), fetal bovine serum (FBS, Gibco), and DMSO (Merck) were used; L929 (mouse fibroblast cells), RAW 264.7 (macrophages), and HUVEC (human umbilical endothelial cells) cell lines were acquired from NCCS (Himedia) and ATCC. A Pure Lab Ultra water system (ELGA, High Wycombe, United Kingdom) was used to obtain ultrapure water (18.2 ΩM) for sample processing.2.2. Methods
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NAPA = 1
NAPA = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, w/w) was dispersed in 2 mL of toluene. Then, DVB and HD were added and bath sonicated (at RT) for 5 minutes followed by stirring (1200 RPM) at RT for 30 minutes. Further, 10 mg of AIBN and 8 mg of SDS in water were added to the above reaction mixture, and this was kept under stirring (1200 RPM) at RT for 1 h. Then the reaction mixture was sonicated (Sonics (Vibra-cell), 750 watt, 45
1, w/w) was dispersed in 2 mL of toluene. Then, DVB and HD were added and bath sonicated (at RT) for 5 minutes followed by stirring (1200 RPM) at RT for 30 minutes. Further, 10 mg of AIBN and 8 mg of SDS in water were added to the above reaction mixture, and this was kept under stirring (1200 RPM) at RT for 1 h. Then the reaction mixture was sonicated (Sonics (Vibra-cell), 750 watt, 45![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 15 s, 40% power) for 10 minutes using an ultra-probe sonicator in an ice bath. The obtained reaction mixture was kept at 65–75 °C in an oil bath for polymerization (16–20 h) under continuous stirring (at 900 RPM). Further, to remove residual toluene, a sufficient amount of water was added, and the mixture was kept under vigorous stirring (1500 RPM) at 85–90 °C in an oil bath. The prepared NPs were washed thoroughly with an ethanol and water mixture (1
15 s, 40% power) for 10 minutes using an ultra-probe sonicator in an ice bath. The obtained reaction mixture was kept at 65–75 °C in an oil bath for polymerization (16–20 h) under continuous stirring (at 900 RPM). Further, to remove residual toluene, a sufficient amount of water was added, and the mixture was kept under vigorous stirring (1500 RPM) at 85–90 °C in an oil bath. The prepared NPs were washed thoroughly with an ethanol and water mixture (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) 8–10 times, followed by centrifugation (14
1) 8–10 times, followed by centrifugation (14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 RPM, 4 °C, 30 minutes) to remove unreacted monomers and SDS. Then, the samples were freeze-dried for 48 h and stored in a tight container for further studies (monomer conversion %: ∼90% and yield: ∼80%). Similarly, p(NAG) and p(NAPA) NPs were synthesized as control NPs. It can be noted that this modified method provides smaller-sized p(NAG-co-NAPA) NPs, which have potential uses for therapeutic purposes.
000 RPM, 4 °C, 30 minutes) to remove unreacted monomers and SDS. Then, the samples were freeze-dried for 48 h and stored in a tight container for further studies (monomer conversion %: ∼90% and yield: ∼80%). Similarly, p(NAG) and p(NAPA) NPs were synthesized as control NPs. It can be noted that this modified method provides smaller-sized p(NAG-co-NAPA) NPs, which have potential uses for therapeutic purposes.
2.2.3. Characterization of NAG, NAPA, p(NAG), p(NAPA), and p(NAG-co-NAPA) NPs
Analysis of the chemical functionality and structures of NAG, NAPA, p(NAG), p(NAPA), and p(NAG-co-NAPA) NPs was carried out using various spectroscopic techniques. Fourier-transform infrared (FT-IR) spectroscopy was performed with a Nicolet iS5 instrument (THERMO Electron Scientific Instruments LLC) utilizing the attenuated total reflectance (ATR) method. For nuclear magnetic resonance (NMR) spectroscopy, both 1H and 13C NMR studies were conducted using a 500 MHz OneBay NMR spectrometer (BRUKER BioSpin INTERNATIONAL AG). Chemical shifts were recorded in parts per million (ppm) relative to deuterated solvents such as CDCl3, DMSO-d6, and a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixture of CDCl3 and DMSO-d6 (co-solvency approach) for p(NAPA), p(NAG), and p(NAG-co-NAPA) NPs, respectively. Data analysis was performed using MestReNova v14.1.2-25024. Matrix-assisted laser desorption/ionization-time of flight (MALDI-ToF) mass spectra were acquired using a Bruker Autoflex instrument, employing dithranol in THF as the matrix, with the analysis method RN_900-4500_Da.par.
1 mixture of CDCl3 and DMSO-d6 (co-solvency approach) for p(NAPA), p(NAG), and p(NAG-co-NAPA) NPs, respectively. Data analysis was performed using MestReNova v14.1.2-25024. Matrix-assisted laser desorption/ionization-time of flight (MALDI-ToF) mass spectra were acquired using a Bruker Autoflex instrument, employing dithranol in THF as the matrix, with the analysis method RN_900-4500_Da.par.
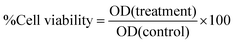 | (1) |
 | (2) |
In this study, the number of embryo (n) was 20 for each dose of NPs and, to ensure exclusivity of ROI for images from the same sample, a few essential steps are followed, such as: (1) unique IDs were assigned to each image; (2) grid mapping of each image was performed to trace origins and exclude duplicates from the same region; and (3) ROIs were limited per sample to 3–5 and random stratified selection was followed during analysis.
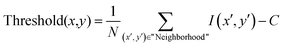 | (3) |
1. Number of pixels in the blood vessels: the total number of black pixels in the segmented image, which correspond to blood vessels.
2. Entropy: measures the randomness or complexity of the texture. An increase in blood vessel density and complexity typically leads to higher entropy, which can be represented as:
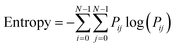 | (4) |
3. Energy: measures the textural uniformity or smoothness of the image. A higher blood vessel density may decrease the energy, indicating a less uniform and more complex texture. The energy can be represented as:
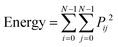 | (5) |
4. Contrast: measures the number of local variations in the image. An increase in contrast generally correlates with a higher density of blood vessels, and it can be represented as:
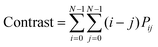 | (6) |
5. Mean: represents the average gray level in the GLCM. In binary images, the mean might not change significantly with an increase in the blood vessel density, but the spatial distribution might shift. The mean can be represented as:
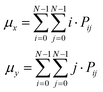 | (7) |
6. Dissimilarity: Higher dissimilarity means more variation between neighbouring pixel values. This can indicate more pronounced blood vessels and can be represented as:
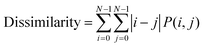 | (8) |
7. Variance: higher variance means greater intensity variation in the image, which could correspond to more varied blood vessels. The variance can be represented as:
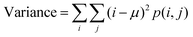 | (9) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2000) was added to each well, incubated for 15 minutes, and then observed using a fluorescence microscope (Evos FL Auto). Images were analyzed using Angiotool (ImageJ) and the GLCM textural-feature-based image processing tool.
2000) was added to each well, incubated for 15 minutes, and then observed using a fluorescence microscope (Evos FL Auto). Images were analyzed using Angiotool (ImageJ) and the GLCM textural-feature-based image processing tool.
3. Results and discussion
3.1. Synthesis and physicochemical characterization of monomers (NAG and NAPA) and model p(NAG-co-NAPA) NPs.
The p(NAG-co-NAPA) copolymer NPs were synthesized in two steps. The first step was the synthesis of the monomers, followed by the synthesis of copolymer NPs. The synthesis of the monomers NAG and NAPA is based on the Scotten–Baumann reaction. In this reaction, the amine group (–NH2) of the monomers reacts with acryloyl chloride in an alkaline medium to form an amide bond (–CONH) via nucleophilic substitution. In the final extraction step, ethyl acetate was used as an organic solvent to improve the percentage yield of monomers. The structures of the monomers were confirmed through 1H-NMR, 13C-NMR and FTIR analysis (Fig. S1–S3), and the obtained results match well with our earlier reports.7,10 In the second step, the synthesized monomers were used to prepare p(NAG-co-NAPA) NPs. The p(NAG-co-NAPA) NPs were achieved through a modified free radical mini-emulsion polymerization method.51 Here, a long-chain HD molecule was used as a co-stabilizer to enhance the stability of the droplets formed during the emulsion process and to prevent Ostwald ripening. The monomers were dispersed in the oil phase (toluene), resulting in the formation of monomer droplets. AIBN helped to initiate polymerization at 65–75 °C within the oil/monomer droplets and at the interface between the oil/monomer droplets and the aqueous phase. The core was stabilized by crosslinking the monomer droplets with DVB, which helps to generate pores and provides mechanical stability to the NPs. DVB is present both within the polymer chains and in between two polymer backbones. The NAG![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) NAPA ratio was maintained at 1
NAPA ratio was maintained at 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (w/w). SDS was used to prevent the coagulation of NPs. This mechanism of copolymer synthesis is straightforward. AIBN initiates the reaction by attacking hydrogen attached to terminal sp2-hybridized primary carbon of NAG, NAPA, and DVB, forming radical ions of both monomers and the cross-linker. These radicals propagate and crosslink the chains, eventually leading to termination when different growing chains meet head-to-head, forming a π bond. The p(NAG-co-NAPA) NPs synthesized here are amphiphilic in nature. The carboxylic acid group (-COOH) in NAG imparts hydrophilic properties to the entire system, while the phenyl rings present in NAPA contribute to the hydrophobic characteristics. This arrangement results in a cross-linked-network-like structure, with the phenyl rings of NAPA and DVB moving towards the core of the micelles and the carboxylic groups oriented towards the outer side. We speculate that this configuration of p(NAG-co-NAPA) may facilitate the formation of a dense core in the NPs, along with the formation of small pores on the shell. It can be noted that NAG, NAPA and DVB can be randomly arranged in the NP network. A probable structure of the copolymer NPs is given in the inset of Fig. 1.
1 (w/w). SDS was used to prevent the coagulation of NPs. This mechanism of copolymer synthesis is straightforward. AIBN initiates the reaction by attacking hydrogen attached to terminal sp2-hybridized primary carbon of NAG, NAPA, and DVB, forming radical ions of both monomers and the cross-linker. These radicals propagate and crosslink the chains, eventually leading to termination when different growing chains meet head-to-head, forming a π bond. The p(NAG-co-NAPA) NPs synthesized here are amphiphilic in nature. The carboxylic acid group (-COOH) in NAG imparts hydrophilic properties to the entire system, while the phenyl rings present in NAPA contribute to the hydrophobic characteristics. This arrangement results in a cross-linked-network-like structure, with the phenyl rings of NAPA and DVB moving towards the core of the micelles and the carboxylic groups oriented towards the outer side. We speculate that this configuration of p(NAG-co-NAPA) may facilitate the formation of a dense core in the NPs, along with the formation of small pores on the shell. It can be noted that NAG, NAPA and DVB can be randomly arranged in the NP network. A probable structure of the copolymer NPs is given in the inset of Fig. 1.
The chemical functionalities of p(NAG-co-NAPA) NPs were confirmed through 1H NMR, 13C NMR, FTIR and MALDI-ToF analysis. The positions and number of protons present in the p(NAG-co-NAPA) NPs were identified through 1H and 13C NMR. The characteristic bands were as follows: 1H-NMR: δ (ppm) = 7.16 (9H, m, aromatic protons), 4.1–3.2 (m, polymer backbone); all other bands from 4.5–2.5 ppm showed the presence of aliphatic protons and the absence of bands in between 5 and 6 ppm confirms the successful crosslinking of DVB with alkene by converting all the alkene protons to alkane protons. Further, as the cross-linked copolymer is sparingly soluble in NMR solvents, residual solvent bands were observed at 8.21 and 2.4 ppm from CDCl3 and DMSO-d6, respectively (Fig. 1). The main 13C solid-state NMR signals were as follows: δ (ppm) = 175.90 (aromatic) and 128.92 (–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O); all other bands around 50–30 ppm showed the presence of carbon present in saturated and unsaturated hydrocarbons (polymer backbone) (Fig. S4). For a comparison and confirmation of the copolymer structure, we have performed a 1H NMR study of copolymer NPs prepared without using the cross linker via the same method (Fig. S5), which are soluble in DMSO-d6. Fig. S5 shows the disappearance of bands in between 5 and 6 ppm, which further confirms the conversion of all alkene protons to alkane protons. Similar to Fig. 1, other than the 2.5 ppm band (DMSO-d6), bands at 2.5 to 4.5 ppm represent alkane protons and successful self-crosslinking. The FTIR spectrum revealed the following characteristic bands: ν (cm−1): 3413 (secondary –NH, s), 3013 (aromatic –CH, s), 2924 and 2846 (alkane –CH, s), 1739 (ester C
O); all other bands around 50–30 ppm showed the presence of carbon present in saturated and unsaturated hydrocarbons (polymer backbone) (Fig. S4). For a comparison and confirmation of the copolymer structure, we have performed a 1H NMR study of copolymer NPs prepared without using the cross linker via the same method (Fig. S5), which are soluble in DMSO-d6. Fig. S5 shows the disappearance of bands in between 5 and 6 ppm, which further confirms the conversion of all alkene protons to alkane protons. Similar to Fig. 1, other than the 2.5 ppm band (DMSO-d6), bands at 2.5 to 4.5 ppm represent alkane protons and successful self-crosslinking. The FTIR spectrum revealed the following characteristic bands: ν (cm−1): 3413 (secondary –NH, s), 3013 (aromatic –CH, s), 2924 and 2846 (alkane –CH, s), 1739 (ester C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O, s), 1659 (–CONH–, s), and 739 (aromatic –CH, b) (Fig. 2(a)). For comparison, the FTIR spectra of p(NAG) and p(NAPA) are also presented in Fig. 2(a), which demonstrate that the distinct bands of p(NAG) and p(NAPA) coexist in the FTIR spectrum of p(NAG-co-NAPA) NPs. MALDI-ToF analysis was performed using dithranol-THF as the matrix to find out the approximate molecular weight of the synthesized copolymer. Since the designed p(NAG-co-NAPA) NPs are a complex cross-linked network-like copolymer, the obtained molecular weight (
O, s), 1659 (–CONH–, s), and 739 (aromatic –CH, b) (Fig. 2(a)). For comparison, the FTIR spectra of p(NAG) and p(NAPA) are also presented in Fig. 2(a), which demonstrate that the distinct bands of p(NAG) and p(NAPA) coexist in the FTIR spectrum of p(NAG-co-NAPA) NPs. MALDI-ToF analysis was performed using dithranol-THF as the matrix to find out the approximate molecular weight of the synthesized copolymer. Since the designed p(NAG-co-NAPA) NPs are a complex cross-linked network-like copolymer, the obtained molecular weight (![[M with combining macron]](https://www.rsc.org/images/entities/i_char_004d_0304.gif) w) represents the chain length between two adjacent crosslinking sites. The heterogeneous fragments in the MALDI-ToF spectrum are indicative of the formation of a random cross-linked block copolymer with
w) represents the chain length between two adjacent crosslinking sites. The heterogeneous fragments in the MALDI-ToF spectrum are indicative of the formation of a random cross-linked block copolymer with ![[M with combining macron]](https://www.rsc.org/images/entities/i_char_004d_0304.gif) w ranging from 1251 to 2510 Da (Fig. S6). The maximum intensity for the p(NAG-co-NAPA) NPs is observed at 1357 Da, with repeated units varying from 100 to 250 Da. MALDI-ToF analysis revealed two prominent fragments: the first at
w ranging from 1251 to 2510 Da (Fig. S6). The maximum intensity for the p(NAG-co-NAPA) NPs is observed at 1357 Da, with repeated units varying from 100 to 250 Da. MALDI-ToF analysis revealed two prominent fragments: the first at ![[M with combining macron]](https://www.rsc.org/images/entities/i_char_004d_0304.gif) w = 1419 Da and
w = 1419 Da and ![[M with combining macron]](https://www.rsc.org/images/entities/i_char_004d_0304.gif) n = 1357 Da, with a polydispersity index (
n = 1357 Da, with a polydispersity index (![[M with combining macron]](https://www.rsc.org/images/entities/i_char_004d_0304.gif) w/
w/![[M with combining macron]](https://www.rsc.org/images/entities/i_char_004d_0304.gif) n) (PDI) of ∼ 1.04; and the second at
n) (PDI) of ∼ 1.04; and the second at ![[M with combining macron]](https://www.rsc.org/images/entities/i_char_004d_0304.gif) w = 1655 Da and
w = 1655 Da and ![[M with combining macron]](https://www.rsc.org/images/entities/i_char_004d_0304.gif) n = 1595 Da, with PDI of around 1.03. These chemical functionality-based observations confirm the successful synthesis of p(NAG-co-NAPA) NPs.
n = 1595 Da, with PDI of around 1.03. These chemical functionality-based observations confirm the successful synthesis of p(NAG-co-NAPA) NPs.
Further, the shape and size of p(NAG-co-NAPA) NPs were investigated using FESEM, HR-TEM and AFM analysis (Fig. 2(b and c) and Fig. S7). FESEM micrographs (Fig. 2(b)) confirmed that the p(NAG-co-NAPA) NPs are nano-sized and spherical in nature. As discussed earlier, the dark core and porous shell-like structure observed in Fig. 2(c) is attributed to the presence of hydrophilic carboxylic groups on the outer surface of the particles, forming a corona, while the hydrophobic phenyl rings from the NAPA monomer and DVB are located inside, contributing to the dense core. The average particle size distribution (Fig. 2(e)) calculated from the FESEM image shown in Fig. 2(b) indicates a high number of p(NAG-co-NAPA) NPs with diameters ranging from 100 nm to 120 nm, while the HR-TEM image in Fig. 2(c) shows particles with diameters ranging from 120 nm to 140 nm (Fig. 2(f)). Further, to find out the optical and chiral properties of p(NAG-co-NAPA) NPs, CD spectroscopy was performed in MilliQ water at RT. p(NAPA) exhibited a distinct absorption band at λmax = 233 nm, which is characteristic of the optically active NAPA monomer (Fig. 2(d)). In contrast, p(NAG) did not show any significant λmax band. However, for p(NAG-co-NAPA) NPs, a broad band was observed at λmax = 222 nm (Fig. 2(d)).60,61 The shift in λmax for p(NAG-co-NAPA) NPs towards a shorter wavelength is attributed to the contribution of the DVB molecules present in the NPs. This behavior demonstrated the chiral property of the p(NAG-co-NAPA) NPs, which can provide insights into NP-drug interactions, the design of specific drugs or vaccines, and the stability, shelf-life, and efficacy of NPs. Further, to assess the roughness of p(NAG-co-NAPA) NPs, 2D and 3D AFM topography images were acquired, and they revealed that the particles have a similar size range and the chains are folded, creating roughness (Fig. S7(a) and (b)), which may enhance protein binding, cellular internalization, and overall cellular delivery performance.62 The images from all three microscopic techniques, FESEM (Fig. 2(b)), HR-TEM (Fig. 2(c) and Fig. S7(c) and (d)), and AFM (Fig. S7(a)), demonstrated that the model p(NAG-co-NAPA) NPs are spherical and porous in nature. The colloidal stability of p(NAG-co-NAPA) NPs was evaluated through zeta potential (ζ) measurements, and size was analyzed through dynamic light scattering (DLS) measurements in MilliQ water at RT (Fig. 2(g) and (h)). The ζ value of p(NAG-co-NAPA) NPs was found to be −44.7 mV, indicating good colloidal stability (Fig. 2(g)). The hydrodynamic diameter obtained from DLS is ∼339.6 nm with a PDI of 0.431 (Fig. 2(h)). This increase in the size of p(NAG-co-NAPA) NPs in comparison to the microscopic results can be attributed to entrapment and swelling. A full size-distribution plot for p(NAG-co-NAPA) NPs is shown in Fig. S8.
3.2. Biocompatibility and hemocompatibility of p(NAG-co-NAPA) NPs
For biomedical applications, copolymer NPs must demonstrate good biocompatibility and hemocompatibility. Cellular viability and the proliferation of normal cells are crucial parameters. At first, the cell viability properties of p(NAG) and p(NAPA) have been checked against three normal cell lines, L929, RAW 264.7, and HUVEC cells, with PBS as a control, at concentrations of 3.125 to 400 µg mL−1 and 5 to 200 µg mL−1 (Fig. 3(a) and (c)). It is observed that p(NAPA) NPs are toxic to the L929 cells at higher concentrations, whereas at lower concentrations they are not toxic (viability is ∼100%). For p(NAG) NPs, at higher concentrations the viability is ∼100%, whereas at lower concentrations (50 to 3.125 µg mL−1) they are proliferative in nature (∼120% viability at 12.5 µg mL−1) (Fig. 3(a)). In the case of RAW 264.7, other than at 400 µg mL−1 p(NAG) NPs, for all other concentrations, both the homopolymers showed cell viability up to ∼80% (Fig. 3(b)). Furthermore, it is observed that p(NAPA) NPs are cytotoxic against HUVEC cells (70% at 5 µg mL−1), whereas p(NAG) NPs are proliferative in nature (126% at 200 µg mL−1) (Fig. 3(c)). Based on the MTT results, cell viability has been further checked for p(NAG-co-NAPA) NPs with the same cell lines (Fig. 3(d) and (e)). In all three cases, the cell viability (%) is observed to be ∼100% or above 100%. Interestingly, p(NAPA-co-NAG) NPs show enhanced metabolic activity toward L929 and RAW 264.7 cells at 100 and 50 µg mL−1 (Fig. 3(d)) and below 100 µg mL−1 for HUVEC cells (Fig. 3(e)). The decrease in cell viability (%) below 50 µg mL−1 (Fig. 3(d)) is due to contact inhibition experienced by cells in the confined space over 24 h. The exact cell viability (%) values for p(NAG-co-NAPA) NPs toward L929, RAW 264.7 and HUVEC cells are listed in Tables S1 and S2 relative to the control group. These studies clearly indicate that p(NAG-co-NAPA) NPs are biocompatible with L929, RAW 264.7, and HUVEC cells, demonstrating the proliferative nature of the NPs at lower concentrations and warranting their possible use in vivo.Hemocompatibility evaluation is essential for biomaterials designed for therapeutic purposes. For a copolymer to be considered as an effective biomaterial, the hemolysis % must remain below 5%. The hemolytic activity of p(NAG-co-NAPA) NPs was assessed in a dose- and time-dependent manner at concentrations of 500 to 7.8125 µg mL−1 (Fig. 3(f)). Distilled water was used as the positive control, while a pure RBC suspension acted as the negative control. A varied degree of hemolysis was observed at two different time points. The exact hemolysis % values at 2 h and 8 h are listed in Table S3. In conclusion, the results indicate that p(NAG-co-NAPA) NPs can be regarded as a safe biomaterial concerning hemocompatibility, demonstrating suitability for therapeutic applications at concentrations up to 500 µg mL−1.
3.3. Angiogenic effects of p(NAG-co-NAPA) NPs studied through the in ovo CAM assay
Angiogenic effects of p(NAG), p(NAPA) and p(NAG-co-NAPA) NPs were assessed through the image processing tool in both dose- and time-dependent manners. The selected doses and times were 1, 10 and 100 µg and 0, 2, 4, 8 and 24 h, respectively, considering PBS as a control. The selection of doses is based on the results obtained from the MTT assay (Fig. 3). Fig. 4(a)–(c) shows the raw images acquired using an optical microscope. It is clear that the images consist of blood vessels. The respective segmented images are shown in Fig. 4(d) and (e). It can be observed that vascularization and sprouting are found more widely in the segmented images. It is clearly evident that the number of blood vessels varies upon varying the dose and time.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600
600![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 to ∼1
000 to ∼1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 888
888![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 is observed at 24 h (Fig. 5(c)). This change in black pixel count demonstrates the pro-angiogenic behavior of p(NAG-co-NAPA) NPs.
000 is observed at 24 h (Fig. 5(c)). This change in black pixel count demonstrates the pro-angiogenic behavior of p(NAG-co-NAPA) NPs.
In the case of entropy, an increasing trend is observed over the time period of 0 to 24 h at the three different concentrations (Fig. 5(d)–(f)). Entropy means a change in the randomness of the system, where an increase in randomness implies that more vessels are formed. It is evident that the entropy is high at 1-, 10- and 100-µg doses compared to the control for the homopolymer NPs. Therefore, the change in vascular patterns may lead to angiogenesis. However, the entropy varies for the homopolymer and copolymer NPs. In the case of p(NAG), the entropy lies above the control group, whereas for p(NAPA), the entropy values for all three doses are close to the control sample (Fig. 5(d) and (e)). However, in the case of p(NAG-co-NAPA) NPs, at 100 µg the entropy lies below the control sample, while for 1- and 10-µg doses, the entropy is above the control value. At a 10-µg dose, at 24 h, a sudden rise in entropy is observed from ∼10.5 to ∼13.5 (Fig. 5(f)). These results further depict the proangiogenic nature of the copolymer NPs. For energy, a decreasing trend is observed for all doses of homopolymer NPs over the entire time period (Fig. 5(g) and (h)). It can also be noted that for p(NAG) NPs at 10- and 100-µg doses, a sudden change in slope is observed (Fig. 5(g)), which demonstrates the prominent angiogenic nature of the p(NAG) NPs compared to the p(NAPA) homopolymer. For copolymer NPs, a quick fall in energy value from 0.86 (8 h) to 0.82 (24 h) is observed at a 10-µg dose. (Fig. 5(i)).
Fig. 6 shows two more textural features, mean and contrast, which are computed to gain a deeper understanding of the angiogenic behavior of homopolymer and copolymer NPs. According to the definition, mean is inversely proportional and contrast is directly proportional to blood-vessel development. In the case of mean, for p(NAG) NPs, at 10- and 100-µg doses a negative change in slope is observed, whereas for p(NAPA), while changes in slope are noticed, a prominent change in mean is, however, not observed. For p(NAG-co-NAPA) NPs, the mean value decreases from 223 to 219 for a 10-µg dose at 24 h (Fig. 6(c)). Further, in the case of contrast, all three considered NPs at the studied doses show an increasing trend up to 24 h, however the values obtained are lower compared to the control sample (Fig. 6(d)–(f)). For p(NAG-co-NAPA) at a 10-µg dose, a sharp rise in contrast value is observed from 8 h to 24 h, i.e. from 2859.2 to 3800.6 (Fig. 6(f)), which further confirms that a 10-µg dose of copolymer NPs can be an optimum dose for promoting angiogenesis. Furthermore, it can be stated that mean and contrast corroborate well with the results obtained for the number of pixels in blood vessels, entropy, and energy discussed above (Fig. 5).
Along with the above-mentioned five textural parameters, two more parameters, dissimilarity and variance, are computed and represented in Fig. 7(a)–(c) and (d)–(f), respectively. The positive change in slope for both parameters represents the angiogenic nature of the NPs. In the case of dissimilarity, for p(NAG) at all three concentrations, increases in dissimilarity values are observed, whereas for p(NAPA) all doses show negligible variation with respect to the control (Fig. 7(a) and (b)). However, for p(NAG-co-NAPA) NPs, both 1-µg and 10-µg doses show sharp increases in the dissimilarity index (Fig. 7(c)). Similarly, with varying time and dose, increases in variance values are observed for all three NPs (Fig. 7(d)–(f)). The variance of an image refers to a statistical measure that quantifies the dispersion or spread of pixel intensity values within an image. In particular, for p(NAG-co-NAPA) NPs at a 10-µg dose variance increases from ∼8000 to ∼9400, while for p(NAPA) and p(NAG) the values are 8816.9 and 7295.4, respectively, at 24 h. It can be concluded that both the dissimilarity and variance results for p(NAG-co-NAPA) NPs match well with the above five textural parameters, demonstrating the proangiogenic nature of the copolymer NPs at a 10-µg dose. Furthermore, it can be highlighted that the textural features obtained from in ovo models fit well with the in vitro cell-based results for the copolymer NPs.
3.5. Validation of GLCM textural-feature-based image processing results with Angiotool through the tube formation assay
Further, to check the accuracy of the GLCM textural-feature-based image processing tool for angiogenesis studies, the tube formation assay is conducted in HUVEC cells, considering p(NAG-co-NAPA) NPs as a model polymer at the same three concentrations (1, 10, and 100 µg) of NPs; subsequently, microscopic images were acquired (Fig. 8(a–h)). HUVECs are considered as a standard model for angiogenesis studies due to their reliable tube formation and response to VEGF, mimicking vascular development processes like migration and sprouting. Their well-characterized nature ensures the reproducible validation of the GLCM method for assessing copolymer NP effects on endothelial behavior.63,64 No treatment with NPs is considered as the control group. It is clearly evident that, compared to the control, for a 1-µg dose, fewer tubes are formed (Fig. 8(e) and (f)), whereas for a 10-µg dose, defined and thick network or mesh-like structures are formed (Fig. 8(e) and (g)). For a 100-µg dose, the p(NAG-co-NAPA) NPs are less angiogenic or toxic toward the cells, where disconnected patchy dead cell structures or thin tubes are observed (Fig. 8(d) and (h)). Additionally, quantitative analysis is performed using two methods: (1) Angiotool with ImageJ (conventional) and (2) the GLCM textural-feature-based image processing tool. From Angiotool data, it is found that for a 10-µg dose, the explant area (116.78 vs. 100.69 and 78.45%), total vessel length (140.73 vs. 126.86 and 78.83%), and total number of junctions (255.55 vs. 222.54 and 142.15%) increase with respect to 1- and 100-µg doses, whereas the mean-e-lacunarity (70.80 vs. 85.07 and 99.01%), or empty space, is less (Fig. 8(i)–(l)). Similarly, the quantitative data obtained from the image processing tool demonstrates that the number of pixels in blood vessels and entropy are highest for the 10-µg dose, whereas energy and mean are lowest, as compared to 1- and 100-µg doses. From these results it can be concluded that a 10-µg dose of p(NAG-co-NAPA) NPs is optimum for promoting angiogenesis, which corroborates well with results obtained from previous experiments. Thus, the image processing tool with GLCM textural features is an alternative solution for the angiogenesis analysis of microscopic images, which can eliminate the shortcomings associated with Angiotool and/or other reported approaches.In this work, p(NAG-co-NAPA) NPs have been successfully synthesized through mini-emulsion polymerization as a model polymer to develop a standard tool for assessing angiogenesis. Angiogenesis, the process of new blood-vessel formation is associated with the treatment of various diseases, and it can be controlled by the addition of biopolymers such as p(NAG), p(NAPA) and p(NAG-co-NAPA) NPs.4 Anti-angiogenic molecules are used for treating cancers and eye diseases, while polymers for neovascularization promote wound-healing efficiency. Additionally, the stimulation of angiogenesis is beneficial for stroke recovery. As a result, the need for analysing angiogenesis has gained significant momentum in the biomedical field and health care technology.65,66 The accurate assessment of angiogenesis relative to dose and time could solve issues related to drug discovery and regenerative medicine. However, the manual quantification of the 3D morphology of reconstituted vasculature using imaging software such as ImageJ, CellProfiler or Angiotool has different limitations, such as the need for manual analysis processes, uneven illumination, noise robustness and loss in fine structures during segmentation, etc.16,20,67 Furthermore, the subjective opinions of analysts may introduce bias into the results.68 Therefore, the present study shows an important direction for dose- and time-dependent treatment by using polymeric nanomaterials as a model system for analysing angiogenesis properties through the GLCM-based textural analysis approach.55,56
After confirmation of particle size, shape, colloidal stability and structure, biocompatibility and hemocompatibility were studied (Fig. 1 and 2). The biocompatibility (∼120%) against three normal cell lines, including HUVEC, (Fig. 3(d) and (e)) and hemocompatibility (<5%) (Fig. 4(f)) results for the copolymer revealed that the synthesized p(NAG-co-NAPA) NPs could have potential for use in therapeutic applications. The cell viability results against L929, RAW 264.7 and HUVEC cells revealed the proliferative nature of p(NAG-co-NAPA) NPs. Further, to achieve the main aim of this work, the CAM assay and tube formation assay have been performed using in ovo and in vitro models, followed by data analysis with image processing tools (an adaptive threshold algorithm with GLCM features) and the conventional Angiotool approach (Fig. 5–8).
By using GLCM texture-based analysis, seven different features have been extracted for p(NAG), p(NAPA) and p(NAG-co-NAPA) NPs and analyzed in a dose- and time-dependent manner. The features studied here are the number of pixels in blood vessels, entropy, energy, mean, contrast, dissimilarity and variance. All the features clearly demonstrate the angiogenic behavior of all three synthesized polymer NPs at the studied doses (Fig. 5–7). However, angiogenesis depends on the concentration of p(NAG), p(NAPA) and p(NAG-co-NAPA) NPs used during treatment and on the period of treatment (Fig. 5–7). To compare and validate the results obtained from the image processing tool, the tube formation assay is also conducted in HUVEC cells, considering p(NAG-co-NAPA) NPs as a model polymer (Fig. 8). Interestingly, it is found that both analysis methods, i.e., Angiotool and the image processing tool, demonstrated similar results, with a 10-µg dose of the model polymer being an optimum dose for promoting angiogenesis via inducing changes in vascular patterns. This further matches well with the textural-feature-based analysis of angiogenesis images obtained from in ovo studies. Additionally, a box chart is presented for better comparison and understanding of the optimal dose for treatment and biomedical applications of the synthesized polymer NPs (Fig. 9). Thus, it can be stated that the image processing tool can be considered as a preliminary framework for the quantitative assessment of angiogenesis and can help to decide a suitable therapeutic dose for specific biomedical applications. Further, it can be concluded that compared to homopolymers, copolymer NPs possess better angiogenic properties and hold potential for various therapeutic applications, including wound management.
4. Conclusions
Despite significant advancements in the development of angiogenesis assessment tools, a fully reliable and automated quantification method remains elusive. In this light, the present work has developed an angiogenesis evaluation tool based on GLCM features that is straightforward and physiologically relevant. To establish this method, amino acid-based copolymer p(NAG-co-NAPA) NPs have been synthesized as a model polymer. As anticipated, hemolysis and in vitro viability assays with various cells, including HUVECs, revealed the biocompatible and proliferative nature of the NPs. The dose- and time-dependent angiogenic properties of p(NAG-co-NAPA) NPs have been computed from microscopic images obtained from an in ovo model and the tube formation assay using a GLCM textural-feature-based image processing tool. For comparison, the tube formation results are cross-verified with Angiotool analysis. Both tools depict that a 10-µg dose of p(NAG-co-NAPA) NPs can be an optimum dose for promoting angiogenesis. These findings suggest that the application of the image processing tool in a wet lab provides an unbiased and high-throughput analysis of various texture-based features from 2D microscopic images compared with existing conventional tools. However, to comprehend the efficacy of p(NAG-co-NAPA) NPs and to develop the GLCM textural-feature-based image processing tool as a preliminary quantitative framework for the analysis of angiogenesis, in vivo studies are being taken under consideration for the future scope of this work.Author contributions
P. P. is the principal investigator (PI) of this project. Ideation, experimental designs, result analysis, and manuscript writing were mainly carried out by S. P. and P. P. In ovo angiogenesis and its analysis through the image processing tool were assisted by A. D. L., G. S., and J. A. F. R. MTT and hemolysis studies were assisted by D. P. S. J. and A. M. helped in carrying out the tube formation assay. The manuscript was written and finalized by S. P. and P. P. All the authors checked and approved the final version for publication.Conflicts of interest
The authors have no conflicts of interest to declare.Data availability
To implement image segmentation and textural-feature extraction, several resources were used. The foundational concepts of image processing, including image normalization, smoothing, and blurring, were studied viahttps://www.geeksforgeeks.org/image-processing-in-python/, which also provided an overview of Python libraries used for image processing. The process of converting RGB images into binary images was implemented using guidelines from https://www.geeksforgeeks.org/convert-image-to-binary-using-python/ and https://docs.opencv.org/4.x/d7/d4d/tutorial_py_thresholding.html, which detailed the use of OpenCV for thresholding operations. Gaussian adaptive thresholding, a crucial step for precise segmentation, was implemented with the help of https://plantcv.readthedocs.io/en/stable/gaussian_threshold/. For the analysis of the tube formation assay, the code used can be found at https://docs.google.com/document/d/1rFYpgmM1jrImflz7sbFEBu4IjM3J1nnBYX7bkMdY61s/edit?tab=t.0. Additional information on GLCM-based textural features, including contrast, correlation, energy, and homogeneity, was sourced from https://support.echoview.com/WebHelp/Windows_And_Dialog_Boxes/Dialog_Boxes/Variable_Properties_Dialog_Box/Operator_Pages/GLCM_Texture_Features.html and https://scikit-image.org/docs/stable/auto_examples/features_detection/plot_glcm.html.Supplementary information (SI): synthesis of NAG and NAPA monomers; 1H and 13C NMR of NAG and NAPA monomers; FTIR spectra of NAG and NAPA monomers; 13C solid state NMR of p(NAG-co-NAPA) NPs; 1H NMR of p(NAG-co-NAPA)wc NPs; MALDI-ToF of p(NAG-co-NAPA) NPs; AFM and HR-TEM images of p(NAG-co-NAPA) NPs; DLS spectra of p(NAG-co-NAPA) NPs; Cell viability % for p(NAG-co-NAPA) NPs with L929, RAW 264.7 and HUVECS; Hemocompatibility % for p(NAG-co-NAPA) NPs; all the error % calculated from cell viability, GLCM based texture feature analysis of angiogenesis and tube formation assay. See DOI: https://doi.org/10.1039/d5ma01126d.
Acknowledgements
The authors acknowledge financial support awarded to Prof. Paik by I-DAPT (File no. I-DAPT/IT (BHU)/2023-24/Project Sanction/47), Indian Council of Medical Research (ICMR), India (Ref: EMDR/SG/12/2023-4724), STARS-IISc Bangalore (Ref. MoE-STARS/STARS-2/2023-0318) and Anusandhan National Research Foundation (ANRF), India (Ref: CRG/2023/005576). Sukanya acknowledges the award of a DST-INSPIRE fellowship (IF180185) and financial support from ICMR to carry out the PhD work. Further, the authors acknowledge Prof. Pradip Maity (CSIR-NCL, Pune) for NMR data analysis and Prof. Yitzhak Mastai (Bar-Ilan University, Israel) for providing the solid-state NMR facility.References
- R. K. Jain, Nat. Med., 2003, 9, 685–693 CrossRef CAS PubMed.
- R. A. Wimmer, A. Leopoldi, M. Aichinger, D. Kerjaschki and J. M. Penninger, Nat. Protoc., 2019, 14, 3082–3100 CrossRef CAS PubMed.
- J.-H. Lee, P. Parthiban, G.-Z. Jin, J. C. Knowles and H.-W. Kim, Prog. Mater. Sci., 2021, 117, 100732 Search PubMed.
- B. A. Nsiah, E. M. Moore, L. C. Roudsari, N. K. Virdone and J. L. West, in Biosynthetic Polymers for Medical Applications, ed. L. Poole-Warren, P. Martens and R. Green, Woodhead Publishing, 2016, pp. 189–203 DOI:10.1016/B978-1-78242-105-4.00008-0.
- P. Carmeliet, Nature, 2005, 438, 932–936 CrossRef CAS PubMed.
- S. K. Smith, Br. J. Obstet. Gynaecol., 2001, 108, 777–783 Search PubMed.
- P. S. Gupta, K. Wasnik, G. Singh, S. Patra, D. Pareek, D. D. Yadav, M. S. Tomar, S. Maiti, M. Singh and P. Paik, Mater. Adv., 2023, 4, 4718–4731 RSC.
- K. Wasnik, P. S. Gupta, G. Singh, S. Maity, S. Patra, D. Pareek, S. Kumar, V. Rai, R. Prakash, A. Acharya, P. Maiti, S. Mukherjee, Y. Mastai and P. Paik, J. Mater. Chem. B, 2024, 12, 6221–6241 Search PubMed.
- P. S. Gupta, K. Wasnik, S. Patra, D. Pareek, G. Singh, D. D. Yadav, S. Maity and P. Paik, Nanoscale, 2024, 16, 1770–1791 RSC.
- K. Wasnik, P. S. Gupta, S. Mukherjee, A. Oviya, R. Prakash, D. Pareek, S. Patra, S. Maity, V. Rai, M. Singh, G. Singh, D. D. Yadav, S. Das, P. Maiti and P. Paik, ACS Appl. Bio Mater., 2023, 6, 5644–5661 CrossRef CAS PubMed.
- M. J. A. S. R. C. Adair TH, Overview of Angiogenesis, 2010 Search PubMed.
- Z. Liu, L. Liu, S. Weng, C. Guo, Q. Dang, H. Xu, L. Wang, T. Lu, Y. Zhang and Z. Sun, Nat. Commun., 2022, 13, 816 Search PubMed.
- Y. A. Nanehkaran, Z. Licai, M. Azarafza, S. Talaei, X. Jinxia, J. Chen and R. Derakhshani, Sci. Rep., 2023, 13, 4126 Search PubMed.
- H. Du, H. Wang, Y. Chen and X. Zhou, Sci. Rep., 2025, 15, 19126 Search PubMed.
- T. M. Vergroesen, V. Vermeulen and R. M. H. Merks, PLoS Comput. Biol., 2025, 21, e1012965 CrossRef CAS PubMed.
- H. S. Rahman, B. L. Tan, H. H. Othman, M. S. Chartrand, Y. Pathak, S. Mohan, R. Abdullah and N. B. Alitheen, BioMed Res. Int., 2020, 2020, 8857428 CrossRef PubMed.
- G. Carpentier, S. Berndt, S. Ferratge, W. Rasband, M. Cuendet, G. Uzan and P. Albanese, Sci. Rep., 2020, 10, 11568 CrossRef CAS PubMed.
- J. Aschauer, M. Klimek, R. Donner, J. Lammer, P. Roberts, M. Schranz and G. Schmidinger, Sci. Rep., 2024, 14, 2124 CrossRef CAS PubMed.
- H. Yanagisawa, M. Sugimoto and T. Miyashita, Sci. Rep., 2021, 11, 419 CrossRef CAS PubMed.
- M. Pereira, J. Pinto, B. Arteaga, A. Guerra, R. N. Jorge, F. J. Monteiro and C. L. Salgado, Int. J. Mol. Sci., 2023, 24, 17625 Search PubMed.
- M. F. Gutknecht, M. E. Seaman, B. Ning, D. A. Cornejo, E. Mugler, P. F. Antkowiak, C. A. Moskaluk, S. Hu, F. H. Epstein and K. A. Kelly, Nat. Commun., 2017, 8, 552 Search PubMed.
- B. A. Corliss, R. W. Doty, C. Mathews, P. A. Yates, T. Zhang and S. M. Peirce, Microcirculation, 2020, 27, e12618 Search PubMed.
- L. Faihs, B. Firouz, P. Slezak, C. Slezak, M. Weißensteiner, T. Ebner, N. Ghaffari Tabrizi-Wizsy, K. Schicho and P. Dungel, Cancers, 2022, 14, 4273 CrossRef PubMed.
- M. S. Hossain, G. M. Shahriar, M. M. M. Syeed, M. F. Uddin, M. Hasan, S. Shivam and S. Advani, Sci. Rep., 2023, 13, 11314 CrossRef CAS PubMed.
- D.-H. Choi, H.-W. Liu, Y. H. Jung, J. Ahn, J.-A. Kim, D. Oh, Y. Jeong, M. Kim, H. Yoon, B. Kang, E. Hong, E. Song and S. Chung, Lab Chip, 2023, 23, 475–484 RSC.
- G. Okoro, P. Wityk, M. B. Nelappana, K. A. Jackiewicz, V. Z. Kucharczyk, A. Tigranyan, C. C. Applegate, I. T. Dobrucki and L. W. Dobrucki, BioData Min., 2025, 18, 69 CrossRef PubMed.
- E. M. Er, M. P. P. González, B. G. Garrido, J. Ho, H.-W. Kim, J. C. Knowles, J. Hua, F. Lali and P. Sawadkar, Adv. Funct. Mater., 2025, 35, e03722 CrossRef CAS.
- J. Cervantes, J. Cervantes, F. García-Lamont, A. Yee-Rendon, J. E. Cabrera and L. D. Jalili, Neurocomputing, 2023, 556, 126626 Search PubMed.
- Y. Tian, Q. Chen, W. Wang, Y. Peng, Q. Wang, F. Duan, Z. Wu and M. Zhou, BioMed Res. Int., 2014, 2014, 106490 Search PubMed.
- W. Wiharto and E. Suryani, Acta Inform. Med., 2020, 28, 42 Search PubMed.
- L. Armi and S. Fekri-Ershad, arXiv, preprint, arXiv:1904.06554, 2019 DOI:10.48550/arXiv.1904.06554.
- S. Alibabaei, M. Rahmani, M. Tahmasbi, M. J. Tahmasebi Birgani and S. Razmjoo, J. Med. Signals Sens., 2023, 13, 261–271 Search PubMed.
- Y. G. Shin, J. Yoo, H. J. Kwon, J. H. Hong, H. S. Lee, J. H. Yoon, E.-K. Kim, H. J. Moon, K. Han and J. Y. Kwak, Comput. Biol. Med., 2016, 75, 257–266 CrossRef PubMed.
- C. Madan, A. Satia, S. Basu, P. Gupta, U. Dutta and C. Arora, Medical Image Computing and Computer Assisted Intervention, 2025, pp. 207–217 Search PubMed.
- S. M. Peirce, Microcirculation, 2008, 15, 739–751 Search PubMed.
- X. Hao, W. Gai, Y. Zhang, D. Zhao, W. Zhou and Y. Feng, Colloids Surf., B, 2024, 241, 114020 CrossRef CAS PubMed.
- X. Hao, W. Gai, F. Ji, J. Zhao, D. Sun, F. Yang, H. Jiang and Y. Feng, Acta Biomater., 2022, 142, 221–241 CrossRef CAS PubMed.
- Q. Xu, C. Li, X. Meng, X. Duo and Y. Feng, Regen. Biomater., 2024, 11 Search PubMed.
- V. Rai, R. Moellmer and D. K. Agrawal, Cells, 2022, 11, 2287 CrossRef CAS PubMed.
- V. Mastrullo, W. Cathery, E. Velliou, P. Madeddu and P. Campagnolo, Front. Bioeng. Biotechnol., 2020, 8, 188 Search PubMed.
- J. Li, J. Wei, A. Li, H. Liu, J. Sun and H. Qiao, Front. Bioeng. Biotechnol., 2021, 9, 657699 Search PubMed.
- M. Li, Y. Wu and L. Ye, Cells, 2022, 11, 1372 CrossRef CAS PubMed.
- R. E. Oberkersch and M. M. Santoro, Vasc. Pharmacol., 2019, 112, 17–23 CrossRef CAS PubMed.
- Z. Wang, C. Cheng, X. Yang and C. Zhang, PLoS One, 2021, 16, e0250126 Search PubMed.
- L. Heikal, A. Starr, D. Hussein, J. Prieto-Lloret, P. Aaronson, L. A. Dailey and M. Nandi, Pain Genet., 2018, 3, 366–377 Search PubMed.
- R. Tan, J. Li, F. Liu, P. Liao, M. Ruiz, J. Dupuis, L. Zhu and Q. Hu, Am. J. Physiol.: Lung Cell. Mol. Physiol., 2020, 319, L1010–L1020 Search PubMed.
- P. Welsh, N. Rankin, Q. Li, P. B. Mark, P. Würtz, M. Ala-Korpela, M. Marre, N. Poulter, P. Hamet, J. Chalmers, M. Woodward and N. Sattar, Diabetologia, 2018, 61, 1581–1591 Search PubMed.
- K. Amin, J. Li, W. R. Chao, M. W. Dewhirst and Z. A. Haroon, Cancer Biol. Ther., 2003, 2, 173–178 CrossRef CAS PubMed.
- M. L. Rose, J. Madren, H. Bunzendahl and R. G. Thurman, Carcinogenesis, 1999, 20, 793–798 CrossRef CAS PubMed.
- D. Guo, C. E. Murdoch, H. Xu, H. Shi, D. D. Duan, A. Ahmed and Y. Gu, Sci. Rep., 2017, 7, 14749 CrossRef PubMed.
- A. K. Yamala, V. Nadella, Y. Mastai, H. Prakash and P. Paik, Nanoscale, 2017, 9, 14006–14014 RSC.
- S. Patra, J. Jyotirmayee, K. Kumar, D. Pareek, P. S. Gupta, A. R. Mourya, T. Das, K. Wasnik, M. Verma, R. Chawla, T. Batra and P. Paik, J. Mater. Chem. B, 2025, 13, 3876–3894 RSC.
- A. Bentolila, I. Vlodavsky, R. Ishai-Michaeli, O. Kovalchuk, C. Haloun and A. J. Domb, J. Med. Chem., 2000, 43, 2591–2600 CrossRef CAS PubMed.
- J. Rogowska, in Handbook of Medical Imaging, ed. I. N. Bankman, Academic Press, San Diego, 2000, pp. 69–85 DOI:10.1016/B978-012077790-7/50009-6.
- P. Mohanaiah, P. Sathyanarayana and L. GuruKumar, Int. J. Sci. Res. Publ., 2013, 3, 1–5 Search PubMed.
- L. Nanni, S. Brahnam, S. Ghidoni, E. Menegatti and T. Barrier, PLoS One, 2013, 8, e83554 CrossRef PubMed.
- L. K. Ramasamy, S. G. Padinjappurathu, S. Kadry and R. Damaševičius, PeerJ Comput. Sci., 2021, 7, e456 CrossRef PubMed.
- E. Dhiravidachelvi, V. Rajamani and C. Manimegalai, Int. J. Adv. Intell. Paradig., 2019, 14, 55–69 Search PubMed.
- R. M. Haralick, K. Shanmugam and I. Dinstein, IEEE Trans. Syst. Man Cybern., 1973, SMC-3, 610–621 Search PubMed.
- L. Werber, L. C. Preiss, K. Landfester, R. Muñoz-Espí and Y. Mastai, Chirality, 2015, 27, 613–618 CrossRef CAS PubMed.
- N. Amdursky and M. M. Stevens, ChemPhysChem, 2015, 16, 2768–2774 CrossRef CAS PubMed.
- X. Wang, X. Fang, X. Liu, Y. Lyu, L. Ma and J. Fu, Next Mater., 2024, 2, 100080 CrossRef.
- A. W. Tan, L. L. Liau, K. H. Chua, R. Ahmad, S. A. Akbar and B. Pingguan-Murphy, Sci. Rep., 2016, 6, 21828 CrossRef CAS PubMed.
- J. Chen, L. Deng, C. Porter, G. Alexander, D. Patel, J. Vines, X. Zhang, D. Chasteen-Boyd, H.-J. Sung, Y.-P. Li, A. Javed, S. Gilbert, K. Cheon and H.-W. Jun, Sci. Rep., 2018, 8, 15749 CrossRef PubMed.
- P. Carmeliet and R. K. Jain, Nature, 2011, 473, 298–307 CrossRef CAS PubMed.
- A. Ergul, A. Alhusban and S. C. Fagan, Stroke, 2012, 43, 2270–2274 CrossRef PubMed.
- W. H. AlMalki, I. Shahid, A. Y. Mehdi and M. H. Hafeez, Indian J. Pharmacol., 2014, 46 CAS.
- S. Webb, Nature, 2018, 554, 555–557 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |









