Open Access Article
Open Access ArticleA novel copper formate-based framework RbCu(HCO2)2Cl: synthesis, crystal structure, thermal, vibrational and magnetic properties and antibacterial activity
Asmae
Ben Abdelhadi
 ab,
Safaa
Hidaoui
a,
Rachid
Ouarsal
a,
Morgane
Poupon
ab,
Safaa
Hidaoui
a,
Rachid
Ouarsal
a,
Morgane
Poupon
 c,
Michal
Dusek
c,
María
de los Llanos Palop Herreros
d,
Marco Antonio
López de la Torre
e,
Luis
Lezama
c,
Michal
Dusek
c,
María
de los Llanos Palop Herreros
d,
Marco Antonio
López de la Torre
e,
Luis
Lezama
 f,
Brahim
El Bali
a,
Mohammed
Lachkar
f,
Brahim
El Bali
a,
Mohammed
Lachkar
 *a and
Abderrazzak
Douhal
*a and
Abderrazzak
Douhal
 *b
*b
aEngineering Laboratory of Organometallic, Molecular Materials, and Environment (LIMOME), Faculty of Sciences, Sidi Mohamed Ben Abdellah University, 30000 Fez, Morocco. E-mail: mohammed.lachkar@usmba.ac.ma
bDepartamento de Química Física, Facultad de Ciencias Ambientales y Bioquímica, e INAMOL, Campus Tecnológico de Toledo, Universidad de Castilla-La Mancha (UCLM), Avenida Carlos III, S.N., 45071 Toledo, Spain. E-mail: Abderrazzak.Douhal@uclm.es
cInstitute of Physics of the Czech Academy of Sciences, Na Slovance 2, 8, Praha 182 21, Czech Republic
dDepartamento de Química Analítica y Tecnología de los Alimentos, Facultad de Ciencias Ambientales y Bioquímica, Universidad de Castilla-La Mancha, (UCLM), Avenida Carlos III, S.N., 45071 Toledo, Spain
eDepartamento de Física, Facultad de Ciencias Ambientales y Bioquímica, e INAMOL, Universidad de Castilla-La Mancha, Avenida Carlos III, S.N., 45071 Toledo, Spain
fDepartamento de Química Orgánica e Inorgánica, Facultad de Ciencia y Tecnología, Universidad del País Vasco, UPV/EHU, B° Sarriena s/n, 48940 Leioa, Spain
First published on 16th December 2025
Abstract
In this contribution, we report on the synthesis and characterization of a novel biologically active Cu(II)-based paddle-wheel (PW) metal–organic framework (MOF), RbCu(HCO2)2Cl (1). Single-crystal X-ray diffraction results confirmed a monoclinic unit cell with space group P21/n and the Rb+ cation as a counter-balanced ion located in the cavities of the framework of the dinuclear copper–copper dimer formed by a PW-arrangement of formate anions in the syn–syn configuration. Each Cu(II) atom has a square-pyramidal environment with a Cu⋯Cu intramolecular distance of 2.7070 (7) Å. The IR spectrum confirms the existence of the formate anion (HCO2−). Magnetic susceptibility experiments, performed from 5 to 300 K, revealed a strong antiferromagnetic coupling (J = −531 cm−1) between the two Cu2+ ions linked by four formate groups. The powder EPR spectra show the typical lines of the triplet state (S = 1) with significant zero-field splitting, attributed to Cu2+–Cu2+ dimers. In vitro antibacterial activity was evaluated against two Gram-positive bacteria (Staphylococcus aureus CECT 86 and Listeria monocytogenes CECT 4031) and two Gram-negative bacteria (Escherichia coli CECT 99 and Klebsiella pneumoniae CECT 143T). The studies revealed that MOF 1 exhibits both bacteriostatic and bactericidal activity against all the microorganisms analyzed, making it a potential candidate for treating bacterial infections. The obtained findings provide more insight into the interesting properties of Cu-based frameworks and antibacterial activity.
1. Introduction
Metal–organic frameworks (MOFs) are an important class of hybrid crystalline materials made of metal centres and organic linkers. They offer adjustable structures, high porosity, and multifunctionality. Their structural flexibility and chemical diversity have enabled many applications, including catalysis, gas storage, sensing, magnetism and increasingly, biological uses.1–3 Recently, research has focused on exploring MOFs with a binuclear paddle wheel (PW) topology. Their outstanding porosity, adjustable design and their unique physicochemical properties make them promising for various applications, such as biologically active agents, electrochemical processes, ferroelectric and dielectric materials, magnetism, catalysis, supramolecular assemblies, and molecular magnetic devices.4–8 Among various metal ions used in PW-based MOF chemistry, copper is one of the most widely employed metal centres for forming paddle-wheel structures, and it is a significant transition metal in biology and coordination chemistry.9–13 Copper is an essential trace element in the human body. In addition, it plays crucial roles in many biological processes and enzyme regulation, as well as in the structural and functional enhancement of proteins; it also exhibits intrinsic antimicrobial and redox activity. Consequently, the combination of biologically relevant copper centres with the structural robustness of the PW motif offers a promising platform for the design of functional materials with enhanced biological activity.14,15 Notably, numerous studies have shown that Cu–PW MOFs are being extensively explored for biological applications. These include antibacterial, anticancer, antifungal, antimicrobial, and antiviral activities, which arise from the unique coordination sphere around central Cu(II) cations, their ability to catalyze the generation of reactive oxygen species (ROS), disrupt membrane integrity, interact with nucleic acids, and inhibit key microbial enzymes.14,16,17 These multifunctional properties make Cu(II) MOFs biologically more relevant and functional than other compounds. The higher biological activity of Cu(II)-based MOFs is significantly influenced by the nature of the ligands used. In particular, N- and O-donor ligands represent a promising class of candidates in the development of new Cu(II)-based antimicrobial materials. These ligands often coordinate the copper metal through different coordination geometries (four, five, or six), forming a stable chelating framework. This chelation process generally reduces the polarity of the copper center, increases the lipophilicity of the compound and thereby improves its ability to penetrate biological membranes, leading to enhanced biological activity.16,18–20 In addition to that, Cu PW-based MOFs have gained significant interest, owing to their magnetic behaviour, facilitated by the rather short formate anion HCO2−, which allows strong magnetic coupling between the Cu(II) centers.21,22 In general, a Cu(II)-based compound contains an unpaired d9 electron configuration that gives its paramagnetic properties. Additionally, it has been reported that the coupling of the d9 unpaired electrons between the Cu(II) centers can result in a triplet (high spin) or a singlet (low spin) ground state or other magnetic properties such as ferromagnetic (FM) and antiferromagnetic (AFM) behavior, respectively. This depends on the types of coordination modes, metal geometry and environment.23,24 In this context, the design and development of novel Cu-based PW frameworks, along with the investigation of their magnetic behaviour and the exploration of their biological activity remain an active and promising area of research for a better understanding of the structure–activity relationship.Herein, we synthesized, fully characterized, examined the magnetic properties, and explored the biological activity of a novel Cu(II)-based MOF, RbCu(HCO2)2Cl (1), containing the largest possible monoatomic A-site cations Rb+. The Cu-based MOF was obtained by a slow diffusion method, with excellent reproducibility and was fully characterized by a combination of single crystal X-ray diffraction (SXRD), thermal stability, EPR and SQUID techniques. At 300 K, 1 crystallizes in the monoclinic system with space group P21/n, and its crystal structure features the well-known paddle-wheel motif typical of copper formates, Cu2(HCOO)4, with an intramolecular Cu⋯Cu distance of 2.7070(7) Å within the dinuclear unit. Magnetic susceptibility data, obtained from 5 to 300 K, revealed that 1 exhibits a strong antiferromagnetic coupling (J = −531 cm−1) between the two Cu2+ ions linked by four formate groups, which form the Cu2 paddle-wheel units that give rise to the anionic network in this compound. Furthermore, we also evaluated the in vitro antibacterial activity of MOF 1 against two Gram-positive bacteria, S. aureus CECT 86 and L. monocytogenes CECT 4031, and two Gram-negative bacteria, E. coli CECT 99 and K. pneumoniae CECT 143T, using both the disc diffusion and the agar well diffusion methods.
2. Experimental section
2.1. Materials
All the chemicals are of reagent grade and were used as supplied. Copper(II) chloride tetrahydrate (CuCl2·4H2O, 99.9%) and formic acid (HCOOH, 98%) were purchased from Fluka (Switzerland). Rubidium carbonate (Rb2CO3, 99.5%) was bought from Sigma-Aldrich (France) and anhydrous methanol (99.7%) was purchased from Panreac (Spain).2.2. Synthesis
The green prismatic single crystals of 1 were obtained using a mild solution chemistry approach at ambient temperature using methanol as solvent, since it is widely known as a non-coordinating medium in synthesizing metal formate frameworks.25 In a typical synthesis procedure, 10 mL of a methanolic solution containing 4 mmol of rubidium carbonate (0.92 g) and 80 mmol of formic acid (4 mL) were placed at the bottom of a glass tube. Upon this solution, 10 mL of a methanolic solution containing 2 mmol of CuCl2·4H2O (0.340 g) were carefully layered on top. The tube was sealed and undisturbed at room temperature for several days until green square-shaped crystals are formed. The crystals of 1 were subsequently isolated by filtration, washed three times with ethanol, and dried at room temperature for 24 h; the resulting crystals were quite stable (Fig. S1). HR-ESI-MS (negative mode) m/z calculated for RbCuC2H2O4Cl: 274.50100; found: 273.94864.2.3. Characterization and measurements
Single-crystal datasets were collected at 300 K, on a suitable green single crystal of 1 with dimensions 0.36 × 0.30 × 0.27 mm3. Data collection was performed using Mo Kα radiation (λ = 0.71073 Å) from a classical sealed tube monochromated by graphite and collimated using a fibre-optics enhanced collimator. As a detector, we used a CCD Atlas S2. CrysAlis PRO software (CrysAlis PRO 1.171.41.117a, Rigaku Oxford Diffraction, 2021) was used to process the data and an empirical multiscan absorption correction was applied using spherical harmonics, implemented in the SCALE3 ABSPACK scaling algorithm. The structure was solved by charge flipping with the SUPERFLIP package26 and refined against F2 by using full-matrix least-squares method with the Jana2020 program package.27 All non-hydrogen atoms were refined with anisotropic thermal displacement parameters. For all the hydrogen atoms, we kept Uiso(H) equal to 1.2Ueq(C/N). Visualization of the structure was made using the Diamond program.28The details concerning the crystal data, data collection, and refinement are given in Table 1. At the same time, the refined atomic positions and displacement parameters are listed in Tables S1 and S2, respectively.
| Crystal data | |
| Chemical formula | C2H2ClCuO4Rb |
| M r (g mol−1) | 274.5 |
| Crystal system; space group | Monoclinic; P21/n |
| Temperature (K) | 300 |
| a, b, and c (Å) | 8.242 (2), 7.979 (2), and 9.723 (2) |
| β (°) | 95.40 (3) |
| V (Å3) | 636.5 (3) |
| Z | 4 |
| Radiation type | Mo Kα |
| μ (mm−1) | 11.38 |
| Data collection | |
| Diffractometer | Xcalibur, AtlasS2, and Gemini ultra |
| T min; Tmax | 0.575; 1 |
| No. of measured, independent and observed [I > 3σ(I)] reflections | 9955, 1645, and 1329 |
| R int | 0.041 |
(sin![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) θ/λ)max (Å−1) θ/λ)max (Å−1) |
0.691 |
| Refinement | |
| R[F2 > 2σ(F2)], wR(F2), and S | 0.023, 0.058, and 1.30 |
| No. of reflections | 1645 |
| No. of parameters | 83 |
| H-atom treatment | H-atom parameters constrained |
| Δρmax; Δρmin (e Å−3) | 0.46; −0.54 |
3. Results and discussion
3.1. Crystal structure
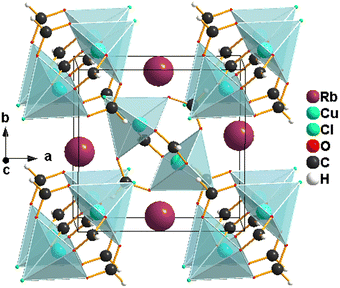
The green single crystals of RbCu(HCO2)2Cl were prepared by a facile slow diffusion solution process at room temperature. Single-crystal X-ray diffraction (SCXRD) studies reveal that the compound crystallizes in a monoclinic system with space group P21/n, a = 8.242(2) Å, b = 7.979(2) Å, c = 9.723(2) Å, β = 95.40(3)° and V = 636.5(3) Å3 at 300.1 K. 1 shows a well-known paddle-wheel structure typical of copper formates (Fig. 1A). The asymmetric unit contains one alkali metal (Rb+) and a binuclear copper(II) paddle-wheel (PW), which contains two Cu2+ ions syn–syn connected via four oxygen atoms from four different formate groups, Cu2(HCOO−)4. Each Cu(II) atom is also coordinated with a disordered chloride ion in the apical position, contributing to the framework's charge-neutrality. The result is a slightly distorted CuO4Cl square-pyramidal environment around each Cu(II) with a Cu⋯Cu intramolecular distance of 2.7070(7) Å in the binuclear unit (Fig. 1B), as described for the isostructural CsCu(HCOO)2Cl29 and close to that of other compounds with similar structures.12,21,22 In these square-pyramidal geometries, the copper ions Cu(II) lie above the basal plane towards the apical chloride ions. In addition, the planes of basal bonds around the copper ions are parallel to one another (Fig. 1C). This typical structure represents the simplest possible paddle-wheel geometry and is widely known as copper formate.15,30,31The values of the bond distances (Å) and bond angles (°) are shown in Table 2. The Cu–Oformate bond lengths range from 1.9624 (2) to 2.0075 (2) Å (Fig. 1B and Table 2). The apical bond distance in Cu–Cl is 2.3981 (12) Å. These bond length values agree with those in other binuclear Cu(II) compounds.13,24,29,32 The bond angles Cl(eq)–Cu–Oformate range from 92.30 (7) to 101.42 (7)° (Table 2). The formate moieties have rather common bond angles, 126.9 (3)° for O2–C1–O3iii and 127.5 (3)° for O1–C2–O4vi (Table 2). The bond angle for the moiety Cl1–Cu2–Cu2 is 168.943 (6)°, showing a small deviation from linearity along the Cu–Cu axis.
| Symmetry codes: (i) −x, −y + 1, −z + 1; (ii) x − 1/2, −y + 3/2, z − 1/2; (iii) −x + 1, −y + 1, −z + 1; (iv) x + 1, y, z; (v) x + 1/2, −y + 3/2, z + 1/2; (vi) x − 1, y, z. | |||
|---|---|---|---|
| Cu2–Cl1 | 2.3981 (12) | Cu2–O2 | 1.962 (2) |
| Cu2–O1i | 2.008 (2) | Cu2–O3 | 1.968 (2) |
| O1–C2 | 1.245 (4) | Cu2–O4 | 1.995 (2) |
| O2–C1 | 1.255 (4) | O3–C1iii | 1.255 (4) |
| C1–H1c1 | 0.96 | O4–C2iv | 1.253 (4) |
| C2–H1c2 | 0.96 | Cl1–Cu2–O2 | 101.42 (7) |
| Cl1–Cu2–O1i | 92.35 (7) | Cl1–Cu2–O3 | 92.30 (7) |
| Cl1–Cu2–O4 | 101.29 (7) | Cu2–O2–C1 | 119.3 (2) |
| O1i–Cu2–O2 | 88.58 (9) | Cu2–O3–C1iii | 127.2 (2) |
| O1i–Cu2–O3 | 91.45 (9) | Cu2–O4–C2iv | 119.4 (2) |
| O1i–Cu2–O4 | 166.34 (9) | O2–C1–O3iii | 126.9 (3) |
| O2–Cu2–O3 | 166.26 (10) | O2–C1–H1c1 | 116.54 |
| O2–Cu2–O4 | 88.07 (9) | O3iii–C1–H1c1 | 116.54 |
| O3–Cu2–O4 | 88.69 (9) | O1–C2–O4vi | 127.5 (3) |
| O1–C2–H1c2 | 116.23 | O4vi–C2–H1c2 | 116.23 |
| Cu2i–O1–C2 | 126.3 (2) | ||
In a unit cell, besides the two binuclear paddle-wheel (PW) copper(II) moieties, there are four Rb+ cations located in the empty spaces within the three-dimensional framework formed by the binuclear paddle-wheel (PW) copper(II) units (Fig. 2). The dinuclear PW units together with the Rb+ cations form a herringbone arrangement when viewed down the a-axis. The Rb+ cations, responsible for the charge balance of the Cu-based compound as counter ions, are bonded to four binuclear paddle-wheel Cu2(HCOO−)4 dimers via formate oxygen and chloride anions through Rb–O and Rb–Cl electrostatic interactions. This results in an extended three-dimensional coordination framework. These coordination linkages connect adjacent PW units into zigzag infinite chains parallel to the a-axis, thereby reinforcing the overall structural stability of the network. The overall supramolecular arrangement, dominated by electrostatic interactions, is comparable to that of CsCu(HCOO)2Cl.29 In contrast, in the [NH3(CH2)nNH3][Cu2(HCOO)4Cl2] metal–organic compounds (n = 3 and 4), structural stability is primarily ensured by N–H⋯O and N–H⋯Cl− H-bond interactions.33
3.2. Vibrational properties
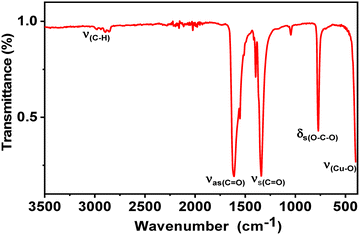
Fig. 3 exhibits the room-temperature IR spectrum of RbCu(HCO2)2Cl, while Table S3 exhibits the values of the IR absorption band frequencies and their assignment. The IR spectrum exhibits all the strongest characteristic formate vibration bands. It is well known that the HCO2− ions have six internal modes: the v1 C–H stretching mode, the v2 symmetric C–O stretching mode, the v4 antisymmetric C–O stretching mode, the v3 symmetric O–C–O bending (scissor) mode, the v5 C–H in-plane bending mode and the v6 C–H out-of-plane bending mode.34,35 These modes were observed in narrow wavenumber ranges for different metal formate frameworks.36,37 In the high wavenumber region, besides the v1(HCO2) mode, others expected bands attributed to some combinations and overtones of other v(HCO2) modes can be observed. From Fig. 3, the vibrations bands at 2895 and 2845 cm−1 are assigned to the stretching mode v1(HCO2), while the IR spectrum shows two very weak vibration bands at 2976 and 2945 cm−1 that are related to the combination of the v4(HCO2) and v5(HCO2) modes, and the overtones 2v5 and 2v2, respectively. The strong bands observed at 1614 and 1552 cm−1 in the medium wavenumber region could belong to the v4(HCOO) asymmetric C–O stretching modes of the CO2− moiety. The C–H in-plane bending mode v5(HCO2) is observed as medium IR bands in the region 1413–1376 cm−1. Additionally, the intense vibration band located at 1337 cm−1 is assignable to the symmetric C–O stretching mode v2(HCO2). In the spectrum, a significant shift in the ν(C–O) and ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) stretching can be observed in reference to the spectra of free formate ions (HCO2−), confirming the coordination of copper(II) ions with the formate groups. The modes observed in the lower wavenumber region at 1045 and 769 cm−1 are assigned to the out-of-plane C–H bending mode v6(HCO2) and the symmetric O–C–O bending mode v3(HCO2), respectively. Finally, the strong vibration band observed at 403 cm−1 is due to the Cu–O bond. These results agree well with those reported for other Cu-based formate frameworks.38–40
O) stretching can be observed in reference to the spectra of free formate ions (HCO2−), confirming the coordination of copper(II) ions with the formate groups. The modes observed in the lower wavenumber region at 1045 and 769 cm−1 are assigned to the out-of-plane C–H bending mode v6(HCO2) and the symmetric O–C–O bending mode v3(HCO2), respectively. Finally, the strong vibration band observed at 403 cm−1 is due to the Cu–O bond. These results agree well with those reported for other Cu-based formate frameworks.38–40
3.3. Thermal properties
The thermal stability of RbCu(HCO2)2Cl was investigated using combined thermogravimetric analysis (TGA) and differential thermal analysis (DTA) results from experiments on a crystalline powder sample in an air atmosphere at a heating rate of 10 K min−1, from 315 to 780 K (Fig. 4). The TGA results reveal that 1 is thermally stable up to 350 K, and above this temperature, it begins to decompose in one main mass loss step, as observed in the TGA curve (red curve in Fig. 4). The experimental percent weight loss is approximately 31.67%, which is close to the expected (calculated) loss of two formate ligands per formula unit (theoretical value: 32.78%). This decomposition is found to be exothermic, as indicated by the DTA, which displays an exothermic peak at 426 K (green curve in Fig. 4). This result is similar to the one reported for CsCu(HCO2)2Cl, in which the degradation occurs in one main weightless step at approximately 425 K.29 During the whole decomposition process of compound 1, the resulting formed residue corresponds to binary (RbCl) and metallic Cu species. These results align well with the observed thermal behavior of other Cu-containing formate-based alkali metals, where decomposition occurs over very small temperature intervals and at low onset temperatures.40,41 Additionally, a small mass degradation is observed in the TGA curve at higher temperatures, which could be attributed to internal redox reactions where copper is reduced. Subsequently, the residue remains nearly stable up to 500 K.3.4. Magnetic properties
Variable temperature magnetic susceptibility measurements were performed on powdered samples of RbCu(HCO2)2Cl in the temperature range 5–300 K. Plots of the thermal evolution of the magnetic molar susceptibility (χm) and the χmT product (χmT = μeff2/8) are shown in Fig. 5. At room temperature, χmT is equal to 0.11 cm3 K mol−1 (μeff = 0.94µB), a value significantly lower than the theoretical spin-only value for an S = 1/2 system (0.375 cm3 K mol−1). Upon cooling, χmT continuously decreases, and it practically vanishes at low temperatures (χmT = 0.002 cm3 K mol−1 at 5 K). The susceptibility decreases from 300 to 80 K and rapidly increases as the temperature is lowered further to 5 K, probably due to the small amount of the paramagnetic Cu(II) compound detected in the EPR study. This behavior is consistent with a strong antiferromagnetic coupling between the two Cu2+ ions linked by four formate groups that form the Cu2 paddle-wheel units that give rise to the anionic network in this compound. Therefore, we tried to fit the susceptibility data using the Bleaney–Bowers equation for a Cu(II) dimer42 (eqn (1)) combined with an additional term accounting for a paramagnetic contribution, assumed to come from a species with the same molecular weight per Cu(II) ion and g-value as the main compound to avoid overparameterization:43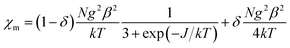 | (1) |
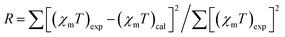 , are J = −531 cm−1, g = 2.125 and δ = 0.0047, with R = 2.8 × 10−5. As shown in Fig. 5, the fitting curves reproduce the experimental magnetic data very well over the whole temperature range investigated. The obtained singlet–triplet energy gap is close to the average value (J = −550 cm−1) determined for copper carboxylate complexes with the same core, although it is somewhat larger than that calculated (J = −425 cm−1) for a model compound [Cu2(μ-HCOO)4(H2O)2] with an intradimeric Cu–Cu distance of 2.707 Å.44 This is due to the presence of chloride ions as axial ligands and to slight differences in the geometry of the carboxylate bridges arising from the Rb–Cl and Rb–O bonds, which stabilize the three-dimensional framework. Despite the short Cu–Cu distance, super exchange is the dominant coupling mechanism in Cu dimers with the syn–syn coordination mode of the formate groups because the unpaired electron on each copper(II) ion occupies parallel dx2−y2 orbitals oriented towards the oxygen atoms of the bridging ligands. Therefore, the coupling constant can be strongly affected by small structural changes in this type of compound.
, are J = −531 cm−1, g = 2.125 and δ = 0.0047, with R = 2.8 × 10−5. As shown in Fig. 5, the fitting curves reproduce the experimental magnetic data very well over the whole temperature range investigated. The obtained singlet–triplet energy gap is close to the average value (J = −550 cm−1) determined for copper carboxylate complexes with the same core, although it is somewhat larger than that calculated (J = −425 cm−1) for a model compound [Cu2(μ-HCOO)4(H2O)2] with an intradimeric Cu–Cu distance of 2.707 Å.44 This is due to the presence of chloride ions as axial ligands and to slight differences in the geometry of the carboxylate bridges arising from the Rb–Cl and Rb–O bonds, which stabilize the three-dimensional framework. Despite the short Cu–Cu distance, super exchange is the dominant coupling mechanism in Cu dimers with the syn–syn coordination mode of the formate groups because the unpaired electron on each copper(II) ion occupies parallel dx2−y2 orbitals oriented towards the oxygen atoms of the bridging ligands. Therefore, the coupling constant can be strongly affected by small structural changes in this type of compound.
 | ||
| Fig. 5 Magnetic behavior of compound RbCu(HCO2)2Cl. The solid lines represent the best least-squares fit to experimental data (see text for details of the fit). | ||
Notably, this compound does not exhibit the complex magnetic behavior observed for the isostructural CsCu(HCOO)2Cl material.29 The susceptibility curves recorded on zero-field cooled (ZFC) and field-cooled (FC) samples show no signs of irreversibility above 5 K. Furthermore, there is no indication of a magnetic transition, such as those observed at 8 and 32 K in the aforementioned compound. At room temperature, the field-dependent magnetization displays a straight line as expected for a classic paramagnetic compound (not shown). A small ferromagnetic anomaly can be detected in the magnetization measurements at 5 K, but it is practically negligible (see Fig. S3). In fact, the magnetization curve at 5 K can be reasonably well described by the Brillouin function for a system with S = 1/2 normalized to account for the percentage of paramagnetic impurity deduced from the susceptibility fit.45
Considering the strong antiferromagnetic coupling between the two metallic ions within paddle-wheel dinuclear copper carboxylates, ferromagnetic orderings are not expected in systems with these building blocks. In fact, the sudden increase in the magnetic susceptibility frequently observed at low temperatures is usually ascribed to paramagnetic impurities resulting from the synthetic procedure.43 However, more complex magnetic properties have been observed in some coordination polymers containing that type of dimeric entity, including spontaneous magnetization.46–48 Such behavior has been interpreted in terms of a weak ferromagnetic coupling between the dimers or canted antiferromagnetic ordering due to the competition of different intra and interdimeric interactions. But these interpretations are questioned by several authors because of the essentially molecular character of the Cu2(COO)4 units in most of these compounds.49 Due to strong antiferromagnetic interactions, dimers do not possess a net spin moment at low temperatures that could give rise to weak ferromagnetism or canted ordering. In fact, theoretical calculations by Shen et al.50 show that the energy barrier necessary to create a net magnetic moment in such dimers would be impossible to overcome at temperatures close to 5 K. These authors have shown that the ferromagnetism observed in certain MOFs with paddle-wheel units linked by conjugated aromatic ligands has its origin in the existence of copper vacancies that allow the existence of localized magnetic moments. The ferromagnetic ordering is achieved via π-electrons of the aromatic linkers that connect the copper-defective dimers. Moreover, it is well known that, in other Cu oxides, Cu or oxygen vacancies produce local moments that enhance ferromagnetism.51,52 According to this model, the presence of a very small number of copper vacancies in the title compound could be the reason why it does not show the weak ferromagnetism described for the isostructural CsCu(HCOO)2Cl. It should be noted that vacancies are an inevitable feature of structurally complex systems such as these, but their concentration is strongly influenced by the preparation procedure.
3.5. Electron paramagnetic resonance behavior
The Q-band (300 K) and X-band (300, 175 and 5 K) powder EPR spectra of RbCu(HCO2)2Cl are displayed in Fig. 6A and B, respectively. A set of five broad anisotropic resonances is observed in the Q-band, centered at approximately 480, 650, 925, 1100 and 1380 mT. The X-band displays two intense signals at magnetic fields of approximately 85 and 500 mT, two weaker signals at around 30 and 590 mT, and a weak, complex band at approximately 315 mT. When the temperature is lowered to 5 K, only the latter can be detected. As is frequently observed for dinuclear copper complexes with carboxylate bridges, the recorded spectra exhibit the expected features of a triplet state (S = 1) with significant zero-field splitting (ZFS). Therefore, the following spin Hamiltonian (eqn (2)), which includes Zeeman electronic and ZFS terms,53 has been used to analyze them: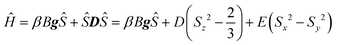 | (2) |
 | ||
| Fig. 6 EPR spectra of RbCu(HCO2)2Cl powder at (A) 33.9 GHz and 295 K and (B) 9.4 GHz at 295, 175 and 5 K. Dotted-red line in A is a simulation with the parameters indicated in the text. | ||
The sign of the D parameter cannot be determined using low-frequency EPR or magnetic susceptibility measurements, particularly in strongly antiferromagnetic systems, which become practically EPR-silent at low temperatures. However, we assume that D is negative for this compound since Ozarowski has shown by high-frequency single-crystal EPR studies that this parameter appears to be negative for all binuclear copper carboxylates exhibiting paddle-wheel structures.56 The zero-field splitting of the S = 1 state in these compounds arises from the combined effect of spin dipolar interactions and anisotropic exchange couplings between one electron in the ground state and another one in an excited state. Therefore, the calculated D value can be evaluated as D = Ddip + Dex. Considering that in this type of dimers the Cu⋯Cu direction practically coincides with the z-axis of the g-tensor, the dipolar contribution can be estimated from the distance between metals (r) using eqn (3).57
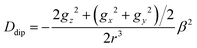 | (3) |
Using the values of g deduced from the EPR fit and the distance r(Cu⋯Cu) = 2.707 Å from the structural resolution, a value of Ddip = −0.149 cm−1 is obtained from the above expression. Therefore, the exchange contribution to the ZFS is Dex = −0.265 cm−1. This value can be used to determine the triplet–singlet separation in dimeric excited states where one cupric ion is in its electronic ground state (dx2−y2) while another one is in an excited state (dxy, dxz, and dyz).58 It is noteworthy that the relationship between Dex and the exchange integral (J) is still unknown, although a correlation between both values has been observed.59
The intense line appearing at around 1100 mT in the Q-band spectrum and labeled “U” does not correspond to any of the expected transitions within a triplet S = 1 state. This signal is too intense to be entirely attributed to the presence of a paramagnetic impurity. Furthermore, its relative intensity decreases with temperature, contrary to what would be expected for such a contribution. Additionally, spectra obtained at various microwave powers suggest that all signals exhibit comparable saturation properties, indicating that they originate from the same magnetic system. Similar resonances have been observed in polymeric structures built on copper dinuclear paddle-wheel units and have been attributed to extended exchange signals originating from the collapse of the fine structure due to interdimeric interactions.60–63 Exchange magnetic couplings between the spins of neighbouring units can average out the zero-field splitting if they are strong enough. Otherwise, the lines could only collapse for certain crystal orientations. Therefore, spectra recorded in powder samples could show both exchange and molecular resonances. Additionally, interdimeric interactions compete with the strong intradimeric antiferromagnetic couplings characteristic of these compounds. As the population of the excited S = 1 state rises with increasing temperature, the distance between the dimers with net spin moment decreases. This leads to a more efficient exchange process between the dimers, which in turn results in an increase in the intensity of the exchange signal “U”, at the expense of the ZFS lines. Considering that the exchange signal is not observable below 100 K in the RbCu(HCOO)2Cl compound, it can be concluded that the increase in magnetic susceptibility observed at low temperatures is not caused by interdimeric interactions.
Finally, it should be noted that, as the temperature decreases, the relaxation time increases and all the peaks of the spectra become narrower, except for the one corresponding to the extended exchange signal. This improves the resolution of the spectra, even though the intensity of the signals decreases (Fig. S4). Below 75 K, the signals corresponding to the triplet state are no longer detectable, confirming the presence of strong intradimeric antiferromagnetic interactions. At 5 K, only the characteristic signal of magnetically isolated Cu(II) ions (labeled “M”) is observed (Fig. S5). The hyperfine structure resulting from the interaction of the S = 1/2 electron spin with the nuclear magnetic moments of the 63Cu and 65Cu isotopes (I = 3/2) is partially resolved on the low-field region of the signal. The spin Hamiltonian parameters obtained from the fitting of the powder spectra are: g1 = 2.360, g2 = 2.111, g3 = 2.044, A1 = 0.0140 cm−1, A2 = 0.0041 cm−1, and A3 < 0.0010 cm−1. The calculated g-values indicate that these ions are in the same ground state (dx2−y2) as those forming the dimer, but the environment is notably different in the equatorial plane. EPR signals from mononuclear Cu2+ ions in compounds with paddle-wheel units are usually observed and attributed to extra-framework impurities. Nevertheless, it cannot be ruled out that the spectrum is due to the existence of copper vacancies that convert some of the dimers into monomers. In any case, the observed increase in magnetic susceptibility below 70 K (Fig. 5) is due to these isolated Cu(II) ions.
3.6. Evaluation of antibacterial activity
The antibacterial activity of RbCu(HCO2)2Cl was evaluated in vitro against two Gram-positive bacteria, S. aureus CECT 86 and L. monocytogenes CECT 4031, and two Gram-negative bacteria, E. coli CECT 99 and K. pneumoniae CECT 143T, using both the disc diffusion and the agar well diffusion methods. For both, 20 µL of a sterile 15 mg mL−1 solution of 1 were used. Table 3 shows the values of the inhibition halo diameter from both methods (see Fig. S6) and those from the commercial discs of antibiotics used as the positive control for each of the bacteria. It is worth noting that the diameters obtained for Gram-positive bacteria were higher than those for Gram-negative in both methods.| Inhibition halo diameter (mm) | ||||
|---|---|---|---|---|
| Gram-positive bacteria | Gram-negative bacteria | |||
| S. aureus CECT 86 | L. monocytogenes CECT 4031 | E. coli CECT 99 | K. pneumoniae CECT 143T | |
| Disc diffusion method | 12 | 15 | 9 | 8 |
| Agar well diffusion method | 25 | 27 | 15 | 15 |
| Norfloxacin (5 µg) | 19 | |||
| Vancomycin (30 µg) | 14.5 | |||
| Gentamicin (10 µg) | 15 | 17.5 | ||
| Ampicillin- sulfactam (10/10 µg) | 16 | |||
| Trimethoprim-sulfamethoxazole (25 µg) | 16 | |||
| Nitrofurantoin (300 µg) | 18 | |||
| Ciprofloxacin (5 µg) | 24.5 | |||
Comparison of values from the disc diffusion method with those from the commercial discs of antibiotics displayed the lower antibacterial activity of 1 against all the bacteria, except for L. monocytogenes CECT 4031, for which the diameters of the halos obtained were similar. The diameters of halos from the agar well diffusion method were, for all the bacteria, greater than those from the disc diffusion method, even though the quantity of 1 used was the same. It could be due to a better diffusion of the compound in the agar from the wells. Fig. 7 shows the results from the quantitative assay to determine the MIC of 1. It is clear that 1 was able to inhibit the growth of both Gram-positive and Gram-negative bacteria, though the decrease of growth varied with the species and with the concentration. For all the bacteria, except for S. aureus CECT 86, the growth was inversely proportional to the concentration of 1, as reported in other studies.64 In the case of S. aureus CECT 86, the maximum decrease was obtained at a concentration of 800 µg mL−1. Using larger concentrations did not produce a significant increase in growth inhibition, with values remaining stable around 20%.
 | ||
| Fig. 7 In vitro antimicrobial activity of RbCu(HCO2)2Cl at different concentrations against two Gram-negative and two Gram-positive bacteria. | ||
The concentrations that were assayed appear to be significantly above the MIC, as the percentages of growth decrease were less than fifty percent for three of the four bacteria tested. Concentrations above 1000 µg mL−1 were more effective against L. monocytogenes CECT 4031, achieving a growth reduction of 54.02% at the highest concentration tested, followed by E. coli CECT 99 and K. pneumoniae CECT 143T. It might be advisable for future studies to use higher concentrations along with toxicity assessments of 1.
In reference to the measurement of the DO600 during the incubation of the strains at different concentrations of 1, it is worth noting that when E. coli CECT 99 was grown in the presence of 2500 µg mL−1 of 1, the DO600 remained constant during the 24 h of incubation and the counts of viable cells at the end of incubation showed that the cells from the inoculum had remained alive without showing growth. This observation suggests that 1 at this concentration has a bacteriostatic effect. A similar result was observed for L. monocytogenes CECT 4031 at concentrations of 2000 and 2500 µg mL−1.
With respect to the antibacterial activity of the copper compounds, some authors reported that it depends on the geometry of the complexes and the nature of the ligands, because both factors influence the lipophilic character of the compounds, affecting their permeability through the cytoplasmic membrane.65,66 The mechanism of action described for these compounds are diverse although in many respects still unknown.67 For example, it has been suggested that the release of copper ions causes the inactivation of enzymes,68 and more recently that the metallic chelates of Cu2+ act disturbing the microbial metabolism and producing their destruction.17 Other possible mechanisms include damage to DNA by reactive oxygen species (ROS) produced through Fenton-type reactions, as well as a decrease in cell membrane integrity.10,17 Differences observed in the activity against Gram-positive and Gram-negative bacteria may stem from variations in the structure and chemical composition of their cell walls, as noted for other metallic compounds.69
The antibacterial activity of metal-based complexes can be rationalized using chelation theory and Tweedie's polarization model.16,70–74 Typically, the chelation modifies metal ion properties, enhancing membrane penetration and polarization-induced reactive oxygen species (ROS) generation, which together account for the superior antibacterial activity of metal complexes.16 For the present compound, the antibacterial activity arises from synergistic structural and electronic effects. We suggest that chelation of Cu(II) by bridging formate ligands delocalizes the positive charge over oxygen atoms, reducing the metal center's polarity and enhancing its lipophilicity relative to the hydrated Cu2+ ion. This favours interaction with bacterial surfaces and membrane penetration. According to Tweedie's model, the Cu–formate environment also promotes polarization of negatively charged bacterial membranes, facilitating copper uptake.70 Additionally, the distorted Cu2+ sites may undergo Cu2+/Cu+ redox cycling, generating ROS that damage key biomolecules.16
4. Conclusion
To conclude, a new copper(II)-based MOF paddle-wheel-like compound, RbCu(HCO2)2Cl, was successfully synthesized and characterized using spectroscopic, thermal, and magnetic techniques. Single-crystal X-ray diffraction revealed that 1 crystallizes in a monoclinic system with a 3D framework built from dinuclear Cu(II) paddle-wheel units, where each Cu(II) center adopts a square-pyramidal geometry with a Cu⋯Cu distance of 2.7070(7) Å. The Rb+ cations serve as counter-ions, ensuring charge balance within the framework. The thermal stability of 1 was studied. Upon further heating of 1, its resulting residue corresponds to binary (RbCl) and metallic Cu species. Magnetic studies revealed strong intradimer antiferromagnetic exchange coupling (J = −531 cm−1) between the Cu2+ centers, corroborated by EPR spectra showing characteristic triplet-state (S = 1) features with zero-field splitting. The in vitro antibacterial tests against S. aureus, L. monocytogenes, E. coli, and K. pneumoniae demonstrated both bacteriostatic and bactericidal activity, suggesting its potential as a promising antimicrobial agent. Overall, these findings highlight the multifunctional nature of this Cu(II)-based paddle-wheel MOF, encouraging further investigation into its magnetic behavior and the development of related bioactive coordination materials for potential biological applications.Author contributions
Asmae Ben Abdelhadi: writing – original draft, methodology, investigation, and formal analysis. Safaa Hidaoui: investigation. Rachid Ouarsal: writing – review & editing, validation, methodology, and formal analysis. Morgane Poupon: software and formal analysis. Michal Dusek: software and formal analysis. María de los Llanos Palop Herreros: investigation, writing – review & editing, validation, methodology, and formal analysis. Marco Antonio López de la Torre: methodology, investigation, and formal analysis. Luis Lezama: writing original draft, validation, methodology, investigation, and formal analysis. Brahim El Bali: conceptualization, validation, formal analysis, and writing – review & editing. Mohammed Lachkar: supervision, project administration, resources, methodology, investigation, and formal analysis, validation and writing – review & editing. Abderrazzak Douhal: supervision, conceptualization, project administration, resources, methodology, investigation, and formal analysis, validation and writing – review & editing. All authors read and approved the manuscript.Conflicts of interest
There are no conflicts of interest to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5ma01055a.CCDC 2468965 (1) contains the supplementary crystallographic data for this paper.75
Acknowledgements
This work was supported by the following grants: PID2020-116519RB-I00 funded by MICIU/AEI/10.13039/501100011033 and the European Union (EU) and SBPLY/23/180225/000196 funded by the JCCM and the EU through Fondo Europeo de Desarrollo Regional (FEDER). A. B. A. is grateful for the grant from the Spanish Service for the Internationalization of Education (SEPIE), through the EU Erasmus+ Key Action program (2020-1-ES01-KA107-079868). The authors A. B. A., S. H., R. O., B. E. B. and M. L. would like to acknowledge the technical assistance from the Interface Regional University Center (University Sidi Mohammed Ben Abdellah, Fez, Morocco). Crystallographic analysis was supported by the project 20-LM2023051 of the Czech Nanolab Infrastructures, supported by MEYS CR.References
- J. Annamalai, P. Murugan, D. Ganapathy, D. Nallaswamy, R. Atchudan, S. Arya, A. Khosla, S. Barathi and A. K. Sundramoorthy, Chemosphere, 2022, 298, 134184 CrossRef CAS PubMed.
- Q. Zhang, S. Yan, X. Yan and Y. Lv, Sci. Total Environ., 2023, 902, 165944 CrossRef CAS PubMed.
- D. Li, A. Yadav, H. Zhou, K. Roy, P. Thanasekaran and C. Lee, Global Challenges, 2024, 8, 2300244 CrossRef PubMed.
- M. Mendt, M. Maliuta, S. Ehrling, F. Schwotzer, I. Senkovska, S. Kaskel and A. Pöppl, J. Phys. Chem. C, 2022, 126, 625–633 CrossRef CAS.
- N. Ahmad, H. A. Younus, A. H. Chughtai, K. Van Hecke, Z. A. Khattak, Z. Gaoke, M. Danish and F. Verpoort, Catal. Sci. Technol., 2018, 8, 4010–4017 RSC.
- T. Zanette, Y. M. Variani, B. P. Nicola, S. Gutiérrez-Tarriño, C. Cerezo-Navarrete, G. Agostini, P. Oña-Burgos, C. W. Lopes and K. Bernardo-Gusmão, Catal. Today, 2025, 444, 114988 CrossRef CAS.
- T.-R. Lin, C.-H. Lee, Y.-C. Lan, S. Mendiratta, L.-L. Lai, J.-Y. Wu, K.-M. Chi and K.-L. Lu, Polymers, 2018, 10, 1398 CrossRef PubMed.
- Z. Zhang, H. Yu, X. Shen, L. Sun, S. Yue and H. Tang, Materials, 2021, 14, 3125 CrossRef CAS PubMed.
- M.-H. Yan, J. Wang, X.-Y. Su, H. Sakiyama, N. Qi, M. Afzal, A. Alarifi, D. Srivastava and A. Kumar, New J. Chem., 2023, 47, 11134–11142 RSC.
- A. L. Pereira, M. A. Vasconcelos, A. L. Andrade, I. M. Martins, A. K. Holanda, A. C. Gondim, D. P. Penha, K. L. Bruno, F. O. Silva and E. H. Teixeira, Curr. Microbiol., 2023, 80, 133 CrossRef CAS PubMed.
- Viola, N. Muhammad, A. Noor, M. Sirajuddin, M. Kubicki, S. Rahim, A. Samad, S. Shujah, A. Wadood and S. Ali, Pharmaceuticals, 2023, 16, 1462 CrossRef CAS PubMed.
- F. Li, K. Liu, S. Fan, T. Qin, S. Xiong and J. Wang, Z. Anorg. Allg. Chem., 2018, 644, 560–563 CrossRef CAS.
- V. Paredes-García, R. C. Santana, R. Madrid, A. Vega, E. Spodine and D. Venegas-Yazigi, Inorg. Chem., 2013, 52, 8369–8377 CrossRef PubMed.
- K. Ngece, V. Khwaza, A. M. Paca and B. A. Aderibigbe, Antibiotics, 2025, 14, 516 CrossRef CAS PubMed.
- M. Premkumar, D. Kaleeswaran, G. Kaviyarasan, D. Prasanth and G. Venkatachalam, ChemistrySelect, 2019, 4, 7507–7511 CrossRef CAS.
- M. S. Alhussaini, A. A. I. Alyahya and A. A. Al-Ghanayem, Dyes Pigm., 2025, 243, 113040 CrossRef CAS.
- B. Sharma, S. Shukla, R. Rattan, M. Fatima, M. Goel, M. Bhat, S. Dutta, R. K. Ranjan and M. Sharma, Int. J. Biomater., 2022, 2022, 6819080 Search PubMed.
- V. J. Tamilpriyai, P. Nagarasu, K. S. Dharshini, P. Dhanaraj, A. Veerappan, D. Moon, S. P. Anthony and V. Madhu, J. Mol. Struct., 2023, 1294, 136414 Search PubMed.
- D. Majumdar, S. Roy, J. Elizabeth Philip, B. Tüzün and S. Hazra, Inorg. Chem. Commun., 2024, 160, 111933 CrossRef CAS.
- S. Notz, K. Müller, R. Thomas, T. Rüffer and H. Lang, Inorg. Chem. Commun., 2024, 167, 112765 Search PubMed.
- J. Soldevila-Sanmartín, J. A. Ayllón, T. Calvet, M. Font-Bardia, C. Domingo and J. Pons, Inorg. Chem. Commun., 2016, 71, 90–93 Search PubMed.
- J. Zhou, L. Du, Z. Li, Y. Qiao, J. Liu, M. Zhu, P. Chen and Q. Zhao, J. Coord. Chem., 2013, 66, 2166–2177 Search PubMed.
- H. Phetmung and A. Nucharoen, Polyhedron, 2019, 173, 114121 CrossRef CAS.
- E. Y. Bivián-Castro, M. Flores-Alamo, R. Escudero, V. Gómez-Vidal, J. J. N. Segoviano-Garfias, J. Castañeda-Contreras and Q. E. Saavedra-Arroyo, Materials, 2023, 16, 4866 CrossRef PubMed.
- S. M. Bovill and P. J. Saines, CrystEngComm, 2015, 17, 8319–8326 RSC.
- L. Palatinus and G. Chapuis, J. Appl. Crystallogr., 2007, 40, 786–790 CrossRef CAS.
- V. Petříček, L. Palatinus, J. Plášil and M. Dušek, Z. Kristallogr. – Cryst. Mater., 2023, 238, 271–282 CrossRef.
- K. Brandenburg and H. Putz, Crystal Impact GbR, Bonn, Germany, 2006.
- J. Zhou, S. Jin, R. Sun, C. Chai, M. Hao, X. Zhong and X. Chen, Phys. Rev. Mater., 2021, 5, 074405 Search PubMed.
- K. Karthik and A. M. Qadir, J. Struct. Chem., 2019, 60, 1126–1132 Search PubMed.
- A Qadir, Orient. J. Chem., 2017, 33, 968–970 Search PubMed.
- B. Saha, M. Bhattacharjee, S. R. Boruah, R. N. Dutta Purkayastha, R. M. Gomila, S. Chowdhury, A. Mandal and A. Frontera, J. Mol. Struct., 2023, 1272, 134104 Search PubMed.
- M. Trzebiatowska-Gusowska and A. Gagor, J. Coord. Chem., 2017, 70, 1–22 Search PubMed.
- C. W. A. Paschoal, M. R. Moura, A. P. Ayala, J. M. Sasaki, P. T. C. Freire, F. E. A. Melo, J. Mendes Filho, I. Guedes, A. G. Leyva, G. Polla, D. Vega and P. K. Perazzo, J. Solid State Chem., 2000, 154, 338–343 CrossRef CAS.
- M. A. Moreno, O. Gálvez, B. Maté, V. J. Herrero and R. Escribano, J. Phys. Chem. A, 2011, 115, 70–75 CrossRef CAS PubMed.
- A. Ciupa and M. Ptak, Vib. Spectrosc., 2016, 86, 67–74 CrossRef CAS.
- M. Mączka, A. Ciupa, A. Gągor, A. Sieradzki, A. Pikul and M. Ptak, J. Mater. Chem. C, 2016, 4, 1186–1193 RSC.
- A. B. Abdelhadi, R. Ouarsal, M. Poupon, M. Dusek, J. P. A. González, L. Lezama, B. El Bali, M. Lachkar and A. Douhal, Mater. Adv., 2024, 5, 6154–6161 RSC.
- A. P. Ayala, J. M. Henriques Neto, C. W. A. Paschoal, I. Guedes, J. M. Sasaki, P. T. C. Freire, F. E. A. Melo, J. Mendes Filho, A. G. Leyva, G. Polla, D. Vega and P. K. Perazzo, J. Raman Spectrosc., 2000, 31, 491–495 CrossRef CAS.
- Y. Abusa, J. Greenfield, G. Viswanathan, S. Goswami, E. Ross, P. Yox, R. Oppong, I. Ojo, J. Liu, A. Ozarowski and K. Kovnir, Chem. Sci., 2025, 16, 11027–11038 Search PubMed.
- E. Eikeland, N. Lock, M. Filsø, M. Stingaciu, Y. Shen, J. Overgaard and B. B. Iversen, Inorg. Chem., 2014, 53, 10178–10188 Search PubMed.
- B. Bleaney and K. Bowers, Proc. R. Soc. London, Ser. A, 1952, 214, 451–465 CAS.
- O. Kahn, Molecular magnetism, VCH Publishers Inc., New York, 1993, pp. 167–174 Search PubMed.
- A. Rodríguez-Fortea, P. Alemany, S. Alvarez and E. Ruiz, Chem. – Eur. J., 2001, 7, 627–637 Search PubMed.
- R. L. Carlin, Magnetochemistry, Springer Science & Business Media, 2012 Search PubMed.
- S. S.-Y. Chui, S. M.-F. Lo, J. P. Charmant, A. G. Orpen and I. D. Williams, Science, 1999, 283, 1148–1150 CrossRef CAS PubMed.
- B. Moulton, J. Lu, R. Hajndl, S. Hariharan and M. J. Zaworotko, Angew. Chem., 2002, 114, 2945–2948 Search PubMed.
- S. Feng, H. Duan, H. Tan, F. Hu, C. Liu, Y. Wang, Z. Li, L. Cai, Y. Cao, C. Wang, Z. Qi, L. Song, X. Liu, Z. Sun and W. Yan, Nat. Commun., 2023, 14, 7063 Search PubMed.
- D. Maspoch, D. Ruiz-Molina and J. Veciana, Chem. Soc. Rev., 2007, 36, 770–818 RSC.
- L. Shen, S.-W. Yang, S. Xiang, T. Liu, B. Zhao, M.-F. Ng, J. R. Göettlicher, J. Yi, S. Li and L. Wang, J. Am. Chem. Soc., 2012, 134, 17286–17290 Search PubMed.
- S. Shi, D. Gao, Q. Xu, Z. Yang and D. Xue, CrystEngComm, 2015, 17, 2118–2122 RSC.
- D. Gao, J. Zhang, J. Zhu, J. Qi, Z. Zhang, W. Sui, H. Shi and D. Xue, Nanoscale Res. Lett., 2010, 5, 769 CrossRef CAS PubMed.
- E. Wasserman, L. C. Snyder and W. A. Yager, J. Chem. Phys., 1964, 41, 1763–1772 CrossRef CAS.
- W. R. Hagen, Int. J. Mol. Sci., 2023, 24, 14793 CrossRef CAS PubMed.
- J. A. Weil and J. R. Bolton, Electron paramagnetic resonance: elementary theory and practical applications, John Wiley & Sons, 2007 Search PubMed.
- A. Ozarowski, Inorg. Chem., 2008, 47, 9760–9762 Search PubMed.
- A. Bencini and D. Gatteschi, Electron paramagnetic resonance of exchange coupled systems, Springer Science & Business Media, 2012 Search PubMed.
- P. K. Ross, M. D. Allendorf and E. I. Solomon, J. Am. Chem. Soc., 1989, 111, 4009–4021 CrossRef CAS.
- L. Gutierrez, G. Alzuet, J. Borrás, A. Castineiras, A. Rodríguez-Fortea and E. Ruiz, Inorg. Chem., 2001, 40, 3089–3096 CrossRef CAS PubMed.
- M. Perec, R. Baggio, R. P. Sartoris, R. C. Santana, O. Peña and R. Calvo, Inorg. Chem., 2010, 49, 695–703 CrossRef CAS PubMed.
- M. Simenas, M. Kobalz, M. Mendt, P. Eckold, H. Krautscheid, J. R. Banys and A. Pöppl, J. Phys. Chem. C, 2015, 119, 4898–4907 CrossRef CAS.
- V. Paredes-García, R. C. Santana, R. Madrid, A. S. Vega, E. Spodine and D. Venegas-Yazigi, Inorg. Chem., 2013, 52, 8369–8377 CrossRef PubMed.
- A. Pöppl, S. Kunz, D. Himsl and M. Hartmann, J. Phys. Chem. C, 2008, 112, 2678–2684 CrossRef.
- S. Reja, D. Sarkar, K. Sarkar, D. Mukherjee, T. Fayaz, P. Sanphui and R. K. Das, Inorg. Chim. Acta, 2024, 560, 121809 Search PubMed.
- G. Borthagaray, M. Mondelli and M. Torre, J. Infect. Dis. Epidemiol., 2016, 2, 10.23937 Search PubMed.
- S. K. Nandanwar and H. J. Kim, ChemistrySelect, 2019, 4, 1706–1721 Search PubMed.
- A. C. Hangan, R. L. Lucaciu, A. Turza, L. Dican, B. Sevastre, E. Páll, L. S. Oprean and G. Borodi, Int. J. Mol. Sci., 2023, 24, 13819 CrossRef CAS PubMed.
- J. A. Lemire, J. J. Harrison and R. J. Turner, Nat. Rev. Microbiol., 2013, 11, 371–384 CrossRef CAS PubMed.
- A. B. Abdelhadi, S. Rodríguez-Sánchez, R. Ouarsal, M. Saadi, L. El Ammari, N. Morley, B. El Bali, Ó. Gómez-Torres, M. Lachkar and A. Douhal, Mater. Adv., 2024, 5, 3058–3066 RSC.
- T. BG, Phytopathology, 1964, 55, 910–918 Search PubMed.
- C. Barbot, L. Gouriou, M. Mignot, M. Sebban, P. Zhang, D. Landy, C.-C. Ling and G. Gouhier, Molecules, 2025, 30, 3263 Search PubMed.
- S. Arulmozhi, G. Sasikumar, A. Subramani, M. K. Mohammed, S. J. A. Ali, S. Ponnusamy, M. S. Jabir, A. M. Elgorban, W. Zhang and H. Natarajan, ACS Omega, 2023, 8, 34458–34470 CrossRef CAS PubMed.
- D. Majumdar, J. E. Philip, S. Roy, B. Gassoumi and H. Ghalla, BMC Chem., 2025, 19, 227 Search PubMed.
- S. S. A. Fathima, M. M. S. Meeran and E. Nagarajan, Struct. Chem., 2020, 31, 521–539 CrossRef.
- CCDC 2468965: Experimental Crystal Structure Determination, 2025 DOI:10.5517/ccdc.csd.cc2nw51s.
| This journal is © The Royal Society of Chemistry 2026 |