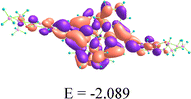Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Harnessing the zinc(II) bis-terpyridine complex to overcome drug resistance: mechanistic insights into antibacterial activity
Sourav
Sutradhar†
 a,
Saurabh
Gupta†
b,
Parnashabari
Sarkar
a,
Dipankar
Das
a,
Kamaldeep
Paul
a,
Saurabh
Gupta†
b,
Parnashabari
Sarkar
a,
Dipankar
Das
a,
Kamaldeep
Paul
 *bc and
Biswa Nath
Ghosh
*bc and
Biswa Nath
Ghosh
 *a
*a
aDepartment of Chemistry, National Institute of Technology Silchar, Silchar, Assam, India. E-mail: bnghosh@che.nits.ac.in
bTIET-VT, Centre of Excellence in Emerging Materials, Thapar Institute of Engineering and Technology, Patiala, Punjab, India 147004
cDepartment of Chemistry and Biochemistry, Thapar Institute of Engineering and Technology, Thapar, Punjab, India. E-mail: kpaul@thapar.edu
First published on 17th February 2026
Abstract
The increasing prevalence of drug-resistant bacteria represents a serious global health challenge, prompting scientists to develop innovative antibacterial compounds. An example of such a compound with significant potential as an antibacterial agent is zinc(II) complex ZnL2 of a terpyridine-based ligand 4′-(3,4-diethoxyphenyl)-2,2′:6′,2″-terpyridine L. The single crystal X-ray diffraction analysis of [ZnL2](ClO4)2·CH3CN·H2O confirms distorted octahedral coordination geometry around zinc(II) in which zinc is coordinated by two terpyridine ligands L. The preliminary investigations identified ZnL2 as a potent antibacterial compound, outperforming widely used antibiotics like amoxicillin, tetracycline, and chloromycin in its action against Bacillus subtilis and Salmonella enterica. The complex ZnL2 efficiently inhibits the development of drug resistance, with 20 passages for B. subtilis and 23 passages for S. enterica, and showed low toxicity towards normal cell lines of Hek293. As part of the extended investigation, biofilm inhibition, membrane disruption, leakage, and intracellular oxidative stress assays were done. Further analysis of the complex's interactions with DNA was conducted to assess its potential for targeting bacterial DNA as a therapeutic agent. The interaction of the complex with transport proteins such as serum albumin was evaluated to gain an initial understanding of the drug's ADME characteristics. The findings of this study emphasise the substantial antibacterial potential of the developed zinc terpyridine complex. Its promising activity could form the basis for future advancement in the treatment of bacterial infections.
1. Introduction
Nitrogen-based heterocyclic compounds have received significant attention due to their extensive range of applications in pharmacology.1,2 Nitrogen-containing heterocycles, including pyridine, pyrimidine, quinoline, and pyrazine, are fundamental structural elements in many phytochemical drugs like codeine, morphine, vinblastine, reserpine, procaine, papaverine, emetine, and digitalis-derived cardiac glycosides.3 These heterocycles are also extensively found in synthetic drugs such as diazepam, isoniazid, chlorpromazine, metronidazole, chloroquine, azidothymidine, antipyrine, etc.3–5 In addition, most enzymes, co-enzymes, hormones, vitamins, nucleic acids and alkaloids have N-heterocyclic skeletons.5 2,2′:6′,2″-Terpyridines are one such category of nitrogen-based heterocyclic ligands, which have been used extensively as anticancer,6 antioxidant7 and antimicrobial agents.8 These ligands have high affinity for binding with biomolecules like serum albumin and DNA. In addition, 2,2′:6′,2″-terpyridines also possess strong binding affinity towards transition metals, due to the presence of three pyridine ring nitrogen atoms. The metal complexes of 2,2′:6′,2″-terpyridines have significant applications in anion and amino acid sensing,9–11 halogen bonding,12,13 gelation,14–17 dye-sensitised solar cells,18,19 catalytic fields,11etc. Among the various transition metal complexes, zinc complexes are of significant interest. Zinc is the second most abundant trace element in the human body, playing a crucial role in maintaining health across all life stages and catalyzes over 300 enzymes, facilitating critical biochemical processes.20 The zinc(II) ion does not participate in oxidation–reduction reactions but its role in metalloenzymes is attributed to its ability to act as a Lewis acid.21 Zinc complexes have been studied for various applications in radioprotection, tumor photosensitizers, antidiabetic agents, anticonvulsants, and anti-inflammatory agents.22 Zinc complexes can also bind with serum albumins and DNA, and thus have potential anticancer activity. In the development of newer antibiotics to overcome the resistance of microbes, zinc complexes have shown promising results against bacteria and fungi, demonstrating potential antimicrobial activity.23 The escalating prevalence of multi-drug-resistant microorganisms, a direct consequence of antibiotic overuse and misuse, poses a significant threat to global health and socioeconomic stability.24 The inability of current antibiotics to effectively combat these infections underscores the importance of developing novel antibacterial agents.25 Despite decades of research, the development of such agents remains a pressing challenge. Among all bacteria accountable for deadly diseases, the treatment of infections caused by Bacillus subtilis and Salmonella enterica has caused a major challenge. B. subtilis is generally considered non-pathogenic and is widely utilized in industrial and probiotic applications. However, certain strains can pose health risks, particularly to immunocompromised individuals. In rare instances, B. subtilis has been associated with serious infections such as bacteremia, meningitis, and cerebral abscesses.26 Similarly, S. enterica is a major cause of foodborne illnesses worldwide. It is divided into numerous serotypes, many of which are associated with various health issues in humans and animals.27 Salmonellosis is the most common illness caused by S. enterica. Typhoid fever, paratyphoid fever, and chronic carrier state are also caused by S. enterica.28 Considering the urgency for innovative therapeutic approaches and to address this critical public health concern in the form of antibiotic resistance, a zinc(II) complex of the terpyridine ligand has been examined against bacterial strains in search of a drug as an antibacterial agent.1.1 Designing of the work
Limited work has been done on antibacterial activity against B. subtilis and S. enterica, and a comprehensive mechanistic evaluation has not been done with any antibacterial agent against both bacterial strains. Over the past few years, researchers have developed many zinc complexes of different ligands. These reported complexes exhibited antibacterial activity against B. subtilis with MIC value >10 µg mL−1; also they showed their antibacterial potential towards other bacterial strains such as E. coli, S. aureus, E. faecalis, and P. aeruginosa, but up to now no zinc complexes have been reported that exhibit excellent antibacterial activity against B. subtilis and S. enterica with low MIC values (Table S1). Moreover, the antibacterial activity of terpyridine zinc complexes has not been explored in depth so far. To address the limited exploration of the antibacterial activity of terpyridine zinc complexes and to fill up this gap, we have synthesised a more drug resistant and less toxic zinc terpyridine complex. Here, in this work, we have synthesized zinc(II) complex ZnL2 of 4′-(3,4-diethoxyphenyl)-2,2′:6′,2″-terpyridine ligand L in a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 ratio (Scheme 1). The zinc complex ZnL2 has been characterized by NMR, UV-vis, HRMS, and SC-XRD studies. The prepared ZnL2 has been examined for antibacterial potential against four Gram-positive and four Gram-negative bacterial strains. It showed good antibacterial activity against B. subtilis and S. enterica with low MIC values of 1.56 µg mL−1 and outperformed the marketed drugs and was further assessed for its mode of action via membrane disruption, protein leakage, biofilm inhibition, metabolic dysfunction, oxidative stress damage, and lipid peroxidation. The interaction of ZnL2 with calf thymus (ct)-DNA was also studied with spectroscopic techniques to build DNA-targeting antibacterial agents. Furthermore, the binding behavior of ZnL2 with human serum albumin (HSA), a transportation protein, was explored to obtain initial information about the drug's ADME (absorption, distribution, metabolism, and excretion) properties.
2 ratio (Scheme 1). The zinc complex ZnL2 has been characterized by NMR, UV-vis, HRMS, and SC-XRD studies. The prepared ZnL2 has been examined for antibacterial potential against four Gram-positive and four Gram-negative bacterial strains. It showed good antibacterial activity against B. subtilis and S. enterica with low MIC values of 1.56 µg mL−1 and outperformed the marketed drugs and was further assessed for its mode of action via membrane disruption, protein leakage, biofilm inhibition, metabolic dysfunction, oxidative stress damage, and lipid peroxidation. The interaction of ZnL2 with calf thymus (ct)-DNA was also studied with spectroscopic techniques to build DNA-targeting antibacterial agents. Furthermore, the binding behavior of ZnL2 with human serum albumin (HSA), a transportation protein, was explored to obtain initial information about the drug's ADME (absorption, distribution, metabolism, and excretion) properties.
2. Results and discussion
2.1 Chemistry
The ligand 4′-(3,4-diethoxyphenyl)-2,2′:6′,2″-terpyridine L was synthesized by the reaction of two equivalents of 2-acetylpyridine and one equivalent of 3,4-diethoxybenzaldehyde in ammonia and methanolic KOH medium, according to the literature method.29 The addition of NaClO4 to a stirred reaction mixture of Zn(NO3)2·6H2O (1 equivalent) and L (2 equivalents) in MeOH afforded the yellow coloured zinc complex ZnL2 (Scheme 1). L and the complex ZnL2 were characterized by NMR, UV-visible, HRMS and FTIR analyses (Fig. S1–S7 in SI). The UV-visible absorption spectra of the zinc complex in acetonitrile show absorption peaks at 230 nm, 277 nm (π → π* transition), and 322 nm (attributed to metal-induced intra-ligand charge transfer).29 In HRMS, the molecular ion peak at m/z 959.2336 corresponds to [ZnL2](ClO4)+. The zinc complex has also been characterized by X-ray single crystal diffraction study. The stability of ZnL2 was measured by UV-visible spectroscopy in PB buffer at pH 7.4. The ZnL2 spectral peaks remained unchanged, apart from a slight variation in intensity after 48 hours at room temperature (Fig. S8).2.2 Crystal structure of the zinc(II) complex
Yellow-coloured single crystals of the zinc complex were obtained by slow diffusion of di-isopropyl ether into an acetonitrile–methanol (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) solution of the complex. The complex was crystallized in the monoclinic (P21/n) space group. X-ray crystallographic data and structural refinement parameters for the complex are reported in Table S2. The crystal structure of the complex consists of a [Zn(L)2]2+ cation (Fig. 1), two perchlorate anions, one solvated water molecule, and one solvated acetonitrile molecule. The crystal structure of the zinc complex confirms the anticipated 1
1) solution of the complex. The complex was crystallized in the monoclinic (P21/n) space group. X-ray crystallographic data and structural refinement parameters for the complex are reported in Table S2. The crystal structure of the complex consists of a [Zn(L)2]2+ cation (Fig. 1), two perchlorate anions, one solvated water molecule, and one solvated acetonitrile molecule. The crystal structure of the zinc complex confirms the anticipated 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 metal to ligand stoichiometry in the cation. The dicationic charge on the complex is balanced by the presence of two perchlorates as counter anions. The zinc ion in the crystal structure shows a distorted octahedral geometry, bound by six pyridine nitrogen atoms of two ligands to give a ZnN6 type coordinated complex with meridional configuration. The central Zn–N bond distances [2.069(5)–2.082(5) Å] are shorter than terminal Zn–N bond distances [2.161(5)–2.204(5) Å] due to the restricted bite angle of the terpyridine core moiety. There is a slight deviation from the ideal octahedral geometry, as evidenced by the bond angle around the zinc ion [N2–Zn1–N5 171.8(2)°]. Selected bond distances and angles are presented in Table S3. Similar bond distances were observed in the previously reported zinc–terpyridine complexes.30,31 The central pyridine ring is attached to diethoxyphenyl showing a slight deviation from planarity with dihedral angles in the range of 7.9(10)–14.2(11)°. In ZnL2 it is observed that on one side of the ligand L, the central pyridine ring remains coplanar with both the terminal pyridine rings, while the other side of the ligand L is slightly tilted by a dihedral angle range of 5.9(11)–12.7(10)°. Three of the ethoxy groups remain coplanar with the phenyl, but one ethoxy group O3–C47–C48 is out of plane. The dihedral angles around the zinc ions for ZnL2 are N1–C5–C6–N2 = 3.6(8)°, N2–C16–C17–N3 = 1.4(8)°, N4–C30–C31–N5 = 6.8(8)°, and N5–C41–C42–N6 = 7.9(8)°.
2 metal to ligand stoichiometry in the cation. The dicationic charge on the complex is balanced by the presence of two perchlorates as counter anions. The zinc ion in the crystal structure shows a distorted octahedral geometry, bound by six pyridine nitrogen atoms of two ligands to give a ZnN6 type coordinated complex with meridional configuration. The central Zn–N bond distances [2.069(5)–2.082(5) Å] are shorter than terminal Zn–N bond distances [2.161(5)–2.204(5) Å] due to the restricted bite angle of the terpyridine core moiety. There is a slight deviation from the ideal octahedral geometry, as evidenced by the bond angle around the zinc ion [N2–Zn1–N5 171.8(2)°]. Selected bond distances and angles are presented in Table S3. Similar bond distances were observed in the previously reported zinc–terpyridine complexes.30,31 The central pyridine ring is attached to diethoxyphenyl showing a slight deviation from planarity with dihedral angles in the range of 7.9(10)–14.2(11)°. In ZnL2 it is observed that on one side of the ligand L, the central pyridine ring remains coplanar with both the terminal pyridine rings, while the other side of the ligand L is slightly tilted by a dihedral angle range of 5.9(11)–12.7(10)°. Three of the ethoxy groups remain coplanar with the phenyl, but one ethoxy group O3–C47–C48 is out of plane. The dihedral angles around the zinc ions for ZnL2 are N1–C5–C6–N2 = 3.6(8)°, N2–C16–C17–N3 = 1.4(8)°, N4–C30–C31–N5 = 6.8(8)°, and N5–C41–C42–N6 = 7.9(8)°.
 | ||
| Fig. 1 Crystal structures of the zinc complex ZnL2. Hydrogen atoms, counter anions and solvent molecules are omitted for clarity. | ||
2.3 Quantum chemical studies
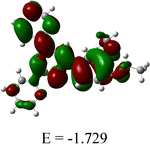
Quantum chemical studies are used to forecast the important pharmacokinetic features and the feasibility of building important interactions with target molecules.32,33 FMO (frontier molecular orbitals) control the interactions between molecules and target sites, and the energy difference between HOMO and LUMO aids in determining a compound's bioactivity. The HOMO and LUMO exert a positive influence on the ligand–receptor interaction process.33 The ligand and ZnL2 displayed energy gaps of 4.221 and 0.556 eV, respectively, indicating that both can exert good bioactivity (Table 1). The HOMO of the ligand is located on the phenyl ring attached with two ethoxy groups, manifesting that it interacts via electrostatic interactions with positively charged biological ions, and the LUMO of the ligand is located over the whole of the ligand, indicating that it binds with negatively charged residues. In the case of ZnL2, both HOMO and LUMO are located over the whole complex, indicating that the whole complex binds with negative as well as positively charged biological ions. The Cartesian coordinates of L and ZnL2 are given in Tables S4 and S5, respectively.2.4 In vitro antibacterial activity
Using a two-fold serial dilution procedure as recommended by CLSI (Clinical and Laboratory Standards Institute), the ligand L and its zinc metal complexes in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 (ZnL) and 1
1 (ZnL) and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 (ZnL2) ratio were evaluated for their ability to suppress bacterial growth.34,35 The ligand and complexes were tested against Gram-positive and Gram-negative bacterial strains, including Bacillus subtilis, Enterococcus faecalis, Listeria species, and Staphylococcus aureus, as well as Salmonella enterica, Escherichia coli, Serratia marcescens, and Acinetobacter calcoaceticus. As represented in Table 2, the ligand displayed moderate activity in inhibiting the growth of most of the tested bacterial strains. When the ligand was complexed with the zinc salt in the form of mono-complex ZnL, it exhibited better activity in suppressing the growth of most of the tested bacterial strains. This might be due to the change in the electron density of the ligand when complexed with Zn(NO3)2. The free nitrate group helped to increase the water solubility of the complex due to which it was able to exhibit better antibacterial properties than the ligand itself. Complex ZnL inhibited the growth of all the tested bacterial strains with MIC values ranging from 3.12 to 25 µg mL−1. The activity of ZnL was enhanced further when the free nitrate group was replaced with the same ligand, thus forming a complex of zinc and the ligand in 1
2 (ZnL2) ratio were evaluated for their ability to suppress bacterial growth.34,35 The ligand and complexes were tested against Gram-positive and Gram-negative bacterial strains, including Bacillus subtilis, Enterococcus faecalis, Listeria species, and Staphylococcus aureus, as well as Salmonella enterica, Escherichia coli, Serratia marcescens, and Acinetobacter calcoaceticus. As represented in Table 2, the ligand displayed moderate activity in inhibiting the growth of most of the tested bacterial strains. When the ligand was complexed with the zinc salt in the form of mono-complex ZnL, it exhibited better activity in suppressing the growth of most of the tested bacterial strains. This might be due to the change in the electron density of the ligand when complexed with Zn(NO3)2. The free nitrate group helped to increase the water solubility of the complex due to which it was able to exhibit better antibacterial properties than the ligand itself. Complex ZnL inhibited the growth of all the tested bacterial strains with MIC values ranging from 3.12 to 25 µg mL−1. The activity of ZnL was enhanced further when the free nitrate group was replaced with the same ligand, thus forming a complex of zinc and the ligand in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 ratio. ZnL2 with perchlorate ion as the counter ion exhibited excellent activity in suppressing the growth of the tested bacterial strains, with MIC values ranging from 1.56 to 6.25 µg mL−1, which were far superior to those of ZnL. The enhanced antibacterial activity of ZnL2 may be attributed to the presence of two electron-donating ligands attached to Zn, which increases the overall electron density of [ZnL2] (ClO4)2. Thus, it has been found that the free ligand exhibited moderate activity, ZnL displayed better activity than the ligand, whereas [ZnL2] (ClO4)2 exhibited excellent activity in inhibiting the growth of bacterial strains. The results were also compared to marketed drugs amoxicillin, tetracycline, chloromycin, and colistin, where [ZnL2] (ClO4)2 outperformed these marketed drugs. Comparison of the results of [ZnL2] (ClO4)2 with those of the modern membrane-targeting drug colistin showed that [ZnL2] (ClO4)2 remarkably outperformed colistin in inhibiting the growth of all the tested strains. Furthermore, to determine whether the antibacterial activity of the [ZnL2] (ClO4)2 complex arises primarily from the metal–ligand framework and is independent of the counterion, the ZnL2 complex was also evaluated with clinically accepted counterions, including Cl−, NO3− and SO42−. The results showed no significant change in antibacterial activity upon replacement of the perchlorate ion with these alternative anions, indicating that the observed antibacterial effect is intrinsic to the ZnL2 complex and not influenced by the nature of the counterion. Based on this counterion-independent antibacterial performance, [ZnL2] (ClO4)2 was selected for further detailed investigations. Subsequent studies focused on evaluating its antibacterial efficacy and elucidating its preliminary mechanism of action against Bacillus subtilis and Salmonella enterica, in view of its potent ability to inhibit bacterial growth.
2 ratio. ZnL2 with perchlorate ion as the counter ion exhibited excellent activity in suppressing the growth of the tested bacterial strains, with MIC values ranging from 1.56 to 6.25 µg mL−1, which were far superior to those of ZnL. The enhanced antibacterial activity of ZnL2 may be attributed to the presence of two electron-donating ligands attached to Zn, which increases the overall electron density of [ZnL2] (ClO4)2. Thus, it has been found that the free ligand exhibited moderate activity, ZnL displayed better activity than the ligand, whereas [ZnL2] (ClO4)2 exhibited excellent activity in inhibiting the growth of bacterial strains. The results were also compared to marketed drugs amoxicillin, tetracycline, chloromycin, and colistin, where [ZnL2] (ClO4)2 outperformed these marketed drugs. Comparison of the results of [ZnL2] (ClO4)2 with those of the modern membrane-targeting drug colistin showed that [ZnL2] (ClO4)2 remarkably outperformed colistin in inhibiting the growth of all the tested strains. Furthermore, to determine whether the antibacterial activity of the [ZnL2] (ClO4)2 complex arises primarily from the metal–ligand framework and is independent of the counterion, the ZnL2 complex was also evaluated with clinically accepted counterions, including Cl−, NO3− and SO42−. The results showed no significant change in antibacterial activity upon replacement of the perchlorate ion with these alternative anions, indicating that the observed antibacterial effect is intrinsic to the ZnL2 complex and not influenced by the nature of the counterion. Based on this counterion-independent antibacterial performance, [ZnL2] (ClO4)2 was selected for further detailed investigations. Subsequent studies focused on evaluating its antibacterial efficacy and elucidating its preliminary mechanism of action against Bacillus subtilis and Salmonella enterica, in view of its potent ability to inhibit bacterial growth.
| Comp. | Gram-positive bacterial strains | Gram-negative bacterial strains | ||||||
|---|---|---|---|---|---|---|---|---|
| E. faecalis | B. subtilis | L. species | S. aureus | E. coli | S. enterica | A. calcoaceticus | S. marcescens | |
| A – amoxicillin; B – tetracycline; C – chloromycin; D – colistin. | ||||||||
| Ligand | 50 | 25 | 50 | 12.5 | 50 | 100 | 12.5 | 25 |
| ZnL | 12.5 | 3.12 | 12.5 | 1.56 | 25 | 12.5 | 25 | 6.25 |
| [ZnL2] (ClO4)2 | 3.12 | 1.56 | 6.25 | 3.12 | 6.25 | 1.56 | 3.12 | 3.12 |
| [ZnL2] Cl2 | 3.12 | 1.56 | 6.25 | 3.12 | 6.25 | 1.56 | 3.12 | 3.12 |
| [ZnL2] (NO3)2 | 3.12 | 1.56 | 6.25 | 3.12 | 6.25 | 1.56 | 3.12 | 3.12 |
| [ZnL2]SO4 | 3.12 | 1.56 | 6.25 | 3.12 | 6.25 | 1.56 | 3.12 | 3.12 |
| A | 6.25 | 50 | 50 | 25 | 100 | 200 | 50 | 1.56 |
| B | 3.12 | 1.56 | 6.25 | 1.56 | 1.56 | 100 | 1.56 | 1.56 |
| C | 3.12 | 1.56 | 1.56 | 3.12 | 3.12 | 50 | 3.12 | 1.56 |
| D | 50 | 6.25 | 6.25 | 100 | 12.5 | 50 | 6.25 | 25 |
2.5 Bactericidal or bacteriostatic action
To ascertain whether the mode of action displayed by active ZnL2 towards B. subtilis and S. enterica is bacteriostatic or bactericidal, minimal bacterial concentrations (MBCs) were investigated following the completion of preliminary experiments. The active ZnL2 displayed MBC/MIC values of 2 and 1 against B. subtilis and S. enterica, respectively (Fig. 2a), suggesting that the complex is bactericidal to both the tested bacterial strains.2.6 Bacterial susceptibility evaluation
Antibiotic abuse has made treating bacterial illnesses extremely difficult since it has led to the development of bacterial resistance, which has reduced the effectiveness of present drugs and increased their dosage.36–38 Using tetracycline, amoxicillin, and chloromycin as positive controls, the drug resistance of ZnL2 was investigated at the MIC value against B. subtilis and S. enterica in up to 25 passages. According to the data shown in Fig. 2b and c, ZnL2 was found to be more effective than commercial antibiotics at delaying the onset of drug resistance against both the tested strains. The MIC values of tetracycline, amoxicillin, and chloromycin increased significantly after 9, 11, and 15 passages, respectively, whereas ZnL2 did not show any discernible change in MIC values until 20 passages against B. subtilis. Compared to this, in the case of S. enterica, ZnL2 delayed the occurrence of drug resistance up to 23 passages, while the MIC of marketed drugs tetracycline, amoxicillin, and chloromycin enhanced dramatically after 7, 10, and 13 passages, respectively. These results suggest that ZnL2 can overcome resistance in treating infections due to its low potential for generating drug resistance in B. subtilis and S. enterica.2.7 Cytotoxic studies of the complex
A key component of the drug candidate to develop as an antibacterial agent is its biosafety, which is evaluated by determining the cytotoxicity of the active drug candidates towards normal cell lines.39 Thus, MTT assay was employed to evaluate the cytotoxic effect of active drug candidate ZnL2 against normal cell line Hek293. At several doses ranging from 0.78 to 12.5 µg mL−1, ZnL2 exhibited only 2.81, 3.85, 4.53, 5.63 and 6.57% cytotoxicity towards normal cell line Hek293 (Fig. 2d). Also, we have evaluated the cytotoxicity of the free ligand towards normal cell lines to understand whether Zn(II) coordination improves the properties compared to the free ligand. The result suggested that the free ligand at various concentrations exhibited 15.2 to 35.4% cytotoxicity towards Hek293, indicating the decreased therapeutic value of the free ligand. Comparing the cytotoxicity results of the free ligand and ZnL2, the complex displayed very low toxicity to normal cell lines compared to the free ligand. The low cytotoxicity of ZnL2 to normal cells over bacterial cells suggested the biosafety of the complex and its potential for further research and development into antibacterial drugs. The cytotoxic experiment was conducted in triplicate.An essential parameter in evaluating the biosafety of synthesized ZnL2 is its propensity to induce hemolysis. In view of this, we examined the hemolytic activity of complex ZnL2 against goat red blood cells (RBCs) at various concentrations (MIC, 2 × MIC, 4 × MIC, and 8 × MIC), with Triton X-100 serving as the positive control. As depicted in Fig. 2e, ZnL2 maintained remarkably low hemolysis rates, which remained below 7% even after 3 h of exposure. Notably, at the highest tested concentration of 8 × MIC, complex ZnL2 induced hemolysis of only 6.2%. These values are well within the acceptable threshold for hemocompatible agents, underscoring its biosafety profile. The absence of significant RBC lysis in the presence of active complex ZnL2 highlights its therapeutic potential. This assay was conducted in triplicate to ensure reproducibility and statistical reliability.
2.8 Kinetics of bactericidal activity
The strong bactericidal effects of antibiotics inhibit the development of bacterial resistance. Therefore, a key test for determining the drug's bactericidal qualities at various concentrations is the time–kill kinetic study.40,41 Within three hours, B. subtilis and S. enterica in the control inoculum entered their exponential phase (Fig. 3a and b), whereas the growth of bacteria was inhibited and cells were destroyed upon treatment with ZnL2 inoculum. Within 4.5 and 3.5 h at MIC, ZnL2 began to exhibit its bactericidal activity against B. subtilis and S. enterica, respectively. At higher concentrations of ZnL2, such as 2 × MIC, 4 × MIC, and 8 × MIC, the time needed to kill bacteria in the inoculum decreased from 4.5 h to 3 h in the case of B. subtilis and from 3.5 to 1.5 h in the case of S. enterica. Consequently, ZnL2 had outstanding bactericidal activities in addition to good antibacterial properties, which shortened the treatment period. | ||
| Fig. 3 Time–killing kinetics of (a) B. subtilis and (b) S. enterica treated with ZnL2 at various concentrations. | ||
2.9 Biofilm inhibition assay
The development of biofilm on the surface of bacteria increases their pathogenicity and virulence and aids in developing bacterial resistance. Hence, bacteria are protected from antibiotic attack, reducing the bactericidal efficacy of antibacterial agents.42,43 These biofilms promote bacterial absorption and reproduction on the surface of medical equipment, making it more difficult to use it clinically.44 Using the crystal violet assay, we evaluated the ability of ZnL2 to suppress biofilm formation by B. subtilis and S. enterica. The inhibition of biofilm by active ZnL2 increases with the increase in concentration. ZnL2 was able to suppress the formation of biofilm of B. subtilis and S. enterica up to 24.3% and 30.2%, respectively, at the MIC value. As the concentration reached 8 × MIC, the complex inhibited biofilm formation by B. subtilis and S. enterica up to 75.2% and 87.1%, respectively (Fig. 4a). On the other hand, colistin inhibited the biofilm of B. subtilis and S. enterica 53.2% and 60.2%, respectively, at 8 × MIC (Fig. S9a). Comparing the biofilm inhibition ability of ZnL2 and colistin, the Zn complex displayed better biofilm inhibition properties than colistin towards both the tested strains. According to these findings, ZnL2 may prevent B. subtilis and S. enterica from forming biofilms, thus delaying the emergence of bacterial resistance. The same experiment was conducted in triplicate.2.10 Biofilm virulence factor
Combining the findings mentioned above, it is concluded that ZnL2 may successfully block the biofilm virulence factor that causes robust biofilm formation. It might also make it easier to break up the established biofilm, which would be beneficial for boosting the antibacterial activity and preventing the emergence of drug resistance.
2.11 Membrane disruption assay
By shielding the bacteria from external materials, their membranes perform a vital function in preserving the many metabolic events that occur in the right order.48 The protective barrier constructed by bacterial membranes is the primary cause of decreased antibiotic permeability, which changes the concentration of medications that enter the bacteria and has a direct impact on their effectiveness and therapeutic values leading to the development of drug resistance.49 Antibacterial agents are increasingly being developed with a focus on the cell membrane. We were motivated to investigate the potential of ZnL2 to disrupt the bacterial cell membrane further due to its excellent antibacterial efficacy, limited potential for drug resistance, and quick bactericidal properties.B. subtilis and S. enterica, with or without ZnL2, were stained with the EtBr dye and examined under a confocal laser scanning microscope to learn more about the antibacterial mechanism of the complex. When B. subtilis and S. enterica were treated with ZnL2, a strong red fluorescence was seen, further suggesting that ZnL2 could destroy B. subtilis and S. enterica cell membranes. In contrast, the control group in the absence of ZnL2 showed no fluorescence, as illustrated in Fig. 7d and e.
Combining the findings mentioned above, it is proposed that ZnL2 may efficiently harm the cell membrane of B. subtilis and S. enterica and induce membrane depolarisation, which in turn may trigger cell apoptosis and, eventually, cell death, contributing to its antibacterial function. All the above experiments were performed in triplicate.
2.12 Leakage of cytoplasmic content
The integrity of their cell membranes maintains the stability and biological activity of bacteria; any damage to these membranes allows the cytoplasmic content, including protein and nucleic acids, to escape, leading to cell death.53 Follin's test was employed to measure the amount of released protein from B. subtilis and S. enterica to verify the membrane damage caused by the ZnL2 complex. As the amount of ZnL2 increased, the concentration of leaked protein increased proportionately, as shown in Fig. 8a. As a result, the data showed conclusive evidence of membrane damage. Additionally, the concentrations of leaked nucleic acid from B. subtilis and S. enterica were measured to demonstrate the damage caused to the integrity of the membrane. ZnL2 exhibited a concentration-dependent increase in nucleic acid leaking from bacterial cells (Fig. 8b). The outcomes were consistent with membrane degradation. Both experiments were performed in triplicate. | ||
| Fig. 8 (a) Protein leakage and (b) DNA leakage from B. subtilis and S. enterica in the presence of ZnL2 at different concentrations. | ||
2.13 Metabolic dysfunction
Bacterial metabolism sustains both the development and survival of microorganisms. Through a variety of metabolic processes, bacteria produce vital macromolecules, maintain cellular homeostasis, and obtain energy.54 When cell membranes are disrupted, proteins on the surface become denatured, which results in metabolic dysfunction and ultimately cell death. The intracellular metabolic activity of B. subtilis and S. enterica in the presence of ZnL2 was assessed using the Almar blue test. After incubation with ZnL2, the metabolic activity of B. subtilis and S. enterica dropped dramatically in a concentration-dependent manner (Fig. 9a). ZnL2 reduced the metabolic activity of B. subtilis and S. enterica by 25.4% and 19.6% at 8 × MIC, respectively, whereas in the presence of colistin, the metabolic activities of B. subtilis and S. enterica were reduced to 40.1% and 35.5%, respectively, at 8 × MIC (Fig. S11a). The result suggested that reduction in metabolic activity in the presence of ZnL2 was more as compared to colistin. The experiment was conducted in triplicate. | ||
| Fig. 9 (a) Loss in metabolic activity; (b) intracellular ROS production; (c) loss in GSH activity and (d) malondialdehyde production in B. subtilis and S. enterica treated with ZnL2. | ||
2.14 Intracellular oxidative stress
2.15 Change in the morphology of B. subtilis and S. enterica cells
The bacterial morphological modification using scanning electron microscopic (SEM) images further validated the damage caused to the B. subtilis and S. enterica membrane by ZnL2. Untreated B. subtilis and S. enterica cells exhibited regular, symmetrical, undamaged, and rod-shaped cells. Both the bacteria showed cell wall rupture and cytoplasmic content leakage after being treated with ZnL2 at 2 × MIC (Fig. 10). Furthermore, ZnL2 was discovered to cause B. subtilis and S. enterica cells to shrink and break. These findings verified that ZnL2 caused membrane damage in B. subtilis and S. enterica.2.16 Interactions with DNA
Deoxyribonucleic acid (DNA), the storehouse of genetic information, is a frequently used molecular target for natural and synthetic drugs such as antitumor, antibacterial, and antiviral. Antibacterial drug development often targets bacterial DNA due to its critical role in cellular processes.59 DNA encodes the genetic instructions for protein synthesis, which is fundamental for bacterial metabolism and function. Thus, exploring the binding affinity of DNA with ZnL2 with the help of absorption and emission spectroscopy can help us to build a potent antibacterial agent.
Fig. 11a displays the UV-visible absorption spectra of the zinc complex ZnL2 upon addition of ct-DNA in aq. HEPES buffer. The complex ZnL2 shows absorption peaks at 230 nm, 277 nm and 322 nm. On incremental addition of ct-DNA to the complex solution, the 230 nm and 277 nm peak show a hyperchromic shift, whereas the 322 nm peak shows hypochromic shift. In addition, a new peak around 365 nm was also observed on incremental addition of ct-DNA. ct-DNA binding with the ligand L was also performed to check the effectiveness in comparison with ZnL2 (Fig. S12). The binding constant for ligand L with ct-DNA was found to be 3.04 × 105 M−1 (Fig. S13). The binding constant for the interaction of ZnL2 with ct-DNA was found to be 4.05 × 105 M−1 (calculated using the Benesi–Hildebrand equation (Fig. S14)). A fluorescence titration by varying the concentration of ct-DNA in a solution of ZnL2 was also performed to understand the interaction between ZnL2 and ct-DNA (Fig. 11b). The zinc complex exhibits weak fluorescence (λmax = 525 nm) (in HEPES buffer, pH 7.4) and on increasing the concentration of ct-DNA, the emission intensity of ZnL2 increases, which suggests the interaction between ZnL2 and ct-DNA (Fig. 11b). To assess the nature of complex's interaction with ct-DNA, a fluorescence titration was performed by incremental addition (0 to 75 µM) of the zinc complex to a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 solution of ct-DNA and ethidium bromide (EtBr) (intercalating agent) in HEPES buffer (pH 7.4). The EtBr–DNA complex exhibited emission maxima at 601 nm (λex = 480 nm) which decreased upon incremental addition of the zinc complex (Fig. 11c), due to the displacement of EtBr from the EtBr–DNA complex by the zinc complex. The results indicated that the complex binds to ct-DNA via an intercalative mode of interaction. The intercalation of ZnL2 into ct-DNA results in reduction of the available EtBr binding sites and further leads to a decrease in fluorescence intensity (Fig. 11c and Fig. S15). The reduction in fluorescence intensity indicates that ZnL2 binds to DNA and causes structural distortion at the tertiary or quaternary levels.
1 solution of ct-DNA and ethidium bromide (EtBr) (intercalating agent) in HEPES buffer (pH 7.4). The EtBr–DNA complex exhibited emission maxima at 601 nm (λex = 480 nm) which decreased upon incremental addition of the zinc complex (Fig. 11c), due to the displacement of EtBr from the EtBr–DNA complex by the zinc complex. The results indicated that the complex binds to ct-DNA via an intercalative mode of interaction. The intercalation of ZnL2 into ct-DNA results in reduction of the available EtBr binding sites and further leads to a decrease in fluorescence intensity (Fig. 11c and Fig. S15). The reduction in fluorescence intensity indicates that ZnL2 binds to DNA and causes structural distortion at the tertiary or quaternary levels.
2.17 HSA binding studies
The study of interaction of potential drugs with human serum albumin (HSA) is crucial for understanding various pharmacokinetic–pharmacodynamic properties, efficacy, and drug safety.60 HSA is the most abundant protein in human plasma, and it plays a key role in the transportation and distribution of drugs throughout the body. The extent of binding can significantly affect the distribution of drugs in the body.61 Highly bound drugs remain in the bloodstream longer, while those with low binding may diffuse more freely into tissues and organs. Spectroscopic techniques such as absorption and emission spectroscopy were employed to study the binding affinities of ZnL2 towards HSA.The absorption spectrum of HSA (10 µM) in PBS buffer shows an absorption peak at 280 nm, which is attributed to aromatic amino acids like phenylalanine (Phen), tyrosine (Tyr), and tryptophan (Trp).62 On gradual addition of ZnL2 (0–30 µM) to the HSA solution, a hyperchromic shift was observed at 280 nm along with a new peak at 332 nm (Fig. 12a). The absorption maxima of HSA at 280 nm remained unchanged even in the presence of ZnL2, suggesting that ZnL2 binds with HSA through non-covalent interactions.63 HSA binding with the ligand L was also performed to compare the efficiency with that of ZnL2 (Fig. S16 and S17). The binding constant (Kb) was determined by the Benesi–Hildebrand plot (Fig. S18) and was found to be 0.87 × 106 M−1.
 | ||
| Fig. 12 (a) Absorption and (b) emission spectra of HSA with increasing concentration of ZnL2 at 298 K in PBS buffer (pH 7.4). | ||
A fluorescence titration experiment was further carried out to study the mode of interaction between the complex and ligand with HSA (Fig. S19–S23). Upon excitation at 280 nm, the free HSA (10 µM) solution in phosphate buffer (pH = 7.4) showed an intrinsic fluorescent band at 350 nm because of the amino acid residues Tyr, Trp, and Phe. The fluorescence intensity of HSA at 350 nm was decreased gradually after the addition of ZnL2 (0–30 µM) to the solution of HSA (10 µM) in phosphate buffer (Fig. 12b). An isosbestic point was observed at 446 nm, representing the equilibrium between HSA and ZnL2 and the rise of a new peak at 507 nm. A slight red shift of 10 nm deviation at 343 nm was observed when the concentration of the complex was increased, which can be speculated due to the increase in the polarity of the microenvironment around the tryptophan residue. The emission intensity of HSA was suppressed entirely at increasing ZnL2 complex concentrations, demonstrating the effective binding of ZnL2 with HSA. When HSA interacts with a foreign ligand, the microenvironment changes, which causes quenching of the fluorescence intensity. The Stern–Volmer equation was utilised to determine the quenching mechanism. The quenching constant (Ksv) and the bimolecular quenching constant (Kq) (Fig. S22 and S23) were analysed and found to be 1.71 × 105 M−1 and 1.71 × 1013 M−1, respectively. As Kq is greater than the maximum value for dynamic quenching of biopolymers (2 × 1010 M−1 s−1), a ground-state complex is likely to be formed from the association of the compound with HSA, attributing to the static quenching mechanism. Furthermore, by calculating the binding constant values, the stability of complexes for the interaction of ZnL2 with HSA was evaluated. The binding constant parameters were thus determined using the modified Stern–Volmer equation and found to be 1.46 × 106 M−1 (Kb), indicating the effective binding of ZnL2 with HSA (Fig. S23). The binding stoichiometry was determined to be approximately 1, indicating that ZnL2 binds with HSA at a single site with 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 stoichiometry. Furthermore, Gibb's free energy was determined and found to be −8.38 kcal mol−1. The spontaneous binding process between ZnL2 and HSA was predicted by this negative value of ΔG, which suggested that hydrogen bonds and van der Waals interactions are in charge of the binding between ZnL2 and HSA. By consolidating the results from various studies, it was concluded that ZnL2 binds effectively to HSA, facilitating its efficient transport to the target site for combating bacterial infections.
1 stoichiometry. Furthermore, Gibb's free energy was determined and found to be −8.38 kcal mol−1. The spontaneous binding process between ZnL2 and HSA was predicted by this negative value of ΔG, which suggested that hydrogen bonds and van der Waals interactions are in charge of the binding between ZnL2 and HSA. By consolidating the results from various studies, it was concluded that ZnL2 binds effectively to HSA, facilitating its efficient transport to the target site for combating bacterial infections.
To determine the binding site of ZnL2 towards HSA, we performed the drug displacement studies with ibuprofen and warfarin as site markers. With incremental additions of ZnL2, the emission bands of complexes of HSA–warfarin and HSA–ibuprofen at 350 nm were quenched (Fig. 13). From the modified Stern–Volmer equation, the binding constant (Kb) for ZnL2–HSA in the presence of warfarin was found to be 1.46 × 106 L mol−1 (Fig. S24). In contrast, in the presence of drug ibuprofen, the binding constant for ZnL2–HSA was found to be 0.54 × 106 L mol−1 (Fig. S25) (Table 3). It has been observed that the binding constant of ZnL2 has been significantly decreased in the presence of site marker ibuprofen. These results demonstrated that ibuprofen prevents the binding of ZnL2, suggesting that ZnL2 binds to HSA Suldow's site II.
 | ||
| Fig. 13 Effect of ZnL2 on the emission spectra of the (a) warfarin–HSA complex and (b) ibuprofen–HSA complex. | ||
| ZnL2 | K b (106 L mol−1) | n |
|---|---|---|
| Blank | 1.46 | 1.00 |
| Warfarin | 1.40 | 1.21 |
| Ibuprofen | 0.54 | 1.10 |
Time-resolved emission studies were also conducted to determine the quenching mechanism responsible for the interaction of ZnL2 with HSA. According to steady-state emission studies, the binding interactions between HSA and ZnL2 are caused by static quenching. Time-resolved spectra of native HSA (10 µM) with and without the progressive addition of ZnL2 (0–30 µM) were recorded (Fig. S26). On addition of ZnL2, the native HSA's time decay profile decreased, suggesting the presence of dynamic quenching (Table 4). Combining the findings from time-resolved and steady-state fluorescence investigations, it is proposed that ZnL2 interacts with HSA through both static and dynamic quenchings.64
| System | Conc. | τ 1 [ns] | τ 2 [ns] | τ 3 [ns] | α 1 | α 2 | α 3 | Tav | χ 2 |
|---|---|---|---|---|---|---|---|---|---|
| HSA | 2.58 | 7.16 | 0.91 | 0.21 | 0.65 | 0.14 | 5.32 | 1.05 | |
| HSA-ZnL2 | 01![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 01 01 |
1.99 | 6.39 | 0.40 | 0.12 | 0.10 | 0.78 | 1.17 | 1.08 |
01![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 02 02 |
1.83 | 6.95 | 0.21 | 0.09 | 0.04 | 0.86 | 0.65 | 1.03 | |
01![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 03 03 |
1.44 | 6.20 | 0.08 | 0.04 | 0.02 | 0.95 | 0.24 | 1.07 |
3. Conclusion
As the threat of multidrug-resistant bacteria rises, the demand for new antibacterial drugs has become critical, but developing a drug with minimal resistance risk remains a considerable challenge. To address this issue, we have developed a zinc complex with 4′-(3,4-diethoxyphenyl)-2,2′:6′,2″-terpyridine (L) in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 ratio (ZnL2) as a promising antibacterial agent to tackle severe drug resistance. The synthesized ZnL2 was characterized by NMR, HRMS, FTIR, and SC-XRD studies. Initial bioactivity tests and structure–activity relationship analysis revealed that ZnL2 exhibited remarkable effectiveness in inhibiting bacterial growth. The complex showed excellent antibacterial activity against most of the tested bacterial strains with low MIC values ranging from 6.25 µg mL−1 to 1.56 µg mL−1 and outperformed the marketed drugs. ZnL2 significantly inhibited the biofilm formation, leading to a delay in the emergence of drug resistance. The complex ZnL2 effectively disrupts the bacterial membrane, reduces metabolic activity, and disrupts the biofilm virulence factor responsible for forming a strong biofilm, thus causing leakage of intercellular protein and nucleic acid contents. Additionally, bio-functional analysis reveals that ZnL2 can generate ROS within cells, which can lead to oxidative stress accumulation and GSH oxidation to GSSG, which in turn causes a decrease in GSH activity and oxidative cell damage. ZnL2 can easily transport to the target region and has good antibacterial activity because it may bind with DNA and HSA to form supramolecular complexes. Superior efficacy against bacterial strains, membrane disruption, drug-likeness, biofilm inhibition, oxidative stress damage, and metabolic dysfunction were all demonstrated by ZnL2 in comparison to marketed drugs. The complex ZnL2 can effectively overcome drug resistance and can be taken further for clinical trials for its development as a potent antibacterial agent.
2 ratio (ZnL2) as a promising antibacterial agent to tackle severe drug resistance. The synthesized ZnL2 was characterized by NMR, HRMS, FTIR, and SC-XRD studies. Initial bioactivity tests and structure–activity relationship analysis revealed that ZnL2 exhibited remarkable effectiveness in inhibiting bacterial growth. The complex showed excellent antibacterial activity against most of the tested bacterial strains with low MIC values ranging from 6.25 µg mL−1 to 1.56 µg mL−1 and outperformed the marketed drugs. ZnL2 significantly inhibited the biofilm formation, leading to a delay in the emergence of drug resistance. The complex ZnL2 effectively disrupts the bacterial membrane, reduces metabolic activity, and disrupts the biofilm virulence factor responsible for forming a strong biofilm, thus causing leakage of intercellular protein and nucleic acid contents. Additionally, bio-functional analysis reveals that ZnL2 can generate ROS within cells, which can lead to oxidative stress accumulation and GSH oxidation to GSSG, which in turn causes a decrease in GSH activity and oxidative cell damage. ZnL2 can easily transport to the target region and has good antibacterial activity because it may bind with DNA and HSA to form supramolecular complexes. Superior efficacy against bacterial strains, membrane disruption, drug-likeness, biofilm inhibition, oxidative stress damage, and metabolic dysfunction were all demonstrated by ZnL2 in comparison to marketed drugs. The complex ZnL2 can effectively overcome drug resistance and can be taken further for clinical trials for its development as a potent antibacterial agent.
4. Experimental section
4.1 Chemistry
The chemicals used for synthesis were purchased from SRL and Sigma Aldrich and used without any further purification. Ligand L was synthesised according to the reported method29 and the synthetic procedure mentioned in the SI. The 1H NMR spectrum was recorded on a 400 MHz Bruker NMR spectrometer to characterise the compounds. High resolution mass spectrometry (HRMS) of the synthesized analogues was performed using the Agilent 6545XT Advance Bio LC/Q-TOF. The FT-IR spectrum was recorded with a Bruker Hyperion 3000 FT-IR spectrometer. A Biotek ELISA plate was used for conducting bacterial studies. All the absorption studies were conducted on a Shimadzu UV-visible spectrophotometer. A HITACHI-F4600 fluorescence spectrophotometer was used for emission studies, and SEM was performed on a Carl Zeiss Sigma 500 FEG-SEM.4.2 Synthesis of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/h3_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/h3_char_2009.gif) 2 Zinc complex ZnL2
2 Zinc complex ZnL2
A methanolic solution of ligand L (0.2 mmol, 79.6 mg) and zinc nitrate hexahydrate (0.1 mmol, 29.7 mg) was heated at 60 °C and stirred for 2 h at room temperature. Further, sodium perchlorate solution (0.1 mmol, 12.20 mg) was added to it and stirred for another 2 h to obtain yellow precipitates. The precipitates were filtered, followed by washing with ethanol and distilled water. Slow diffusion of di-isopropyl ether into an acetonitrile–methanol (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) solution of ZnL2 gave yellow-coloured crystals. Yield (85%). HRMS [C50H46N6O4Zn + ClO4]+m/z 959.2336 (calcd. 959.7820); 1H NMR (400 MHz, DMSO-d6) δ/ppm 9.26 (s, 4H), 9.14–9.12 (d, 4H), 8.30–8.26 (t, 4H), 8.03–8.01 (d, 2H), 7.95–7.90 (m, 6H), 7.52–7.48 (t, 4H), 7.32–7.30 (d, 2H), 4.35–4.30 (q, 4H), 4.25–4.20 (q, 4H), 1.47–1.41 (dt, 12H). FT-IR (υmax/cm−1): 3593(H2O), 3093 (aromatic sp2-C–H str), 2982 (sp3-C–H asym str of –CH3), 2931 (sp3-C–H asym str of –CH2), 2874 (sp3-C–H sym str of –CH3), 1602, 1523, 1474, 1434 (aromatic C
1) solution of ZnL2 gave yellow-coloured crystals. Yield (85%). HRMS [C50H46N6O4Zn + ClO4]+m/z 959.2336 (calcd. 959.7820); 1H NMR (400 MHz, DMSO-d6) δ/ppm 9.26 (s, 4H), 9.14–9.12 (d, 4H), 8.30–8.26 (t, 4H), 8.03–8.01 (d, 2H), 7.95–7.90 (m, 6H), 7.52–7.48 (t, 4H), 7.32–7.30 (d, 2H), 4.35–4.30 (q, 4H), 4.25–4.20 (q, 4H), 1.47–1.41 (dt, 12H). FT-IR (υmax/cm−1): 3593(H2O), 3093 (aromatic sp2-C–H str), 2982 (sp3-C–H asym str of –CH3), 2931 (sp3-C–H asym str of –CH2), 2874 (sp3-C–H sym str of –CH3), 1602, 1523, 1474, 1434 (aromatic C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C, C
C, C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N str), 1407, 1257 (aromatic C–O asym str), 1206, 1153 (aliphatic C–O str), 1081 (aromatic C–O sym str), 859 (C–H bending, p-sub), 791 (C–H bending, m-sub), 689 (C–H bending, m-sub), 622.
N str), 1407, 1257 (aromatic C–O asym str), 1206, 1153 (aliphatic C–O str), 1081 (aromatic C–O sym str), 859 (C–H bending, p-sub), 791 (C–H bending, m-sub), 689 (C–H bending, m-sub), 622.
4.3 Quantum chemical studies
The 3D structures of ligand L and ZnL2 were built in the GaussView. The Gaussian 16W suite of programs was used to execute theoretical calculations of the ligand L and ZnL2, and these were optimized with DFT calculations using the B3LYP level of the 6–311++G(d,p) basis set for C, O, N, and H, whereas the LANL2DZ basis set was used for zinc atoms with no symmetry constraints in the gas phase. The frontier molecular orbitals, namely the highest occupied molecular orbital (HOMO) and the lowest unoccupied molecular orbital (LUMO), were generated using the cubegen module and visualized with Chemcraft software. These orbital plots were used to qualitatively examine the electronic distribution and nature of frontier orbitals, providing supportive insight into the electronic characteristics of ligand L and the ZnL2 complex.Author contributions
S.S.: synthesis, characterisation, and methodology; S. G.: conceptualization, methodology, and drafting; P.S.: data curation; D.D.: data curation; K. P.: visualization, investigation, supervision and editing; B.N.G.: conceptualization, investigation, supervision and editing.Conflicts of interest
The authors declare no competing financial interest.Data availability
Data shall be available on request.Supplementary information (SI) is available. Supporting information: materials and methods; crystallographic data (CCDC 2424188); additional tables and figures of related biological studies; spectral and other characterization data for key target compounds. A separate.zip folder containing the raw data images for Fig. 7(d) and (e) is provided – the software used for image analysis is FV-10 ASW 4.2 Viewer. See DOI: https://doi.org/10.1039/d5ma00502g.
CCDC 2424188 contains the supplementary crystallographic data for this paper.65
Acknowledgements
The authors acknowledge IIT Patna for providing the NMR data and NIT Rourkela for the HRMS data. The authors also acknowledge DST and SC-XRD Lab, SAIF-IIT Madras and crystallographer Dr Shobhana Krishnaswamy for data collection, structure solution and refinement. The authors S.S., P.S. and D.D. extend their gratitude to NIT Silchar for the Institute Fellowship and the facilities provided. K.P. thanks the SERB, New Delhi (CRG/2023/004080), and CEEMS (Project No: TIET/CEEMS/Regular/2021/018), VT-India, for providing funds.References
- A. Kumar, A. K. Singh, H. Singh, V. Vijayan, D. Kumar, J. Naik, S. Thareja, J. P. Yadav, P. Pathak and M. Grishina, Nitrogen Containing Heterocycles as Anticancer Agents: A Medicinal Chemistry Perspective, Pharmaceuticals, 2023, 16(2), 299 Search PubMed.
- J. Hu, W. Wu and C. Yang, Recent Advances in the Synthesis and Applications of Nitrogen-Containing Macrocyclic Arenes, Molecules, 2025, 30(17), 3646 Search PubMed.
- P. V. Ledade, T. L. Lambat, J. K. Gunjate, P. K. P. G. Chopra, A. V. Bhute, M. R. Lanjewar, P. M. Kadu, U. J. Dongre and S. H. Mahmood, Nitrogen-Containing Fused Heterocycles: Organic Synthesis and Applications as Potential Anticancer Agents, Curr. Org. Chem., 2023, 27(3), 206–222 CrossRef CAS.
- R. Ranjith, The Chemistry and Biological Significance of Imidazole, Benzimidazole, Benzoxazole, Tetrazole and Quinazolinone Nucleus, J. Chem. Pharm. Res., 2016, 8(5), 505–526 Search PubMed.
- R. Pal, G. S. P. Matada, G. Teli, M. Saha and R. Patel, Therapeutic Potential of Anticancer Activity of Nitrogen-Containing Heterocyclic Scaffolds as Janus Kinase (JAK) Inhibitor: Biological Activity, Selectivity, and Structure–Activity Relationship, Bioorg. Chem., 2024, 107696 CrossRef CAS PubMed.
- R. Musiol, P. Malecki, M. Pacholczyk and J. Mularski, Terpyridines as Promising Antitumor Agents: An Overview of Their Discovery and Development, Expert Opin. Drug Discovery, 2022, 17(3), 259–271 CrossRef CAS PubMed.
- K. Velugula, A. Kumar and J. P. Chinta, Nuclease and Anticancer Activity of Antioxidant Conjugated Terpyridine Metal Complexes, Inorg. Chim. Acta, 2020, 507, 119596 Search PubMed.
- S. Behera, P. P. Dash, A. K. Bishoyi, K. Dash, P. Mohanty, C. R. Sahoo, R. N. Padhy, M. Mishra, B. N. Ghosh and H. Sahoo, Protein Interactions, Molecular Docking, Antimicrobial and Antifungal Studies of Terpyridine Ligands, J. Biomol. Struct. Dyn., 2023, 41(20), 11274–11285 CrossRef CAS PubMed.
- D. Chao and Y. Zhang, Aggregation Enhanced Luminescent Detection of Homocysteine in Water with Terpyridine-Based Cu2+ Complexes, Sens. Actuators, B, 2017, 245, 146–155 CrossRef CAS.
- K. Velugula and J. P. Chinta, Silver Nanoparticles Ensemble with Zn(II) Complex of Terpyridine as a Highly Sensitive Colorimetric Assay for the Detection of Arginine, Biosens. Bioelectron., 2017, 87(Ii), 271–277 CrossRef CAS PubMed.
- S. F. Kainat, M. B. Hawsawi, E. U. Mughal, N. Naeem, A. M. Almohyawi, H. M. Altass, E. M. Hussein, A. Sadiq, Z. Moussa and A. S. Abd-El-Aziz, Recent Developments in the Synthesis and Applications of Terpyridine-Based Metal Complexes: A Systematic Review, RSC Adv., 2024, 14(30), 21464–21537 RSC.
- D. Das, S. Sutradhar, K. Rissanen and B. N. Ghosh, Synthesis and Structure of Trimethylplatinum(IV) Iodide Complex of 4′-(4-Methoxyphenyl)-2,2′:6′,2″-Terpyridine Ligand and Its Halogen Bonding Property, Z. Anorg. Allg. Chem., 2020, 646(5), 301–306 CrossRef CAS.
- B. N. Ghosh, M. Lahtinen, E. Kalenius, P. Mal and K. Rissanen, 2, 2′: 6′, 2 ″-Terpyridine Trimethylplatinum (Iv) Iodide Complexes as Bifunctional Halogen Bond Acceptors, Cryst. Growth Des., 2016, 16(5), 2527–2534 CrossRef CAS.
- S. Sutradhar, D. Das and B. N. Ghosh, Copper(II) and Cadmium(II) Triggered Hydrogelation of a Simple Trimethoxy Terpyridine Ligand, J. Mol. Struct., 2022, 1265, 133442 CrossRef CAS.
- S. Sutradhar, S. Basak, D. Das and B. N. Ghosh, Hydrogelation Behaviour of Methoxy Terpyridine Ligand Induced by Transition Metal Ions, Polyhedron, 2023, 236, 116344 CrossRef CAS.
- S. Sutradhar, G. K. Suresh, S. Sarma, D. Das, P. Sarkar, S. G. Patra and B. N. Ghosh, Sulphide and Iodide Anion Recognition by a Selective Copper Hydrogel of a Chloro Substituted Terpyridine Ligand, Inorg. Chem. Commun., 2024, 170, 113263 CrossRef CAS.
- S. Sutradhar, S. Sarma, S. Palo, P. Sarkar, T. K. Sarma and B. N. Ghosh, Copper-Induced Tert-Butoxy-Substituted Terpyridine Hydrogel for Enhanced Peroxidase-like Activity and Colourimetric Glutathione Sensing, New J. Chem., 2025, 49(38), 16713–16722 RSC.
- E. Athanasopoulos and J. Conradie, DFT Study of the Spectroscopic Behaviour of Different Iron(II)-Terpyridine Derivatives with Application in DSSCs, J. Mol. Graphics Modell., 2024, 129, 108753 CrossRef CAS PubMed.
- D. El-Sherbiny, H. Cheema, F. El-Essawy, A. Abdel-Megied and A. El-Shafei, Synthesis and Characterization of Novel Carbazole-Based Terpyridyl Photosensitizers for Dye-Sensitized Solar Cells (DSSCs), Dyes Pigm., 2015, 115, 81–87 CrossRef CAS.
- F. S. Al-Fartusie and S. N. Mohssan, Essential Trace Elements and Their Vital Roles in Human Body. Indian J Adv, Chem. Sci., 2017, 5(3), 127–136 CAS.
- G. Bahr; P. E. Tomatis and A. J. Vila, The Biochemistry and Enzymology of Zinc Enzymes, 2023 Search PubMed.
- M. Porchia, M. Pellei, F. Del Bello and C. Santini, Zinc Complexes with Nitrogen Donor Ligands as Anticancer Agents, Molecules, 2020, 25(24), 5814 CrossRef CAS PubMed.
- S. Nasiri Sovari and F. Zobi, Recent Studies on the Antimicrobial Activity of Transition Metal Complexes of Groups 6–12, Chemistry, 2020, 2(2), 418–452 CrossRef.
- J.-S. Zhao, N. Ahmad, S. Li and C.-H. Zhou, Hydrazyl Hydroxycoumarins as New Potential Conquerors towards Pseudomonas Aeruginosa, Bioorg. Med. Chem. Lett., 2024, 103, 129709 CrossRef CAS PubMed.
- G. Muteeb, M. T. Rehman, M. Shahwan and M. Aatif, Origin of Antibiotics and Antibiotic Resistance, and Their Impacts on Drug Development: A Narrative Review, Pharmaceuticals, 2023, 16(11), 1615 CrossRef CAS PubMed.
- I. Tsonis, L. Karamani, P. Xaplanteri, F. Kolonitsiou, P. Zampakis, G. Gatzounis, M. Marangos and S. F. Assimakopoulos, Spontaneous Cerebral Abscess Due to Bacillus Subtilis in an Immunocompetent Male Patient: A Case Report and Review of Literature, World J. Clin. Cases, 2018, 6(16), 1169 CrossRef PubMed.
- S. M. Jajere, A Review of Salmonella Enterica with Particular Focus on the Pathogenicity and Virulence Factors, Host Specificity and Antimicrobial Resistance Including Multidrug Resistance, Vet. world, 2019, 12(4), 504 Search PubMed.
- A. K. Bhunia and A. K. Bhunia, Salmonella Enterica, in Foodborne Microbial Pathogens: Mechanisms and Pathogenesis, 2018, pp. 271–287 Search PubMed.
- D. Das, S. Sutradhar, A. Singh and B. N. Ghosh, Zinc-Terpyridine Based Chemosensor for Detection of Pyrophosphate Anion in Aqueous Medium, Z. Anorg. Allg. Chem., 2021, 647(11), 1234–1238 CrossRef CAS.
- B. N. Ghosh, F. Topić, P. K. Sahoo, P. Mal, J. Linnera, E. Kalenius, H. M. Tuononen and K. Rissanen, Synthesis, Structure and Photophysical Properties of a Highly Luminescent Terpyridine-Diphenylacetylene Hybrid Fluorophore and Its Metal Complexes, Dalton Trans., 2015, 44(1), 254–267 RSC.
- B. N. Ghosh, R. Puttreddy and K. Rissanen, Synthesis and Structural Characterization of New Transition Metal Complexes of a Highly Luminescent Amino-Terpyridine Ligand, Polyhedron, 2020, 177, 114304 CrossRef CAS.
- Y.-M. Tan, J. Zhang, Y.-J. Wei, Y.-G. Hu, S.-R. Li, S.-L. Zhang and C.-H. Zhou, Cyanomethylquinolones as a New Class of Potential Multitargeting Broad-Spectrum Antibacterial Agents, J. Med. Chem., 2024, 67(11), 9028–9053 CrossRef CAS PubMed.
- T. J. Harrison, X. Chen, K. Yasoshima and D. Bauer, Phototoxicity—Medicinal Chemistry Strategies for Risk Mitigation in Drug Discovery, J. Med. Chem., 2023, 66(14), 9345–9362 Search PubMed.
- H. Sun, M. F. Ansari, N. Battini, R. R. Y. Bheemanaboina and C.-H. Zhou, Novel Potential Artificial MRSA DNA Intercalators: Synthesis and Biological Evaluation of Berberine-Derived Thiazolidinediones, Org. Chem. Front., 2019, 6(3), 319–334 Search PubMed.
- S.-F. Cui, D. Addla and C.-H. Zhou, Novel 3-Aminothiazolquinolones: Design, Synthesis, Bioactive Evaluation, SARs, and Preliminary Antibacterial Mechanism, J. Med. Chem., 2016, 59(10), 4488–4510 CrossRef CAS PubMed.
- Y.-G. Hu, N. Battini, B. Fang and C.-H. Zhou, Discovery of Indolylacryloyl-Derived Oxacins as Novel Potential Broad-Spectrum Antibacterial Candidates, Eur. J. Med. Chem., 2024, 270, 116392 Search PubMed.
- H. Huo, W. Dan, L. Qin, J. Bo, X. Zhang, C. Yang, B. Bai, J. Ren, B. Shi and J. Li, Novel Steroidal β-Carboline Derivatives as Promising Antibacterial Candidates against Methicillin-Resistant Staphylococcus Aureus, Eur. J. Med. Chem., 2025, 283, 117187 CrossRef CAS PubMed.
- H. Huo, W. Dan, M. Li, Y. Chen, C. Yang, L. Wu, B. Shi and J. Li, Design, Synthesis, and Biological Evaluation of Steroidal Indole Derivatives as Membrane-Targeting Antibacterial Candidates, Eur. J. Med. Chem., 2025, 283, 117156 CrossRef CAS PubMed.
- S. Rani and K. Paul, Triphenylethylene Analogues: Design, Synthesis and Evaluation of Antitumor Activity and Topoisomerase Inhibitors, Eur. J. Med. Chem., 2020, 208, 112775 CrossRef CAS PubMed.
- M. Shakir, A. Ali, S. Lakshmi, M. Garg, H. T. A. Almuqdadi, I. Irfan, M. Kamthan, M. C. Joshi, S. Javed and D. S. Rawat, Synthesis and Mechanistic Studies of 4-Aminoquinoline-Isatin Molecular Hybrids and Schiff's Bases as Promising Antimicrobial Agents, Eur. J. Med. Chem., 2025, 283, 117127 CrossRef CAS PubMed.
- Y. Yi, J. Zhang, S. Lin, H. Wang, G. Li, S. Yang, R. Shang, R. Zhang and F. Li, Design, Synthesis, and Biological Evaluation of Novel Pleuromutilin Derivatives with Methicillin-Resistant Staphylococcus Aureus-Targeting Phenol Linker Groups, Eur. J. Med. Chem., 2025, 282, 117061 CrossRef CAS PubMed.
- W. Guo, Z. Yang, K. Wang, W. Li, Y. Zhao, Y. Yang, W. Chang, Z. Gong, Z. Liu and Y. Chen, Discovery of Unique Bis-Substituted Aromatic Amide Derivatives as Novel Highly Potent Antibiotics for Combating Methicillin-Resistant Staphylococcus Aureus (MRSA), J. Med. Chem., 2024, 67(3), 2129–2151 CrossRef CAS PubMed.
- A. Vishwakarma; F. Dang; A. Ferrell; H. A. Barton and A. Joy, Peptidomimetic Polyurethanes Disrupt Surface Established Bacterial Biofilms and Prevent Biofilm Formation, 2021 Search PubMed.
- S. Guo, Q. Huang, Y. Chen, J. Wei, J. Zheng, L. Wang, Y. Wang and R. Wang, Synthesis and Bioactivity of Guanidinium-Functionalized Pillar [5] Arene as a Biofilm Disruptor, Angew. Chem., Int. Ed., 2021, 60(2), 618–623 CrossRef CAS PubMed.
- Q.-W. Lin, J.-Q. Lu, Y.-S. Huang, J.-J. Liu, W.-M. Chen and J. Lin, Cyclic Diguanylate G-Quadruplex Inducer-Nitric Oxide Donor Conjugate as a Bifunctional Antibiofilm Agent and Antibacterial Synergist against Pseudomonas Aeruginosa with a Hyperbiofilm Phenotype, J. Med. Chem., 2023, 66(17), 11927–11939 CrossRef CAS PubMed.
- O. Ciofu, C. Moser, P. Ø. Jensen and N. Høiby, Tolerance and Resistance of Microbial Biofilms, Nat. Rev. Microbiol., 2022, 20(10), 621–635 Search PubMed.
- J. Zhang, C. Chen, J. Chen, S. Zhou, Y. Zhao, M. Xu and H. Xu, Dual Mode of Anti-Biofilm Action of G3 against Streptococcus Mutans, ACS Appl. Mater. Interfaces, 2020, 12(25), 27866–27875 Search PubMed.
- S. Gupta and K. Paul, Membrane-Active Substituted Triazines as Antibacterial Agents against Staphylococcus Aureus with Potential for Low Drug Resistance and Broad Activity, Eur. J. Med. Chem., 2023, 258, 115551 CrossRef CAS PubMed.
- S. He, X. Li, Y. Wei, L. Zhang, J. Wang, Z. Yang and A. Shan, Cuminaldehyde Potentiates Antiproteolytic Peptide Efficacy via Parallel Pathways of Enhanced Inner Membrane-Damaging Activity and Inhibition of Bacterial Energy Metabolism, J. Med. Chem., 2024, 68(1), 776–791 CrossRef PubMed.
- S. Akhtar, M. M. Ansari, R. D. Verma, J. Sharma, A. Gupta, R. K. Dhuriya, D. P. Verma, J. Saroj, M. Ali and N. K. Verma, Generating a Peptide Library Using the Repeats of Amino Acid Scaffolds Created by Sliding the Framework of a 7-Mer Human Chemerin Segment and Discovery of Potent Antibacterial and Antimycobacterial Peptides, J. Med. Chem., 2024, 68(1), 566–589 CrossRef PubMed.
- J. Liu, H. Li, Q. He, K. Chen, Y. Chen, R. Zhong, H. Li, S. Fang, S. Liu and S. Lin, Design, Synthesis, and Biological Evaluation of Tetrahydroquinoline Amphiphiles as Membrane-Targeting Antimicrobials against Pathogenic Bacteria and Fungi, Eur. J. Med. Chem., 2022, 243, 114734 CrossRef CAS PubMed.
- S. Gupta, V. Luxami and K. Paul, Unlocking the Antibacterial Potential of Naphthalimide–Coumarins to Overcome Drug Resistance with Antibiofilm and Membrane Disruption Ability against Escherichia Coli, ACS Appl. Mater. Interfaces, 2025, 17(3), 4380–4399 CrossRef PubMed.
- J. Zhang, N. Battini, J.-M. Ou, S.-L. Zhang, L. Zhang and C.-H. Zhou, New Efforts toward Aminothiazolylquinolones with Multitargeting Antibacterial Potential, J. Agric. Food Chem., 2023, 71(5), 2322–2332 CrossRef CAS PubMed.
- X. Lu, X. Xu, Y. Ding, X. Gong, L. Ming, X. Dai, C. Gu, J. Wang, J. Zhao and M. Gao, Discovery and Optimization of Tetrahydroacridine Derivatives as a Novel Class of Antibiotics against Multidrug-Resistant Gram-positive Pathogens by Targeting Type I Signal Peptidase and Disrupting Bacterial Membrane, Eur. J. Med. Chem., 2025, 283, 117101 CrossRef CAS PubMed.
- F. He, X. Liu, S. Yang, H. Tan, L.-P. Yang and L.-L. Wang, Guanidinium-Functionalized Carbon Dots: An Efficient Antibacterial Agent against Multidrug-Resistant ESKAPE Pathogens, ACS Appl. Mater. Interfaces, 2024, 16(48), 65955–65969 Search PubMed.
- R. Yang, W. Cheng, M. Huang, T. Xu, M. Zhang, J. Liu, S. Qin and Y. Guo, Novel Membrane-Targeting Isoxanthohumol-Amine Conjugates for Combating Methicillin-Resistant Staphylococcus Aureus (MRSA) Infections, Eur. J. Med. Chem., 2024, 268, 116274 CrossRef CAS PubMed.
- Y.-F. Sui, M. F. Ansari, B. Fang, S.-L. Zhang and C.-H. Zhou, Discovery of Novel Purinylthiazolylethanone Derivatives as Anti-Candida Albicans Agents through Possible Multifaceted Mechanisms, Eur. J. Med. Chem., 2021, 221, 113557 CrossRef CAS PubMed.
- S. Roy, A. Mondal, V. Yadav, A. Sarkar, R. Banerjee, P. Sanpui and A. Jaiswal, Mechanistic Insight into the Antibacterial Activity of Chitosan Exfoliated MoS2 Nanosheets: Membrane Damage, Metabolic Inactivation, and Oxidative Stress, ACS Appl. Bio Mater., 2019, 2(7), 2738–2755 CrossRef CAS PubMed.
- S. Gupta and K. Paul, DNA Damage and Intercalation by Elinafide Modified Bis-Naphthalimides for Their Anticancer Activity, J. Mol. Liq., 2023, 382, 121980 CrossRef CAS.
- S. Gupta, V. Luxami and K. Paul, Deciphering Binding Potential of Naphthalimide–Coumarin Conjugate with c-MYC G-Quadruplex for Developing Anticancer Agents: A Spectroscopic and Molecular Modeling Approach, ACS Appl. Bio Mater., 2025, 8(2), 1077–1096, DOI:10.1021/acsabm.4c01388.
- P. Sharma and K. Paul, Selective Recognition of Oncogene Promoter C-Myc G-Quadruplex: Design, Synthesis, and In Vitro Evaluation of Naphthalimide and Imidazo[1,2-a]Pyrazines for Their Anticancer Activity, ACS Appl. Bio Mater., 2025, 8(2), 1377–1396 CrossRef CAS PubMed.
- P. Sarkar, S. Gupta, A. H. Udaya Kumar, D. Das, S. Sutradhar, K. Paul, N. K. Lokanath and B. N. Ghosh, Protein Interactions and Drug Displacement Studies of Novel Copper(II) and Zinc(II) Complexes of a Dipyrazinylpyridine Ligand, J. Mol. Liq., 2023, 387, 122561, DOI:10.1016/j.molliq.2023.122561.
- S. Gupta, V. Luxami and K. Paul, Bacterial Cell Death to Overcome Drug Resistance with Multitargeting Bis-Naphthalimides as Potent Antibacterial Agents against Enterococcus Faecalis, J. Mater. Chem. B, 2024, 12(23), 5645–5660 RSC.
- S. Rani, S. Gupta, V. Luxami and K. Paul, A Novel Target and Biomarker Benzothiazolyl–Naphthalimide Probes for Precise and Selective Detection of Serum Albumin and Anticancer Activity, New J. Chem., 2022, 46(25), 12082–12092, 10.1039/D1NJ03650E.
- CCDC 2424188: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2mckm6.
Footnote |
| † Equally contributed. |
| This journal is © The Royal Society of Chemistry 2026 |