Open Access Article
Open Access ArticleEngineering deuterated bacterial cellulose via biosynthesis for neutron applications
Amanda Muñoz-Juan†
a,
Daniela Diaza,
Judith Marin-Ortegaa,
Nerea Murugarren‡
a,
Mariano Campoy-Quiles a,
Felipe Zamorano
a,
Felipe Zamorano b,
Consuelo Guardiola
b,
Consuelo Guardiola b,
Martín Pérez§
b,
Martín Pérez§
 bc and
Anna Laromaine
bc and
Anna Laromaine *a
*a
aInstitut de Ciència de Materials de Barcelona, ICMAB-CSIC, Bellaterra 08193, Spain. E-mail: alaromaine@icmab.es
bInstitute of Microelectronics of Barcelona, IMB-CNM (CSIC), Bellaterra 08193, Barcelona, Spain
cConsejo Nacional de Investigaciones Científicas y Técnicas (CONICET), Buenos Aires, Argentina
First published on 8th May 2026
Abstract
Deuterated bacterial nanocellulose (dBNC) combines the biocompatibility and versatility of cellulose with the distinctive properties of deuterium, thereby facilitating the development of functional materials and their potential future applications in neutron science. We applied a scalable film-to-film biosynthesis protocol that reduces costs and time while achieving controlled deuterium incorporation into the biosynthesis of bacterial nanocellulose (BNC) using Komagataeibacter xylinus, adapted to D2O and deuterated glycerol. Spectroscopic analyses (FTIR, Raman, NIR, SR-µFTIR) confirmed the presence and homogeneous distribution of O–D and C–D bonds in the membrane. At the same time, physical properties, such as crystallinity and surface charge, remained comparable to those of native BNC. Neutron irradiation experiments demonstrated that dBNC films interact with fast neutrons, producing recoil deuterons, which supports their potential as biofriendly materials of interest for neutron science. This study paves the way for future use of dBNC as a promising material in neutron-based technologies and confirms the applicability of the optimized approach for its production and characterization.
Introduction
The pursuit of biocompatible materials is gaining momentum across various disciplines, with cellulose emerging as a particularly prominent material due to its natural abundance and properties, including biocompatibility, biodegradability, and the ability to be readily functionalized via its numerous hydroxyl groups.1 This biopolymer is being extensively investigated for applications in biomedicine, electronics, materials science, and food science.2–6 Furthermore, its deuterated variants are also being examined, enlarging the applications of this material.7–11 Initially, deuterated cellulose was used in fundamental research to elucidate cellulose structure and its interactions with other molecules. Following this strategy, deuterated cellulose has been used to gain insight into its interactions with biomolecules and solvents, thereby improving its use as a drug delivery system. More recently, it has been proposed as an anticounterfeiting material10 due to its different absorption properties in the infrared and near-infrared regions.Deuterated cellulose can be obtained through different chemical reactions, the main limitation of which is that only accessible hydrogens, those found in hydroxyl groups, are interchanged with deuterium.10,12 For that reason, more studies have focused on producing deuterated cellulose directly by culturing plants or microorganisms with deuterium-enriched nutrients, thereby incorporating deuterium as –CD, and –OD chemical bonds in the structure of cellulose.9,13 Bacteria are more resistant to deuterium toxicity than plants and grow faster, which speeds the production of deuterated cellulose.14 It has been reported that the production of deuterated bacterial cellulose (dBNC) is primarily achieved using Komagataeibacter xylinus and Gluconacetobacter hansenii strains.7,14 Some studies have opted to replace all nutrients with their deuterated counterparts completely.9,14 Although this approach increases the deuterium concentration in the final pellicle, it also significantly increases production costs and duration. Alternatively, other studies have substituted H2O with D2O and employed a deuterated carbon source, such as deuterated glucose or glycerol.7,11,15 In all cases, bacteria are acclimated to deuterated media by gradually increasing the concentration of deuterated compounds. The bacteria cultivated at each stage are subsequently used to inoculate the next medium with a higher deuterium concentration, typically via a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5 dilution and 50 ml volumes.9 Deuterium atoms form bonds that exhibit different vibrational frequencies, stability, and reactivity from hydrogen bonds.16 Therefore, dBNC has been used to understand polymer structure, biosynthesis, and to visualize biomacromolecules’ absorption.8,11,13,17
5 dilution and 50 ml volumes.9 Deuterium atoms form bonds that exhibit different vibrational frequencies, stability, and reactivity from hydrogen bonds.16 Therefore, dBNC has been used to understand polymer structure, biosynthesis, and to visualize biomacromolecules’ absorption.8,11,13,17
Additionally, the potential of deuterated polymers in areas such as active-layer materials for clean energy, future electronics, and neutron science is emerging.14,18–20 Among them, neutron moderators are materials used in neutron imaging or therapy that interact with neutrons, slowing fast neutrons (>1 MeV) and promoting recoil, while ideally avoiding capture.21 The slowed neutrons are used in various neutron-based techniques, including neutron imaging, small-angle neutron scattering (SANS), and neutron diffractometry.22,23 Neutron moderator materials contain small atoms with a high scattering cross-section. Therefore, H2O is widely used as a moderator material because hydrogen has the smallest atomic size. Other moderators include heavy water (D2O), sodium, and graphite.24 These materials have limitations depending on the application, such as a high reduction signal due to material self-absorption and limited tunability. One of the optimization strategies is to substitute hydrogen atoms with deuterium, which has a lower thermal neutron capture section (5.5 × 10−4 b vs. 0.33 b for hydrogen),25,26 which translates to a 600-fold reduction in self-absorption.27–32 Consequently, deuterated bacterial cellulose could be produced with different degrees of deuteration and final shapes, providing design flexibility in comparison with current materials used as moderators.25–27
In this study, we applied a protocol that minimizes the cost and time required to produce bacterial nanocellulose (BNC), named “film to film”.33 We evaluated whether it could be used to produce different degrees of deuterated bacterial nanocellulose (dBNC). Posteriorly, dBNC films were fully characterized, including the final and intermediate membranes, with special interest in techniques such as FT-IR, NIR, and Raman spectroscopy to quickly and non-destructively confirm the presence of deuterium. Finally, we evaluate the interaction of dBNC films with neutrons. Consequently, this paper aims to present and discuss experimental observations of deuterated cellulose, focusing on H/D effects and distribution, and to encourage further practical applications of deuterated polymers.
Results and discussion
Deuterium incorporation into BNC films during the adaptation process
The cellulose produced by bacteria, known as bacterial nanocellulose (BNC), can be made by different bacterial strains and carbon sources (Fig. 1A).15,34,35 The production of BNC is hindered by the biosynthesis process, which can take time and incur reagent costs, especially for deuterated cellulose. Deuterated glycerol (D-gly) is usually preferred over deuterated glucose9,11,17 because its metabolism acidifies the media less, increasing cellulose yield, and it is cheaper than deuterated glucose.14,36 To avoid the toxic effects of deuterium on bacteria while producing deuterated bacterial nanocellulose, K. xylinus is gradually acclimated to deuterium by culturing the bacteria in media with low deuterium levels and transferring those adapted to a medium with a higher deuterium concentration. To reduce production time and costs, we used the “film-to-film” technique applied for BNC.33 This technique involves placing freshly harvested BNC films rich in bacteria into fresh media, allowing the growth of a new membrane at the air-media interface (Fig. 1B).33 The production time depends on the bacteria concentration, the culture media volume, and the bacteria's acclimatization. The film-to-film technique offers two main advantages: the presence of adapted bacteria already in the film formed at the interface and the lower media volumes required, resulting in faster production time. In our hands, we adapted the bacteria into 2 ml of medium, and it took 7–10 days to harvest a new film from the culture surface. | ||
| Fig. 1 Synthesis of deuterated bacterial nanocellulose (dBNC). (A) Different carbon sources can be used to produce dBNC. (B) Deuteration process through the film-to-film protocol. | ||
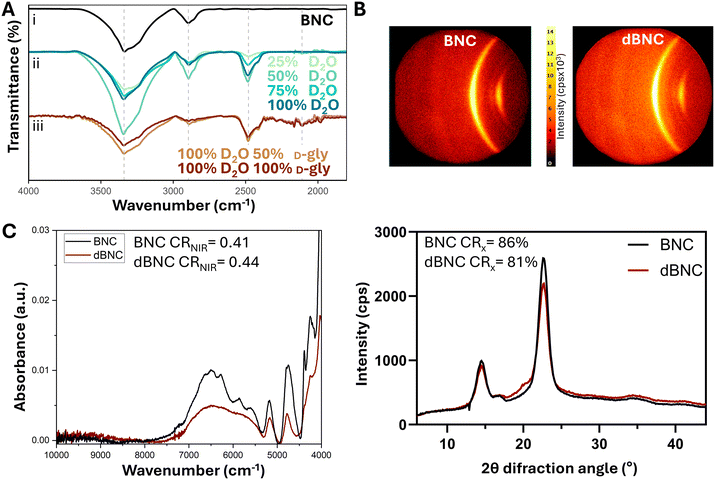
Using the film-to-film protocol, H2O was gradually replaced by deuterium oxide (D2O) in 25% increments until the medium reached 100% D2O. Films formed in this medium displayed the ν(O–D) at 2480 cm−1, not present in natural BNC films (Fig. 2A-i, ii). The ν(O–H) at 3340 cm−1 was also observed in dBNC, indicating that OD bonds did not fully replace hydroxyl groups.11 To compare trends in deuterium incorporation across different growth conditions, we calculated the ratio between the maximum absorption of OD (2480 cm−1) and OH (3340 cm−1) peaks to be interpreted as semi-quantitative indicators of relative deuterium incorporation levels. During the adaptation process, we observed that increasing D2O, we augmented the ratio (OD/OH) from 0.11 ± 0.05 (25% D2O growth media) to 0.48 ± 0.08 when cultured with medium with 100% D2O (Table 1).
| Ratios | Growth media | |||||
|---|---|---|---|---|---|---|
| 25% D2O | 50% D2O | 75% D2O | 100% D2O | 100% D2O 50% D-gly | 100% D2O 100% D-gly | |
| OD/OH A2480/A3340 | 0.11 ± 0.05 | 0.34 ± 0.05 | 0.29 ± 0.03 | 0.48 ± 0.08 | 0.65 ± 0.03 | 0.76 ± 0.10 |
| CD/CH A2110/A2895 | — | — | — | — | 1.27 ± 0.80 | 2.32 ± 0.70 |
Once bacteria adapted to grow in D2O, we introduced deuterated glycerol (D-gly) into the media in two steps, increasing its concentration by 50%. In both steps, we observed the O–D bond at 2480 cm−1 and the carbon–deuterium (C–D) bond at 2110 cm−1 (Fig. 2A-iii), which confirmed the incorporation of deuterium in the film.11 The ratio CD/CH was 1.27 ± 0.80 in dBNC films grown in medium with 50% D-gly and 100% D2O, indicating that nearly every C–H bond has a corresponding C–D bond. When D-gly was the main carbon source in the media (100% D-gly + 100% D2O), the ratio CD/CH increased to 2.32 ± 0.70, showing a rise in C–D bonds compared with C–H bonds. The presence of C–H and O–H bonds in deuterated films is expected, since there are still compounds in the media (e.g., peptone, citric acid, yeast extract) that contain hydrogen. A full substitution of C–H and O–H bonds by their deuterium counterparts is possible when all reagents are deuterated, but the cost of production also increases steadily.9,14
Physical properties of dBNC films
The incorporation of deuterium in BNC films could produce changes in cellulose's crystallinity. Cellulose is a polycrystalline structure with crystalline and amorphous regions, which was reflected in the general area detector diffraction system (GADDS) image of both natural and deuterated BNC (Fig. 2B). The X-ray diffraction pattern of both natural and deuterated BNC (Fig. 2B) exhibits the characteristic peaks of cellulose at 2θ = 14.5°, 16.5°, and 22.7° corresponding with planes (10), (110), and (200).17 According to the literature, we calculated the crystallinity index (CRX) by the Segal peak height method17,37,38 and found that BNC and dBNC have CRX values of 86% and 81%, respectively (Fig. 2B). Using near-infrared (NIR) spectroscopy, we also detected differences between BNC and dBNC spectra, especially in the 5500–7000 cm−1 region, where the O–H first overtone is present in cellulose.39 We calculated the crystallinity index (CRNIR) for BNC and dBNC following the method described by Inagaki, T. et al.39 by measuring the area under the peak curves of the amorphous region peaks at 6950 cm−1, and the crystalline regions at 6450 cm−1 and 6287 cm−1. BNC and dBNC have CRNIR values of 0.41 and 0.44, respectively (Fig. 2C). Although CRX and CRNIR are derived from different measurement principles and are therefore not directly comparable, a consistent correlation between them has been shown.39 Both methods indicate that the differences between BNC and dBNC fall within the error of the technique. In accordance with previous studies,17,39 these results indicate that BNC and dBNC present comparable levels of crystallinity within the resolution of the applied methods.Films were blended to obtain fibers, and their surface charge was characterized by measuring the Z-potential. Natural and deuterated BNC fibers exhibit moderate stability with similar Z-potentials, −17 ± 0.7 mV and −18 ± 0.4 mV, respectively. Therefore, deuterium incorporation did not affect the surface charge or the colloidal stability of these fibers.
Distribution of O–D and C–D chemical bonds
Raman Microspectroscopy and synchrotron radiation-based Fourier-transform infrared microspectroscopy (SR-µFTIR), two complementary techniques, were employed to assess the biodistribution of C–D and O–D chemical bonds within the samples. We used Raman spectroscopy to obtain a 25 × 25 point image, where each point corresponded to a single spectrum. The average spectrum is shown in Fig. 3A. The cellulose Raman spectrum is usually divided into three regions based on the chemical fingerprints of each region. Hydrogen bond (–OH) vibrations are found above 3000 cm−1;40 whereas carbon bonds are located between 1000–1600 cm−1, except for the –CH vibration that is found at 2900 cm−1.41,42 The third region between 1600–2900 cm−1 is denominated the silent region due to the absence of absorption peaks in protonated cellulose. In this last region, two peaks appear due to the deuteration process: the O–D bond at 2500 cm−1 and the C–D bond between 2000–2400 cm−1.16,43In our case, dBNC films presented one peak at ca. 2100 cm−1, which we assigned to the C–D bond,43 and another peak at ca. 2470 cm−1, assigned to O–D.16 Unlike FTIR, Raman spectroscopy could detect a weak signal from C–D bonds in the dBNC produced with just 100% D2O in the media. To analyze the distribution of the deuterated peaks in the sample, we performed a statistical analysis to quantify the number of pixels for each absorption intensity at each peak, obtaining histograms of intensities for O–D and C–D bonds (Fig. 3B); and we mapped the C–D and O–D peaks in a 25 × 25 point film image (Fig. 3C), observing a homogenous distribution of chemical bonds. The distribution of pixels for non-deuterated films is around zero, as expected. The incorporation of C–D and O–D bonds into the film shifts the histogram to the right, indicating a greater presence of pixels with higher absorption intensity at each deuterated peak (Fig. 3B). The average intensity of pixels at 2473 cm−1 (O–D bond) increased to 60 cts in films produced in 100% D2O media and to 150 cts in films produced in media with 100% D2O and 100% D-glycerol. The introduction of a deuterated carbon source increases the incorporation of O–D bonds, likely due to a more abundant deuterium environment. Reassuringly, the average intensity of the C–D bond increased by almost one order of magnitude when films were produced in media containing 100% D-gly, underscoring the importance of using a deuterated carbon source.
We complemented the previous spatial characterization of deuterated films by evaluating the spatial distribution of deuterated chemical bonds in deuterated fibers using SR-µFTIR analysis. The C–D and O–D peaks appeared at 2110 cm−1 and 2480 cm−1, respectively (Fig. 4A).11 The C–D intensity was low, preventing the acquisition of an image map. On the other hand, the O–D intensity peak enabled localization of O–D along dBNC fibers and complemented the previous Raman mapping. During sample preparation, a drop of dBNC fibers is placed on a CaF2 window and dried overnight under vacuum. At this step, fibers can deposit onto other fibers, increasing the amount of material analyzed. Therefore, single-fiber resolution is not achieved with this technique, and a larger sample amount, a darker sample (Fig. 4B), is correlated with a greater increase in O–D intensity in the map (red-orange region in Fig. 4C).
Neutron radiation fingerprint of deuterated cellulose
Natural and dBNC membranes were irradiated at the Centro Nacional de Aceleradores (CNA) with neutrons up to 8 MeV of energy, for 45 min, and compared them to evaluate their interaction with neutrons. The 50 µm thick films were placed on top of a 3 × 3 mm silicon carbide (SiC) detector, with a 50 µm epitaxial layer, as shown in the insight of Fig. 5A.44 The number of detected events is plotted in Fig. 5A as a function of energy for both BNC and dBNC, showing an increased number of detected counts in the range, compared to the bare detector. In the absence of a membrane, the detected signal arises from neutron interactions within the SiC active volume, primarily via elastic scattering with silicon and carbon nuclei. These interactions produce recoil Si and C ions; however, due to their relatively high mass, the energy transferred per collision is limited, resulting in a lower detected signal. When a membrane is placed on top of the detector, additional interaction channels are introduced through elastic scattering of neutrons with hydrogen or deuterium nuclei.The difference between BNC and dBNC is a shoulder observed in the 1.85–2.15 MeV range, which is attributed to the deuterium contribution, indicating an even higher ionization yield than hydrogen. This data confirms the presence of deuterium in dBNC membranes and its interaction with neutrons. Considering the nature of both films and the neutron field, the experimental setup was simulated using the Particle and Heavy Ion Transport code System (PHITS) code, based on the Monte Carlo method.45 The overall behavior observed in the simulations is consistent with the experimental trends (Fig. 5A), confirming that above 2 MeV dBNC membranes interact with neutrons, producing a higher number of counts per incident neutron than BNC membranes, indicating a stronger contribution from recoil deuterons than from recoil protons. This is consistent with the extended high-energy tail observed in spectra from the deuterated membrane.
The contribution of each particle to the total simulated spectrum of dBNC membrane is shown in Fig. 5B. Protons and deuterons are significantly lighter than silicon and carbon ions, which results in a more efficient energy transfer per collision. As expected from two-body elastic scattering kinematics, a larger fraction of the incident neutron energy can be transferred to these light nuclei. This process produces recoil protons and deuterons with relatively high kinetic energies, which in turn deposit more energy within the detector's active volume. In this configuration, the maximum energy transferred by neutrons to deuterons and protons is approximately 7.1 MeV and 8 MeV, respectively (Fig. S1). Their corresponding ranges in SiC are about 300 µm for protons and 150 µm for deuterons. This implies that both particles can traverse the entire active region of the detector. However, the linear energy transfer (LET) of deuterons is higher than that of protons (Fig. S1), resulting in a greater maximum energy deposition, as observed in Fig. 5B.
Conclusion
Cellulose deuteration offers significant potential to enhance material properties and broaden functional applications. However, traditional biosynthesis methods need refinement to boost efficiency and resource utilization. The film-to-film protocol introduced here effectively addresses these challenges by reducing the volume of deuterated media and accelerating deuteration. This method not only streamlines production but also enables detailed monitoring of bacterial adaptation during intermediate stages, an aspect largely underexplored in previous studies. Comprehensive spectroscopic analyses confirmed the successful incorporation of O–D and C–D bonds, indicating a uniform distribution of deuterium within the cellulose matrix. The partial retention of O–H and C–H bonds reflects incomplete substitution in the culture media, potentially offering a tunable balance between deuterated and non-deuterated components. Deuterium incorporation did not produce detectable changes in cellulose crystallinity within the resolution of the applied techniques. The preservation of crystallinity suggests that key structural features of native cellulose are retained. The manipulation of the deuterated films was qualitatively similar to that of the non-deuterated counterparts, indicating that the films seem to have similar mechanical properties.The neutron experiments confirmed the presence of deuterium in the membranes and the different recoil behavior with fast neutrons. The interaction between deuterium and fast neutrons opens avenues for their use in neutron-based analytical techniques, such as neutron imaging, and suggests the potential of these films for low-absorption applications as neutron-interacting material. This capability not only underscores the functional innovation enabled by the optimized biosynthesis method but also provides a basis for assessing scalability, encouraging further exploration of the practical applications of deuterated polymers, with broader implications for materials science and neutron technology.
Experimental
Materials and strains
Komagataebacter xylinus NCIMB 5346 (K. xylinus) was purchased from the Spanish Type Culture Collection (CECT, Spain).Standard Hestrin-Schramn (HS) media: 20 g l−1 of dextrose, 5 g l−1 of peptone, 5 g l−1 of yeast extract (Conda Lab.), 6.8 g l−1 of sodium phosphate dodecahydrate, and 1.15 g l−1 of citric acid monohydrate (Merck) and MilliQ water. For the cleaning process, ethanol (Panreac) and 0.1 M NaOH (Merck) were used.
Deuterated compounds: D2O and D-glycerol UD8-99% (Eurisotop).
Synthesis of deuterated BNC films
We followed the “Film-to-film” protocol.33 This consists of placing freshly harvested BNC films at the bottom of the recipient with HS media. We used 24-well plates containing 2 mL of standard HS medium or HS medium supplemented with deuterated compounds. Initially, we adapted K. xylinus to D2O by increasing the heavy water concentration by 25% at each step. BNC films rich in K. xylinus were placed at the bottom of a 24-well plate containing 2 ml of HS media with 25% D2O. After 7–10 days, another film was formed in the air–liquid interface. Following the same procedure, the newly formed film was used to adapt bacteria to HS media with 50% D2O. This step was repeated until adaptation to HS media with 100% D2O was achieved. Once bacteria were adapted to heavy water, we adapted bacteria to the deuterated carbon source, D-glycerol (D-gly). We adapted in two steps, 50% D-gly and 100% D-gly. The media for each step were sterilized using a 0.22 µm (MillexTM MP filter Sterile) filter and two 15 minute UV sterilization cycles. Bacterial cellulose films acquired the shape of wells in a 24-well plate, with a final area of approximately 1.93 cm2.Bacterial debris from the films was removed using a previously established protocol33,46 to characterize the membranes further. BNC films were introduced into 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ethanol
1 ethanol![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MilliQ water under stirring for 10 minutes, followed by 10 minutes in MilliQ water. Then, the films were subjected to two rounds of 20 minutes boiling in MilliQ water and two rounds of 20 minutes boiling in 0.1 M NaOH (Sigma). Finally, the films were transferred to MilliQ water baths to achieve a neutral pH (pH 6.5). Films were introduced in D2O and autoclaved at 121 °C, 30 minutes. Films were placed between Teflon surfaces, pressed with one kg weight, and dried at 60 °C overnight (17 h). dBNC fibers (dBNCf) were obtained by blending 1 mg dBNC films in 100 ml MilliQ water for 30 minutes with a commercial blender. Fibers were filtered from MilliQ water and redispersed into 5 ml D2O, yielding a final concentration of 200 µg ml−1.
MilliQ water under stirring for 10 minutes, followed by 10 minutes in MilliQ water. Then, the films were subjected to two rounds of 20 minutes boiling in MilliQ water and two rounds of 20 minutes boiling in 0.1 M NaOH (Sigma). Finally, the films were transferred to MilliQ water baths to achieve a neutral pH (pH 6.5). Films were introduced in D2O and autoclaved at 121 °C, 30 minutes. Films were placed between Teflon surfaces, pressed with one kg weight, and dried at 60 °C overnight (17 h). dBNC fibers (dBNCf) were obtained by blending 1 mg dBNC films in 100 ml MilliQ water for 30 minutes with a commercial blender. Fibers were filtered from MilliQ water and redispersed into 5 ml D2O, yielding a final concentration of 200 µg ml−1.
FT-IR
BNC and dBNC films obtained after each adaptation step were dried overnight at 60 °C and folded twice to enhance the signal from chemical bonds. Analysis was conducted using FTIR (JASCO FTIR 4700, Madrid, Spain) in transmission mode at a spectral resolution of 4 cm−1; 512 scans were averaged, and the measurement range was 400–4000 cm−1. The spectra were processed with Spectra Manager™ Suite software to reduce CO2 and H2O noise, perform baseline correction, and smooth the data. The spectrum shown is the average of three different scans. Ratios were calculated by converting transmittance values to absorbance, then dividing by the average absorbance for each chemical bond.NIR FTIR spectra were recorded on a Bruker Vertex 70 spectrometer equipped with a DLaTGS detector and a KBr beamsplitter. The spectral resolution was 4 cm−1, and 32 scans were averaged over a measurement range of 3000–9996 cm−1. Baseline correction and smoothing were applied using Origin software. To estimate the crystallinity in each sample, we followed the equation (eqn 1) proposed by Inagaki et al.39 The crystallinity index using NIR spectra (CRNIR) was calculated as follows:
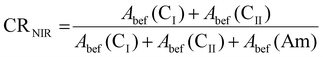 | (1) |
where Abef(CI) is the integral intensity of the OH band CI region (6429–6469 cm−1); Abef(CII) is the integral intensity of the OH band CII region (6267–6317 cm−1); and Abef(Am) is the integral intensity of the Am region (6949–7532 cm−1).
X-ray diffraction
The D8 Advance X-ray diffractometer (Bruker, Germany) with the two-dimensional (2D) Vantec-500 detector (GADDS-General Detector Diffraction System) and a KFL CU 2K (λ(CuKα) = 1.541840 Å) tube was used for X-ray characterization of natural and deuterated membranes. The diffraction patterns were obtained with 40 kV, 40 mA, with a scanner amplitude of 15.00 (300–900 s) and a collimator of 0.5 mm. Crystallinity index (CRX) was calculated using (eqn 2) according to the method described by Segal et al.38
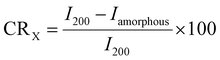 | (2) |
I200 represents the maximum intensity of the peak 2θ = 22.7°, which corresponds to the (200) plane.
Iamorphous represents the lowest intensity between the (200) plane and the (110) peak at 2θ = 14.5, which is the background, and it correlates with the amorphous region of cellulose.17,38
Raman spectroscopy
Dried BNC and dBNC films were cut into four parts and mounted, with 3 µl MilliQ drop between each piece. These sandwiches were dried at 60 °C for an additional 3 hours before evaluation. Raman spectroscopy was performed using Witec alpha300RA equipment, which features a solid-state laser at 785 nm to minimize luminescence, a 300 g mm−1 grating, and a 40× objective. The chemical mapping was created using 25 × 25 spectra, with an exposure time of 500 ms per spectrum and a power of 25 mW. The data was corrected for cosmic rays and background. The images were produced by integrating the areas under the deuterium-related peaks, representing the intensities of the C–D bond at 2148 cm−1 and the O–D bond at 2473 cm−1. Witec Project Five was used to analyze the data and perform the statistical analysis.SR-µFTIR
One drop of 3 µL of 68 µg mL−1 dBNCf was deposited onto a CaF2 window and dried under vacuum at RT for 16 h. Samples were kept under vacuum until the measurement day to avoid water absorption. A Hyperion 3000 microscope coupled to a Vertex 80 spectrometer (Brucker, Billerica, MA) equipped with a 36× magnification objective was used for Synchrotron Radiation Fourier Transform Infrared microspectroscopy (SR-µFTIR) at the MIRAS beamline at ALBA Synchrotron (Cerdanyola del Vallés, Barcelona, Spain). The spectra were collected in transmission mode using an MCT detector at 4 cm−1 spectral resolution, 5 × 5 µm aperture dimensions. A focal plane array (FPA) sensor was used for imaging measurement using the internal source, co-adding 128 scans in the 900–4000 cm−1 range. Data was analyzed using OPUS 7.5 (Bruker), Unscrambler X 10.5, and MATLAB R2010b. The color scale ranges from the lowest intensity (blue) to the highest (pink).Zeta potential
The zeta potential of fibers was analyzed using a Zetasizer ULTRA (Malvern Instruments, UK) after dilution in MilliQ water at a concentration of 68 µg ml−1. Three measurements were performed, and the average Z potential was calculated.Neutron irradiation
The experiments were carried out at the HISPaNoS facility of the National Accelerator Center (CNA) in Seville.47 The membranes were evaluated through neutron irradiations using a silicon carbide (SiC) detector fabricated at the National Center for Microelectronics in Barcelona. This device is described in the references.44,48,49 Two irradiations, each with a duration of 45 minutes, were performed using a continuous fast neutron spectrum (up to 8 MeV), produced by the Be(d,n) nuclear reaction with 4 MeV deuteron beams.47 The detector was operated at a bias voltage of 200 V, resulting in an active thickness of 20 µm. During the first irradiation, the device was covered with a 50 µm-thick dBNC membrane, as shown in Fig. 5A. In the second irradiation, it was replaced with a non-deuterated membrane of similar characteristics. The detector response was simulated using the PHITS code, based on the Monte Carlo method.45Author contributions
Amanda Muñoz-Juan: writing – original draft, review & editing, validation, methodology, investigation, formal analysis, conceptualization. Daniela Díaz, Judith Marin-Ortega, and Nerea Murugarren: methodology, investigation. Mariano Campoy-Quiles: Raman experiment and analysis, review. Felipe Zamorano, Martín Pérez neutron experiments and formal analysis. Consuelo Guardiola, neutron experiments, formal analysis, and funding acquisition. Anna Laromaine: writing – original draft, review & editing, supervision, resources, funding acquisition, conceptualization, validation.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have influenced the work reported in this paper.Abbreviations
| BNC | Bacterial nanocellulose |
| dBNC | Deuterated BNC |
| D-gly | Deuterated glycerol |
| FT-IR | Fourier transform infrared |
| SR-µFTIR | Synchrotron radiation-based Fourier transform infrared microspectroscopy |
| HS | Hestrin-Schramn media |
| SiC | Silicon carbide |
| PHITS | Particle and heavy ion transport code system |
| LET | Linear energy transfer |
| SEM | Standard error of the mean |
| CRX | Cristallinity index from X-ray data |
| CRNIR | Cristallinity indez from NIR measurements |
| CNA | Centro nacional de aceleradores |
| b (unit) | Barn |
Data availability
All data supporting the findings of this study is available in the repository digital.csic. https://digital.csic.es/handle/10261/431274 with the DOI: https://doi.org/10.20350/digitalCSIC/18345.Acknowledgements
SR-µFTIR experiments were performed at the BL01-MIRAS beamline at ALBA Synchrotron with the collaboration of ALBA staff (ID:2020094625). We are grateful to Agustín Mihi (ICMAB-CSIC) for providing access to the NIR spectrometer, supervising the measurements, and guiding in data analysis. This research was supported by the Spanish Ministry of Science and Innovation through the RTI2018-096273-B-I00 and PID2021-122645OB-100 projects, ‘Severo Ochoa’ Program for Center of Excellence in R&D (CEX2019-000917), the Generalitat de Catalunya (2017SGR765 grant). A. M. J. acknowledges the Ph.D. scholarship (FPU18/05190) in the framework of the Biotechnology Ph.D. program of the UAB. A. M. J. and A. L. participate in the Spanish National Research Council (CSIC) interdisciplinary platform for sustainable plastics towards a circular economy (SusPlast), the Aerogels COST ACTION (CA 18125), the EPNOE network, Conexión Nanomedicina del CSIC, and Red Nanocare 2.0. M. P. thanks funding from the European Union Horizon 2022 research and innovation program under the Marie Sklodowska-Curie grant agreement No. 101106191. C. G. acknowledges the Subvenciones 2022 SUB-08 program from the Spanish Nuclear Safety Council (CSN) and the Spanish government through the ‘María de Maeztu Unit of Excellence’ accreditation (CEX2023-001397-M). Finally, the authors would like acknowledge the assistance of the ICMAB-CSIC Scientific & Technical Services: X-ray Diffraction Laboratory (Anna Crespi), Spectroscopy Lab and Soft Matter Unit/Unit 6 of the ICTS “NANBIOSIS”; and CNA for their technical support.References
- H. Seddiqi, E. Oliaei, H. Honarkar, J. Jin, L. C. Geonzon, R. G. Bacabac and J. Klein-Nulend, Cellulose, 2021, 28, 1893–1931 CrossRef CAS.
- P. Singh, H. Baniasadi, S. Gupta, R. Ghosh, S. Shaikh, J. Seppälä and A. Kumar, Int. J. Biol. Macromol., 2024, 278, 134402 CrossRef CAS PubMed.
- S. Yang, Q. Ding, Y. Li and W. Han, Int. J. Biol. Macromol., 2024, 259, 129203 CrossRef CAS PubMed.
- A. Muñoz-Juan, A. Assié, A. Esteve-Codina, M. Gut, N. Benseny-Cases, B. S. Samuel, E. Dalfó and A. Laromaine, Carbohydr. Polym., 2024, 331, 121815 CrossRef PubMed.
- Z. Wu, B. Wang, J. Li, R. Wu, M. Jin, H. Zhao, S. Chen and H. Wang, Nano Lett., 2022, 22, 8152–8160 CrossRef CAS PubMed.
- K. Nyamayaro, P. Keyvani, F. D'Acierno, J. Poisson, Z. M. Hudson, C. A. Michal, J. D. W. Madden, S. G. Hatzikiriakos and P. Mehrkhodavandi, ACS Appl. Mater. Interfaces, 2020, 12, 52182–52191 CrossRef CAS PubMed.
- V. S. Raghuwanshi, Y. Cohen, G. Garnier, C. J. Garvey and G. Garnier, Macromolecules, 2021, 54, 6982–6989 CrossRef CAS.
- T. Imai, M. Naruse, Y. Horikawa, K. Yaoi, K. Miyazaki and J. Sugiyama, Cellulose, 2023, 30, 8423–8438 CrossRef CAS.
- J. He, S. V. Pingali, S. P. S. Chundawat, A. Pack, A. D. Jones, P. Langan, B. H. Davison, V. Urban, B. Evans and H. O'Neill, Cellulose, 2014, 21, 927–936 CrossRef CAS.
- Y. Song, S. Liu, H. Ben, Y. Zhang, G. Han, A. J. Ragauskas and W. Jiang, Front. Plant Sci., 2021, 12, 709692 CrossRef PubMed.
- J. Su, V. S. Raghuwanshi, W. Raverty, C. J. Garvey, P. J. Holden, M. Gillon, S. A. Holt, R. Tabor, W. Batchelor and G. Garnier, Sci. Rep., 2016, 6, 36119 CrossRef CAS PubMed.
- Y. Song, X. Meng, W. Jiang, B. R. Evans, H. Ben, Y. Zhang, Y. Pu, S. V. Pingali, B. H. Davison, S. Zhang, G. Han and A. J. Ragauskas, Cellulose, 2022, 29, 4269–4286 CrossRef CAS.
- S. Cui, Y. Xie, X. Wei, K. Zhang and X. Chen, Ind. Crops Prod., 2022, 187, 115452 CrossRef CAS.
- H. O'Neill, R. Shah, B. R. Evans, J. He, S. V. Pingali, S. P. S. Chundawat, A. D. Jones, P. Langan, B. H. Davison and V. Urban, in Methods in Enzymology, Academic Press Inc., 2015, vol. 565, pp. 123–146 Search PubMed.
- V. S. Raghuwanshi, J. Su, C. J. Garvey, S. A. Holt, W. Raverty, R. F. Tabor, P. J. Holden, M. Gillon, W. Batchelor and G. Garnier, Cellulose, 2017, 24, 11–20 CrossRef CAS.
- Y. Song, W. Jiang, H. Ben, Y. Zhang, G. Han and A. J. Ragauskas, Cellulose, 2021, 28, 5351–5361 CrossRef CAS.
- G. Bali, M. B. Foston, H. M. O'Neill, B. R. Evans, J. He and A. J. Ragauskas, Carbohydr. Res., 2013, 374, 82–88 CrossRef CAS PubMed.
- A. Uali and A. Aslanbek, Cellulose, 2025, 32, 6463–6475 CrossRef CAS.
- M. Shao, J. Keum, J. Chen, Y. He, W. Chen, J. F. Browning, J. Jakowski, B. G. Sumpter, I. N. Ivanov, Y. Z. Ma, C. M. Rouleau, S. C. Smith, D. B. Geohegan, K. Hong and K. Xiao, Nat. Commun., 2014, 5, 4180 CrossRef PubMed.
- L. Li, J. Jakowski, C. Do and K. Hong, Macromolecules, 2021, 54, 3555–3584 CrossRef CAS.
- M. M. Hosamani, A. Vinayak, G. B. Hiremath, P. N. Patil and N. M. Badiger, Ann. Nucl. Energy, 2022, 171, 109045 CrossRef CAS.
- A. Radulescu, R. Biehl and A. Papagiannopoulos, J. Appl. Crystallogr., 2025, 58, 1582–1594 CrossRef CAS PubMed.
- B. An, Y. Deng, Z. Jin and S. Sun, Adv. Funct. Mater., 2025, 35, 2422522 CrossRef CAS.
- S. Ghosh, Comprehensive Materials Processing, Elsevier, 2nd edn, 2024, vol. 8, pp. 87–98 Search PubMed.
- B. Hamermesh, G. R. Ringo and S. Wexler, Phys. Rev., 1953, 90, 603–606 CrossRef CAS.
- L. Kaplan, G. R. Ringo and K. E. Wilzbach, Phys. Rev., 1952, 87, 785–786 CrossRef CAS.
- J. Terreni, M. Trottmann, R. Delmelle, A. Heel, P. Trtik, E. H. Lehmann and A. Borgschulte, J. Phys. Chem. C, 2018, 122, 23574–23581 CrossRef CAS.
- P. Boillat, J. Biesdorf, P. Oberholzer, A. Kaestner and T. J. Schmidt, J. Electrochem. Soc., 2014, 161, F192–F198 CrossRef CAS.
- M. Zarebanadkouki, E. Kroener, A. Kaestner and A. Carminati, Plant Physiol., 2014, 166, 487–499 CrossRef PubMed.
- M. Caresana, M. Ferrarini, G. P. Manessi, M. Silari and V. Varoli, Nucl. Instrum. Methods Phys. Res., Sect. A, 2013, 712, 15–26 CrossRef CAS.
- R. H. Olsher, H.-H. Hsu, A. Beverding, J. H. Kleck, W. H. Casson, D. G. Vasilik and R. T. Devine, Health Phys., 2000, 79, 170–181 CrossRef CAS PubMed.
- S. Yue, S. Tabbassum, E. H. Jaye, C. A. M. Anderson and L. H. Nie, Phys. Eng. Sci. Med., 2025, 48, 35–46 CrossRef PubMed.
- N. Murugarren, S. Roig-Sanchez, I. Antón-Sales, N. Malandain, K. Xu, E. Solano, J. S. Reparaz and A. Laromaine, Adv. Sci., 2022, 9, 2201947 CrossRef CAS PubMed.
- Y. Li, C. Tian, H. Tian, J. Zhang, X. He, W. Ping and H. Lei, Appl. Microbiol. Biotechnol., 2012, 96, 1479–1487 CrossRef CAS PubMed.
- S. S. Wang, Y. H. Han, J. L. Chen, D. C. Zhang, X. X. Shi, Y. X. Ye, D. L. Chen and M. Li, Polymers, 2018, 10, 963 CrossRef PubMed.
- C. Recsei, M. Cagnes, R. A. Russell, R. E. Sonstrom and T. Darwish, Org. Biomol. Chem., 2025, 23, 3380–3392 RSC.
- K. S. Salem, N. K. Kasera, M. A. Rahman, H. Jameel, Y. Habibi, S. J. Eichhorn, A. D. French, L. Pal and L. A. Lucia, Chem. Soc. Rev., 2023, 52, 6417–6446 RSC.
- L. Segal, J. J. Creely, A. E. Martin and C. M. Conrad, Text. Res. J., 1959, 786–794 CrossRef CAS.
- T. Inagaki, H. W. Siesler, K. Mitsui and S. Tsuchikawa, Biomacromolecules, 2010, 11, 2300–2305 CrossRef CAS PubMed.
- M. Szymańska-Chargot, J. Cybulska and A. Zdunek, Sensors, 2011, 11, 5543–5560 CrossRef PubMed.
- N. Gierlinger, Appl. Spectrosc. Rev., 2018, 53, 517–551 CrossRef PubMed.
- H. Satha, I. Kouadri and D. Benachour, J. Polym. Environ., 2020, 28, 1914–1920 CrossRef CAS.
- H. Kubota, K. Sakamoto and T. Matsui, Sci. Rep., 2020, 10, 16426 CrossRef CAS PubMed.
- M. Pérez, F. Zamorano, C. Fleta, B. Fernández, C. Guerrero, P. Godignon, G. Pellegrini, P. Pérez-Maroto and C. Guardiola, Nucl. Instrum. Methods Phys. Res., Sect. A, 2024, 1069, 169968 CrossRef.
- K. Niita, T. Sato, H. Iwase, H. Nose, H. Nakashima and L. Sihver, Radiat. Meas., 2006, 41, 1080–1090 CrossRef CAS.
- J. Singh, T. W. J. Steele and S. Lim, Biomater. Adv., 2023, 144, 213174 CrossRef CAS PubMed.
- M. A. Millán-Callado, C. Guerrero, B. Fernández, J. Gómez-Camacho, M. Macías and J. M. Quesada, Radiat. Phys. Chem., 2024, 217, 111464 CrossRef.
- M. Pérez, F. Zamorano, C. Fleta, C. Muñoz-Montplet, D. Jurado-Bruggeman, M. Jiménez, P. Guardia, G. Grabulosa-Morera, R. Morales-Pérez, G. Rius, P. Godignon, G. Pellegrini and C. Guardiola, Sci. Rep., 2025, 15, 30543 CrossRef PubMed.
- M. Pérez, J. J. Blostein, F. Zamorano, C. Fleta, J. Marín, G. Pellegrini and C. Guardiola, Sci. Rep., 2026, 16, 2322 CrossRef PubMed.
Footnotes |
| † Current position: Danish Research Institute of Translational Neuroscience - DANDRITE, Nordic-EMBL Partnership for Molecular Medicine. Aarhus University, Department of Molecular Biology and Genetics, 8000 Aarhus, Denmark. |
| ‡ Current position: Centro de Biología Molecular Severo Ochoa, CSIC-UAM, Nicolás Cabrera 1, Cantoblanco, 28049, Madrid, Spain. |
| § Current position: Comisión Nacional de Energía Atómica (CNEA), Av. Bustillo 9500, San Carlos de Bariloche, Río Negro, Argentina, and Instituto Balseiro, Universidad Nacional de Cuyo, Argentina. |
| This journal is © The Royal Society of Chemistry 2026 |