Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Design of tuneable, biocompatible anionic polymers by L-norvaline amino acid post-polymerisation modification of pentafluorophenyl acrylate copolymers
Michael Ringleb ab,
Michael Streiber
ab,
Michael Streiber ab,
Theresa M. Lutzc,
Lena Rotherc,
Elisa-Maria Bachingerc,
Jonas De Breuckc,
Christopher Kuenneth
ab,
Theresa M. Lutzc,
Lena Rotherc,
Elisa-Maria Bachingerc,
Jonas De Breuckc,
Christopher Kuenneth d,
Ulrich S. Schubert
d,
Ulrich S. Schubert abef,
Stefan Zechelabef,
Anja Traeger
abef,
Stefan Zechelabef,
Anja Traeger *ab and
Meike N. Leiske
*ab and
Meike N. Leiske *cg
*cg
aLaboratory of Organic and Macromolecular Chemistry (IOMC), Friedrich Schiller University Jena, Humboldtstrasse 10, 07743 Jena, Germany. E-mail: anja.traeger@uni-jena
bJena Center for Soft Matter (JCSM), Friedrich Schiller University Jena, Philosophenweg 7, 07743 Jena, Germany
cMacromolecular Chemistry, University of Bayreuth, Universitätsstraße 30, 95447 Bayreuth, Germany. E-mail: meike.leiske@uni-bayreuth.de
dFaculty of Engineering Science, University of Bayreuth, Universitätsstraße 30, 95447 Bayreuth, Germany
eHelmholtz Institute for Polymers in Energy Applications Jena (HIPOLE Jena), Lessingstraße 12-14, 07743, Jena, Germany
fHelmholtz-Zentrum Berlin für Materialien und Energie (HZB), Hahn-Meitner-Platz 1, 14109, Berlin, Germany
gBavarian Polymer Institute, Universitätsstraße 30, 95447 Bayreuth, Germany
First published on 12th February 2026
Abstract
Based on their high cytocompatibility and unique interactions with biological matter, polyanions are attracting increasing research interest. The hydrophilicity–hydrophobicity ratio of these substances exerts a significant influence on their applicability as delivery systems, i.e., for particle formation and their interaction with biological matter. In this study a library of anionic copolymers with tailored hydrophilicity–hydrophobicity ratio is generated: First, random copolymers of pentafluorophenyl acrylate (PFPA) and hydrophilic N-acryloylmorpholine (NAM) or hydrophobic methyl acrylate (MA), respectively, are synthesised. In a second step, the PFPA units are substituted by L-norvaline (Nva) via post-polymerisation modification. We found that the comonomer ratio within the polymer has an impact on the hydrophobicity of the copolymers – in particular the Nva moiety significantly influences the hydrophobicity. Moreover, all investigated polymers feature good biocompatibility, while an increase in NAM content only slightly affects the hemolytic activity. Furthermore, an exemplary tested NAM-copolymer shows fast intracellular uptake in just 4 h. Our results obtained with Nva modified copolymers of different hydrophobicity-hydrophilicity ratio indicate that those polyanions could have potential for further biomedical applications such as drug delivery.
Introduction
Polymeric (nano)carriers have massively influenced material and medicinal sciences, opening versatile application scenarios across the biomedical, agricultural and environmental fields.1–4 These delivery routes provide substantial advantages compared to traditional approaches, exemplarily improved biocompatibility, reduced toxicity and precisely controlled substance release kinetics.4–7 The nanocarrier systems owe their immense potential to the customisability of the polymeric structures such as different available surface chemistries, size distribution and structural adaptability.8,9 In this way, it is possible to design particles with exceptional properties for specific applications, such as crossing biological barriers while minimising side effects for the organism.4,5,9 Different studies highlight their potential in the field of drug delivery, cancer and gene therapy, as well as nanomedicine.4,10–12 For the effective design of polymeric nanocarriers, their amphiphilic properties play a vital role. They influence the formation, stability and loading capacity as well as the interaction with biological systems.7,13–15 A balance between hydrophilic and hydrophobic properties is crucial for enhanced biocompatibility.16–19 Recent studies have shown that even small variations in this amphiphilic ratio of a polymeric nanocarrier can massively affect the performance such as cell association, which, in turn, highlights the significance of the ability to precisely adjust this property.9,20,21An option to address this challenge is the utilization of post-polymerisation modification (PPM) methods. It shows significant potential for the introduction of functional diversity into polymers without changes to the polymeric backbone,22–26 degree of polymerisation or dispersity of the synthesised polymers.27–29 The utilisation of polymer libraries designed by PPM promotes the investigation of structure–property relationships as it allows for the interpretation of the influence of slight modifications of functional groups onto the properties.30 A variety of efficient and selective reactions for PPM have been established in the past.23,25,26,31–35 Using these methods the stability, hydrophobicity, stimuli-responsiveness and biological interactions, among other parameters, can be precisely tuned to investigate the structure–property–function relationships of polymeric nanocarriers.34,36,37 Of the various precursors for PPM, poly(pentafluorophenyl acrylate) (PPFPA) is among the most versatile.9,36–38 Exemplarily, PPFPA is applied to achieve functional polyacrylamides or polyacrylates.9,22 Recently Théato, Warren and coworkers presented a flow-platform for the automated synthesis and PPM of PPFPA by amines.39 Due to the activated ester moiety, PPFPA is easily convertible by nucleophilic substitution with (primary) amines promoting the introduction of diverse functional side chains.9,38,40–42 The synthesis of PPFPA-containing (co)polymers can be carried out by radical polymerisations, e.g., by reversible-deactivation radical polymerisation techniques such as reversible addition–fragmentation chain-transfer (RAFT) polymerisation to obtain polymers with a rather defined molar mass and a narrow dispersity.43–49
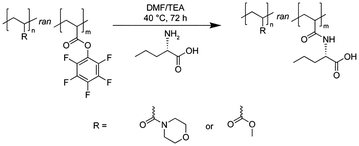
The innate reactivity of PPFPA towards nucleophiles such as amines renders it suitable for the PPM with peptides, proteins or amino acids to create a wide range of potentially bioactive polymers.9,42 Recently, our groups investigated the synthesis of PPFPA-derived polycarboxylates of varying hydrophobicity by PPM with different amino acids.9 Interestingly, more hydrophobic derivatives revealed unique cell membrane interactions and cellular uptake, which was attributed to the balance of anionic charge and hydrophobicity.9 Previously, it was not possible to tailor the pH-dependent membrane disruptive properties of these homopolymers, which are indispensable for intracellular endosomal release.7,49,50 Here, the use of copolymers might be beneficial as the combination of monomers exhibiting different characteristics results in materials with tailored properties such as adjusted charge density or hydrophobicity (i.e., amphiphilic copolymers).7,15 The design of copolymers of PFPA with comonomers such as N-acryloylmorpholine (NAM, hydrophilic) or methyl acrylate (MA, hydrophobic) constitutes a novel approach for the creation of adaptable polyanion platforms for nanocarrier design.
The current study aims to present amino-acid-derived anionic copolymers as a platform for the design of materials with tailored biological properties. By combining PFPA with either NAM or MA, the generation of libraries of polymers with precisely controlled compositions and, hence, hydrophobicity through modifications with amino acids (i.e., L-norvaline (Nva)) is enabled. Via this approach, it is shown that anionic copolymers can be further tailored to enhance the colloidal stability of ionic polymeric nanocarriers to modify the cell uptake of polyanions. The insights gathered by this systematic investigation will contribute to the understanding of structure–property relationships for polymeric nanocarriers, facilitating the rational design of materials.
Results and discussion
Synthesis of amino-acid-derived anionic copolymers
Recently, our groups demonstrated the modification of activated ester polymers with multiple amino acids of varying hydrophobicity to evaluate their membrane interactions and create polycarboxylates for drug delivery purposes.9 While anionic charges are known to be highly biocompatible, their cell interaction is commonly restricted due to charge repulsion with the negatively charged cell membrane.51 To balance this drawback, the inclusion of hydrophobic moieties is believed to be beneficial. Hydrophobic polyanions containing Nva moieties revealed unique cell membrane interactions and internalisation by eukaryotic cells. However, these polymers did not show any pH-dependent membrane disruptive properties, which are required for endosomal release. Based on these results, we decided to explore the potential of Nva-containing polycarboxylates further by creating a copolymer library with hydrophobic MA or hydrophilic NAM moieties.In detail, PFPA and a comonomer (i.e., NAM or MA) were copolymerised using 2-(dodecylthiocarbonothioylthio)-2-methyl propionic acid as chain-transfer agent (CTA) and 2,2′-azobis(2,4-dimethylvaleronitril) (V65) as initiator in anhydrous acetonitrile (see Scheme 1). Fig. 1A shows that the time-dependent monomer consumptions of NAM and PFPA did not differ drastically at equal monomer ratios (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) in the initial feed, thus, indicating the formation of random copolymers instead of the expected preferred incorporation of NAM. A potential reasoning for this could be connected to the different structure of the propagating chain ends which is influenced by both incorporated monomer moieties.52–54 A similar observation was made for copolymerisations with an excess of NAM (4
1) in the initial feed, thus, indicating the formation of random copolymers instead of the expected preferred incorporation of NAM. A potential reasoning for this could be connected to the different structure of the propagating chain ends which is influenced by both incorporated monomer moieties.52–54 A similar observation was made for copolymerisations with an excess of NAM (4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) (Fig. S4). This result is, furthermore, in accordance with a previous literature report applying another chain-transfer agent (CTA) for the RAFT copolymerisation of NAM and PFPA.47 For the hydrophobic MA as well, investigations on the copolymerisation kinetics with PFPA (1
1) (Fig. S4). This result is, furthermore, in accordance with a previous literature report applying another chain-transfer agent (CTA) for the RAFT copolymerisation of NAM and PFPA.47 For the hydrophobic MA as well, investigations on the copolymerisation kinetics with PFPA (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) (Fig. 1B) suggested the formation of random copolymers of these two monomers.
1) (Fig. 1B) suggested the formation of random copolymers of these two monomers.
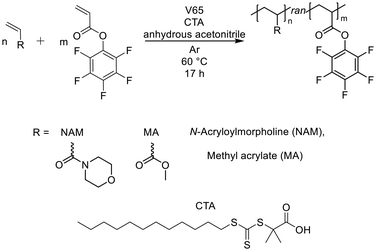 | ||
| Scheme 1 Schematic representation of the RAFT-copolymerisation of PFPA and a comonomer (NAM or MA) to synthesise a library of random copolymers. | ||
Since MA and NAM both formed random copolymers with PFPA these two monomers were confirmed to be ideal candidates as hydrophobic/hydrophilic comonomers for the preparation of amino-acid-derived polycarboxylates as the incorporation rate of the individual monomers correlates with the initial monomer feed.
Two libraries of random PFPA copolymers with narrow dispersity and a calculated degree of polymerisation (Xn,NMR) between ca. 150 to 250 (Table 1) were produced. The Xn,NMR was calculated for each individual monomer based on the 1H NMR (MA and NAM) or 19F NMR (PFPA) spectra directly after 17 h of polymerisation time.23,55 The monomer conversions for MA, NAM and PFPA were determined from their respective NMR spectra by comparing characteristic proton or fluorine signal integrals of monomer and polymer species according to eqn (1)–(3). For more details see the experimental part. These values were then used to calculate the degree of polymerisation per comonomer based on the initial monomer-to-CTA ratio as a basis for the determination of the NMR-derived molar mass (Mn,NMR) per comonomer. The final Xn,NMR and Mn,NMR were obtained by adding the values per comonomer. An overview of the conversions as well as the Xn,NMR and Mn,NMR for the individual comonomers is presented in the SI (Table S7). The targeted degrees of polymerisation (Xn,theo) were selected based on preliminary optimisation of the monomer-to-CTA ratio for each monomer system, as PFPA required higher ratios (260![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) for controlled polymerisation than MA or NAM (ca. 160
1) for controlled polymerisation than MA or NAM (ca. 160![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1). Consequently, copolymerisations were performed using compromise ratios, leading to systematic variations in Xn with copolymer composition, as reflected by the theoretical and experimental values in Table 1. In summary, two libraries of nine copolymers of differing composition were obtained. All copolymers exhibited low dispersity. The 1H NMR spectra of the polymers are presented in the SI (Fig. S8 and S9).
1). Consequently, copolymerisations were performed using compromise ratios, leading to systematic variations in Xn with copolymer composition, as reflected by the theoretical and experimental values in Table 1. In summary, two libraries of nine copolymers of differing composition were obtained. All copolymers exhibited low dispersity. The 1H NMR spectra of the polymers are presented in the SI (Fig. S8 and S9).
| Copolymer | Mn,appa (g mol−1) | Đa | Xn,NMR | Mn,NMR (g mol−1) | Xn,theo |
|---|---|---|---|---|---|
| a SEC in CHCl3/i-PrOH/NEt3 [94/4/2] (standard: PMMA). | |||||
| P(NAM10-ran-PFPA90)-CTA | 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600 600 |
1.42 | 241 | 54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
260 |
| P(NAM20-ran-PFPA80)-CTA | 29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
1.32 | 244 | 53![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
260 |
| P(NAM30-ran-PFPA70)-CTA | 26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.35 | 248 | 51![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600 600 |
260 |
| P(NAM40-ran-PFPA60)-CTA | 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 700 700 |
1.43 | 245 | 48![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
260 |
| P(NAM50-ran-PFPA50)-CTA | 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.35 | 209 | 39![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
220 |
| P(NAM60-ran-PFPA40)-CTA | 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.18 | 154 | 27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
160 |
| P(NAM70-ran-PFPA30)-CTA | 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600 600 |
1.23 | 165 | 28![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
169 |
| P(NAM80-ran-PFPA20)-CTA | 19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
1.25 | 157 | 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
160 |
| P(NAM90-ran-PFPA10)-CTA | 19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
1.23 | 158 | 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
160 |
| P(MA10-ran-PFPA90)-CTA | 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.30 | 240 | 54![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
260 |
| P(MA20-ran-PFPA80)-CTA | 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
1.32 | 239 | 52![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
260 |
| P(MA30-ran-PFPA70)-CTA | 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
1.4 | 221 | 46![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
240 |
| P(MA40-ran-PFPA60)-CTA | 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
1.26 | 221 | 44![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
240 |
| P(MA50-ran-PFPA50)-CTA | 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
1.29 | 210 | 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
230 |
| P(MA60-ran-PFPA40)-CTA | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
1.28 | 167 | 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
180 |
| P(MA70-ran-PFPA30)-CTA | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
1.27 | 164 | 28![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
180 |
| P(MA80-ran-PFPA20)-CTA | 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
1.31 | 166 | 26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
180 |
| P(MA90-ran-PFPA10)-CTA | 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
1.23 | 146 | 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
160 |
In our previous study, we demonstrated the successful PPM of PPFPA with various amino acids via transamidation of the activated ester moieties in the polymer side chain.9 Notably, this reaction also results in an aminolysis of the CTA Z-group to thiols and, consequently, may favour chain coupling by disulfide formation. However, this unwanted coupling was previously not observed. For this reason, initial PPM attempts of P(NAMn-ran-PFPAm) copolymers were conducted in a similar manner. Still, after purification via dialysis in aqueous media, SEC analysis revealed the formation of high molar mass byproducts (Fig. S10 and S11).
Interestingly, the coupling efficiency increased with increasing NAM-content in the copolymers. Since chain coupling was not observed before dialysis, we assume that an assembly of the copolymers in water during the dialysis favoured the proximity of thiol end-groups and, thus, chain coupling. Here, an increasing NAM-content may have led to an increased stability of assemblies due to better hydration of the outer shell.
While these results already indicated the potential of copolymers for nanoparticle preparation, uncontrolled chain-coupling would hinder the determination of structure–property relationships and, hence, had to be avoided. For example, when polymeric-bound, terminal CTA is present,9 polymer assembly occurred via intermolecular disulphide bridges. For this reason, in the current study, we permanently removed the Z-group by radical treatment as previously reported.56,57 The successful deprotection was illustrated by SEC measurements (Fig. S10 and S11). The traces with and without CTA measured by the refractive index (RI) hardly differed, whereas the absorption measurements (λ = 310 nm) of the eluted products – independent if P(NAMn-ran-PFPAm) or P(MAn-ran-PFPAm) – revealed that successful radical CTA removal remarkably reduced the absorbance signal. Confirmed by monomodal distribution of the polymers, bimodal curves – e.g., induced by chain coupling by intermolecular polymer reaction – were absent.56
For each individual polymer, we found qualitatively similar results when analysing the 1H NMR and FTIR spectroscopy data. The measurements are exemplarily shown for Nva-modified copolymers (1H NMR; Fig. 2A and B) and P(NAM10-ran-Nva-OH-AAm90) as well as P(MA10-ran-Nva-OH-AAm90) (FTIR; Fig. 2C and D), whereas all other spectra can be found in the SI (Fig. S12–S16). 1H NMR spectra provided information about the PPM efficiency of all copolymers. Here, the ratio of non-ionic moieties was used as a reference point. In particular, the cyclic methylene moieties of NAM (δ = 3.2 to 3.7 ppm) and the characteristic methoxy group of MA (δ = 3.6 ppm) were used. In both cases, the additional signals at δ = 4.0 ppm were obtained from the replacement of PFPA groups by Nva during PPM and were attributed to the CH group of Nva neighbouring the newly formed amide bond in alignment with our previous report.9 In addition, the terminal CH3 group of Nva was clearly observed at δ ≈ 1.0 ppm confirming the successful PPM by the amino acid. Once the PPM efficiency was calculated, the conversion of PFPA to Nva moieties was assessed to be ≥70% (Table 2). At this point, it is very important to emphasize that the calculation basis for the post-polymerization efficiency with Nva is derived from the copolymer compositions. This means that the percentual determination of the Nva modification depends on the PFPA repeating units present in each polymer.
| Copolymer | Mn,appa (g mol−1) | Đa | Nvab (%) | PPM efficiency (%) |
|---|---|---|---|---|
| a Determined via SEC in 0.07 M Na2HPO4 (calibration with poly(methacrylic acid) sodium salt homopolymers, PSS calibration kit).b Determined via 1H NMR (300 MHz) of postmodified copolymers in dPBS. | ||||
| P(NAM10-ran-Nva-OH-AAm90) | 36![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
1.27 | 64 | 71 |
| P(NAM20-ran-Nva-OH-AAm80) | 38![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.24 | 49 | 61 |
| P(NAM30-ran-Nva-OH-AAm70) | 37![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
1.20 | 61 | 88 |
| P(NAM40-ran-Nva-OH-AAm60) | 37![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
1.24 | 43 | 71 |
| P(NAM50-ran-Nva-OH-AAm50) | 28![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
1.24 | 37 | 74 |
| P(NAM60-ran-Nva-OH-AAm40) | 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
1.19 | 31 | 78 |
| P(NAM70-ran-Nva-OH-AAm30) | 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
1.13 | 22 | 74 |
| P(NAM80-ran-Nva-OH-AAm20) | 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
1.12 | 16 | 82 |
| P(NAM90-ran-Nva-OH-AAm10) | 9400 | 1.09 | 9 | 93 |
| P(MA10-ran-Nva-OH-AAm90) | 38![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
1.18 | 62 | 69 |
| P(MA20-ran-Nva-OH-AAm80) | 40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
1.17 | 66 | 82 |
| P(MA30-ran-Nva-OH-AAm70) | 33![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
1.09 | 60 | 85 |
| P(MA40-ran-Nva-OH-AAm60) | 32![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.17 | 53 | 89 |
| P(MA50-ran-Nva-OH-AAm50) | 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
1.21 | 45 | 89 |
| P(MA60-ran-Nva-OH-AAm40) | 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600 600 |
1.31 | 36 | 91 |
| P(MA70-ran-Nva-OH-AAm30) | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.41 | 29 | 97 |
| P(MA80-ran-Nva-OH-AAm20) | 8100 | 1.43 | 17 | 83 |
In addition to PPM with Nva only, polymers with the fluorescent label 6-aminofluoresceine (6-AF) were synthesised via sequential transamidation as previously reported.9 Fluorescent labelling played an important role in providing detailed information about, e.g., the polarity of respective synthesised polymers, later analysed in further assays.9 Similar to the Nva modification, the covalent interaction with amino acid after modification with 6-AF was found to be over 80% (Table 9). All copolymer datasets – whether modified with the fluorescent dye and amino acid or amino acid only – indicated an incomplete PPM efficiency below 100%. To ensure that the copolymers did not undergo hydrolysis during functionalization, this conversion data was additionally verified by FT-IR measurements. For this purpose, the difference between the absorption bands of relevant groups in the PFPA-based copolymer (P(NAM10-ran-PFPA90), P(MA10-ran-PFPA90)) and the post-modified Nva variant (P(NAM10-ran-Nva-OH-AAm90), P(MA10-ran-Nva-OH-AAm90)) was analysed (Fig. 2C and D).
In detail, the groups typical for PFPA, e.g., the carbonyl group at ṽ = 1780 cm−1 was right-shifted in the Nva-modified polymer (P(Nva-OH-AAm): ṽ = 1550 cm−1) and, thus, overlapped with the newly formed amide bond of Nva (ṽ = 1589 cm−1). In addition, Nva-modification was distinguished from PFPA-containing polymers by the free carboxylic acid group (ṽ = 1410 cm−1). Hydrolysis of Nva and 6-AF was absent since the characteristic peak for poly(acrylic acid) at ṽ = 1738 cm−1 was not visible, indicating the absence of unwanted hydrolysis. Dynamic light-scattering measurements of the polymers (data not shown) revealed self-assembly in aqueous media. We assume that this assembly impacted the NMR results. Since we use 10 to 15 mg mL−1 polymer per NMR measurement and the critical micelle concentrations (Fig. S16) range between those concentrations, the Nva modification might be shielded and accessibility is required for detection. With rising NAM content, an intensity increase in the peak at ṽ = 1410 cm−1 was detectable and, in comparison, the same holds true for MA variants (ṽ = 1750 cm−1).
Moreover, Nva-modified and Nva/6-AF labelled polymers were analysed with SEC measurements (Fig. 3 and Fig. S17). Monomodal distribution of NAM and MA copolymers was observed after PPM with Nva (Fig. 3) and Nva/6-AF (Fig. S17). Additionally, the absorbance signal during SEC measurements allowed for recording that the copolymers are successfully modified with the fluorescent dye. The molar mass variables, as well as the PPM modification efficiency obtained from the SEC data are listed in Table 2. Further SEC results are presented in the SI (Fig. S17).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P) based on the Wildman and Crippen scheme for estimating partition coefficients,59 at different pH-values (Fig. 4A). Furthermore, we performed high-performance liquid chromatography (HPLC) experiments and analysed the partition coefficients of 6-AF labelled copolymers in chloroform and water.
P) based on the Wildman and Crippen scheme for estimating partition coefficients,59 at different pH-values (Fig. 4A). Furthermore, we performed high-performance liquid chromatography (HPLC) experiments and analysed the partition coefficients of 6-AF labelled copolymers in chloroform and water.
A larger c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value indicates lower hydrophilicity of the polymers. The obtained data (Fig. 4B) showed that the computed c
P value indicates lower hydrophilicity of the polymers. The obtained data (Fig. 4B) showed that the computed c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value of protonated P(NAM10-ran-Nva-OH-AAm90) is approximately −88 and the hydrophilicity increases with higher NAM content (P(NAM90-ran-Nva-OH-AAm10) approximately −164) because of the impact of the protonated copolymeric Nva on the polymeric hydrophobicity is low. In comparison, deprotonated Nva has a strong effect on the polymeric hydrophobicity, i.e., the computed c
P value of protonated P(NAM10-ran-Nva-OH-AAm90) is approximately −88 and the hydrophilicity increases with higher NAM content (P(NAM90-ran-Nva-OH-AAm10) approximately −164) because of the impact of the protonated copolymeric Nva on the polymeric hydrophobicity is low. In comparison, deprotonated Nva has a strong effect on the polymeric hydrophobicity, i.e., the computed c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value increases from −328 (P(NAM10-ran-Nva-OH-AAm90)) to around −190 (P(NAM90-ran-Nva-OH-AAm10)). Becoming obvious from the increasing theoretical log
P value increases from −328 (P(NAM10-ran-Nva-OH-AAm90)) to around −190 (P(NAM90-ran-Nva-OH-AAm10)). Becoming obvious from the increasing theoretical log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value for deprotonated (around −20; 90% MA content) and protonated (around 6; 90% MA content) P(MAn-ran-Nva-OH-AAmm) by increasing the MA content, those polymers exhibited more hydrophobic properties. Similar to the NAM-based copolymers, the copolymeric Nva contributed more to the c
P value for deprotonated (around −20; 90% MA content) and protonated (around 6; 90% MA content) P(MAn-ran-Nva-OH-AAmm) by increasing the MA content, those polymers exhibited more hydrophobic properties. Similar to the NAM-based copolymers, the copolymeric Nva contributed more to the c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value (in deprotonated state) and thus to the hydrophobicity of the copolymers. The cheminformatic computations support experimental investigations which showed the influence of copolymeric Nva on the hydrophobicity properties as well.
P value (in deprotonated state) and thus to the hydrophobicity of the copolymers. The cheminformatic computations support experimental investigations which showed the influence of copolymeric Nva on the hydrophobicity properties as well.
In detail, HPLC measurements with an acetonitrile/water gradient acidified with TFA (pH ≈ 2) (see Table 3) revealed that NAM-based copolymers eluted earlier (Fig. 4C), while the MA-containing copolymers eluted later (up to ca. 21 min Fig. 4D). The variable content of NAM or MA within the copolymers did only affect the hydrophobicity minorly. However, the data revealed the following scenario: the carboxyl groups of Nva were protonated during the measurements at acidic pH-values. Thus, the copolymers were neutral, however, still hydrophobic and interacted more strongly with the column material via hydrophobic interactions. Only in NAM-based polymers with a high Nva content (>60%) the hydrophilic interaction of protonated carboxylic groups and thus the interaction with water is more pronounced. Overall, increasing MA content in combination with protonated Nva boosted the hydrophobic interactions with the column and thus, the polymers eluted simultaneously or later compared to their NAM counterparts. In comparison, the polymer hydrophobicity decreases with increasing NAM content which results in a shorter retention time and earlier elution from the column.
| Time (min) | Fraction of 0.1% (v/v) | Fraction of acetonitrile (%) |
|---|---|---|
| TFA in water (%) | ||
| 0 | 98 | 2 |
| 5 | 50 | 50 |
| 10 | 0 | 100 |
| 17 | 0 | 100 |
| 25 | 98 | 2 |
| 35 | 98 | 2 |
This HPLC data were further corroborated by measuring the partition coefficient of the individual polymers at pH = 4, as well as under physiological pH-conditions using DPBS to discuss the influence of the NAM, MA, and Nva groups on the polymeric hydrophobicity. Each polymer was dissolved in the corresponding aqueous solvent, mixed with organic chloroform and the fluorescence reduction in the water phase was quantified (Fig. 4E and F). The average fluorescence intensity reduction was more pronounced in acidic environment compared to experimental conditions at higher pH-values. Interestingly, the fluorescence decreased up to ∼90% with Nva contents <80%, i.e., the polymeric hydrophobicity state was hardly dependent on NAM (hydrophilic) and MA (hydrophobic), respectively. However, Nva revealed the strongest influence on hydrophobicity depending on the pH-value. A reason for this is that Nvas were protonated at acidic pH-values and the hydrophobic interaction was more pronounced. Thus, the copolymer solubility in chloroform was improved with increased Nva contents ≥50% since the fluorescence reduction in the water phase is not detectable anymore. In the case of DPBS, the contained salt promoted polymer–polymer interactions via charged Nva (ionic interactions) groups since the Debye shielding is weakened based on low salt concentrations. Hence, the hydrophilicity within the polymer was stronger and the fluorescence reduction less apparent with up to 40% fluorescence reduction for NAM-modified polymers and 80% fluorescence reduction for MA-modified polymers. A similar trend can be observed for both polymers. However, MA in combination with Nva affected the hydrophobicity of the individual copolymers more strongly since the influence of the more hydrophilic NAM was less pronounced.
Interaction of amino-acid-derived polyanions with cells
The copolymers investigated in this study were rationally designed by systematically modifying the amount of the anionic component. To evaluate their membrane interaction, erythrocytes were used as a model. For this purpose, blood from human donors was purified, and isolated red blood cells (RBCs) were incubated in DPBS with the polymers at two different pH-values (6.0 and 7.4). The extent of haemolysis was determined by measuring the released haemoglobin. Haemolysis levels were quantified using a Triton-X control (representing 100% haemolysis), with results categorized as follows: 0 to 2% haemolysis was considered non-haemolytic, 2 to 5% as mildly haemolytic, and values exceeding 5% as strongly haemolytic. No significant haemolysis was observed in these studies. Interestingly, the polymers with a decreasing proportion of anionic repeating units, such as P(NAM80-ran-Nva-OH-AAm20) and P(NAM90-ran-Nva-OH-AAm10), exhibited slight haemolysis at pH = 7.4 (Fig. 6A). This could be due to the enhanced hydrophobic character of the copolymers, forcing, membrane interaction, influencing their haemolytic potential. However, the balance between hydrophobicity and other structural factors likely plays a key role in determining the extent of haemolysis induced by such materials.60 E.g., despite their higher hydrophobicity, MA-modified polymers did not induce haemolysis. This unexpected result may be attributed, in part, to the reduced solubility of the corresponding copolymers in the cellular environment, which could limit direct membrane interaction. It is also noteworthy that no enhanced haemolysis was detected at pH = 6, emphasizing a favourable balance between anionic character and hydrophobicity. This is particularly remarkable, as anionic polymers typically exhibit stronger haemolysis under acidic conditions.61 This effect can be explained by the good solubility of these polymers due to their high NAM content, combined with the hydrophobic nature of the protonated carboxylic acid groups (Fig. 4).
In such copolymers, the small proportion of hydrophobic units can readily interact with the cellular membrane. In contrast, copolymers with a higher fraction of hydrophobic monomers tend to self-associate, forming aggregates in which the hydrophobic segments are partially shielded and, thus, less accessible to the membrane. This explains why the more water-soluble variants exhibit stronger membrane interactions than the more hydrophobic ones, even though this may seem counterintuitive at first. It can be assumed that this effect was not observed for copolymers with increasing MA content because their lower water solubility may limit the accessibility of hydrophobic regions to the membrane. This suggests that both the physicochemical properties of individual monomers and the overall polymer composition could be important factors in tuning membrane interactions.
In parallel, the aggregation behaviour was investigated as well. Alongside haemolysis, RBC aggregation represents another potential interaction between the polymers and the RBCs. In contrast to the cationic branched polyethyleneimine (b-PEI), which was used as positive control, the copolymers did not exhibit any aggregation behaviour (Fig. 6B–D and Fig. S18).
Thus, the copolymers demonstrated excellent biocompatibility and low amounts of MA and NAM within the polymer structure enable a pH dependent membrane interaction. The post-polymerisation reaction influenced the biological properties of P(Nva-OH-AAm) copolymers in such a way that a higher degree of functionalization with PNAM slightly altered its haemolytic properties. In future studies, efforts could be made to further enhance the anionic character to potentially enable pH-dependent membrane interactions.
As illustrated in Fig. 7A, the cells were seeded in 24-well plates and treated with the fluorescently labelled polymer. DPBS served as a negative control (NC). After 4 h of incubation, flow cytometry analysis was conducted to quantify the extent of polymer association with cells.
Representative, normalised fluorescence histograms (Fig. 7B and C) revealed a clear shift in fluorescence intensity in the cells treated with the polymer compared to the negative control, indicating successful interaction. Each histogram represents one biological replicate, compiled from three technical replicates. Quantitative analysis of mean fluorescence intensity (MFI) and the percentage of positive cells determined in the fluorescein isothiocyanate (FITC) channel (FITC pos. cells) (Fig. 7D and E) confirmed a statistically significant increase in fluorescence in the polymer-treated group compared to the control group (***p ≤ 0.001). These results suggest that the 6AF-labelled polymer efficiently associates with cells under the tested conditions. To complement the flow cytometry data, confocal laser scanning microscopy (CLSM) was employed to visualize intracellular polymer localization at the single-cell level. The cell membrane was stained red with CellMask and the nuclei were stained blue with Hoechst 33342. Single fluorescence channels are shown in Fig. S20. While control cells revealed no green fluorescence (Fig. 7F), polymer-treated cells displayed green-fluorescent signals (Fig. 7G), indicating the presence of the 6AF-labelled polymer. Moreover, CLSM observations proved an internalization of the polymer, as indicated by the green spots inside the cell. Together, these data confirm the strong cellular association of the P(NAM60-ran-Nva-OH-AAm40)-6-AF. While our previous study with P(Nva-OH-AAm)-6AF revealed fluorescence after 4 h primarily associated with the cell membrane of L929 cells,9 the copolymer investigated in this study exhibited a markedly different uptake profile. Distinct intracellular fluorescent spots were observed after just 4 h of incubation, suggesting a significantly accelerated and more efficient internalization mechanism.
Conclusion
Two libraries of copolymers were successfully synthesised from PFPA with hydrophilic (NAM) and hydrophobic (MA) comonomers. In comparison to the homopolymerisations of the respective comonomers, a reactivity change could be observed leading to roughly identical incorporation rates of the comonomers (PFPA and MA/NAM) and, hence, the synthesis of random copolymers. Successful PPM of the copolymers with Nva delivered two libraries of polyanions. Characterisation of the properties showed that the ratio of the non-PFPA comonomer did not have a significant influence on the hydrophobicity of the final polyanions, while the incorporation rate of Nva through PPM had a more pronounced effect, in particular at acidic pH-value. The copolymers exhibit excellent biocompatibility, with no visible toxicity to cells. Moreover, an exemplary NAM-copolymer was shown to associate with and internalize into cells within 4 hours. This highlights the potential of the presented copolymers for a wide range of applications, particularly in the field of anionic polymer-based delivery systems, such as drug delivery. Our observations further suggest that the balance between hydrophilic and hydrophobic monomers plays a critical role in mediating polymer–membrane interactions. Thus, a small fraction of hydrophobic units as part of a water-soluble polymer facilitates direct interactions with cellular membranes, whereas higher hydrophobic content could promote self-association and aggregation, reducing polymer–membrane interactions. This highlights that the physicochemical properties of individual monomers and overall polymer composition can be tuned to control cellular interactions.Experimental part
Materials and instrumentation
The solvents and chemicals for end group removal on the polymers and post-polymerisation reactions were obtained from the following manufacturers: 2,2′-azobis(2-methylpropionitrile) (AIBN; Fluka; 98% purity) purified by recrystallization from methanol (Fisher Scientific, 99.9%), chloroform (Fisher Scientific, 99.8%), diethyl ether (Fisher Scientific; 99.5%), dilauroyl peroxide (Luperox; Thermo Fisher Scientific; 97%), N,N-dimethylformamide (DMF; VWR; 99.9%), n-hexane (VWR; 95%), toluene (VWR; 99%), 6-amino-3′,6′-dihydroxy-3H-spiro[isobenzofuran-1,9′-xanthen]-3-one (6-AF; BLDpharm; 95%), Dulbecco's phosphate buffered saline (DPBS; VWR), L-norvaline (Nva; Carbolution Chemicals; 97%), and triethylamine (TEA; VWR; 99%).
For analytical methods and polymer characterisation studies 0.1 v/v% trifluoroacetic acid (TFA; VWR), hydrochloric acid (HCl; Carl Roth; 37% solution), sodium hydroxide (NaOH; Carl Roth; >98%), D2O (Deutero; 99.95% deuterium), DMSO-d6 (Deutero; 99.8% deuterium) and CDCl3 (Deutero; 99.95% deuterium) were used as purchased.
The following commercially available materials were used for the biological tests: Dulbecco's modified Eagle's medium (DMEM) low glucose (1 g L−1) (Capricorn; Catalogue number: DMEM-LPSTA), fetal bovin serum (FBS) (Capricorn; Catalogue number: FBS-11A), penicillin (Capricorn; Catalog number: PS-B), streptomycin (Capricorn; Catalogue number: PS-B), N-2-hydroxyethylpiperazine-N-2-ethane sulfonic acid (HEPES) buffer (Capricorn; Catalogue number: HEP-B), PrestoBlue cell viability reagent (Thermo Fisher Scientific; Catalogue number: A13262), branched Poly(ethylene imine) (bPEI 10000) (Polyscience Inc; Catalogue number: 19850), 24-well plates (imaging) (Ibidi; Catalogue number: 82426), Hoechst 33342 (Invitrogen; Catalogue number: H3570), CellMask™ Plasma Membrane Stain (Invitrogen; Catalogue number: C10046), DMEM without phenol red (Gibco; Catalogue number: A1443001).
SEC measurements for the investigation of the (co)polymerisation kinetics were performed on the following setup:
Agilent 1200 series, DG-2080-53 (degasser), G1310A (pump), G1329A (autosampler), Techlab (oven), RID-10A (RI detector), SPD-20A (UV detector), PSS GRAM guard/30/1000 Å column (10 μm particle size), N,N-dimethyl acetamide (DMAc) with 0.21 wt% LiCl as eluent at 1 mL min−1 at 40 °C, poly(methyl methacrylate) (PMMA) standard.
SEC measurements of the amino acid modified polyanions were performed on an instrument consisting of a column set with a Suprema pre-column (particle size = 5 μm) and three Suprema main columns (particle size = 5 μm, 1 Å × 30 Å; 2 Å × 1000 Å) with a separation range from 100 to 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 Da (PSS, Mainz, Germany) together with a variable wavelength detector (1200 Series, Agilent Technologies). As solvent, 0.07 M aqueous Na2HPO4 was used (for dissolving the polymers and as an eluting solvent) with a flow rate of 0.8 mL min−1 and the columns were maintained at room temperature. As internal standard, ethylene glycol (HPLC grade) was used. The calibration was performed with narrowly distributed poly(methacrylic acid) sodium salt homopolymers (PMA Na salt; PSS calibration kit). An injection volume of 60 μL was used for the measurements. The samples were dissolved with a concentration of 2 mg mL−1 and filtered through a 0.22 μm PTFE Nylon filter before analysis. The UV detector was set to λ = 490 nm for measurements of the 6AF-labelled polymers.
000 Da (PSS, Mainz, Germany) together with a variable wavelength detector (1200 Series, Agilent Technologies). As solvent, 0.07 M aqueous Na2HPO4 was used (for dissolving the polymers and as an eluting solvent) with a flow rate of 0.8 mL min−1 and the columns were maintained at room temperature. As internal standard, ethylene glycol (HPLC grade) was used. The calibration was performed with narrowly distributed poly(methacrylic acid) sodium salt homopolymers (PMA Na salt; PSS calibration kit). An injection volume of 60 μL was used for the measurements. The samples were dissolved with a concentration of 2 mg mL−1 and filtered through a 0.22 μm PTFE Nylon filter before analysis. The UV detector was set to λ = 490 nm for measurements of the 6AF-labelled polymers.
For SEC measurements with DMF, a GRAM 10 µm 3000 Å gel column (separation range of 5000 to 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 Da) was used. The sample solvent and eluent was DMF with lithium bromide (LiBr) (5 g L−1) and a flow rate of 0.5 mL min−1 was selected using polystyrene as standard (polystyrene homopolymer: PSS calibration kit).
000 Da) was used. The sample solvent and eluent was DMF with lithium bromide (LiBr) (5 g L−1) and a flow rate of 0.5 mL min−1 was selected using polystyrene as standard (polystyrene homopolymer: PSS calibration kit).
The temperature program of the GC was set depending on the substances. For the methyl acrylate copolymers, the temperature program of the GC was set as follows:
2 min at 40 °C, heating with 15 K min−1, 4 min at 100 °C, FID temperature: 270 °C.
For the NAM-containing polymers, the program was set as follows: 2 min at 80 °C, heating with 15 K min−1, 5 min at 240 °C, FID temperature: 270 °C.
Synthesis and characterisation
| Copolymer | [M]![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) [CTA] [CTA]![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) [V65] [V65] |
nCTA (mmol) | nV65 (mmol) | nNAM (mmol) | nPFPA (mmol) | VCH3CN (mL) | Voverall (mL) | Yield (%) |
|---|---|---|---|---|---|---|---|---|
| P(NAM10-ran-PFPA90)-CTA | 260![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.135 | 0.033 | 3.5 | 31.5 | 11.09 | 17.50 | 32 (2.62 g) |
| P(NAM20-ran-PFPA80)-CTA | 260![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.139 | 0.035 | 7.2 | 28.8 | 11.54 | 18.00 | 34 (2.65 g) |
| P(NAM30-ran-PFPA70)-CTA | 260![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.173 | 0.043 | 13.5 | 31.5 | 14.59 | 22.50 | 45 (4.24 g) |
| P(NAM40-ran-PFPA60)-CTA | 260![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.173 | 0.043 | 18.0 | 27.0 | 14.76 | 22.50 | 56 (5.00 g) |
| P(NAM50-ran-PFPA50)-CTA | 220![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.218 | 0.055 | 24.0 | 24.0 | 15.71 | 24.00 | 39 (3.54 g) |
| P(NAM60-ran-PFPA40)-CTA | 160![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.375 | 0.094 | 36.0 | 24.0 | 19.18 | 27.50 | 44 (4.74 g) |
| P(NAM70-ran-PFPA30)-CTA | 169![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.338 | 0.084 | 39.9 | 17.1 | 17.55 | 30.00 | 49 (4.79 g) |
| P(NAM80-ran-PFPA20)-CTA | 160![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.344 | 0.086 | 44.0 | 11.0 | 18.04 | 27.50 | 62 (5.54 g) |
| P(NAM90-ran-PFPA10)-CTA | 160![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.375 | 0.094 | 54.0 | 6.0 | 19.90 | 30.00 | 87 (7.91 g) |
| Copolymer | [M]![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) [CTA] [CTA]![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) [V65] [V65] |
nCTA (mmol) | nV65 (mmol) | nMA (mmol) | nPFPA (mmol) | VCH3CN (mL) | Voverall (mL) | Yield (%) |
|---|---|---|---|---|---|---|---|---|
| P(MA10-ran-PFPA90)-CTA | 260![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.177 | 0.044 | 4.6 | 41.4 | 14.73 | 23.00 | 87 (8.89 g) |
| P(MA20-ran-PFPA80)-CTA | 260![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.190 | 0.048 | 9.9 | 39.6 | 16.212 | 24.75 | 86 (8.81 g) |
| P(MA30-ran-PFPA70)-CTA | 240![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.188 | 0.047 | 13.5 | 31.5 | 14.975 | 22.50 | 39 (3.41 g) |
| P(MA40-ran-PFPA60)-CTA | 240![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.200 | 0.050 | 19.2 | 28.8 | 16.321 | 24.00 | 68 (5.78 g) |
| P(MA50-ran-PFPA50)-CTA | 230![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.270 | 0.067 | 31.0 | 31.0 | 21.461 | 31.00 | 69 (6.97 g) |
| P(MA60-ran-PFPA40)-CTA | 180![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.381 | 0.095 | 41.1 | 27.4 | 23.694 | 34.25 | 83 (8.34 g) |
| P(MA70-ran-PFPA30)-CTA | 180![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.419 | 0.105 | 52.9 | 22.7 | 26.662 | 37.75 | 57 (5.71 g) |
| P(MA80-ran-PFPA20)-CTA | 180![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.433 | 0.108 | 62.4 | 15.6 | 28.11 | 39.00 | 46 (4.18 g) |
| P(MA90-ran-PFPA10)-CTA | 160![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0.25 0.25 |
0.406 | 0.102 | 58.5 | 6.5 | 23.616 | 32.50 | 82 (5.39 g) |
Pentafluorophenyl acrylate (PFPA) was transferred into a previously inerted vial and stirred for at least 40 min with inhibitor remover. For the NAM copolymers, at this point, also the NAM was destabilized by filtration through a small column of neutral aluminium oxide. In the meantime, a 25 or 50 mL round bottom flask (depending on the total volume of the resulting polymerisation mixture) equipped with a rare earth stirring bar was heated with a heat gun and the atmosphere inside the flask was changed to an argon atmosphere by three cycles of filling the flask with argon and subsequently applying a vacuum onto the flask. Furthermore, a 10 mL glass vial was inerted in the same way. After the flask cooled back down to room temperature, a rubber septum was placed on the flask and anhydrous acetonitrile (16.76 g) was added to the flask gravimetrically. Subsequently, the CTA (98.3 mg, 0.27 mmol, 0.0087 equiv.) was added to the flask and the solution was stirred until the whole CTA was dissolved. In the meantime, V65 was added to a previously inerted vial and dissolved in acetonitrile to yield a solution of β = 10 mg mL−1. At this point, for the MA copolymers, the MA was destabilized by filtration through a short column of neutral aluminium oxide. After the complete dissolution of the CTA, both monomers were added to the flask gravimetrically (PFPA: 7.381 g, 31.0 mmol, 1 equiv.; MA: 2.669 g, 31.0 mmol, 1 equiv.) to yield a certain ratio of [PFPA + comonomer]![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) [CTA] (230
[CTA] (230![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) and a total monomer concentration of 2 M. Furthermore, the ratio of one monomer to the other was chosen as the ratio in the final copolymer was envisaged (10 to 90 mol% monomer 1 (NAM or MA) to 90 to 10 mol% monomer 2 (PFPA)); in the currently described case: 50 mol% to 50 mol%.
1) and a total monomer concentration of 2 M. Furthermore, the ratio of one monomer to the other was chosen as the ratio in the final copolymer was envisaged (10 to 90 mol% monomer 1 (NAM or MA) to 90 to 10 mol% monomer 2 (PFPA)); in the currently described case: 50 mol% to 50 mol%.
Subsequently, the initiator solution (1.674 mL, 0.067 mmol V65, 0.0022 equiv.) was added to the flask utilizing an Eppendorf-pipette (1 mL nominal volume). The overall ratio of CTA to initiator was 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. Afterwards, the solution was deoxygenized by sparging with argon for ca. 20 min. The yellow mixture was then transferred to a preheated oil bath thermostated to 60 °C and stirred for 17 h at 500 revolutions per minute (rpm). The mixture was subsequently transferred to a dialysis tubing (Spectrum Spectra/Por regenerated cellulose ester (RC), molecular weight cutoff (MWCO): 8 kDa) and dialysis against THF was performed for three days with daily solvent change and without stirring. Subsequently, the solvent was removed under reduced pressure, and the polymer was dried in a vacuum oven at 60 °C until mass constancy. The products were obtained in a range from a yellow solid (samples with high amount of NAM) to yellow viscous resins (high amount of MA). The yields and the exact masses of all polymerisations are presented in Tables 4 and 5.
1. Afterwards, the solution was deoxygenized by sparging with argon for ca. 20 min. The yellow mixture was then transferred to a preheated oil bath thermostated to 60 °C and stirred for 17 h at 500 revolutions per minute (rpm). The mixture was subsequently transferred to a dialysis tubing (Spectrum Spectra/Por regenerated cellulose ester (RC), molecular weight cutoff (MWCO): 8 kDa) and dialysis against THF was performed for three days with daily solvent change and without stirring. Subsequently, the solvent was removed under reduced pressure, and the polymer was dried in a vacuum oven at 60 °C until mass constancy. The products were obtained in a range from a yellow solid (samples with high amount of NAM) to yellow viscous resins (high amount of MA). The yields and the exact masses of all polymerisations are presented in Tables 4 and 5.
The SEC samples of the different polymers were prepared and then stored intermittently in the dark at 5 °C. The subsequent measurements were performed within a timeframe of ca. 20 min after preparation to avoid potential cleavage of the active ester moiety in the chloroform SEC eluent containing triethylamine and iso-propanol during waiting times.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) [CTA]0) according to eqn (1)–(3).
[CTA]0) according to eqn (1)–(3).
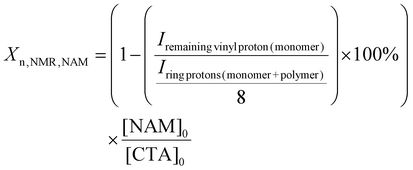 | (1) |
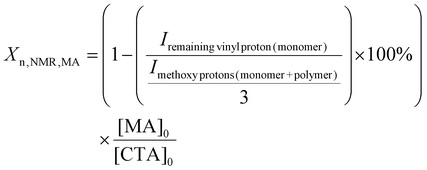 | (2) |
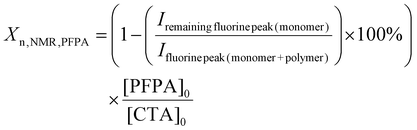 | (3) |
The total Xn,NMR for the respective copolymer was afterwards calculated by adding the Xns of both comonomers. The molar masses of the copolymers were calculated by multiplying the Xn of the individual comonomers with the molar mass of the corresponding monomer and then adding them up to receive one molar mass (Mn,NMR).
| Copolymer | mpolymer (g) | mAIBN (mg) | mluperox (mg) | VDMF (mL) | Vtoluene (mL) |
|---|---|---|---|---|---|
| P(NAM10-ran-PFPA90) | 1.0 | 119.8 | 29.1 | 13.6 | — |
| P(NAM20-ran-PFPA80) | 1.0 | 125.1 | 30.4 | 14.2 | — |
| P(NAM30-ran-PFPA70) | 1.0 | 130.9 | 31.8 | 14.9 | — |
| P(NAM40-ran-PFPA60) | 1.0 | 137.3 | 33.3 | 15.6 | — |
| P(NAM50-ran-PFPA50) | 1.0 | 144.3 | 35.0 | 16.4 | — |
| P(NAM60-ran-PFPA40) | 1.0 | 152.1 | 36.9 | 17.3 | — |
| P(NAM70-ran-PFPA30) | 1.0 | 160.8 | 39.0 | 18.3 | — |
| P(NAM80-ran-PFPA20) | 1.0 | 170.5 | 41.4 | 19.4 | — |
| P(NAM90-ran-PFPA10) | 1.0 | 181.4 | 44.0 | 20.6 | — |
| P(MA10-ran-PFPA90) | 1.0 | 98.2 | 23.8 | — | 11.2 |
| P(MA20-ran-PFPA80) | 1.0 | 105.4 | 25.6 | — | 12 |
| P(MA30-ran-PFPA70) | 1.0 | 113.7 | 27.6 | — | 12.9 |
| P(MA40-ran-PFPA60) | 1.0 | 123.5 | 30.0 | — | 14 |
| P(MA50-ran-PFPA50) | 1.0 | 135.1 | 32.8 | — | 15.3 |
| P(MA60-ran-PFPA40) | 1.0 | 149.0 | 36.2 | — | 16.9 |
| P(MA70-ran-PFPA30) | 1.0 | 166.3 | 40.4 | — | 18.9 |
| P(MA80-ran-PFPA20) | 1.0 | 187.9 | 45.6 | — | 21.4 |
| P(MA90-ran-PFPA10) | 1.0 | 216.2 | 52.5 | — | 24.6 |
The reaction vial was closed with a rubber septum and deoxygenized by argon sparging for 30 min while cooling the reaction mixture in ice. After stirring at 80 °C for 2.5 h, the mixture was cooled to room temperature and precipitated in ice-cold diethyl ether (10-fold excess relative to DMF; centrifugation at 5000 rpm for 1 min). For copolymers which did not precipitate: DMF was removed by reduced pressure. The remaining product was dissolved in 10 mL chloroform and precipitated with 90 mL ice-cold n-hexane (centrifugation at 5000 rpm for 1 min). The copolymers were dried overnight in a vacuum oven to obtain the crude solids. The end-group removal was confirmed via SEC measurements in DMF (sample eluent: DMF with lithium bromide (5 g L−1); PS standard), revealing the decrease of the UV trace at λ = 310 nm (see Table 7 for details).
| Copolymer | Mn,app (g mol−1) | Mw,app (g mol−1) | Đ |
|---|---|---|---|
| P(NAM10-ran-PFPA90) | 8300 | 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.26 |
| P(NAM20-ran-PFPA80) | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.32 |
| P(NAM30-ran-PFPA70) | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
1.37 |
| P(NAM40-ran-PFPA60) | 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.35 |
| P(NAM50-ran-PFPA50) | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600 600 |
1.36 |
| P(NAM60-ran-PFPA40) | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 700 700 |
16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.3 |
| P(NAM70-ran-PFPA30) | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
1.35 |
| P(NAM80-ran-PFPA20) | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 700 700 |
16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
1.28 |
| P(NAM90-ran-PFPA10) | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 700 700 |
1.27 |
| P(MA10-ran-PFPA90) | 9900 | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.30 |
| P(MA20-ran-PFPA80) | 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
1.29 |
| P(MA30-ran-PFPA70) | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
1.27 |
| P(MA40-ran-PFPA60) | 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.27 |
| P(MA50-ran-PFPA50) | 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.20 |
| P(MA60-ran-PFPA40) | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.23 |
| P(MA70-ran-PFPA30) | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.20 |
| P(MA80-ran-PFPA20) | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
1.24 |
| P(MA90-ran-PFPA10) | 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 700 700 |
14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.18 |
Post-polymerisation modification of PPFPA with aliphatic amino acids for cell viability studies
To obtain Nva-modified copolymers, the activated ester in P(NAMn-ran-PFPAm) and P(MAn-ran-PFPAm) was reacted in an aminolysis with the amino acid to form the corresponding acryl amide P(NAMn-ran-Nva-OH-AAmm) and P(MAn-ran-Nva-OH-Aamm), respectively.9 The post-polymerisation modification is described exemplarily for P(NAM10-ran-PFPA90) and the chemicals required for further copolymer modifications are listed in Table 8: for the reaction, 200 mg copolymer (1.0 equiv., based on PFPA units), 500 µl TEA and 277 mg Nva (3.0 equiv. related to the reactive polymer units) were dissolved in 10 mL anhydrous DMF. After sealing the reaction vial, the suspension was incubated at 40 °C for 72 h and then cooled down to room temperature. To remove unreacted amino acids, the product was filtered and dialyzed against deionized water (Spectrum Spectra/Por, RC, MWCO: 3.5 kDa) for one week with daily water exchange. Polymers dissolved in distilled water (diH2O) were run over a column of basic aluminium oxide to remove residual triethylammonium salt impurities. After lyophilization, the product was obtained as a white or off-white solid. Polymers were analysed by SEC, 1H NMR and FTIR measurements to obtain information about the molar mass distribution as well as degree of modification and potential hydrolysis. After a freeze-drying step, the copolymer properties were analysed. Those polymers were investigated by SEC, NMR, HPLC and used for cell viability studies and hemolysis experiments.| Copolymer | P(NAMn-ran-PFPAm−100) | 6-AF | Nva | DMF/TEA | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| P(MAn-ran-PFPAm−100) | ||||||||||
| m (mg) | n (mmol) | equiv.a | m (mg) | n (mmol) | equiv.a | m (mg) | n (mmol) | equiv.a | V (mL) | |
| a The equivalent is based on the PFPA units. | ||||||||||
| P(NAM10-ran-Nva-OH-AAm90) | 100 | 3.6 × 10−3 | 1 | 1.39 | 4.0 × 10−3 | 1.1 | 138.5 | 1.18 | 3 | 5/0.25 |
| P(NAM20-ran-Nva-OH-AAm80) | 100 | 3.8 × 10−3 | 1 | 1.46 | 4.2 × 10−3 | 1.1 | 128.6 | 1.10 | 3 | 5/0.25 |
| P(NAM30-ran-Nva-OH-AAm70) | 100 | 4.0 × 10−3 | 1 | 1.52 | 4.4 × 10−3 | 1.1 | 117.7 | 1.01 | 3 | 5/0.25 |
| P(NAM40-ran-Nva-OH-AAm60) | 100 | 4.2 × 10−3 | 1 | 1.60 | 4.6 × 10−3 | 1.1 | 105.8 | 0.90 | 3 | 5/0.25 |
| P(NAM50-ran-Nva-OH-AAm50) | 100 | 4.4 × 10−3 | 1 | 1.68 | 4.8 × 10−3 | 1.1 | 92.7 | 0.79 | 3 | 5/0.25 |
| P(NAM60-ran-Nva-OH-AAm40) | 100 | 4.6 × 10−3 | 1 | 1.77 | 5.1 × 10−3 | 1.1 | 78.1 | 0.67 | 3 | 5/0.25 |
| P(NAM70-ran-Nva-OH-AAm30) | 100 | 4.9 × 10−3 | 1 | 1.87 | 5.4 × 10−3 | 1.1 | 61.9 | 0.53 | 3 | 5/0.25 |
| P(NAM80-ran-Nva-OH-AAm20) | 100 | 5.2 × 10−3 | 1 | 1.98 | 5.7 × 10−3 | 1.1 | 43.8 | 0.37 | 3 | 5/0.25 |
| P(NAM90-ran-Nva-OH-AAm10) | 100 | 5.5 × 10−3 | 1 | 2.11 | 6.1 × 10−3 | 1.1 | 23.3 | 0.2 | 3 | 5/0.25 |
| P(MA10-ran-Nva-OH-AAm90) | 100 | 3.0 × 10−3 | 1 | 1.14 | 3.3 × 10−3 | 1.1 | 141.9 | 1.21 | 3 | 5/0.25 |
| P(MA20-ran-Nva-OH-AAm80) | 100 | 3.2 × 10−3 | 1 | 1.23 | 3.5 × 10−3 | 1.1 | 135.4 | 1.16 | 3 | 5/0.25 |
| P(MA30-ran-Nva-OH-AAm70) | 100 | 3.5 × 10−3 | 1 | 1.32 | 3.8 × 10−3 | 1.1 | 127.8 | 1.09 | 3 | 5/0.25 |
| P(MA40-ran-Nva-OH-AAm60) | 100 | 3.8 × 10−3 | 1 | 1.44 | 4.1 × 10−3 | 1.1 | 118.9 | 1.02 | 3 | 5/0.25 |
| P(MA50-ran-Nva-OH-AAm50) | 100 | 4.1 × 10−3 | 1 | 1.57 | 4.5 × 10−3 | 1.1 | 108.4 | 0.93 | 3 | 5/0.25 |
| P(MA60-ran-Nva-OH-AAm40) | 100 | 4.5 × 10−3 | 1 | 1.73 | 5.0 × 10−3 | 1.1 | 95.7 | 0.82 | 3 | 5/0.25 |
| P(MA70-ran-Nva-OH-AAm30) | 100 | 5.1 × 10−3 | 1 | 1.93 | 5.6 × 10−3 | 1.1 | 80.1 | 0.68 | 3 | 5/0.25 |
| P(MA80-ran-Nva-OH-AAm20) | 100 | 5.7 × 10−3 | 1 | 2.19 | 6.3 × 10−3 | 1.1 | 60.3 | 0.52 | 3 | 5/0.25 |
| P(MA90-ran-Nva-OH-AAm10) | 100 | 6.6 × 10−3 | 1 | 2.51 | 7.2 × 10−3 | 1.1 | 34.7 | 0.3 | 3 | 5/0.25 |
After dissolving 100 mg (1.0 equiv.) copolymer and 1 mg 6-AF (1.1 equiv.) in 5 mL anhydrous DMF and 25 µl TEA, the reaction vial was sealed and stirred at 40 °C for 24 h in the dark. Then, 139 mg Nva (3.0 equiv. related to the reactive polymer units) and 225 µl TEA were added to the solution and the reaction was allowed to stir further for 72 h. The mixture was cooled down to room temperature, the unreacted Nva was separated by filtration and a dialysis step in deionized water (Spectrum Spectra/Por, RC, MWCO 3.5 kDa; daily water exchange) was performed for seven days. After freeze-drying, the resulting copolymers were characterised (Table 9). The formed polymers were investigated by SEC, NMR (see Fig. S14) and used for cell interaction studies and the hydrophobicity assay. Moreover, the post-polymerization (PPM) efficiency of the NAM- and MA-based polymers with Nva was determined in Table 9 by NMR measurements.
| Copolymer | Mn, app (g mol−1) | Mw, app (g mol−1) | Đ | Nva (%) | PPM efficiency (%) |
|---|---|---|---|---|---|
| P(NAM10-ran-Nva-OH-AAm90)-6-AF | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
1.38 | 47 | 52 |
| P(NAM20-ran-Nva-OH-AAm80)-6AF | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
1.82 | 44 | 55 |
| P(NAM30-ran-Nva-OH-AAm70)-6-AF | 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
33![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600 600 |
1.35 | 39 | 56 |
| P(NAM40-ran-Nva-OH-AAm60)-6-AF | 24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 600 600 |
31![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
1.26 | 34 | 56 |
| P(NAM50-ran-Nva-OH-AAm50)-6-AF | 31![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
43![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
1.36 | 40 | 80 |
| P(NAM60-ran-Nva-OH-AAm40)-6-AF | 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
29![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 800 800 |
1.46 | 27 | 67 |
| P(NAM70-ran-Nva-OH-AAm30)-6-AF | 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
1.47 | 22 | 72 |
| P(NAM80-ran-Nva-OH-AAm20)-6-AF | 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
19![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 100 |
1.45 | 17 | 83 |
| P(NAM90-ran-Nva-OH-AAm10)-6-AF | 6800 | 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
1.65 | 9 | 93 |
| P(MA10-ran-Nva-OH-AAm90)-6-AF | 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.42 | 46 | 51 |
| P(MA20-ran-Nva-OH-AAm80)-6-AF | 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
1.44 | 48 | 60 |
| P(MA30-ran-Nva-OH-AAm70)-6-AF | 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
1.31 | 46 | 65 |
| P(MA40-ran-Nva-OH-AAm60)-6-AF | 23![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300 300 |
1.13 | 35 | 59 |
| P(MA50-ran-Nva-OH-AAm50)-6-AF | 26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 200 |
36![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
1.39 | 38 | 75 |
| P(MA60-ran-Nva-OH-AAm40)-6-AF | 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
1.52 | 32 | 79 |
| P(MA70-ran-Nva-OH-AAm30)-6-AF | 7700 | 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 900 |
2.2 | 29 | 96 |
| P(MA80-ran-Nva-OH-AAm20)-6-AF | 5700 | 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
2.51 | 15 | 75 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio with chloroform and vortexed for 1 min at maximum speed. After the visible separation of two phases (a few minutes up to 24 h) was obtained, 100 µl of each sample of the upper aqueous phase was taken and quantified spectrophotometrically. The fluorescence intensity reduction of each copolymer in the aqueous layer was calculated as follows (eqn (4)):
1 ratio with chloroform and vortexed for 1 min at maximum speed. After the visible separation of two phases (a few minutes up to 24 h) was obtained, 100 µl of each sample of the upper aqueous phase was taken and quantified spectrophotometrically. The fluorescence intensity reduction of each copolymer in the aqueous layer was calculated as follows (eqn (4)):
 | (4) |
All copolymer tests were conducted in triplicates.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) P values. The partition coefficients (c
P values. The partition coefficients (c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values) were computed using the cheminformatics tool RDKit58 based on the Wildman and Crippen scheme.59,66
P values) were computed using the cheminformatics tool RDKit58 based on the Wildman and Crippen scheme.59,66In detail, the c![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P value calculations are conducted for ideal polymers– polymers with 100% post-polymerization modification efficiency and polymer lengths of Xn = 200 – whereas experimentally obtained data is not considered since the data correlation is linear and the experimental results will fit to the plotted dashed lines.
P value calculations are conducted for ideal polymers– polymers with 100% post-polymerization modification efficiency and polymer lengths of Xn = 200 – whereas experimentally obtained data is not considered since the data correlation is linear and the experimental results will fit to the plotted dashed lines.
Biological evaluation
For biological observations, the polymers were dissolved in PBS at a concentration of 2 mg mL−1. The polymer suspensions were stored at 4 °C, equilibrated to room temperature, and resuspended before use. No large aggregates were observed.Determination of cytotoxicity in L929 cells via PrestoBlue assay
The biological assay was adapted from Solomun et al.67 To determine cytotoxicity adapted to ISO10993-5 (L929, MTT) L929 cells (CLS Eppelheim, Catalogue number: 400260) were seeded in a 96-well plate with a cell density of 0.1 × 106 cells mL−1 in 100 µL and incubated for 24 h at 37 °C in humidified 5% (v/v) CO2 atmosphere. The cells were cultivated in DMEM low glucose (1 g L−1). The media was supplemented with 10% (v/v) FBS, 100 U mL−1 penicillin and 100 µg mL−1 streptomycin (L929 culture medium). For the experiments 10 mM HEPES was added (L929 test medium). 1 h prior treatment the medium was changed to 90 µL fresh L929 test medium. Cells were treated with polymers at various concentrations ranging from 0 to 200 µg mL−1. 24 h after treatment, the medium was replaced by a 10% (v/v) solution of PrestoBlue cell viability reagent diluted with L929 culture medium. The cells were incubated for 45 min at 37 °C.Fluorescence intensity (λex = 560 nm, λem = 590 nm) was measured with an Tecan Infinite M200 Pro plate reader. Cells treated with buffer were used as a positive control (F0, no cytotoxicity, 100% metabolic activity), and the viability was calculated relative to the buffer control after subtracting the blank (Fblank, PrestoBlue diluted in medium 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 without cells), eqn (5) as follows:
10 without cells), eqn (5) as follows:
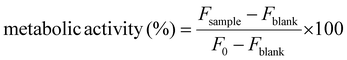 | (5) |
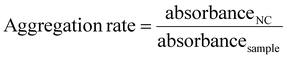 | (6) |
Exemplarily microscopy images of aggregation are provided in Fig. 6.
In parallel, the remaining (700 µL) test solution was incubated for 1 h at 37 °C. To determine haemoglobin release, the tubes were centrifuged at 2400g for 5 min and the supernatant was transferred to a 96-well plate in three technical replicates. To measure the haemolytic effect of the polymers the absorbance was measured at λ = 544 nm with λ = 630 nm as reference bandwidth. As a positive control 1% Triton X-100 (T-X) was used (100% haemolysis), and pure DPBS was used as negative control. The haemolysis (%) was calculated using eqn (7).
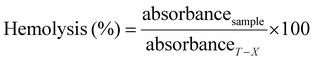 | (7) |
Values below 2% haemolysis are classified as non-haemolytic, values between 2 to 5% are slightly haemolytic and values above 5% are haemolytic.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 single cells were measured and analysed by forward and sideward scatter (FSC/SSC). Fluorescence was measured at λex = 488 nm with a 525/40 nm bandpass filter (FITC channel). Positive cells were identified by gating against NC. A detailed gating strategy is provided in Fig. S20. Analysis was conducted with Kaluza V 2.2.1.
000 single cells were measured and analysed by forward and sideward scatter (FSC/SSC). Fluorescence was measured at λex = 488 nm with a 525/40 nm bandpass filter (FITC channel). Positive cells were identified by gating against NC. A detailed gating strategy is provided in Fig. S20. Analysis was conducted with Kaluza V 2.2.1.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1000 in DMEM without phenol red. After 10 min of incubation, the staining solution was removed and the cells were washed twice with prewarmed DPBS. Finally, DMEM without phenol red was added for imaging. Imaging was performed using a LSM980 (Zeiss) applying laser for excitation at 405 nm (0.5%) (emission filter 412 to 509 nm, detection of nuclei), a laser at 488 nm (1.5%) (emission filter 492 to 678 nm, detection of 6AF) and a laser at 639 nm (0.2%) (emission filter 643 to 696 nm, detection of cell membrane). The experiments were performed at least twice. All images were processed with ZEN software, version 3.7 (ZEN lite) (Zeiss). The same values were applied to all images.
1000 in DMEM without phenol red. After 10 min of incubation, the staining solution was removed and the cells were washed twice with prewarmed DPBS. Finally, DMEM without phenol red was added for imaging. Imaging was performed using a LSM980 (Zeiss) applying laser for excitation at 405 nm (0.5%) (emission filter 412 to 509 nm, detection of nuclei), a laser at 488 nm (1.5%) (emission filter 492 to 678 nm, detection of 6AF) and a laser at 639 nm (0.2%) (emission filter 643 to 696 nm, detection of cell membrane). The experiments were performed at least twice. All images were processed with ZEN software, version 3.7 (ZEN lite) (Zeiss). The same values were applied to all images.Statistical analysis
Statistical analyses were performed with GraphPad Prism (version 10.3.1). Biological data are presented as mean ± standard deviation (s.d.) with individual data points as described in the figure legends. Two-way analysis of variance (ANOVA) was performed to compare multiple groups. An unpaired t-test with Welch's correction was performed on normally distributed, ungrouped data. Statistical significance is indicated as follows: ns p > 0.05, *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001.Author contributions
Conceptualisation: MR, MS, TML, MNL, SZ and AT; data curation: MR, TML, LR, EMB, MS, JDB and CK; investigation: MR, TML, LR, EMB, JDB and MS; creation of the copolymer libraries: MR; post-polymerisation modification: TML, LR and EMB; performing of biotests: MS; performing of kinetic experiments: MR; formal analysis: MR, LR, TML, MS and JDB; funding acquisition: MNL, AT and USS; methodology: MNL, MR, TML, LR and MS; project administration: MR, TML, MS, LR, MNL and AT; software for log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P calculation: CK; writing – original draft: MR, TML and MS; writing – review & editing: MR, MS, TML, LR, CK, AT, SZ, MNL and USS; visualisation: MR, JDB, TML, CK and MS; supervision: MNL, AT, TML, SZ and USS; resources: CK, AT, MNL and USS. All authors have read and agreed to the published version of the manuscript.
P calculation: CK; writing – original draft: MR, TML and MS; writing – review & editing: MR, MS, TML, LR, CK, AT, SZ, MNL and USS; visualisation: MR, JDB, TML, CK and MS; supervision: MNL, AT, TML, SZ and USS; resources: CK, AT, MNL and USS. All authors have read and agreed to the published version of the manuscript.
Conflicts of interest
There are no conflicts to declare.Data availability
The original and primary data supporting the findings of this work are available in a separate Zenodo archive under: https://doi.org/10.5281/zenodo.17663120.The calculation for the computed log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P values is stored in a separate GitHub repository under: https://github.com/kuennethgroup/logp_copo_nam_nva_pma.
P values is stored in a separate GitHub repository under: https://github.com/kuennethgroup/logp_copo_nam_nva_pma.
Supplementary information (SI): experimental details of the PNAM homopolymer; data on the kinetic investigations of homo- and copolymerisation kinetics; information on utilized 1H NMR peaks for the determination of the conversion during copolymer library synthesis; 1H NMR spectra of the copolymers, SEC curves, 1H NMR spectra and FTIR spectra of copolymers and further details on the biological investigations. See DOI: https://doi.org/10.1039/d5lp00376h.
Acknowledgements
The authors would like to acknowledge Prof Dr Ruth Freitag, Prof Dr Andreas Greiner and Prof Dr Thomas Scheibel for providing access to their laboratories and equipment. We would like to thank Maria Müller for conducting supporting chemical reactions. The work was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research foundation), Collaborative Research Center SFB/TRR225 ‘From the fundamentals of Biofabrication to functional tissue models’ – project number 326998133; research training group, GRK 3040, COIN, project A2 and B1 – project number: 527537972 as well as under the regime of the priority program SPP 2363 “Utilization and Development of Machine Learning for Molecular Applications – Molecular Machine Learning” (SCHU 1229/63-1; project number 497115849), and project number 316213987, CRC 1278 (projects B01 and C06). Further funding was provided by the funding scheme Exzellenzverbünde und Universitätskooperationen (EVUK) within the project “Function by Design: Cellular Hybrids” by the Bavarian Ministry of the Science and Art”.M. N. L. acknowledges financial support from the “Fonds der Chemischen Industrie im Verband der Chemischen Industrie”. A. T. acknowledges funding by the DFG Heisenberg Programme (514006196). We thank the Microverse Imaging Center (funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany's Excellence Strategy – EXC 2051 – Project-ID 390713860), Patrick Then and Aurélie Jost for providing microscope facility support for data acquisition. The LSM 980 was funded by the Free State of Thuringia with grant number 2019 FGI 0001. This work was performed within the Joint Lab for Polymers Jena-Bayreuth. The project underlying these results was funded by the Free State of Thuringia under number 2016 IZN 0009 and co-financed by the European Union under the European Regional Development Fund (ERDF). The TOC as well as additional parts of some graphics in this manuscript were prepared with BioRender.com. Readability and language of parts of the original draft of this manuscript were improved utilizing the tools DeepL and ChatGPT.
References
- S. Sharma, T. M. Perring, S.-J. Jeon, H. Huang, W. Xu, E. Islamovic, B. Sharma, Y. M. Giraldo and J. P. Giraldo, Nat. Commun., 2024, 15, 9737 CrossRef CAS PubMed.
- S. Shakiba, C. E. Astete, S. Paudel, C. M. Sabliov, D. F. Rodrigues and S. M. Louie, Environ. Sci. Nano, 2020, 7, 37–67 RSC.
- M. N. Leiske and K. Kempe, Macromol. Rapid Commun., 2022, 43, e2100615 CrossRef PubMed.
- A. Karabasz, M. Bzowska and K. Szczepanowicz, Int. J. Nanomed., 2020, 15, 8673–8696 CrossRef CAS PubMed.
- D. S. Idris, A. Roy, S. Pandit, S. Alghamdi, M. Almehmadi, A. A. Alsaiari, O. Abdulaziz, A. Alsharif, M. U. Khandaker and M. R. I. Faruque, e-Polymer, 2023, 23, 20230049 CrossRef CAS.
- A. Sultana, M. Zare, V. Thomas, T. S. Kumar and S. Ramakrishna, Med. Drug Discov., 2022, 15, 100134–1001354 Search PubMed.
- V. Kumar, B. Gupta, G. Kumar, M. K. Pandey, E. Aiazian, V. S. Parmar, J. Kumar and A. C. Watterson, J. Macromol. Sci., Part A:Pure Appl. Chem., 2010, 47, 1154–1160 Search PubMed.
- V. Manimaran, R. P. Nivetha, T. Tamilanban, J. Narayanan, S. Vetriselvan, N. K. Fuloria, S. V. Chinni, M. Sekar, S. Fuloria, L. S. Wong, A. Biswas, G. Ramachawolran and S. Selvaraj, Front. Mol. Biosci., 2023, 10, 1232109–1232126 Search PubMed.
- J. de Breuck, M. Streiber, M. Ringleb, D. Schröder, N. Herzog, U. S. Schubert, S. Zechel, A. Traeger and M. N. Leiske, ACS Polym. Au, 2024, 4, 222–234 CrossRef CAS PubMed.
- J. S. Chawla and M. M. Amiji, Int. J. Pharm., 2002, 249, 127–138 CrossRef CAS PubMed.
- F. U. Din, W. Aman, I. Ullah, O. S. Qureshi, O. Mustapha, S. Shafique and A. Zeb, Int. J. Nanomed., 2017, 12, 7291–7309 CrossRef PubMed.
- R. De, M. K. Mahata and K.-T. Kim, Adv. Sci., 2022, 9, e2105373 CrossRef PubMed.
- F. Meng, Z. Zhong and J. Feijen, Biomacromolecules, 2009, 10, 197–209 CrossRef CAS PubMed.
- R. P. Brinkhuis, F. P. J. T. Rutjes and J. C. M. van Hest, Polym. Chem., 2011, 2, 1449–1462 RSC.
- J. K. Elter, J. Eichhorn, M. Ringleb and F. H. Schacher, Polym. Chem., 2021, 12, 3900–3916 RSC.
- B. Charleux, G. Delaittre, J. Rieger and F. D'Agosto, Macromolecules, 2012, 45, 6753–6765 CrossRef CAS.
- F. Richter, K. Leer, L. Martin, P. Mapfumo, J. I. Solomun, M. T. Kuchenbrod, S. Hoeppener, J. C. Brendel and A. Traeger, J. Nanobiotechnol., 2021, 19, 1–15 Search PubMed.
- J. I. Solomun, L. Martin, P. Mapfumo, E. Moek, E. Amro, F. Becker, S. Tuempel, S. Hoeppener, K. L. Rudolph and A. Traeger, J. Nanobiotechnol., 2022, 20, 1–17 CrossRef PubMed.
- M. Streiber, N. E. Göppert, V. Bachmann, B. Schulze, R. Gläßer, I. Anufriev, P. Schädel, I. Nischang, C. Weber, O. Werz, U. S. Schubert and A. Traeger, Mater. Today Chem., 2025, 44, 102549–102560 CrossRef CAS.
- K. Kuperkar, D. Patel, L. I. Atanase and P. Bahadur, Polymers, 2022, 14, 4702 CrossRef CAS PubMed.
- C. Biglione, T. M. P. Neumann-Tran, S. Kanwal and D. Klinger, J. Polym. Sci., 2021, 59, 2665–2703 CrossRef CAS.
- W. Xue, H. Mutlu and P. Theato, Eur. Polym. J., 2020, 130, 109660–109669 CrossRef CAS.
- C. Battistella, Y. Yang, J. Chen and H.-A. Klok, ACS Omega, 2018, 3, 9710–9721 CrossRef CAS PubMed.
- Y. Pinyakit, T. Palaga, S. Kiatkamjornwong and V. P. Hoven, J. Mater. Chem. B, 2020, 8, 454–464 RSC.
- M. A. Gauthier, M. I. Gibson and H.-A. Klok, Angew. Chem., Int. Ed., 2009, 48, 48–58 CrossRef CAS PubMed.
- X. Chen and T. Michinobu, Macromol. Chem. Phys., 2022, 223, 2100370 CrossRef CAS.
- L. Yuan, L. He, Y. Wang, X. Lang, F. Yang, Y. Zhao and H. Zhao, Macromolecules, 2020, 53, 3175–3181 CrossRef CAS.
- H. Chen, L. Wang, S. Feng and L. Li, Eur. Polym. J., 2024, 220, 113463–113470 CrossRef CAS.
- A. Y. Dzhuzha, I. I. Tarasenko, L. I. Atanase, A. Lavrentieva and E. G. Korzhikova-Vlakh, Int. J. Mol. Sci., 2023, 24, 1049–1068 Search PubMed.
- E. M. Muzammil, A. Khan and M. C. Stuparu, RSC Adv., 2017, 7, 55874–55884 RSC.
- S. Agar, E. Baysak, G. Hizal, U. Tunca and H. Durmaz, J. Polym. Sci., Part A: Polym. Chem., 2018, 56, 1181–1198 Search PubMed.
- Y. Wang, S. Zhang, L. Wang, W. Zhang, N. Zhou, Z. Zhang and X. Zhu, Polym. Chem., 2015, 6, 4669–4677 RSC.
- M. Rimmele, F. Glöcklhofer and M. Heeney, Mater. Horiz., 2022, 9, 2678–2697 RSC.
- R. J. Kopiasz, D. Kozon, J. Pachla, Ł. Skórka and D. Jańczewski, Polymer, 2021, 217, 123452 CrossRef CAS.
- R. Kakuchi and P. Theato, ACS Macro Lett., 2013, 2, 419–422 CrossRef CAS PubMed.
- W. Graisuwan, H. Zhao, S. Kiatkamjornwong, P. Theato and V. P. Hoven, J. Polym. Sci., Part A: Polym. Chem., 2015, 53, 1103–1113 CrossRef CAS.
- M. Liu, D. Miao, X. Wang, C. Wang and W. Deng, J. Polym. Sci., 2020, 58, 2074–2087 CrossRef CAS.
- H. Son, Y. Jang, J. Koo, J.-S. Lee, P. Theato and K. Char, Polym. J., 2016, 48, 487–495 CrossRef CAS.
- A. P. Grimm, S. T. Knox, C. Y. P. Wilding, H. A. Jones, B. Schmidt, O. Piskljonow, D. Voll, C. W. Schmitt, N. J. Warren and P. Théato, Macromol. Rapid Commun., 2025, 46, e2500264 CrossRef PubMed.
- M. I. Gibson, E. Fröhlich and H.-A. Klok, J. Polym. Sci., Part A: Polym. Chem., 2009, 47, 4332–4345 CrossRef CAS.
- K. A. Günay, N. Schüwer and H.-A. Klok, Polym. Chem., 2012, 3, 2186–2192 RSC.
- H. Son, J. Ku, Y. Kim, S. Li and K. Char, Biomacromolecules, 2018, 19, 951–961 CrossRef CAS PubMed.
- M. Ringleb, Y. Köster, S. Zechel and U. S. Schubert, Digital Discovery, 2025, 4(11), 3292–3303 RSC.
- J.-M. Noy, M. Koldevitz and P. J. Roth, Polym. Chem., 2015, 6, 436–447 RSC.
- J. Alex, J. Ulbrich, M. Rosales-Guzmán, C. Weber, U. S. Schubert and C. Guerrero-Sanchez, Eur. Polym. J., 2021, 143, 110175 CrossRef CAS.
- F. D. Jochum and P. Theato, Macromolecules, 2009, 42, 5941–5945 CrossRef CAS.
- Z. Hou, W. M. Nau and R. Hoogenboom, Polym. Chem., 2021, 12, 307–315 RSC.
- M. Eberhardt and P. Théato, Macromol. Rapid Commun., 2005, 26, 1488–1493 CrossRef CAS.
- R. Bou Zerdan, Z. Geng, B. Narupai, Y. J. Diaz, M. W. Bates, D. S. Laitar, B. Souvagya, A. K. van Dyk and C. J. Hawker, J. Polym. Sci., 2020, 58, 1989–1997 CrossRef CAS.
- D. Babikova, R. Kalinova, I. Zhelezova, D. Momekova, S. Konstantinov, G. Momekov and I. Dimitrov, RSC Adv., 2016, 6, 84634–84644 RSC.
- A. Traeger and M. N. Leiske, Biomacromolecules, 2025, 26, 5–32 CrossRef CAS PubMed.
- O. Sedlacek, K. Lava, B. Verbraeken, S. Kasmi, B. G. de Geest and R. Hoogenboom, J. Am. Chem. Soc., 2019, 141, 9617–9622 CrossRef CAS PubMed.
- M. R. Elzes, I. Mertens, O. Sedlacek, B. Verbraeken, A. C. A. Doensen, M. A. Mees, M. Glassner, S. Jana, J. M. J. Paulusse and R. Hoogenboom, Biomacromolecules, 2022, 23, 2459–2470 CrossRef CAS PubMed.
- K. P. Schumann, T. M. Lutz and M. N. Leiske, J. Polym. Sci., 2025, 63(21), 4476–4487 CrossRef CAS.
- M. G. Hogben and W. A. G. Graham, J. Am. Chem. Soc., 1969, 91, 283–291 CrossRef CAS.
- M. N. Leiske, Z. A. I. Mazrad, A. Zelcak, K. Wahi, T. P. Davis, J. A. McCarroll, J. Holst and K. Kempe, Biomacromolecules, 2022, 23, 2374–2387 Search PubMed.
- S. Perrier, Macromolecules, 2017, 50, 7433–7447 Search PubMed.
- G. Landrum, P. Tosco, B. Kelley, R. Rodriguez, D. Cosgrove, R. Vianello, sriniker, P. Gedeck, G. Jones, Kawashima, Eisuke, N. Schneider, D. Nealschneider, A. Dalke, T. Hurst, M. Swain, B. Cole, S. Turk, A. Savelev, A. Vaucher, M. Wójcikowski, N. Maeder, H. Faara, I. Take, Walker, Rachel, Scalfani, F. Vincent, D. Probst, K. Ujihara, Pahl, Axel, Godin, Guillaume, Lehtivarjo, Juuso, J. Adamczyk, C. von Bargen, A. Dirksen, J. A. Mitchell, Pirie, Rachael, Tubert-Brohman, Ivan, marlon2468morales-hue and spparel, RDKit: Open-source cheminformatics, https://www.rdkit.org Search PubMed.
- S. A. Wildman and G. M. Crippen, J. Chem. Inf. Comput. Sci., 1999, 39, 868–873 CrossRef CAS.
- K. Kuroda, G. A. Caputo and W. F. DeGrado, Chem. – Eur. J., 2009, 15, 1123–1133 Search PubMed.
- R. A. Jones, C. Y. Cheung, F. E. Black, J. K. Zia, P. S. Stayton, A. S. Hoffman and M. R. Wilson, Biochem. J., 2003, 372, 65–75 CrossRef CAS PubMed.
- A. M. Mahmoud, P. A. J. M. de Jongh, S. Briere, M. Chen, C. J. Nowell, A. P. R. Johnston, T. P. Davis, D. M. Haddleton and K. Kempe, ACS Appl. Mater. Interfaces, 2019, 11, 31302–31310 Search PubMed.
- Z. Jiang, H. Liu, H. He, N. Yadava, J. J. Chambers and S. Thayumanavan, Bioconjugate Chem., 2020, 31, 1344–1353 CrossRef CAS PubMed.
- M. Ringleb, T. Schuett, S. Zechel and U. S. Schubert, Digit. Discov., 2023, 2, 1883–1893 Search PubMed.
- M. Chen, G. Moad and E. Rizzardo, J. Polym. Sci., Part A: Polym. Chem., 2009, 47, 6704–6714 CrossRef CAS.
- M. N. Leiske, C. Kuenneth, J. de Breuck, B. G. de Geest and R. Hoogenboom, Macromol. Chem. Phys., 2023, 224, 2300200 CrossRef CAS.
- J. I. Solomun, G. Cinar, P. Mapfumo, F. Richter, E. Moek, F. Hausig, L. Martin, S. Hoeppener, I. Nischang and A. Traeger, Int. J. Pharm., 2021, 593, 120080–120094 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |