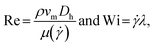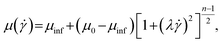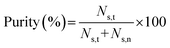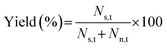Open Access Article
Open Access ArticleIntelligent image-activated sorting of large cells enabled by elasto-inertial focusing
Yuzuki
Nagasaka
 a,
Akihiro
Isozaki
a,
Akihiro
Isozaki
 ab,
Hiroki
Matsumura
ab,
Hiroki
Matsumura
 ac,
Natsumi Tiffany
Ishii
ac,
Natsumi Tiffany
Ishii
 a,
Norah
Roels
a,
Norah
Roels
 a,
Mina
Rassuli
a,
Mina
Rassuli
 d,
Kelvin C. M.
Lee
d,
Kelvin C. M.
Lee
 a,
Walker
Peterson
a,
Walker
Peterson
 a,
Tianben
Ding
a,
Tianben
Ding
 *a and
Keisuke
Goda
*a and
Keisuke
Goda
 *aefghi
*aefghi
aDepartment of Chemistry, Graduate School of Science, The University of Tokyo, Tokyo 113-0033, Japan. E-mail: tding@g.ecc.u-tokyo.ac.jp; goda@chem.s.u-tokyo.ac.jp
bDepartment of Mechanical Engineering, College of Science and Engineering, Ritsumeikan University, Shiga 525-8577, Japan
cDepartment of Chemical and Biomolecular Engineering, Georgia Institute of Technology, Atlanta, Georgia 30322, USA
dDepartment of Chemistry, RWTH Aachen University, Aachen 52074, Germany
eSiRIUS Institute of Medical Research, Tohoku University, Miyagi 980-0872, Japan
fGraduate School of Medicine, Tohoku University, Miyagi 980-0872, Japan
gInternational Centre for Synchrotron Radiation Innovation Smart, Tohoku University, Miyagi 980-8572, Japan
hInstitute of Technological Sciences, Wuhan University, Wuhan, Hubei 430072, China
iDepartment of Bioengineering, University of California, Los Angeles, California 90095, USA
First published on 19th February 2026
Abstract
Image-activated cell sorting (IACS) enables high-speed sorting of cells based on real-time analysis of their images, providing a powerful means to link cellular morphology and function at the single-cell level on a large scale. Although IACS has been demonstrated for a variety of small-to-medium-size cells, applying it to large cells, cell clusters, and other large objects remains challenging, despite the scientific and industrial value in morphology-based sorting of such objects. The main difficulty lies in controlling large and complex cells throughout a microfluidic chip, from inlet to outlet especially the image acquisition and sorting regions. In particular, conventional IACS systems based on hydrodynamic focusing struggle to maintain stable focusing of large objects over the intervals required for processing images, leading to reduced sorting purity, yield, or event rate. To address these limitations, here we report an IACS system based on elasto-inertial focusing which enables IACS of large cells at high flow speeds of ∼1 m s−1. We validated our developed elasto-inertial focuser by demonstrating that particles with a large diameter of >20 μm maintained their positions in the center of the focuser over a long distance of ≥35 mm. We integrated the focuser into the IACS system and sorted size-mixed particles (50% target) using a convolutional neural network-based classifier, demonstrating 96.0% purity and 80.5% yield at an event rate of 172 events per second (eps). Finally, we realized elasto-inertial focusing-based IACS of Euglena gracilis, a large microalgal cell species, based on intracellular lipid droplet formation, demonstrating 4.5-fold enrichment of target cells from 11.9% to 53.8% at 128 eps. Our work highlights that IACS based on elasto-inertial focusing enables the sorting of large objects based on high-content real-time image analysis without compromising sorting purity, yield, or event rate.
Introduction
Image-activated cell sorting (IACS)1–11 enables high-speed sorting of cells based on real-time analysis of their images, providing a powerful means to link cellular morphology and function at the single-cell level on a large scale.12 Going beyond the limits of fluorescence-activated cell sorting (FACS) which is typically based on single-point fluorescence intensity measurements combined with threshold-based real-time analysis,13,14 IACS integrates high-speed microscopy7,15–17 and real-time digital image processing including advanced deep learning-based analysis11 with microfluidics18–21 to extract and analyze morphological and cytochemical features which inform accurate and rapid cell sorting.22,23 IACS has exhibited significant potential in a wide range of applications across biology, medicine, environmental science, and beyond. In the field of hematology, IACS of hemocytes based on cell type or composition is advancing diagnostics and drug treatment of hematological diseases.1,3,6 In genetics, researchers identified regulators in the nuclear factor κB pathway by the combination of CRISPR-pooled screen and IACS, showing the capability of highly-scalable phenotype-based genetic screening.2 In environmental science and biology, researchers made strides in elucidating detailed mechanisms of carbon fixation by microalgal cells via IACS of Chlamydomonas reinhardtii cells based on intracellular mitochondrial localization.24 These achievements underscore the broad scientific value of IACS as a method which opens up multiple important directions for cell-based studies.12Despite these advances, current IACS systems face critical limitations. In particular, IACS of large objects (>20 μm diameter) has been challenging, with prior reports demonstrating IACS of cells with diameters only up to about 20 μm. This limitation is highly consequential, as numerous areas of biomedical and industrial research rely on larger objects including tumor clusters,25–27 spheroids,28 organoids,29 microalgae,30 and hydrogel cell microcarriers.31,32 As a result, key morphological–functional relationships in these systems remain inaccessible. The primary hurdle for applying IACS to large cells comes from the latency between imaging and sorting, which is inherent to the high-content image analysis in IACS. Implementation of deep learning-based analysis, which increases classification accuracy,33 exacerbates the temporal cost of analysis. In order to maintain a high event rate (≥100 events per second or eps; event rate is the total number of sorted and unsorted objects flowing in a system per second),34 a high flow speed (≥1 m s−1) is required, such that the sample travel distance between imaging and sorting actuation must be long (≥30 mm). Over this long interval, objects must remain well focused to achieve a uniform flow velocity and thus precise sort timing, which is essential for sorting with high purity and yield. Conventional IACS systems, however, rely on hydrodynamic focusing19 and thus struggle to maintain stable focusing of large objects over long distances. Shear-gradient lift force, an inertial force that scales nonlinearly with object size, drives large objects towards the microchannel wall,35,36 where they experience slower velocities according to the Hagen–Poiseuille flow profile. This results in stochastic variation in their arrival time at the sorting region. Therefore, existing systems must either restrict the focusing distance and flow speed, thereby sacrifice sorting throughput,3,4,6 or operate at high flow speeds without effective focusing, thereby sacrificing sorting purity and yield.1,2,5 To date, no IACS system has demonstrated high-purity, high-yield, and high-event-rate sorting of large objects under these constraints.
To overcome these limitations, we report elasto-inertial focusing-based IACS, which enables IACS of large objects at a high flow speed of 1 m s−1, without a loss of purity or yield. We achieved this via our design and construction of an IACS system featuring an elasto-inertial focuser, which is a long, straight, 66 mm microchannel filled with viscoelastic fluids. The focuser generates elastic forces that counteract the unavoidable inertial forces, tightly focusing cells at the centerline of the long focuser and ensuring uniform flow velocities of large objects.37–39 Notably, this focusing mechanism becomes increasingly effective as object size increases, allowing elasto-inertial focusing to maintain stable alignment of large objects even over extended distances. We first tested the focusing performance of our 66 mm microchannel by quantifying the spatial distribution of polystyrene (PS) particles as they passed through the focuser, demonstrating improved focusing with increasing particle size. We then applied elasto-inertial focusing-based image-activated sorting to size-mixed PS particles (50% target) using real-time convolutional neural network (CNN)-based image analysis, achieving 96.0% purity and 80.5% yield at an event rate of 172 eps. Finally, we demonstrated elasto-inertial focusing-based IACS of Euglena gracilis (E. gracilis), a unicellular microalgal species with body lengths of 20–50 μm, analyzed and sorted based on intracellular lipid droplet formation. This achieved a 4.54-fold enrichment of target cells, from 11.9% to 53.8% at 128 eps. To the best of our knowledge, this work represents image-activated sorting of the largest particles and largest cells reported to date while maintaining high event rates without compromising spatial resolution. Elasto-inertial focusing-based IACS thus enables high-content, real-time image analysis and sorting of large objects while maintaining sorting purity, yield, and throughput. The system we have developed holds promise for new applications in microalgal metabolic engineering, microalgal strain engineering for biofuel, and beyond.
Results
Elasto-inertial focusing-based IACS system
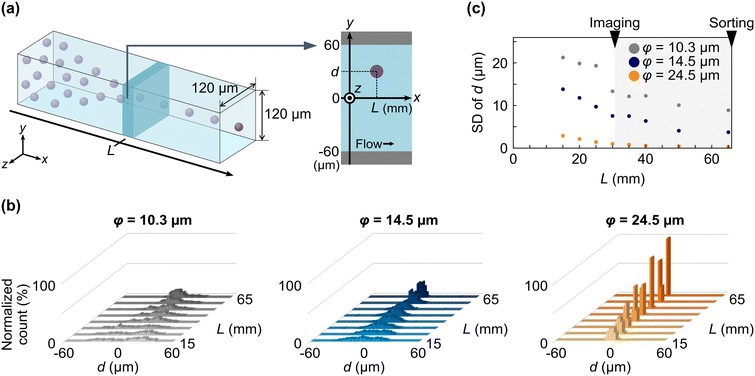
The schematic of elasto-inertial focusing-based IACS developed in this study for image-activated large cell sorting is shown in Fig. 1. This system is based on our previously reported intelligent IACS5 and seamlessly integrates several key components: (i) an elasto-inertial focuser for maintaining the focus of large objects at the center of a long microchannel, (ii) a virtual-freezing fluorescence imaging16-based optomechanical microscope for acquiring blur-free two-channel fluorescence images of cells flowing at a high speed of ∼1 m s−1 at a resolution of 0.361 μm per pixel, (iii) a speed meter for measuring the flowing speed of cells, (iv) an intelligent image processor for processing cell images with the size of 88 pixel × 88 pixel and making sort decision in real time, and (v) on-chip dual-membrane push–pull pumps20 for rapidly isolating target cells during flow through the microfluidic chip. The elasto-inertial focuser we introduce here is a 66 mm-long straight square cross-sectional microchannel with a side length of 120 μm. The on-chip focuser was made of a cycloolefin polymer (COP), together with wells of the dual-membrane pumps. As the focuser maintains cell flow in the center of the microchannel, the cells are imaged by the optomechanical microscope 31–32 mm from the inlet, and the resulting images are analyzed by the intelligent image processor. Using sort timing predictions by the speed meter, target cells are sorted by the dual-membrane pumps. Details of the design of the microfluidic chip, calculation of sort timing, and functionality of the dual membrane pumps are described in the Materials and methods section, and Fig. S1–S4 of the SI. In this study, we used 0.1% (w/v) sodium hyaluronate (HA) solutions for focusing particles and E. gracilis cells, inspired by a previous report38 that demonstrated elasto-inertial focusing in moderate to exceptionally high Reynolds regimes that cover our high-flow-speed condition. HA is a biocompatible polymer, and its diluted solutions have widely been used for elasto-inertial focusing-based microfluidic particle and cell manipulations.39–41 We set the flow rate at 420 μL min−1, corresponding to a maximum particle speed of ∼1 m s−1 at the channel center. Details of the rheological parameters of the HA solutions and simulations of the flow profile are described in the Materials and methods section and Fig. S5 of the SI. Although HA was the only material tested in this study, different shear-thinning solutions of polymers, such as polyethylene glycol and polyvinylpyrrolidone, could also be selected, under the condition that the resulting fluid pressure inside the focuser does not exceed the maximum durable pressure of the microfluidic chip.Evaluation of the elasto-inertial focuser
We evaluated the performance of the elasto-inertial focuser using PS particles of different sizes. Three suspensions were prepared with particle diameters (φ) of 10.3 μm, 14.5 μm, and 24.5 μm in 0.1% sodium hyaluronate (HA) dissolved in Dulbecco's phosphate-buffered saline without calcium and magnesium (D-PBS(−)). Each suspension was injected into the focuser at a flow rate of 420 μL min−1, corresponding to a maximum flow speed of ∼1 m s−1 at the channel center. Sample concentrations were adjusted to 3.8 × 104 cells per mL, resulting in an event rate of ∼250 eps. Assuming that the particle flow in the center of the focuser is governed by Poisson process, the probability of the distance of successive focused particles larger than S m is given by P(s > S) = e−250S. Therefore, the particles in this evaluation were sparsely distributed in the focuser with S = 205 μm at 95% probability (P = 0.95), corresponding to >8.3 times the diameter of the largest particles (24.5 μm). We recorded particles flowing in the microchannel at different distances from the inlet (L) by a high-speed camera. Using the recordings, we measured each particle's vertical distance from the microchannel center (d) at various Ls [Fig. 2(a); see the Materials and methods section for the experimental details]. Fig. 2(b) shows histograms of d as a function of L, measured for each PS particle size. As L increased, all particle sizes showed progressively narrower single-peak distributions centered at −2 μm < d < 2 μm, indicating effective focusing in the elasto-inertial focuser. For more quantitative analysis, the standard deviation (SD) of d was plotted against L for each particle sample [Fig. 2(c)]. The SD of d for 24.5 μm and 14.5 μm particles decreased as L increased, reaching <2 μm and <8 μm, respectively, at L > 30 mm. This L range covers the entire distance from the imaging area around L = 31–32 mm to the upstream edge of the sorting area at L = 66 mm. While 10.3 μm particles also showed a gradual reduction in SD with L, the overall focusing was less effective than that of the larger particles, only achieving an SD of < 12 μm at L > 45 mm. These results indicate that our focuser can reliably maintain the focusing of objects with a diameter of at least 14.5 μm for 36 ms at a flow speed of 1 m s−1, including particles as large as 24.5 μm, exceeding the object sizes previously demonstrated with IACS.To validate elasto-inertial focusing for biological cells, we next flowed E. gracilis cells through the focuser at the same flow rate and compared their focusing performance to that of the 14.5 μm particles, whose diameter is comparable to the cell width (minor axis of the ellipsoidal cell shape). The distribution of displacement d of the cells [Fig. 3(a)] followed a similar trend to that of the particles, exhibiting progressively narrower peaks within −2 μm < d < 2 μm, as L increased. However, the SD of d for the cells plateaued at approximately 10 μm for L > 30 mm, more than twice the value observed for the particles [Fig. 3(b)]. The reduced focusing performance of the cells can be mainly attributed to greater variability in cell morphology [Fig. S6(a)]. The minor and major axes of the cells were 14.9 ± 2.0 μm (SD) and 28.7 ± 3.7 μm, respectively, representing broader distributions than those of the particles, which exhibited minor and major axes of 15.4 ± 1.3 μm (SD) and 16.7 ± 1.6 μm, respectively, as calculated from the video frames. Further analysis of the SDs of d as a function of cell area, aspect ratio, minor axis, and major axis [Fig. S6(b)] revealed that larger aspect ratios and shorter minor axes were associated with impaired focusing performance.
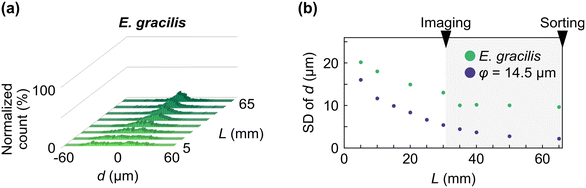 | ||
| Fig. 3 Evaluation of elasto-inertial focusing with E. gracilis cells. (a) Histogram of d for E. gracilis cells measured at various L values. (b) SD of d measured at different L values. | ||
Image-activated particle sorting
To validate our elasto-inertial focusing-based IACS system, we performed image-activated sorting of fluorescent PS particles (Fig. 4 and Video S1). A 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mixture of 24.5 μm and 14.5 μm PS particles was prepared, with the 24.5 μm particles designated as the sorting target. Prior to sorting, each particle population was imaged separately by the IACS system [Fig. 5(a)]. A LeNet-5-dropout CNN model42 was pre-trained with 3635 target and 6863 non-target IACS images. As shown in Fig. 5(b), both training and validation accuracy approached nearly 100% after 20 epochs. The model trained for 20 epochs achieved a precision of 99.88% and a recall of 99.89% when tested on held-out data [Fig. 5(c)]. This model was then applied for real-time classification during IACS of the particle mixture. Fig. 5(d) shows the rank-ordered processing time for image analysis and data transfer of each event during sorting. At an event rate of 172 eps, a total of 96.9% of events were processed within 32 ms, the practical minimum time in our IACS system required for image processing and decision-making before particles travel from the imaging to the sorting area, indicating that most particle images were classified before sorting actuation. At this event rate, 852 target particles and 36 non-target particles flowed into the upper or the lower microchannel branches connected to the sorted fraction in 10.72 s, while 206 target particles and 750 non-target particles passed through the middle branch to the not-sorted collection (Table 1). The calculated purity and yield were 96.0% and 80.5%, respectively. Purity remained high at increased event rates of 264 eps (92.6%) and 312 eps (92.6%). On the other hand, as the event rate increased, the fraction of events processed within 32 ms decreased to 61.0% at 264 eps and 32.7% at 312 eps [Fig. 5(d)], resulting in a corresponding reduction in sorting yield to 51.9% and 23.1%, respectively (Table 1). These results highlight the requirement for longer processing times in IACS systems using deep learning-based classification models and thus the importance of maintaining extended focused flow of large objects.
1 mixture of 24.5 μm and 14.5 μm PS particles was prepared, with the 24.5 μm particles designated as the sorting target. Prior to sorting, each particle population was imaged separately by the IACS system [Fig. 5(a)]. A LeNet-5-dropout CNN model42 was pre-trained with 3635 target and 6863 non-target IACS images. As shown in Fig. 5(b), both training and validation accuracy approached nearly 100% after 20 epochs. The model trained for 20 epochs achieved a precision of 99.88% and a recall of 99.89% when tested on held-out data [Fig. 5(c)]. This model was then applied for real-time classification during IACS of the particle mixture. Fig. 5(d) shows the rank-ordered processing time for image analysis and data transfer of each event during sorting. At an event rate of 172 eps, a total of 96.9% of events were processed within 32 ms, the practical minimum time in our IACS system required for image processing and decision-making before particles travel from the imaging to the sorting area, indicating that most particle images were classified before sorting actuation. At this event rate, 852 target particles and 36 non-target particles flowed into the upper or the lower microchannel branches connected to the sorted fraction in 10.72 s, while 206 target particles and 750 non-target particles passed through the middle branch to the not-sorted collection (Table 1). The calculated purity and yield were 96.0% and 80.5%, respectively. Purity remained high at increased event rates of 264 eps (92.6%) and 312 eps (92.6%). On the other hand, as the event rate increased, the fraction of events processed within 32 ms decreased to 61.0% at 264 eps and 32.7% at 312 eps [Fig. 5(d)], resulting in a corresponding reduction in sorting yield to 51.9% and 23.1%, respectively (Table 1). These results highlight the requirement for longer processing times in IACS systems using deep learning-based classification models and thus the importance of maintaining extended focused flow of large objects.
 | ||
| Fig. 4 Sequential photos of image-activated particle sorting. Red: target particles. Blue: non-target particles. Scale bar, 200 μm. | ||
| Event rate (eps) | N s,t | N s,n | N n,t | N n,n | Purity (%) | Yield (%) |
|---|---|---|---|---|---|---|
| 172 | 852 | 36 | 206 | 750 | 96.0 | 80.5 |
| 264 | 833 | 66 | 761 | 1176 | 92.6 | 51.9 |
| 312 | 446 | 37 | 1488 | 1376 | 92.3 | 23.1 |
Image-activated sorting of E. gracilis cells
Finally, we performed IACS of E. gracilis cells (microalgal cells with widths of 5–20 μm and lengths of 20–50 μm) based on intracellular lipid droplet content, a morphological feature requiring images for cell classification, judged by the system's CNN model. To prepare the image datasets for the model, we developed a custom python program that detects lipid droplet centers by thresholding local intensity maxima of BODIPY 505/515 fluorescence. This program was applied to cell images acquired by our IACS system. Cells in which lipid droplets were detected were labeled as sorting targets, while those without droplets were labeled as non-targets for CNN model construction [Fig. 6(a)]. An 8-layer CNN1 was trained using 3635 target images and 6863 non-target images [Fig. 6(b)]. The trained model achieved 82.0% precision and 97.5% recall at the 217th epoch [Fig. 6(c)].We then applied this trained CNN model for image-activated sorting of E. gracilis cells. Image libraries of target and non-target cells acquired in real time during IACS are displayed in Fig. 7 and S7. Cells classified as targets exhibit stronger and more distinct droplet patterns in the BODIPY fluorescence channel, whereas cells categorized as non-targets show little signal in the BODIPY channel or cropping artifacts. At an event rate of 128 eps, 88.2% of the images were processed within 32 ms by the CNN-based classification [Fig. 8(a)]. After sorting, we evaluated the CNN model's decisions by applying the same custom lipid droplet counting program described above. As shown in Fig. 8(b), 73.67% of target cells (3638/4938 cells) were correctly classified by the CNN model as containing ≥1 droplets, while 5.12% of non-target cells (915/17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 867 cells) were misclassified as containing ≥1 droplets.
867 cells) were misclassified as containing ≥1 droplets.
We further assessed aliquots of the sorted collections from the IACS system. Pre-sorted and post-sorted cells collected from the microfluidic chip outlets were separately concentrated by centrifugation, transferred onto slides, and imaged under a commercial fluorescence microscope. The same program used to label IACS images for CNN model construction was applied to these microscope images to count cells containing lipid droplets. Among 160 post-sorted cells examined, 86 cells (53.8%) contained lipid droplets, whereas only 32 of 270 pre-sorted cells (11.9%) contained droplets [Fig. 8(c)], corresponding to a 4.54-fold enrichment achieved by our elasto-inertial focusing-based IACS of E. gracilis cells. These results indicate that our system enables high-speed sorting of large cells based on content-rich and real-time image analysis. It should be noted that the lower purity compared with the particle sorting at equivalent or higher event rates can be attributed to reduced focusing performance. As shown in Fig. 3(b), the focusing of E. gracilis cells was less stable than that of 14.5 μm particles. Fluctuations in focusing lead to variability in cell flow velocity, which complicates precise synchronization of the sorting actuation and consequently degrades sorting accuracy.
Discussion
In this study, we proposed and experimentally demonstrated elasto-inertial focusing-based IACS, which enabled high-speed sorting of large cells based on high-content and real-time image analysis without compromising sorting purity, yield, or throughput. By combining elasto-inertial focusing with a long microchannel, large cells could be focused over extended distances, allowing sufficient computational time for deep learning-based image analysis while maintaining a high flow speed. We demonstrated the capability of our system first via image-activated sorting of large PS particles at an event rate of 172 eps with 96.0% purity and 80.5% yield, and then via IACS of E. gracilis cells at an event rate of 128 eps with a real-time image classification accuracy of 73.67% and a 4.54-fold enrichment ratio. For additional context, we provide a tabulated comparison (Table S4) of our system with other reported IACS systems compatible with deep learning-based, content-rich image analysis. Our system uniquely demonstrates IACS of the largest cells to date while maintaining a high event rate of >100 eps. Furthermore, our elasto-inertial focuser operates without sheath flow, eliminating the need for complex microchannel designs or sophisticated flow-control modules.The technical capability of our elasto-inertial focusing-based IACS system can be further extended. First, throughput could be improved by optimizing the computational network. In this work, limited processing speed resulted in a reduced proportion of images processed within the defined 32 ms window, decreasing yield to 51.9% at 264 eps and 23.1% at 312 eps [Fig. 5(d) and Table 1]. Enhancing computer hardware or incorporating parallel computing would boost computational capability, allowing a higher proportion of images to be processed within the defined time. Improvements in microfluidic channel design, including the elasto-inertial focuser and dual membrane pump wells, could also enhance performance. For example, lengthening the focuser would give additional time for image processing. In this study, we employed COP-based on-chip dual-membrane push–pull pumps for sorting actuation. Although COP-based microfluidic chips are relatively cost-effective, the maximum achievable actuation frequency is limited by the material's Young's modulus. This limitation could be addressed by fabricating chips from stiffer materials (e.g., borosilicate glass) or by adopting faster sorting mechanisms (e.g., acoustic-based sorting or microbubble-based sorting).43–45 It is worth noting that our work represents the first demonstration of combining viscoelastic fluid focusing with piezoelectric actuator-based active sorting (i.e., sorting activated by forces applied external to the microchannel). Optimizing the design of the sorting area could further improve throughput, yield, and purity. We anticipate that these computational and microfluidic improvements will also enable the integration of classification methods capable of higher-content images, such as multi-channel, three-dimensional (3D), and super-resolution images, without compromising throughput. In particular, implementing 3D image will be essential for large objects, as 3D images reflect complete morphological features with greater accuracy and reproducibility than single-plane 2D images.46 One potential limitation to achieving stable sorting performance for large objects is inter-particle spacing. As object size and volumetric concentration increase, neighboring objects are more likely to undergo coupled oscillatory motion,47 leading to increased flow-velocity fluctuations that can degrade sorting yield and purity. Further investigation and application-specific optimization of sorting performance may be required.
In terms of further applications, we expect our IACS system to be applicable to a variety of large cells or cell clusters to achieve IACS based on diverse morphological features. In the case of E. gracilis, we demonstrated IACS based on the existence of condensed lipid droplets, a feature that previous high-throughput microfluidic sorting techniques20,48,49 could not exploit. This capability could potentially enable comprehensive investigation of phenotype–genotype correlations related to lipid droplet condensation, which would broaden insights into E. gracilis and supporting applications in metabolic engineering and strain engineering for biofuel production. Moreover, our IACS system could sort E. gracilis cells based on combinations of morphological features to enrich for subpopulations specialized in specific biochemical compound production, photosynthetic activities, and heavy metal removal.50,51 Beyond E. gracilis, we anticipate that our system could be extended to high-throughput sorting of larger biological objects, particularly spheroids and organoids cultured from patients' cells. Although they are expected to serve as scalable, individualized disease models to evaluate the efficacy of therapeutic treatments in vitro,52–54 their intrinsic morphological and compositional variations cause low reproducibility and reliability in efficacy testing, positing a challenge toward their clinical applications.55–58 Isolating organoids or spheroids with specific morphological features could reduce variation in drug responses, enabling more reliable personalized prognosis. While the focuser developed in this study still has limitations, including inefficient focusing of small single cells (<10 μm diameter) or incompatibility with objects approaching or exceeding the microchannel side length (120 μm), these challenges can be addressed through design optimization. For example, fabricating a focuser with a large cross section would extend its applicability to larger sorting targets. We tested focuser designs with a wider cross-section (150 μm side length) and confirmed effective elasto-inertial focusing over long distances for particles 14.5 μm, 24.5 μm, and 51.0 μm in diameter [Fig. S8(a)]. We anticipate that even larger microchannels could be integrated into the IACS system for sorting larger objects, since elasto-inertial focusing has been demonstrated across broad ranges of rheological parameters.38,59,60 In live cell sorting applications, cell viability under elasto-inertial focusing is an important consideration. A previous study61 reported reduced viability of E. gracilis cells after flow through a narrow microchannel in a FACS system, indicating susceptibility to high shear stress. Thus, careful control of stress will be essential when modifying focuser and sorter designs for live-cell applications. Through such adjustments, our IACS system could be adapted to more versatile needs, enabling comprehensive correlation analysis between morphological features and cellular functions in large cells or cell clusters across biomedical and environmental applications.
Materials and methods
Principle of elasto-inertial focusing
Elasto-inertial focusing arises from the balance between fluid inertia and elasticity. Specifically, the relationship between Reynolds number (Re) and Weissenberg number (Wi), which index the relative strength of inertia and elasticity to fluid viscosity, respectively, satisfies 0 < Wi/Re and is not Wi/Re ≫ 1 in elasto-inertial focusing.39 Re and Wi are given bywhere ρ, vm, Dh, μ(
![[small gamma, Greek, dot above]](https://www.rsc.org/images/entities/i_char_e0a2.gif) ),
), ![[small gamma, Greek, dot above]](https://www.rsc.org/images/entities/i_char_e0a2.gif) , and λ are density, average flow velocity, hydraulic diameter, viscosity, characteristic (average) shear rate, and relaxation time of the fluid, respectively. In the case of a microchannel with a square cross-section, Dh corresponds to the microchannel side length w. In this study, elasto-inertial focusing was achieved using shear-thinning 0.1% (v/w) HA solutions prepared from HA powder with a molecular weight of 1.01–1.8 MDa (Lifecore Biomedical, USA) dissolved in D-PBS(−) (Wako Chemicals, Japan) for the particle samples or in 1
, and λ are density, average flow velocity, hydraulic diameter, viscosity, characteristic (average) shear rate, and relaxation time of the fluid, respectively. In the case of a microchannel with a square cross-section, Dh corresponds to the microchannel side length w. In this study, elasto-inertial focusing was achieved using shear-thinning 0.1% (v/w) HA solutions prepared from HA powder with a molecular weight of 1.01–1.8 MDa (Lifecore Biomedical, USA) dissolved in D-PBS(−) (Wako Chemicals, Japan) for the particle samples or in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 volume ratio of D-PBS(−) and AF-6 without carbon sources (National Institute for Environmental Studies, Japan) for the E. gracilis cell samples.38
1 volume ratio of D-PBS(−) and AF-6 without carbon sources (National Institute for Environmental Studies, Japan) for the E. gracilis cell samples.38
Construction of the microfluidic chip and its integration into the IACS
The microfluidic chip in this study was specifically designed for the IACS system and fabricated by ZEON Japan Co. from a 2 mm-thick COP plate (ZEONEX690R) with the microchannel structure, an inlet hole, and outlet holes, and a 0.188 mm-thick COP film (ZE14) to seal the microchannel. The detailed chip dimensions are depicted in Fig. S1 and Design S1. COP-based chips offer high transparency to visible light and can be fabricated simply and cost-effectively. The microfluidic chip was mounted between custom-made aluminum chip holders that secured its position relative to piezoelectric actuators (AE0203D04DF, Tokin, Japan) using parallel pins. Samples were introduced into the chip via tube connectors (M-3AU-2, SMC, Japan) and sealed with O-rings (SS-005-1A, Musashi Oil Seal, Japan) [Fig. S2(a)]. The holder unit was set to the chip holder slot [Fig. S2(b)], integrating the chip into our IACS system. The 3D model of the holder unit and the chip holder is available as Design S2.Measurement of cell flow speed and determination of sort timing in the IACS system
The timing of sort actuation was determined using forward-scattered signals from three infrared (IR) laser spots [Fig. S3(a)]. Two of these (“IR1” and “IR2”) are positioned in the imaging area at L = 31–32 mm with IR2 400 μm downstream of IR1, and the other (“IR3”) was placed at L = 66 mm, 34–35 mm downstream of IR1 (>32 ms for particles flowing at ∼1 m s−1). The passage times of each particle or cell at the three laser positions (tIR1, tIR2, and tIR3) were recorded and plotted in real time on a graph of tIR2–tIR1versus tIR3–tIR1. A straight line with slope k was determined which was fitted to the populated area of the plot. The sort actuation timing for the n-th particle (tout(n)) was calculated as: tout(n) = tIR1(n) + k(tIR2(n) − tIR1(n)). For example, Fig. S3(b) shows the scatter plot of IR timings measured from particles classified as target during sorting, with the fitted line corresponding to k = 95.1.Functionality of dual-membrane push–pull pumps
Fig. S4(a) schematically shows the components and functionality of the dual-membrane push–pull pumps, which consist of piezoelectric actuators, attachment balls (SBM-SUJ-2, Tsubaki Nakashima, Japan), custom-made aluminum plates (diameter: 3 mm, thickness: 0.5 mm), and COP-based film membranes (diameter: 4 mm, thickness: 0.188 mm). The membrane pumps sort target cells from the flowing cell population as follows: at the sort timing of the n-th target particle, a linear increase of voltage (V) is applied to one of the piezoelectric actuators (piezo A), causing its membrane to be pushed via the attachment ball and aluminum plate. Simultaneously, the other actuator (piezo B) receives a decreased voltage, pulling its membrane [Fig. S4(b)]. This coordinated push–pull action generates a local flow from piezo A to piezo B, directing the target cell into the upper channel at the junction. At the next sorting event, the voltages are reversed, piezo A pulls while piezo B pushes, producing a flow from piezo B to piezo A and directing the target cell into the lower channel [Fig. S4(a)]. Repetition of these alternating push–pull movements enables successive sorting of target cells.20 In this study, the applied voltage (V) and the voltage change duration (τ) were set to 40 V and 400 μs, respectively.Simulation of flow velocity and shear rate distributions in the elasto-inertial focuser
To characterize the flow field in the elasto-inertial focuser, the flow velocity and shear rate distributions where simulated based on the finite-element method implemented in COMSOL Multiphysics (version 6.0, COMSOL, USA). As shown in Fig. S5(a), the computational domain was a straight microchannel modeled as a cuboid with a length (x direction) of 500 μm and a square cross-section with a height (y direction), and depth (z direction) of 120 μm. A tetrahedral mesh was applied with minimum and maximum sizes of 1 μm and 3 μm, respectively. The simulated fluids were water and a 0.1% (w/v) HA (1650 kDa) D-PBS(−) solution, corresponding to the fluid used in the particle focusing and sorting experiments. Steady-state flow was assumed between the inlet plane at x = −250 μm and the outlet plane x = 250 μm, with a fixed volumetric flow rate of 420 μL min−1. The HA solution was modeled as a shear-thinning fluid using the Carreau model:38where μinf, μ0, λ, and n are the infinite rate viscosity, zero-shear-rate viscosity, relaxation time, and power-exponent factor, respectively. The rheological parameters used for water and the HA solution are listed in Table S1. Flow velocity and shear-rate profiles were evaluated on the plane x = 0 μm, along the horizontal line (x, y) = (0, 0), and along the diagonal line defined by x = 0 and y = z [Fig. S5(b)]. As shown in Fig. S5(c), water exhibited a parabolic velocity profile with velocities ranging from 0 m s−1 at the channel walls to a maximum of 1.0 m s−1 at the microchannel center. In contrast, the HA solution showed a plug-like velocity profile with a lower maximum centerline velocity of 0.85 m s−1. The corresponding shear-rate distributions [Fig. S5(d)] showed that the shear rate is near zero at the channel centerline and at the channel corners, while reaching its maximum near the midpoints of the channel side walls (4 × 104 s−1 for water and 5 × 104 s−1 for HA).
Experimental evaluation of elasto-inertial focusing
The microfluidic chip, mounted between the chip holders, was set on an inverted microscope (IX73, Olympus, Japan) equipped with a high-speed camera (Phantom v9154, Vision Research, USA). Particles flowed at different positions along the microchannel were recorded at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 frames per s with an exposure time of 10 μs. Using ImageJ, the positions of gravity center of each particle were measured from the video frames, and each particle's vertical distance from the microchannel center (d) and standard deviation (SD) of d at different positions from the inlet (L) were calculated. In addition, ellipse approximation was performed on each cell image on the video frames, and minor and major axes of the ellipses were extracted for the analysis of correlations between cell shapes and focusing performance shown in Fig. S6.
000 frames per s with an exposure time of 10 μs. Using ImageJ, the positions of gravity center of each particle were measured from the video frames, and each particle's vertical distance from the microchannel center (d) and standard deviation (SD) of d at different positions from the inlet (L) were calculated. In addition, ellipse approximation was performed on each cell image on the video frames, and minor and major axes of the ellipses were extracted for the analysis of correlations between cell shapes and focusing performance shown in Fig. S6.
Construction of CNN models for sorting particles and E. gracilis cells
Target and non-target image datasets were used to train and validate the CNN models (LeNet-5-dropout model for particle images and 8-layer CNN model for E. gracilis cell images) on AIDeveloper,62 a python-based open-source software. For the CNN-based sorting demonstration, the trained model was integrated into the C++-based image analysis algorithm in the previously reported intelligent IACS system11 for real-time communication. The images used for CNN training and sort decision-making were 88 × 88 pixels in size.Measurement of sorting performance in particle sorting
Sorting performance was evaluated in terms of throughput, purity, and yield, based on counts of target and non-target particles flowing through the sort area, recorded by a high-speed camera (Phantom v1211, Vision Research, USA). For each sorting experiment, videos were recorded for a total duration T s. The throughput was found by the total number of particles (C) in the video divided by T. Here, T was fixed to 10.72 s among all the particle sorting experiments. Purity and yield were given by:where Ns,t is the number of sorted target particles, Ns,n the number of sorted non-target particles, and Nn,t the number of not-sorted target particles.
Preparation of E. gracilis cells
E. gracilis cells (NIES-48, Microbial Culture Collection at the National Institute for Environmental Studies, Japan) were cultured in AF-6 medium free of carbon sources and incubated at 25 °C under a 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 light–dark cycle. One month prior to imaging and sorting experiments, cells in the growth phase were transferred to AF-6 medium without nitrogen (AF-6N-). The AF-6N- medium was prepared from scratch (its components are listed in Table S3). Intracellular lipids were stained with BODIPY 505/515 (Thermo Fisher Scientific, USA), a green-fluorescent nonpolar lipid dye with minimal spectral overlap with autofluorescence of chlorophyll. BODIPY 505/515 powder was dissolved in dimethyl sulfoxide at 1 mM, stored at −20 °C in the dark as a stock solution, and diluted by deionized water to 10 μM immediately before staining. Cells were washed by deionized water, resuspended in the 10 μM BODIPY solution at a density of 1 × 105 cells per mL, and incubated overnight in the dark at 25 °C. On the following day, cells were washed with deionized water and resuspended in commercial AF-6 medium at twice the final density. For elasto-inertial focusing, a 0.2% HA solution in D-PBS(−) was added to the cell suspension at a 1
10 light–dark cycle. One month prior to imaging and sorting experiments, cells in the growth phase were transferred to AF-6 medium without nitrogen (AF-6N-). The AF-6N- medium was prepared from scratch (its components are listed in Table S3). Intracellular lipids were stained with BODIPY 505/515 (Thermo Fisher Scientific, USA), a green-fluorescent nonpolar lipid dye with minimal spectral overlap with autofluorescence of chlorophyll. BODIPY 505/515 powder was dissolved in dimethyl sulfoxide at 1 mM, stored at −20 °C in the dark as a stock solution, and diluted by deionized water to 10 μM immediately before staining. Cells were washed by deionized water, resuspended in the 10 μM BODIPY solution at a density of 1 × 105 cells per mL, and incubated overnight in the dark at 25 °C. On the following day, cells were washed with deionized water and resuspended in commercial AF-6 medium at twice the final density. For elasto-inertial focusing, a 0.2% HA solution in D-PBS(−) was added to the cell suspension at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 volume ratio, resulting in a final suspension of cells in 0.1% HA solution with a 1
1 volume ratio, resulting in a final suspension of cells in 0.1% HA solution with a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 volume ratio of AF-6 and D-PBS(−). We confirmed using particle samples that elasto-inertial focusing performance in this 0.1% HA, AF-6
1 volume ratio of AF-6 and D-PBS(−). We confirmed using particle samples that elasto-inertial focusing performance in this 0.1% HA, AF-6![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D-PBS(−) mixture was comparable to that obtained with 0.1% HA in D-PBS(−) alone [Fig. S8(b)].
D-PBS(−) mixture was comparable to that obtained with 0.1% HA in D-PBS(−) alone [Fig. S8(b)].
Author contributions
Yuzuki Nagasaka: conceptualization (equal), methodology (lead), software (lead), validation (lead), formal analysis (lead), investigation (lead), writing – original draft (lead), writing – review & editing (equal), visualization (lead), project administration (equal). Akihiro Isozaki: conceptualization (lead), methodology (equal), writing – review & editing (equal), supervision (equal), funding acquisition (equal). Hiroki Matsumura: investigation (equal), writing – review & editing (equal). Natsumi Tiffany Ishii: investigation (equal), writing – review & editing (equal). Norah Roels: validation (equal), writing – review & editing (equal). Mina Rassuli: validation (equal), writing – review & editing (equal). Kelvin C. M. Lee: methodology (equal), investigation (equal), writing – review & editing (equal), Walker Peterson: writing – review & editing (equal), supervision (equal). Tianben Ding: conceptualization (equal), writing – review & editing (equal), supervision (lead), project administration (lead), funding acquisition (equal). Keisuke Goda: conceptualization (equal), writing – review & editing (lead), project administration (equal), supervision (equal), funding acquisition (lead).Conflicts of interest
K. G. is an inventor on patents covering the virtual-freezing fluorescence imaging microscope and the data analysis and display method. K. G. is a shareholder of CYBO.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Additional raw data are available from the corresponding author upon reasonable request.Supplementary information is available. See DOI: https://doi.org/10.1039/d5lc01143d.
Acknowledgements
This work was supported by JSPS Core-to-Core Program JPJSCCA20190007, JSPS KAKENHI grant numbers 19H05633, 20H00317, 21H01778 and 24K17800, JST Spring GX JPMJSP2108, The Mazda Foundation, Institute for Fermentation Osaka, Mishima Kaiun Memorial Foundation, Bay Spec Inc., and White Rock Foundation. We thank Dr. Yuta Nakagawa for insightful discussion regarding microfluidic chip design and fabrication. We also extend our gratitude to Serendipity Lab for facilitating collaboration opportunities.References
- N. Nitta, T. Sugimura, A. Isozaki, H. Mikami, K. Hiraki, S. Sakuma, T. Iino, F. Arai, T. Endo, Y. Fujiwaki, H. Fukuzawa, M. Hase, T. Hayakawa, K. Hiramatsu, Y. Hoshino, M. Inaba, T. Ito, H. Karakawa, Y. Kasai, K. Koizumi, S. W. Lee, C. Lei, M. Li, T. Maeno, S. Matsusaka, D. Murakami, A. Nakagawa, Y. Oguchi, M. Oikawa, T. Ota, K. Shiba, H. Shintaku, Y. Shirasaki, K. Suga, Y. Suzuki, N. Suzuki, Y. Tanaka, H. Tezuka, C. Toyokawa, Y. Yalikun, M. Yamada, M. Yamagishi, T. Yamano, A. Yasumoto, Y. Yatomi, M. Yazawa, D. Di Carlo, Y. Hosokawa, S. Uemura, Y. Ozeki and K. Goda, Cell, 2018, 175, 266–276 CrossRef CAS PubMed.
- D. Schraivogel, T. M. Kuhn, B. Rauscher, M. Rodríguez-Martínez, M. Paulsen, K. Owsley, A. Middlebrook, C. Tischer, B. Ramasz, D. Ordoñez-Rueda, M. Dees, S. Cuylen-Haering, E. Diebold and L. M. Steinmetz, Science, 2022, 375, 315–320 CrossRef CAS PubMed.
- R. Tang, L. Xia, B. Gutierrez, I. Gagne, A. Munoz, K. Eribez, N. Jagnandan, X. Chen, Z. Zhang, L. Waller, W. Alaynick, S. H. Cho, C. An and Y. H. Lo, Biosens. Bioelectron., 2022, 220, 114865 CrossRef PubMed.
- K. Lee, S.-E. Kim, J. Doh, K. Kim and W. K. Chung, Lab Chip, 2021, 21, 1798–1810 RSC.
- A. Isozaki, H. Mikami, H. Tezuka, H. Matsumura, K. Huang, M. Akamine, K. Hiramatsu, T. Iino, T. Ito, H. Karakawa, Y. Kasai, Y. Li, Y. Nakagawa, S. Ohnuki, T. Ota, Y. Qian, S. Sakuma, T. Sekiya, Y. Shirasaki, N. Suzuki, E. Tayyabi, T. Wakamiya, M. Xu, M. Yamagishi, H. Yan, Q. Yu, S. Yan, D. Yuan, W. Zhang, Y. Zhao, F. Arai, R. E. Campbell, C. Danelon, D. Di Carlo, K. Hiraki, Y. Hoshino, Y. Hosokawa, M. Inaba, A. Nakagawa, Y. Ohya, M. Oikawa, S. Uemura, Y. Ozeki, T. Sugimura, N. Nitta and K. Goda, Lab Chip, 2020, 20, 2263–2273 Search PubMed.
- M. Salek, N. Li, H.-P. Chou, K. Saini, A. Jovic, K. B. Jacobs, C. Johnson, V. Lu, E. J. Lee, C. Chang, P. Nguyen, J. Mei, K. P. Pant, A. Y. Wong-Thai, Q. F. Smith, S. Huang, R. Chow, J. Cruz, J. Walker, B. Chan, T. J. Musci, E. A. Ashley and M. Masaeli, Commun. Biol., 2023, 6, 971 CrossRef PubMed.
- X. Chen, Y. Gu, J. Chen, C.-H. Lee, I. Gagne, R. Tang, L. Waller, Z. Zhang, A. C. Zhang, Y. Han, W. Wang, I. Y. Lian, S. H. Cho and Y.-H. Lo, APL Photonics, 2020, 5, 040801 Search PubMed.
- T. Gerling, N. Godino, F. Pfisterer, N. Hupf and M. Kirschbaum, Lab Chip, 2023, 23, 3172–3185 RSC.
- A. A. Nawaz, M. Urbanska, M. Herbig, M. Nötzel, M. Kräter, P. Rosendahl, C. Herold, N. Toepfner, M. Kubánková, R. Goswami, S. Abuhattum, F. Reichel, P. Müller, A. Taubenberger, S. Girardo, A. Jacobi and J. Guck, Nat. Methods, 2020, 17, 595–599 CrossRef CAS PubMed.
- N. Nitta, T. Iino, A. Isozaki, M. Yamagishi, Y. Kitahama, S. Sakuma, Y. Suzuki, H. Tezuka, M. Oikawa, F. Arai, T. Asai, D. Deng, H. Fukuzawa, M. Hase, T. Hasunuma, T. Hayakawa, K. Hiraki, K. Hiramatsu, Y. Hoshino, M. Inaba, Y. Inoue, T. Ito, M. Kajikawa, H. Karakawa, Y. Kasai, Y. Kato, H. Kobayashi, C. Lei, S. Matsusaka, H. Mikami, A. Nakagawa, K. Numata, T. Ota, T. Sekiya, K. Shiba, Y. Shirasaki, N. Suzuki, S. Tanaka, S. Ueno, H. Watarai, T. Yamano, M. Yazawa, Y. Yonamine, D. Di Carlo, Y. Hosokawa, S. Uemura, T. Sugimura, Y. Ozeki and K. Goda, Nat. Commun., 2020, 11, 3452 CrossRef CAS PubMed.
- A. Isozaki, H. Mikami, K. Hiramatsu, S. Sakuma, Y. Kasai, T. Iino, T. Yamano, A. Yasumoto, Y. Oguchi, N. Suzuki, Y. Shirasaki, T. Endo, T. Ito, K. Hiraki, M. Yamada, S. Matsusaka, T. Hayakawa, H. Fukuzawa, Y. Yatomi, F. Arai, D. Di Carlo, A. Nakagawa, Y. Hoshino, Y. Hosokawa, S. Uemura, T. Sugimura, Y. Ozeki, N. Nitta and K. Goda, Nat. Protoc., 2019, 14, 2370–2415 CrossRef CAS PubMed.
- T. Ding, K. C. M. Lee, K. K. Tsia, T. N. Siegel, D. Di Carlo and K. Goda, Nat. Rev. Bioeng., 2025, 3, 890–907 CrossRef CAS.
- L. A. Herzenberg, D. Parks, B. Sahaf, O. Perez, M. Roederer and L. A. Herzenberg, Clin. Chem., 2002, 48, 1819–1827 CrossRef CAS.
- A. Cossarizza, H. D. Chang, A. Radbruch, S. Abrignani, R. Addo, M. Akdis, I. Andrä, F. Andreata, F. Annunziato, E. Arranz, P. Bacher, S. Bari, V. Barnaba, J. Barros-Martins, D. Baumjohann, C. G. Beccaria, D. Bernardo, D. A. Boardman, J. Borger, C. Böttcher, L. Brockmann, M. Burns, D. H. Busch, G. Cameron, I. Cammarata, A. Cassotta, Y. Chang, F. G. Chirdo, E. Christakou, L. Čičin-Šain, L. Cook, A. J. Corbett, R. Cornelis, L. Cosmi, M. S. Davey, S. De Biasi, G. De Simone, G. del Zotto, M. Delacher, F. Di Rosa, J. Di Santo, A. Diefenbach, J. Dong, T. Dörner, R. J. Dress, C. A. Dutertre, S. B. G. Eckle, P. Eede, M. Evrard, C. S. Falk, M. Feuerer, S. Fillatreau, A. Fiz-Lopez, M. Follo, G. A. Foulds, J. Fröbel, N. Gagliani, G. Galletti, A. Gangaev, N. Garbi, J. A. Garrote, J. Geginat, N. A. Gherardin, L. Gibellini, F. Ginhoux, D. I. Godfrey, P. Gruarin, C. Haftmann, L. Hansmann, C. M. Harpur, A. C. Hayday, G. Heine, D. C. Hernández, M. Herrmann, O. Hoelsken, Q. Huang, S. Huber, J. E. Huber, J. Huehn, M. Hundemer, W. Y. K. Hwang, M. Iannacone, S. M. Ivison, H. M. Jäck, P. K. Jani, B. Keller, N. Kessler, S. Ketelaars, L. Knop, J. Knopf, H. F. Koay, K. Kobow, K. Kriegsmann, H. Kristyanto, A. Krueger, J. F. Kuehne, H. Kunze-Schumacher, P. Kvistborg, I. Kwok, D. Latorre, D. Lenz, M. K. Levings, A. C. Lino, F. Liotta, H. M. Long, E. Lugli, K. N. MacDonald, L. Maggi, M. K. Maini, F. Mair, C. Manta, R. A. Manz, M. F. Mashreghi, A. Mazzoni, J. McCluskey, H. E. Mei, F. Melchers, S. Melzer, D. Mielenz, L. Monin, L. Moretta, G. Multhoff, L. E. Muñoz, M. Muñoz-Ruiz, F. Muscate, A. Natalini, K. Neumann, L. G. Ng, A. Niedobitek, J. Niemz, L. N. Almeida, S. Notarbartolo, L. Ostendorf, L. J. Pallett, A. A. Patel, G. I. Percin, G. Peruzzi, M. Pinti, A. G. Pockley, K. Pracht, I. Prinz, I. Pujol-Autonell, N. Pulvirenti, L. Quatrini, K. M. Quinn, H. Radbruch, H. Rhys, M. B. Rodrigo, C. Romagnani, C. Saggau, S. Sakaguchi, F. Sallusto, L. Sanderink, I. Sandrock, C. Schauer, A. Scheffold, H. U. Scherer, M. Schiemann, F. A. Schildberg, K. Schober, J. Schoen, W. Schuh, T. Schüler, A. R. Schulz, S. Schulz, J. Schulze, S. Simonetti, J. Singh, K. M. Sitnik, R. Stark, S. Starossom, C. Stehle, F. Szelinski, L. Tan, A. Tarnok, J. Tornack, T. I. M. Tree, J. J. P. van Beek, W. van de Veen, K. van Gisbergen, C. Vasco, N. A. Verheyden, A. von Borstel, K. A. Ward-Hartstonge, K. Warnatz, C. Waskow, A. Wiedemann, A. Wilharm, J. Wing, O. Wirz, J. Wittner, J. H. M. Yang and J. Yang, Eur. J. Immunol., 2021, 51, 2708–3145 CrossRef CAS PubMed.
- E. D. Diebold, B. W. Buckley, D. R. Gossett and B. Jalali, Nat. Photonics, 2013, 7, 806–810 CrossRef CAS.
- H. Mikami, M. Kawaguchi, C. J. Huang, H. Matsumura, T. Sugimura, K. Huang, C. Lei, S. Ueno, T. Miura, T. Ito, K. Nagasawa, T. Maeno, H. Watarai, M. Yamagishi, S. Uemura, S. Ohnuki, Y. Ohya, H. Kurokawa, S. Matsusaka, C. W. Sun, Y. Ozeki and K. Goda, Nat. Commun., 2020, 11, 1162 CrossRef CAS PubMed.
- H. Mikami, J. Harmon, H. Kobayashi, S. Hamad, Y. Wang, O. Iwata, K. Suzuki, T. Ito, Y. Aisawa, N. Kutsuna, K. Nagasawa, H. Watarai, Y. Ozeki and K. Goda, Optica, 2018, 5, 117–126 CrossRef CAS.
- C. H. Chen, S. H. Cho, F. Tsai, A. Erten and Y. H. Lo, Biomed. Microdevices, 2009, 11, 1223–1231 CrossRef CAS PubMed.
- Y. J. Chiu, S. H. Cho, Z. Mei, V. Lien, T. F. Wu and Y. H. Lo, Lab Chip, 2013, 13, 1803–1809 Search PubMed.
- S. Sakuma, Y. Kasai, T. Hayakawa and F. Arai, Lab Chip, 2017, 17, 2760–2767 RSC.
- A. A. Nawaz, Y. Chen, N. Nama, R. H. Nissly, L. Ren, A. Ozcelik, L. Wang, J. P. McCoy, S. J. Levine and T. J. Huang, Anal. Chem., 2015, 87, 12051–12058 CrossRef CAS PubMed.
- D. Schraivogel and L. M. Steinmetz, Mol. Syst. Biol., 2023, 19, e11254 CrossRef PubMed.
- T. M. Kuhn, M. Paulsen and S. Cuylen-Haering, Trends Cell Biol., 2024, 34, 657–670 CrossRef CAS PubMed.
- J. Harmon, J. Findinier, N. T. Ishii, M. Herbig, A. Isozaki, A. Grossman and K. Goda, Cytometry, Part A, 2022, 101, 1027–1034 CrossRef CAS PubMed.
- H. Matsumura, L. T.-W. Shen, A. Isozaki, H. Mikami, D. Yuan, T. Miura, Y. Kondo, T. Mori, Y. Kusumoto, M. Nishikawa, A. Yasumoto, A. Ueda, H. Bando, H. Hara, Y. Liu, Y. Deng, M. Sonoshita, Y. Yatomi, K. Goda and S. Matsusaka, Lab Chip, 2023, 23, 1561–1575 RSC.
- B. M. Szczerba, F. Castro-Giner, M. Vetter, I. Krol, S. Gkountela, J. Landin, M. C. Scheidmann, C. Donato, R. Scherrer, J. Singer, C. Beisel, C. Kurzeder, V. Heinzelmann-Schwarz, C. Rochlitz, W. P. Weber, N. Beerenwinkel and N. Aceto, Nature, 2019, 566, 553–557 CrossRef CAS PubMed.
- N. Aceto, A. Bardia, D. T. Miyamoto, M. C. Donaldson, B. S. Wittner, J. A. Spencer, M. Yu, A. Pely, A. Engstrom, H. Zhu, B. W. Brannigan, R. Kapur, S. L. Stott, T. Shioda, S. Ramaswamy, D. T. Ting, C. P. Lin, M. Toner, D. A. Haber and S. Maheswaran, Cell, 2014, 158, 1110–1122 CrossRef CAS PubMed.
- A. Senrung, S. Lalwani, D. Janjua, T. Tripathi, J. Kaur, N. Ghuratia, N. Aggarwal, A. Chhokar, J. Yadav, A. Chaudhary, U. Joshi and A. C. Bharti, In Vitro Models, 2023, 2, 219–248 CrossRef CAS PubMed.
- Z. Zhao, X. Chen, A. M. Dowbaj, A. Sljukic, K. Bratlie, L. Lin, E. L. S. Fong, G. M. Balachander, Z. Chen, A. Soragni, M. Huch, Y. A. Zeng, Q. Wang and H. Yu, Nat. Rev. Methods Primers, 2022, 2, 94 CrossRef CAS PubMed.
- C. Lei, Y. Nakagawa, Y. Nagasaka, T. Ding, H. Kanno, C. Toyokawa, K. Niizuma, K. Suzuki, M. Li, A. Sunna, V. Hampl and K. Goda, TrAC, Trends Anal. Chem., 2024, 180, 117938 CrossRef CAS.
- J. de Rutte, R. Dimatteo, M. M. Archang, M. van Zee, D. Koo, S. Lee, A. C. Sharrow, P. J. Krohl, M. Mellody, S. Zhu, J. V. Eichenbaum, M. Kizerwetter, S. Udani, K. Ha, R. C. Willson, A. L. Bertozzi, J. B. Spangler, R. Damoiseaux and D. Di Carlo, ACS Nano, 2022, 16, 7242–7257 CrossRef CAS PubMed.
- C. Y. Wu, D. Stoecklein, A. Kommajosula, J. Lin, K. Owsley, B. Ganapathysubramanian and D. Di Carlo, Microsyst. Nanoeng., 2018, 4, 21 CrossRef PubMed.
- M. Hayashi, S. Ohnuki, Y. Tsai, N. Kondo, Y. Zhou, H. Zhang, N. T. Ishii, T. Ding, M. Herbig, A. Isozaki, Y. Ohya and K. Goda, Lab Chip, 2023, 23, 4232–4244 RSC.
- M. Herbig, A. Isozaki, D. Di Carlo, J. Guck, N. Nitta, R. Damoiseaux, S. Kamikawaji, E. Suyama, H. Shintaku, A. R. Wu, I. Nikaido and K. Goda, Nat. Methods, 2022, 19, 633–634 CrossRef CAS PubMed.
- D. Di Carlo, D. Irimia, R. G. Tompkins and M. Toner, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 18892–18897 CrossRef CAS PubMed.
- J. Zhou and I. Papautsky, Lab Chip, 2013, 13, 1121–1132 Search PubMed.
- H. Jeon, S. H. Lee, J. Shin, K. Song, N. Ahn and J. Park, Microsyst. Nanoeng., 2024, 10, 15 CrossRef CAS PubMed.
- E. J. Lim, T. J. Ober, J. F. Edd, S. P. Desai, D. Neal, K. W. Bong, P. S. Doyle, G. H. McKinley and M. Toner, Nat. Commun., 2014, 5, 4120 CrossRef CAS PubMed.
- J. Zhou and I. Papautsky, Microsyst. Nanoeng., 2020, 6, 113 CrossRef CAS PubMed.
- N. Xiang, Z. Ni and D. Wu, Sens. Diagn., 2023, 2, 929–937 Search PubMed.
- J. Nam, Y. Shin, J. K. S. Tan, Y. B. Lim, C. T. Lim and S. Kim, Lab Chip, 2016, 16, 2086–2092 Search PubMed.
- Y. Lecun, E. Bottou, Y. Bengio and P. Haffner, Proc. IEEE, 1998, 86, 2278–2324 CrossRef.
- Y. Kasai, C. Leipe, M. Saito, H. Kitagawa, S. Lauterbach, A. Brauer, P. E. Tarasov, T. Goslar, F. Arai and S. Sakuma, Sci. Adv., 2021, 7, abe7327 Search PubMed.
- W. L. Ung, K. Mutafopulos, P. Spink, R. W. Rambach, T. Franke and D. A. Weitz, Lab Chip, 2017, 17, 4059–4069 RSC.
- T. Iino, K. Okano, S. W. Lee, T. Yamakawa, H. Hagihara, Z. Y. Hong, T. Maeno, Y. Kasai, S. Sakuma, T. Hayakawa, F. Arai, Y. Ozeki, K. Goda and Y. Hosokawa, Lab Chip, 2019, 19, 2669–2677 Search PubMed.
- J. T. C. Liu, A. K. Glaser, K. Bera, L. D. True, N. P. Reder, K. W. Eliceiri and A. Madabhushi, Nat. Biomed. Eng., 2021, 5, 203–218 Search PubMed.
- A. Hubman, I. Plazl and T. Urbic, J. Mol. Liq., 2021, 328, 115410 CrossRef CAS.
- M. Li, H. E. Muñoz, K. Goda and D. Di Carlo, Sci. Rep., 2017, 7, 10802 CrossRef PubMed.
- A. Isozaki, Y. Nakagawa, M. H. Loo, Y. Shibata, N. Tanaka, D. L. Setyaningrum, J.-W. Park, Y. Shirasaki, H. Mikami, D. Huang, H. Tsoi, C. T. Riche, T. Ota, H. Miwa, Y. Kanda, T. Ito, K. Yamada, O. Iwata, K. Suzuki, S. Ohnuki, Y. Ohya, Y. Kato, T. Hasunuma, S. Matsusaka, M. Yamagishi, M. Yazawa, S. Uemura, K. Nagasawa, H. Watarai, D. Di Carlo and K. Goda, Sci. Adv., 2020, 6, aba6712 Search PubMed.
- M. Xu, J. Harmon, D. Yuan, S. Yan, C. Lei, K. Hiramatsu, Y. Zhou, M. H. Loo, T. Hasunuma, A. Isozaki and K. Goda, Environ. Sci. Technol., 2021, 55, 7880–7889 Search PubMed.
- H. E. Muñoz, M. Li, C. T. Riche, N. Nitta, E. Diebold, J. Lin, K. Owsley, M. Bahr, K. Goda and D. Di Carlo, Anal. Chem., 2018, 90, 11280–11289 CrossRef PubMed.
- E. C. Costa, A. F. Moreira, D. de Melo-Diogo, V. M. Gaspar, M. P. Carvalho and I. J. Correia, Biotechnol. Adv., 2016, 34, 1427–1441 Search PubMed.
- J. F. Dekkers, G. Berkers, E. Kruisselbrink, A. Vonk, H. R. De Jonge, H. M. Janssens, I. Bronsveld, E. A. Van De Graaf, E. E. S. Nieuwenhuis, R. H. J. Houwen, F. P. Vleggaar, J. C. Escher, Y. B. De Rijke, C. J. Majoor, H. G. M. Heijerman, K. M. De Winter-De Groot, H. Clevers, C. K. Van Der Ent and J. M. Beekman, Sci. Transl. Med., 2016, 8, 344a84 Search PubMed.
- D. Tuveson and H. Clevers, Science, 2019, 364, 952–955 CrossRef CAS PubMed.
- T. Boerstler, D. Kachkin, E. Gerasimova, N. Zagha, F. Furlanetto, N. Nayebzade, L. Zappia, M. Boisvert, M. Farrell, S. Ploetz, I. Prots, M. Regensburger, C. Günther, J. Winkler, P. Gupta, F. Theis, M. Karow, S. Falk, B. Winner and F. Krach, Commun. Biol., 2025, 8, 1412 CrossRef PubMed.
- K. B. Jensen and M. H. Little, Stem Cell Rep., 2023, 18, 1255–1270 Search PubMed.
- B. Phipson, P. X. Er, A. N. Combes, T. A. Forbes, S. E. Howden, L. Zappia, H. J. Yen, K. T. Lawlor, L. J. Hale, J. Sun, E. Wolvetang, M. Takasato, A. Oshlack and M. H. Little, Nat. Methods, 2019, 16, 79–87 CrossRef CAS PubMed.
- S. Jiang, H. Zhao, W. Zhang, J. Wang, Y. Liu, Y. Cao, H. Zheng, Z. Hu, S. Wang, Y. Zhu, W. Wang, S. Cui, P. E. Lobie, L. Huang and S. Ma, Cell Rep. Med., 2020, 1, 100161 CrossRef CAS PubMed.
- K. W. Seo, Y. J. Kang and S. J. Lee, Phys. Fluids, 2014, 26, 063301 CrossRef.
- S. Yang, J. Y. Kim, S. J. Lee, S. S. Lee and J. M. Kim, Lab Chip, 2011, 11, 266–273 Search PubMed.
- K. Yamada, H. Suzuki, T. Takeuchi, Y. Kazama, S. Mitra, T. Abe, K. Goda, K. Suzuki and O. Iwata, Sci. Rep., 2016, 6, 26327 CrossRef CAS PubMed.
- M. Kräter, S. Abuhattum, D. Soteriou, A. Jacobi, T. Krüger, J. Guck and M. Herbig, Adv. Sci., 2021, 8, 2003743 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |