Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
In vitro space of Disse model for exploration of drug induced hepatotoxicity
Ana
Mesic
 *a,
Antonietta
Messina
b,
Zoe
Tiprez
b,
Benoit
Charlot
*a,
Antonietta
Messina
b,
Zoe
Tiprez
b,
Benoit
Charlot
 c,
Safa Mohamed
Ismail
d,
Nicolas
Huang
d,
Sakina
Bensalem
a,
Jean-Charles
Duclos-Vallee
b and
Bruno
Le Pioufle
a
c,
Safa Mohamed
Ismail
d,
Nicolas
Huang
d,
Sakina
Bensalem
a,
Jean-Charles
Duclos-Vallee
b and
Bruno
Le Pioufle
a
aUniversity Paris Saclay, ENS Paris Saclay, CNRS LUMIN, France
bLaboratory of Physio-pathogenesis and Liver Disease Treatment, INSERM U1193, France
cIES, Institute of Electronics and Systems, CNRS University of Montpellier, France
dUniversité Paris-Saclay, CNRS, Institut Galien Paris-Saclay, Orsay, France
First published on 21st March 2026
Abstract
The limited predictive power of animal models remains a major bottleneck in drug development, particularly in assessing drug-induced liver injury (DILI). To address this, we developed a novel in vitro liver-on-a-chip platform focused on modelling the space of Disse (sD) – a key microenvironment, yet insufficiently studied, which mediates cross-talk between endothelial and hepatic systems. The system integrates a biocompatible sodium alginate hydrogel whose mechanical properties were optimised to mimic physiological liver stiffness, enabling molecular cross-talk between human liver sinusoidal endothelial cells (LSECs) and HepaRG hepatocytes without direct contact. Under dynamic perfusion, the co-culture showed viability and polarisation for 8 days, forming organised tissues with functional bile canaliculi. The presence of LSECs markedly enhanced hepatic performance, reflected by increased albumin and urea secretion and activation of pro-regenerative secretory pathways. Prolonged acetaminophen exposure demonstrated the model's capacity to reproduce hepatotoxic responses, confirming its predictive relevance. Such versatile microphysiological platform might become a powerful tool for studying endothelial–hepatic communication, modelling liver pathologies, and improving preclinical DILI testing.
Introduction
The efficiency of pharmaceutical research and development (R&D) has been in continuous decline for decades, with productivity estimated to have dropped nearly 80-fold since the 1950s when adjusted for inflation.1 Bringing a single drug to the market today requires on average $1.56 billion, equivalent to 6500–15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 person-years of work and about 10 to 12 years.2 Moreover, drug-induced liver injury (DILI) is a leading cause of late-stage failure and post-marketing withdrawals.3 A major driver of these inefficiencies is the limited predictive power of animal models used during preclinical testing. Despite showing apparent efficacy and safety in preclinical studies, 92% of candidate drugs ultimately fail during clinical trials.4 Beyond ethical concerns, animal testing is 1.5–30 times more expensive than in vitro methods and often poorly predictive of human physiology.5 Together, these limitations underscore the urgent need for advanced in vitro platforms that combine human relevance with efficiency.
000 person-years of work and about 10 to 12 years.2 Moreover, drug-induced liver injury (DILI) is a leading cause of late-stage failure and post-marketing withdrawals.3 A major driver of these inefficiencies is the limited predictive power of animal models used during preclinical testing. Despite showing apparent efficacy and safety in preclinical studies, 92% of candidate drugs ultimately fail during clinical trials.4 Beyond ethical concerns, animal testing is 1.5–30 times more expensive than in vitro methods and often poorly predictive of human physiology.5 Together, these limitations underscore the urgent need for advanced in vitro platforms that combine human relevance with efficiency.
Organ-on-a-chip technologies have emerged as a promising solution to the limitations of conventional preclinical models.6 These platforms replicate key physiological cues using a minimal set of cells, through several complementary strategies. In the case of the liver-on-a-chip, they enable the co-culture of multiple cell types, where hepatocytes maintained alongside endothelial cells display enhanced viability and functionality, reflected in increased albumin and bile secretion.7–9 Besides, the incorporation of biomimetic materials within the organ-on-a-chip structure permits the fine-tuning of extracellular matrix (ECM) properties. Indeed, hepatocytes respond through their gene expression and functionality to substrates of varying viscoelasticity, mirroring the pathological stiffening of the ECM during fibrosis.10 In addition, the application of dynamic perfusion introduces physiologically relevant shear stress, which has been shown to promote the differentiation of both hepatocytes and sinusoidal endothelial cells. Finally, micro compartments allow packing of hepatocytes at high cellular density, facilitating differentiation and the formation of bile canaliculi.11,12 Despite these advances, current liver-on-a-chip models fail to reproduce the space of Disse (sD) – the extracellular region bridging hepatocytes and liver sinusoidal endothelial cells – as they do not establish spatial separation between different cell types and/or replicating mechanical properties, such as elasticity or porosity.
The sD is a thin perisinusoidal region between liver sinusoidal endothelial cells (LSECs) and the basal membrane of hepatocytes, typically only 500–1000 nm thick.13,14 Its essential function is the maintenance of liver homeostasis through exchange of nutrients, metabolites and signalling molecules, between sinusoid and hepatocytes. Additionally, it is a niche for mesenchymal stem cells involved in liver regeneration,15 hepatic stellate cells responsible for the formation of scar tissue during fibrosis16,17 and target localisation for activated Kupffer cells during inflammatory responses.18 Being such a complex microenvironment, it is highly sensitive to pathological remodelling. In pathologies such as liver fibrosis, cirrhosis, or DILI, the sD goes through multiple cellular, structural and biochemical changes: hepatocytes experience loss of polarity, reducing their ability for molecule intake,19 LSECs undergo defenestration,20,21 stellate cells become activated, proliferate, and deposit excessive ECM; Kupffer cells secrete proinflammatory cytokines; the composition and mechanical properties of the ECM significantly change.22
Despite its physiological and pathological importance, modelling the sD in vitro remains extremely challenging. Its nanometre-scale width requires advanced microfabrication techniques for spatial separation of multiple cell types, while maintaining correct biochemical cues and functional interactions. Furthermore, replicating a continuous plasma flow, cellular diversity, and ECM complexity adds layers of technical difficulty. Currently, there are three main strategies employed in in vitro sD models: use of microfluidic channels, porous membranes or hydrogels:
1. Microfluidic channels allow dynamic perfusion, chemical gradient formation, and high-density hepatocyte culture, promoting bile canaliculi formation and functional alignment; however, direct exposure to shear stress and limited multicellular complexity remain challenges.23,24
2. Membrane-based systems enable the co-culture of hepatocytes and endothelial cells on opposite sides of a semipermeable barrier, allowing differential media, controlled molecular exchange, and partial replication of the sD width (1–10 μm), yet ECM properties tunability is limited.25,26
3. Hydrogel-based approaches provide a tuneable 3D extracellular matrix that supports hepatocyte aggregation, spheroid formation, and endothelial network integration; when combined with microfluidics, hydrogels allow perfusion and shear stress, enhancing physiological relevance and enabling vascularised hepatic constructs.27–29
Here, we report on the development of an in vitro sD model on a microfluidic chip with an integrated biocompatible hydrogel possessing tuneable mechanical properties. The platform integrates two cell types: hepatocytes and endothelial cells, spatially separated by a sodium alginate hydrogel barrier that reproduces the mechanical properties of the extracellular matrix while enabling molecular cross-talk. This architecture allows physiological interactions without direct cell contact, providing a more faithful representation of the in vivo sinusoidal microenvironment. Furthermore, we demonstrate the advantages of this system in comparison to single cell liver-on-a-chip models, and its utility in hepatotoxicity assays using 72-hour acetaminophen exposure. This article introduces a physiologically relevant sD model that addresses a major unmet need in liver tissue engineering and toxicology testing.
Materials and methods
Design and fabrication of the microfluidic device
Microfluidic chip designs were created using L-Edit IC (Siemens EDA Solutions) software. Silicon master-mould was fabricated by standard photolithography. SU-8 2100 photoresist (Kayaku Advanced Materials, USA) was spin-coated onto silicon wafers at 3000 rpm for 24 s, exposed for 24 s to 365 nm UV light through a chrome mask (Toppan™) and hard baked for 5 min at 95 °C. Uncross linked resin was removed by immersion in SU-8 Developer for 4 min. SU-8 pattern height and consistency was measured with Dektak 150 Surface Profiler (Veeco, USA).PDMS (polydimethylsiloxane, Sylgard 184, Dow Corning) was mixed in a 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 base-to-curing agent ratio, degassed under vacuum for 1 h, poured over the moulds, and cured at 90 °C for 2 h. The cured PDMS replicas were peeled, cut, and punched to create inlets and outlets (1.25 mm for side channels, 0.5 mm for the middle channel). PDMS layers and glass slides were plasma-activated (Diener Atto, 50% power, 6 s, 6 × 104 Pa O2) and bonded. Devices were finalised by a 10 min post-bonding bake at 150 °C.
1 base-to-curing agent ratio, degassed under vacuum for 1 h, poured over the moulds, and cured at 90 °C for 2 h. The cured PDMS replicas were peeled, cut, and punched to create inlets and outlets (1.25 mm for side channels, 0.5 mm for the middle channel). PDMS layers and glass slides were plasma-activated (Diener Atto, 50% power, 6 s, 6 × 104 Pa O2) and bonded. Devices were finalised by a 10 min post-bonding bake at 150 °C.
Hydrogel loading and characterisation
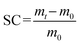 | (1) |
| G* = G′ + iG″ | (2) |
| E = 2(1 + ν)G* | (3) |
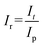 | (4) |
Cell culture on a chip
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2.
2.
Biological characterisation of the model
Statistical and image analysis
All images were analysed in the ImageJ (Fiji) 1.54f software33 (more details in the SI). All variables were expressed as means ± standard deviations. Statistical significance between groups was evaluated by a Student's t-test. P-Value inferior than 0.05 was regarded as statistically significant (p-value notations: * < 0.05, ** < 0.01, *** < 0.001, **** < 0.0001).Results
Microfluidic chip design
The microfluidic chip was designed to replicate the spatial organisation of sinusoidal cells, hepatocytes and the sD in the liver sinusoid, based on previously published work.34,35 The device consists of three parallel channels separated by two arrays of 34 polygonal pillars. This chip geometry allows mimicry a layout of the endothelial capillary, hepatocytes and space of Disse in a liver sinusoid (Fig. 1A and B). Hydrogel loaded in the middle channel mechanically mimics the ECM in the sD, while each cell type cultured in one side channel. This enables compartmentalised culture of two cell types through the mediation of a controlled molecular exchange across the middle channel. All channels have a height of 100 μm. The central effective culturing area of the chip is 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 μm long and 2000 μm wide (Fig. 1C and D). The hydrogel channel has a width of 1000 μm, while the flanking side channels measure 500 μm each. Within the arrays, PDMS pillars are spaced 15 μm, creating a stable retention zone for the hydrogel (Fig. 1E). Models of fluid flow in the chip were developed using finite element simulation (SI).
000 μm long and 2000 μm wide (Fig. 1C and D). The hydrogel channel has a width of 1000 μm, while the flanking side channels measure 500 μm each. Within the arrays, PDMS pillars are spaced 15 μm, creating a stable retention zone for the hydrogel (Fig. 1E). Models of fluid flow in the chip were developed using finite element simulation (SI).
Hydrogel characterisation
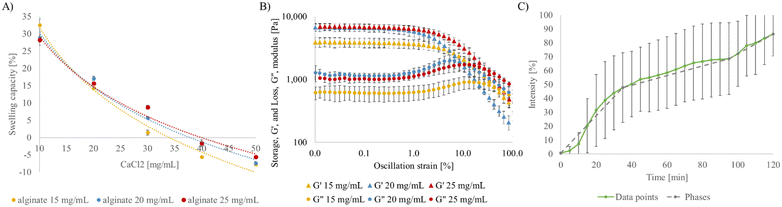
Sodium alginate is a hydrogel widely used in tissue engineering. It was chosen to mimic the sD area on the chip. It is chemically inert, derived from non-animal sources, and offers tuneable mechanical properties. Liquid sodium alginate resin was integrated in the middle channel where PDMS pillars allowed precise loading and retention. Crosslinking with CaCl2 through the side channels resulted in the formation of a stable, rigid matrix. The hydrogel remained intact after cell loading in the side channels and 8 days of culture at 37 °C, 5% CO2 saturation and 95% humidity conditions.This combination of biocompatibility, structural robustness and controlled permeability encouraged us to choose the sodium alginate as hydrogel for subsequent co-culture experiments. The swelling of sodium alginate hydrogels was systematically evaluated across varying crosslinker CaCl2 concentrations to identify the formulation that maintains structural integrity of the hydrogel barrier. An optimal CaCl2 concentration of 30 mg mL−1 was selected, yielding a controlled swelling range of 1.4–7.8%, which ensured that the hydrogel remained stable without swelling in the adjacent channels (Fig. 2A). Rheological testing revealed that increasing alginate concentration enhanced gel stiffness. Young's modulus values varied from 12 ± 3 kPa at 15 mg mL−1 to 21 ± 3 kPa at 20–25 mg mL−1 (crosslinked with CaCl2 30 mg mL−1) (Fig. 2B). The 15 mg mL−1 formulation corresponded most closely to healthy liver tissue, while higher concentrations resembled the stiffer profiles of fibrotic or pathologically remodelled regions.36 Transport of the small fluorescent molecule across sodium alginate barrier was examined by perfusion on the system with CDFA solution. CDFA molecule was selected for its fluorescent properties, spherical structure and relatively small molecular mass comparable to glucose (445.21 Da). On-chip assays showed that 80% of total saturation was reached across the hydrogel barrier after 110 min. Molecular transport data points show three distinct phases in the transport kinetics marked with a grey dashed line on the Fig. 2C: initial diffusion-dominated transport (0–35 min), a lag phase (35–95 min) due to barrier resistance, and a new steady-state (95–120 min) after overcoming initial resistance. Slopes of these phases are 1.18, 0.3 and 0.71 respectively.
Endothelial tube characterisation
LSEC cells were loaded in one of the side channels so that they attach to all channel walls. They were subsequently cultured in direct exposure to the shear stress induced by medium perfusion. Immunofluorescent staining of specific markers was assessed after 8 days of dynamic culture (1 μL min−1). F-actin staining revealed cytoskeletal alignment along the axis of medium flow. Quantitative analysis of angular fibre distribution confirmed a significant orientation of LSEC cells in the direction of shear stress. As demonstrated in Fig. 3A, the majority of cytoskeletal vectors aligned with the flow direction, with 45.4% of angles distributed within the −30° to +30° and 12.7% within the −5° to +5° range relative to the channel axis (0° corresponding to medium flow).To evaluate tissue identity and barrier integrity, immunofluorescent staining was performed for key endothelial markers. ZO1 and VE-cadherin (CD144) were localised at cell junctions, indicating the establishment of tight junctions essential for the endothelial barrier function. PECAM-1 (CD31) was strongly expressed on the cell membrane confirming differentiation status of the tissue and endothelial-specific adhesion properties (Fig. 3C). Together, these markers confirmed both the differentiation and cohesion of the endothelial sheet along the full length of the channel (full channel immunofluorescent staining in the SI). Three-dimensional reconstruction from z-stack images further demonstrated the formation of a continuous, vessel-like structure. Endothelial cells lined the bottom, top, and vertical walls of the channel, enclosing a well-defined lumen in the middle of the channel (Fig. 3B). This endothelial distribution was confirmed along the full length of the channel.
Hepatic tissue characterisation
HepaRG cells were seeded in the hepatic channel of the microfluidic chip and cultured under dynamic perfusion (1 μL min−1) for 8 days. To characterise the developed tissue, cellular quantification, viability assessment and immunofluorescence staining were performed. Quantification of DNA, RNA, and protein content after cellular extraction indicated an average of 35 ng of DNA, 485 ng of RNA, and 4.37 μg of protein per chip, corresponding to approximately 6000 hepatocytes per device.37 Live-dead staining revealed high viability of the tissue with 94 ± 5% of live cells (chips n = 3, SI). Immunofluorescent staining confirmed successful differentiation and polarisation of the hepatic tissue. Nuclear expression of HNF4α, a key transcription factor for hepatocyte development, confirmed differentiation into functional hepatocytes. Albumin, a hallmark secreted protein of hepatocytes, was detected in the cytoplasm surrounding the nuclei, highlighting active synthetic function. Further evidence of functional polarisation and bile canaliculi formation was provided by membrane expression of BSEP, a bile acid transporter, and MRP2, an efflux pump for organic anions and bilirubin (Fig. 4 and S4). | ||
| Fig. 4 Immunofluorescent staining for hepatocyte specific markers of HepaRG cells on the chip. Blue – DAPI, magenta – HNF4α, red – albumin, yellow – BSEP, cyan – MRP2 (scale bar = 100 μm). | ||
Impact of LSEC–HepaRG co-culture on hepatic expression
LSECs have been shown to positively influence hepatocyte function, enhancing the secretion of functional markers such as albumin and urea, while at the same time improving resistance to toxicants.8,9,38 To investigate these effects in our model, HepaRG cells were cultured either as monocultures or in co-culture with LSECs. Hepatic functionality was then evaluated after the culture period based on three approaches: quantification of key liver biomarkers, characterisation of hepatocyte polarisation, and probe-based assessment of protein expression.Liver biomarkers, albumin and urea, were evaluated through ELISA quantification from the medium perfused through the chip. Albumin secretion remained stable in both monoculture and co-culture conditions, but levels were consistently higher in the LSEC–HepaRG co-culture. Indeed, average albumin production reached 1248 ± 214 ng per chip in co-culture compared with 392 ± 168 ng in monoculture (p < 0.0001). Urea secretion followed the same trend, with monoculture chips secretion at 7.84 × 10−2 ± 3.42 × 10−4 nmol compared with 1.69 × 10−1 ± 1.83 × 10−3 nmol in co-culture chips. These values indicate more than a twofold increase in urea production consistent throughout the whole culture duration (Fig. 5A and B). These results demonstrate a consistent enhancement of hepatocyte metabolic activity in the presence of LSECs. Bile canaliculi formation was assessed using the live fluorescent probe Fluobile™, which marks functional canalicular networks in hepatocytes. After 8 days of culture, quantification revealed a significantly higher number and a larger total fluorescent area of bile canaliculi in LSEC–HepaRG co-culture compared to monoculture (Fig. 5C). High-magnification imaging further showed that canaliculi formation preferentially occurred in regions of increased cellular density and within 3D structures, often extending along the membranes of adjacent hepatocytes (Fig. 5D). These observations indicate that endothelial co-culture promotes hepatocyte polarisation and the organisation of functional bile canaliculi.
To evaluate the impact of the co-culture on the hepatic secretome, proteomic profiling was performed using the Olink Target 96 Inflammation panel and analysed on log2-transformed NPX values.
Hierarchical clustering of z-scored NPX values (Fig. 6A) revealed a clear separation between HepaRG monocultures and LSEC–HepaRG co-cultures: co-culture samples clustered together and showed a broad, coordinated increase in signal intensity across the pre-selected analytes (selection criterion: p < 0.05; full NPX table in the SI).
The PCA loading plot identified the analytes that most strongly contributed to sample separation. The cluster of proteins positioned on the far right of the plot – including VEGFA, CD40, MMP-1, and CSF-1 – were the major drivers along PC1, reflecting the distinction between monoculture and co-culture conditions. In contrast, proteins in the cluster located on the upper left of the plot, predominantly DNER, MCP-4, and IL-5, accounted for within-group variability along PC2, representing secondary axes of separation. Finally, principal component analysis showed robust separation of conditions along PC1 (≈70% variance explained), with co-culture samples projecting positively on PC1 and monocultures projecting negatively (Fig. 6C). Together, these results indicate that inclusion of LSECs substantially remodelled the hepatic secretome, yielding a reproducible protein signature dominated by growth/regenerative and inflammatory/remodelling signals.
Toxicologic assay
Due to enhanced performance of the model in terms of hepatic function and polarity, the model containing LSEC–HepaRG co-culture was chosen for hepatotoxicity assay. Hepatic viability and metabolic activity were assessed under prolonged 2 mM acetaminophen (APAP) treatment by live mitochondrial and CYP3A4 probe staining. Image analysis of fluorescent MitoTracker staining images, revealed a pronounced reduction in mitochondrial activity following APAP exposure, with APAP-treated samples showing a 77.2 ± 5.6% decrease in live mitochondrial signal compared to controls (Fig. 7A). This indicates severe impairment of mitochondrial function, consistent with oxidative stress-induced toxicity. CYP3A4 activity was also significantly affected by APAP exposure. In control tissues, CYP3A4 signal covered 9.71 ± 4.71% of the culture area, while in APAP-treated samples it increased to 22.81 ± 4.93% (Fig. 7B), reflecting a stress-induced alteration of drug metabolism pathways. Notably, the APAP-treated hepatic tissue displayed uneven signal distribution, fragmentation, and localised shrinkage, consistent with hepatocyte damage and detachment. These morphological alterations were consistent with hallmark features of APAP-induced hepatocyte injury reported in previous studies.39 | ||
| Fig. 7 Markers of hepatic viability and function in toxicity conditions. A) Live mitochondria (MitoTracker staining, *p < 0.05) and B) CYP3A4 expression (P450 Glo CYP3A4, ****p < 0.0001). | ||
Discussion
This study presents the development of a microfluidic chip used for biomimetic reproduction of the architecture and cellular interactions in a sD. The platform integrated a sodium alginate hydrogel with easily tuneable mechanical properties as an ECM analogue, enabling compartmentalised dynamic co-culture of HepaRG and LSECs. HepaRG cells were intentionally selected as a robust and well-characterised hepatic model that offers a practical compromise between physiological relevance and experimental reproducibility for proof-of-concept microphysiological systems. Their use enabled stable long-term co-culture under dynamic conditions while minimising donor-to-donor variability and experimental complexity associated with primary human hepatocytes. The system maintained high viability and structural integrity of both tissues for at least 8 days of dynamic culture, supporting stable hepatic function and endothelial organisation. Together, these results showed the establishment of a physiological model capable of sustaining endothelial–hepatic interactions in a controlled, physiologically relevant environment.The microfluidic design of the device directly influenced its functionality. The three parallel channel configuration allowed spatial separation of cell types while allowing diffusional exchange through a hydrogel-filled central compartment that mimics native sD. Although the absolute width of the hydrogel channel (1000 μm) exceeds the in vivo thickness of the sD (approximately 1 μm), this model reproduced its physiological role as a mechanical and biochemical communication interface, while preserving structural hierarchy of the liver sinusoid. Hepatocytes and endothelial cells were allowed intercellular communication while in indirect contact. Furthermore, the design allowed a special set of conditions, such as shear stress exposure, be applied to LSEC cells, thus closely mimicking in vivo situation. Current microfluidic models replicate the sD by integrating microchannels called slits, scaffolds or membranes.25,40 These structures often fail to reproduce its essential mechanical characteristics, such as physiological elasticity, correct shear stress exposure and controlled porosity. The use of PDMS pillars in our chip ensured mechanical retention of the hydrogel and sustained perfusion without noticeable effect on the gel positioning within the chip. To do so, the microchannel dimensions were optimised to maintain stable hydrogel confinement and laminar flow across physiologic shear ranges.
Hydrogel optimisation was as important for the model's biomimetic performance as was the microfluidic chip design. In the majority of liver-on-a-chip models in the literature, the hydrogel of choice was typically collagen at high concentrations. These models also rely on direct embedding of hepatocytes within collagen, which often causes fibrotic phenotype due to biochemical cues of excessive ECM.41 In contrast, the alginate hydrogel used here was chosen for its chemical inertness, non-animal origin, and mechanical tunability. Its porous structure enabled passive diffusion of signalling molecules while avoiding active biochemical interaction with hepatocyte or endothelial-secreted molecules, thereby preserving native intercellular communication. Crosslinking parameters were optimised to yield a Young's modulus of approximately 12 kPa, corresponding to the physiological stiffness of healthy human liver tissue.36 Thereby, the non-physiological stiffening associated with fibrosis was avoided, and this allowed mechanical but not biochemical signalling from the matrix to drive cell behaviour. The hydrogel also demonstrated structural robustness and controlled permeability, with the approximate porosity of 30% estimated from tracking small-molecule transport across the barrier, thus closely resembling diffusional dynamics expected across the native sD.
Comprehensive characterisation of both endothelial and hepatic tissues confirmed successful tissue-specific organisation and functionality. Under continuous flow, LSECs aligned their actin cytoskeleton along the direction of shear stress, indicating flow-induced polarisation. While a uniform distribution of orientations would yield approximately 33.4% of vectors within −30° to +30° and 5.6% within −5° to +5°, our data showed 45.4% and 12.7%, respectively (p = 0.001), demonstrating a significant enrichment of alignment along the flow axis. Immunofluorescent staining for endothelial junctional markers ZO1, VE-cadherin, and PECAM-1 demonstrated the establishment of a cohesive and differentiated endothelial layer forming a tube with a lumen. The HepaRG tissue exhibited high viability (94 ± 5%) and strong expression of hepatocyte-specific markers, including HNF4α and albumin, confirming hepatic differentiation. The presence of apical transporters (BSEP, MRP2) indicated polarisation and bile canaliculi formation. Together, these observations demonstrated that the microfluidic environment and matrix mechanics supported physiologically relevant cell organisation and function.
Beyond single-tissue characterisation, the co-culturing of HepaRG cells with LSECs substantially enhanced hepatic performance. Co-culture resulted in a threefold increase in albumin secretion and a twofold elevation in urea synthesis compared to monocultures, consistent with previous reports of endothelial-mediated support of hepatocyte function. The presence of LSECs also promoted hepatocyte polarisation, reflected by a higher number and surface area of functional bile canaliculi, which is consistent with earlier studies.8,9,38 It is likely that the enhanced hepatic functionality in the LSEC–HepaRG co-culture arises from multiple complementary mechanisms, including paracrine secretion of hepatocyte-supportive growth factors and cytokines, improved nutrient and waste exchange mediated by the endothelial layer, and direct intercellular communication through physical and biochemical signalling pathways that collectively promote polarisation and metabolic stability. Secretome profiling further revealed that co-culture triggered a coordinated remodelling of the molecular microenvironment. Biological interpretation of the analytes showed that expressed proteins were grouped into six groups based on cellular function (Table 1). Principal component and hierarchical clustering analyses showed clear separation between monoculture and co-culture samples, driven by distinct groups of proteins associated with angiogenesis, immune modulation, and extracellular matrix remodelling. Upregulation of factors such as VEGFA, HGF, SCF, and Flt3L likely contributed to enhanced hepatocyte viability and metabolic output, while increased secretion of cytokines including IL-6, IL-8, and CXCL1 suggested activation of regenerative signalling cascades. Concurrent elevation of MMP-1, MMP-10, and LAP TGF-β1 indicates ongoing ECM remodelling and possible low-grade activation of wound-healing pathways.42 In conclusion, our secretome screening revealed activation of both pro-regenerative and pro-remodelling components, maintained through endothelial–hepatocyte crosstalk. Further application of this analytical approach should enable the identification of LSEC secreted biomarkers to determine the key signalling molecular triggers in this intercellular communication.
| Cellular functions | Analytes |
|---|---|
| Angiogenesis and hepatocyte survival | VEGFA, HGF, SCF, Flt3L |
| Inflammatory and immune response | IL-6, IL-8, IL-10RA, CXCL1, CCL23, MCP-4, CSF-1, PD-L1, IL-18R1 |
| ECM remodelling and fibrosis | MMP-1, MMP-10, LAP TGF-β1 |
| Hepatic regeneration factors | FGF-5, FGF-19, FGF-21, LIF, LIF-R |
| Apoptosis and survival regulation | TRAIL, CASP-8 |
| Metabolic/immune modulators | IL-33, ADA, NT-3 |
To demonstrate the functional relevance of the system for drug testing, a prolonged hepatotoxicity assay using acetaminophen was performed. APAP was selected as a reference compound due to its established role as the gold standard in hepatotoxicity screening, with extensively characterised molecular mechanisms. Following 2 mM APAP exposure from day 5 to day 8, MitoTracker staining revealed a 77% reduction in live mitochondrial signal, directly indicating reduction in cell viability. This observation aligns with the known mechanism of APAP toxicity mediated by oxidative stress-induced mitochondrial dysfunction through the reactive metabolite NAPQI, which depletes intracellular glutathione and disrupts mitochondrial respiration.43 CYP3A4 activity, assessed via live luminescent assay, also decreased markedly after treatment, indicating impaired hepatic metabolism. Morphological assessment confirmed tissue shrinkage and detachment; features typical of hepatocyte death. Together, these results validate the capacity of the LSEC–HepaRG co-culture model to detect functional and metabolic hallmarks of drug-induced hepatotoxicity under 72-hour exposure conditions, thus serving as a proof of concept for DILI modelling.
Overall, this study introduces a liver-on-a-chip model that combines structural, mechanical, and biochemical fidelity to the native hepatic microenvironment. The integration of a tuneable alginate hydrogel enables reliable mimicking of sD mechanical properties without fibrotic phenotype induction. The model reproduces key structural and functional characteristics of the liver sinusoid, including endothelial cells organised in a vessel-like structure, hepatocyte polarisation, bile canaliculi formation, and responsiveness to drug induced toxicity. While APAP was used here for model validation, future investigation will focus on screening additional drug classes with distinct hepatotoxic mechanisms.
Conclusion
We have developed and characterised a novel in vitro liver-on-a-chip platform focused on modelling of the sD. The system integrates a biocompatible sodium alginate hydrogel whose mechanical properties are optimised to closely mimic those of physiological human liver tissue. The model successfully maintains the viability and functionality of both LSECs and HepaRG cells under dynamic co-culture conditions for 8 days, supporting tissue organisation and functional polarisation. Co-culturing of HepaRG with LSECs significantly enhances hepatic performance, leading to increased albumin and urea secretion, improved bile canaliculi formation, and secretion of pro-regenerative and remodelling factors. The predictive capacity of the model is further confirmed through acetaminophen exposure. Future perspectives will focus on the development of the model in two main directions. First, addition of Kupffer and stellate cells, which would increase the predictive power and reliability of the platform in conditions such as chronic inflammation. Second, screening a library of multiple drug classes and characterising the model's response in reaction to them. Altogether, this work establishes a robust and human-relevant microphysiological system for studying endothelial–hepatic interactions and drug-induced liver injury.Author contributions
Conceptualisation, A. M. and A. M.; methodology, A. M., A. M., Z. T. and S. M. I.; validation, A. M.; formal analysis, A. M. and S. M. I.; investigation, A. M.; resources, A. M.; data curation, A. M.; software, A. M.; writing – original draft preparation, A. M.; writing – review and editing, A. M., A. M., S. B., J. C. D. V. and B. L. P.; visualisation, A. M.; supervision, A. M., S. B., J. C. D. V. and B. L. P.; funding acquisition, J. C. D. V. and B. L. P.; project administration, A. M., S. B., J. C. D. V. and B. L. P. All authors have read and agreed to the published version of the manuscript.Conflicts of interest
There are no conflicts of interest to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI): a detailed protocol for biological characterisation and image analysis; immunofluorescent images for LSEC alignment quantification, images of hepatic tissue viability staining, NPX values table of secretome.Supplementary information is available. See DOI: https://doi.org/10.1039/d5lc01139f.
Acknowledgements
This work was supported by ENS Paris Saclay, INSERM U1193 and University Paris Saclay, Faculty of Pharmacy and ANR-21-CE-19-0025 DILI-on-chip project. AM's PhD fellowship and AM's researcher position were funded by ANR (Agence Nationale de Recherche) program DILI-on-chip through a project ANR-21-CE-19-0025.References
- J. W. Scannell, A. Blanckley, H. Boldon and B. Warrington, Diagnosing the decline in pharmaceutical R&D efficiency, Nat. Rev. Drug Discovery, 2012, 11, 191–200 CrossRef CAS PubMed.
- O. J. Wouters, M. McKee and J. Luyten, Estimated Research and Development Investment Needed to Bring a New Medicine to Market, 2009-2018, JAMA, J. Am. Med. Assoc., 2020, 323, 844–853 CrossRef PubMed.
- I. J. Onakpoya, C. J. Heneghan and J. K. Aronson, Post-marketing withdrawal of 462 medicinal products because of adverse drug reactions: a systematic review of the world literature, BMC Med., 2016, 14, 10 CrossRef PubMed.
- J. Arrowsmith and P. Miller, Phase II and phase III attrition rates 2011-2012, Nat. Rev. Drug Discovery, 2013, 12, 569–570 CrossRef CAS PubMed.
- D. Taylor, The Pharmaceutical Industry and the Future of Drug Development, Royal Society of Chemistry, 2016 Search PubMed.
- L. Ewart, et al., Performance assessment and economic analysis of a human Liver-Chip for predictive toxicology, Commun. Med., 2022, 2, 154 CrossRef PubMed.
- C. Ma, et al., On-Chip Construction of Liver Lobule-like Microtissue and Its Application for Adverse Drug Reaction Assay, Anal. Chem., 2016, 88, 1719–1727 CrossRef CAS PubMed.
- Y. B. Kang, S. Rawat, J. Cirillo, M. Bouchard and H. Noh, Layered long-term co-culture of hepatocytes and endothelial cells on a transwell membrane: toward engineering the liver sinusoid, Biofabrication, 2013, 5, 045008 CrossRef CAS PubMed.
- H. Lee and D.-W. Cho, One-step fabrication of an organ-on-a-chip with spatial heterogeneity using a 3D bioprinting technology, Lab Chip, 2016, 16, 2618–2625 RSC.
- Z. You, L. Zhou, W. Li, C. Huang and Y. Du, Mechanical microenvironment as a key cellular regulator in the liver, Acta Mech. Sin., 2019, 35, 289–298 CrossRef.
- P. J. Lee, P. J. Hung and L. P. Lee, An artificial liver sinusoid with a microfluidic endothelial-like barrier for primary hepatocyte culture, Biotechnol. Bioeng., 2007, 97, 1340–1346 CrossRef CAS PubMed.
- Y. Nakao, H. Kimura, Y. Sakai and T. Fujii, Bile canaliculi formation by aligning rat primary hepatocytes in a microfluidic device, Biomicrofluidics, 2011, 5, 022212 CrossRef PubMed.
- P. Nagy, S. S. Thorgeirsson and J. W. Grisham, et al., Organizational Principles of the Liver, in The Liver, ed. I. M. Arias, Wiley, 2020, pp. 1–13 Search PubMed.
- The Liver: Biology and Pathobiology, ed. I. M. Arias, H. J. Alter, J. L. Boyer, D. E. Cohen, D. A. Shafritz, S. S. Thorgeirsson and A. W. Wolkoff, Wiley, 2020 Search PubMed.
- D. Häussinger and C. Kordes, Space of Disse: a stem cell niche in the liver, Biol. Chem., 2019, 401, 81–95 CrossRef PubMed.
- H. Senoo, et al., Hepatic stellate cell (vitamin A-storing cell) and its relative – past, present and future, Cell Biol. Int., 2010, 34, 1247–1272 CrossRef CAS PubMed.
- K. Wake, Hepatic stellate cells: Three-dimensional structure, localization, heterogeneity and development, Proc. Jpn. Acad., Ser. B, 2006, 82, 155–164 CrossRef CAS PubMed.
- A.-A. La, A. Ma, A. J and J. Mérida-Velasco Jr, Disse and his Space, Indian Journal of Anatomy, 2018, 7(1), 99–102 CrossRef.
- C. Sanz-García, et al., The Space of Disse: The Liver Hub in Health and Disease, Livers, 2021, 1, 3–26 CrossRef.
- T. Mori, et al., Defenestration of the sinusoidal endothelial cell in a rat model of cirrhosis, Hepatology, 1993, 17, 891–897 CrossRef CAS PubMed.
- L. DeLeve, Hepatic Microvasculature in Liver Injury, Semin. Liver Dis., 2007, 27, 390–400 CrossRef CAS PubMed.
- P. Bedossa and V. Paradis, Liver extracellular matrix in health and disease: Liver extracellular matrix, J. Pathol., 2003, 200, 504–515 CrossRef PubMed.
- W. Li, et al., Direct mechanical exposure initiates hepatocyte proliferation, JHEP Rep., 2023, 5, 100905 CrossRef PubMed.
- Y. B. Kang, J. Eo, B. Bulutoglu, M. L. Yarmush and O. B. Usta, Progressive hypoxia-on-a-chip: An in vitro oxygen gradient model for capturing the effects of hypoxia on primary hepatocytes in health and disease, Biotechnol. Bioeng., 2020, 117, 763–775 CrossRef CAS PubMed.
- Y. Du, et al., Mimicking liver sinusoidal structures and functions using a 3D-configured microfluidic chip, Lab Chip, 2017, 17, 782–794 RSC.
- J. Kim, et al., Three-Dimensional Human Liver-Chip Emulating Premetastatic Niche Formation by Breast Cancer-Derived Extracellular Vesicles, ACS Nano, 2020, 14, 14971–14988 Search PubMed.
- M. Jang, P. Neuzil, T. Volk, A. Manz and A. Kleber, On-chip three-dimensional cell culture in phaseguides improves hepatocyte functions in vitro, Biomicrofluidics, 2015, 9, 034113 CrossRef PubMed.
- J. Liu, C. Feng, M. Zhang, F. Song and H. Liu, Design and Fabrication of a Liver-on-a-chip Reconstructing Tissue-tissue Interfaces, Front. Oncol., 2022, 12, 959299 CrossRef CAS PubMed.
- F. Bonanini, et al., In vitro grafting of hepatic spheroids and organoids on a microfluidic vascular bed, Angiogenesis, 2022, 25, 455–470 CrossRef CAS PubMed.
- A. Pourjavadi, H. Ghasemzadeh and R. Soleyman, Synthesis, characterization, and swelling behavior of alginate-g-poly(sodium acrylate)/kaolin superabsorbent hydrogel composites, J. Appl. Polym. Sci., 2007, 105, 2631–2639 CrossRef CAS.
- E. Ferrari, et al., A method to generate perfusable physiologic-like vascular channels within a liver-on-chip model, Biomicrofluidics, 2023, 17, 064103 CrossRef CAS PubMed.
- M. Boul, et al., A versatile microfluidic tool for the 3D culture of HepaRG cells seeded at various stages of differentiation, Sci. Rep., 2021, 11, 14075 CrossRef CAS PubMed.
- J. Schindelin, et al., Fiji: an open-source platform for biological-image analysis, Nat. Methods, 2012, 9, 676–682 CrossRef CAS PubMed.
- I. K. Zervantonakis, et al., Three-dimensional microfluidic model for tumor cell intravasation and endothelial barrier function, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 13515–13520 CrossRef PubMed.
- M. B. Chen, et al., On-chip human microvasculature assay for visualization and quantification of tumor cell extravasation dynamics, Nat. Protoc., 2017, 12, 865–880 CrossRef CAS PubMed.
- A. Karimi and A. Shojaei, An Experimental Study to Measure the Mechanical Properties of the Human Liver, Dig. Dis., 2018, 36, 150–155 CrossRef PubMed.
- C. Brun, et al., Extended lifespan and improved genome stability in HepaRG-derived cell lines through reprogramming by high-density stress, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2219298120 CrossRef CAS PubMed.
- M. Ortega-Ribera, et al., Resemblance of the human liver sinusoid in a fluidic device with biomedical and pharmaceutical applications, Biotechnol. Bioeng., 2018, 115, 2585–2594 CrossRef CAS PubMed.
- M. M. Heldring, A. H. Shaw and J. B. Beltman, Unraveling the effect of intra- and intercellular processes on acetaminophen-induced liver injury, npj Syst. Biol. Appl., 2022, 8, 27 CrossRef CAS PubMed.
- K.-J. Jang, et al., Reproducing human and cross-species drug toxicities using a Liver-Chip, Sci. Transl. Med., 2019, 11, eaax5516 CrossRef CAS PubMed.
- T. Guo, et al., Regulators, functions, and mechanotransduction pathways of matrix stiffness in hepatic disease, Front. Physiol., 2023, 14, 1098129 CrossRef PubMed.
- A. Sánchez and I. Fabregat, Growth factor- and cytokine-driven pathways governing liver stemness and differentiation, World J. Gastroenterol., 2010, 16, 5148–5161 CrossRef PubMed.
- A. Ramachandran and H. Jaeschke, Oxidant Stress and Acetaminophen Hepatotoxicity: Mechanism-Based Drug Development, Antioxid. Redox Signaling, 2021, 35, 718–733 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |