Open Access Article
Open Access ArticleTumor-on-chip platforms for transport phenotyping: decoding CAF-driven barriers to drug delivery
Doriane
Le Manach
 *a,
Vincent
Senez†
*a,
Vincent
Senez†
 b and
Matthias
Nees†
b and
Matthias
Nees†
 *ac
*ac
aDepartment of Biochemistry and Molecular Biology, Medical University of Lublin, W. Chodźki 1 Street, 20-093 Lublin, Poland. E-mail: matthias.nees@umlub.edu.pl
bUniv. Lille, Inserm, CHU Lille, CNRS, U1366-UMR9020 – CRCLille – Cancer Research Center of Lille, F-59000 Lille, France
cFICAN West Cancer Centre Laboratory, Cancer Research Unit, Institute of Biomedicine, Turku University Hospital, University of Turku, Turku, Finland
First published on 13th April 2026
Abstract
Physical barriers within solid tumors constitute a fundamental but often overlooked mechanism of therapeutic resistance, contributing to the poor success rate of cancer drug translation. Therapeutic molecules often fail to reach their intended targets due to mass-transport limitations imposed by the remodeled, spatially heterogeneous tumor microenvironment (TME). Cancer-associated fibroblasts (CAFs) drive dynamic remodeling of the extracellular matrix (ECM), generating local variations in stiffness, porosity, and intrinsic permeability that, together, shape evolving transport phenotypes that govern drug accessibility. We focus on mechanical pathways of stromal mechanotransduction, tracing the sequence from CAF activation through ECM remodeling, to barrier formation, and show how these processes collectively govern therapeutic outcomes. We also evaluate advanced microfluidic and tumor-on-chip (ToC) platforms that reproduce stromal heterogeneity under controlled conditions, mimicking tissue architecture, transport behavior, and therapeutic response. By enabling patient-specific profiling of CAF-driven transport phenotypes, these systems demonstrate that transport barriers are not fixed obstacles but dynamically modifiable therapeutic targets. “Transport phenotyping” could complement genomic profiling in clinical oncology by integrating heterogeneity, biophysics, and precision medicine, potentially transforming personalized treatment strategies for patients whose tumors remain refractory to current therapies.
1. Introduction
Contemporary cancer therapeutics face a fundamental paradox: despite unprecedented molecular characterization of oncogenic processes and the sophisticated development of targeted agents, recurrent and metastatic solid tumors remain largely refractory to treatment. This paradox contributes to poor patient survival outcomes. Annual approval statistics show that new chemical entities often display remarkable preclinical efficacy, yet around 95% fail to achieve meaningful survival benefits in clinical trials, with most failures occurring in phase II and III due to lack of effectiveness.1,2 This pattern of early discovery success but late-stage attrition cannot be explained solely by biological complexity or tumor heterogeneity and plasticity. An increasingly recognized obstacle lies in the physical architecture of solid tumors: a dense, fibroblastic (desmoplastic) stroma that prevents drugs, from small molecules to antibodies and nanoparticles, from reaching malignant cells.3 Intrinsic drug resistance reflects cell-autonomous mechanisms such as mutations in molecular targets, activation of efflux pumps, or enhanced DNA repair pathways. Extrinsic resistance arises from the tumor microenvironment (TME), which acts as both a physical and biochemical barrier to treatment. While genetic and molecular resistance mechanisms have been extensively reviewed,4,5 we focus here on the physical microenvironment, notably the extracellular matrix (ECM), as a critical determinant of extrinsic, transport-mediated resistance.‡ This stromal barrier is not generated by tumor cells themselves but by the cellular components of the TME, cancer-associated fibroblasts (CAFs), through ECM remodeling and mechanical stress. Tumor heterogeneity and cell plasticity occur across all solid cancers and vary not only between patients but also among regions of the same lesion.6 Beyond cellular variability, the ECM undergoes continuous remodeling, generating spatially and temporally heterogeneous stiffness, porosity, and intrinsic permeability that govern drug accessibility.7,8 Here, we use the term intrinsic permeability (k) to denote the matrix property of the ECM, distinguishing it from hydraulic conductivity, which additionally depends on fluid viscosity and is often used inconsistently in the biological literature. Such remodeling is dynamic, nonlinear, and shaped by CAF heterogeneity and biochemical–mechanical feedbacks.9–11 CAF-driven matrix remodeling thus shapes evolving transport landscapes that determine whether therapeutic agents and immune cells can reach malignant cells.12,13Patients with stroma-rich, fibrotic, and desmoplastic tumors such as pancreatic ductal adenocarcinoma (PDAC), and subsets of breast and ovarian adenocarcinomas or head-and-neck squamous carcinomas (HNSCC) face particularly poor outcomes.14–16 PDAC, in which the stromal compartment can constitute up to 90% of tumor volume, remains among the deadliest tumor entities despite significant therapeutic progress.17–19 Standard therapies and even modern immunotherapies often fail not because molecular targets are absent, but because dense stroma prevents drugs and immune cells from accessing tumor islands, creating immune-excluded or ‘cold’ tumors.20–26
CAFs orchestrate this transport barrier through ECM deposition, fiber reorganization, and cytokine programs that spatially segregate immune cells and therapeutic agents.27 For example, CXCL12 secretion sequesters T-cells in stromal compartments, a mechanism reversible by CXCR4 antagonists such as Plerixafor, revealing that matrix remodeling defines an immune transport dimension in tumors.28 The TME therefore behaves as a dynamic and self-adapting transport system rather than a passive scaffold. CAFs, a heterogeneous population of tumor-reprogrammed stromal cells, continuously deposit and reorganize ECM components such as collagen and proteoglycans, establishing stiffness gradients and fiber architectures that sustain invasion and therapy resistance.18,21 Proliferation of cancer cells and CAFs generates excessive ECM and solid stress (SS) that compresses blood and lymphatic vessels, elevates interstitial fluid pressure (IFP), and disrupts perfusion, producing heterogeneous advective-diffusive transport regimes that limit drug penetration.29–33 Quantitative studies indicate that only a minute fraction of administered drugs reaches malignant cells, underscoring the dominant role of physical barriers.33
Both cancer cells and CAFs sense and respond to these mechanical cues via mechanotransduction pathways (including integrins, the cytoskeleton, ion channels, and YAP/TAZ, MAPK signaling), converting forces into biochemical programs that further drive ECM remodeling and barrier reinforcement.34–40 This establishes a bidirectional mechanochemical feedback loop in which matrix mechanics and transport properties continuously co-evolve with stromal activation.38,41–45
Consequently, each tumor develops a unique and evolving transport phenotype, defined by ECM architecture, intrinsic permeability, and interstitial flow patterns that govern the accessibility of drugs and immune cells. However, identifying and quantifying which features of this remodeled matrix impede therapeutic transport remains a major challenge. Microfluidics and tumor-on-chip (ToC) technologies have emerged to address this problem by combining physiological relevance with experimental control and transparency. Unlike 2D cultures or animal models, ToC platforms can reproduce stromal architecture, pressure gradients, oxygen and nutrient transport, and CAF-mediated ECM remodeling under defined conditions, while enabling direct measurement of stiffness, porosity, and molecular gradients.46–52 These systems therefore provide a unique opportunity to characterize patient-specific transport phenotypes and to link CAF-driven matrix remodeling to therapeutic accessibility. The central motivation of this review is to examine how CAF-mediated mechanotransduction drives ECM remodeling and transport barrier formation, and how microfluidic and computational models can be used to quantify and perturb these transport phenotypes. Each approach provides distinct mechanistic insights and translational opportunities. We therefore structure this review around a causal and transport-centered framework. Section 2 establishes the biological foundations by defining the functional states of CAFs and their roles in ECM remodeling and mechanical feedback. Section 3 introduces the physical principles of mass transport in dynamically remodeling tissues and shows how these concepts can be experimentally accessed using microfluidic tumor-on-chip platforms. Section 4 integrates these biological and physical dimensions to analyze transport-limited, extrinsic mechanisms of therapeutic resistance driven by stromal remodeling. Finally, section 5 discusses how transport phenotyping can be leveraged as a functional complement to genomic profiling, with implications for patient stratification, therapeutic sequencing, and clinical translation.
2. The causality chain: a unifying framework for the impact of matrix remodeling on tumor biology
2.1 Conceptual foundations
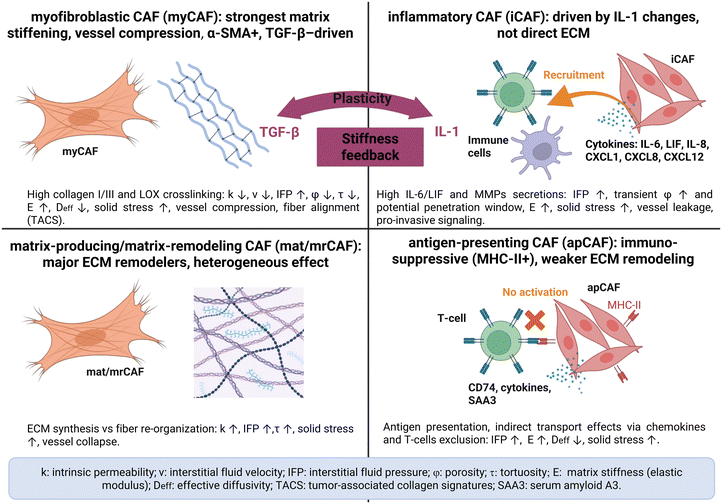
The complexity of tumor–stroma interactions has historically resisted purely reductionist approaches. CAF heterogeneity, ECM composition, chemical gradients, and mechanical forces are coupled through nonlinear feedback loops that blur simple cause-and-effect relationships.10,11,53–55 This complexity calls for conceptual frameworks that integrate cellular diversity, matrix mechanics, and dynamic transport. CAF populations form a functional spectrum of overlapping states that jointly shape the TME.56 Among these, we define matrix-producing and matrix-remodeling CAFs (mat/mrCAFs) as the CAF populations whose dominant functional program is ECM deposition, crosslinking, and fiber reorganization. This operational definition largely overlaps with the classical myofibroblastic CAF (myCAF) phenotype, characterized by high α-smooth muscle actin (α-SMA) expression and contractility, as well as strong expression of fibrillar collagens (COL1A1, COL3A1), fibronectin (FN1), and crosslinking enzymes such as LOX and LOXL1/2. These enzymes generate covalent collagen crosslinks that increase matrix stiffness while reducing porosity and intrinsic permeability.57,58 In contrast, inflammatory CAFs (iCAFs) display dominant cytokine and chemokine programs (e.g., IL-6, CXCL12) that primarily modulate paracrine signaling, immune recruitment, and vascular function, while antigen-presenting CAFs (apCAFs) express MHC-II transcripts and participate in immune interactions.59–62 Although these CAF states overlap biologically and individual CAFs may display mixed programs, their distinction is useful because their dominant functions have different consequences for tissue transport: mat/mrCAF matrix deposition reduces porosity and increases tortuosity; myCAF contractility generates solid stress and vessel compression; iCAF and apCAF programs indirectly modulate transport through the immune and vascular pathway. For clarity, we use the mat/mrCAF label as a shorthand when discussing matrix to transport coupling.CAF heterogeneity extends beyond molecular markers to functional consequences on matrix architecture and transport properties, with distinct CAF states differentially shaping stiffness, porosity, and immune-cell accessibility (see Box 1).
These CAF populations are embedded in reciprocal mechanochemical feedback loops. Activated mat/mrCAFs deposit and reorganize ECM, increasing stiffness and generating fiber alignment and gel contraction. The remodeled matrix, in turn, alters the mechanical and biochemical environment sensed by CAFs, further reinforcing their activated state. Transport processes are an integral part of this loop: ECM remodeling modifies diffusivity, intrinsic permeability, and interstitial flow, thereby reshaping the spatial distributions of oxygen, nutrients, cytokines, immune cells, and therapeutic agents.63 These altered gradients feedback on CAF activation, cancer cell plasticity, and immune behavior, creating emergent transport barriers that are not fixed but dynamically maintained. Single-cell and functional studies illustrate how distinct CAF subtypes contribute to this process through complementary mechanical strategies. For example, CAF-S4/myCAF-like cells exhibit a highly contractile, pericyte-like program with elevated strain-energy density sufficient to compact local collagen and mechanically promote invasion, whereas CAF-S1/myCAF-like cells emphasize proliferative and secretory programs that further drive ECM production and cytokine-mediated crosstalk.64 Together, these CAF subsets generate spatially organized mechanical and matrix architectures that define transport pathways within the tumor.65,66 At the tissue level, CAF-driven ECM deposition and contractility produce desmoplasia, increased solid stress, and vessel compression, leading to elevated interstitial fluid pressure and heterogeneous advective-diffusive transport regimes.67 These physical changes translate directly into spatial gradients of oxygen, metabolites, drugs, and immune cells, which in turn govern cancer cell proliferation, migration, and survival. Histologically, this process manifests fibrotic stromal networks surrounding distorted tumor islands, abnormal vasculature, and infiltrative growth patterns that correlate with clinical aggressiveness and therapeutic resistance.65,66,68–75
Box 1 Schematic overview of representative CAF functional states and their dominant effects on extracellular matrix (ECM) structure and mass transport in stroma-rich carcinomasUpper left (myCAF): primary matrix-stiffening subtype; upper right (iCAF): inflammatory subtype with indirect transport effects via MMPs and vascular leakage; lower left (mat/mrCAF): heterogeneous ECM remodelers; lower right (apCAF): immunosuppressive subtype with indirect transport effects via T-cell exclusion. The central element depicts bidirectional plasticity driven by TGF-β/IL-1 between subtypes. Myofibroblastic CAFs (myCAFs) are characterized by high α-smooth muscle actin (α-SMA) expression, contractility, and TGF-β-driven matrix crosslinking, leading to increased tissue stiffness, vessel compression, elevated interstitial fluid pressure, and reduced effective diffusivity and intrinsic permeability. Matrix-producing and matrix-remodeling CAFs (mat/mrCAFs) deposit, reorganize, and degrade ECM fibers, dynamically modulating porosity, tortuosity, and permeability, thereby generating spatially heterogeneous transport pathways. Inflammatory CAFs (iCAFs) secrete cytokines and chemokines (e.g., IL-6, CXCL12) that primarily affect immune-cell localization and vascular function, indirectly influencing transport. Antigen-presenting CAFs (apCAFs) express MHC class II molecules and modulate immunity, with secondary effects on immune infiltration and spatial drug distribution. Not all CAF states are present or dominant across all tumor types. The schematic illustrates typical functional programs reported in desmoplastic cancers and highlights how distinct CAF activities contribute, directly or indirectly, to the emergence of CAF-driven transport phenotypes that regulate drug and immune-cell accessibility. |
Together, these observations motivate a causal framework in which CAF activation, ECM remodeling, and transport are tightly coupled (Fig. 1). Rather than representing linear cause-and-effect relationships, the arrows in this framework denote bidirectional feedback: CAFs remodel the matrix, the matrix alters transport, and transport reshapes cellular states. This reciprocal coupling can generate stable or evolving stromal–tumor configurations whose defining feature is a specific transport phenotype. Understanding and perturbing these CAF-driven transport phenotypes is therefore central to deciphering therapy resistance and to designing effective tumor-on-chip experiments. Indeed, this framework naturally enables hypothesis-driven ‘virtual experiments’ by inverting inputs and outputs along the causality chain. Forward causality can be tested by perturbing upstream drivers of CAF activation and remodeling (e.g., TGF-β signaling or LOX activity) while tracking transport readouts, whereas reverse causality can be probed by directly modulating ECM-level variables (stiffness, porosity, intrinsic permeability, or interstitial pressure) and quantifying cellular responses. Microfluidic systems are particularly well suited for such bidirectional causal tests because they allow controlled tuning of boundary conditions and matrix properties together with real-time, multiparametric measurements, thereby providing a quantitative bridge between stromal mechanics and drug transport.76–87 When the ECM is considered as the primary causal input, relevant structural, mechanical, and molecular parameters, including stiffness, composition, porosity, permeability, and mechanotransduction readouts, can be quantified experimentally. A detailed list of input and output variables and their measurement modalities is provided in section S1.
2.2 Transport physics in dynamic biological systems
Eqn (1)–(7) together form a compact mathematical framework that links solute transport and tissue mechanics in CAF-remodeled tumors. Numerical and analytical solutions yield spatial and temporal maps of solute concentration, interstitial pressure, and tissue deformation, which directly reflect how ECM density, crosslinking, and solid stress shape drug and immune-cell accessibility. These maps address biologically central questions: where will a drug penetrate within a CAF-rich tissue? Is poor delivery dominated by limited diffusion through a dense collagen network, by advective wash-out driven by elevated interstitial fluid pressure, or by rapid cellular uptake? How will interventions that modulate CAF activity, alter matrix structure, or relieve solid stress reshape tissue exposure? For readers less familiar with mathematical notation, the equations can be read as a quantitative blueprint that connects CAF-driven ECM remodeling to measurable transport parameters such as effective diffusivity, intrinsic permeability, and interstitial pressure, and from there to spatial patterns of drug distribution that can be directly compared with microfluidic and tumor-on-chip experiments.12,13,88–92 A complete list of symbols, units, and their corresponding experimental readouts in tumor-on-chip systems is provided in Table S1.
Box 2 Mathematical framework for CAF-driven transport phenotypingUnderstanding solute transport through tumor tissue requires applying physical principles to dynamic, evolving biological systems. In solid tumors, CAF activity, ECM remodeling, vascular leakiness, and cell contractility continuously reshape diffusion, advection, and tissue deformation. Unlike static engineered porous media, tumor transport therefore occurs in a matrix whose porosity, stiffness, and permeability evolve over time. The coupled advection–diffusion–reaction and poroelastic equations introduced in section 2.2 provide a compact but mechanistically meaningful way to formalize this complexity. Rather than being abstract, each variable appearing in the equations corresponds to a biologically and experimentally accessible property of CAF-remodeled tissue. For example, the effective diffusivity Deff reflects how collagen density, fiber alignment, and crosslinking by mat/mrCAFs restrict molecular motion; the intrinsic permeability k captures how ECM compaction, LOX-mediated crosslinking, and cellular contraction regulate interstitial fluid flow; and the interstitial fluid pressure p reports the combined effects of vascular leakiness and CAF-driven solid stress. |
In this formulation, matrix remodeling and mechanotransduction directly enter the transport problem through their impact on these coefficients. Increased matrix deposition and crosslinking by mat/mrCAFs reduce ϕ, increase tortuosity, and lower both Deff and k, while myCAF contractility elevates solid stress and interstitial pressure, thereby modifying advective transport. Conversely, enzymatic matrix degradation, mechanical decompression, or vascular normalization act by shifting the same parameters in the opposite direction. The mathematical framework therefore translates qualitative descriptors such as ‘dense’, ‘stiff’, or ‘fibrotic’ into quantitative variables that can be measured, quantitative variables that can be measured, perturbed, and used to test biological hypotheses. By linking CAF activity and ECM structure to transport coefficients that determine drug and immune-cell access, this formalism provides a quantitative bridge between stromal biology and therapeutic delivery and defines the basis for transport phenotyping in tumor-on-chip systems.
For readers less familiar with mathematical formalism, the conceptual role of the equations introduced below and their relationship to experimentally measurable biological quantities are summarized in Box 2.
 | (1) |
Here, c is the local solute concentration, Deff is the effective diffusivity in the CAF-remodeled extracellular matrix, vp is the interstitial fluid velocity, and R represents cellular uptake, degradation, or local secretion.
In CAF-rich tumors, none of these terms are spatially or temporally constant.94,95 The effective diffusivity Deff reflects how matrix-producing and matrix-remodeling CAFs alter collagen density, fiber alignment, and crosslinking. Increased deposition of fibrillar collagens and LOX-mediated crosslinking reduce matrix porosity ϕ and increase tortuosity λ, thereby lowering Deff = D0ϕ/λ2 relative to free diffusion in the interstitial fluid.96–99 As a result, even small molecules experience slowed penetration in desmoplastic regions, while large biologics or nanoparticles are much more strongly hindered.
The advective term ∇·(vpc) captures the contribution of interstitial fluid flow to solute transport. In tumors, CAF-driven matrix contraction, vessel compression, and vascular leakiness generate elevated and spatially heterogeneous interstitial fluid pressure, producing local fluid velocities that can either enhance delivery along pressure gradients or promote convective wash-out from poorly perfused tumor regions. Thus, CAF activity enters the transport equation not only through diffusion but also through pressure-driven advection.
The reaction term R(x, y, z, t) accounts for cellular uptake, binding, or degradation of the solute. In addition to cancer-cell–intrinsic drug consumption, CAFs themselves can sequester, metabolize, or exclude therapeutic agents through ECM binding and pericellular matrix deposition. Consequently, the apparent rate of drug loss within a CAF-rich microenvironment reflects both cellular and matrix-associated sinks.
Unlike engineered porous materials, tumor tissues evolve continuously: CAF activation, ECM remodeling, vascular collapse or normalization, and cellular turnover all modify Deff, vp, and R over time. The advection–diffusion–reaction equation therefore provides a dynamic, mechanistically grounded description of how CAF-driven matrix remodeling gives rise to heterogeneous and time-dependent transport phenotypes that govern drug accessibility.97,98
| q = −k(ϕ, x, y, z, t)/μ·∇p | (2) |
The interstitial fluid velocity vp entering eqn (1) is described by:
| vp = q/ϕ(x, y, z, t) = −(1/ϕ)k(ϕ, x, y, z, t)·∇p. | (3) |
Darcy's law thus provides a mechanistic link between stromal remodeling and drug delivery: CAF-mediated matrix densification and vessel compression simultaneously reduce permeability and elevate pressure, shifting transport from diffusion-dominated regimes toward outward, pressure-driven flow. These coupled effects are central to the emergence of CAF-driven transport barriers that limit therapeutic and immune-cell access.
| ∇·σ + b = 0, | (4) |
To relate stress to deformation, poroelastic or poroviscoelastic constitutive laws are used. In this framework, tissue stress depends both on solid deformation and on pore pressure.103,104 A Kelvin–Voigt-type formulation,
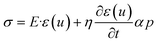 | (5) |
In CAF-remodeled tissues, these coefficients are not material constants but biological variables. Matrix deposition and LOX-mediated crosslinking by mat/mrCAFs increase the effective elastic modulus E, while myCAF contractility and cell proliferation generate compressive strain ε that compacts the matrix. Elevated pore pressure p, resulting from vascular leakiness and vessel compression, further contributes to solid stress. Together, these effects lead to progressive densification of the ECM, reduction of pore volume, and distortion of fluid pathways.
Through this coupling, mechanical remodeling feeds directly back into transport.92,95 Local increases in stiffness and compressive strain reduce porosity ϕ and intrinsic permeability k, while elevated pore pressure modifies advective flow. The poro(visco)elastic formulation therefore provides a mechanistic description of how CAF-driven force generation and matrix stiffening translate into dynamic, spatially heterogeneous transport barriers within solid tumors.105,106
| ∇·q + ∂/∂t[α(∇·u) + (1/M)p] = Qf | (6) |
In CAF-rich tumors, this equation formalizes how matrix compaction and vessel compression alter fluid storage. MyCAF-driven contraction and proliferative growth of tumor cells reduce pore volume, expelling fluid and raising interstitial pressure, whereas vascular leakiness continuously injects fluid into the tissue. The balance of these processes determines whether interstitial flow stagnates or whether pressure-driven fluid motion dominates.
Because porosity changes with deformation, solute storage must evolve accordingly. A generalized solute mass balance including sorption is:
| ∂/∂t[ϕc + (1 − ϕ)cs] + ∇·(cq − ϕDeff(ϕ, x, y, z, t)·∇c) = R(c, p, u, x, t) + Qc | (7) |
Here, ϕ is the local porosity, c the mobile solute concentration, and cs the fraction bound or immobilized within the solid matrix (for example through binding to collagen, proteoglycans, or pericellular ECM deposited by CAFs). Qc represents local solute sources or sinks.
CAF-driven ECM deposition, crosslinking, and compression decrease ϕ, reducing the volume available for fluid and solute transport while increasing the fraction of drug that becomes trapped or immobilized within the dense matrix. Conversely, enzymatic matrix degradation or mechanical decompression increases pore volume and reduces solute sequestration, transiently enhancing drug penetration and retention.
This formulation therefore captures an important biological reality: CAF-mediated matrix remodeling does not only alter how fast drugs move, but also how much of a drug can be stored, retained, or excluded within the tumor microenvironment, contributing to heterogeneous and time-dependent therapeutic exposure.93,95,97
For example, collagen content and LOX-mediated crosslinking determine matrix stiffness and pore architecture, which can be quantified by rheology, AFM, and imaging-based measurements of porosity and fiber organization.77,78,82,110,111 These structural properties directly constrain Deff and k, which can be extracted from tracer diffusion, FRAP, or pressure-driven flow assays in microfluidic devices.78,82 Similarly, myCAF contractility and tissue compression modulate solid stress and interstitial pressure, which can be measured through pressure sensors, deformation tracking, or poroelastic relaxation experiments.84,112,113
By fitting the transport and poroelastic equations to experimental data, one can infer spatially and temporally resolved transport phenotypes that reflect the underlying CAF and ECM state. Importantly, this calibration is not limited to static conditions: microfluidic systems allow perturbation of CAF activity (e.g., TGF-β inhibition, LOX blockade), matrix structure (enzymatic degradation, crosslinking), or mechanical loading, enabling direct tests of how specific stromal interventions reshape Deff, k, and p, and thereby drug accessibility.
In this way, tumor-on-chip platforms transform the mathematical framework into a practical tool for CAF-driven transport phenotyping, linking stromal biology, physical transport parameters, and therapeutic response in a quantitatively testable manner.
2.3 Designing experiments to extract CAF-driven transport phenotypes
The central challenge in tumor transport modeling is not mathematical formulation but experimental calibration: the deliberate design of experiments that yield biologically meaningful values for effective diffusivity Deff, intrinsic permeability k, and reaction terms R in CAF-remodeled tissues. Because these parameters emerge from the cellular and stromal architecture of tumors, experimental systems must capture both matrix mechanics and CAF-driven remodeling.12,13,34,114,115Microfluidic tumor–stroma platforms provide a uniquely suited environment for this task. By controlling CAF composition, ECM density, and interstitial pressure while continuously measuring tracer transport and tissue deformation, these systems enable direct mapping of stromal state to transport phenotype. In practice, this means that CAF populations (for example, myCAF-enriched versus mat/mrCAF-enriched mixtures) can be embedded in collagenous matrices, exposed to defined mechanical or biochemical perturbations, and probed using fluorescent tracers or therapeutic molecules to quantify penetration and retention over time. A practical entry point is to extract spatially averaged transport coefficients by fitting advection–diffusion–reaction models to tracer uptake or release data. For example, time-resolved penetration of fluorescent dextran derivatives, albumin, or chemotherapeutics in CAF-conditioned matrices can be fitted to infer Deff and apparent uptake rates.107–109,116
Similarly, pressure-relaxation or flow-through assays yield estimates of intrinsic permeability and elastic moduli, directly reporting how CAF-driven matrix densification and contraction modulate interstitial transport.117–119
Importantly, CAF-mediated remodeling is inherently heterogeneous and dynamic. As CAFs deposit, crosslink, and reorganize collagen, local values of Deff and k evolve over days to weeks. Microfluidic platforms allow these temporal changes to be tracked and, when combined with imaging of collagen architecture or CAF localization, provide spatially resolved transport phenotypes that reflect the underlying stromal organization.
Detailed descriptions of hydrogel systems, tracer-based diffusion measurements, pressure–flow assays, and sources of experimental variability, as well as their use for quantitative model calibration, are provided in section S2, which presents the full methodological and materials-science framework supporting this approach.
By systematically perturbing CAF activity (e.g., TGF-β blockade, LOX inhibition), matrix structure (enzymatic degradation, crosslinking), or mechanical loading (compression, stress relief), one can establish causal links between stromal biology and transport.120,121 In this way, tumor-on-chip systems enable bidirectional testing: CAF-driven matrix remodeling predicts transport behavior, and measured transport phenotypes, in turn, reveal the dominant stromal mechanisms that limit drug delivery. These CAF-driven transport parameters do not change monotonically but evolve through characteristic temporal phases during tumor progression, yielding transient windows of enhanced accessibility, followed by the establishment of persistent transport barriers. A conceptual synthesis of these dynamics, integrating experimental observations and tumor-on-chip studies, is illustrated in Fig. 2.
2.4 Clinical implications: from CAF-driven transport phenotypes to therapeutic stratification
Modular in vitro tumor–stroma models provide a practical route to translate CAF-driven transport biology into clinically actionable information. By explicitly manipulating fibroblast populations, ECM composition, and mechanical loading, these systems allow direct measurements of how stromal remodeling shapes drug and immune accessibility.Patient-derived CAFs or defined CAF subtypes can be combined with controlled ECM formulations to generate tissues with distinct transport phenotypes, characterized by their effective diffusivity, intrinsic permeability, and interstitial pressure.122,123 Fluorescent tracers or therapeutic agents introduced into these systems reveal how rapidly and how deeply molecules penetrate, providing a functional readout of stromal barrier strength. When integrated with transport models, these measurements identify transient windows of enhanced accessibility or persistent regions of exclusion. This approach has immediate clinical implications. Tumors that appear similar at the genomic or histological level can differ markedly in their CAF composition and matrix architecture, leading to very different transport phenotypes. Such differences help explain why drugs that are highly potent in vitro or in genetically stratified patients fail to achieve sufficient exposure in desmoplastic tumors such as PDAC, breast, or ovarian carcinomas.13,29,124,125 Transport phenotyping therefore provides a complementary axis of patient stratification, orthogonal to molecular profiling.
Importantly, CAF-driven transport barriers are not fixed. Enzymatic matrix remodeling, mechanical stress relief, or pharmacological reprogramming of CAFs can transiently increase permeability and diffusivity, thereby improving drug penetration. Tumor-on-chip platforms allow these interventions to be tested in a controlled and patient-specific manner, enabling rational sequencing of stromal-modulating and cytotoxic or immunotherapeutic agents.125–127
A detailed overview of experimental and pharmacological strategies to modulate CAF-driven matrix barriers, including enzymatic, mechanical, and stromal reprogramming approaches, is provided in section S3.
By linking CAF biology to measurable transport parameters, and by providing experimental systems to perturb both, transport phenotyping offers a framework to identify which patients are likely to benefit from stromal targeting, and when such interventions should be applied to maximize therapeutic access.
3. Microfluidic platforms: bridging scales and transport complexity
3.1 From model hierarchy to transport-resolved tumor-on-chip systems
The evolution from conventional in vitro tumor models to microfluidic ToC platforms has been driven by a central limitation of standard systems: their inability to resolve spatially heterogeneous transport under dynamic stromal remodeling. While 2D monolayers, spheroids, and organoids reproduce selected aspects of tumor organization, they fail to capture the coupled gradients of nutrients, oxygen, drugs, and mechanical stresses that emerge in confined, perfused, and remodeling matrices.128 Microfluidic ToC systems uniquely address this gap by combining controlled geometry, perfusion, and mechanical confinement with three-dimensional cell–matrix organization. These platforms enable the direct measurement of interstitial flow, pressure gradients, and solute transport within evolving stromal architectures, thereby linking CAF activity, matrix remodeling, and drug accessibility in a single experimental framework.129 In desmoplastic tumors, where CAF-driven matrix densification and solid stress dominate transport resistance, this capability is essential.71–73,130,131 An overview of the hierarchy of tumor models used for mass transport studies, including sealed and openable tumor-on-chip configurations, is provided in Fig. S1. Material-dependent transport artefacts and their implications for quantitative transport assays are summarized in Table S2.3.2 Core microfluidic architectures for transport phenotyping
Despite wide design diversity, most transport-focused ToC platforms fall into three functional classes, defined by their ability to impose and measure transport-relevant fields.Material-specific transport artefacts and surface functionalization strategies are discussed in section S4.
3.3 Toward longitudinal, transport-resolved tumor models
A recurring limitation of current ToC platforms is their predominantly static or endpoint characterization of transport. CAF-driven remodeling, however, is inherently dynamic: intrinsic permeability, porosity, and solid stress evolve over days to weeks as matrix deposition, contraction, and reorganization progress.9,10,115,141–144 Capturing these dynamics, therefore requires longitudinal platforms capable of repeated or continuous measurements of key transport parameters, including intrinsic permeability, interstitial velocity, and solute diffusion.51,52,86,107–109,116Because transport properties are inseparably linked to the physicochemical and mechanical characteristics of the surrounding extracellular matrix, the choice of hydrogel and ECM formulation becomes a central experimental determinant rather than a neutral scaffold. Different matrices offer distinct ranges of stiffness, degradability, and pore architecture, which directly condition the accessible transport regimes and the interpretation of CAF-driven remodeling effects. The implications of hydrogel and ECM choice for CAF-driven transport phenotyping, including tunability of transport parameters and associated experimental pitfalls, are summarized in Table S3.
However, dynamic CAF-driven remodeling not only alters transport coefficients; it can also induce matrix contraction, densification, and geometric distortion of the gel compartment itself.48,145,146 If not adequately controlled, these effects confound transport measurements by modifying flow paths, pressure gradients, and effective diffusion distances over time. Consequently, reliable longitudinal transport phenotyping requires actively remodeling matrices to be mechanically and chemically stabilized within the microfluidic device. Detailed hydrogel stabilization and anchoring strategies are described in sections S5 and S6.
Future ToC systems should therefore prioritize (i) stable confinement of actively remodeling matrices, (ii) reproducible control of pressure and flow boundary conditions, and (iii) integration of transport assays with structural and mechanical characterization. When combined with continuum or poroelastic transport models, such platforms enable the extraction of physically meaningful parameters that define a tumor's transport phenotype rather than a single drug response.147,148
3.4 Technical limitations and methodological caveats
Despite their increasing sophistication, tumor-on-chip platforms remain constrained by methodological and technical limitations that must be considered when interpreting transport measurements. No single experimental technique simultaneously captures solute transport, matrix structure, and mechanics without bias. Fluorescent-based methods such as FRAP and fluorescent correlation spectroscopy (FCS) enable estimation of effective diffusivity but can locally perturb the system through photobleaching or photochemical damage and are limited to a small spatial sampling volume.116,149 Single-particle tracking (SPT) approaches often rely on nanometer-scale tracers (20–100 nm) that exceed the effective pore size relevant for small-molecule drugs, complicating quantitative extrapolation.150Structural imaging techniques, including second-harmonic generation (SHG) and confocal reflectance microscopy, provide detailed information on collagen organization and fiber alignment but do not directly report transport parameters.151,152 Mechanical mapping approaches such as atomic force microscopy (AFM), Brillouin microscopy, and elastography probe distinct mechanical regimes and length scales, each with intrinsic limitations in depth penetration, spatial resolution, or biological interpretation.153 Moreover, labeling strategies can alter tracer size, charge, or hydrophobicity, thereby modifying transport behavior relative to unlabeled compounds.154 These constraints underscore the necessity of multimodal experimental strategies that combine direct transport assays, structural and mechanical characterization, and mechanistic modeling. Cross-validation across techniques is essential to disentangle biological effects from methodological artefacts and to derive physically meaningful estimates of effective diffusivity, intrinsic permeability and poroelastic properties in evolving tumor microenvironments. Detailed methodological considerations and reporting recommendations are provided in section S7. Beyond these technical aspects, several unresolved conceptual and experimental challenges currently limit the quantitative interpretation and comparison of transport measurements across tumor-on-chip platforms; these are summarized in Box 3.
Box 3 Open questions on transport and mechanical confinement in tumor-on-chip systemsThis Box highlights unresolved conceptual and experimental challenges that currently limit the quantitative interpretation of transport phenotypes in tumor-on-chip systems, particularly in the context of CAF-driven matrix remodeling and mechanical confinement.1. Dynamic evolution of transport parameters under CAF-driven remodeling How do key transport parameters, such as intrinsic permeability (k(ϕ, x, y, z, t)) and effective diffusivity (Deff(ϕ, x, y, z, t)), evolve over time as CAFs remodel the extracellular matrix and generate solid stress? How can these dynamics be captured without assuming static tissue properties, given the continuous and nonlinear nature of stromal remodeling? 2. Separating device-induced artifacts from biological transport barriers How can transport limitations arising from device-related effects, such as material sorption, gas permeability, or surface chemistry, be disentangled from genuine biological barriers created by CAF-driven ECM densification, fiber alignment, and stress-induced confinement? 3. Independent and coupled roles of solid stress and interstitial pressure How can solid stress and interstitial fluid pressure be independently imposed or to identify their respective and combined contributions to drug, oxygen, and immune-cell transport within confined tumor microenvironments? 4. Coupling vascular dysfunction with interstitial transport How can perfusion, endothelial permeability, and impaired lymphatic drainage be integrated with stromal remodeling to reproduce hypoperfusion-high IFP regimes characteristic of desmoplastic tumors? 5. Immune-cell transport in mechanically confined, CAF-remodeled matrices How do chemotaxis, infiltration, and persistence of immune cells change as CAF-driven matrix density, fiber alignment, and compressive stress are systematically varied? To what extent does mechanical confinement contribute to immune exclusion independently of immunosuppressive signaling? 6. Toward comparable transport phenotypes across platforms Which minimal set of transport- and mechanics-related parameters, such as ECM stiffness, intrinsic permeability, effective diffusivity, pressure gradients, porosity, fiber anisotropy, or confinement indices, is required to enable meaningful comparison and interpretation of transport phenotypes across tumor-on-chip studies? |
3.5 Patient-derived systems and translational relevance
Patient-derived ToC platforms extend transport phenotyping beyond generic models by preserving native stromal composition, CAF subtype balance, and ECM architecture. Unlike genomic profiling, which identifies molecular vulnerabilities, these systems directly probe whether therapeutic agents can physically reach malignant cells within their microenvironment.Experimental evidence increasingly shows that tumors with higher effective permeability or lower matrix-induced resistance exhibit faster and stronger responses to therapy, whereas dense, CAF-rich matrices delay drug accumulation and blunt efficacy.155 By quantifying transport parameters under physiologically relevant flow conditions, patient-derived ToC models provide a functional complement to molecular biomarkers and offer a framework to stratify tumors based on accessibility rather than target expression alone.
Despite encouraging validation studies, clinical translation remains limited by assay duration, technical complexity, and regulatory constraints. Nevertheless, transport-focused ToC systems represent a promising intermediate scale between mechanistic modeling and clinical pharmacokinetics, particularly for desmoplastic tumors where physical barriers dominate therapeutic failure.
Given the strong sensitivity of transport measurements to experimental conditions and inter-laboratory variability, recommended minimum reporting standards and reproducibility considerations for tumor-on-chip transport studies are summarized in section S8.
The ability of tumor-on-chip platforms to quantify evolving transport barriers provides the experimental foundation for transport phenotyping. In the following section, we examine how these platforms are used to interrogate CAF-driven matrix remodeling as a modifiable therapeutic target.
The value of experimental tumor models lies less in predicting clinical outcomes with absolute precision, a goal unattainable given the multifactorial nature of efficacy, toxicity, and clearance, but rather in identifying the key biophysical determinants of drug accessibility.
Microfluidic systems incorporating patient-derived components show exceptional promise in this regard.156,157 Beyond the use of primary patient cells, their advantage lies in retaining and reproducing native heterogeneity, including the relative abundance of CAF subtypes (mat/mrCAFs, iCAFs, pCAFs) and the ECM properties they generate, such as composition, stiffness, and spatial organization. Because both cellular composition and ECM architecture regulate interstitial transport, patient-derived ToC platforms offer a practical means to recreate clinically relevant barriers to drug penetration and chemosensitivity. Indeed, CAF heterogeneity produces stromal microenvironments that differentially modulate tumor growth and therapy response,158 while increased collagen crosslinking elevates solid stress and reduces pore size, thus limiting drug delivery. By maintaining controlled tumor–stroma ratios and ECM features, these platforms enable the systematic analysis of how CAF subtype balance matrix density and crosslinking may influence drug distribution and efficacy.159–161
Perfused microfluidic tumor platforms using primary patient-derived tissue now enable quantitative mapping of intratumoral transport under physiologically relevant flow conditions. For instance, interstitial fluid velocities of ∼0.1–10 μm s−1, comparable to those measured in vivo, have been successfully reproduced in 3D ToC models, eliciting similar transport responses.162
While genomic biomarkers identify molecular susceptibilities within the tumor cells, they provide no information about how drugs may reach those cells within their physical microenvironment. Patient-derived microfluidic systems complement these molecular assays by offering a functional transport-based phenotype. They permit direct ex vivo testing of chemosensitivity and reveal how tissue architecture, stromal density, vascular leakiness, and perfusion collectively govern drug distribution and efficacy.95,163,164 Tumors exhibiting higher permeability respond more rapidly and strongly to treatment. In contrast, dense and transport-limiting matrices delay drug accumulation and require more prolonged exposure to achieve comparable therapeutic effects.
Clinical and preclinical validation studies have shown that ToC platforms can reproduce therapeutic responses with remarkable fidelity across biological scales. In an esophageal adenocarcinoma model, neoadjuvant chemotherapy responses were accurately predicted for all eight patients within 12 days, matching the parallel clinical results.165 Comparable results were obtained in breast and prostate tumor slice cultures, which reproduced the in vivo sensitivity to cisplatin and apalutamide, respectively.166 Similarly, pancreatic cancer tumor chips based on hydrogel microcapsules generated patient-specific drug-response profiles consistent with clinical follow-up data.167 Preclinical models reinforce these findings. Petreus et al. (2021) reported almost identical irinotecan efficacy between microfluidic tumor spheroids (52% growth inhibition) and xenograft (53%), confirming quantitative alignment between these two very different platforms.168 Similarly, Toley et al. (2013) showed that reproducing doxorubicin release from liposomal formulations (Doxil) in a tumor-mimetic chip required explicit modeling of diffusion, binding, and clearance to match clinical outcomes.169
Collectively, these studies illustrate how patient-derived and preclinical ToC platforms can bridge the gap between mechanistic experimentation and clinical pharmacokinetics, complementing molecular biomarker data by providing direct, physical insight into drug accessibility within the TME. Despite these advances, substantial barriers still limit the clinical integration of ToC systems. To date, no ToC platform has received formal FDA clearance or CLIA certification for clinical use with patients (referring to the Clinical Laboratory Improvement Amendments, the U.S. regulatory framework that governs the quality standards and accreditation of clinical laboratories authorized to perform diagnostic testing on human specimens). Although both the FDA and CDC are actively evaluating these technologies through pilot initiatives such as the ISTAND program (Innovative Science and Technology Approaches for New Drugs (ISTAND) Program|FDA), broader regulatory adoption remains constrained by challenges in standardization, validation, and the definition of clear contexts of use.170,171
Beyond the regulatory dimension, practical barriers persist. Assay turnaround time remains a significant limitation: while microfluidic drug transport assays based on established cell lines can provide readouts within 5–7 days,168 workflows involving patient-derived material often require 2–4 weeks, or even up to 6–8 weeks, when patient-derived organoids (PDOs) are used, due to the extended time needed for organoid establishment, growth, and maturation.172,173 Success rates also vary widely: for example, only ∼31% of esophageal adenocarcinoma biopsies successfully yielded organoid lines,174 and a recent meta-analysis in gastric cancer reported an average success rate of ∼66.6% across multiple studies.175 Such variability reflects both biological heterogeneity and technical attrition. They also indicate that only specific subtypes of the above-mentioned cancer entities may preferentially support success in vitro cultures, thereby introducing a bias in representability.
In practice, organoids remain valuable for cell banking or expansion when sample material is limited. Still, for truly time-sensitive applications, the most direct strategy is to introduce freshly collected patient tissue into a ToC system, thus bypassing the lengthy organoid-generation step. Even so, this approach faces significant challenges, including the technical complexity of microfluidic handling, the need for skilled personnel, and rigorous quality-control standards, factors that currently restrict clinical use to well-equipped research centers. Finally, the regulatory frameworks for validating and approving physical transport-based biomarkers remain far less mature than those for molecular diagnostics, posing a critical bottleneck to complete clinical translation.
4. Mechanisms of resistance: beyond cellular autonomy
Building on the CAF-driven causality framework established in section 2, we now examine how the spatial and temporal organization of the TME governs transport and contributes to therapy resistance beyond cell-autonomous mechanisms.4.1 Temporal dynamics of transport and therapeutic windows
Time is a critical yet often underappreciated dimension of tumor biology. Microfluidic and theoretical studies consistently report biphasic transport behavior, characterized by an early advection-facilitated phase followed by diffusion-limited penetration as stromal remodeling progresses. Elliott and Yuan first formalized this transition, describing an initial permeability-dominated regime followed by transport restriction.176 Similar phase-dependent dynamics have since been reported for small molecules,177 and immune-cell trafficking.178 These observations challenge static views of drug delivery and highlight the need to consider temporal evolution when interpreting therapeutic efficacy.During early tumor development, a transient degradative phase, often dominated by MMP activity, can increase matrix permeability and reduce transport resistance.179 Drug delivery during this short window may result in relatively homogeneous penetration, even in confined tumor volumes. However, this permissive state is short-lived. As tumors mature, collagen deposition and enzymatic crosslinking progressively transform the ECM into a dense, transport-resistant network.33 Simultaneously, vascular permeability declines and size-exclusion thresholds emerge, hindering diffusion even for small-molecules.3,180 Drug regimens that are effective during early stages may thus fail once stromal barriers are established.
Hypoxia plays a central role in orchestrating these transitions.181 Oxygen depletion activates hypoxia-inducible factor 1-alpha (HIF-1α) signaling, upregulating MMPs while concurrently inducing ECM-modifying enzymes such as prolyl 4-hydroxylase α-subunit (P4HA), procollagen-lysine 2-oxyglutarate 5-dioxygenase (PLOD), and the LOX/LOXL family.31,181 Although hypoxia-driven angiogenesis transiently restores oxygenation, this relief ultimately favors matrix stabilization, increased IFP, and tissue stiffening.127,182 As oxygenation improves, metabolic shifts suppress acidic protease activity, further stabilizing the ECM.183
These dynamics have been directly observed in tumor-on-chip systems. In PDAC-on-chip models, collagen accumulation and declining intrinsic permeability occurred primarily between days 7 and 21, and correlated with reduced gemcitabine efficacy.134 Such observations illustrate that therapeutic windows are governed not only by drug properties but also by the kinetics of stromal remodeling, which vary across tumor types and patients.
4.2 Mechanical stress as a spatial regulator of transport
Tumor growth inherently generates mechanical stress through the combined effects of cellular proliferation, ECM deposition, and interstitial fluid accumulation within confined tissue. Biomechanically, solid tumors behave as biphasic materials, comprising a solid matrix and an interstitial fluid phase, whose coupled behavior is well described by poroelastic or poroviscoelastic models.Collective CAF-mediated matrix contraction generates macroscopic solid stress in the kilopascal range (∼1.3–13 kPa) in human tumors.164,184 Matrix densification simultaneously reduces intrinsic permeability, and elevates IFP, leading to vessel compression and pore constriction across multiple length scales.29,34,164,180,185–187 These stresses are spatially heterogeneous and peak at the tumor-stroma interface, where transport resistance is maximal.164,186 As a result, drug and nutrient delivery becomes highly uneven, generating intratumoral transport heterogeneity.
4.3 Mechanotransduction and feedback amplification of resistance
Mechanical stress is not a passive consequence of tumor growth but an active regulatory signal. Through mechanotransduction pathways, stromal and tumor cells convert physical forces into transcriptional programs that reinforce resistance.30,44,188Tensile strain promotes integrin clustering and focal adhesion formation, activating focal adhesion kinase (FAK)-mediated signaling cascades that promote AKT-driven survival under stress.30,127,182 Matrix stiffening induces YAP/TAZ nuclear translocation, driving expression of ECM genes such as COL1A1/COL1A2, FN1, LOX, and connective tissue growth factor (CTGF).30,127,182,189,190 This establishes a positive feedback loop in which stiffening promotes further matrix deposition and crosslinking.
Importantly, mechanical cues also influence CAF plasticity. Chronic stress can induce transition between CAF phenotypes,9,55,191 thus complicating therapeutic strategies aimed at selectively targeting specific fibroblast subpopulations. These feedback mechanisms integrate mechanical heterogeneity, transport limitation, and stromal diversity into a self-sustaining resistance program.
4.4 Extrinsic resistance revealed by patient-derived transport phenotypes
Therapy resistance in solid tumors arises from both intrinsic (cell-autonomous) and extrinsic (transport-limited) mechanisms. While intrinsic resistance, such as drug-target mutations or enhanced drug efflux, has been extensively reviewed,4 extrinsic resistance arises from the physical constraints of the TME. Dense ECM deposition, enzymatic crosslinking, and elevated IFP can prevent drugs from reaching otherwise sensitive tumor cells.22,31,124,180,192Patient-derived tumor-on-chip systems provide direct experimental evidence for transport-limited resistance. By preserving native CAF composition and ECM architecture, these platforms reproduce clinically relevant barriers to drug penetration.193 Across multiple tumor entities, effective permeability and matrix organization consistently emerge as dominant determinants of therapeutic response.124,194,195 Tumors with higher permeability respond faster and more strongly to treatment, whereas dense, CAF-rich matrices delay drug accumulation and blunt efficacy.
Several studies demonstrate the translational relevance of this transport phenotype. Multiplexed microfluidic platforms that culture intact tumor slices preserve native ECM architecture and stromal composition and generate spatially resolved drug-response readouts within days, directly linking tissue penetration to efficacy.196 Patient-derived pancreatic tumor-on-chip systems incorporating organoids with stellate cells and macrophages recapitulate desmoplastic matrix deposition and show altered chemotherapy responses when stromal composition is experimentally modified, supporting a causal role for matrix-mediated transport barriers.197 Fully 3D-printed and microfluidic tumor-on-chip devices sustaining patient spheroids or explants further demonstrate concordance with matched in vivo models, indicating translational relevance of transport-aware functional testing.198 Complementary in vivo work in pancreatic cancer shows that enzymatic hyaluronan depletion reduces interstitial fluid pressure, restores vascular perfusion and increases intratumoral drug accumulation, converting a poorly perfused, drug-refractory tumor into a treatment-responsive one.124,199,200
Together, these findings demonstrate that extrinsic resistance is not a fixed property but a modifiable consequence of transport limitation. ToC platforms therefore provide a functional complement to molecular biomarkers by revealing whether therapeutic agents can physically access their targets within the tumor microenvironment.
4.5 Scope and outlook
Despite these advances, clinical translation of transport-based phenotyping remains constrained by assay duration, technical complexity, and regulatory uncertainty. While organoids remain valuable for cell expansion, direct use of freshly resected tissue in ToC systems offers a more time-efficient route for assessing transport phenotypes, albeit with logistical challenges. Regulatory frameworks for physical, transport-based biomarkers are still less mature than those for molecular diagnostics, representing a key bottleneck for broader adoption.Nonetheless, the convergence of mechanistic modeling, microfluidic experimentation, and patient-derived systems provides a strong foundation for targeting stromal barriers as a therapeutic strategy. The following section examines how these insights are being translated into interventions that actively modulate CAF-driven matrix remodeling to improve drug delivery.
Extended discussion of clinical validation studies, regulatory considerations, and practical constraints is provided in sections S9–S13.
5. Therapeutic strategies: targeting the physical microenvironment
5.1 Matrix normalization and mechanical modulation approaches
Therapies targeting the tumor stroma rather than the tumor cells usually aim to restore transport-permissive properties while preserving the tissue architecture. Because excessive matrix ablation can paradoxically accelerate tumor progression by releasing mechanical growth constraints, most current strategies focus on matrix normalization or selective modulation rather than complete removal.13,201In this context, stromal targeting is best viewed to rebalance transport rather than to eliminate the extracellular matrix.
Beyond enzymatic degradation, several pharmacological agents have demonstrated unexpected anti-fibrotic effects. Losartan, an angiotensin II receptor blocker, suppresses the expression of profibrotic mediators such as insulin-like growth factor-1 (IGF1), TGFβ, CCL11 (eotaxin), MMP9, COL1A1, and thrombospondin (THBS2), thereby inhibiting pathological TGF-β-driven fibroblast activation.125 By reducing collagen synthesis and crosslinking while sparing normal stromal homeostasis, Losartan exemplifies a normalization strategy that selectively targets tumor-associated fibrosis. Other approaches modulate tissue mechanics without disrupting ECM architecture. Halofuginone, initially developed for autoimmune diseases, interferes with multiple matrix-related signaling pathways, including TGF-β/Smad3/Nrf2, Akt/mTORC1/Wnt/β-catenin, and HDAC2 signaling, reducing pathological matrix crosslinking and softening the stroma while preserving its structural integrity.202 Similarly, rho-associated protein kinase (ROCK) inhibitors reduce actomyosin contractility, relieving vessel compression, improving, and enhancing drug delivery.203,204 Together, these approaches demonstrate that redistributing mechanical stress can improve transport as effectively as bulk matrix degradation.
Despite these benefits, stromal modulation carries inherent risks. Excessive increases in vascular permeability may inadvertently facilitate tumor cell dissemination or compromise local containment.205 Consequently, patient stratification based on stromal features such as HA content or tissue stiffness, together with careful risk–benefit assessment, is essential for safe clinical translation.126
Additional stromal-modulation strategies are discussed in section S14.
5.2 Sequential therapy design
Sequential therapy strategies exploit the temporal dimension of stromal remodeling by conditioning the TME prior to cytotoxic or immunomodulatory treatment. Rather than co-administering all agents simultaneously, a first phase aims to reduce stromal resistance and improve transport, followed by a second phase that delivers the therapeutic payload under more permissive conditions. Preclinical studies have demonstrated that such conditioning significantly enhances intratumoral drug penetration and efficacy. For example, Zhang et al. (2024) designed a dendritic polymer system that initially delivered dasatinib-loaded nanoparticles to reprogram CAF metabolism and reduce collagen synthesis, followed by chemotherapeutic administration that achieved deeper penetration and greater tumor cell kill in the remodeled TME.32Sequential strategies are particularly compelling in immunotherapy. Reducing matrix stiffness or relieving physical barriers can restore CD8+ T-cells infiltration into tumor cores, effectively converting immune-excluded (cold) tumors into inflamed, therapy-responsive states. Preclinical studies have shown that stromal normalization enhances the efficacy of immune checkpoint blockade by facilitating T-cell access to previously-excluded regions.206,207 Inhibition of LOX/LOXL-mediated collagen crosslinking follows the same principle, improving CD8+ T-cell infiltration and sensitizing tumors to anti-PD-L1 therapy.208
These approaches highlight the importance of timing and context: the therapeutic window for the second, effector phase is determined not only by drug pharmacokinetics but also by the dynamics of stromal remodeling. Defining this window requires imaging-guided assessment of transport and mechanical properties, reinforcing the relevance of transport phenotyping in therapy design.
Expanded examples of sequential and conditioning therapies are provided in section S15.
5.3 Clinical trial landscape and physical biomarkers
Several stromal-targeting strategies have progressed to clinical trials, revealing both their therapeutic potential and their limitations (Table S4). In PDAC, PEGPH20 trials showed that patient stratification by HA content is essential: clinical benefit was restricted to HA-high tumors, underscoring the need for pre-treatment biomarker selection.126,209Functional imaging is increasingly central to identifying patients likely to benefit from transport-targeted therapies and to monitoring treatment response. Magnetic resonance elastography (MRE) provides a non-invasive measure of tumor stiffness and has been shown to predict or stratify neoadjuvant response in breast cancer cohorts.210 Dynamic contrast-enhanced MRI (DCE-MRI) offers complementary information on vascular function. The volume transfer constant (Ktrans) integrates blood flow, vascular permeability, and surface area, with reduced Ktrans values reflecting perfusion impairment caused by vessel compression or limited conductance. In PDAC, PEGPH20 treatment rapidly increased Ktrans, correlating with HA depletion, vascular decompression, and improved drug delivery.211–213
When applied longitudinally, these imaging modalities enable stratification of patients and response-adaptive therapy by aligning treatment decisions with improvements in perfusion or reductions in stiffness. However, broader clinical adoption remains limited by variability in imaging infrastructure, the absence of standardized physical biomarker thresholds, and safety concerns associated with stromal modulation. Extended clinical imaging and biomarker studies are summarized in section S16.
5.4 Integrating physical microenvironment targeting into precision oncology
Taken together, the strategies outlined above form a therapeutic continuum, ranging from matrix normalization and mechanical modulation to sequential conditioning protocols and imaging-guided adaptive therapies. Enzymatic degradation can induce rapid increases in permeability but requires stringent patient selection; mechanotherapeutics approaches achieve more gradual and reversible normalization; and sequential strategies exploit transient windows of enhanced accessibility to boost the efficacy of cytotoxic and immunotherapies.Successful clinical translation will depend on integrating robust physical biomarkers such as stiffness and perfusion into trial design and therapeutic decision-making. In this framework, physical microenvironment-targeted therapies complement molecular precision medicine by restoring drug accessibility and overcoming transport-limited resistance rather than by directly targeting tumor-cell-intrinsic vulnerabilities.
6. Critical assessment and future directions
6.1 Current consensus and open controversies
There is now broad agreement that fibroblast-driven ECM remodeling creates measurable transport limitations in desmoplastic tumors. Mechanistic, pathological, and biophysical studies converge on the view that increased matrix density and crosslinking elevate tissue stiffness while reducing intrinsic permeability, thereby limiting molecular and cellular access to malignant cells.3,31,182 These effects are particularly prominent in stroma-rich malignancies such as pancreatic ductal adenocarcinoma and selected breast and head-and-neck cancers, where the stromal compartment dominates tissue volume and mechanical behavior.Importantly, the transport barrier is neither homogeneous nor static. Instead, tumors exhibit a mosaic of regions ranging from relatively compliant to highly compacted and crosslinked matrices, whose microscale organization governs local transport. ECM composition, alignment, porosity, and crosslinking dynamically evolve in response to CAF activity, therapy, and perfusion. These parameters are experimentally accessible, and can be measured, modeled, and even perturbed using microfluidic ToC platforms and complementary in vivo approaches.192,214 Representative ongoing clinical trials that explicitly or implicitly target physical and stromal transport barriers are summarized in Table S4.
Despite this consensus, several mechanistic controversies remain with direct translational implications. First, interstitial flow can exert opposing effects depending on its magnitude, geometry, and the size of the carried agents. Modest interstitial flow may enhance the advective delivery of macromolecules, whereas elevated flow can reverse transcapillary gradients and impair the extravasation of most therapeutics.215,216 This duality explains why identical interventions may improve the delivery of one agent while reducing the efficacy of another. Second, stromal targeting strategies, including CAF ablation, reprogramming, enzymatic degradation, or mechanical modulation, cannot be applied uniformly. Their efficacy depends on CAF subtype composition, spatial organization, ECM composition, and disease stage.217 Consequently, stromal interventions must be matched to the tumor's biophysical phenotype, reinforcing the need for individualized, transport-informed strategies rather than one-size-fits-all approaches. These debates are not merely conceptual: they directly shape trial design, patient stratification, and therapeutic sequencing.
6.2 Methodological, translational, and reporting gaps
Progress in quantitatively linking matrix remodeling to transport physics represents a conceptual shift in understanding therapeutic resistance. It extends intervention strategies beyond molecular targets to the physical infrastructure of tumors, recognizing stromal-mediated transport barriers as resistance mechanisms equivalent in importance to intrinsic cellular ones.The causality framework linking fibroblast activation, ECM remodeling, and transport barrier formation provides mechanistic clarity for the design of rational interventions. By transforming descriptive observations into predictive models, this framework identifies optimal intervention points within pathological cascades and supports evidence-based sequencing and combination strategies targeting both physical and molecular resistance mechanisms. Microfluidic platforms have evolved from proof-of-concept tools into validated experimental systems with increasing clinical relevance. By recapitulating patient-specific tumor architectures under controlled conditions, they bridge the gap between reductionist in vitro assays and complex in vivo systems. This progress is illustrated by the growing integration of commercial ToC technologies, such as Emulate's Zoë-CM2® culture module,218,219 InSphero's Akura™ organ-on-chip,220,221 and MIMETAS's OrganoPlate®,222 into oncology pipelines, where they reproduce clinically relevant drug-response phenotypes more faithfully than conventional 2D assays.
Nevertheless, major gaps remain. Standardization of experimental protocols, reporting of transport parameters, and definition of clinically actionable contexts of use are still lacking. A step-by-step overview of practical workflow constraints, key experimental decision points, and transport-relevant quality controls in patient-derived tumor-on-chip studies is provided in Table S5. Biological studies must increasingly treat physical properties as mechanistically equivalent to molecular determinants of tumor behavior, while regulatory frameworks must evolve to accommodate physical biomarkers under validation standards comparable to genomic markers.
6.3 Outlook: from transport barriers to therapeutic vulnerabilities
Within the next decade, transport profiling could complement genomic characterization in clinical decision-making. Patient stratification would then integrate both molecular alterations and drug-accessibility profiles, enabling sequential strategies that normalize physical properties before targeted delivery. The translation of transport phenotyping from experimental platforms to routine clinical use will require coordinated progress across technology development, standardization, clinical validation, and implementation. A prospective, scenario-based roadmap outlining these interdependent milestones over the next decade is summarized in Fig. 3. Mechanical modulation, stromal reprogramming, and externally applied physical interventions may become standard components of multimodal therapy. In this context, computational and AI-assisted approaches may further support transport phenotyping by guiding experimental design, parameter exploration, and sequential intervention strategies. Representative examples of such optional frameworks and their minimal data requirements are summarized in Table S6.Beyond improving existing treatments, early intervention during matrix remodeling could prevent barrier formation altogether. Detection of pre-barrier states may enable prophylactic normalization strategies, eliminating malignancies before transport limitation arises. These principles extend beyond oncology to other matrix-mediated pathologies such as fibrosis, atherosclerosis, and neurodegeneration. For patients with transport-limited malignancies refractory to current therapies, integrating transport physics with precision oncology offers a promising therapeutic avenue. Physical barriers that currently protect tumors may ultimately become exploitable therapeutic vulnerabilities.
7. Concluding remarks
This review highlights mass-transport phenotyping as a unifying framework for understanding how stromal architecture, mechanical confinement, and interstitial fluid dynamics jointly regulate therapeutic accessibility in solid tumors. Rather than treating biochemical, mechanical, and transport parameters as independent features, tumor-on-chip platforms enable these dimensions to be made experimentally commensurate and quantitatively comparable, thereby complementing molecular and genomic profiling with a functional, biophysical perspective on drug accessibility. By articulating the conceptual links between stromal remodeling and transport barriers, and by outlining practical priorities for platform design and reporting, we aim to support the development of tumor-on-chip systems that are both biologically meaningful and experimentally interoperable. To translate these principles into actionable design criteria, Table 1 summarizes practical development directions for next-generation ToC platforms explicitly dedicated to mass-transport phenotyping.| Capability | Why it matters? | Metrics/readouts |
|---|---|---|
| Controlled modulation of mechanical confinement | Tests how solid stress affects drug/oxygen/immune access | Applied compressive stress (kPa), strain, ECM porosity, cell viability maps |
| Independent tuning of interstitial pressure and perfusion | Distinguishes advective vs. diffusive transport contributions | ΔP (Pa), perfusion flow rate (μL min−1), tracer velocity, perfusion fraction |
| ECM that supports physiologically relevant remodeling | Recapitulates desmoplasia and dynamic barrier formation | Time-course of ECM density, fiber alignment (SHG), crosslink markers, stiffness (kPa) |
| Non-invasive readouts of transport and mechanics | Enables longitudinal experiments without perturbation | FRAP, fluorescent tracer diffusion (Deff), microbead-based or elastography stiffness maps, pressure sensors |
| Device materials compatible with quantitative assays | Avoids artefactual adsorption or loss of analytes | Adsorption tests (percent recovery), surface treatment documented, control tracer mass balance |
| Compatibility with standard cell-culture workflows | Promotes adoption and reproducibility | Multi-well footprint, pipet access, common plate reader/microscope compatibility |
| Transport-centered reporting standards | Enables cross-platform, quantitative comparison | Minimum dataset: ECM stiffness, k, Deff, ΔP, tracer properties, device geometry, and access to the analysis code used to compute transport parameters |
Such platforms are not intended to replace existing experimental models, but rather to serve as a complementary analytical layer. By enabling mechanistic questions related to drug, nutrient, and immune-cell access to be addressed quantitatively, transport-aware ToC systems provide an intermediate scale between reductionist in vitro assays and complex in vivo models, where physical constraints often remain difficult to disentangle. Ultimately, adopting mass-transport-aware ToC approaches offers the opportunity to move beyond descriptive observations of treatment resistance toward a predictive, parameter-based understanding of tumor response.
This shift has the potential to refine experimental strategies, guide the rational design and sequencing of therapies, and improve the interpretability and reproducibility of preclinical studies across laboratories. For transport-limited malignancies that remain refractory to current treatments, integrating transport physics into precision oncology may help transform physical barriers from passive obstacles into actionable therapeutic vulnerabilities.
Glossary
Cancer-associated fibroblasts (CAFs): activated stromal fibroblasts within tumors that produce and remodel ECM. Heterogeneous population including matrix-producing (mpCAFs) and matrix-remodeling (mrCAFs) subtypes.Intrinsic permeability (k): a geometric property of a porous medium that quantifies its ability to transmit fluid independently of the fluid viscosity. It is expressed in m2 and depends on ECM porosity, pore size, fiber alignment, tortuosity, and matrix compaction. In tumors, k typically decreases by several orders of magnitude during CAF-driven desmoplastic remodeling, directly limiting interstitial flow and advective transport.
Fluid viscosity (μ): dynamic viscosity of the interstitial fluid (Pa s), assumed close to that of aqueous culture media and treated as constant in this review. Variations in advective transport are therefore attributed primarily to ECM-controlled changes in intrinsic permeability k.
Effective diffusivity (Deff): apparent diffusion coefficient of a solute within a tissue or ECM, accounting for porosity, tortuosity, binding, and confinement. Deff is lower than the free diffusivity in solution (D0) and dynamically evolves with matrix remodeling and solid stress.
Interstitial fluid pressure (IFP): pressure within the fluid-filled spaces between cells and matrix fibers. Elevated in tumors (1–8 kPa) compared to normal tissues (−1–+1 kPa), reducing transcapillary pressure gradients and limiting advective transport despite intact vasculature.31,223
Porosity (ϕ): fraction of tissue volume occupied by fluid-filled voids. In tumors, porosity decreases with ECM densification and compression, directly reducing diffusivity and intrinsic permeability.
Mechanotransduction: the process by which cells convert mechanical stimuli into biochemical signals. Key pathways involved include integrin clustering, focal adhesion kinase (FAK) activation, and the nuclear translocation of YAP/TAZ.
Solid stress: compressive and tensile forces generated within growing tumors as cells proliferate against surrounding tissue constraints. It can reach 1.3–13.0 kPa, sufficient to collapse blood vessels.164,184
Tortuosity: dimensionless parameter describing the geometric complexity of transport pathways in a porous medium. In biological tissues, higher tortuosity reflects elongated and convoluted diffusion or flow paths imposed by ECM architecture. It is commonly defined as 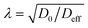 , where D0 is the free diffusivity and Deff the effective diffusivity in tissue.
, where D0 is the free diffusivity and Deff the effective diffusivity in tissue.
Mass-transport phenotyping: functional characterization of tumors based on quantitative transport properties (e.g., diffusivity, permeability, interstitial flow) that determine drug, nutrient, and immune-cell accessibility, independently of molecular target expression.
Tumor-on-chip (ToC): microfluidic in vitro platforms that recapitulate key aspects of tumor architecture, perfusion, and stromal remodeling under controlled conditions, enabling quantitative analysis of transport and mechanical phenomena.
Extrinsic (transport-limited) resistance: apparent therapeutic resistance arising from limited drug delivery due to physical barriers in the tumor microenvironment, despite intrinsic cellular sensitivity to the drug.
Author contributions
Doriane Le Manach: conceptualization, visualization, project administration, writing – original draft, writing – review & editing. Vincent Senez: conceptualization, funding acquisition, supervision, review & editing. Matthias Nees: funding acquisition, resources, supervision, review & editing.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, experimental data, software or code have been included, and no new data were generated or analyzed as part of this review.Supplementary information: Section S1 and Table S1 provide a comprehensive list of causal input parameters, transport-related outputs, symbols, units, and corresponding experimental readouts for CAF-driven transport phenotyping. (Sections 2.1 & 2.2); Sections S2 and S3 detail experimental, computational, and interventional strategies for calibrating transport parameters and modulating CAF-driven matrix barriers within transport-phenotyping frameworks. (Sections 2.3 & 2.4); Fig. S1 presents a hierarchical overview of tumor model systems used for mass-transport studies, positioning tumor-on-chip platforms relative to conventional in vitro and in vivo models. (Section 3.1); Section S4 and Table S2 provide a comparative overview of microfluidic device structural materials, surface functionalization strategies, and their implications for quantitative transport assays. (Sections 3.1 & 3.2); Sections S5 and S6, together with Table S3, summarize hydrogel and ECM choices, matrix stabilization strategies, and confinement approaches for CAF-driven transport phenotyping, including common experimental pitfalls. (Sections 3.2 & 3.3); Section S7 discusses measurement techniques, technical limitations, and multimodal strategies for quantifying diffusion, advection, and matrix remodeling in tumor-on-chip systems. (Section 3.4); Section S8 outlines recommended minimum reporting standards and reproducibility considerations for transport-resolved tumor-on-chip studies. (Section 3.5); Sections S9–S13 address practical, regulatory, and translational constraints for transport-based phenotyping, including representability, success rates, validation examples, and comparison with molecular biomarkers. (Section 4.5); Sections S14–S16 expand on stromal-targeting strategies, sequential therapy concepts, and imaging biomarkers relevant to transport-guided therapeutic interventions. (Sections 5.1, 5.2 & 5.3); Table S4 summarizes representative ongoing clinical trials targeting physical and stromal barriers to improve drug transport in solid tumors. (Section 6.1); Tables S5 and S6 detail workflow constraints for patient-derived tumor-on-chip transport phenotyping and optional AI-assisted experimental design strategies (Sections 6.2 & 6.3). See DOI: https://doi.org/10.1039/d5lc01131k.
Acknowledgements
This work was supported by the Polish National Science Centre (NCN): UMO-2020/37/B/N24/03920, and UMO-2021/41/B/NZ7/03786, the Cancéropole Nord-Ouest, the Région Hauts-de-France, the Centre National de la Recherche Scientifique (CNRS), the Ligue Nationale Contre le Cancer (France). Box 1 and Fig. 1 were created with https://BioRender.com.Notes and references
- D. Sun, C. Macedonia, Z. Chen, S. Chandrasekaran, K. Najarian, S. Zhou, T. Cernak, V. L. Ellingrod, H. V. Jagadish, B. Marini, M. Pai, A. Violi, J. C. Rech, S. Wang, Y. Li, B. Athey and G. S. Omenn, J. Med. Chem., 2024, 67, 16035–16055 CrossRef CAS PubMed.
- M. Hay, D. W. Thomas, J. L. Craighead, C. Economides and J. Rosenthal, Nat. Biotechnol., 2014, 32, 40–51 CrossRef CAS PubMed.
- A. I. Minchinton and I. F. Tannock, Nat. Rev. Cancer, 2006, 6, 583–592 CrossRef CAS PubMed.
- N. Vasan, J. Baselga and D. M. Hyman, Nature, 2019, 575, 299–309 CrossRef CAS PubMed.
- A. Soragni, E. S. Knudsen, T. N. O'Connor, C. E. Tognon, J. W. Tyner, B. Gini, D. Kim, T. G. Bivona, X. Zang, A. K. Witkiewicz, D. W. Goodrich, D. Jiang, S. T. Gammon, C. D. Willey, P. C. Boutros, V. C. Sandulache, A. A. Osman, J. N. Myers, K. Mehla, P. K. Singh, K. S. Chan, H. Gao and H. Marathe, Nat. Rev. Cancer, 2025, 25, 613–633 CrossRef CAS PubMed.
- M. Gerlinger, A. J. Rowan, S. Horswell, J. Larkin, D. Endesfelder, E. Gronroos, P. Martinez, N. Matthews, A. Stewart, P. Tarpey, I. Varela, B. Phillimore, S. Begum, N. Q. McDonald, A. Butler, D. Jones, K. Raine, C. Latimer, C. R. Santos, M. Nohadani, A. C. Eklund, B. Spencer-Dene, G. Clark, L. Pickering, G. Stamp, M. Gore, Z. Szallasi, J. Downward, P. A. Futreal and C. Swanton, N. Engl. J. Med., 2012, 366, 883–892 CrossRef CAS PubMed.
- P. Lu, V. M. Weaver and Z. Werb, J. Cell Biol., 2012, 196, 395–406 CrossRef CAS PubMed.
- M. Egeblad, M. G. Rasch and V. M. Weaver, Curr. Opin. Cell Biol., 2010, 22, 697–706 CrossRef CAS PubMed.
- G. Biffi, T. E. Oni, B. Spielman, Y. Hao, E. Elyada, Y. Park, J. Preall and D. A. Tuveson, Cancer Discovery, 2019, 9, 282–301 CrossRef PubMed.
- D. Öhlund, A. Handly-Santana, G. Biffi, E. Elyada, A. S. Almeida, M. Ponz-Sarvise, V. Corbo, T. E. Oni, S. A. Hearn, E. J. Lee, I. I. C. Chio, C.-I. Hwang, H. Tiriac, L. A. Baker, D. D. Engle, C. Feig, A. Kultti, M. Egeblad, D. T. Fearon, J. M. Crawford, H. Clevers, Y. Park and D. A. Tuveson, J. Exp. Med., 2017, 214, 579–596 CrossRef PubMed.
- R. Kalluri, Nat. Rev. Cancer, 2016, 16, 582–598 CrossRef CAS PubMed.
- P. A. Netti, D. A. Berk, M. A. Swartz, A. J. Grodzinsky and R. K. Jain, Cancer Res., 2000, 60, 2497–2503 CAS.
- R. K. Jain and T. Stylianopoulos, Nat. Rev. Clin. Oncol., 2010, 7, 653–664 CrossRef CAS PubMed.
- F. Di Maggio and K. H. El-Shakankery, Pancreas, 2020, 49, 313 CrossRef PubMed.
- K. Okuyama, M. Tsuchiya, K. C. Debnath, S. Islam and S. Yanamoto, Ther. Adv. Med. Oncol., 2025, 17, 17588359251317144 CrossRef CAS PubMed.
- S. Jenkins, M. E. Kachur, K. Rechache, J. M. Wells and S. Lipkowitz, Curr. Oncol. Rep., 2021, 23, 54 CrossRef PubMed.
- J. Kota, J. Hancock, J. Kwon and M. Korc, Cancer Lett., 2017, 391, 38–49 CrossRef CAS PubMed.
- M. A. Saad, W. Zhung, M. E. Stanley, S. Formica, S. Grimaldo-Garcia, G. Obaid and T. Hasan, Mol. Pharmaceutics, 2022, 19, 2549–2563 CrossRef CAS PubMed.
- C.-C. Lin and M. Korc, Cancer Lett., 2018, 436, 22–27 CrossRef CAS PubMed.
- P. P. Adiseshaiah, R. M. Crist, S. S. Hook and S. E. McNeil, Nat. Rev. Clin. Oncol., 2016, 13, 750–765 CrossRef CAS PubMed.
- G. S. Offeddu, E. Cambria, S. E. Shelton, K. Haase, Z. Wan, L. Possenti, H. T. Nguyen, M. R. Gillrie, D. Hickman, C. G. Knutson and R. D. Kamm, Adv. Sci., 2024, 11, 2402757 CrossRef PubMed.
- L. Miao, C. M. Lin and L. Huang, J. Controlled Release, 2015, 219, 192–204 CrossRef CAS PubMed.
- M. Binnewies, E. W. Roberts, K. Kersten, V. Chan, D. F. Fearon, M. Merad, L. M. Coussens, D. I. Gabrilovich, S. Ostrand-Rosenberg, C. C. Hedrick, R. H. Vonderheide, M. J. Pittet, R. K. Jain, W. Zou, T. K. Howcroft, E. C. Woodhouse, R. A. Weinberg and M. F. Krummel, Nat. Med., 2018, 24, 541–550 CrossRef CAS PubMed.
- Y. Ju, D. Xu, M. Liao, Y. Sun, W. Bao, F. Yao and L. Ma, npj Precis. Oncol., 2024, 8, 199 CrossRef PubMed.
- S. Azadi, H. Aboulkheyr Es, S. Razavi Bazaz, J. P. Thiery, M. Asadnia and M. Ebrahimi Warkiani, Biochim. Biophys. Acta, Mol. Cell Res., 2019, 1866, 118526 CrossRef CAS PubMed.
- C. Dextras, M. Dashnyam, L. A. M. Griner, J. Sundaresan, B. Chim, Z. Yu, S. Vodnala, C.-C. R. Lee, X. Hu, N. Southall, J. J. Marugan, A. Jadhav, N. P. Restifo, N. Acquavella, M. Ferrer and A. Singh, Sci. Rep., 2020, 10, 5688 CrossRef CAS PubMed.
- J. A. Joyce and D. T. Fearon, Science, 2015, 348, 74–80 CrossRef CAS PubMed.
- C. Feig, J. O. Jones, M. Kraman, R. J. B. Wells, A. Deonarine, D. S. Chan, C. M. Connell, E. W. Roberts, Q. Zhao, O. L. Caballero, S. A. Teichmann, T. Janowitz, D. I. Jodrell, D. A. Tuveson and D. T. Fearon, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 20212–20217 CrossRef CAS PubMed.
- T. Stylianopoulos, J. D. Martin, M. Snuderl, F. Mpekris, S. R. Jain and R. K. Jain, Cancer Res., 2013, 73, 3833–3841 CrossRef CAS PubMed.
- B. Blanco, H. Gomez, J. Melchor, R. Palma, J. Soler and G. Rus, Phys. Life Rev., 2023, 44, 279–301 CrossRef CAS PubMed.
- C.-H. Heldin, K. Rubin, K. Pietras and A. Östman, Nat. Rev. Cancer, 2004, 4, 806–813 CrossRef CAS PubMed.
- Y. Zhang, Z. Fang, D. Pan, Y. Li, J. Zhou, H. Chen, Z. Li, M. Zhu, C. Li, L. Qin, X. Ren, Q. Gong and K. Luo, Adv. Mater., 2024, 36, 2401304 CrossRef CAS PubMed.
- X. Zhang, X. Zhang, T. Yong, L. Gan and X. Yang, EBioMedicine, 2024, 105, 105200 CrossRef CAS PubMed.
- T. Stylianopoulos, L. L. Munn and R. K. Jain, Trends Cancer, 2018, 4, 292–319 CrossRef CAS PubMed.
- C. A. Horta, K. Doan and J. Yang, Curr. Opin. Cell Biol., 2023, 85, 102245 CrossRef CAS PubMed.
- T. Panciera, L. Azzolin, M. Cordenonsi and S. Piccolo, Nat. Rev. Mol. Cell Biol., 2017, 18, 758–770 CrossRef CAS PubMed.
- S. Piccolo, T. Panciera, P. Contessotto and M. Cordenonsi, Nat. Cancer, 2023, 4, 9–26 CAS.
- S. Dupont, L. Morsut, M. Aragona, E. Enzo, S. Giulitti, M. Cordenonsi, F. Zanconato, J. Le Digabel, M. Forcato, S. Bicciato, N. Elvassore and S. Piccolo, Nature, 2011, 474, 179–183 CrossRef CAS PubMed.
- T. Bertero and C. Gaggioli, Mol. Cell. Oncol., 2019, 6, 1592945 CrossRef PubMed.
- F. Kai, A. P. Drain and V. M. Weaver, Dev. Cell, 2019, 49, 332–346 CrossRef CAS PubMed.
- B. D. Matthews, C. K. Thodeti, J. D. Tytell, A. Mammoto, D. R. Overby and D. E. Ingber, Integr. Biol., 2010, 2, 435–442 CrossRef CAS PubMed.
- A. M. Goldyn, P. Kaiser, J. P. Spatz, C. Ballestrem, R. Kemkemer and P. Lappalainen, Cytoskeleton, 2010, 67, 241–250 CrossRef CAS PubMed.
- B. G. Kim, J. S. Sung, Y. Jang, Y. J. Cha, S. Kang, H. H. Han, J. H. Lee and N. H. Cho, Commun. Biol., 2019, 2, 313 CrossRef PubMed.
- M. R. Zanotelli, J. Zhang and C. A. Reinhart-King, Cell Metab., 2021, 33, 1307–1321 CrossRef CAS PubMed.
- T. R. Cox, Nat. Rev. Cancer, 2021, 21, 217–238 CrossRef CAS PubMed.
- Y. Yu, T. Zhou and L. Cao, J. Cell Commun. Signaling, 2023, 17, 1163–1179 CrossRef CAS PubMed.
- J. J. F. Sleeboom, H. Eslami Amirabadi, P. Nair, C. M. Sahlgren and J. M. J. den Toonder, Dis. Models Mech., 2018, 11, dmm033100 CrossRef PubMed.
- S.-Y. Jeong, J.-H. Lee, Y. Shin, S. Chung and H.-J. Kuh, PLoS One, 2016, 11, e0159013 CrossRef PubMed.
- Y. L. Huang, C. Tung, A. Zheng, B. J. Kim and M. Wu, Integr. Biol., 2015, 7, 1402–1411 CrossRef CAS PubMed.
- K. M. Lugo-Cintrón, M. M. Gong, J. M. Ayuso, L. A. Tomko, D. J. Beebe, M. Virumbrales-Muñoz and S. M. Ponik, Cancers, 2020, 12, 1173 CrossRef PubMed.
- I. Druzhkova, E. Nikonova, N. Ignatova, I. Koryakina, M. Zyuzin, A. Mozherov, D. Kozlov, D. Krylov, D. Kuznetsova, U. Lisitsa, V. Shcheslavskiy, E. A. Shirshin, E. Zagaynova and M. Shirmanova, Cancers, 2022, 14, 5487 CrossRef CAS PubMed.
- Q. Sun, S. H. Tan, Q. Chen, R. Ran, Y. Hui, D. Chen and C.-X. Zhao, ACS Biomater. Sci. Eng., 2018, 4, 4425–4433 CrossRef CAS PubMed.
- E. Wershof, D. Park, R. P. Jenkins, D. J. Barry, E. Sahai and P. A. Bates, PLoS Comput. Biol., 2019, 15, e1007251 CrossRef CAS PubMed.
- E. Elyada, M. Bolisetty, P. Laise, W. F. Flynn, E. T. Courtois, R. A. Burkhart, J. A. Teinor, P. Belleau, G. Biffi, M. S. Lucito, S. Sivajothi, T. D. Armstrong, D. D. Engle, K. H. Yu, Y. Hao, C. L. Wolfgang, Y. Park, J. Preall, E. M. Jaffee, A. Califano, P. Robson and D. A. Tuveson, Cancer Discovery, 2019, 9, 1102–1123 CrossRef CAS PubMed.
- E. Sahai, I. Astsaturov, E. Cukierman, D. G. DeNardo, M. Egeblad, R. M. Evans, D. Fearon, F. R. Greten, S. R. Hingorani, T. Hunter, R. O. Hynes, R. K. Jain, T. Janowitz, C. Jorgensen, A. C. Kimmelman, M. G. Kolonin, R. G. Maki, R. S. Powers, E. Puré, D. C. Ramirez, R. Scherz-Shouval, M. H. Sherman, S. Stewart, T. D. Tlsty, D. A. Tuveson, F. M. Watt, V. Weaver, A. T. Weeraratna and Z. Werb, Nat. Rev. Cancer, 2020, 20, 174–186 CrossRef CAS PubMed.
- C. Neuzillet, A. Tijeras-Raballand, C. Ragulan, J. Cros, Y. Patil, M. Martinet, M. Erkan, J. Kleeff, J. Wilson, M. Apte, M. Tosolini, A. S. Wilson, F. R. Delvecchio, C. Bousquet, V. Paradis, P. Hammel, A. Sadanandam and H. M. Kocher, J. Pathol., 2019, 248, 51–65 CrossRef CAS PubMed.
- Y. Xu, J. Li, J. Wang and F. Deng, J. Transl. Med., 2024, 22, 645 CrossRef CAS PubMed.
- C. Zhang, Y. Song, X. Cui, Y. Wang, J. Liu and Z. Shen, Sci. Rep., 2025, 15, 20533 CrossRef PubMed.
- G. Caligiuri and D. A. Tuveson, Cancer Cell, 2023, 41, 434–449 CrossRef CAS PubMed.
- Z. Liu, Y. Ba, D. Shan, X. Zhou, A. Zuo, Y. Zhang, H. Xu, S. Liu, B. Liu, Y. Zhao, S. Weng, R. Wang, J. Deng, P. Luo, Q. Cheng, X. Hu, S. Yang, F. Wang and X. Han, Cell Rep., 2025, 44(4) DOI:10.1016/j.celrep.2025.115555.
- R. Melchionna, P. Trono, A. Di Carlo, F. Di Modugno and P. Nisticò, J. Exp. Clin. Cancer Res., 2023, 42, 347 CrossRef CAS PubMed.
- L. Arpinati, G. Carradori and R. Scherz-Shouval, Nat. Rev. Cancer, 2024, 24, 676–693 CrossRef CAS PubMed.
- L. Cords, S. Engler, M. Haberecker, J. H. Rüschoff, H. Moch, N. de Souza and B. Bodenmiller, Cancer Cell, 2024, 42, 396–412.e5 CrossRef CAS PubMed.
- F. Pelon, B. Bourachot, Y. Kieffer, I. Magagna, F. Mermet-Meillon, I. Bonnet, A. Costa, A.-M. Givel, Y. Attieh, J. Barbazan, C. Bonneau, L. Fuhrmann, S. Descroix, D. Vignjevic, P. Silberzan, M. C. Parrini, A. Vincent-Salomon and F. Mechta-Grigoriou, Nat. Commun., 2020, 11, 404 CrossRef CAS PubMed.
- G. J. Laurent, R. C. Chambers, M. R. Hill and R. J. McAnulty, Biochem. Soc. Trans., 2007, 35, 647–651 CrossRef CAS PubMed.
- S. Knoedler, S. Broichhausen, R. Guo, R. Dai, L. Knoedler, M. Kauke-Navarro, F. Diatta, B. Pomahac, H.-G. Machens, D. Jiang and Y. Rinkevich, Front. Immunol., 2023, 14, 1233800 CrossRef CAS PubMed.
- P. P. Provenzano, D. R. Inman, K. W. Eliceiri, J. G. Knittel, L. Yan, C. T. Rueden, J. G. White and P. J. Keely, BMC Med., 2008, 6, 11 CrossRef PubMed.
- R. A. Gatenby and R. J. Gillies, Nat. Rev. Cancer, 2004, 4, 891–899 CrossRef CAS PubMed.
- P. Vaupel and L. Harrison, Oncologist, 2004, 9(Suppl 5), 4–9 CrossRef PubMed.
- N. A. Bhowmick, E. G. Neilson and H. L. Moses, Nature, 2004, 432, 332–337 CrossRef CAS PubMed.
- T. Casey, J. Bond, S. Tighe, T. Hunter, L. Lintault, O. Patel, J. Eneman, A. Crocker, J. White, J. Tessitore, M. Stanley, S. Harlow, D. Weaver, H. Muss and K. Plaut, Breast Cancer Res. Treat., 2009, 114, 47–62 CrossRef CAS PubMed.
- H. Ueno, A. M. Jones, K. H. Wilkinson, J. R. Jass and I. C. Talbot, Gut, 2004, 53, 581–586 CrossRef CAS PubMed.
- M. W. Conklin, J. C. Eickhoff, K. M. Riching, C. A. Pehlke, K. W. Eliceiri, P. P. Provenzano, A. Friedl and P. J. Keely, Am. J. Pathol., 2011, 178, 1221–1232 CrossRef PubMed.
- M. K. Sewell-Loftin, S. V. H. Bayer, E. Crist, T. Hughes, S. M. Joison, G. D. Longmore and S. C. George, Sci. Rep., 2017, 7, 12574 CrossRef PubMed.
- A. Orimo, P. B. Gupta, D. C. Sgroi, F. Arenzana-Seisdedos, T. Delaunay, R. Naeem, V. J. Carey, A. L. Richardson and R. A. Weinberg, Cell, 2005, 121, 335–348 CrossRef CAS PubMed.
- B. Hinz, G. Celetta, J. J. Tomasek, G. Gabbiani and C. Chaponnier, Mol. Biol. Cell, 2001, 12, 2730–2741 CrossRef CAS PubMed.
- J. Najera, M. R. Rosenberger and M. Datta, Cancers, 2023, 15, 3285 CrossRef CAS PubMed.
- R. L. Blackmon, R. Sandhu, B. S. Chapman, P. Casbas-Hernandez, J. B. Tracy, M. A. Troester and A. L. Oldenburg, Biophys. J., 2016, 110, 1858–1868 CrossRef CAS PubMed.
- A. Avendano, J. J. Chang, M. G. Cortes-Medina, A. J. Seibel, B. R. Admasu, C. M. Boutelle, A. R. Bushman, A. A. Garg, C. M. DeShetler, S. L. Cole and J. W. Song, ACS Biomater. Sci. Eng., 2020, 6, 1408–1417 CrossRef CAS PubMed.
- J. Sapudom, C. D. Müller, K.-T. Nguyen, S. Martin, U. Anderegg and T. Pompe, Gels, 2020, 6, 33 CrossRef CAS PubMed.
- T. Imamura, H. Iguchi, T. Manabe, G. Ohshio, T. Yoshimura, Z. H. Wang, H. Suwa, S. Ishigami and M. Imamura, Pancreas, 1995, 11, 357–364 CrossRef CAS PubMed.
- J. R. Staunton, B. L. Doss, S. Lindsay and R. Ros, Sci. Rep., 2016, 6, 19686 CrossRef CAS PubMed.
- C. S. Vidmar, M. Bazzi and V. K. Lai, J. Mech. Behav. Biomed. Mater., 2022, 128, 105107 CrossRef CAS PubMed.
- V. Serpooshan, M. Julien, O. Nguyen, H. Wang, A. Li, N. Muja, J. E. Henderson and S. N. Nazhat, Acta Biomater., 2010, 6, 3978–3987 CrossRef CAS PubMed.
- L. Cords, S. Tietscher, T. Anzeneder, C. Langwieder, M. Rees, N. de Souza and B. Bodenmiller, Nat. Commun., 2023, 14, 4294 CrossRef CAS PubMed.
- I. Calejo, M. A. Heinrich, G. Zambito, L. Mezzanotte, J. Prakash and L. Moreira Teixeira, Adv. Exp. Med. Biol., 2022, 1379, 171–203 CrossRef CAS PubMed.
- S. Chen, H. Qi, Y. Kuang, Q. Li, X. Chen and Y. Wang, Lab Chip, 2025, 25, 3858–3867 RSC.
- M. Soltani and P. Chen, PLoS One, 2013, 8, e67025 CrossRef CAS PubMed.
- T. Stylianopoulos and R. K. Jain, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 18632–18637 CrossRef CAS PubMed.
- F. Moradi Kashkooli, M. Soltani, M. M. Momeni and A. Rahmim, Front. Oncol., 2021, 11, 655781 CrossRef PubMed.
- R. K. Jain, Annu. Rev. Biomed. Eng., 1999, 1, 241–263 CrossRef CAS PubMed.
- R. K. Jain, J. D. Martin and T. Stylianopoulos, Annu. Rev. Biomed. Eng., 2014, 16, 321–346 CrossRef CAS PubMed.
- L. T. Baxter and R. K. Jain, Microvasc. Res., 1989, 37, 77–104 CrossRef CAS PubMed.
- R. K. Jain, Annu. Rev. Biomed. Eng., 1999, 1, 241–263 CrossRef CAS PubMed.
- P. A. Netti, L. T. Baxter, Y. Boucher, R. Skalak and R. K. Jain, Cancer Res., 1995, 55, 5451–5458 CAS.
- C. Nicholson and E. Syková, Trends Neurosci., 1998, 21, 207–215 CrossRef CAS PubMed.
- S. Ramanujan, A. Pluen, T. D. McKee, E. B. Brown, Y. Boucher and R. K. Jain, Biophys. J., 2002, 83, 1650 CrossRef CAS PubMed.
- G. A. Busby, M. H. Grant, S. P. Mackay and P. E. Riches, J. Biomech., 2013, 46, 837–840 CrossRef PubMed.
- A. G. Ogston, B. N. Preston and J. D. Wells, Proc. R. Soc. London, Ser., 1973, 333, 297–316 CAS.
- P. W. Sweeney, A. d'Esposito, S. Walker-Samuel and R. J. Shipley, PLoS Comput. Biol., 2019, 15, e1006751 CrossRef CAS PubMed.
- X. Zheng, K. Zhao, T. Jackson and J. Lowengrub, J. Math. Biol., 2022, 85, 5 CrossRef PubMed.
- C. Pozrikidis and D. A. Farrow, Ann. Biomed. Eng., 2003, 31, 181–194 CrossRef CAS PubMed.
- M. A. Biot, J. Appl. Phys., 1941, 12, 155–164 CrossRef.
- M. A. Biot, J. Appl. Phys., 1955, 26, 182–185 CrossRef CAS.
- W. Zhan, M. Alamer and X. Y. Xu, Adv. Drug Delivery Rev., 2018, 132, 81–103 CrossRef CAS PubMed.
- M. Soltani and P. Chen, J. Biol. Eng., 2012, 6, 4 CrossRef CAS PubMed.
- E. Axpe, D. Chan, G. S. Offeddu, Y. Chang, D. Merida, H. L. Hernandez and E. A. Appel, Macromolecules, 2019, 52, 6889–6897 CrossRef CAS PubMed.
- N. R. Richbourg and N. A. Peppas, Macromolecules, 2021, 54, 10477–10486 CrossRef CAS PubMed.
- M. H. Hettiaratchi, A. Schudel, T. Rouse, A. J. García, S. N. Thomas, R. E. Guldberg and T. C. McDevitt, APL Bioeng., 2018, 2, 026110 CrossRef PubMed.
- H. Haslene-Hox, E. Oveland, K. Woie, H. B. Salvesen, O. Tenstad and H. Wiig, Am. J. Physiol., 2015, 308, H18–H28 CAS.
- K. Burke and E. Brown, Intravital, 2015, 3, e984509 CrossRef PubMed.
- A. Avendano, J. J. Chang, M. G. Cortes-Medina, A. J. Seibel, B. R. Admasu, C. M. Boutelle, A. R. Bushman, A. A. Garg, C. M. DeShetler, S. L. Cole and J. W. Song, ACS Biomater. Sci. Eng., 2020, 6, 1408–1417 CrossRef CAS PubMed.
- C. S. Vidmar, M. Bazzi and V. K. Lai, J. Mech. Behav. Biomed. Mater., 2022, 128, 105107 CrossRef CAS PubMed.
- B. Dey and G. P. R. Sekhar, J. Theor. Biol., 2016, 395, 62–86 CrossRef CAS PubMed.
- M. Soltani and P. Chen, PLoS One, 2011, 6, e20344 CrossRef CAS PubMed.
- H. Ishikawa-Ankerhold, R. Ankerhold and G. Drummen, in eLS, John Wiley & Sons, Ltd, 2014 Search PubMed.
- J. Cacheux, J. Ordonez-Miranda, A. Bancaud, L. Jalabert, D. Alcaide, M. Nomura and Y. T. Matsunaga, Sci. Adv., 2023, 9, eadf9775 CrossRef CAS PubMed.
- T. Kihara, J. Ito and J. Miyake, PLoS One, 2013, 8, e82382 CrossRef PubMed.
- J. J. Mack, K. Youssef, O. D. V. Noel, M. P. Lake, A. Wu, M. L. Iruela-Arispe and L.-S. Bouchard, Biomaterials, 2013, 34, 1980–1986 CrossRef CAS PubMed.
- C. Gaggioli, S. Hooper, C. Hidalgo-Carcedo, R. Grosse, J. F. Marshall, K. Harrington and E. Sahai, Nat. Cell Biol., 2007, 9, 1392–1400 CrossRef CAS PubMed.
- Z. Rahman, A. Deep Bordoloi, H. Rouhana, M. Tavasso, G. van der Zon, V. Garbin, P. ten Dijke and P. E. Boukany, Lab Chip, 2024, 24, 422–433 RSC.
- K. R. Levental, H. Yu, L. Kass, J. N. Lakins, M. Egeblad, J. T. Erler, S. F. T. Fong, K. Csiszar, A. Giaccia, W. Weninger, M. Yamauchi, D. L. Gasser and V. M. Weaver, Cell, 2009, 139, 891–906 CrossRef CAS PubMed.
- H. Jiang, M. Zheng, X. Liu, S. Zhang, X. Wang, Y. Chen, M. Hou and J. Zhu, ACS Omega, 2019, 4, 12606–12615 CrossRef CAS PubMed.
- P. P. Provenzano, C. Cuevas, A. E. Chang, V. K. Goel, D. D. Von Hoff and S. R. Hingorani, Cancer Cell, 2012, 21, 418–429 CrossRef CAS PubMed.
- B. Diop-Frimpong, V. P. Chauhan, S. Krane, Y. Boucher and R. K. Jain, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 2909–2914 CrossRef CAS PubMed.
- S. R. Hingorani, L. Zheng, A. J. Bullock, T. E. Seery, W. P. Harris, D. S. Sigal, F. Braiteh, P. S. Ritch, M. M. Zalupski, N. Bahary, P. E. Oberstein, A. Wang-Gillam, W. Wu, D. Chondros, P. Jiang, S. Khelifa, J. Pu, C. Aldrich and A. E. Hendifar, J. Clin. Oncol., 2018, 36, 359–366 CrossRef CAS PubMed.
- J. Prakash and Y. Shaked, Cancer Discovery, 2024, 14, 1375–1388 CrossRef CAS PubMed.
- T. Chauhdari, S. A. Zaidi, J. Su and Y. Ding, In Vitro Models, 2025, 4, 71–88 CrossRef PubMed.
- L. E. Charelli, J. P. D. Ferreira, C. P. Naveira-Cotta and T. A. Balbino, J. Tissue Eng. Regener. Med., 2021, 15, 883–899 CrossRef CAS PubMed.
- L. Onfroy-Roy, D. Hamel, L. Malaquin and A. Ferrand, Cancers, 2021, 13, 1749 CrossRef CAS PubMed.
- R. Borst, L. Meyaard and M. I. Pascoal Ramos, J. Transl. Med., 2024, 22, 382 CrossRef PubMed.
- J. M. Lee, J. W. Choi, C. D. Ahrberg, H. W. Choi, J. H. Ha, S. G. Mun, S. J. Mo and B. G. Chung, Microsyst. Nanoeng., 2020, 6, 52 CrossRef CAS PubMed.
- S. I. Lee, Y. Y. Choi, S. G. Kang, T. H. Kim, J. W. Choi, Y. J. Kim, T.-H. Kim, T. Kang and B. G. Chung, Polymers, 2022, 14, 3752 CrossRef CAS PubMed.
- D. Kpeglo, M. Haddrick, M. A. Knowles, S. D. Evans and S. A. Peyman, Lab Chip, 2024, 24, 854–868 RSC.
- T. Meynard, F. Royer, R. Houssier, O. Bajeux, S. Paget, F. Lahdaoui, A. M. Valdivia, N. Maubon, J. Vicogne, I. V. Seuningen and V. Senez, Lab Chip, 2025, 25, 4119–4137 RSC.
- M. D. Mohan, N. Latifi, R. Flick, C. A. Simmons and E. W. K. Young, ACS Appl. Mater. Interfaces, 2024, 16, 20169–20185 CAS.
- J.-H. Lee, S.-K. Kim, I. A. Khawar, S.-Y. Jeong, S. Chung and H.-J. Kuh, J. Exp. Clin. Cancer Res., 2018, 37, 4 CrossRef PubMed.
- B. Kramer, L. de Haan, M. Vermeer, T. Olivier, T. Hankemeier, P. Vulto, J. Joore and H. L. Lanz, Int. J. Mol. Sci., 2019, 20, 4647 CrossRef CAS PubMed.
- F. L. Lai Benjamin, X. Lu Rick, Y. Hu, H. L. Davenport, W. Dou, E. Y. Wang, N. Radulovich, M. S. Tsao, Y. Sun and M. Radisic, Adv. Funct. Mater., 2020, 30, 2000545 CrossRef PubMed.
- M. Geyer, L.-M. Gaul, S. L. D. Agosto, V. Corbo and K. Queiroz, Front. Immunol., 2023, 14, 1155085 CrossRef CAS PubMed.
- M. Sefidgar, M. Soltani, K. Raahemifar, M. Sadeghi, H. Bazmara, M. Bazargan and M. Mousavi Naeenian, Microvasc. Res., 2015, 99, 43–56 CrossRef CAS PubMed.
- A. A. Rakhimov, A. A. Valiev, A. A. Akhmetov and K. V. Danilko, St. Petersburg Polytech. Univ. J. Phys. Math., 2023, 16(1.2), 253–259 Search PubMed.
- B. Wirthl, J. Kremheller, B. A. Schrefler and W. A. Wall, PLoS One, 2020, 15, e0228443 CrossRef CAS PubMed.
- T. Zhang, Y. Ren, P. Yang, J. Wang and H. Zhou, Cell Death Dis., 2022, 13, 897 CrossRef CAS PubMed.
- E. Bell, B. Ivarsson and C. Merrill, Proc. Natl. Acad. Sci. U. S. A., 1979, 76, 1274–1278 CrossRef CAS PubMed.
- F. Calvo, N. Ege, A. Grande-Garcia, S. Hooper, R. P. Jenkins, S. I. Chaudhry, K. Harrington, P. Williamson, E. Moeendarbary, G. Charras and E. Sahai, Nat. Cell Biol., 2013, 15(6), 637–646 CrossRef CAS PubMed.
- M. E. Smithmyer, L. A. Sawicki and A. M. Kloxin, Biomater. Sci., 2014, 2, 634–650 RSC.
- M. Horie, A. Saito, Y. Yamaguchi, M. Ohshima and T. Nagase, J. Visualized Exp., 2015, 52469 Search PubMed.
- J. Li, C. Dong and J. Ren, TrAC, Trends Anal. Chem., 2017, 89, 181–189 CrossRef CAS.
- M. Kawai, H. Higuchi, M. Takeda, Y. Kobayashi and N. Ohuchi, Breast Cancer Res., 2009, 11, R43 CrossRef PubMed.
- X. Chen, O. Nadiarynkh, S. Plotnikov and P. J. Campagnola, Nat. Protoc., 2012, 7, 654–669 CrossRef CAS PubMed.
- E. S. Hwang, D. J. Morgan, J. Sun, M. E. Hartnett, K. C. Toussaint and B. Coats, Biomed. Opt. Express, 2023, 14, 932–944 CrossRef PubMed.
- M. Zhu, K. Zhang, E. C. Thomas, R. Xu, B. Ciruna, S. Hopyan and Y. Sun, Sci. Adv., 2025, 11, eadt7274 CrossRef PubMed.
- T. Sun, H. Zhao, L. Hu, X. Shao, Z. Lu, Y. Wang, P. Ling, Y. Li, K. Zeng and Q. Chen, Acta Pharm. Sin. B, 2024, 14, 2428–2446 CrossRef CAS PubMed.
- D. K. Logsdon, G. F. Beeghly and J. M. Munson, Cell. Mol. Bioeng., 2017, 10, 463–481 CrossRef CAS PubMed.
- K. Seaman, Y. Sun and L. You, Med-X, 2023, 1, 11 CrossRef CAS.
- A. Sontheimer-Phelps, B. A. Hassell and D. E. Ingber, Nat. Rev. Cancer, 2019, 19, 65–81 CrossRef CAS PubMed.
- P. S. W. Cheng, M. Zaccaria and G. Biffi, Trends Cancer, 2025, 11, 135–153 CrossRef CAS PubMed.
- A. La Rocca, V. De Gregorio, E. Lagreca, R. Vecchione, P. A. Netti and G. Imparato, Int. J. Mol. Sci., 2023, 24, 5678 CrossRef CAS PubMed.
- E. Steinberg, R. Friedman, Y. Goldstein, N. Friedman, O. Beharier, J. A. Demma, G. Zamir, A. Hubert and O. Benny, Commun. Biol., 2023, 6, 1157 CrossRef PubMed.
- T. Miti, B. Desai, D. Miroshnychenko, D. Basanta and A. Marusyk, Cancers, 2024, 16, 2405 CrossRef CAS PubMed.
- M. A. Winkelman, D. Y. Kim, S. Kakarla, A. Grath, N. Silvia and G. Dai, Lab Chip, 2022, 22, 170–192 RSC.
- R. K. Jain, Sci. Am., 1994, 271, 58–65 CrossRef CAS PubMed.
- T. Stylianopoulos, J. D. Martin, V. P. Chauhan, S. R. Jain, B. Diop-Frimpong, N. Bardeesy, B. L. Smith, C. R. Ferrone, F. J. Hornicek, Y. Boucher, L. L. Munn and R. K. Jain, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 15101–15108 CrossRef CAS PubMed.
- S. Pal, E. Shimshoni, S. F. Torres, M. Kong, K. Tai, V. Sangwan, N. Bertos, S. D. Bailey, J. Bérubé, D. E. Ingber and L. Ferri, J. Transl. Med., 2025, 23, 577 CrossRef CAS PubMed.
- S. Chakrabarty, W. F. Quiros-Solano, M. M. P. Kuijten, B. Haspels, S. Mallya, C. S. Y. Lo, A. Othman, C. Silvestri, A. van de Stolpe, N. Gaio, H. Odijk, M. van de Ven, C. M. A. de Ridder, W. M. van Weerden, J. Jonkers, R. Dekker, N. Taneja, R. Kanaar and D. C. van Gent, Cancer Res., 2022, 82, 510–520 CrossRef CAS PubMed.
- T. Song, H. Zhang, Z. Luo, L. Shang and Y. Zhao, Adv. Sci., 2023, 10, e2206004 CrossRef PubMed.
- T. Petreus, E. Cadogan, G. Hughes, A. Smith, V. Pilla Reddy, A. Lau, M. J. O'Connor, S. Critchlow, M. Ashford and L. Oplustil O'Connor, Commun. Biol., 2021, 4, 1001 CrossRef CAS PubMed.
- B. J. Toley, Z. T. Lovatt, J. L. Harrington and N. S. Forbes, Integr. Biol., 2013, 5, 1184–1196 CrossRef CAS PubMed.
- L. R. Avula and P. Grodzinski, Front. Lab Chip Technol., 2024, 3 DOI:10.3389/frlct.2024.1487377.
- N. Isoherranen, R. Madabushi and S.-M. Huang, Clin. Transl. Sci., 2019, 12, 113–121 CrossRef PubMed.
- T. Tan, D. Mouradov, P. Gibbs and O. M. Sieber, STAR Protoc., 2024, 5, 103090 CrossRef CAS PubMed.
- S. Bose, H. Clevers and X. Shen, Med, 2021, 2, 1011–1026 CrossRef PubMed.
- X. Li, H. E. Francies, M. Secrier, J. Perner, A. Miremadi, N. Galeano-Dalmau, W. J. Barendt, L. Letchford, G. M. Leyden, E. K. Goffin, A. Barthorpe, H. Lightfoot, E. Chen, J. Gilbert, A. Noorani, G. Devonshire, L. Bower, A. Grantham, S. MacRae, N. Grehan, D. C. Wedge, R. C. Fitzgerald and M. J. Garnett, Nat. Commun., 2018, 9, 2983 CrossRef PubMed.
- K.-L. Jiang, X.-X. Wang, X.-J. Liu, L.-K. Guo, Y.-Q. Chen, Q.-L. Jia, K.-M. Yang and J.-H. Ling, World J. Gastrointest. Oncol., 2024, 16, 1626–1646 CrossRef PubMed.
- N. T. Elliott and F. Yuan, Biotechnol. Bioeng., 2012, 109, 1326–1335 CrossRef CAS PubMed.
- S. Pradhan, A. M. Smith, C. J. Garson, I. Hassani, W. J. Seeto, K. Pant, R. D. Arnold, B. Prabhakarpandian and E. A. Lipke, Sci. Rep., 2018, 8, 3171 CrossRef PubMed.
- Y. Zhao, Y. Wu, K. Islam, R. Paul, Y. Zhou, X. Qin, Q. Li and Y. Liu, ACS Appl. Mater. Interfaces, 2024, 16, 22839–22849 CAS.
- L. A. Shuman Moss, S. Jensen-Taubman and W. G. Stetler-Stevenson, Am. J. Pathol., 2012, 181, 1895–1899 CrossRef PubMed.
- V. P. Chauhan, T. Stylianopoulos, Y. Boucher and R. K. Jain, Annu. Rev. Chem. Biomol. Eng., 2011, 2, 281–298 CrossRef CAS PubMed.
- D. M. Gilkes, G. L. Semenza and D. Wirtz, Nat. Rev. Cancer, 2014, 14, 430–439 CrossRef CAS PubMed.
- Z. Yuan, Y. Li, S. Zhang, X. Wang, H. Dou, X. Yu, Z. Zhang, S. Yang and M. Xiao, Mol. Cancer, 2023, 22, 48 CrossRef CAS PubMed.
- S. Ricard-Blum and S. D. Vallet, Front. Pharmacol., 2016, 7, 11 Search PubMed.
- H. T. Nia, H. Liu, G. Seano, M. Datta, D. Jones, N. Rahbari, J. Incio, V. P. Chauhan, K. Jung, J. D. Martin, V. Askoxylakis, T. P. Padera, D. Fukumura, Y. Boucher, F. J. Hornicek, A. J. Grodzinsky, J. W. Baish, L. L. Munn and R. K. Jain, Nat. Biomed. Eng., 2016, 1, 0004 CrossRef PubMed.
- C. Wong, T. Stylianopoulos, J. Cui, J. Martin, V. P. Chauhan, W. Jiang, Z. Popović, R. K. Jain, M. G. Bawendi and D. Fukumura, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 2426–2431 CrossRef CAS PubMed.
- M. Kalli and T. Stylianopoulos, Front. Oncol., 2018, 8, 55 CrossRef PubMed.
- F. Bordeleau, B. N. Mason, E. M. Lollis, M. Mazzola, M. R. Zanotelli, S. Somasegar, J. P. Califano, C. Montague, D. J. LaValley, J. Huynh, N. Mencia-Trinchant, Y. L. Negrón Abril, D. C. Hassane, L. J. Bonassar, J. T. Butcher, R. S. Weiss and C. A. Reinhart-King, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 492–497 CrossRef CAS PubMed.
- J. Barbazan, C. Pérez-González, M. Gómez-González, M. Dedenon, S. Richon, E. Latorre, M. Serra, P. Mariani, S. Descroix, P. Sens, X. Trepat and D. M. Vignjevic, Nat. Commun., 2023, 14, 6966 CrossRef CAS PubMed.
- J. Winkler, A. Abisoye-Ogunniyan, K. J. Metcalf and Z. Werb, Nat. Commun., 2020, 11, 5120 CrossRef CAS PubMed.
- P. Tenti and L. Vannucci, Cancer Immunol., Immunother., 2019, 69, 223–235 CrossRef PubMed.
- A. Chakravarthy, L. Khan, N. P. Bensler, P. Bose and D. D. De Carvalho, Nat. Commun., 2018, 9, 4692 CrossRef PubMed.
- J. M. Munson and A. C. Shieh, Cancer Manage. Res., 2014, 6, 317–328 CrossRef PubMed.
- S. Schuth, S. Le Blanc, T. G. Krieger, J. Jabs, M. Schenk, N. A. Giese, M. W. Büchler, R. Eils, C. Conrad and O. Strobel, J. Exp. Clin. Cancer Res., 2022, 41, 312 CrossRef CAS PubMed.
- A. Pluen, Y. Boucher, S. Ramanujan, T. D. McKee, T. Gohongi, E. di Tomaso, E. B. Brown, Y. Izumi, R. B. Campbell, D. A. Berk and R. K. Jain, Proc. Natl. Acad. Sci. U. S. A., 2001, 98, 4628–4633 CrossRef CAS PubMed.
- J. C. Pease, M. Brewer and J. S. Tirnauer, Biol. Open, 2012, 1, 622–628 CrossRef CAS PubMed.
- L. F. Horowitz, A. D. Rodriguez, Z. Dereli-Korkut, R. Lin, K. Castro, A. M. Mikheev, R. J. Monnat, A. Folch and R. C. Rostomily, npj Precis. Oncol., 2020, 4, 12 CrossRef CAS PubMed.
- M. R. Haque, C. R. Wessel, D. D. Leary, C. Wang, A. Bhushan and F. Bishehsari, Microsyst. Nanoeng., 2022, 8, 36 CrossRef PubMed.
- E. Steinberg, R. Friedman, Y. Goldstein, N. Friedman, O. Beharier, J. A. Demma, G. Zamir, A. Hubert and O. Benny, Commun. Biol., 2023, 6, 1157 CrossRef PubMed.
- A. D. Rodriguez, L. F. Horowitz, K. Castro, H. Kenerson, N. Bhattacharjee, G. Gandhe, A. Raman, R. J. Monnat, R. Yeung, R. C. Rostomily and A. Folch, Lab Chip, 2020, 20, 1658–1675 RSC.
- I. Veith, M. Nurmik, A. Mencattini, I. Damei, C. Lansche, S. Brosseau, G. Gropplero, S. Corgnac, J. Filippi, N. Poté, E. Guenzi, A. Chassac, P. Mordant, J. Tosello, C. Sedlik, E. Piaggio, N. Girard, J. Camonis, H. Shirvani, F. Mami-Chouaib, F. Mechta-Grigoriou, S. Descroix, E. Martinelli, G. Zalcman and M. C. Parrini, Cell Rep. Med., 2024, 5, 101549 CrossRef CAS PubMed.
- M. Magzoub, S. Jin and A. S. Verkman, FASEB J., 2008, 22, 276–284 CrossRef CAS PubMed.
- W. Li, Y. Wu, Y. Zhang, W. Gao, X. Li, H. Luo, M. Lu, Z. Liu and A. Luo, Adv. Sci., 2025, 12, 2416523 CrossRef CAS PubMed.
- M. J. Paszek, N. Zahir, K. R. Johnson, J. N. Lakins, G. I. Rozenberg, A. Gefen, C. A. Reinhart-King, S. S. Margulies, M. Dembo, D. Boettiger, D. A. Hammer and V. M. Weaver, Cancer Cell, 2005, 8, 241–254 CrossRef CAS PubMed.
- C. Vennin, V. T. Chin, S. C. Warren, M. C. Lucas, D. Herrmann, A. Magenau, P. Melenec, S. N. Walters, G. Del Monte-Nieto, J. R. W. Conway, M. Nobis, A. H. Allam, R. A. McCloy, N. Currey, M. Pinese, A. Boulghourjian, A. Zaratzian, A. A. S. Adam, C. Heu, A. M. Nagrial, A. Chou, A. Steinmann, A. Drury, D. Froio, M. Giry-Laterriere, N. L. E. Harris, T. Phan, R. Jain, W. Weninger, E. J. McGhee, R. Whan, A. L. Johns, J. S. Samra, L. Chantrill, A. J. Gill, M. Kohonen-Corish, R. P. Harvey, A. V. Biankin, Australian Pancreatic Cancer Genome Initiative (APGI), T. R. J. Evans, K. I. Anderson, S. T. Grey, C. J. Ormandy, D. Gallego-Ortega, Y. Wang, M. S. Samuel, O. J. Sansom, A. Burgess, T. R. Cox, J. P. Morton, M. Pajic and P. Timpson, Sci. Transl. Med., 2017, 9, eaai8504 CrossRef PubMed.
- T. Tomita, M. Kato and S. Hiratsuka, Cancer Sci., 2021, 112, 2966–2974 CrossRef CAS PubMed.
- J. A. Grout, P. Sirven, A. M. Leader, S. Maskey, E. Hector, I. Puisieux, F. Steffan, E. Cheng, N. Tung, M. Maurin, R. Vaineau, L. Karpf, M. Plaud, A.-L. Bègue, K. Ganesh, J. Mesple, M. Casanova-Acebes, A. Tabachnikova, S. Keerthivasan, A. Lansky, J. L. Bérichel, L. Walker, A. H. Rahman, S. Gnjatic, N. Girard, M. Lefèvre, D. Damotte, J. Adam, J. C. Martin, A. Wolf, R. M. Flores, M. B. Beasley, R. Pradhan, S. Müller, T. U. Marron, S. J. Turley, M. Merad, E. Kenigsberg and H. Salmon, Cancer Discovery, 2022, 12, 2606–2625 CrossRef CAS PubMed.
- L. Jenkins, U. Jungwirth, A. Avgustinova, M. Iravani, A. Mills, S. Haider, J. Harper and C. M. Isacke, Cancer Res., 2022, 82, 2904–2917 CrossRef CAS PubMed.
- D. H. Peng, B. L. Rodriguez, L. Diao, L. Chen, J. Wang, L. A. Byers, Y. Wei, H. A. Chapman, M. Yamauchi, C. Behrens, G. Raso, L. M. S. Soto, E. R. P. Cuentes, I. I. Wistuba, J. M. Kurie and D. L. Gibbons, Nat. Commun., 2020, 11, 4520 CrossRef CAS PubMed.
- C. B. Thompson, H. M. Shepard, P. M. O'Connor, S. Kadhim, P. Jiang, R. J. Osgood, L. H. Bookbinder, X. Li, B. J. Sugarman, R. J. Connor, S. Nadjsombati and G. I. Frost, Mol. Cancer Ther., 2010, 9, 3052–3064 CrossRef CAS PubMed.
- A. P. Sinha, P. Jurrius, A.-S. van Schelt, O. Darwish, B. Shifa, G. Annio, Z. Peterson, H. Jeffery, K. Welsh, A. Metafa, J. Spence, A. Kothari, H. Hamed, G. Bitsakou, V. Karydakis, M. Thorat, E. Shaari, A. Sever, A. Rigg, T. Ng, S. Pinder, R. Sinkus and A. Purushotham, Radiol. Imaging Cancer, 2025, 7, e240138 CrossRef PubMed.
- J. Cao, S. Pickup, C. Clendenin, B. Blouw, H. Choi, D. Kang, M. Rosen, P. J. O'Dwyer and R. Zhou, Clin. Cancer Res., 2019, 25, 2314–2322 CrossRef CAS PubMed.
- J. Xiao, H. Rahbar, D. S. Hippe, M. H. Rendi, E. U. Parker, N. Shekar, M. Hirano, K. J. Cheung and S. C. Partridge, npj Breast Cancer, 2021, 7, 42 CrossRef CAS PubMed.
- X. Yin, P. Liu, C. Zhang, C. Pan, Y. Lu, Z. Wang, B. Luo, B. Du, A. Yang, Q. Wu, C. Gu and Y. Shi, J. Magn. Reson. Imaging, 2025, 62(6), 1826–1840 CrossRef PubMed.
- A. Avendano, M. Cortes-Medina and J. W. Song, Front. Bioeng. Biotechnol., 2019, 7 DOI:10.3389/fbioe.2019.00006.
- M. Welter and H. Rieger, PLoS One, 2013, 8, e70395 CrossRef CAS PubMed.
- M. Wu, H. B. Frieboes, S. R. McDougall, M. A. J. Chaplain, V. Cristini and J. Lowengrub, J. Theor. Biol., 2013, 320, 131–151 CrossRef PubMed.
- Z. Cao, S. Quazi, S. Arora, L. D. Osellame, I. J. Burvenich, P. W. Janes and A. M. Scott, J. Biomed. Sci., 2025, 32, 7 CrossRef PubMed.
- Solid Tumor Cell Therapy Assessment with Organ-Chips, https://emulatebio.com/solid-tumor-cell-therapy-organ-chips/, (accessed 23 September 2025).
- C. Strelez, S. Chilakala, K. Ghaffarian, R. Lau, E. Spiller, N. Ung, D. Hixon, A. Y. Yoon, R. X. Sun, H.-J. Lenz, J. E. Katz and S. M. Mumenthaler, iScience, 2021, 24, 102509 CrossRef CAS PubMed.
- O. T. P. Nguyen, P. M. Misun, C. Lohasz, J. Lee, W. Wang, T. Schroeder and A. Hierlemann, Front. Immunol., 2021, 12, 781337 CrossRef CAS PubMed.
- Akura™ Organ-on-Chip Technology for In Vitro Assays, https://insphero.com/technology/akura-organ-on-chip-technology/, (accessed 23 September 2025).
- K. M. Wisdom, J. Suijker, L. Van den Broek, B. Sridharan, T. S. P. Grandhi, A. Cheng, M. Lamb, S. A. Titus, D. Poore, N. Shah, S.-H. Cheng, E. Kim, S. Griffin and J. Ekert, ALTEX, 2023, 40, 649–664 Search PubMed.
- R. K. Jain, Cancer Res., 1988, 48, 2641–2658 CAS.
Footnotes |
| † These authors contributed equally. |
| ‡ Hereafter, the term “transport” refers to molecular movement within TME. |
| This journal is © The Royal Society of Chemistry 2026 |