Ultra-high throughput droplet microfluidics for cultivation and functional screening of environmental microbial strains and consortia
Luca
Potenza
,
Jozef
Krzak
,
Maciej S.
Andrzejewski
,
Adam
Pyzik
 and
Tomasz S.
Kaminski
and
Tomasz S.
Kaminski
 *
*
Department of Molecular Biology, Institute of Biochemistry, Faculty of Biology, University of Warsaw, Miecznikowa 1, 02-096 Warsaw, Poland. E-mail: ts.kaminski2@uw.edu.pl
First published on 15th April 2026
Abstract
Screening of microbial diversity is critical for discovering novel strains with enhanced biocatalytic capabilities. Traditional cultivation techniques often miss rare or slow-growing microorganisms, limiting our understanding of environmental microbiomes and their practical applications. Droplet-based microfluidics has emerged as a powerful platform for ultra-high-throughput screening of single cells or microbial consortia, encapsulated within microscale droplets that act as precisely controlled bioreactors. This tutorial review focuses on the application of droplet microfluidics for microbial cultivation and isolation, emphasizing advantages such as high throughput and the ability to perform functional assays on cultures originated from individual cells. Unlike previous reviews that broadly address microfluidic technologies or target specific applications like antibiotic resistance or enzyme discovery, we concentrate on protocols specifically designed for the clonal and parallel cultivation of microorganisms from environmental samples. Droplet microfluidics has already shown significant potential in environmental biotechnology, bioremediation, and microbial ecology. Reducing assay volumes and costs while increasing screening efficiency, positions droplet microfluidics as a key technology for the future of environmental microbiology research.
I. Introduction
Microbial strains and consortia may possess unique biochemical properties with valuable applications yet often remain undetected by conventional cultivation and screening techniques. Precise screening is essential not only for exploring biodiversity but also for identifying strains with desirable traits that can be harnessed across various biotechnology sectors, such as pollutant degradation and the production of enzymes or bioactive molecules.1,2 A promising alternative has emerged from recent advancements in droplet microfluidic screening technologies.3,4 These systems enable the execution of complex protocols within stable emulsion microbioreactors, significantly reducing both time and cost per screening campaign. Droplet-based assays present clear advantages over traditional methods by allowing the encapsulation, incubation, sorting, and analysis of cultures originating from individual cells within microscale bioreactors.5–7 This approach enables the isolation of strains based on growth dynamics8,9 or biocatalytic activity,10 with one of its most significant benefits being the ability to recover rare or slow-growing taxa that are difficult to enrich using classical cultivation techniques.8,11,12Characterizing complex microbial consortia from unknown environmental samples - such as those derived from soil, water, or air – remains a significant challenge, even with modern screening technologies. For example, a single gram of soil can contain up to 1010 bacterial cells and an estimated species diversity ranging from 4 × 103 to 5 × 104 species.13,14 Accurately representing the entire population and diversity of such samples without introducing cultivation bias is central to the so-called uncultured microbiome challenge. This term refers to the substantial fraction of microorganisms that cannot be isolated using classical laboratory techniques,15 such as cultivation on selective media in Petri dishes. Technically biased taxonomic representation and the frequent loss of highly active strains significantly limit our understanding of microbial diversity, function, and ecological interactions. Addressing this challenge requires methodological innovations, including the application of droplet-based ultrahigh-throughput screening (uHTS) assays.
Recent technological advancements in droplet-based microfluidics, including novel detection modalities and increased automation, have significantly enhanced its utility as screening platforms.16 Ultra-high-throughput screening (uHTS) in droplets is increasingly relevant across diverse areas of modern biotechnology, with applications ranging from the functional screening of microbial activity17,18 to diagnostics,19 directed enzyme evolution,20 and drug discovery.10,21 This review specifically focuses on the use of droplets as bioreactors for clonal cultivation and functional characterization of microorganisms. As a novel extension of uHTS, droplet microfluidics has been widely recognized for its growing role in microbiological research.22
In this context, uHTS refers to screening capacities exceeding 105 to 107 discrete units per assay, typically characterized by droplet generation or sorting rates in the kilohertz (kHz) range. However, a critical distinction must be made between nominal throughput, the raw speed of droplet formation, and effective throughput, which accounts for sorting constraints, and downstream recovery. While droplet generation rates can readily exceed 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 droplets per second (10 kHz), the practical decision rate in active sorting systems (e.g. fluorescence-activated droplet sorting, FADS) is often constrained by the physical ability to deflect droplets within the flow and in some cases by sensor integration time. As a result, high-accuracy sorting typically operates within a range of ∼0.1 to 2 kHz.23,24 Although sorting rates approaching 30 kHz have been demonstrated, these are generally limited to very small droplets and may not be broadly applicable. Additionally, sorting throughputs of 10 kHz or higher can be achieved using fluorescence-activated cell sorting (FACS) when working with double emulsions, microgels, or microcapsule formats. However, the effective number of screened units per unit time must also consider the Poisson distribution governing cell encapsulation. To ensure predominantly single-cell occupancy, a significant fraction of droplets remains empty, thereby reducing biological throughput relative to the technical droplet rate.25,26 Furthermore, practical throughput is further reduced by incubation times and the operational delays associated with transferring samples between sequential devices. Therefore, a comprehensive evaluation of microfluidic platform performance must consider key parameters such as droplet generation rate and sorting efficiency.
000 droplets per second (10 kHz), the practical decision rate in active sorting systems (e.g. fluorescence-activated droplet sorting, FADS) is often constrained by the physical ability to deflect droplets within the flow and in some cases by sensor integration time. As a result, high-accuracy sorting typically operates within a range of ∼0.1 to 2 kHz.23,24 Although sorting rates approaching 30 kHz have been demonstrated, these are generally limited to very small droplets and may not be broadly applicable. Additionally, sorting throughputs of 10 kHz or higher can be achieved using fluorescence-activated cell sorting (FACS) when working with double emulsions, microgels, or microcapsule formats. However, the effective number of screened units per unit time must also consider the Poisson distribution governing cell encapsulation. To ensure predominantly single-cell occupancy, a significant fraction of droplets remains empty, thereby reducing biological throughput relative to the technical droplet rate.25,26 Furthermore, practical throughput is further reduced by incubation times and the operational delays associated with transferring samples between sequential devices. Therefore, a comprehensive evaluation of microfluidic platform performance must consider key parameters such as droplet generation rate and sorting efficiency.
In this tutorial review, we present a contemporary and comprehensive summary of various microfluidic systems designed for selecting microbial colonies encapsulated in droplets based on their activity. Unlike other recent microfluidic reviews, which often emphasize general technical challenges and developments in droplet-based systems16,27 or focus on specific applications such as antimicrobial susceptibility testing,28 enzyme engineering,20,29 or the human microbiome,30 we specifically detail protocols that enable clonal and parallel cultivation of environmental microorganisms, often followed by functional selection. Considering the immense diversity of microbial communities, we argue that droplet microfluidic platforms are indispensable for investigating hard-to-cultivate species. In this context, single-cell droplet microfluidic methods are particularly valuable, as they enable precise screening of complex and heterogeneous microbial samples.
II. Advantages of screening microbial consortia with microdroplets
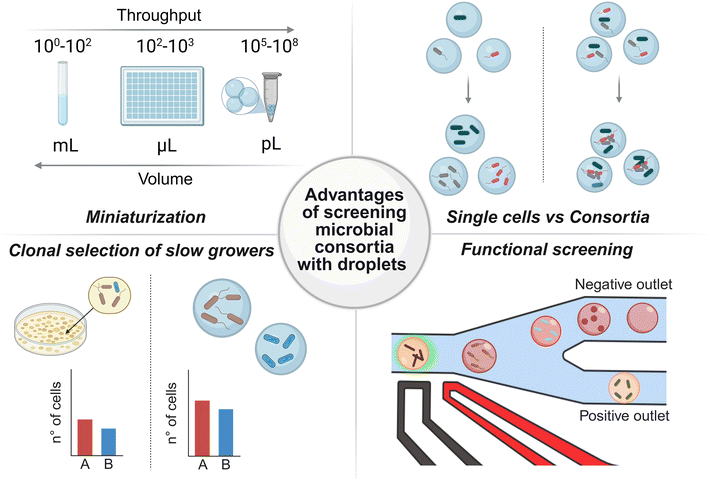
Miniaturizing microbial microcultures into picoliter droplets enables cost-efficient, high-throughput parallel screening. These capabilities make droplet microfluidics especially valuable for environmental microbiology and bioprospecting. In this section, we highlight the key advantages of the microdroplet format for microbial cultivation and functional screening (Fig. 1).Increase of throughput, miniaturization and cost reduction
Microfluidic systems provide an effective platform for the miniaturization of biological and chemical assays.31 In droplet-based approaches, aqueous compartments with volumes typically ranging from picolitres (10−12 L) to nanolitres (10−9 L) are generated at kilohertz frequencies, yielding millions of discrete microreactors for parallel screening.32 At these volumes, droplets can be inoculated at the single-cell level by controlling encapsulation according to Poisson statistics.26 Alternatively, multiple cells per droplet may be encapsulated when the aim is to investigate microbial consortia or communities. Following encapsulation, microorganisms are cultivated directly within droplets, with growth times generally ranging from several hours to a few days, commonly 12–72 hours,33,34 depending on the strain, cultivation media composition and incubation methods. This timescale is sufficient for many bacteria species to reach detectable densities, for instance, on the order of 102–103 cells per droplet, or to exhibit measurable metabolic or enzymatic activities relevant to environmental screening applications. Alternatively, if emulsion stability is sufficient, droplets can be incubated for extended periods, e.g. a month. Stable compartmentalization of microorganisms supports prolonged cultivation time further enabling the growth of slow-growing strains.9Aqueous droplets dispersed in oil serve as picoliter-volume reaction vessels that can be screened at thousands per second, significantly reducing the assay time, reagent use, and overall cost.35 These droplet-based reactions can be performed in inexpensive, single-use devices, further minimizing costs, labor, and processing time.36 A major advancement that enabled the emergence of droplet microfluidics was the development of soft lithography, which enables rapid replica molding of microfluidic devices.37 Rather than etching microchannels into costly substrates, a master mold can be used to fabricate numerous chips from transparent elastomers such as polydimethylsiloxane (PDMS). This method facilitates low-cost, custom production of microfluidic devices, making the technology more accessible and scalable. Crucially, the success of droplet microfluidics is not only attributed to soft lithography but also to the development of stable emulsions by selection of proper oil and surfactant compositions.38,39 These combinations are essential for generating and maintaining the high stability of the picoliter-volume aqueous compartments, which act as individual microreactors. Despite the low cost of individual microfluidic chips, the instrumentation required to operate droplet-based systems can still be expensive and, in some cases, prohibitive for small laboratories. To address this, passive droplet sorting techniques are emerging as attractive alternatives to active detection methods such as fluorescence or absorbance-based systems.40,41 By exploiting physical properties such as surface tension, deformability, or droplet size, passive systems enable selective enrichment without the need for complex instrumentation. These approaches simplify device design, reduce operating costs, and improve ease of use.42
Screening of clonal and complex microbial consortia
As a result of developments in complex experimental design, droplet microfluidics has emerged as an indispensable platform for investigating microbial interactions with higher precision and throughput than conventional culture-based techniques.32 Programmable devices like kChip43 allow for the controlled merging of fluorescently-tagged droplets containing different microorganisms, enabling detailed investigation of both positive and negative interactions, such as cross-feeding between auxotrophs or quorum-sensing inhibition. Droplet microfluidics can also be used to map interactions directly in microbial consortia. By randomly co-encapsulating cells, diverse synthetic communities can be generated and tested in parallel to reveal interactions between the taxa without predefining the partners44 and to assess the effect of external factors on community structure.45 Similarly, parallel droplet-based approaches make it possible to investigate how cultivation media properties, such as pH and buffering capacity, influence nutrient-driven interspecies interactions.46Clonal growth and isolation of slow growers
The characterization of rare and slow-growing microorganisms is particularly important, as these taxa are often underrepresented in conventional culture-based studies yet play critical roles in natural and engineered ecosystems. Slow growers may harbor unique metabolic pathways, secondary metabolites, or enzymatic activities that are valuable for strain selection, bioprospecting, bioremediation, drug discovery, and microbial ecology. For example, studies on the so-called ‘microbial dark matter’ have revealed that slow-growing or uncultivable lineages contribute disproportionately to phylogenetic and functional diversity.47,48 Moreover, rare taxa have been shown to stabilize community dynamics and mediate key ecological processes.49 From a biotechnological perspective, many bioactive compounds and novel enzymes are produced by slow-growing microorganisms that are difficult to access using standard cultivation approaches.10,50–52Traditional lab-scale methods, including dilution-to-extinction, most probable number (MPN), and colony-forming unit (CFU) plating, rely on spatial separation to limit resource competition in the cultivation medium.9,53 However, these techniques are low-throughput and often ineffective at recovering slow- or poorly-growing microorganisms or those unable to grow on conventional solid media and air–liquid interfaces.54,55 Microfluidic cell encapsulation offers a promising solution to these limitations. It is especially advantageous for studying rare microbial taxa, slow-growing species, and organisms with unique metabolic traits. For example, as early as two decades ago, simple emulsification techniques made it possible to encapsulate and cultivate single cells within droplets. This advancement led to the isolation of previously uncultured microorganisms from environmental samples, organisms that could not be accessed using conventional methods. Although not as advanced as modern high-throughput droplet microfluidics, these early approaches represented a significant milestone in the field.56 In another study8 slow-growing bacteria from the gut microbiome were enriched using cultivation in microdroplets followed by an image-based droplet sorting. This active selection approach enhanced the recovery of rare taxa beyond what was achievable through droplet cultivation alone.
Functional screening
Conventional methods for studying microbial function are often labor-intensive and limited in scope, restricting the discovery of novel strains, natural products, or enzymes. In contrast, functional droplet-based screening in microfluidic enables the rapid analysis of large microbial libraries.52 These assays can evaluate diverse functions, including enzyme activity, antimicrobial production, or metabolite synthesis and consumption,57 using the target product itself as a functional marker. By encapsulating single cells or microbial consortia, functional assays can be performed with high precision, facilitating the detection of rare activities, microbial interactions, and high-performing strains with potential for biotechnological application.58,59 Beyond throughput, droplet microfluidics enables quantitative functional assays, allowing microbial or enzymatic activity to be measured with high sensitivity at the single-droplet level. To increase robustness, growth amplification strategies have been employed to reduce phenotypic noise, thereby improving the accuracy of functional comparisons between individual clones.60 In addition, droplets can serve as microreactors, where coupled reactions and kinetic analyses of enzymatic processes can be executed.17,61 The utility of droplet systems has been further extended by the development of advanced sorting strategies. Label-free approaches, such as sorting by interfacial tension (SIFT), enable direct selection of metabolically active cells without external reporters,42 while active sorting systems such as fluorescence-activated droplet sorting (FADS)23 and absorbance activated droplet sorting (AADS),62,63 provide ultra-high throughput selection based on optical readouts. Furthermore, picoinjection allows the controlled addition of reagents into droplets prior to sorting, enabling multi-step assays and expanding the scope of functional screening.34,64 In summary, droplet-based functional screening offers unmatched throughput and sensitivity. However, the broader adoption of these assays still lags behind more established non-microfluidic techniques. Ongoing research is actively addressing this gap through the development of new Lab-on-a-Chip platforms and protocols.III. Modular components of microfluidics systems for screening of environmental microbial strains and consortia
Handling and processing of pico- or nanoliter-scale microvolumes differ fundamentally from standard pipetting in microplate-based workflows. Early studies established core functional modules that underpin modern droplet microfluidics. For example, on-chip valve technology enabled single-cell genome amplification of uncultivated microbial taxa directly from complex communities, providing access to previously unexplored microbial ‘dark matter’.65 Passive size-based droplet separation via deterministic lateral displacement arrays demonstrated high-throughput sorting of droplets according to biologically induced diameter differences.66 Another label-free strategy to quantify metabolic activity of individual cells was based on buoyancy-related volume changes within emulsions, resulting in osmotically driven droplet shrinkage.67 Controlled electrocoalescence of droplets synchronized by microfluidic flow enabled programmable, high-frequency droplet fusion for biochemical assays.68These early developments established the modular framework that formed the basis of modern systems, in which generation and manipulation of microdroplets containing microbial single cells and microcolonies require specialized modules, which can be integrated into a single microfluidic chip or distributed across multiple devices to enable multi-step functional assays.
Droplet generation
In lab-on-a-chip devices, droplets are formed through various mechanisms, including squeezing, dripping, jetting, tip-streaming, and tip-multi-breaking.69,70 Common microfluidic geometries include crossflow (e.g., T-junction, Y-junction), co-flow (2D and 3D), and flow-focusing geometries, where viscous shear forces in the continuous oil phase pinch off aqueous droplets. Alternatively, step emulsification produces monodisperse droplets by exploiting variations in channel width or depth that govern fluid flow. The choice of droplet generation module is typically guided by parameters such as flow rate, channel dimensions, and device design. A critical aspect of most microfluidic applications is droplet monodispersity, as it ensures that downstream operations such as reagent addition, quantitative readout and droplet sorting, proceed in a controlled and predictable manner. In contrast, polydisperse emulsions can compromise the reliability and reproducibility of these processes. A recent innovation, known as particle templated emulsification, uses solid particles as templates to produce uniform droplets.71 In this approach, microfluidic devices are used to generate monodisperse template particles prior to the main experiment with cell encapsulation, which can then be carried out using simple vortexing. This strategy increases the accessibility of droplet-based assays and holds promise for broad application in microbial cultivation and screening. As droplet generation techniques have been comprehensively reviewed elsewhere,69,70 and microbial encapsulation generally does not require non-standard droplet formation methods, we do not discuss this topic further in the present tutorial review.Droplet incubation
Microbial assays in microdroplet format primarily involve the use of specialized incubation techniques to support microbial growth. After droplet generation, incubation can be performed either off-chip, for example in external chambers, or on-chip, where droplets can be confined within the chip - trapped in microarrays or stored in integrated microchambers. A commonly used approach is off-chip static incubation, where droplets are stored in Eppendorf tubes or glass syringes, allowing relatively large volumes of emulsions to be maintained, with droplets floating on top of the oil phase. However, this strategy may not be optimal for aerobic organisms, as localized oxygen depletion can occur in densely packed droplets.72 An alternative to static incubation is off-chip dynamic incubation, in which droplets are continuously oxygenated by circulating oil through the emulsion by a flow from syringe or peristaltic pump within a dedicated incubation chamber.73 This method has been shown to enhance microbial growth and boost protein expression72 or biosurfactants secretion.50 In contrast, on-chip incubation employs microfluidic architectures specifically designed to trap and optionally release droplets in array-based formats under flow control. Picoliter-sized droplets can be confined and optically monitored over extended periods, allowing precise tracking of microbial growth and metabolic activity.74 Notably, on-chip systems also enable the full integration of droplet generation, incubation, and detection within a single microfluidic platform.5Imaging
Microbial growth within microfluidic systems can be monitored using a variety of imaging techniques. Bright-field microscopy is the most accessible and commonly used method for observing microbial growth inside droplets. Visualization can be performed on microscope slides, commercial microchambers, or directly on-chip, as PDMS devices are fully transparent. Time-lapse microscopy captures images at defined intervals, allowing dynamic observation of microbial processes such as cell division, growth, and biofilm formation over time. In contrast, fluorescence microscopy enables the selective labeling and real-time tracking of microbial cells, offering high sensitivity and specificity. Platforms such as the kChip have been adopted for droplet-based microbial assays. Implementing fluorophores into droplets makes it possible to encode and track their compositions, enabling parallel screening of complex interactions, such as formation of microbial communities.43,58 Furthermore, confocal laser scanning microscopy provides high-resolution, three-dimensional imaging, enabling detailed spatial analysis of droplets and the microbes within them.74,75 Additionally, Raman spectroscopy and surface-enhanced Raman scattering (SERS) enable label-free molecular readouts of droplet contents.76 However, it faces challenges related to signal intensity and optical integration. Despite these limitations, it has been successfully applied to the screening of single microalgal cells.87 In this approach, Raman microspectroscopy analysis of individual cells is performed prior to their encapsulation in microdroplets, which are subsequently subjected to droplet sorting.77Other modules: picoinjection and droplet fusion
In addition to droplet sorting, various microfluidic chip designs support complex screening workflows for microbial activity. In multi-step protocols, picoinjection is a valuable module that enables the high-throughput addition of buffers, reagents, or substrates into existing droplets.78 A typical picoinjector consists of two sections: a droplet spacing region, which uses oil from side channels to separate droplets, followed by the injection region, where an aqueous phase is introduced to each droplet. The merging of the injected fluid with the droplet is continuous or can be triggered by an electric field applied via embedded electrodes in the chip.79 Similarly, droplet fusion can be used to introduce defined volumes of liquid into droplets by merging two droplets. This process is essential for mixing reagents or combining samples at significantly different droplet volumes. Fusion is traditionally induced using an electric field in dedicated chambers or wider channels. Droplets can also be passively merged in structures such as pillar traps,64,80 static arrays,81 or through flow manipulation using rails and anchors.82,83 However, in some multi-step protocols, unintended droplet merging can occur, for example after prolonged incubation or during PCR. These undesired fusions can be addressed by routing the droplets through a droplet-size filter microfluidic device to sort and remove irregularly merged droplets.84Droplet sorting
In addition to enabling imaging, the use of transparent PDMS in microfluidic chip fabrication has enabled the development of a wide range of techniques for droplet analysis and sorting (Table 1). Current microfluidic sorting protocols exhibit substantial variability in throughput, sensitivity, and cost, as well as in their methodological requirements, ranging from dependence on fluorescent reporters or dyes to fully label-free approaches (Fig. 2). These differences reflect trade-offs between analytical precision, complexity of the setup, and practical applicability.| Detection method | Applied to environmental microbial screening? | Sorting frequency | Droplet volume | Reference | ||
|---|---|---|---|---|---|---|
| Labelled | Fluorescence activated droplet sorting | FADS | ✓ | Low kHz | Mid pL | 23 |
| Fluorescence activated cell sorting | FACS | ✓ | Mid kHz | Mid pL | 98 , 117 | |
| Absorbance activated droplet sorting | AADS | ✗ | Low kHz | Mid pL | 62 , 63 | |
| Label-free | ||||||
| Surface-enhanced Raman spectroscopy | SERS | ✗ | High Hz | Mid pL | 76 | |
| Passive | Deformability | ✓ | Mid Hz | Mid pL | 51 , 113 | |
| Droplet density | ✗ | NR | Low nL | 92 | ||
| Surface tension | ✗ | High Hz | High pL | 93 | ||
| Size | ✗ | NR | Mid-high pL | 136 | ||
| Magnetic | ✗ | NR | Low nL | 7 | ||
| Image analysis | ✗ | Mid Hz | Mid pL | 89 | ||
| Electrochemical | ✗ | Low Hz | Mid nL | 90 | ||
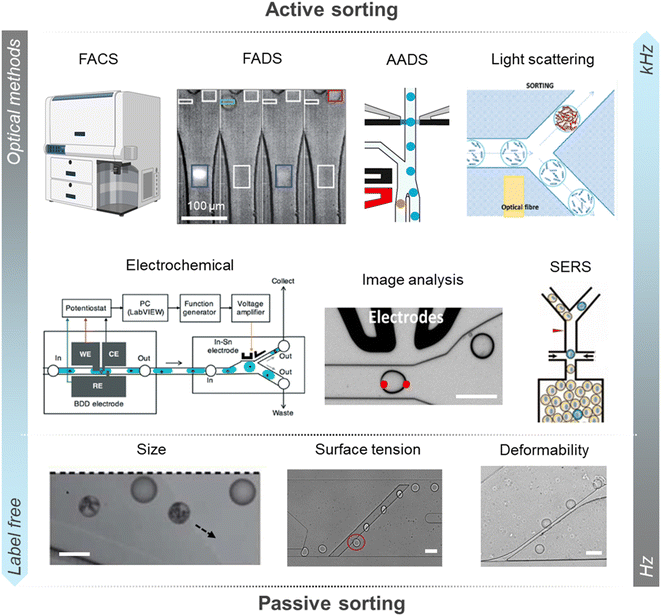 | ||
| Fig. 2 Schematic overview of droplet-based sorting technologies for the enrichment of microbial consortia. State-of-the-art microfluidic platforms employ a wide spectrum of readout modalities and sorting mechanisms, which differ in sensitivity, scalability, and cost, and can be broadly classified into active and passive approaches. At the highest throughput levels (103–104 Hz), active optical methods such as FADS and FACS are the most established protocols, offering robust and versatile implementations across diverse biological applications. Although costly, FACS remains a viable option for droplet formats such as double emulsions. Light scattering detection in droplets can also be performed with up to kilohertz frequencies, providing a rapid and label-free option. Intermediate-throughput approaches (102–103 Hz) integrate alternative readout principles that enable higher analytical flexibility. Electrochemical detection affords sensitive quantification, while image-based classification supports real-time assessment of droplet morphology and content. Implementation of SERS also represents a label-free identification of biochemical signatures of droplets. At the lower throughput scale (101–102 Hz), passive enrichment strategies exploit intrinsic droplet properties such as size, biosurfactant production, or deformability to achieve selective screening. Scale bars shown in the FADS, image analysis, size, surface tension, and deformability panels are representative of 100 μm. Fig. 2 was reproduced with permission from the following references: ref. 98, Wiley, 2022, CC BY 4.0; ref. 23, RSC, 2009; ref. 62, ACS, 2023, CC BY 4.0; ref. 88, RSC, 2016; ref. 90, RSC, 2020; ref. 8, eLife, 2020, CC BY 4.0; ref. 76, Front. Bioeng. Biotechnol., 2024, CC BY 4.0; ref. 135, RSC, 2018; ref. 50, RSC, 2024; and ref. 51 ACS, 2024, CC BY 4.0. | ||
Among the most widely adopted methods is FADS, which is frequently used alongside the increasingly popular AADS. Recent advancements in FADS have moved beyond binary selection, introducing multiplexed digital sorters that utilize multiple fluorescence intensity thresholds to isolate mutants with varying levels of enzymatic activity, such as glucoamylase in yeast strains.85,86
Building upon these FADS-based strategies, recent advancements have moved beyond binary ‘hit-or-miss’ selection by introducing multiplexed digital sorters. These systems utilize multiple fluorescence intensity thresholds to isolate mutants with varying levels of activity, such as glucoamylase in yeast strains, rather than focusing exclusively on top performers. To further enhance this precision, novel architectures like the SeParate platform have been developed,86 combining serial and parallel triaging concepts to achieve high-accuracy multiway sorting into multiple output channels. This nuanced approach to selection is of particular strategic importance, as identifying intermediate or ‘medium-performing’ strains provides a more comprehensive map of the fitness landscape. Such variants may harbor unique genetic traits, such as superior robustness or alternative metabolic flux patterns, that are essential for long-term strain stability and industrial scalability, yet are often prematurely discarded in traditional high-yield-only pipelines.85
Several other analytical techniques, though less conventional, have shown promise including Raman spectroscopy,87 light scattering analysis,88 and real-time image-based droplet analysis.8,89 Beyond optical techniques, alternative analytical approaches, such as mass spectrometry, electrochemical detection,90 have also been explored for active sorting and characterizing droplet contents.20 While active droplet sorting systems dominate current workflows, recent studies have introduced passive microfluidic strategies that enable droplet selection based on intrinsic physicochemical properties such as surface tension, buoyancy, and droplet deformability.42,50,51,91–93 Microvalve-based approaches can complement other methods, offering deterministic control over droplet trajectories, mechanical actuation, and straightforward integration with on-chip incubation modules. These features are particularly advantageous for workflows involving long-term cultivation or environmentally sensitive microorganisms. Pneumatic microvalves have been shown to enhance sorting performance through precise control of droplet motion and reduced misrouting errors,94 and have been applied for capture and enrichment of pathogenic cells.95 More recently, pneumatic microvalve-driven multi-way fluorescence-activated sorting platforms have enabled simultaneous fractionation of multiple target populations with high specificity and throughput.96
Despite the upfront costs of implementing approaches based on microfluidics, the capacity to conduct ultra-high throughput, parallelized experiments enables faster discovery and optimization, making the technology cost effective over time, especially considering the reduced reagent consumption (per individual experiment). Combining microfluidics with FACS further enhances efficiency by enabling sorting and deposition of individual target cells based on fluorescent markers or functional activity, allowing the isolation of rare or high-performing strains for biotechnological applications.97–99 As in other fields, ongoing technological advancements and increased accessibility are expected to reduce operational barriers and costs, expanding the reach of this innovative screening approach.
IV. Types of screening approaches
Droplet-based assays enable a wide range of screening strategies for the analysis and selection of microbial consortia (Fig. 3). Depending on the experimental objective, different workflows can be applied. For example, growth-based assays often rely on microscopy and genomic analysis to evaluate the outcomes of droplet screening experiments. In contrast, functional activity assays typically require the integration of droplet sorting strategies. The following sections highlight recent advances in these screening modalities, illustrating how microfluidic platforms are being used to dissect community dynamics, identify rare strains, and optimize microbial functions for both research and biotechnological applications.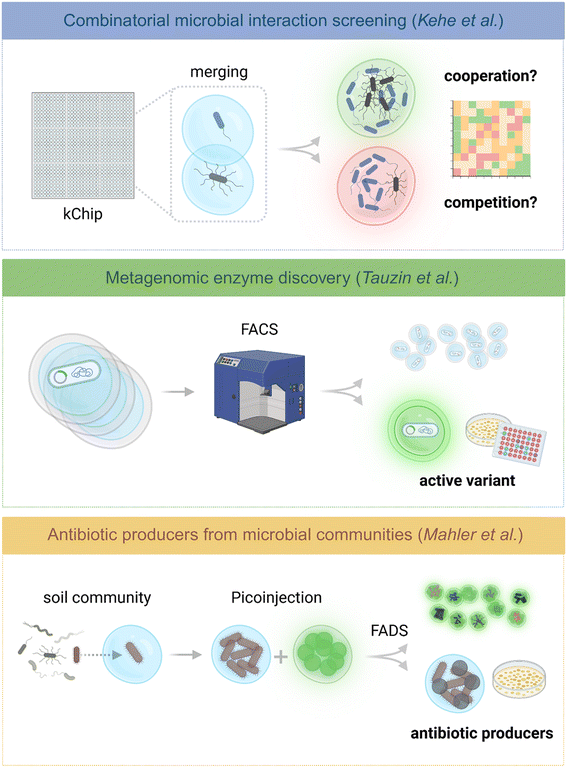 | ||
| Fig. 3 Droplet microfluidics enables the development of diverse HTS approaches. Compartmentalization of cells, enzymes, or microbial communities into droplets functioning as independent microreactors enables massively parallel assays beyond conventional protocols. Here we highlight three applications showcasing the diverse range of uses for droplet-based microfluidic methods. Top – blue panel: the kChip platform investigates microbial interactions by merging fluorescent color-encoded droplets in microarrays to assemble defined communities (Kehe et al.). Middle – green panel: this screening encapsulated bacteria expressing variants of a target enzyme in double emulsions. Upon enzyme production, the reaction generates a fluorogenic product, and active droplets are enriched by FACS and recovered and upscaled (Tauzin et al.). Bottom – yellow panel: single-cell encapsulation enables microbial growth in droplets. After incubation, droplets are merged with fluorescent reporter bacteria, forming co-cultures. Based on the resulting fluorescence signal, droplets are sorted to identify antibiotic-producing strains (Mahler et al.). Fig. 3 was adapted with permission from the following references: ref. 58 Copyright 2021, AAAS, CC BY-NC 4.0, ref. 111 Copyright 2020, Springer, CC BY 4.0, ref. 9 Copyright 2021, eLife, CC BY 4.0. | ||
Droplet cultivation and microscopic analysis
Microscopy-based observation of droplet contents is one of the earliest strategies in microfluidic microbial assays, enabling monitoring of cell morphology and population dynamics within individual droplets.100,101 In addition, freely available software facilitates cost- and time-efficient image-based droplet detection.102,103 The kChip platform enables the massively parallel analysis of microbial interactions by generating fluorescently barcoded nanoliter droplets, each containing specific microbial strains or media components. These droplets are randomly loaded into microwells and merged via electrofusion to create diverse synthetic communities. Subsequent time-lapse imaging quantifies microbial growth or inhibition based on fluorescence or optical readouts, allowing high-throughput mapping of interaction networks across thousands of combinations.43 DropVIST further integrates droplet microfluidics with wide-field, real-time imaging to continuously monitor microbial growth and metabolic activity within droplets. Its automated image analysis and droplet-tracking pipeline provides dynamic, high-throughput quantification of microbial behavior, making the system particularly suitable for rapid phenotyping and functional screening.104 Beyond content- or color-based readouts, droplet morphology provides functional information: deformability has been used to assess microbial proteolytic activity via induced-droplet-ovalization (IDO) analysis.33 Although highly sensitive and label free, such methods generally lack direct integration with high throughput enrichment of positive droplets, apart from an image based sorting algorithm that classified droplets by colony density, enabling enrichment of droplet cultures with slowly growing colonies.8Droplet microfluidics for microbial genomics
Recent advances in droplet microfluidics have enabled high-throughput clonal cultivation of complex microbiomes, followed by genomic characterization, offering a powerful strategy for dissecting microbial diversity, strain-level variation, and interspecies interactions.105Clonal cultivation of complex input microbiomes was applied to fecal microbiota transplant donor stool under multiple media conditions. This approach significantly increased taxonomic richness and enhanced the recovery of rare and clinically relevant taxa compared to conventional plate-based cultivation methods.8 Another recent example of a growth-based assay followed by genomic characterization is the MicDrop platform, which integrates droplet microfluidics with DNA sequencing to study the human gut microbiota. MicDrop enabled the cultivation of 2.8 times more bacterial taxa compared to conventional methods. Designed specifically to investigate carbohydrate-degrading bacteria across individuals, the platform revealed notable variations in species richness and abundance. Additionally, lifestyle factors were found to correlate with microbial diversity. This approach offers significant potential for advancing our understanding of microbiota ecology and evolution, and for guiding the development of microbiota-targeted therapies.11
A similar approach was used to generate millions of droplets, each encapsulating single bacterial cells extracted from honeybee guts in droplets composed of various media. Metagenomic analysis revealed significant shifts in community structure, with certain species exhibiting greater strain-level diversity compared to bulk gut samples. Unique clusters within Bifidobacterium and distinctive genes in Lactobacillus panisapium were identified. By isolating individual cells in droplets, interspecific competition was minimized, enabling high-resolution shotgun sequencing. Comparative genomic analysis provided new insights into host-specific adaptations and microbe–microbe interactions, highlighting the utility of this method for studying complex animal-associated microbial communities at scale.106
In another example, a microfluidic platform was recently developed for screening bacterial growth phenotypes within multispecies consortia. Interspecies interactions play a critical role in shaping microbial communities, with positive interactions significantly influencing both diversity and productivity. Using the kChip platform, one study examined over 1.8 × 104 pairwise interactions among 20 soil bacterial strains across 40 environmental conditions. Positive interactions were found to be common, primarily in the form of parasitism, in which slow-growing strains benefit from the presence of fast-growing taxa with distinct carbon utilization profiles. Remarkably, more than 85% of non-growing strains were promoted by co-culture with faster-growing partners.58
Functional screening
Functional screening of bacteria using microfluidics involves analyzing microbial activity within individual droplets. This approach enables high-throughput assessment of various phenotypes, including cellulolytic, lipolytic, and proteolytic activities; antibiotic susceptibility; biofilm formation; and quorum sensing dynamics.107 In the table below we highlight recent studies that demonstrate modern and innovative applications of droplet microfluidics for the functional screening of microbial growth and activity (Table 2).| Target | Format | Screening | Sorting | Reference | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Clonal | Multispecies community | Enrichment | Growth analysis | Strain improvement | Interactions | FADS | FACS | AADS | Image-based | Passive | None | |||
| Growth based | Compositions of symbionts | ■ | ■ | ■ | 126 | |||||||||
| Effect of antibiotics on community assembly | ■ | ■ | ■ | 45 | ||||||||||
| Human gut microbes | ■ | ■ | ■ | 131 | ||||||||||
| Antibiotic resistant gut microbes | ■ | ■ | ■ | 8 | ||||||||||
| Soil strains | ■ | ■ | ■ | 60 | ||||||||||
| ■ | ■ | ■ | 58 | |||||||||||
| Water microbiome | ■ | ■ | ■ | 110 | ||||||||||
| Honeybee gut microbiome | ■ | ■ | ■ | 106 | ||||||||||
| Enzymatic activity | Xylose-degraders, L-lactate producers | ■ | ■ | ■ | 57 | |||||||||
| Cellulases | ■ | ■ | ■ | 10 | ||||||||||
| ■ | ■ | ■ | 34 | |||||||||||
| ■ | ■ | ■ | ■ | 108 | ||||||||||
| β-N-acetylgalactosaminidase | ■ | ■ | ■ | 111 | ||||||||||
| Amino acid racemase | ■ | ■ | ■ | 122 | ||||||||||
| Degradation of poly- and monosaccharide | ■ | ■ | ■ | 11 | ||||||||||
| Amino acid dehydrogenase | ■ | ■ | 72 | |||||||||||
| α-Amylases | ■ | ■ | ■ | ■ | 109 | |||||||||
| Agarases | ■ | ■ | ■ | 91 | ||||||||||
| Proteases | ■ | ■ | ■ | 33 | ||||||||||
| ■ | ■ | ■ | 51 | |||||||||||
| Chitinases, glucanases, NAc-galactosaminidases | ■ | ■ | ■ | 130 | ||||||||||
| Natural products | Amicoumacin A | ■ | ■ | ■ | ■ | 52 | ||||||||
| Antimicrobial compounds | ■ | ■ | ■ | 9 | ||||||||||
| Biosurfactants | ■ | ■ | ■ | 50 | ||||||||||
One of the first droplet microfluidic platforms for ultra-high-throughput discovery of cellulolytic microorganisms encapsulated single environmental cells in picoliter droplets with a fluorogenic substrate, enabling enzymatic activity detection at the single-cell level. Screening over 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells in less than 20 minutes, the system achieved a 17-fold and 7-fold enrichment in cellobiohydrolase and endoglucanase activities, respectively, and revealed microbial community profiles distinct from conventional cultivation. This study represented an interesting example of a proof-of-concept demonstration toward practical discovery of microorganisms with specific enzymes. Its developmental impact lies in proving that droplet microfluidics can rapidly explore functional microbial diversity, establishing a robust foundation for industrial bioprospecting and demonstrating the transformative potential of high-throughput microfluidic technologies in biotechnology.10
000 cells in less than 20 minutes, the system achieved a 17-fold and 7-fold enrichment in cellobiohydrolase and endoglucanase activities, respectively, and revealed microbial community profiles distinct from conventional cultivation. This study represented an interesting example of a proof-of-concept demonstration toward practical discovery of microorganisms with specific enzymes. Its developmental impact lies in proving that droplet microfluidics can rapidly explore functional microbial diversity, establishing a robust foundation for industrial bioprospecting and demonstrating the transformative potential of high-throughput microfluidic technologies in biotechnology.10
A droplet-based high-throughput screening platform was later developed to enhance cellulase production in Pichia pastoris through whole-cell directed evolution. The system relied on a cellulase-catalyzed reaction that released a fluorescent signal, enabling sorting of methanol-grown P. pastoris cells with 94.4% efficiency at a rate of 0.3 kHz. After five iterative cycles of enzyme mutagenesis and droplet-based screening, cellulase activity in evolved mutants progressively increased. The top variant showed approximately a two-fold improvement over the parental strain, with whole-genome sequencing revealing a consistent point mutation across high-performing clones. Developmentally, this work showcased how microfluidics can directly integrate with whole cell directed evolution, advancing industrial strain engineering toward efficient enzyme production pipelines.108
Recently, a droplet-based microfluidic platform employing FADS has been developed for ultrahigh-throughput screening of enzyme-producing microorganisms. The system integrates single-cell encapsulation, long-term droplet incubation, and fluorescence-based sorting at rates up to 0.3 kHz, thereby accelerating strain development and directed evolution. In a model enrichment experiment, it achieved a 45-fold increase in the proportion of active variants, demonstrating the precision and efficiency of the workflow. When applied to a mutant library, it isolated Bacillus licheniformis strains with more than a 50% increase in α-amylase production.109 This work is significant in demonstrating how droplet microfluidics can be combined with directed evolution, particularly using atmospheric and room-temperature plasma (ARTP) mutagenesis, thereby moving beyond simple cultivation toward fully integrated, high-throughput platforms for strain optimization and industrial biotechnology. Building upon these FADS-based strategies, recent advancements have moved beyond binary ‘hit-or-miss’ selection by introducing multiplexed digital sorters. These systems utilize multiple fluorescence intensity thresholds to isolate mutants with varying levels of activity, such as glucoamylase in yeast strains, rather than focusing exclusively on top performers. To further enhance this precision, novel architectures like the SeParate platform have been developed,86 combining serial and parallel triaging approaches to achieve high-accuracy multiway sorting into multiple output channels. This nuanced approach to selection is of particular importance, as identifying intermediate or ‘medium-performing’ strains provides a more comprehensive map of the fitness landscape. Such variants may harbor unique genetic traits, such as superior robustness or alternative metabolic flux patterns, that are essential for long-term strain stability and industrial scalability, yet are often prematurely discarded in traditional high-yield-only pipelines.85
Another important advancement was the development of FNAP-sort, which integrates microfluidic cultivation with on-chip sorting using a fluorescent nucleic acid probe to detect RNase activity in growing cells. In this system, microbes are encapsulated with a FRET-quenched RNA probe that emits fluorescence upon cleavage by RNases during cell growth. Droplets exceeding a defined fluorescence threshold are sorted and subsequently dispensed for cultivation and 16S rRNA sequencing to identify isolates. The study demonstrated the recovery of nearly pure cultures of rare environmental taxa from complex microbiome samples, including organisms present at very low abundance in the source population. Notably, the entire workflow was implemented using commercially available components, enhancing its accessibility to other laboratories. By enabling the selective recovery of viable microbes independent of prior labeling or growth rate, FNAP-sort mitigates biases toward fast-growing species and facilitates the isolation of slow-growing or rare taxa that are often overlooked in conventional cultivation methods.110
An important extension of droplet microfluidics enabled the investigation of host–microbiome interactions by coupling fosmid libraries with functional screening of metagenomic DNA derived from mucosal microbiota. This platform enabled high-throughput detection of β-N-acetylgalactosaminidase activity, a key glycosidase involved in host–microbe communication and mucus layer homeostasis, with relevance to inflammatory bowel diseases (IBD). From 1.9 × 103 droplets sorted by FACS, 3569 colonies were recovered and 372 rescreened, yielding 31 highly active clones corresponding to 15 unique sequence hits. Several activities were traced to horizontal gene transfers involving Bacteroides species, highlighting the evolutionary plasticity of gut microbial functions. This work demonstrated how droplet microfluidics can bridge functional enzyme discovery with microbiome ecology, enabling precise dissection of host-associated microbial activities at single-cell resolution.111
Droplet microfluidics has also been adapted for anaerobic cultivation of gut microbes, allowing high-throughput encapsulation and growth monitoring via image analysis. This approach enriched rare and slow-growing taxa overlooked by traditional plate cultures and successfully recovered clinically relevant Bacteroides species absent in standard methods. When the cultures were supplemented with antibiotics, droplet cultivation revealed 21 resistant populations that remained undetected using conventional techniques. The impact of this work lies in extending droplet technology to clinically important anaerobic ecosystems, linking cultivation with resistance profiling. By overcoming cultivation biases and uncovering hidden microbial reservoirs, this study positioned droplet microfluidics as a powerful platform for translational microbiome research, diagnostics, and public health surveillance.8
Finally, the technology has been applied to prioritize antibiotic producers from soil microbial communities. After encapsulation and growth, droplets were infused with susceptible reporter strains, enabling identification of inhibitory activity within individual compartments. This strategy captured a broader diversity of producers than plate cultures and led to the isolation of Bacillus-related strains producing bacillaenes antibacterial polyketides and surfactin-like lipopeptides. The study shifted droplet microfluidics from merely expanding cultivable diversity toward direct functional bioactivity screening. Its impact lies in embedding antimicrobial discovery directly into droplet workflows, streamlining natural product screening, and accelerating the search for novel antibiotics at ultrahigh-throughput.9
In contrast to the aforementioned screening methods, HTS can also be performed passively. A recent example demonstrated the development of a passive droplet microfluidic platform for high-throughput screening and enrichment of proteolytic microorganisms based on droplet deformability. The barrier-driven sorting mechanism of the DPDS chip (deformability-based passive droplet sorter) operates by exploiting the solid-to-liquid transition of the gelatine medium that occurs in the presence of proteolytic strains.51 The system encapsulated single bacterial cells in 100 pL gelatine droplets, enabling clonal growth and sorting without the need for labels, dyes or fluorescence. Optimization of droplet size, gelatine concentration, and temperature allowed for stable and precise sorting at rates of up to 50 Hz. Under optimal conditions, the DPDS achieved high accuracy, sensitivity, and specificity. When applied to a mock consortium of P. aeruginosa (proteolytic, environmental strain) and E. coli (non-proteolytic), the platform enriched P. aeruginosa from an initial 0.5% abundance to over 99% purity within 90 minutes. Compared to previous deformability-based approaches,81 this system improved throughput by 50-fold while maintaining excellent sorting precision. Overall, this study demonstrates a simple, cost-effective alternative to active sorting, enabling label-free, high-throughput selection of proteolytic microbes from environmental sample for biotechnological and environmental applications.113
V. Planning of experimental workflow
The successful identification of beneficial traits from environmental microbial communities requires a well-structured experimental workflow that addresses the inherent complexity and variability of natural samples. The experiment planning phase is especially critical and should integrate considerations of sampling strategy, cultivation format, and selection of screening approach to maximize the likelihood of identifying functionally relevant phenotypes within droplets.Across droplet-based and microscale platforms, system design is governed by key trade-offs. Incubation time inversely affects throughput, as longer cultivation reduces experimental pace; however, this step can be readily parallelized and typically requires only simple instrumentation and minimal resources.9,51 A balance also exists between environmental fidelity and analytical depth. Smaller droplets may not fully capture complex microbial consortia, but compartments containing hundreds of cells are typically sufficient to reproduce relevant functional interactions. Finally, screening speed must be weighed against biological representativeness. While droplet systems favor rapid analysis, they can simplify ecological complexity, yet similar limitations are also present in macroscale methods, making this a general experimental constraint rather than a platform-specific drawback. Conversely, droplets enable the generation of thousands of precisely defined synthetic consortia,43,58 offering a powerful approach to dissect biological complexity and interactions within microbial communities.
Sampling strategy
In microfluidic workflows, the sampling strategy must ensure the preparation of a uniform, single-cell suspension that is compatible with droplet encapsulation. For complex sources such as soil or biofilm-rich environments, pre-treatment steps, such as mechanical disruption, gentle filtration, or the use of chemical dispersants (e.g., Tween 20), may be necessary to disaggregate microbial clusters and minimize debris that can obstruct microfluidic channels. It is essential to evaluate the potential impact of any additives on emulsion stability following droplet generation. Achieving the appropriate cell density is vital to obtain the desired droplet occupancy, typically modeled by the Poisson distribution. Plating environmental samples and preparing initial droplet emulsions can help identify optimal cell loading conditions and assess microbial growth, providing essential insights for optimising dilution strategies.Droplet formats
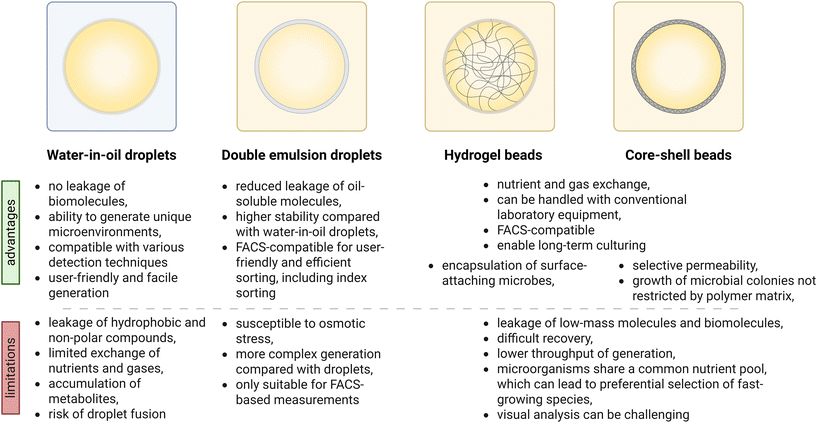
The use of microfluidics enables generation of a diverse range of microemulsions. While water-in-oil droplets are most commonly used for microbial screening, implementation of other emulsion formats is possible and broadly implemented. Each one of them (water-in-oil droplets, double emulsions, hydrogel beads, or core–shell beads) presents distinct advantages and trade-offs (Fig. 4). Choosing an appropriate format is a crucial aspect of microfluidic experimental design, as it not only influences the compatibility with downstream screening methods, but also determines factors such as emulsion stability, nutrient exchange, substrate retention, crosstalk between compartments, and the throughput of a screening campaign. The selected format must align closely with the screening objectives and desired selection strategy.Droplet microfluidics focuses on the precise manipulation of fluids at the microscale. A clear understanding of droplet formats, such as their composition, size, and functionality, is essential for designing effective experiments and enabling downstream applications. In a typical format, microfluidic workflows commonly use single emulsion droplets. These systems involve two immiscible phases: a biologically inert liquid, often fluorinated oil, serves as the continuous phase of the emulsion. The second phase is an aqueous solution, typically a cultivation medium, which acts as the dispersed phase and forms droplets upon introduction into the microfluidic system. Within the chip, it is critical that the aqueous droplets do not adhere to the channel walls where the oil flows, as this can lead to cross-contamination between droplets. Emulsion systems enable the compartmentalization of cells or biochemical reactions and allow for the encapsulation of biomolecules.112
Double emulsions, which are more complex than single emulsions, and they provide hierarchical compartmentalization, supporting the co-encapsulation of cells, reagents, and other components. These structures are typically generated through sequential emulsifications in separate microfluidic devices or specialized single chips designed with regions of differing wettability.114 Double emulsions can also serve as templates for the production of microcapsules and microparticles with core and shell architectures. These formats are highly valuable in pharmaceutical, biological, and industrial applications.115,116 In the case of microbial screening, the main advantage of double emulsions is their compatibility with widely accessible FACS instruments.117,118 FACS enables multiparameter sorting, which is difficult to achieve using custom-built fluorescence activated droplet sorters. Additionally, index sorting allows for the deposition of single droplets or microgels into individual wells of a multiwell plate.
Recently, the integration of hydrogels into microfluidic systems has led to significant advancements.119 Due to their biophysical similarity to soft biological tissues, hydrogels are highly suitable for use in microfluidic systems for screening,91 cell culture,120 and tissue engineering.121 Hydrogels and hydrogel microcapsules also enable FACS-based sorting and, unlike double emulsions, they allow for long-term cultivation of various cells, including microorganisms, due to the unrestricted exchange of small nutrients and metabolites. This can enable growth-based assays, where growth is an indicator of an improved catalytic activity of a strain, especially when coupled with antimetabolite selection.122 However, this benefit comes at the cost of substrate leakage. The diffusion of low molecular weight compounds can significantly limit the application of these formats in high throughput screening.
This limitation can be partially addressed through implementing microcapsules with semi-permeable shells. The pore size of the shell can be fine-tuned, allowing for semi-controlled transport in and out of the capsule.123 This makes it possible to run more complicated workflows using standard laboratory equipment, while omitting highly technical microfluidic steps. Microcapsules enable isolation and cultivation of microbes from environmental samples, achieving improved recovery of rare taxa and enrichment of slow-growers using FACS.124
Importantly, microorganisms encapsulated within microgels and capsules share a common nutrient pool, which may lead to competitive dynamics and favor the proliferation of fast-growing species, thereby suppressing slower-growing or rare taxa. Looking forward, we anticipate that both droplet-based and microgel-based formats can be combined for synergistic benefits. For example, microbial cultivation could be performed in a microgel format, followed by particle templated emulsification for functional assays in a droplet format.
Cultivation approach
The next key consideration is the cultivation strategy. Since many environmental microbes remain difficult to grow using traditional methods, the workflow should incorporate multiple approaches to improve their cultivability and recovery.Aside from cultivation media optimization,12 this challenge can be largely addressed by using the droplet format, which has been shown to promote taxonomic enrichment in culture.8 In addition, microbial diversity can be enhanced by employing low nutrient or conditionally enriched media, co-cultivation strategies, or by optimizing in-droplet cultivation, such as incubation time and aeration. Where applicable, semi-defined media that mimic native environments, such as soil extract or host-derived components, may promote the growth of otherwise unculturable species.125 Importantly, all cultivation steps must be evaluated for their compatibility with droplet encapsulation and downstream analyses. This includes avoiding media components or screening substrates that may destabilize emulsions or increase leakage of substrates and reaction products between droplets. A critical design decision is whether to screen single strains or microbial consortia. Screening consortia for synergistic activity requires droplet conditions that support co-cultivation126 careful control of droplet volume, which affects nutrient availability, and precise tuning of inoculum density. In either case, seeding density should be optimized to promote monoclonal occupancy or defined co-culture cell ratios, depending on the experimental objectives. This can be achieved using the principles of the Poisson distribution, which relate cell occupancy probability to the input cell density and final droplet size. Both parameters must be tightly regulated throughout the droplet generation process.
Additionally, while the use of liquid media remains the dominant choice for droplet-based cultivation, several approaches incorporate a solid or phase within droplets to better emulate surface-associated growth and to support organisms that require solid media. One common strategy for bacteria is to encapsulate cells in hydrogel beads, commonly referred to as gel microdroplets (GMDs), enabling microcolony formation and downstream high-throughput screening. This approach has been implemented in metagenomic screening,127 whole-genome sequencing128 from microcolonies and cultivation of fungal strains.129 In parallel, implementation of solid-phase media within droplets have proven useful for filamentous fungi, where hyphal growth can destabilize conventional droplets. For example, a core–shell droplet platform has been used for high-throughput fungal screening, where a solid-phase droplet exterior composed of gelatin methacrylate enables multi-day incubation and sorting without droplet merging.129 Droplets can also be supplemented with solid-phase media, such as colloidal chitin to enable functional screening of cell-wall degrading enzymes.130
Selection of sorting methods
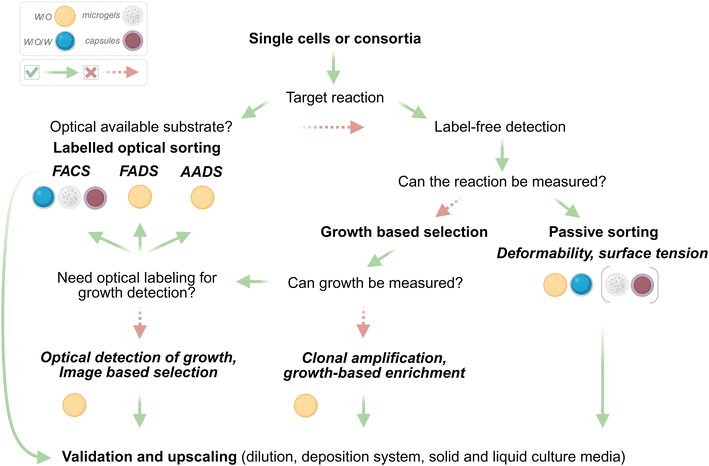
The most critical decision in experimental design is the choice of the sorting approach (Fig. 5). As outlined, the first question in this process is whether the target reaction is known and detectable. If the target can be monitored using an optically active substrate, methods such as FACS, FADS, or AADS become suitable options. These techniques enable selection based on fluorescence or absorbance, depending on the substrate and the specific droplet format used, such as water in oil, water in oil in water, microgels, or microcapsules. However, assays based on optically active substrates are associated with challenges such as substrate or product leakage, high background signals from substrate hydrolysis, or instability of the optical signal within droplets. Potential solutions include chemical modification of substrates to increase their polarity and decrease their hydrophobicity, coupling reactions to more stable and less leaky compounds, or incorporating advanced microfluidic operations such as picoinjection to introduce assay components at controlled time points. If optically active substrates are inadequate or unavailable, the screening workflow can shift toward label-free detection strategies. Light scattering techniques offer viable alternatives, particularly when screening for growth-based enrichment or for morphological characteristics. Passive sorting approaches can serve as effective alternatives to the previously described active methods, particularly when screening for activity on native substrates in the absence of reporters or bait substrates. In such cases, droplet selection can rely on indirect metrics such as deformability,46,81 surface tension,45 or other physicochemical features to infer biological activity. These methods become especially powerful when integrated with advanced image analysis algorithms for phenotypic classification.Validation and upscaling of selected cultures
A complete workflow must also include strategies for hit validation and recovery. This involves ensuring that emulsion-breaking procedures and nucleic acid extraction protocols preserve the identity of positive hits. Depending on the downstream objective, such as taxonomic identification, functional rescreening, or genetic engineering, suitable approaches may include barcoded sequencing, regrowth of encapsulated strains, or direct recovery of genetic material. Following hit identification, the final step involves deposition and potential analysis and scale-up of the droplets or cultures of interest. Transitioning from microscale screening to preparative or application-oriented standard workflows requires maintaining the genotype–phenotype linkage established during droplet-based selection, while enabling microbial growth under controlled conditions. The deposition of droplets into larger volumes can be achieved using commercially available sorting platforms, such as FACS that deposits double emulsions or microgels onto microwell plates,131 through controlled spreading of droplets on agar plates60 or using dedicated droplet deposition systems.61 Crucially, the feasibility of deposition and upscaling depends on the droplet format used in the screening pipeline, as only certain formats are compatible with FACS. The success of a droplet-based discovery and recovery pipeline depends on the careful integration of all stages of the workflow, from sampling and cultivation to screening and downstream recovery, to ensure that detected hits can be reliably scaled and applied. To address the common pitfalls of microfluidic workflows, we provide a concise troubleshooting guide (Table 3) linking commonly observed assay artifacts to their most likely causes and mitigation strategies across droplet incubation, and screening.| Observed issue | Step | Likely cause | Changes to implement |
|---|---|---|---|
| Droplet coalescence | Any | Interfacial instability, shear | Surfactant optimization, lower shear in microfluidic devices, filtration and size selection following problematic steps |
| Heterogenous growth | Incubation | High inoculum density, nutrient depletion in long incubation cycles | Lowering inoculum density, increased droplet size to increase nutrient availability, decreased incubation time |
| Limited growth | Incubation | Limited oxygen, low nutrient availability | Dynamic incubation with aeration, capsule format to enable nutrient exchange, implementing biologically inert compounds in the microfluidic systems |
| Droplet shrinkage | Incubation | Evaporation, osmosis | Sealing the incubation chambers, collection of droplets in sealed tips for incubation, lower dead volume in incubation containers |
| Growth stops after initial burst | Incubation | Waste accumulation | Earlier sorting, lower cell density, encapsulation in beads/core–shell capsules for waste removal |
| Homogenous signal | Screening | Droplet cross-talk, reporter leakage | Implementation of hydrophylic reporters for functional assays, increase of surfactant concentration in emulsion |
| Apparent selection bias | Screening | Incubation length that favors fast growers | Use of limiting nutrients in media, tune incubation time to target slow growers, reassess incubation conditions |
VI. Challenges and opportunities for further development
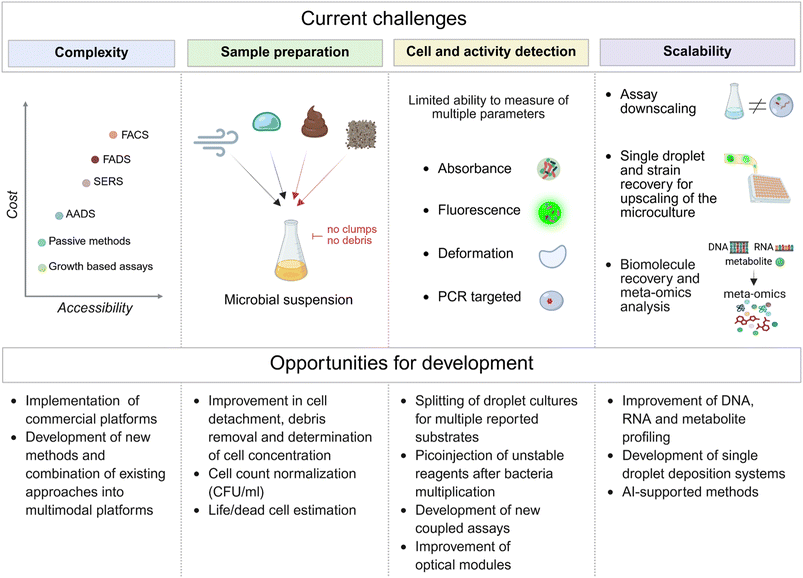
In the last section, we review the most pressing challenges currently facing the field of microbial functional screening using microfluidic technologies. We also highlight key opportunities for researchers to overcome these barriers and advance the development of microdroplet-based platforms, making them more efficient, scalable, and accessible to a broader community of microbiology researchers (Fig. 6).Strategies for cell resuspension from complex biological substrates
Environmental samples present diverse challenges for single cell encapsulation, depending on their origin and composition. Aqueous samples, e.g. from natural reservoirs, are relatively straightforward to process and typically require only basic steps such as shaking, centrifugation, and/or filtration to remove particulates. In contrast, airborne microbiomes may require sample concentration prior to analysis, and methods for separation and detection of microbial cells from these samples still need further optimization.132 The most significant challenges, however, arise from complex matrices such as soil and fecal matter. These sample types contain highly diverse microbial populations embedded in dense, heterogeneous substrates. Microorganisms in these environments tend to form aggregates or adhere to solid particles, complicating the preparation of uniform single cell suspensions and increasing the risk of encapsulating cell clusters instead of individual cells. To overcome these bottlenecks, tailored protocols must be developed based on the specific properties of each sample.133 For instance, the presence of biofilms introduces additional complexity, as cells are embedded in a dense extracellular matrix. Disrupting this matrix is essential for effective resuspension and can be achieved through enzymatic digestion, chemical treatments with detergents, or mechanical disruption methods such as sonication. After disaggregation, thorough washing steps are necessary to remove residual matrix components and debris, resulting in a cleaner bacterial suspension suitable for downstream applications, including droplet generation.Counting strategies and cell encapsulation in microdroplets
Encapsulating single cells from unknown and complex samples, such as those derived from environmental sources, is a challenging but essential step that requires optimization of both cell resuspension and accurate enumeration methods. Direct counting typically involves visualizing bacterial cells under a microscope. This is commonly performed using a small volume of the suspension loaded into a counting chamber, such as a Petroff-Hausser or Thoma chambers, where cells are manually or automatically enumerated. This straightforward technique offers rapid results and, when combined with viability stains, can distinguish between live and dead cells. However, in highly diverse samples containing bacteria, archaea, and fungi, it becomes difficult to achieve precise enumeration, which complicates subsequent single cell encapsulation. To improve accuracy in single cell droplet encapsulation, we recommend generating calibration curves to determine cell counts in terms of CFU per milliliter or CFU per gram. While conventional plating on solid media is widely used, it may be misleading, as many environmental microbes are not cultivable under standard conditions. Therefore, to estimate the actual number of viable cells and to standardize clonal cultivation in droplets, we suggest performing parallel tests by generating emulsions from various dilutions of the sample. After incubation, the number of droplets containing clonal microcolonies can be counted and converted to CFU per sample. Alternatively, advanced techniques such as impedance flow cytometry can provide rapid and accurate estimates of cell counts in environmental samples.134Limited scope of detection methods
A variety of detection modes have been developed to support functional assays of microbial activity in droplet microfluidic systems. These approaches often rely on fluorescent probes, reporter molecules, dyes, or microfluidic imaging to monitor microbial growth, metabolism, or other cellular functions within individual droplets. In addition, passive and label-free sorting methods have been described, utilizing properties such as surface tension,50 viscoelasticity,51,91 or droplet size135,136 to enable selective enrichment. Despite these advancements, the range of effective detection modes for screening microbial consortia remains limited, with most assays relying on fluorescence or absorbance measurement. Functional detection of microbial activity in droplets presents two major challenges. The first is detection accuracy. Implementing conventional optical assays, such as absorbance or optical density measurements, at the picoliter scale is technically difficult. These methods are often not scalable to droplet volumes used in microfluidics, requiring the development of new chip designs and optimized detection strategies.137 The second challenge is multiplexing capability. Simultaneous measurement of multiple parameters or analytes within a single droplet is hindered by limited droplet volume, signal sensitivity, and potential interference between various signals, for example from fluorophores with overlapping spectra. As a result, designing comprehensive assays that capture multiple aspects of microbial function in droplet systems remains a significant technical hurdle. Recent advances, such as the NOVAsort droplet microfluidic platform, which integrates droplet fluorescence and droplet size detection via light scattering, demonstrate that combining multiple parameters can enhance signal discrimination and sorting fidelity. Although this approach has not yet been applied to microbial screening, it represents a promising direction for environmental applications, where challenges such as droplet merging or shrinkage, as well as mass transfer issues and emulsion polydispersity, can all hinder the execution of droplet-based HTS.138Culture upscaling after selection
Droplet screening has gained significant momentum over the past decade, driving a growing demand for reliable methods to recover genetic material from individual sorted droplets for downstream studies and applications. Advances in genomics have enabled droplet barcoding and in-droplet PCR, making it possible to label and track individual positive clones efficiently. New approaches such as biotinylated TaqMan probes can offer an alternative strategy, enabling the selective targeting and recovery of droplets containing specific microbial species or genes.133 In microbiological workflows, however, it remains critical to cultivate cells at larger volumes and ideally store retrieved positive isolates after screening. Combining genetic destructive assays, such as PCR, with cell recovery requires splitting each droplet into two or more replicates while maintaining the genotype–phenotype link. This can be achieved using a SlipChip, a method that has been successfully applied to the targeted isolation of rare gut bacteria.139 Alternatively, single droplets containing colonies can first be deposited, then expanded, and finally analyzed once the cell population is sufficient for various assays at macroscale. One of such demonstrated approaches involves displacing individual droplet cultures into separate wells of a multiwell plate,61 where each sorted clone can then be grown under laboratory conditions.140 Alternatively, droplets can be deposited in a circular array on a petri dish covered with oil. This format allows individual sessile droplets to be picked for further cultivation or analysis.60 However, these are early demonstrations of relatively complex experimental setups that are not yet ready for widespread implementation. Further microfluidics research is needed to further expand and develop new generations of droplet deposition technologies compatible with various droplet sorting modules.Access to microfluidic technologies
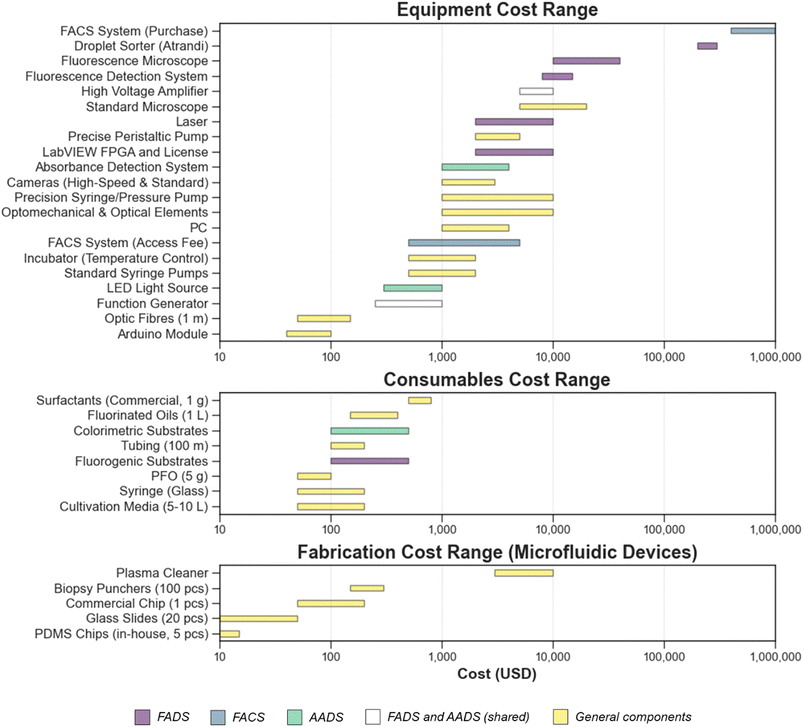
Despite the low cost of individual microfluidic chips, the instrumentation required to operate droplet-based systems can still be expensive and, in some cases, prohibitive for small laboratories (Fig. 7). Even though monodisperse droplets can be generated using low-cost methods,141 other microfluidic techniques, such as picoinjection and sorting, still require a substantial initial investment in specialized equipment. Access to droplet microfluidics is steadily improving, driven by the growing availability of commercial platforms, open-source hardware designs, and user-friendly software tools. However, limited access to advanced instrumentation and microfabrication facilities in certain regions or institutions continues to pose a barrier to broader adoption. The complexity and cost of equipment for droplet-based experiments can vary widely. Basic systems may include components such as syringe pumps and standard microfluidic chips, while more advanced setups integrate high speed cameras, precision controllers, and automated modules. As a result, total system costs can range from several hundred to several hundred thousand dollars, depending on functionality and scale. One potential limitation of droplet-microfluidic systems is the limited supply of fluorinated oils, which key manufacturers, such as 3M, are phasing out alongside other PFAS (per- and polyfluoroalkyl substances) products. In response to 3M's announcement, alternative fluorinated oils compatible with commonly used fluorosurfactants have appeared on the market, helping to mitigate this issue. Nevertheless, an increase in oil prices is still anticipated. However droplet microfluidics is unlikely to be significantly affected, as the low volumes of oil used per assay mean that assay reagents and the fluorosurfactant will still continue to be the highest-cost compounds. Despite the upfront costs, droplet microfluidics provide substantial long-term benefits, including reduced reagent consumption, smaller sample volumes, and significantly increased experimental throughput compared to traditional benchtop methods. The capacity to conduct ultra-high throughput, parallelized experiments enables faster discovery and optimization, making the technology cost effective over time. Combining microfluidics with FACS further enhances efficiency by enabling sorting and deposition of individual target cells based on fluorescent markers or functional activity, allowing the isolation of rare or high performing strains for biotechnological applications.97,98 As in other fields, ongoing technological advancements and increased accessibility are expected to reduce operational barriers and costs, expanding the reach of this innovative screening approach.VII. Future applications and perspectives
Droplet microfluidic platforms are increasingly being integrated with single-cell genomics or transcriptomics and mass-spectrometry-based metabolomics, generating rich datasets that link genotype to phenotype. These data could feed into predictive models for strain design and bioprocess optimization, exemplifying a convergence of microfluidics, omics, and AI. At the same time, broader adoption of these technologies would benefit from greater transparency in reporting device architectures and experimental workflows. Open sharing of microfluidic designs could significantly improve reproducibility and accelerate methodological development within the research community. Beyond transparency, the field would also benefit from a closer alignment between biological objectives and microfabrication protocols. Researchers entering this area frequently face a complex selection problem: choosing among different droplet-based systems and identifying appropriate protocols for a given cell type or assay chemistry to meet specific experimental needs.Combination with genomics and transcriptomics
The integration of microfluidics with genomics, particularly single cell genomics and transcriptomics, has significantly enhanced our ability to study biological systems with high precision.36,142 Recent advances in microfluidic genomics have exemplified this integration. High-throughput single-cell genome sequencing in droplets has enabled strain-resolved profiling of complex microbial communities. The Microbe-seq platform amplified and barcoded over 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 single-amplified genomes from gut microbiome samples and co-assembled nearly 100 bacterial genomes, revealing extensive strain diversity, horizontal gene transfer, and phage-host associations.84 Complementing this, the EASi-seq workflow repurposed a commercial single-cell DNA platform to generate large-scale microbial genome atlases, integrating barcoded reads with metagenomic assemblies to improve strain detection and gene annotation.143 These droplet-based workflows allow efficient separation and analysis of rare or low-abundance microbes, supporting future applications in microbiome monitoring, subpopulation tracking, and multi-omic integration for predictive modeling.
000 single-amplified genomes from gut microbiome samples and co-assembled nearly 100 bacterial genomes, revealing extensive strain diversity, horizontal gene transfer, and phage-host associations.84 Complementing this, the EASi-seq workflow repurposed a commercial single-cell DNA platform to generate large-scale microbial genome atlases, integrating barcoded reads with metagenomic assemblies to improve strain detection and gene annotation.143 These droplet-based workflows allow efficient separation and analysis of rare or low-abundance microbes, supporting future applications in microbiome monitoring, subpopulation tracking, and multi-omic integration for predictive modeling.
In contrast, methods such as bacterial single-cell RNA sequencing (scRNA-seq) remains challenging due to difficult lysis, low and unstable mRNA, and the lack of polyadenylated tails needed for standard eukaryotic protocols. Nonetheless, recent advances have made bacterial scRNA-seq increasingly viable,144 and droplet-based microfluidic platforms now enable high-resolution, single-cell gene expression profiling across diverse environments. These systems also capture dynamic transcriptional responses to stimuli, supporting studies of microbial adaptation and regulation.145 A notable example is BacDrop, which analyzes bacterial populations at scale, revealing heterogeneous antibiotic responses in Klebsiella pneumoniae and identifying mobile-element-driven cellular states linked to resistance evolution.146 BacDrop further resolves antibiotic-induced subpopulations that remain undetectable in bulk RNA-seq. Together, these systems extend microfluidic genomics beyond cultivation, providing scalable access to the genetic structure and heterogeneity of complex microbial communities such as soil microbiota, where diversity and functional complexity are high.
Although genomics and transcriptomics are inherently destructive assays, we anticipate that future high-throughput methods will complement functional droplet screening by using optogenetic barcodes144,147 to link microbial phenotypes and genotypes at scale. Alternatively, selected droplets could be split into two daughter droplets: one sacrificed for genomic analysis and the other preserved for downstream cultivation or upscaling. Implementing such strategies will require more advanced microfluidic modules capable of precise droplet division and routing, creating new opportunities and design challenges for the lab-on-a-chip research community.
Metabolome analysis
While most current research studying the metabolism of microorganisms relies on growth-based functional screening, such as assays for carbohydrate metabolism,11 direct metabolome analysis in droplet-based microfluidic systems remains relatively rare. A recently developed platform combining epifluorescence microscopy coupled with electrospray ionization mass spectrometry (ESI-MS) was used to study secondary metabolites produced by single Streptomyces cells in ∼200 pL droplets, successfully detecting streptomycin produced in situ.144 Similarly, on-chip cultivation of chemically mutagenized strains with ultra-high performance liquid chromatography tandem mass spectrometry (UHPLC-MS/MS) analysis of their metabolites146 illustrates that MS can detect virtually any microbial product or enzymatic reaction output in droplets without labels. More recently, the mass-activated droplet sorting (MADS) system introduced an assay in which each nanoliter droplet is split into two streams, one analyzed by ESI-MS and the other sorted based on the MS result.148 This label-free approach achieved ∼0.7 droplets per second and screened ∼15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 samples in hours, significantly expanding the droplet toolkit for drug discovery and biocatalyst screening. Each droplet thus acts as an independent microreactor, enabling massively parallel single-cell assays that capture phenotypic variability. Despite the limited use of metabolome research, droplet microfluidics provides a promising platform for validating metabolic modeling predictions. Major challenges in microbial metabolome modeling, highlight the need for experimental systems capable of capturing spatial, temporal, and phenotypic variability,149 criteria that are well addressed by droplet microfluidic platforms functioning as high throughput microreactors for single cell or small consortia experiments. The potential value of such systems has been suggested for exploring microbial phenotypic heterogeneity under controlled conditions. In the future, such MS-coupled microreactors could directly validate genome-scale metabolic models at the single-cell level and, in combination with AI-driven analysis, enable data-rich metabolic engineering workflows.
000 samples in hours, significantly expanding the droplet toolkit for drug discovery and biocatalyst screening. Each droplet thus acts as an independent microreactor, enabling massively parallel single-cell assays that capture phenotypic variability. Despite the limited use of metabolome research, droplet microfluidics provides a promising platform for validating metabolic modeling predictions. Major challenges in microbial metabolome modeling, highlight the need for experimental systems capable of capturing spatial, temporal, and phenotypic variability,149 criteria that are well addressed by droplet microfluidic platforms functioning as high throughput microreactors for single cell or small consortia experiments. The potential value of such systems has been suggested for exploring microbial phenotypic heterogeneity under controlled conditions. In the future, such MS-coupled microreactors could directly validate genome-scale metabolic models at the single-cell level and, in combination with AI-driven analysis, enable data-rich metabolic engineering workflows.
Interfacing with artificial intelligence
The consolidation of droplet microfluidics and artificial intelligence (AI) offers exceptional potential to enhance experimental design, data analysis and interpretation in biological research. AI is particularly well suited for processing the vast datasets produced by high-throughput microfluidic systems, enabling the extraction of meaningful insights from complex experiments, as already demonstrated for example in protein engineering.150Artificial intelligence technologies can also improve real-time control and automation. AI-driven systems dynamically tune microfluidic parameters using real-time sensor or imaging feedback, boosting precision, reducing variability, and increasing workflow efficiency. Custom-trained convolutional neural networks (CNNs) further adapt analyses to specific experimental conditions. Platforms such as DropAI151 integrate droplet microfluidics with machine learning to screen thousands of cell-free gene expression reactions while minimizing reagent use. Models predict reaction efficiency, optimize reagent compositions, and generalize results across biological systems via transfer learning. In the coming years, AI methodologies such as deep learning and pattern recognition are expected to further accelerate high-throughput image analysis at the single-droplet level. Convolutional neural networks, in particular, have become indispensable for interpreting images generated in microfluidic experiments. By automatically detecting and classifying complex morphological features, CNNs enable rapid and reliable assessment of biological assay outcomes within droplets. For instance, a YOLOv11 deep learning model have been successfully applied in an environmental screening experiment to determine whether bacteria encapsulated in droplets exhibited proteolytic activity, based on droplet deformation caused by bacterial enzyme secretion.30 Similarly, CNN architectures such as YOLOv4 have been employed for sorting droplets containing mammalian cells.152 In the future, similar approaches could be used to analyze the morphology of microbial colonies within microdroplets, aiding in the selection of those of interest or supporting image-based droplet sorting8 for label free selection of environmental strains.
Looking ahead, the integration of AI and droplet microfluidics is poised to revolutionize experimental automation, adaptive control, and data-driven discovery in bioengineering. Deep learning-based image analysis will continue to play a pivotal role, enabling rapid, high-accuracy interpretation of complex biological phenomena within individual droplets.
VIII. Conclusions
Ultra-high throughput droplet microfluidics is transforming how environmental microbial strains and consortia are cultivated and screened, offering unprecedented resolution for accessing rare, slow growing, and previously uncultivable taxa. Its applications include functional screening for enzymatic activity, antimicrobial resistance, metabolic traits, and interspecies interactions, all performed within massively parallel and miniaturized workflows. The field is advancing rapidly, driven by innovations in droplet generation, surface chemistry, manipulation strategies as well as sorting mechanisms. These developments have expanded droplet formats beyond simple single emulsions and introduced novel sorting methods that enable more sophisticated and tailored screening approaches. Importantly, these innovations are opening opportunities well beyond environmental microbiology. Applications now span biology, chemistry, materials science, and medicine, greatly extending the reach and impact of droplet microfluidics. As the technology continues to mature, establishing standardized, modular workflows will be essential for ensuring reproducibility, scalability, and integration with complementary platforms such as genomics and machine learning. Together, these developments position droplet microfluidics not only as a transformative tool for microbial cultivation and screening, but also as a foundational technology for next generation microbial biotechnology and interdisciplinary research.Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
This research was funded by the National Science Centre, Poland grant SONATA BIS no. 2023/50/E/ST4/00545 and SONATA no. 2024/55/D/NZ9/03060, and TEAM-NET programme of the Foundation for Polish Science, project no. POIR.04.04.00-00-14E6/18-00 as a part of Measure 4.4 of the 2014–2020 Smart Growth Operational Programme, EU. Fig. 1–5 prepared with https://www.biorender.com/ (academic licence, assigned to the University of Warsaw).References
- X. Qian, L. Chen, Y. Sui, C. Chen, W. Zhang and J. Zhou, et al., Biotechnological potential and applications of microbial consortia, Biotechnol. Adv., 2020, 40, 107500, DOI:10.1016/j.biotechadv.2019.107500.
- M. Gavrilescu, Environmental biotechnology: achievements, opportunities and challenges, Dynamic Biochemistry, Process Biotechnology and Molecular Biology, 2010, vol. 4, pp. 1–36, http://www.globalsciencebooks.info/Online/GSBOnline/images/2010/DBPBMB_4(1)/DBPBMB_4(1)1-36o.pdf Search PubMed.
- K. Alain and J. Querellou, Cultivating the uncultured: limits, advances and future challenges, Extremophiles, 2009, 13, 583–594, DOI:10.1007/s00792-009-0261-3.
- S. M. Prakadan, A. K. Shalek and D. A. Weitz, Scaling by shrinking: empowering single-cell “omics” with microfluidic devices, Nat. Rev. Genet., 2017, 18, 345–361, DOI:10.1038/nrg.2017.15.
- A. M. Kaushik, K. Hsieh, L. Chen, D. J. Shin, J. C. Liao and T.-H. Wang, Accelerating bacterial growth detection and antimicrobial susceptibility assessment in integrated picoliter droplet platform, Biosens. Bioelectron., 2017, 97, 260–266, DOI:10.1016/j.bios.2017.06.006.
- Y. Ding, P. D. Howes and A. J. deMello, Recent Advances in Droplet Microfluidics, Anal. Chem., 2020, 92, 132–149, DOI:10.1021/acs.analchem.9b05047.
- Y. J. Sung, J. Y. H. Kim, H. I. Choi, H. S. Kwak and S. J. Sim, Magnetophoretic sorting of microdroplets with different microalgal cell densities for rapid isolation of fast growing strains, Sci. Rep., 2017, 7, 1–11, DOI:10.1038/s41598-017-10764-6.
- W. J. Watterson, M. Tanyeri, A. R. Watson, C. M. Cham, Y. Shan and E. B. Chang, et al., Droplet-based high-throughput cultivation for accurate screening of antibiotic resistant gut microbes, eLife, 2020, 9, e56998, DOI:10.7554/eLife.56998.
- L. Mahler, S. Niehs, K. Martin, T. Weber, K. Scherlach and C. Hertweck, et al., Highly parallelized droplet cultivation and prioritization on antibiotic producers from natural microbial communities, eLife, 2021, 10, e64774, DOI:10.7554/eLife.64774.
- M. Najah, R. Calbrix, I. P. Mahendra-Wijaya, T. Beneyton, A. D. Griffiths and A. Drevelle, Droplet-Based Microfluidics Platform for Ultra-High-Throughput Bioprospecting of Cellulolytic Microorganisms, Chem. Biol., 2014, 21, 1722–1732, DOI:10.1016/j.chembiol.2014.10.020.
- M. M. Villa, R. J. Bloom, J. D. Silverman, H. K. Durand, S. Jiang and A. Wu, et al., Interindividual Variation in Dietary Carbohydrate Metabolism by Gut Bacteria Revealed with Droplet Microfluidic Culture, mSystems, 2020, 5, e00864-19, DOI:10.1128/msystems.00864-19.
- J. Dai, Y. Ouyang, R. Gupte, X. J. A. Liu, Y. Li and F. Yang, et al., Microfluidic droplets with amended culture media cultivate a greater diversity of soil microorganisms, Appl. Environ. Microbiol., 2025, 91, e01794-24, DOI:10.1128/aem.01794-24.
- C. Lauber, M. Strickland, M. Bradford and N. Fierer, The influence of soil properties on the structure of bacterial and fungal communities across land-use types, Soil Biol. Biochem., 2008, 40, 2407–2415, DOI:10.1016/j.soilbio.2008.05.021.
- V. Torsvik, J. Goksøyr and F. Daae, High diversity in DNA of soil bacteria, Appl. Environ. Microbiol., 1990, 56, 782–787, DOI:10.1128/AEM.56.3.782-787.1990.
- S. Liu, C. D. Moon, N. Zheng, S. Huws, S. Zhao and J. Wang, Opportunities and challenges of using metagenomic data to bring uncultured microbes into cultivation, Microbiome, 2022, 10, 76, DOI:10.1186/s40168-022-01272-5.
- T. S. Kaminski, O. Scheler and P. Garstecki, Droplet microfluidics for microbiology: techniques, applications and challenges, Lab Chip, 2016, 16, 2168–2187, 10.1039/C6LC00367B.
- S. Hengoju, M. A. Tovar, D. Man, S. Buchheim and M. Rosenbaum, Droplet Microfluidics for Microbial Biotechnology, Adv. Biochem. Eng./Biotechnol., 2020, 179, 129–157, DOI:10.1007/10_2020_140.
- R. Ostafe, R. Prodanovic, W. Ung, D. Weitz and R. Fischer, A high-throughput cellulase screening system based on droplet microfluidics, Biomicrofluidics, 2014, 8, 041102, DOI:10.1063/1.4886771.
- M. Dhar, J. N. Lam, T. Walser, S. M. Dubinett, M. B. Rettig and D. Di Carlo, Functional profiling of circulating tumor cells with an integrated vortex capture and single-cell protease activity assay, Proc. Natl. Acad. Sci. U. S. A., 2018, 115, 9986–9991, DOI:10.1073/pnas.1803884115.
- M. Gantz, S. Neun, E. J. Medcalf, L. D. van Vliet and F. Hollfelder, Ultrahigh-Throughput Enzyme Engineering and Discovery in In Vitro Compartments, Chem. Rev., 2023, 123, 5571–5611, DOI:10.1021/acs.chemrev.2c00910.
- N. Shembekar, C. Chaipan, R. Utharala and C. A. Merten, Droplet-based microfluidics in drug discovery, transcriptomics and high-throughput molecular genetics, Lab Chip, 2016, 16, 1314–1331, 10.1039/C6LC00249H.
- Z. He, H. Wu, X. Yan and W. Liu, Recent advances in droplet microfluidics for microbiology, Chin. Chem. Lett., 2022, 33, 1729–1742 CrossRef CAS , https://www.sciencedirect.com/science/article/abs/pii/S1001841721006331.
- J.-C. Baret, O. J. Miller, V. Taly, M. Ryckelynck, A. El-Harrak and L. Frenz, et al., Fluorescence-activated droplet sorting (FADS): efficient microfluidic cell sorting based on enzymatic activity, Lab Chip, 2009, 9, 1850–1858, 10.1039/b902504a.
- A. Sciambi and A. R. Abate, Accurate microfluidic sorting of droplets at 30 kHz, Lab Chip, 2015, 15, 47–51, 10.1039/C4LC01194E.
- L. Mazutis, J. Gilbert, W. L. Ung, D. A. Weitz, A. D. Griffiths and J. A. Heyman, Single-cell analysis and sorting using droplet-based microfluidics, Nat. Protoc., 2013, 8, 870–891, DOI:10.1038/nprot.2013.046.
- D. J. Collins, A. Neild, A. DeMello, A.-Q. Liu and Y. Ai, The Poisson distribution and beyond: methods for microfluidic droplet production and single cell encapsulation, Lab Chip, 2015, 15, 3439–3459, 10.1039/C5LC00614G.
- E. M. Payne, D. A. Holland-Moritz, S. Sun and R. T. Kennedy, High-throughput screening by droplet microfluidics: perspective into key challenges and future prospects, Lab Chip, 2020, 20, 2247–2262, 10.1039/D0LC00347F.
- A. Ruszczak, S. Bartkova, M. Zapotoczna, O. Scheler and P. Garstecki, Droplet-based methods for tackling antimicrobial resistance, Curr. Opin. Biotechnol., 2022, 76, 102755, DOI:10.1016/j.copbio.2022.102755.
- S. Neun, P. J. Zurek, T. S. Kaminski and F. Hollfelder, Ultrahigh throughput screening for enzyme function in droplets, Methods in enzymology, Elsevier, 2020, vol. 643, pp. 317–343, DOI:10.1016/bs.mie.2020.06.002.
- Y. Xu, Z. Wang, C. Li, S. Tian and W. Du, Droplet microfluidics: unveiling the hidden complexity of the human microbiome, Lab Chip, 2025, 25, 1128–1148, 10.1039/D4LC00877D.
- S.-Y. Teh, R. Lin, L.-H. Hung and A. P. Lee, Droplet microfluidics, Lab Chip, 2008, 8, 198–220, 10.1039/B715524G.
- M. T. Guo, A. Rotem, J. A. Heyman and D. A. Weitz, Droplet microfluidics for high-throughput biological assays, Lab Chip, 2012, 12, 2146–2155, 10.1039/C2LC21147E.
- L. Potenza, M. S. Andrzejewski and T. S. Kaminski, Induced droplet ovalization (IDO): Image-based microfluidic method for high-throughput and label-free characterization of microbial proteolytic strains from wastewater sludge, Sens. Actuators, B, 2025, 445, 138554, DOI:10.1016/j.snb.2025.138554.
- L. Potenza, L. Drewniak and L. Kozon, Droplet microfluidic multistep platform for high-throughput active selection of cellulolytic microorganisms, Sensors and Actuators Reports, 2026 DOI:10.1016/j.snr.2026.100468.
- J. J. Agresti, E. Antipov, A. R. Abate, K. Ahn, A. C. Rowat and J.-C. Baret, et al., Ultrahigh-throughput screening in drop-based microfluidics for directed evolution, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 4004–4009, DOI:10.1073/pnas.0910781107.
- M. Hosokawa, Y. Nishikawa, M. Kogawa and H. Takeyama, Massively parallel whole genome amplification for single-cell sequencing using droplet microfluidics, Sci. Rep., 2017, 7, 5199, DOI:10.1038/s41598-017-05436-4.
- D. Qin, Y. Xia and G. M. Whitesides, Soft lithography for micro- and nanoscale patterning, Nat. Protoc., 2010, 5, 491–502, DOI:10.1038/nprot.2009.234.
- X. Li, C. Zhang, H. Peng, S. Ma, T. P. Davis and R. Qiao, Design and synthesis of fluorosurfactants for microfluidic droplet-based bioapplications, Polym. Sci. Technol., 2025, 2, 66–78, DOI:10.1021/polymscitech.5c00092.
- J.-C. Baret, Surfactants in droplet-based microfluidics, Lab Chip, 2012, 12, 422–433, 10.1039/C1LC20582J.
- A. Nakamura, Y. Suzuki, N. Homma, Y. Shida, R. Sato and H. Takaku, et al., Ultrahigh-throughput screening of environmental bacteria for proteolytic activity using droplet-based microfluidics, Appl. Environ. Microbiol., 2025, e00109–e00125, DOI:10.1128/aem.00109-25.
- A. M. Kaba, S. Gounel, T. Beneyton, L. Buisson, J.-C. Baret and N. Mano, Confocal Absorbance-Activated Droplet Sorting (cAADS) for Enzyme Engineering. Advanced, Science, 2025, e05324, DOI:10.1002/advs.202505324.
- C. W. Pan, D. G. Horvath, S. Braza, T. Moore, A. Lynch and C. Feit, et al., Sorting by interfacial tension (SIFT): label-free selection of live cells based on single-cell metabolism, Lab Chip, 2019, 19, 1344–1351, 10.1039/C8LC01328D.
- J. Kehe, A. Kulesa, A. Ortiz, C. M. Ackerman, S. G. Thakku and D. Sellers, et al., Massively parallel screening of synthetic microbial communities, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 12804–12809, DOI:10.1073/pnas.1900102116.
- M.-Z. Jiang, H.-Z. Zhu, N. Zhou, C. Liu, C.-Y. Jiang and Y. Wang, et al., Droplet microfluidics-based high-throughput bacterial cultivation for validation of taxon pairs in microbial co-occurrence networks, Sci. Rep., 2022, 12, 18145, DOI:10.1038/s41598-022-23000-7.
- R. H. Hsu, R. L. Clark, J. W. Tan, J. C. Ahn, S. Gupta and P. A. Romero, et al., Microbial Interaction Network Inference in Microfluidic Droplets, Cell Syst., 2019, 9, 229–242.e4, DOI:10.1016/j.cels.2019.06.008.
- C. Gopalakrishnappa, Z. Li and S. Kuehn, Environmental modulators of algae-bacteria interactions at scale, Cell Syst., 2024, 15(838–853), e13, DOI:10.1016/j.cels.2024.08.002.
- C. Lok, Mining the microbial dark matter, Nature, 2015, 522, 270, DOI:10.1038/522270a.
- L. A. Hug, B. J. Baker, K. Anantharaman, C. T. Brown, A. J. Probst and C. J. Castelle, et al., A new view of the tree of life, Nat. Microbiol., 2016, 1, 1–6, DOI:10.1038/nmicrobiol.2016.48.
- A. Jousset, C. Bienhold, A. Chatzinotas, L. Gallien, A. Gobet and V. Kurm, et al., Where less may be more: how the rare biosphere pulls ecosystems strings, ISME J., 2017, 11, 853–862, DOI:10.1038/ismej.2016.174.
- K. Staskiewicz, M. Dabrowska-Zawada, L. Kozon, Z. Olszewska, L. Drewniak and T. S. Kaminski, Droplet microfluidic system for high throughput and passive selection of bacteria producing biosurfactants, Lab Chip, 2024, 24, 1947–1956, 10.1039/D3LC00656E.
- L. Potenza, L. Kozon, L. Drewniak and T. S. Kaminski, Passive Droplet Microfluidic Platform for High-Throughput Screening of Microbial Proteolytic Activity, Anal. Chem., 2024, 96, 15931–15940, DOI:10.1021/acs.analchem.4c02979.
- S. S. Terekhov, I. V. Smirnov, M. V. Malakhova, A. E. Samoilov, A. I. Manolov and A. S. Nazarov, et al., Ultrahigh-throughput functional profiling of microbiota communities, Proc. Natl. Acad. Sci. U. S. A., 2018, 115, 9551–9556, DOI:10.1073/pnas.1811250115.
- D. K. Button, F. Schut, P. Quang, R. Martin and B. R. Robertson, Viability and isolation of marine bacteria by dilution culture: theory, procedures, and initial results, Appl. Environ. Microbiol., 1993, 59, 881–891, DOI:10.1128/aem.59.3.881-891.1993.
- S. Wai, Y. Mizunoe, A. Takade and S. Yoshida, A comparison of solid and liquid media for resuscitation of starvation-and low-temperature-induced nonculturable cells of Aeromonas hydrophila, Arch. Microbiol., 2000, 173, 307–310, DOI:10.1007/s002030000142.
- N. Hanišáková, M. Vítězová and S. K.-M. Rittmann, The historical development of cultivation techniques for methanogens and other strict anaerobes and their application in modern microbiology, Microorganisms, 2022, 10, 412, DOI:10.3390/microorganisms10020412.
- K. Zengler, G. Toledo, M. Rappé, J. Elkins, E. J. Mathur and J. M. Short, et al., Cultivating the uncultured, Proc. Natl. Acad. Sci. U. S. A., 2002, 99, 15681–15686, DOI:10.1073/pnas.252630999.
- B. L. Wang, A. Ghaderi, H. Zhou, J. Agresti, D. A. Weitz and G. R. Fink, et al., Microfluidic high-throughput culturing of single cells for selection based on extracellular metabolite production or consumption, Nat. Biotechnol., 2014, 32, 473–478, DOI:10.1038/nbt.2857.
- J. Kehe, A. Ortiz, A. Kulesa, J. Gore, P. C. Blainey and J. Friedman, Positive interactions are common among culturable bacteria, Sci. Adv., 2021, 7, eabi7159, DOI:10.1126/sciadv.abi7159.
- Z. Yu, K. Geisler, T. Leontidou, R. E. B. Young, S. E. Vonlanthen and S. Purton, et al., Droplet-based microfluidic screening and sorting of microalgal populations for strain engineering applications, Algal Res., 2021, 56, 102293, DOI:10.1016/j.algal.2021.102293.
- C.-Y. Jiang, L. Dong, J.-K. Zhao, X. Hu, C. Shen and Y. Qiao, et al., High-Throughput Single-Cell Cultivation on Microfluidic Streak Plates, Appl. Environ. Microbiol., 2016, 82, 2210–2218, DOI:10.1128/AEM.03588-15.
- T. Weber, S. Hengoju, A. Samimi, M. Roth, M. Tovar and M. A. Rosenbaum, Recovery and isolation of individual microfluidic picoliter droplets by triggered deposition, Sens. Actuators, B, 2022, 369, 132289, DOI:10.1016/j.snb.2022.132289.
- E. J. Medcalf, M. Gantz, T. S. Kaminski and F. Hollfelder, Ultra-High-Throughput Absorbance-Activated Droplet Sorting for Enzyme Screening at Kilohertz Frequencies, Anal. Chem., 2023, 95, 4597–4604, DOI:10.1021/acs.analchem.2c04144.
- F. Gielen, R. Hours, S. Emond, M. Fischlechner, U. Schell and F. Hollfelder, Ultrahigh-throughput–directed enzyme evolution by absorbance-activated droplet sorting (AADS), Proc. Natl. Acad. Sci. U. S. A., 2016, 113, E7383–E7389, DOI:10.1073/pnas.1606927113.
- X. Niu, S. Gulati, J. B. Edel and A. J. Demello, Pillar-induced droplet merging in microfluidic circuits, Lab Chip, 2008, 8, 1837–1841, 10.1039/B813325E.
- Y. Marcy, C. Ouverney, E. M. Bik, T. Lösekann, N. Ivanova and H. G. Martin, et al., Dissecting biological “dark matter” with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth, Proc. Natl. Acad. Sci. U. S. A., 2007, 104, 11889–11894, DOI:10.1073/pnas.0704662104.
- H. N. Joensson, M. Uhlén and H. A. Svahn, Droplet size based separation by deterministic lateral displacement—separating droplets by cell-induced shrinking, Lab Chip, 2011, 11, 1305–1310, 10.1039/C0LC00688B.
- L. Boitard, D. Cottinet, C. Kleinschmitt, N. Bremond, J. Baudry and G. Yvert, et al., Monitoring single-cell bioenergetics via the coarsening of emulsion droplets, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 7181–7186, DOI:10.1073/pnas.1200894109.
- K. Ahn, J. Agresti, H. Chong, M. Marquez and D. A. Weitz, Electrocoalescence of drops synchronized by size-dependent flow in microfluidic channels, Appl. Phys. Lett., 2006, 88, 264105, DOI:10.1063/1.2218058.
- P. Zhu and L. Wang, Passive and active droplet generation with microfluidics: a review, Lab Chip, 2017, 17, 34–75, 10.1039/C6LC01018K.
- Z. Z. Chong, S. H. Tan, A. M. Gañán-Calvo, S. B. Tor, N. H. Loh and N.-T. Nguyen, Active droplet generation in microfluidics, Lab Chip, 2016, 16, 35–58, 10.1039/C5LC01012H.
- M. N. Hatori, S. C. Kim and A. R. Abate, Particle-templated emulsification for microfluidics-free digital biology, Anal. Chem., 2018, 90, 9813–9820, DOI:10.1021/acs.analchem.8b01759.
- P. J. Zurek, R. Hours, U. Schell, A. Pushpanath and F. Hollfelder, Growth amplification in ultrahigh-throughput microdroplet screening increases sensitivity of clonal enzyme assays and minimizes phenotypic variation, Lab Chip, 2021, 21, 163–173, 10.1039/D0LC00830C.
- L. Mahler, M. Tovar, T. Weber, S. Brandes, M. M. Rudolph and J. Ehgartner, et al., Enhanced and homogeneous oxygen availability during incubation of microfluidic droplets, RSC Adv., 2015, 5, 101871–101878, 10.1039/C5RA20118G.
- A. Huebner, D. Bratton, G. Whyte, M. Yang, A. J. Demello and C. Abell, et al., Static microdroplet arrays: a microfluidic device for droplet trapping, incubation and release for enzymatic and cell-based assays, Lab Chip, 2009, 9, 692–698, 10.1039/B813709A.
- S. L. Pratt, G. K. Zath, T. Akiyama, K. S. Williamson, M. J. Franklin and C. B. Chang, DropSOAC: stabilizing microfluidic drops for time-lapse quantification of single-cell bacterial physiology, Front. Microbiol., 2019, 10, 2112, DOI:10.3389/fmicb.2019.02112.
- L. Hu, R. Luo, D. Wang, F. Lin, K. Xiao and Y. Kang, SERS-based microdroplet platform for high-throughput screening of Escherichia coli strains for the efficient biosynthesis of D-phenyllactic acid, Front. Bioeng. Biotechnol., 2024, 12, 1470830, DOI:10.3389/fbioe.2024.1470830.
- C. B. Chang, J. N. Wilking, S. Kim, H. C. Shum and D. A. Weitz, Monodisperse emulsion drop microenvironments for bacterial biofilm growth, Small, 2015, 11, 3954–3961, DOI:10.1002/smll.201403125.
- J. M. Holstein, C. Gylstorff and F. Hollfelder, Cell-free Directed Evolution of a Protease in Microdroplets at Ultrahigh Throughput, ACS Synth. Biol., 2021, 10, 252–257, DOI:10.1021/acssynbio.0c00538.
- J. Clausell-Tormos, A. D. Griffiths and C. A. Merten, An automated two-phase microfluidic system for kinetic analyses and the screening of compound libraries, Lab Chip, 2010, 10, 1302–1307, 10.1039/B921754A.
- A. R. Abate, T. Hung, P. Mary, J. J. Agresti and D. A. Weitz, High-throughput injection with microfluidics using picoinjectors, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 19163–19166, DOI:10.1073/pnas.1006888107.
- M. Sun, S. S. Bithi and S. A. Vanapalli, Microfluidic static droplet arrays with tuneable gradients in material composition, Lab Chip, 2011, 11, 3949–3952, 10.1039/C1LC20709A.
- P. Abbyad, R. Dangla, A. Alexandrou and C. N. Baroud, Rails and anchors: guiding and trapping droplet microreactors in two dimensions, Lab Chip, 2011, 11, 813–821, 10.1039/C0LC00104J.
- E. Fradet, P. Abbyad, M. H. Vos and C. N. Baroud, Parallel measurements of reaction kinetics using ultralow-volumes, Lab Chip, 2013, 13, 4326–44330, 10.1039/C3LC50768H.
- W. Zheng, S. Zhao, Y. Yin, H. Zhang, D. M. Needham and E. D. Evans, et al., High-throughput, single-microbe genomics with strain resolution, applied to a human gut microbiome, Science, 2022, 376, eabm1483, DOI:10.1126/science.abm1483.
- C. Leal-Alves, S. Dumont, Z. Deng, S. Alkhaldi, Z. Leung and M. Oeser, et al., A microfluidic multiplex sorter for strain development, Adv. Mater. Technol., 2025, 10, 2401209, DOI:10.1002/admt.202401209.
- W. Verbist, J. Breukers, S. Sharma, I. Rutten, H. Gerstmans and L. Coelmont, et al., SeParate: multiway fluorescence-activated droplet sorting based on integration of serial and parallel triaging concepts, Lab Chip, 2024, 24, 2107–2121, 10.1039/d3lc01075a.
- X. Wang, L. Ren, Y. Su, Y. Ji, Y. Liu and C. Li, et al., Raman-activated droplet sorting (RADS) for label-free high-throughput screening of microalgal single-cells, Anal. Chem., 2017, 89, 12569–12577, DOI:10.1021/acs.analchem.7b03884.
- X. Liu, R. E. Painter, K. Enesa, D. Holmes, G. Whyte and C. G. Garlisi, et al., High-throughput screening of antibiotic-resistant bacteria in picodroplets, Lab Chip, 2016, 16, 1636–1643, 10.1039/C6LC00180G.
- E. Zang, S. Brandes, M. Tovar, K. Martin, F. Mech and P. Horbert, et al., Real-time image processing for label-free enrichment of Actinobacteria cultivated in picolitre droplets, Lab Chip, 2013, 13, 3707–3713, 10.1039/C3LC50572C.
- H. Goto, Y. Kanai, A. Yotsui, S. Shimokihara, S. Shitara and R. Oyobiki, et al., Microfluidic screening system based on boron-doped diamond electrodes and dielectrophoretic sorting for directed evolution of NAD (P)-dependent oxidoreductases, Lab Chip, 2020, 20, 852–861, 10.1039/C9LC01263J.
- M. Muta, W. Kawakubo, D. H. Yoon, D. Tanaka, T. Sekiguchi and S. Shoji, et al., Deformability-Based Microfluidic Microdroplet Screening to Obtain Agarolytic Bacterial Cells, Anal. Chem., 2023, 95, 16107–16114, DOI:10.1021/acs.analchem.3c02174.
- E. Rousounelou, S. Schmitt, L. Pestalozzi, M. Held, T. Roberts and S. Panke, Ultra-high throughput screening for novel protease specificities, Methods Enzymol., 2020, 644, 169–189, DOI:10.1016/bs.mie.2020.06.005.
- A. C. Hatch, A. Patel, N. R. Beer and A. P. Lee, Passive droplet sorting using viscoelastic flow focusing, Lab Chip, 2013, 13, 1308–1315, 10.1039/c2lc41160a.
- S. H. Jin, B. Lee, J. S. Kim and C.-S. Lee, Improvement strategy of a microfluidic sorter using a pneumatic bilayer valve, Chem. Eng. Sci., 2021, 245, 116834, DOI:10.1016/j.ces.2021.116834.
- H. J. Choi, G. Ahn, U. S. Yu, E. J. Kim, J.-Y. Ahn and J. O. Chan, Pneumatically driven microfluidic platform and fully automated particle concentration system for the capture and enrichment of pathogens, ACS Omega, 2023, 8, 28344–28354, DOI:10.1021/acsomega.3c02264.
- B. Lee, S. H. Jin, J. S. Kim, S. Jang, G. Y. Jung and C.-S. Lee, Pneumatic microvalve-driven multi-way fluorescence-activated sorting platform, Chem. Eng. J., 2026, 172733, DOI:10.1016/j.cej.2026.172733.
- R. Tu, Y. Zhang, E. Hua, L. Bai, H. Huang and K. Yun, et al., Droplet-based microfluidic platform for high-throughput screening of Streptomyces, Commun. Biol., 2021, 4, 647, DOI:10.1038/s42003-021-02186-y.
- M. Oberpaul, S. Brinkmann, M. Marner, S. Mihajlovic, B. Leis and M. A. Patras, et al., Combination of high-throughput microfluidics and FACS technologies to leverage the numbers game in natural product discovery, Microb. Biotechnol., 2022, 15, 415–430, DOI:10.1111/1751-7915.13872.
- D. Yanakieva, A. Elter, J. Bratsch, K. Friedrich, S. Becker and H. Kolmar, FACS-based functional protein screening via microfluidic co-encapsulation of yeast secretor and mammalian reporter cells, Sci. Rep., 2020, 10, 10182, DOI:10.1038/s41598-020-66927-5.
- H. Wu, X. Chen, X. Gao, M. Zhang, J. Wu and W. Wen, High-Throughput Generation of Durable Droplet Arrays for Single-Cell Encapsulation, Culture, and Monitoring, Anal. Chem., 2018, 90, 4303–4309, DOI:10.1021/acs.analchem.8b00048.
- J. Wang, L. Du, Y. Han, D. Zhang and D. Jing, Advancing in situ single-cell microbiological analysis through a microwell droplet array with gradual open sidewall, Lab Chip, 2023, 23, 5165–5172, 10.1039/D3LC00590A.
- I. Sanka, S. Bartkova, P. Pata, O.-P. Smolander and O. Scheler, Investigation of different free image analysis software for high-throughput droplet detection, ACS Omega, 2021, 6, 22625–22634, DOI:10.1021/acsomega.1c02664.
- M. Saar-Abroi, K. Lindpere, D. Kácsor, T. Olman, D. Gonzalez, F. L. Sulp, K. Kiir, I. Sanka, S. Bartkova and O. Scheler, High-throughput bacterial aggregation analysis in droplets, Sci. Rep., 2025, 15, 42980 CrossRef CAS PubMed.
- S. Ki, H. Shin and D.-K. Kang, Integration of wide-field imaging system with droplet microfluidics for monitoring living bacteria, Sens. Actuators, B, 2024, 412, 135765, DOI:10.1016/j.snb.2024.135765.
- S. Hengoju, K. Abdissa, S. T. Boto, A. Samimi, K. Martin and I. D. Jacobsen, et al., A droplet microfluidic strategy for cultivation, investigation, and high-throughput isolation of mouse gut microbiome bacteria, Appl. Environ. Microbiol., 2025, 91, e00695-25, DOI:10.1128/aem.00695-25.
- Y. Meng, S. Li, C. Zhang and H. Zheng, Strain-level profiling with picodroplet microfluidic cultivation reveals host-specific adaption of honeybee gut symbionts, Microbiome, 2022, 10, 1–15, DOI:10.1186/s40168-022-01333-9.
- H.-H. Jeong, S. H. Jin, B. J. Lee, T. Kim and C.-S. Lee, Microfluidic static droplet array for analyzing microbial communication on a population gradient, Lab Chip, 2015, 15, 889–899, 10.1039/C4LC01097C.
- H. Yuan, Y. Zhou, Y. Lin, R. Tu, Y. Guo and Y. Zhang, et al., Microfluidic screening and genomic mutation identification for enhancing cellulase production in Pichia pastoris, Biotechnol. Biofuels Bioprod., 2022, 15, 1–16, DOI:10.1186/s13068-022-02150-w.
- H. Yuan, R. Tu, X. Tong, Y. Lin, Y. Zhang and Q. Wang, Ultrahigh-throughput screening of industrial enzyme-producing strains by droplet-based microfluidic system, J. Ind. Microbiol. Biotechnol., 2022, 49, kuac007, DOI:10.1093/jimb/kuac007.
- K. Saito, Y. Ota, D. M. Tourlousse, S. Matsukura, H. Fujitani and M. Morita, et al., Microdroplet-based system for culturing of environmental microorganisms using FNAP-sort, Sci. Rep., 2021, 11, 9506, DOI:10.1038/s41598-021-88974-2.
- A. S. Tauzin, M. R. Pereira, L. D. Van Vliet, P.-Y. Colin, E. Laville and J. Esque, et al., Investigating host-microbiome interactions by droplet based microfluidics, Microbiome, 2020, 8, 1–20, DOI:10.1186/s40168-020-00911-z.
- A. Rakszewska, J. Tel, V. Chokkalingam and W. T. Huck, One drop at a time: toward droplet microfluidics as a versatile tool for single-cell analysis, NPG Asia Mater., 2014, 6, e133, DOI:10.1038/am.2014.86.
- L. Potenza, V. Smacchia, Ł. Drewniak and T. S. Kaminski, Microfluidic enrichment of proteolytic microbial consortia from sewage sludge, bioRxiv, 2026, preprint, 2026.02.10.705040, DOI:10.64898/2026.02.10.705040.
- D. Chong, X. Liu, H. Ma, G. Huang, Y. L. Han, X. Cui, J. Yan and F. Xu, Advances in fabricating double-emulsion droplets and their biomedical applications, Microfluid. Nanofluid., 2015, 19, 1071–1090, DOI:10.1007/s10404-015-1635-8.
- T. Harada and D. E. Discher, Bubble wrap of cell-like aggregates, Nature, 2011, 471, 172–173, DOI:10.1038/471172a.
- M. Palamoor and M. M. Jablonski, Comparative study on diffusion and evaporation emulsion methods used to load hydrophilic drugs in poly (ortho ester) nanoparticle emulsions, Powder Technol., 2014, 253, 53–62, DOI:10.1016/j.powtec.2013.11.014.
- K. K. Brower, C. Carswell-Crumpton, S. Klemm, B. Cruz, G. Kim and S. G. Calhoun, et al., Double emulsion flow cytometry with high-throughput single droplet isolation and nucleic acid recovery, Lab Chip, 2020, 20, 2062–2074, 10.1039/D0LC00261E.
- J. M. Duarte, I. Barbier and Y. Schaerli, Bacterial microcolonies in gel beads for high-throughput screening of libraries in synthetic biology, ACS Synth. Biol., 2017, 6, 1988–1995, DOI:10.1021/acssynbio.7b00111.
- Z. Zhu and C. J. Yang, Hydrogel droplet microfluidics for high-throughput single molecule/cell analysis, Acc. Chem. Res., 2017, 50, 22–31, DOI:10.1021/acs.accounts.6b00370.
- K. Higashi and N. Miki, Hydrogel Fiber Cultivation Method for Forming Bacterial Cellulose Microspheres, Micromachines, 2018, 9, 36, DOI:10.3390/mi9010036.
- D. Velasco, E. Tumarkin and E. Kumacheva, Microfluidic encapsulation of cells in polymer microgels, Small, 2012, 8, 1633–1642, DOI:10.1002/smll.201102464.
- C. Femmer, M. Bechtold, M. Held and S. Panke, In vivo directed enzyme evolution in nanoliter reactors with antimetabolite selection, Metab. Eng., 2020, 59, 15–23, DOI:10.1016/j.ymben.2020.01.003.
- M. van Zee, J. de Rutte, R. Rumyan, C. Williamson, T. Burnes and R. Radakovits, et al., High-throughput selection of cells based on accumulated growth and division using PicoShell particles, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2109430119, DOI:10.1073/pnas.2109430119.
- H. Sallet, M. Calvo, M. Titus, N. Jacquemin, K. L. Meibom and R. Bernier-Latmani, High-throughput cultivation and isolation of environmental anaerobes using selectively permeable hydrogel capsules, ISME Commun., 2025, 5, ycaf117, DOI:10.1093/ismeco/ycaf117.
- D. Nichols, N. Cahoon, E. M. Trakhtenberg, L. Pham, A. Mehta and A. Belanger, et al., Use of ichip for high-throughput in situ cultivation of “uncultivable” microbial species, Appl. Environ. Microbiol., 2010, 76, 2445–2450, DOI:10.1128/AEM.01754-09.
- J. Park, A. Kerner, M. A. Burns and X. N. Lin, Microdroplet-enabled highly parallel co-cultivation of microbial communities, PLoS One, 2011, 6, e17019, DOI:10.1371/journal.pone.0017019.
- A. E. Dichosa, A. R. Daughton, K. G. Reitenga, M. S. Fitzsimons and C. S. Han, Capturing and cultivating single bacterial cells in gel microdroplets to obtain near-complete genomes, Nat. Protoc., 2014, 9, 608–621, DOI:10.1038/nprot.2014.034.
- A. Manome, H. Zhang, Y. Tani, T. Katsuragi, R. Kurane and T. Tsuchida, Application of gel microdroplet and flow cytometry techniques to selective enrichment of non-growing bacterial cells, FEMS Microbiol. Lett., 2001, 197, 29–33, DOI:10.1111/j.1574-6968.2001.tb10578.x.
- C. Zhang, X. Wu, F. Song, S. Liu, S. Yu and J. Zhou, Core–shell droplet-based microfluidic screening system for filamentous fungi, ACS Sens., 2023, 8, 3468–3477, DOI:10.1021/acssensors.3c01018.
- K. Samlali, C. L. Alves, M. Jezernik and S. C. Shih, Droplet digital microfluidic system for screening filamentous fungi based on enzymatic activity, Microsyst. Nanoeng., 2022, 8, 123, DOI:10.1038/s41378-022-00456-1.
- A. L. McCully, M. Loop Yao, K. K. Brower, P. M. Fordyce and A. M. Spormann, Double emulsions as a high-throughput enrichment and isolation platform for slower-growing microbes, ISME Commun., 2023, 3, 47, DOI:10.1038/s43705-023-00241-9.
- J. Wang, L. Yang, H. Wang and L. Wang, Application of microfluidic chips in the detection of airborne microorganisms, Micromachines, 2022, 13, 1576, DOI:10.3390/mi13101576.
- A. Pryszlak, T. Wenzel, K. W. Seitz, F. Hildebrand, E. Kartal and M. R. Cosenza, et al., Enrichment of gut microbiome strains for cultivation-free genome sequencing using droplet microfluidics, Cells Rep. Methods, 2022, 2, 100137, DOI:10.1016/j.crmeth.2021.100137.
- M. Ling, J. Szarvas, V. Kurmauskaitė, V. Kiseliovas, R. Žilionis and B. Avot, et al., High throughput single cell metagenomic sequencing with semi-permeable capsules: unraveling microbial diversity at the single-cell level in sewage and fecal microbiomes, Front. Microbiol., 2025, 15, 1516656, DOI:10.3389/fmicb.2024.1516656.
- M. Li, M. van Zee, K. Goda and D. Di Carlo, Size-based sorting of hydrogel droplets using inertial microfluidics, Lab Chip, 2018, 18, 2575–2582, 10.1039/C8LC00568K.
- D. H. Yoon, D. Tanaka, T. Sekiguchi and S. Shoji, Size-dependent and property-independent passive microdroplet sorting by droplet transfer on dot rails, Micromachines, 2018, 9, 513, DOI:10.3390/mi9100513.
- C. L. Nogueira, J. C. Sousa, R. Afonso, D. R. Santos, P. Rodrigues and L. Albuquerque, et al., Combining droplet microfluidics and magnetoresistive sensors for the rapid and quantitative detection of Klebsiella pneumoniae in urinary tract infections, Sens. Actuators, B, 2024, 404, 135175, DOI:10.1016/j.snb.2023.135175.
- H. Zhang, R. Gupte, Y. Li, C. Huang, A. R. Guzman and J. J. Han, et al., NOVAsort for error-free droplet microfluidics, Nat. Commun., 2024, 15, 9444 CrossRef CAS PubMed , https://www.nature.com/articles/s41467-024-52932-z.
- L. Ma, J. Kim, R. Hatzenpichler, M. A. Karymov, N. Hubert and I. M. Hanan, et al., Gene-targeted microfluidic cultivation validated by isolation of a gut bacterium listed in Human Microbiome Project's Most Wanted taxa, Proc. Natl. Acad. Sci. U. S. A., 2014, 111, 9768–9773, DOI:10.1073/pnas.1404753111.
- M. Weiss, S. Hasan, R. Genth, M. Mollah, E. Robert and A. Gil, et al., A single droplet dispensing system for high-throughput screening and reliable recovery of rare events, Lab Chip, 2025, 600–612, 10.1039/D4LC00536H.
- N.-N. Deng, Z.-J. Meng, R. Xie, X.-J. Ju, C.-L. Mou and W. Wang, et al., Simple and cheap microfluidic devices for the preparation of monodisperse emulsions, Lab Chip, 2011, 11, 3963–3969, 10.1039/C1LC20629J.
- F. Lan, B. Demaree, N. Ahmed and A. R. Abate, Single-cell genome sequencing at ultra-high-throughput with microfluidic droplet barcoding, Nat. Biotechnol., 2017, 35, 640–646, DOI:10.1038/nbt.3880.
- X. Li, L. Xu, B. Demaree, C. Noecker, J. E. Bisanz and D. W. Weisgerber, et al., Microbiome single cell atlases generated with a commercial instrument, Adv. Sci., 2025, 12, 2409338, DOI:10.1002/advs.202409338.
- A. Tsuchida, T. Kaneko, K. Nishikawa, M. Kawasaki, R. Yokokawa and H. Shintaku, Opto-combinatorial indexing enables high-content transcriptomics by linking cell images and transcriptomes, Lab Chip, 2024, 24, 2287–2297, 10.1039/D3LC00866E.
- K. Wink, L. Mahler, J. R. Beulig, S. K. Piendl, M. Roth and D. Belder, An integrated chip-mass spectrometry and epifluorescence approach for online monitoring of bioactive metabolites from incubated Actinobacteria in picoliter droplets, Anal. Bioanal. Chem., 2018, 410, 7679–7687, DOI:10.1007/s00216-018-1383-1.
- B. Gao, G. Li, L. Zhou, Y. Guo, Q. Tu and J. Wang, On-chip screening of chemically induced mutant strains and assay of their metabolites by liquid chromatography tandem mass spectrometry, Sens. Actuators, B, 2021, 348, 130655, DOI:10.1016/j.snb.2021.130655.
- F. Kawasaki, Y. Mori, T. Mimori, I. Sato and S. Ota, Identification of In-Droplet Multicellular Communities by Light-Induced Combinatorial DNA Barcoding, Chem. – Eur. J., 2023, 29, e202301133, DOI:10.1002/chem.202301133.
- D. A. Holland-Moritz, M. K. Wismer, B. F. Mann, I. Farasat, P. Devine and E. D. Guetschow, et al., Mass activated droplet sorting (mads) enables high-throughput screening of enzymatic reactions at nanoliter scale, Angew. Chem., Int. Ed., 2020, 59, 4470–4477, DOI:10.1002/anie.201913203.
- L. Raajaraam and K. Raman, Modeling microbial communities: perspective and challenges, ACS Synth. Biol., 2024, 13, 2260–2270, DOI:10.1021/acssynbio.4c00116.
- M. Gantz, S. V. Mathis, F. E. H. Nintzel, M. Penner, P. J. Zurek and T. Knaus, Microdroplet screening rapidly profiles a biocatalyst to enable its AI-assisted engineering, bioRxiv, 2025, preprint, 2024.04.08.588565, DOI:10.1101/2024.04.08.588565.
- J. Zhu, Y. Meng, W. Gao, S. Yang, W. Zhu and X. Ji, et al., AI-driven high-throughput droplet screening of cell-free gene expression, Nat. Commun., 2025, 16, 2720, DOI:10.1038/s41467-025-58139-0.
- L. Howell, V. Anagnostidis and F. Gielen, Multi-object detector YOLOv4-tiny enables high-throughput combinatorial and spatially-resolved sorting of cells in microdroplets, Adv. Mater. Technol., 2022, 7, 2101053, DOI:10.1002/admt.202101053.
| This journal is © The Royal Society of Chemistry 2026 |