 Open Access Article
Open Access ArticleProgress toward real-world diagnostic applications of microfluidic paper-based analytical devices (μPADs)
Yohei
Tanifuji
a and
Daniel
Citterio
 *ab
*ab
aDepartment of Applied Chemistry, Faculty of Science and Technology, Keio University, 3-14-1 Hiyoshi, Kohoku-ku, Yokohama 223-8522, Japan. E-mail: citterio@applc.keio.ac.jp; Tel: +81 45 566 1568
bFaculty of Science, Ubon Ratchathani University, Ubon Ratchathani 34190, Thailand
First published on 13th February 2026
Abstract
Since their first report in 2007, microfluidic paper-based analytical devices (μPADs) have continued to gain attention as promising tools for point-of-care diagnostics due to their low cost, portability, ease of operation, and design flexibility. This review summarizes and discusses recent advances in the field, mostly based on works published between 2017 and 2025, with a focus on progress and remaining challenges in bridging the gap between proof-of-concept demonstrations in academic laboratories and real-world implementation. Special emphasis is placed on devices validated with clinical samples and capable of true sample-in–answer-out operation. To comprehensively assess recent developments, nearly one hundred reported examples were analysed not only in terms of analytical figures of merit but also with respect to practical criteria such as real-sample testing, long-term storage stability, the need for off-device sample pretreatment, reagent handling complexity, time-control requirements, and the number of operation steps. In parallel, topics of ongoing academic interest are highlighted, including automated sequential reagent delivery, strategies for accelerating liquid flow, and robust signal readout methods going beyond purely qualitative approaches to enhance assay sensitivity, precision, rapidity, and instrument-free usability. Finally, the review introduces emerging analytical technologies newly integrated into μPAD platforms, such as surface-enhanced Raman scattering (SERS), bioluminescence, CRISPR-based assays, and machine learning-driven data interpretation, which further expand the analytical capabilities and scope of μPADs.
Introduction
Starting from the earliest litmus paper pH tests in the 17th century, paper has been used as a versatile substrate for analytical assays for centuries. The introduction of urinary glucose dipsticks in the 1950s1,2 and lateral flow immunoassays (LFIAs) in the 1980s3,4 represent just two examples for the success in commercialization, emphasizing how a low-cost, mass-producible, and disposable material could be transformed into practical real-world analytical tools, including diagnostic devices. The currently ongoing massive development of paper-based analytical devices (PADs) in academic research laboratories was triggered back in 2007, when the Whitesides group introduced the concept of microfluidic paper-based analytical devices (μPADs) by patterning of fluidic microchannels delimited by hydrophobic barriers on filter paper via photolithography.5 Their pioneering work demonstrated that paper could be used not just as a passive substrate for reagent immobilization, but as an engineered microfluidic platform capable of multiplexed assays, solid–liquid separation, and capillary-driven sample transport, eliminating the need for external pumps or power sources.Following this breakthrough and continuing at present, μPADs-related research has expanded rapidly across various fields of application including medical diagnostics, environmental monitoring and food safety testing, among others.6–8 The relatively low technical and economical barrier to entry in terms of device fabrication and cost, as well as the design flexibility has driven an explosion of proof-of-concept studies in the scientific literature, with the wax printing method as a representative and dominant paper patterning method.9 Reported works cover the full variety of signal detection techniques known from general analytical chemistry adapted to PADs, reaching from colorimetry to electrochemistry, fluorescence, bioluminescence and surface-enhanced Raman scattering, as well as a broad spectrum of targeted analytes ranging from small molecules, electrolytes, metabolites, nucleic acids, to pathogens. However, the enthusiasm and research activities in academia were not exactly matched by the adoption to real-world applications. The majority of reported μPADs have only been evaluated under controlled laboratory conditions, often using simple aqueous solutions or target analyte spiked artificial samples just marginally mimicking real-world situations. This tendency limited the understanding of how such devices would perform when applied to complex, heterogeneous clinical specimens, where factors like viscosity, interfering substances, and matrix variability can significantly alter assay performance.
In 2017, we have published a comprehensive review summarizing and discussing the state-of-the-art in μPADs for medical diagnostics in this journal.10 At that time, the technology was still in a developmental state. It was beyond the initial proof-of-concept stage but mostly not yet mature enough for broad deployment into clinical practice. We systematically analysed the reasons why, despite intense academic research activity, μPADs still lagged in practical uptake compared with already established paper-based diagnostics such as lateral flow immunoassays (LFIAs). Drawing from both the literature and publicly available market information, we identified several critical areas requiring focused improvement:
• Operational complexity and susceptibility to user error.
• Limited validation with real clinical samples and lack of interference studies.
• Insufficiently investigated long-term storage stability.
• Dependence on specialized detection equipment.
• Inconsistent and difficult-to-quantify signal readouts.
• Regulatory hurdles and competition with established diagnostics.
Since that time, the global context for rapid diagnostics has shifted dramatically. The COVID-19 pandemic brought LFIAs into the daily routine of millions worldwide, normalizing self-administered assays and increasing public familiarity with their operation and interpretation.8,11,12 This societal shift has not only lowered the psychological threshold for adopting paper-based devices but also raised expectations for their reliability, convenience, and affordability.
In terms of technological progress, most of the above-listed challenges have seen incremental rather than groundbreaking advances. However, a notable shift has emerged in the general development of μPADs, marked by:
• A growing number of studies evaluating performance using real clinical specimens and thereby investigating potential interference from complex matrices.
• An increasing number of studies aiming at sample-in–answer-out operation, achieved by combining existing microfluidic architectures, detection chemistries, and sample pretreatment modules.
Even without fundamental breakthroughs, several performance-oriented areas have seen active development, particularly:
• Sensitivity and precision: improved through more advanced sequential reagent delivery strategies, with emerging designs automating reagent delivery via passive or active flow-control elements, reducing user intervention steps and improving analytical reproducibility.
• Rapidity: enabled by optimized or alternative fluidic pathways, capillary force control, and preloaded reagents, allowing to obtain results within minutes without sacrificing analytical performance.
• Long-term storage stability: supported by integration of nanozymes as replacement of natural enzymes.
Beyond operational performance, progress has also been made in signal interpretation. Initial μPAD approaches were mostly reliant on external bulky devices (e.g. scanners, electrochemical workstations, etc.) used in combination with computers for quantitative data processing. On the other end of the spectrum remained the qualitative naked eye “yes/no” signal readout schemes (e.g. LFIAS), or simple comparisons with colour reference charts for semi-quantitative interpretation (e.g. urinary dipsticks). Recent years have seen a trend where researchers increasingly aim at either less subjective, hence more reliable semi-quantitative systems, or quantitative analysis without use of bulky specialized equipment. This trend includes:
• (Semi-)quantitative systems: achieved by incorporating internal calibration zones and smartphone-based image analysis, to reduce user subjectivity and improve interpretation near decision thresholds.
In addition to these advancements, newly emerged analytical technologies have been integrated with μPADs, such as:
• Signal detection methods including surface-enhanced Raman scattering (SERS) and bioluminescence.
• CRISPR-based assay techniques.
• Artificial intelligence integration represented by machine learning (ML).
In contrast to recent comprehensive reviews on μPADs,7,13–15 the current critical review focuses on post-2017 developments mainly in the field of medical diagnostics. It highlights attempts to shift toward integrated, application-ready platforms and advancements in sensitivity, rapidity, and semi-quantitative readouts, as well as integration of newly emerged analytical technologies with μPADs. Since the well-developed lateral flow immunoassay approaches are only marginally touched in this work, the reader is referred to related reviews for details of advancements in that field.11,16,17
General progress in the field of μPADs
Validation testing for clinical applications
A growing number of recent studies has moved beyond proof-of-concept demonstrations to report integrated μPAD platforms capable of sample-in–answer-out operation, particularly in but not limited to the field of immunoassays and nucleic acid testing (NAT).18,19 The basic characteristics and the analytical performance of these devices, including the aspects of real sample pretreatment, are summarized in Tables 1–3, along with further details in Tables S1–S3, separated into devices for immunoassays, NAT, and other targets. As it is well known, biofluid samples represent complex matrices containing many compounds potentially interfering with a specific assay, such as for example proteins.20 While the degree of clinical validation varies widely, the approaches employed for clinical sample testing with μPADs can be broadly classified into four categories: (1) spiking known amounts of target analytes into artificial matrices mimicking clinical samples; (2) spiking known amounts of target analytes into real clinical sample matrices to evaluate recovery; (3) diluting clinical specimens with water or buffer; (4) applying off-device pretreated sample fractions such as serum, plasma, or extracted nucleic acid samples; and (5) directly analysing the raw clinical sample without any off-device pretreatment. From a point-of-care testing perspective, the last approach offers the greatest real-world relevance and acts as proof for eliminating the need for off-device sample preparation.| Analyte | Sample matrix | Real sample test | Sample type category | Detection technique | Detection range | Detection limit | Storage stability | Assay time | Off-device treatment | Step # | Reagent handling | External equipment | Ref. |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| TNF-α: tumour necrosis factor-α, IL-6: human interleukin-6, HBV: hepatitis B virus, HCV: hepatitis C virus, cTnI: cardiac troponin I, anti-Bb Abs: anti-Borrelia burgdorferi antibodies, CK-MB: creatine kinase-MB, FABP: fatty acid binding protein, CRP: C-reactive protein, anti-IFN-γ abs: anti-interferon-γ autoantibodies, BNP: brain natriuretic peptide, HIV1: human immunodeficiency virus type 1, HA: hemagglutinin, DEN1: dengue virus type 1, PfHRP2: plasmodium falciparum histidine-rich protein 2, DPV: differential pulse voltammetry, CA: chronoamperometry, EIS: electrochemical impedance spectroscopy, CL: chemiluminescence, FL: fluorescence, BL: bioluminescence, Echem: electrochemistry, AR: augmented reality, MS: mass spectrometry, DL: deep learning, r.t.: room temperature, N.A.: not available; sample type categories are: (1) spiking known amounts of target analytes into artificial matrices mimicking clinical samples; (2) spiking known amounts of target analytes into real clinical sample matrices; (3) diluting clinical specimens with water or buffer; (4) applying off-device pretreated sample fractions; and (5) directly analysing the raw clinical sample without any off-device pretreatment. | |||||||||||||
| TNF-α | Rat tissue sample | Clinical sample (rat) | 5 | Colorimetric | N.A. | 22 pM | N.A. | 31 min | No | 2 | No | Yes | 21 |
| IL-6 | Human saliva | Clinical sample | 5 | Coloured length | 0.05–25.0 pg mL−1 | 0.05 pg mL−1 | 4 °C | 35 min | No | 5 | Yes | No | 44 |
| Human urine | 0.05 to 25.0 pg mL−1 | 0.05 pg mL−1 | <12 weeks | ||||||||||
| SARS-CoV-2 | Exhaled breath | 4 | Echem (DPV) | 3.7–10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 ng mL−1 000 ng mL−1 |
1 ng mL−1 | 4 °C | 60 min | Yes | 4 | Yes | Yes | 202 | |
| (Exhaled breath) | (Exhaled breath) | >33 days | |||||||||||
| HBV | Human serum | 4 | Echem (CA) | 0.1–250 ng mL−1 | 8.2 pg mL−1 | −20 °C | <500 s | Yes | 3 | No | Yes | 69 | |
| HCV | 0.001–250 ng mL−1 | 1.19 pg mL−1 | >30 days | ||||||||||
| SARS-CoV-2 | 4 | Colorimetric | N.A. | 5 nM | N.A. | 15 min | Yes | 5 | Yes | Yes | 203 | ||
| (Serum) | |||||||||||||
| 4 | Colorimetric (DL assisted) | N.A. | N.A. | N.A. | <20 min | Yes | 8 | Yes | Yes | 183 | |||
| (89.5% accuracy, serum) | |||||||||||||
| cTnI | 4 | Colorimetric (DL assisted) | Up to 100 ng mL−1 | 0.2 pg mL−1 | N.A. | <15 min | Yes | 8 | Yes | Yes | 185 | ||
| (Serum) | (Serum) | ||||||||||||
| 4 | CL (DL assisted) | 0.001–100 ng mL−1 | 0.16 pg mL−1 | N.A. | 25 min | Yes | 9 | Yes | Yes | 193 | |||
| (Serum) | (Serum) | ||||||||||||
| Anti-Bb Abs | 4 | Colorimetric (DL assisted) | N.A. | N.A. (sensitivity 90.5%, specificity 87.0%, serum) | N.A. | <15 min | Yes | 6 | No | Yes | 178 | ||
| 4 | Colorimetric (DL assisted) | N.A. | N.A. (sensitivity 95.5%, specificity 100%, serum) | r.t. | <20 min | Yes | 6 | Yes | Yes | 184 | |||
| >60 days | |||||||||||||
| Myoglobin | 4 | FL (DL assisted) | N.A. | 0.52 ng mL−1 | N.A. | <15 min | Yes | 7 | Yes | Yes | 182 | ||
| (Serum) | |||||||||||||
| CK-MB | 0.3 ng mL−1 | ||||||||||||
| (Serum) | |||||||||||||
| FABP | 0.49 ng mL−1 | ||||||||||||
| (Serum) | |||||||||||||
| CRP | Human serum | Diluted clinical sample | 3 | Colorimetric (DL assisted) | 0–10 mg L−1 | N.A. | N.A. | <12 min | Yes | 6 | Yes | Yes | 179 |
| (Serum) | |||||||||||||
| Anti-IFN-γ abs | Human serum | Diluted clinical sample | 3 | Colorimetric | 0.003–3 μg mL−1 | 0.001 μg mL−1 (smartphone) | N.A. | <10 min | Yes | 3 | No | Yes/no | 22 |
| (PBS) | 0.03 μg mL−1 (naked eye) | ||||||||||||
| (PBS) | |||||||||||||
| SARS-CoV-2 | Human saliva | Spiked into biofluid sample | 2 | Colorimetric | 10–160 pg mL−1 | 10 pg mL−1 | N.A. | 8 min | No | 8 | Yes | Yes | 204 |
| (Saliva) | (Saliva) | ||||||||||||
| cTnI | Human whole blood | 2 | Echem (EIS) | 90 pg mL−1 to 90 ng mL−1 | 4.6 pg mL−1 (190 fM) | N.A. | 46 min | No | 9 | Yes | Yes | 55 | |
| (Artificial blood plasma) | (Artificial blood plasma) | ||||||||||||
| BNP | Echem (EIS) | 6 pg mL−1 to 6 ng mL−1 | 1.2 pg mL−1 (40 fM) | ||||||||||
| (Artificial blood plasma) | (Artificial blood plasma) | ||||||||||||
| D-dimer | Echem (EIS) | 91 ng mL−1 to 91 μg mL−1 | 146 pg mL−1 (730 fM) | ||||||||||
| (Artificial blood plasma) | (Artificial blood plasma) | ||||||||||||
| HIV1 | Porcine whole blood | 2 | BL | N.A. | 2.8 nM | −20 °C | 21 min | No | 2 | No | Yes | 23 | |
| (Porcine serum) | >2 months | ||||||||||||
| Hemagglutinin | 7.1 nM | ||||||||||||
| (Porcine serum) | |||||||||||||
| HBV | 19.3 nM | ||||||||||||
| (Porcine serum) | |||||||||||||
| HIV1 | Human plasma | 2 | Colorimetric | 0.03–3 ng mL−1 | 0.03 ng mL−1 | N.A. | <10 min | Yes | 6 | Yes | Yes | 30 | |
| (Plasma) | (Plasma) | ||||||||||||
| α-Fetoprotein | Human serum | Spiked into biofluid sample | 2 | Echem (EIS) | 0.01–500 ng mL−1 (9 min) | 1.65 pg mL−1 (9 min) | 4 °C | 9 min | Yes | 7 | No | Yes | 31 |
| 0.5 pg mL−1 to 10 ng mL−1 (30 min) | 3.54 fg mL−1 (30 min) | <15 days | 30 min | ||||||||||
| Procalcitonin | Synthetic serum | Spiked into artificial biofluid sample | 1 | AR (smartphone) | 2–10 μg mL−1 | 2 μg mL−1 | N.A. | 84 min | Yes | 8 | Yes | Yes | 121 |
| (Serum) | (Serum) | ||||||||||||
| α-Fetoprotein | Buffer | N.A. | — | Echem (CA) | 10–100 ng mL−1 | 0.63 ng mL−1 | N.A. | 33 min | N.A. | 5 | Yes | Yes | 41 |
| (KNO3) | (KNO3) | ||||||||||||
| PfHRP2 | PBS | Spiked into biofluid sample | — | MS | N.A. | 5 nM | N.A. | >10 min | N.A. | 6 | Yes | Yes | 205 |
| (PBS) | |||||||||||||
| Analyte | Sample matrix | Real sample test | Sample type category | Detection technique | Detection range | Detection limit | Storage stability | Assay time | Off-device treatment | Step # | Reagent handling | External equipment | Ref. |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ctDNA: circulating tumour DNA, HPV: human papillomavirus, RSV: respiratory syncytial virus, HBV: hepatitis B virus, miRNA: micro-RNA, HIV1: human immunodeficiency virus type 1, MRSA: methicillin-resistant Staphylococcus aureus, SWV: square wave voltammetry, EIS: electrochemical impedance spectroscopy, FL: fluorescence, BL: bioluminescence, Echem: electrochemistry, r.t.: room temperature, N.A.: not available; sample type categories: (1) spiking known amounts of target analytes into artificial matrices mimicking clinical samples; (2) spiking known amounts of target analytes into real clinical sample matrices; (3) diluting clinical specimens with water or buffer; (4) applying off-device pretreated sample fractions; and (5) directly analysing the raw clinical sample without any off-device pretreatment. | |||||||||||||
| ctDNA | Mice serum | Clinical sample (mice) | 4 | Colorimetric | 200–1500 pM | 95.81 pM | 4 °C | 63 min | Yes | 6 | Yes | Yes | 56 |
| (Serum) | (Serum) | >15 days | |||||||||||
| Pressure | 79.88 pM | ||||||||||||
| (Serum) | |||||||||||||
| SARS-CoV-2 | Human saliva | Clinical sample | 5 | Colorimetric | N.A. | 50 copies μL−1 | N.A. | 60 min | No | 5 | Yes | Yes | 24 |
| (Saliva) | |||||||||||||
| Swab | 4 | FL | 102–104 copies | 102 copies | r.t. | 40 min | Yes | 4 | Yes | Yes | 39 | ||
| >30 days | |||||||||||||
| 4 | Echem (amperometry) | 0.05–100 nM | 0.12 pM | 4 °C | 52 min | Yes | 5 | No | Yes | 79 | |||
| (PBS) | (PBS) | >4 weeks | |||||||||||
| SARS-CoV-2 | 4 | Colorimetric | N.A. | 400 copies per μl | N.A. | <30 min | Yes | 5 | Yes | Yes | 28 | ||
| (Alpha, beta, delta, and gamma) | (Smartphone) | ||||||||||||
| 2400 copies per μl | |||||||||||||
| (Naked eye) | |||||||||||||
| HPV | 4 | FL | 1–105 fM | 1 fM | N.A. | 35 min | Yes | 6 | Yes | Yes | 59 | ||
| HPV 14 subtypes | 4 | FL | N.A. | 1–100 aM | N.A. | 40 min | Yes | 3 | Yes | Yes | 169 | ||
| Influenza virus subtype H1N1 | 4 | Echem (amperometry) | 0.05–1000 nM | 0.35 pM | 4 °C | 52 min | Yes | 5 | Yes | Yes | 79 | ||
| (PBS) | (PBS) | >4 weeks | |||||||||||
| RSV | 10–1000 nM | 0.36 pM | |||||||||||
| (PBS) | (PBS) | ||||||||||||
| HBV | Human serum | 4 | Echem (SWV) | 10 pM to 2 μM | 7.23 pM | 4 °C | 7 min | Yes | 2 | No | Yes | 38 | |
| (PBS) | (PBS) | <1 month | |||||||||||
| Dengue virus | 4 | Colorimetric | N.A. | 5 copies | <30 °C | ∼30 min | Yes | 9 | Yes | Yes | 58 | ||
| (Serum) | >30 h | ||||||||||||
| 4 | FL | N.A. | 10 copies | N.A. | <1 h | Yes | 8 | Yes | Yes | 206 | |||
| (Serum) | |||||||||||||
| Zika virus | 4 | ||||||||||||
| Let-7a (miRNA) | Diluted clinical sample | 3 | BL | 2 fM to 50 pM | 1.7 fM | r.t. | 150 min | Yes | 4 | Yes | Yes | 86 | |
| >3 months | |||||||||||||
| SARS-CoV-2 | Human saliva | Spiked into biofluid sample | 2 | BL | N.A. | 40 nM | r.t. | 7–12 min | (Yes) | 3 | No | Yes | 150 |
| (Saliva) | <7 weeks | ||||||||||||
| Influenza A virus | 2 | Colorimetric | N.A. | 50 copies per μL | N.A. | 60 min | No | 5 | Yes | Yes | 24 | ||
| (Saliva) | |||||||||||||
| Influenza B virus | |||||||||||||
| HIV1 | Human whole blood | Spiked into biofluid sample | 2 | FL | N.A. | 3 × 103 copies per mL | N.A. | 30 min (on-chip) | Yes | 7 | Yes | Yes | 207 |
| (Whole blood) | >10 min (post amplification) | ||||||||||||
| MRSA | Blood | Spiked into biofluid sample | 2 | FL | 1–104 copies (35 min) | 1 copy (42 min) | r.t. | 36 or 43 min | Yes | 4 | No | Yes | 208 |
| 105–108 copies (42 min) | >35 days | ||||||||||||
| Chikungunya | Human serum | Clinical sample | 2 | FL | N.A. | 10 copies | N.A. | <1 h | Yes | 8 | Yes | Yes | 206 |
| (Serum) | |||||||||||||
| SARS-CoV-2 | N.A. (aerosol generated by breathing simulator) | Spiked into artificial biofluid sample | 1 | Colorimetric | N.A. | 17 aM | N.A. | 30 min (sample collection) | No | 2 | No | No | 25 |
| ∼1.5 h (on-chip) | |||||||||||||
| Five pathogens (E. coli, S. aureus, P. aeruginosa, K. pneumoniae, A. baumannii) | Surface swab (mimicking bacterial contamination on bench) | 1 | FL | 100–106 copies per μL | 1 copy per μL | N.A. | <60 min | No | 5 | No | Yes | 209 | |
| (PBS) | (PBS) | ||||||||||||
| HIV1 | Human whole blood | Spiked into diluted biofluid sample | 1 | Colorimetric | N.A. | 3 × 105 copies (whole blood) | r.t. | 90 min | Yes | 4 | No | Yes | 210 |
| >3 weeks | |||||||||||||
| Mpox virus | Artificial urine or saliva | 1 | Colorimetric | 10–1000 copies per mL | ≈5 copies per mL | 55 °C | <30 min | Yes | 5 | Yes | Yes | 90 | |
| (Urine, saliva) | (Urine, saliva) | >30 days | |||||||||||
| HPV16 | N.A. | N.A. | — | Echem (SWV) | N.A. | 1 copy per μL | N.A. | 130 min | Yes | 6 | Yes | Yes | 211 |
| HPV | Water | — | Echem (personal glucose meter) | N.A. | 57 pM | −20 °C | 120 min | (Yes) | 4 | No | Yes | 177 | |
| >4 weeks | |||||||||||||
| Ebola virus | Water | — | Coloured length | N.A. | 11 aM | N.A. | 285 min | Yes | 4 | Yes | Yes | 171 | |
| Analyte | Sample matrix | Real sample test | Sample type category | Detection technique | Detection range | Detection limit | Storage stability | Assay time | Off-device treatment | Step # | Reagent handling | External equipment | Ref. |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| cTnI: cardiac troponin I, ALP: alkaline phosphatase, Dam MTase: DNA adenine methylation MTase, NADPH: nicotinamide adenine dinucleotide phosphate, FL: fluorescence, echem: electrochemistry, CA: chronoamperometry, r.t.: room temperature, N.A.: not available; sample type categories: (1) spiking known amounts of target analytes into artificial matrices mimicking clinical samples; (2) spiking known amounts of target analytes into real clinical sample matrices; (3) diluting clinical specimens with water or buffer; (4) applying off-device pretreated sample fractions; and (5) directly analysing the raw clinical sample without any off-device pretreatment. | |||||||||||||
| Uric acid | Human saliva | Clinical sample | 4 | Colorimetric | 0–214.8 ppm | 0.1 ppm | 4 °C | <10 min | Yes | 4 | No | Yes | 92 |
| (Artificial saliva) | (Artificial saliva) | >3 months | |||||||||||
| Hemoglobin | 5 | Colorimetric | 1.57–2000 μg mL−1 | 1.57 μg mL−1 | r.t. | 3 min | No | 2 | No | Yes | 195 | ||
| >3 weeks | |||||||||||||
| Coloured distance | 3–200, 200–2000 μg mL−1 | 3 μg mL−1 | |||||||||||
| Albumin | Human urine | 5 | Coloured distance | 0–1000 mg L−1 | N.A. | r.t. | 15 min | No | 1 | No | No | 212 | |
| (Artificial urine) | >2 months | ||||||||||||
| Creatinine | 50.0–750 μg L−1 | ||||||||||||
| (Artificial urine) | |||||||||||||
| Glucose | Human whole blood | 5 | Colorimetric | 2.5–20 mM | 0.3 mM | N.A. | 10 min | No | 2 | No | Yes | 213 | |
| (Whole blood) | (Whole blood) | ||||||||||||
| 5 | Colorimetric | 50–365 mg dL−1 | N.A. | N.A. | 30 min | No | 3 | No | Yes | 214 | |||
| (PBS) | |||||||||||||
| 5 | Colorimetric | 2.20–15.00 mM | 0.12 mM | −4 °C | 12 min | No | 2 | No | Yes | 215 | |||
| (Whole blood) | (Whole blood) | <40 days | |||||||||||
| Urea | 5 | Colorimetric | 2.46–38.14 mM | 0.03 mM | 35 °C | <2 min | No | 1 | No | Yes | 216 | ||
| (Whole blood) | (Whole blood) | >8 days | |||||||||||
| Uric acid | 5 | Colorimetric | 0–1.5 mM | 0.1127 mmol L−1 | 4 °C | >10 min | No | 3 | No | Yes | 83 | ||
| (Whole blood) | (Whole blood) | <28 days | |||||||||||
| Creatinine | 0–100 μM | 0.2978 μmol L−1 | >13 min | ||||||||||
| (Whole blood) | (Whole blood) | ||||||||||||
| Albumin | 0–100 mg mL−1 | 0.7696 mg mL−1 | >6 min | ||||||||||
| (Whole blood) | (Whole blood) | ||||||||||||
| Hemoglobin | 5 | Colorimetric | 2.5–15 g dL−1 | 2.5 g dL−1 | r.t. | <5 min | No | 2 | No | Yes | 29 | ||
| >8 h | |||||||||||||
| cTnI | 4 | Coloured distance | 0.025–2.5 ng mL−1 | 0.025 ng mL−1 | r.t. | 15–30 min | Yes | 2 | No | No | 85 | ||
| (Whole blood) | (Whole blood) | >10 weeks | |||||||||||
| Urea | Human saliva | Diluted clinical sample | 3 | Colorimetric | 0.10–5.0 mM | 0.032 mM | r.t. | 35 min | Yes | 3 | No | Yes | 84 |
| <1 months | |||||||||||||
| NHx | 3 | 0.16–5.0 mM | 0.049 mM | r.t. | 15 min | Yes | |||||||
| <2 months | |||||||||||||
| Zinc(II) | Human urine | Spiked into biofluid sample | 2 | Colorimetric | 50.0–750 μg L−1 | 10 μg L−1 | r.t. | >5 min | Yes | 2 | No | Yes | 217 |
| (Urine) | (Urine) | <7 days | |||||||||||
| Histidine | Human urine | 2 | FL-pixel | 0.5–20 mM | 0.021 mM | N.A. | >10 min | No | 5 | Yes | Yes | 218 | |
| FL-distance | 0.5 mM | ||||||||||||
| Glucose | Human serum | 2 | Colorimetric | 5–11 mM | 0.3 mM | N.A. | 5 min (buffer) | (Yes) | 2 | No | Yes | 78 | |
| (Serum) | (PBS) | 8 min (serum) | |||||||||||
| Cholesterol | 2.6–6.7 mM | 0.2 mM | |||||||||||
| (Serum) | (PBS) | ||||||||||||
| Triglyceride | 1–2.3 mM | 0.3 mM | |||||||||||
| (Serum) | (PBS) | ||||||||||||
| Secnidazole | 2 | FL | 0.1–300 μM | N.A. | N.A. | N.A. | (Yes) | 3 | No | Yes | 219 | ||
| (117 nM in bulk) | |||||||||||||
| ALP | Calf, human, goat serum | 2 | Echem (CA) | N.A. | 1.4 U L−1 | r.t. | 20 min (pre-reaction) | Yes | 3 | Yes | Yes | 211 | |
| >4 weeks | 20 min (flow measurement) | ||||||||||||
| H2O2 | Artificial saliva | Spiked into artificial biofluid sample | 1 | Echem (CA) | 0.25 μM to 5 mM | 0.15 μM | >4 weeks | 110 s | No | 2 | No | Yes | 220 |
| Glucose | 1 | 2.5 μM to 3 mM | 1.5 μM | 4 °C | |||||||||
| <4 weeks | |||||||||||||
| 1 | Coloured distance (barcode) | 15–200 mg dL−1 | 15 mg dL−1 | N.A. | 90 min | No | 3 | No | Yes | 117 | |||
| (Artificial saliva) | (Artificial saliva) | ||||||||||||
| Artificial urine | 1 | Colorimetric | 0–20 mM | 1 mM | N.A. | 10–30 min | No | 1 | No | No | 120 | ||
| (Artificial urine) | (Artificial urine) | ||||||||||||
| Ascorbic acid | 1 | Echem (CA) | 0.15–0.8 mM | 92.8 μM | N.A. | 250 s | No | 2 | No | Yes | 41 | ||
| Lithium ion | Human whole blood | 2 | Colorimetric | 0–2.0 mM | 0.281 mM | N.A. | 10 s | No | 3 | No | (Yes) | 221 | |
| (Whole blood) | (Whole blood) | ||||||||||||
| Dam MTase | Human serum | Spiked into diluted biofluid sample | 2 | FL | 1.0–90 U mL−1 | 0.11 U mL−1 | >10 days | 35 min | (Yes) | 4 | Yes | Yes | 60 |
| NADPH | Simulated urine | Spiked into diluted biofluid sample | 2 | Echem (CA) | 0.01–1.0 mM, 1.0–4.0 mM | 3.75 μM | 55 °C | <10 min | No | 3 | No | Yes | 91 |
| Human serum | (Urine and serum) | >7 days | (5 samples with flow measurement) | ||||||||||
| L-Tryptophan | Human serum | 2 | Colorimetric | 0.25–1.5 mM | 220.7 μM | N.A. | 15 min | (Yes) | 5 | Yes | Yes | 222 | |
| Human urine | |||||||||||||
| Albumin | Human serum | 2 | Coloured distance, angle | 1–25 g L−1 | 0.8 g L−1 | N.A. | 30 min | (Yes) | 2 | No | No | 223 | |
| ALP | 2 | 5–50 U L−1 and 50–200 U L−1 | 5 U L−1 | ||||||||||
| Serotonin | Buffer | N.A. | — | Echem (CA) | 1–20 mM | 0.15 mM | N.A. | 13 min | N.A. | 5 | Yes | Yes | 41 |
| cTnI | PBS | — | SERS | 0–0.5 ng mL−1 | 0.02 ng mL−1 | N.A. | <90 min | N.A. | 3 | No | Yes | 224 | |
Looking at Tables 1–3, most reports demonstrated real sample applicability at least at the level of the above-mentioned category (1), by spiking target analyte into artificial biofluids. Factors contributing to successful real sample matrix tests without dilution include multiple washing steps in immunoassays, off-device nucleic acid extraction and purification, and the application of an additional running buffer after the raw sample is applied. In this context, paper-based immunoassays requiring neither off-device sample pretreatment nor multiple operation steps are outstanding. Examples rely on automated sequential reagent addition methods,21,22 or on bioluminescence resonance energy transfer (BRET) signaling,23 details of which are introduced in following sections. In addition, paper-based NAT functioning without a need for off-device nucleic acid extraction are also noteworthy.24,25 However, there is still a room for improvement in terms of reagent handling and stability, long assay times, as well as a lack of real sample testing. When focusing on clinical sample matrices, blood serum is mostly used in immunoassays (Table 1), while blood serum, nasal swab samples and less frequently saliva are used in NAT (Table 2). On the other hand, whole blood samples are more commonly used in assays targeting routine health check parameters, such as glucose. This is partially attributed to biosafety and logistic challenges associated with obtaining and handling infectious disease samples, as well as the instability of whole blood samples, which makes it difficult to handle clinical samples outside of hospitals. When it comes to spiked samples, it is of importance to consider the practically relevant concentration range for real clinical samples (Tables S1–S3), a fact that is sometimes insufficiently addressed in publications.
Evaluation of sample-in–answer-out μPADs
Reagent handling and device operation involved in assays are important aspects to evaluate their user-friendliness. Ideally, operations needed for end-users should be limited to only sample application (more ideally, volume-independent) and signal readout without any external equipment. Tables 1–3 also summarize the number of device operation steps, the absence or presence of reagent handling requirements and external equipment, and the assay time. Additional details are provided in Tables S1–S3. Table 1 summarizing μPADs for diagnostic immunoassays reveals that they generally require more operation steps, such as washing and reagent addition, compared with other types of assays. Additionally, precise time control is often necessary during these multiple operation steps. On the other hand, as shown in Tables 2 and S2 covering NAT, it is often required to use external equipment, such as heaters or detection instruments to enhance assay sensitivity, in addition to relatively longer assay times. Compared with immunoassays and NAT, μPADs relying on simple reaction mechanisms as shown in Table 3 require fewer operation steps and shorter assay times, since they eliminate the necessity for off-device sample pretreatment and reagent handling, demonstrating significant progress toward sample-in–answer-out systems.When focusing on reagent handling across all types of assays, most μPADs store at least some of the required reagents on the device, while potentially unstable reagents such as biological reagents, hydrogen peroxide, and ferricyanide/ferrocyanide are often not stored. Even when such reagents are pre-deposited on a device, their long-term storage stability is typically not investigated. Thus, parallel evaluation of both reagent handling and long-term storage stability is of great importance. For hydrogen peroxide, methods to prolong its shelf-life when stored on μPADs using poly(vinyl alcohol) and titanium dioxide nanoparticles have recently been reported, enabling H2O2 storage for 30 days at 4 °C and at 25 °C, respectively.26,27 Similar basic evaluations to extend the storage stability of other specific reagents on μPADs are also needed, if they play a critical role in assay reactions.
Similarly, the number of operation steps should be considered together with the necessity of off-device sample pretreatment. Off-device sample pretreatment varies widely, including simple dilution, mixing with reagents, centrifugation, pre-incubation, or nucleic acid (NA) extraction requiring multiple steps (e.g., heating, washing, elution, etc.) (Tables S1–S3). Among the most common off-device pretreatments in clinical diagnostics, blood separation and NA extraction, numerous studies have reported their successful on-device integration. Readers are referred to related reviews for further details.10,18 Even if it is difficult to integrate sample pretreatment, selection of appropriate off-device pretreatment methods is at least important to enhance user-friendliness. For example, while conventional NA extraction by means of commercial kits usually takes around 20 min and requires centrifugation, Zhang et al. adopted a simple thermal lysis method instead, which completes within 5 min and eliminates the need for centrifugation.28 For these reasons and considering the ongoing efforts to on-device integration of sample pretreatment, we believe that fully integrated μPADs eliminating needs for off-device sample pretreatment deserve greater attention.
The necessity of precise time control is summarized in Tables S1–S3. For assays involving only a single reaction step, precise time control can be avoided by signal readout at saturation, or immediately after sample application, the latter being mostly limited to electrochemical measurements. To determine the feasibility of such strategies, the investigation of reaction kinetics would be of high importance. However, this is often not sufficiently addressed in the literature. On the other hand, assays that unavoidably need precise time control can be broadly classified into two categories: in the first one, the signal evolves over time following a single- or multi-step reaction, requiring signal readout at a fixed time after sample application. The second category includes assays involving multiple user interventions at defined time intervals. The former type has greater potential for adaptation to point-of-care testing (POCT), since automated signal acquisition systems can be implemented to eliminate the requirement for exact time control by the user.29 In contrast, for the latter category, apart from automated reagent or buffer addition described in the following sections, integration of a built-in timer into μPADs is the only practical way to enhance usability.30,31
The most commonly reported external equipment for use with μPADs is nowadays the smartphone, often combined with self-made controlled light boxes or 3D-printed attachments. Such setups are generally more user-friendly than scanners, RGB colour sensors, or benchtop fluorometers. Readers are referred to a recent review summarizing advances in smartphone integration with μPADs.32 On the other hand, some reports on colorimetric signal detection do not investigate the feasibility of naked-eye signal interpretation, although the shown figures indicate such possibility. To maximize the potential of instrument-free signal detection, the determination of visual detection limits (ideally evaluated by multiple observers) is of high importance.22,28,33 Apart from external equipment for signal detection, heaters or incubators are often involved, especially in NAT, as mentioned earlier (Table S2). Among these, Lee et al. reported an interesting approach using a power supply-free heating module based on the exothermic reaction of calcium oxide with water. That system achieves precise temperature control using a wax melting near the optimal temperature for isothermal amplification, thereby enhancing the usability in resource-limited areas.24
To summarize the general progress in μPAD development, assays targeting routine health checks or relying on simple reaction mechanisms have nearly reached true sample-in–answer-out functionality. In contrast, immunoassays and NAT, which require multiple operation steps or higher sensitivity, still demand further advancements before reaching that level of integration. The greatest remaining challenge lies in the necessity of off-device sample pretreatment. Most clinical validation testing has been conducted using serum or NA-extracted samples, rather than raw biological fluids such as whole blood. While such off-device sample pretreatment may be acceptable for μPAD applications in small clinics equipped with compact centrifuges or incubators, it remains impractical for bedside diagnostics, at-home testing, or use in resource-limited settings. Accordingly, the development of fully integrated sample-in–answer-out μPADs capable of integrating on-device sample pretreatment is expected to remain a key research trend. However, when assessing whether μPADs described by authors as “fully integrated devices” truly live up to that designation, it is important to simultaneously look at multiple parameters summarized in Tables S1–S3. For example, an excellent long-term storage stability might be reported for a μPAD, but a closer look reveals the omission of pre-deposition of sensitive reagents needed in the assay. In other cases, a significant reduction in μPAD operation steps might be reported but accompanied by extensive off-device sample pretreatment. Moreover, for cases reporting a comparable number of device operation steps, whether precise time control is required can greatly affect the overall complexity of the assay. In other words, we believe the parameters listed in Tables 1–3 and S1–S3 are useful to evaluate and effectively judge user-friendliness of μPADs.
Improvement in performance-oriented parameters
In the following sections, we describe fundamental improvements in performance-oriented parameters of general μPADs, including sensitivity, precision, rapidity, and long-term storage stability. Although the examples introduced below do not exclusively target diagnostic applications, the demonstrated proof-of-concept approaches are readily applicable to diagnostic applications, as they primarily focus on μPAD design itself rather than on specific reaction mechanisms or specific target analytes.Sensitivity and precision
The detection ranges achievable with various analytical methods for μPADs, including colorimetric, fluorescence, electrochemical, and chemiluminescent approaches, were discussed in detail in our 2017 review.10 In general, improving sensitivity can be approached through multiple assay design strategies. For immunoassay formats, for example, enhancing washing efficiency to reduce background noise and applying enzyme-mediated signal amplification are both effective means of increasing sensitivity.34 However, these improvements typically require additional operational steps or sequential reagent addition, which in turn compromise user-friendliness and can limit applicability in resource-limited or self-testing environments. In this context, considerable effort has been devoted to incorporating automated fluid control systems into μPADs, enabling sequential delivery of reagents or buffers with minimal user intervention. This technology is expected to facilitate the translation of highly sensitive laboratory-based clinical assays that require multiple operational steps into bedside or at-home point-of-care diagnostic applications. Even in 2017, a handful of approaches aimed at simplifying these operations have already been reported,35–37 but since then, a broader variety of novel strategies has emerged, as introduced in the following subsections.First, it is worth noting that the early concepts already discussed in our previous review, such as maze-like channel layouts,38 sucrose-based dissolvable barriers,39 and origami-type folding valves,40 have continued to be used in later years, either with further optimization or by introducing alternative or mixed sugars to refine liquid flow delay profiles. Approaches requiring user intervention and/or external equipment have also appeared in recent years, including sliding or movable reagent pads,41,42 manually rotatable platforms,31 centrifugal paper-fluidic devices,43 thermally actuated shape-memory-polymer valves,21 and manually controlled hydrophilic bridges.44 While these methods enable precise timing control, they still depend on user intervention or additional hardware, thereby limiting their applicability in fully autonomous point-of-care devices.
A growing number of passive strategies not requiring user intervention has been developed, enabling single-step assays with self-sequenced liquid delivery. These include mechanisms based on aqueous two-phase systems,45 hydrogel- or swelling polymer-based valves,46,47 and mechanical delay elements such as accordion-folded paper.48 However, these approaches can suffer from certain limitations: their applicability may be restricted to specific assay types, the reproducibility of delay times is sometimes relatively low, and the increased fluid viscosity resulting from dissolved valving materials may affect downstream fluidic transport and enzymatic reactions. On the other hand, Park et al. reported a mechanical compression-based delay method in 2017, where deforming part of a paper channel by applying pressure slowed the fluid flow, with delays tuneable up to approximately 3.5 minutes depending on compression strength (Fig. 1a).49 This approach was then integrated in a sequential ELISA (enzyme-linked immunosorbent assay) workflow, illustrating its utility for immunoassays without requiring additional chemical barriers. Subsequently, Strong et al. developed a wax-printed time-delay channel method by printing wax from both sides of the paper substrate (Fig. 1b).50 This design partially sealed the channel and increased capillary flow resistance, achieving precise liquid flow delays of up to around 800 seconds by changing the coverage percentage of wax deposited on the channel (shown as yellow areas in Fig. 1b, top). The technique was applied to multi-step biochemical assays, demonstrating that wax patterning can serve as a low-cost and reproducible timing mechanism in fully integrated devices. More recently, Lee et al. introduced a delamination timer concept by printing a hydrophobic ink with lower water resistance than that used to fabricate the main channels.51 Upon wetting, this interface gradually separated from an adhesive film, thereby opening the flow path (Fig. 1c, top). By adjusting the printed barrier width and the number of barriers, delays ranging from 30 seconds to over 18 hours were achieved. This timing mechanism was successfully integrated into automated ELISA and DNA purification workflows, showcasing its versatility for both protein- and nucleic acid-based assays. In a very recent application of this system by Dai et al., adjustments of the delamination ink deposition widths (Fig. 1c, bottom left) enabled control of flow delays between 150 s and 900 s (Fig. 1c, bottom right).52 In addition to these systems, several flow-control strategies have been reported by accelerating flow speed through gap-controlled dual-layer paper designs53 or hollow-channel structures embedded within paper.54 These concepts will be described in greater detail in the following section. Along with it, controlling the microfluidic channel length provides another means to realize sequential reagent delivery. Chien et al. reported a one-step vertical flow paper-based immunoassay with on-device signal amplification, in which pre-deposition of reagents combined with control of channel lengths enabled a fully automated reaction without any user intervention.22 Specifically, they achieved around 5 min delay of amplification reagent delivery (blue arrows in Fig. 1d) after initial sample flow (red arrows) through design of the total channel length in 3D-μPADs.
 | ||
| Fig. 1 Examples of passive strategies enabling automated sequential reagent delivery on μPADs by creation of liquid flow delays: a) top left: schematic of a 3D paper-based microfluidic device for measuring liquid flow delay time induced by compression of paper; top right: relationship between applied pressure and time at which blue color appeared in the outlet; bottom: operation principle for the measurement of flow delayed. Adapted from Park et al.49 Copyright 2017, with permission from Elsevier. b) Top: Schematic representation of wax deposition on μPADs; black wax was used to define the border of the hydrophilic flow channel, while yellow wax was applied for fluid time delay creation (indicated as % coverage of the paper channel); bottom: wicking distance over time depending on coverage percentage of yellow wax on channel. Adapted from Strong et al.;50 published by MDPI. c) Schematic illustration of the timer mechanism and fluid flow process in channels with timers opening sequentially from left to right, and the relationship between timer widths and corresponding delays. Adapted from Dai et al.52 Copyright 2026, with permission from Elsevier. d) Folding diagram of μPAD and flow process between 0 to 10 min displays the different assay stages from sample injection, detection, and signal amplification, with the reagent flow at various time points indicated; red arrows: sample detection flow; blue arrows: signal enhancement reagent flow. Adapted from Chien et al.22 Copyright 2025, with permission from Elsevier. | ||
Overall, it should be noted that automatic time-delay and sequential reagent delivery strategies that do not rely on dissolvable materials tend to offer higher reproducibility compared with sugar- or polymer-based approaches. Improvements in reproducibility for assays requiring sequential reagent delivery are important to enhance the potential of μPADs to bridge the gap between highly sensitive, multistep lab-based clinical assays and real-world point-of-care diagnostic applications. However, to date, most such developments have been limited to lateral flow assay formats, whereas in vertical flow assays, delay control is still typically achieved using sugars or polymers,25,39 or by employing origami-type devices30,55–60 that require user intervention. Therefore, this area would profit from further development to extend the applicability of highly reproducible delay mechanisms free of any valving materials to vertical flow assay platforms.
Rapidity
In paper-based microfluidics, the speed of fluid transport driven by capillary action is strongly influenced by the pore size of the paper substrate, as described by the classical Washburn equation.61 While slower transport rates can be advantageous in certain contexts, for example enabling devices with distance-based readout that rely on time-dependent flow62 or facilitating antigen–antibody binding requiring a certain interaction time,63 they have generally been regarded as a limiting factor for achieving rapid diagnostics. Accelerating liquid flow not only shortens assay turnaround time but can also enhance analytical performance by reducing diffusion-driven dispersion of analytes during transit. One of the most prominent approaches involves introducing a gap between two layers, at least one of them being paper. This concept, originally introduced in 2012 by Jahanshahi-Anbuhi et al.,64 has been further investigated by other research groups even prior to 2017.65,66 In 2018, Channon et al. demonstrated that introducing an optimized height of interlayer gap between two paper layers in taped devices increased the flow rate to 1.56 cm s−1, which is higher by around 145-fold in comparison to the classical liquid wicking within a single paper layer following the Washburn behaviour (Fig. 2a).53 This strategy was subsequently applied to enhanced electrochemical flow injection analysis of cadmium ions and virus particles.53,67 This method has since been used in various μPAD applications, continuing to be reported in studies up to the present.68–74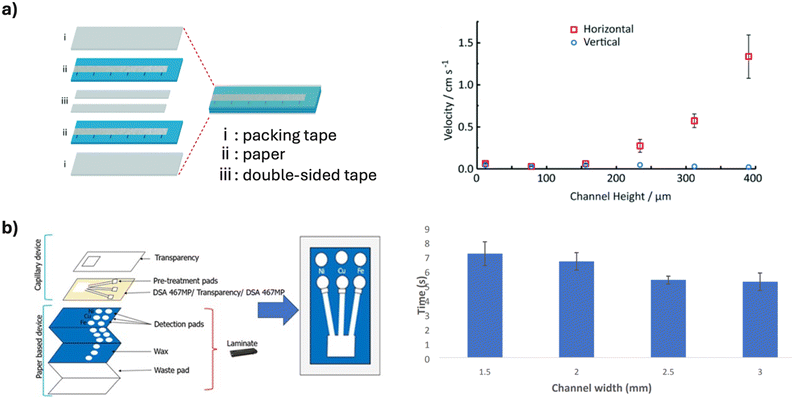 | ||
| Fig. 2 Selected approaches towards μPADs with increased sample liquid flow rates: a) illustration of straight channel multilayer μPAD with a gap defined by the thickness of double-sided tape (iii) in between two hydrophobic wax-patterned paper layers, and effect of channel height on flow rate for horizontal and vertical 2-layer μPAD. Adapted from Channon et al.53 b) Illustration of assembly of device made from paper and transparency film, and its operation, and effect of channel width on time for sample to reach detection zone. Adapted with permission from Aryal et al.75 Copyright 2023 American Chemical Society. | ||
More recently, there has been a noticeable shift toward hybrid device architectures that do not rely solely on paper for fluid transport. Instead, paper is strategically positioned as a reagent chamber or detection zone, while fluid transport is handled by alternative materials with inherently higher wicking or flow rates. Examples include polymer films,75 polydimethylsiloxane (PDMS) microchannels,76 cellophane,77 and 3D-printed substrates,78 each integrated to form continuous flow pathways that bypass the inherent speed limitations of cellulose-based materials. In the following, some examples not necessarily related to diagnostic applications are discussed, since they represent approaches that are potentially applicable to that field. Arun et al. combined PDMS and paper channels, achieving a significant enhancement of fluid transport rate and mixing performance.76 Aryal et al. reported a capillary-driven PAD combined with transparency film for heavy metal detection in 2023 (Fig. 2b).75 While conventional μPADs require minutes to deliver samples to the detection zone, the developed device took only 8 s until colour signals appeared, pointing to a flow rate of roughly 0.3 cm s−1. This velocity is about 30-fold higher than the classical liquid wicking process within a single paper layer. Although designed for environmental monitoring, the mechanism itself could readily be adapted for diagnostic purposes.79–81 Later, Macleod Briongos et al. reported a 59–74% improvement in liquid flow rate by the fabrication of laser-cut grooves on paper.82 Lastly, Park et al. fabricated flow channels on a paper substrate using a digital light processing (DLP) 3D printer.78 While quantitative flow speed data was not provided, the assay time of the device with 3D printed channel was shorter than that of typical 2D μPADs, which was attributed to an increase of fluid flow speed in the small channels embedded within the paper. Through such approaches combining paper with non-paper materials into single devices, researchers have demonstrated the feasibility of significantly reduced assay times, while retaining key advantages of paper, such as the low cost and biochemical compatibility. The examples highlighted in Fig. 2 demonstrate that designs to accelerate the liquid flow speed, including interlayer gaps and hybrid fluidic pathways, can simultaneously shorten assay time and improve analytical performance by limiting diffusion-driven analyte dispersion. These combined effects enhance the practicality of μPADs for rapid and reliable diagnostic applications, particularly in point-of-care settings where both speed and accuracy are critical.
Long-term storage stability
Over the last few years, the storage stability of μPADs has received greater attention than in the past, with many recent studies assessing device performance over storage periods of mostly one month (Tables 1–3). Notably, some reports that extended storage tests beyond one month observed significant declines in stability,26,38,83,84 highlighting the importance of truly long-term investigations. The longest storage durations among the analysed publications were 10 weeks, 12 weeks, and 3 months, which still fall short of commercial requirements.44,85,86 However, such long-term studies present a fundamental challenge in academic settings, where publication timelines and the pressure for novelty often discourage a year-long or multi-year data collection. One potential solution is accelerated stability testing based on the Arrhenius equation.87 By exposing the devices to a harsher than usually encountered environment (typically at elevated temperature), the equivalent of long-term storage can be simulated within a much shorter period. While this method is often used in the biosensor field,88,89 there are few reports adopting it to μPADs. Yakoh's group recently introduced accelerated shelf-life testing for μPADs by storing devices at 55 °C, thereby reducing the required testing period to approximately one-eighth of the normal duration. This approach provides a practical way for more efficient evaluation of long-term storage stability.90,91Looking at more details of the storage stability studies presented in Tables S1–S3, it is revealed that the thresholds used to define acceptable signal changes vary widely, typically ranging from 5% to 20%. Some reports do not even mention quantitative results, which makes it difficult to compare across different studies. To address this issue, we believe that as long as commonly accepted standards are not available, setting multiple threshold levels of acceptable signal change and indicating the corresponding storage stability for each of those would be the most practical approach. Furthermore, while most studies monitor the signal stability in the absence and presence of target analyte, some investigations use calibration plots covering a wide target analyte concentration range.57,92 The latter approach is more effective for assessing device behaviour over storage time, especially in the lower analyte concentration range, where performance degradation may have a stronger negative impact.
Regarding strategies to extend μPAD storage stability, most remain consistent with those described in our 2017 review. Namely, incorporating stabilizing agents such as small molecules,93 proteins,94,95 polymers,26,96 sugars,97 or micro/nanoparticles27 into the device matrix. Particular attention continues to be paid to the stability of enzymes, which are often the most degradation-prone components in μPAD assays.98 In this context, the use of nanozymes as enzyme mimics has emerged as a noteworthy trend.99,100 These nanomaterial-based catalysts generally exhibit superior thermal stability and prolonged activity, potentially alleviating the shelf-life bottleneck. While some studies confirmed the long-term storage stability of nanozyme-assisted PADs for more than two months,101,102 others reported stability for about 1 month,103,104 with certain nanozymes retaining activity for as little as 2–3 weeks.105–107 Since there has been no report directly comparing long-term storage stability of natural enzymes and nanozymes on paper substrates, more comprehensive research in this direction is needed in the future. It should also be noted that the use of nanozymes involves a trade-off in target specificity, which could compromise analytical accuracy unless compensated by further innovations in nanozyme design. Thus, nanozyme-assisted PADs are sometimes combined with natural enzymes such as glucose oxidase.108 However, by combining nanozymes and natural enzymes in a single PAD, the advantages of the former in terms of device storage stability are possibly sacrificed. Meaningful progress in this field will depend not only on assay engineering but also on advancements in the broader field of nanozyme chemistry.
As summarized earlier, long-term storage stability testing is still often overlooked even in simple assays and is further complicated by inconsistencies in the evaluation criteria. Hence, it is important to remain critical of these aspects when considering the real-world applicability of μPADs.
Simplified signal interpretation beyond purely qualitative readouts
As already mentioned in the introduction, early μPADs primarily relied on benchtop sized equipment and computer software for quantitative data interpretation, or on qualitative binary visual indications, such as the simple presence or absence of a coloured line. The former offered limited accessibility for POCT, while the latter were vulnerable to user subjectivity, particularly near clinically relevant threshold concentration values. In our previous review, we already introduced several concepts for semi-quantitative assays, which do not sacrifice user-friendliness, including counting-based,109 timing-based,110 distance-based,62 and text-based signal readout formats.111 Since 2017, however, a range of new approaches has been reported, further advancing this field.One of the most widely explored concepts has been distance-based formats, in which the migration length of a coloured front in a paperfluidic channel correlates with analyte concentration. Lee et al. converted the originally straight channel design to angular readouts,112 thereby enabling more compact device layouts while retaining the concentration resolution. Subsequently, Teprek et al. and Khachornsakkul et al. introduced counting-based schemes in which discrete reaction spots along the channel sequentially change colour as the sample liquid progresses in vertical and lateral flow configurations, respectively (Fig. 3a and b).113,114 Multiple paper layers in an origami-type device have been modified with reagents, for example for the titrimetric determination of Ca2+ levels. Sample liquid passing vertically through the device, thereby reacted with the pre-deposited reagents on each circular reaction zone, resulting in depletion of the target analyte and hence, changes in the number of spots showing a colorimetric reaction (Fig. 3a). In an alternative design based on a similar reaction principle, results are interpreted by counting colored zones in a circular lateral flow arrangement. Also building upon the distance-based readout concept, others have incorporated machine-readable patterns such as for example QR codes,115 enabling smartphones to decode the reaction pattern and translate it into concentration data (Fig. 3c). Multiple QR codes composed of colorimetric indicators were placed in series into a paperfluidic channel. In absence of analyte, all QR codes are hidden, but an increasing number becomes readable with increasing analyte concentrations. Although the latter study targeted heavy metal ions for proof-of-concept, the same approach is also applicable to diagnostic assays. More recently, Manmana et al. introduced a system converting a distance-based concept into a 1D barcode-based signal readout116 for a semi-quantitative glucose assay in artificial urine matrix (Fig. 3d).117 The barcode-based readout offers the advantage of single-step result interpretation from a single code, compared to the QR code-based system requiring the scanning of multiple QR codes to obtain semi-quantitative results. Only specifically designed QR code patterns offer the possibility for a semi-quantitative single QR code assay.118 Both QR- and 1D barcode-based signalling strategies offer clear advantages in terms of simplifying data digitization and storage, features not that readily achieved with more conventional readout methods.
 | ||
| Fig. 3 Examples of μPADs enabling naked eye (semi-)quantitative signal readout: a) illustration of a folded origami-type μPAD with sample application, and standard colour scale used for interpretation of experimental results on μPAD for end point detection in titrimetric analysis; increasing number of purple spots indicates higher target analyte levels (e.g. Ca2+). Adapted from Teprek et al.113 Copyright 2020, with permission from Elsevier. b) Illustration of a device and method to interpret results by counting colored zones in a circular lateral flow arrangement. Adapted with permission from Khachornsakkul et al.114 Copyright 2022 American Chemical Society. c) Photograph of QR code array PADs relying on the distance-based signalling mechanism after application of various Cu2+ concentrations. Reproduced from Katoh et al.115 d) Schematic illustration explaining the principle of semi-quantitative 1D barcode readout PADs; red arrows in the right panel indicate bar positions that change from normal to wide state with increasing analyte concentration, thereby converting the distance-based concept into a changing barcode signals. e) Schematic showing the working principle of a text-displaying device; overlaying a transparent film with screening colour on a paper layer with inkjet-deposited reagents for colorimetric protein detection (human serum albumin) results in the readability of text depending on the urinary albumin level. Adapted from Yamada et al.33 Copyright 2017 American Chemical Society. f) Schematic showing the working principle of a “dip-and-read” distance-based detection method; overlaying a transparent film with colour gradient on a paper layer with a colorimetric reagent deposited in the form of a line results in a target analyte concentration-dependent change in the visible line length. Adapted from Yamada et al.119 (g) Photographs of origami-μPAD, schematic cross-sectional view of multilayer origami-type traffic light μPAD and illustration of varying liquid flow paths depending on a hydrogen peroxide-responsive hydrophobic to hydrophilic phase-switching reagent (hydrophobic wall) resulting in H2O2 concentration-dependent traffic light colour display. Adapted from Ohta et al.120 | ||
In parallel, transparent film-supported strategies have emerged as an alternative means of interpreting results of colorimetric assays. In 2017, Yamada et al. reported a text-displaying colorimetric PAD, in which chemical reactions trigger colour development of indicators inkjet-deposited onto paper in the form of text. When overlaid with a transparent film featuring coloured zones of varying colour shades and intensities, the underlaying text symbols (e.g., “trace”, “1+”, “2+”) become readable only when their colour intensity surpasses a threshold corresponding to a specific urinary albumin concentration (Fig. 3e).33 Later on, in 2018, a “dip-and-read” distance-based detection method was developed by modifying the transparent film overlay concept. Using a film with a colour gradient, a line of colorimetric indicator printed on a paper substrate becomes visible to the naked eye only in regions where its colour exceeds that of the overlaid film (Fig. 3f).119 Although so far only demonstrated for Ni quantification, this method is expected to be equally applicable to diagnostic assays. This “quasi distance-based” readout approach offers advantages over most conventional distance-based assays in terms of speed and independence from the applied sample volume. Because the entire PAD is dipped into the sample liquid, rather than relying on the application of a fixed sample volume to an inlet area, it is not necessary to wait for the completion of fluid wicking through the device. However, the method is more likely to be affected by ambient light conditions.
Finally, several additional innovations exploit microchannel engineering to encode concentration information. Ohta et al. reported a multi-layer origami traffic light μPAD for semi-quantitative glucose detection in artificial urine, achieved by varying liquid flow paths, channel geometry and width, and the concentration of a hydrogen peroxide-responsive hydrophobic to hydrophilic phase-switching material (Fig. 3g).120 While earlier traffic light type μPADs relied on smartphones for augmented reality-based result display121 or fluorescence-based signal output,122 the origami device enabled naked-eye readout via coloured spots composed of food dyes that maintain a stable time-independent colour appearance resistant to fading, in contrast to many chromogenic indicators. The approaches shown in Fig. 3 demonstrate how advances in semi-quantitative visual and encoded signal outputs can transform μPAD readouts beyond purely qualitative analysis. Such improvements in interpretability and consistency are crucial for diagnostic applications in resource-limited settings, thereby strengthening the applicability of μPADs for real-world point-of-care testing.
On the other hand, smartphone-based signal detection approaches represent one of the most important strategies for enabling target quantification. They have already been explored prior to 2017, as discussed in our previous review. However, many early smartphone readout methods suffered from the influence of external light variability, requiring dedicated light sources or enclosed imaging boxes to ensure consistent measurements.123,124 Chen et al. addressed this limitation by scattering the smartphone's built-in flash to achieve uniform illumination (Fig. 4a).125 Fan et al. mitigated the issue by adapting image processing algorithms to correct for ambient light fluctuations (Fig. 4b).92 Alongside these hardware- and software-based improvements, there has been a rise in studies explicitly validating smartphone compatibility as part of μPAD design, positioning mobile phone-based detection as a key enabler for achieving true POCT functionality.13,32
 | ||
| Fig. 4 Examples of approaches mitigating the influence of external light variability in smartphone-assisted μPAD signal readout: a) photographs of 3D printed assembled imaging box with smartphone and illustration of light diffusion upon scattering of the smartphone's built-in flash to achieve uniform illumination for readout of μPADs. Adapted from Chen et al.125 Copyright 2019, with permission from Elsevier. b) Principle of colour correction using a standard colour bar: automatic recognition of colour bar region, measurement of RGB colour intensities, and reconstruction of image. Adapted with permission from Fan et al.92 Copyright 2022 American Chemical Society. | ||
Signal detection methods and analytical techniques recently integrated into μPADs
While much of the development in μPADs since 2017 has focused on refining existing methodologies, the broader field of analytical chemistry has also witnessed the emergence of entirely new technologies with transformative potential. Several of these innovations originally developed without μPADs in mind have begun to find their way into paper-based platforms, enabling capabilities that were previously unattainable in such simple formats. Integration efforts remain relatively limited in number, but their trajectory suggests a growing interest in leveraging these advances for portable, low-cost diagnostics. In this section, we highlight three representative areas where analytical technologies have newly intersected with μPAD research: 1. detection techniques involving surface-enhanced Raman scattering (SERS)126,127 or bioluminescence128 that push detection sensitivity towards low analyte amounts, 2. techniques represented by CRISPR-based assays,129,130 and 3. machine learning (ML)-assisted image analysis and data interpretation, that enhance accuracy and selectivity.131,132 These examples illustrate how the convergence of emerging analytical tools with μPADs could redefine the boundaries of point-of-care testing in the coming years.Surface-enhanced Raman scattering (SERS)
SERS is a highly sensitive detection technique that enhances the weak Raman scattering of molecules adsorbed onto corrugated metallic surfaces through localized surface plasmon resonance.126,127 This technique has enabled ultrasensitive, label-free molecular detection through distinct peaks in Raman spectra representing “fingerprints” of target analytes. Raman signals are enhanced by factors up to 108 or even larger,133 when target molecules are located within hot spots, formed through metallic nanostructures on various solid substrates. Owing to this feature, SERS has gained considerable attraction in the analytical chemistry field in the 2000s, particularly for applications that demand high sensitivity, such as in biomedicine, environmental monitoring, and chemical reaction monitoring.134 The combination of SERS with paper substrates has attracted interest since the 2010s,135,136 largely due to the inherent advantages of paper materials, including biocompatibility, physical flexibility, passive fluid transport through capillary action, low cost, and portability as summarized in several review articles devoted to this specific topic.137,138However, most reports adopting paper as a SERS substrate are not directly related to diagnostics and just marginally mention such application potential as a future perspective, with the exception of some LFIAs. A key bottleneck lies in the detection system itself. For example, Lim et al.139 and Mabbott et al.140 reported paper-based SERS for the detection of acute myocardial infarction biomarkers and a microRNA associated with myocardial infarction. The former one achieved multi-target detection through a multi-channel design fabricated on a single paper (Fig. 5a), and the latter one successfully simplified the assay procedure by integrating a multi-step reaction assay into a multi-layered vertical μPAD (Fig. 5b). Both approaches use gold nanoparticles modified with Raman reporter molecules and either capture antibodies or DNA oligomers complementary to the target RNA. On the other hand, their work still relied on laboratory-scale SERS detecting modules. Therefore, paper-based SERS approaches must be developed along with the miniaturization of Raman spectrophotometers including laser source, detectors, and optical components, before achieving true POCT capabilty.141 While there are many reports devoted to the development of portable Raman spectrophotometers, including those compatible with smartphones, the trade-off between instrument size and analytical sensitivity remains a significant challenge.141 Moreover, publications describing the successful integration of paper-based SERS and portable spectrophotometers, as for example reported by Zeng and co-workers, are currently mostly limited to typical model targets including rhodamine 6G, malachite green and crystal violet with their distinctive C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N spectral features (Fig. 5c).142,143 Considering this fact, as well as the still relatively high costs of portable Raman spectrometers, more work is required looking at both aspects of potential practical application to meaningful analytical targets and portable and low-cost, but sensitive Raman signal detecting instruments.
N spectral features (Fig. 5c).142,143 Considering this fact, as well as the still relatively high costs of portable Raman spectrometers, more work is required looking at both aspects of potential practical application to meaningful analytical targets and portable and low-cost, but sensitive Raman signal detecting instruments.
 | ||
| Fig. 5 Examples of μPAD signal readout approaches relying on surface-enhanced Raman scattering (SERS): a) schematic illustration of multiplexed μPAD with signal acquisition by SERS for detection of acute myocardial infarction biomarkers. Adapted from Lim et al.139 Copyright 2020, with permission from Elsevier. b) Illustration of detection method for microRNA miR-29a in a 3D μPAD and representative images of sample readout using both colorimetric and SERS methods for samples containing miR-29a (+ve) and samples without miR-29a (−ve). Adapted from Mabbott et al.140 c) Schematic illustration of sample application on paper substrate and detection using a miniaturised smartphone-based Raman analyser. Adapted with permission from Zeng et al.142 Copyright 2019 American Chemical Society. | ||
Bioluminescence
Bioluminescence (BL) is a subcategory of chemiluminescence, with light emission produced by an oxidation reaction involving an enzyme (luciferase) and its substrate (luciferin) in living organisms. Compared to other optical detection techniques applied to μPADs relying on colorimetric or fluorescence signals, chemiluminescence and BL have the advantage of achieving superior signal-to-noise ratios due to the nearly zero background signal. At the same time, they eliminate interference caused by environmental light conditions as in case of colorimetric approaches, or the necessity of external light sources for fluorescence-based detection. BL is particularly attractive due to the enzyme reaction resulting in generally higher quantum yields compared with chemiluminescent systems, allowing ultrasensitive target detection down to attomole levels.128While BL-based analytical assays have been reported for decades in laboratory settings, their application to paper-based analytical devices (PADs) started more recently in the late 2010s. One of the earliest reports appeared in 2014 by the Johnsson group, where a BL-based reaction was performed on a filter paper substrate with a 96-wellplate-like design.144 Their system relies on ratiometric BL sensor molecules referred to as luciferase-based indicators of drugs (LUCIDs), enabling the detection of several types of drugs by monitoring changes in the emission colour caused by a bioluminescence resonance energy transfer (BRET) process. However, this approach was still remote from a POCT applicable system, since all reagents were applied during the assay, with the paper platform solely acting as a replacement for plastic microwell plates for the purpose of reducing sample and reagent volumes. Several years later, the same group demonstrated a related BRET-based assay with sensing proteins lyophilized on paper substrates,145 a first step towards reducing the burden of reagent handling by the assay user. Subsequently, Tenda et al. reported fully-integrated BRET-based μPADs using a different type of BL molecular sensor named LUMABS (luminescent antibody sensor) (Fig. 6a).23 By pre-deposition of both LUMABS and its substrate furimazine on spatially separated paper layers, simultaneous detection of antibodies against HIV, hemagglutinin, and dengue virus was achieved by the application of a single drop of whole blood and subsequent acquisition of a photograph in the dark, without any reagent handling. However, when compared to other paper-based immunoassays, the achieved limit of detection in the nanomolar order is relatively high (Table 1). A major advantage of all BRET-based approaches is that BL signals are recorded as changes in emission colour, rather than the highly time-dependent BL emission intensity. The latter necessitates stricter control over signal readout timing.
 | ||
| Fig. 6 Paper- and polymer-based POCT devices with bioluminescence signal readout: a) schematic of the luminescent antibody sensor (LUMABS) working principle, its integration into a multi-layer 3D-μPAD, and its use for simultaneous detection of three different antibodies. Reproduced from Tenda et al.23 b) Comparison of chemical structures of native luciferin coelenterazine with an imidazopyrazinone derivative (Hulumino1) used as human serum albumin (HSA)-responsive luciferin, and measurement procedure for serum HSA with Hulumino1 pre-deposited on polyethylene strip in a dry film state. Adapted from Kurita et al.153 Copyright 2025, with permission from Elsevier. | ||
Since then, a growing number of BL-based μPADs has been reported. However, the greatest challenge in applying BL to μPADs lies in the limited stability of bioluminescent substrates under ambient and even refrigerated conditions.95,146 As a consequence, these substrates are often not pre-deposited on the μPAD but added along with or after sample application, limiting user-friendliness.86,147–149 In some cases, long-term storage stability has not been adequately investigated.150 BL bacteria-based assays are superior in terms of stability, but their assay mechanism might limit analyte selectivity, since it commonly relies on the inhibition of bacterial enzymatic reactions by the target molecules.151,152 While not using paper as the assay platform, Kurita et al. reported a BL substrate named Hulumino1, a coelenterazine analogue (Fig. 6b, top).153 They targeted human serum albumin in serum samples, which behaves as a luciferase catalysing the oxidation of Hulumino1 pre-deposited on a polyethylene strip in a dry film state, producing a corresponding BL signal (Fig. 6b, bottom). While BL intensity rapidly decreased to 68% when devices were left at room temperature overnight after fabrication, approximately 50% of the initial luminescence intensity was still retained after one month stored at room temperature due to the premature substrate oxidation being limited to the surface of the dry film. But given that for many reported BL-based assay devices even storage under vacuum condition could not achieve long-term stability under ambient conditions,154 chemical encapsulation or molecular structure modification of BL substrates are expected to offer more promising solutions.155 However, it should be noted that extension of storage stability often comes at the expense of assay sensitivity, since increased resistance to air oxidation potentially reduces the reactivity of the BL substrates. Overall, continued efforts in this direction are required to lead to fully integrated μPADs with high long-term storage stability under ambient conditions.
CRISPR-based assay techniques
The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and CRISPR-associated (Cas) protein systems initially attracted attention in the field of gene editing,156 with their applications in analytical chemistry emerging in the late 2010s. CRISPR/Cas systems are classified into several subtypes according to their nucleic acid targeting properties, among which Cas9, Cas12, and Cas13 are most widely used.157,158 Common to all systems, a ribonucleoprotein (RNP) complex consisting of a Cas protein and a programmable guide RNA referred to as CRISPR RNA (crRNA), recognizes a specific nucleic acid sequence, after which the RNP complex becomes activated. Owing to their simple design, high specificity capable of distinguishing single-nucleotide mismatches, and the ability of signal amplification through trans-cleavage turn-over reactions (limited to Cas12 and Cas13), CRISPR-based assay techniques have been widely investigated up to now.In the classical Cas9 system, the RNP complex targets double-stranded DNA (dsDNA) and cleaves it through the so-called cis-cleavage reaction. In 2016, Pardee et al. reported a Nucleic Acid Sequence-Based Amplification (NASBA)-CRISPR cleavage assay named NASBACC for Zika virus detection. In this system, isothermal nucleic acid amplification following reverse transcription of isolated Zika RNA, combined with the sequence-specific nuclease activity of CRISPR/Cas9, enabled discrimination of viral strains with single-base resolution.159 They further demonstrated the feasibility of freeze-drying NASBACC reagents on circular-shaped paper discs, suggesting the applicability of this approach on paper-based assay platforms. On the other hand, the Cas13 system targets RNA, where the RNP performs cis-cleavage of the target RNA, followed by collateral indiscriminate cleavage of surrounding RNA molecules, a process referred to as trans-cleavage reaction. The first Cas13-assisted assay was reported by Gootenberg et al. in 2017, termed Specific High-Sensitivity Enzymatic Reporter UnLOCKing (SHERLOCK).129 By using a quenched fluorescent RNA reporter, the trans-cleavage activity of Cas13 triggered by RNA that was amplified through reverse-transcribed recombinase polymerase amplification (RPA) followed by T7 transcription was transduced into a fluorescence signal. This enabled the successful detection of specific strains of Zika and dengue virus, discrimination of pathogenic bacteria, as well as genotyping of human DNA, and identification of mutations in cell-free tumour DNA. Importantly, they also demonstrated the feasibility of freeze-drying SHERLOCK reagents on paper discs. Similarly, in the Cas12 system, the RNP complex targets and cleaves dsDNA, followed by collateral trans-cleavage of surrounding single-stranded DNA (ssDNA). In 2018, Chen et al. reported a Cas12a-based assay combined with RPA for the detection of human papillomavirus in patient samples.130 Their assay method became known as DNA endonuclease-targeted CRISPR trans reporter (DETECTR). The detection mechanism closely resembles that of SHERLOCK, except that a quenched fluorescent ssDNA reporter is used instead of an RNA reporter. Subsequently, both SHERLOCK and DETECTR were integrated with paper-based lateral flow assays in 2018 and 2020, respectively.160–162 Based on this history of CRISPR-based diagnostics, further integration with paper platforms is expected to offer strong potential for POCT applications.
It is therefore no surprise that such integration has increasingly progressed. Since comprehensive reviews on this topic have been published recently,163,164 we here focus on highlighting the crucial and critical aspects. While many μPAD implementations currently employ CRISPR primarily at the signal detection stage,165–170 there is a growing trend toward fully integrated devices in which sample preparation, amplification, and CRISPR-mediated detection occur within a single paper-based platform, as described in an earlier section. A CRISPR-based μPAD going beyond simple paper discs was first reported by English and Soenksen et al. in 2019 for the detection of Ebola ssRNA (Fig. 7a).171 The assay mechanism relies on viscosity changes in an ssDNA-crosslinked hydrogel in response to Cas12a-mediated trans-cleavage. The assay was carried out by adding a pre-incubated Cas12a RNP mixture together with RT-RPA-amplified nucleic acid and the DNA hydrogel on different paper layers. The resulting signal is obtained from dye flow in the bottom paper channel. The first fully integrated CRISPR-based μPAD was reported by Nguyen and Soenksen et al. in 2021 (Fig. 7b).25 Integration of sample collection, lysis, isothermal nucleic acid amplification, a SHERLOCK reaction zone, and a lateral flow strip minimized user-intervention, limited to just pushing an activation button to initiate fluid flow and subsequently reading the result by the naked eye. Furthermore, CRISPR-based diagnostics are not limited to nucleic acid detection, but also applicable to small molecules or metal ion targets by use of aptamers or antibodies through ELISA-like formats. Suzuki et al. reported a CRISPR-assisted origami paper-based ELISA for the detection of hepatitis B virus surface antigen in whole blood in 2025 (Fig. 7c).57 The Cas12a system was used for signal amplification, in combination with DNA network-labelled antibodies initiating trans-cleavage of an ssDNA fluorescent reporter, resulting in higher sensitivity than conventional HRP-labelled ELISA. Moreover, by integrating a blood separation membrane and all necessary reagents onto paper layers, required end-user operation was reduced to paper flap folding and adding buffer solutions.
 | ||
| Fig. 7 Examples of μPADs relying on CRISPR-based assay techniques: a) schematic of the stackable μPAD design for operation with probe ssDNA-crosslinked CRISPR-responsive hydrogels for electrical readout, and SEM images of paperfluidic channels in the absence (top) and presence (bottom) of target dsDNA. From English et al.171 Adapted with permission from AAAS. b) Schematic of the sensor components of a face-mask-integrated SARS-CoV-2 wearable diagnostic μPAD and key steps of the freeze-dried reactions, each separated by a poly(vinyl alcohol) (PVA) time delay. Nguyen et al.25 Springer Nature, 2021, adapted with permission from SNCSC. c) Design of a paper-based origami immunoassay device with pre-deposited reagents on multiple pads, and assay procedure. Adapted with permission from Suzuki et al.57 Copyright 2025 American Chemical Society. | ||
However, aside from the publications mentioned above, to the best of our knowledge, there have been no further reports of fully integrated CRISPR/Cas-related μPADs in the diagnostic field. This gap can be attributed to several factors: the use of paper limited to a substrate material for signal detection (as seen in SERS-based approaches),172,173 the requirement for off-device sample pretreatment59 or off-device nucleic acid amplification,39,169,174 the need for user-handled CRISPR-related reagents,175,176 or the absence of validation with biofluid sample matrices.94,177 Since these techniques are ongoingly developed, truly “fully-integrated μPADs” making use of CRISPR/Cas systems remain highly anticipated.
Artificial intelligence integration
The integration of artificial intelligence (AI) technologies such as machine learning (ML), neural networks (NN), and deep learning (DL) has begun playing an increasingly important role in advancing POCT, particularly given their rapid development in recent years.131 Since the early 2020s, these technologies have been integrated with μPADs primarily for the following purposes:• Enhancement of sensitivity, accuracy, and selectivity.
• Reduction of assay time.
• Improvement of image capture in colorimetric approaches by minimizing ambient light interference.
• Mitigation of interference from complex biofluid sample matrices.
• Simplification of experimental optimization steps.
This field, especially the development of ML-assisted fully integrated μPADs, has been mainly advanced by the Ozcan group. Here, we introduce some relevant publications and discuss how their approaches achieved the objective listed above.
Joung et al. reported a DL-assisted vertical-flow μPAD for serological diagnosis of early-stage Lyme disease in 2020 (Fig. 8a).178 The nitrocellulose sensing membrane incorporated 13 immuno-reaction spots with different capture antigens or antigen-epitope containing peptides, as well as negative and positive controls. Two types of 3D-printed top cases were used for the uniform distribution of applied serum samples to all sensing spots and to generate colour signals through embedded gold nanoparticles (AuNPs) conjugated to anti-human IgM or IgG antibodies. By selecting an optimal subset of detection antigens from the 13 spots through ML and using batch-specific standardization and threshold tuning, they achieved an area-under-the-curve (AUC), sensitivity, and specificity of 0.963, 96.3%, and 85.7%, respectively. In the same year, Ballard et al. reported a similarly structured μPAD for C-reactive protein (hsCRP) testing.179 The device featured 81 immunoreaction spots of different antigens with different concentrations, from which optimal subsets were selected through ML as well. As a result, they successfully quantified hsCRP with an expanded dynamic range. As demonstrated by the examples above, increasing the number of result parameters enhances the performance of ML-based analysis. In this regard, signal detection techniques producing large scan-based datasets, such as electrochemical or SERS measurements, are particularly useful for ML or DL to enhance assay accuracy.180,181 On the other hand, simplification of multi analyte detecting assays using distinct reaction spots for each analyte is another strength of μPADs. However, Goncharov et al. introduced a different approach utilizing non-specific colour changes from multiple sensing spots for the quantification of three biomarkers, myoglobin, creatine kinase-MB (CK-MB), and heart-type fatty acid binding protein (FABP) using DL (Fig. 8b).182 While multivariable linear regression models only yielded coefficients of determination <0.85, neural networks improved these values to 0.92, 0.93, and 0.95 for myoglobin, CK-MB, and FABP, respectively. Similarly, in 2024, Eryilmaz et al. categorized COVID-19 immunity levels based on the IgG and IgM concentration levels.183 Furthermore, Ghosh et al. achieved single-tier serodiagnosis of Lyme disease by a single DL classifier trained to simultaneously interpret the response to IgM and IgG antibodies.184 In addition, inclusion of time-dependent colour change data into training datasets has been shown to contribute to sensitivity enhancement,185 as well as to reduction of assay time without sacrificing the analytical performance (Fig. 8c).185–187
 | ||
| Fig. 8 Examples of μPAD approaches with artificial intelligence integration: a) illustration of 13-immuno-reaction spots occurring on the nitrocellulose sensing membrane of a multiplexed paper-based immunoassay device for Lyme disease, expanded diagram showing paper layers of the device with two types of top holder, and image processing and deep learning (DL)-based analysis of the multiplexed sensing membrane. Adapted with permission from Joung et al.178 Copyright 2025 American Chemical Society. b) Overview of the multiplexed paper-based fluorescence vertical flow assay with multiple sensing spots, and neural network-based biomarker quantification of myoglobin, CK-MB and FABP. Adapted from Goncharov et al.182 c) Flow of image processing starting with image frame extraction from recorded video and architecture of the temporal sequence DL method in an ELISA-type μPAD for cardiac troponin I (cTnI) detection. Adapted with permission from Zhu et al.187 Copyright 2025 American Chemical Society. d) Neural network analysis-based correction of outlier clinical samples (yellow plots in left figure) caused by blood serum matrix interference in a chemiluminescence readout μPAD for cTnI. Adapted from Han et al.193 | ||
Besides data interpretation, ML and DL approaches have also been used to assist the capture and processing of images in colorimetric assays. Mercan et al. used ML to automatically select the optimal analysis region of an image in cases where colorimetric reagents were heterogeneously distributed across a circular assay spot.188 Compared with conventional colour threshold-based analysis, the ML-assisted approach was not affected by variations in seven different illumination conditions and four different smartphone models. Ning et al. further demonstrated result robustness not only to ambient light conditions but also to variations of image shooting angles.189 Based on other similar reports,190 these findings suggest that ML or DL-assisted μPADs can enhance user-friendliness and reliability of smartphone-based assays.
As noted above, ML and DL have also been adopted to mitigate the interference from complex biofluid sample matrices. Low et al. reported an ML-assisted μPAD for accurate acute early diagnosis and prognosis of myocardial infarction.191 After evaluating eleven machine-learning algorithms, they achieved an area under the receiver operating curves (AUROC) of 0.97 ± 0.018 for all possible classification thresholds. Similarly, Jang et al. reported DL-assisted μPADs integrated with field-effect transistors (FET) for cholesterol testing.192 In this system, cholesterol oxidase reacts with cholesterol to generate hydrogen peroxide, which modulates the FET's threshold voltage (ΔVth). While relying on ΔVth alone yielded a coefficient of determination of 0.81, the ML-integrated approach after training with optimal subset of specific range of time and gate voltage achieved high precision (CV < 6.46%) and reasonable correlation (r2 > 0.976) compared against results obtained from a certified laboratory. In another example, Han et al. focused on a small number of outlier clinical serum samples in cardiac troponin I detection using chemiluminescence (CL)-based vertical-flow μPADs with a sandwich immunoassay format.193 Three samples deviated significantly from the calibration curve based on normalized CL intensity (Fig. 8d). These deviations were attributed to serum matrix interference, which was effectively corrected by neural network-based analysis.
On the other hand, integrating ML or DL with μPADs resulted in the identification of unique features. Ning et al. noticed that including reagent batch IDs and fabrication batch IDs as input features during neural network training improved the final accuracy from 80% to 85%, highlighting the influence of batch-to-batch variations in both device fabrication and assay operation.189 Moreover, when developing optimization models for μPADs, flow-related behaviour must be incorporated into the governing equations, given the intrinsic dependence of paper-based microfluidics on capillary-driven transport.194
Very recently, a ML-assisted optimization approach was also extended to distance-based μPADs by Lv et al. in 2025.195 They utilized ML for parameter optimization and engineering manufacturing of distance-based μPADs, resulting in the reduction of experimental time and effort.
While ML- or DL-assisted μPADs contribute to unlock the full analytical potential of these devices as hardware, software, and assay chemistry continue to co-evolve, several considerations must be kept in mind. These include the complex regulatory landscape, the possibility of erroneous or illogical outputs caused by unrepresented or underrepresented cases in training datasets (e.g., rare diseases or specific patient subgroups), and the limited transparency of some ML systems.132 Implementing rigorous quality-control measures and recognizing that certain exceptional cases may remain unpredictable for ML or DL systems are therefore essential steps in responsible application of these techniques.
Conclusions
Since our previous review on μPADs published in this journal in 2017, the field has continued to advance toward real-world applications. At that time, we pointed out three major challenges for μPADs aimed at routine health checks: the necessity of complicated user operations, insufficient evaluation on long-term device storage stability, and reliance on detection equipment unfamiliar to general users. Thanks to the progress made in on-device sample pretreatment and smartphone integration, most studies targeting routine health checks have now achieved sample-in–answer-out functionality, and research focus has shifted towards the simplification of more complex assays, including immunoassays or nucleic acid testing. While assay operation has been simplified through automated sequential reagent delivery and device design innovation, there is still room for improvement in integration of off-device sample pretreatment steps such as for example blood separation and nucleic acid extraction. To comprehensively evaluate future reports on sample-in–answer-out μPAD systems, we emphasize the importance of assessing in parallel the trade-off between the necessity for end-user reagent handling and long-term device storage stability, and the relationship between the number of operation steps and off-device pretreatment requirements, together with time control needs. Meanwhile, long-term device storage stability, one of the challenges highlighted in our 2017 review, remains insufficiently addressed, even for μPADs intended for routine health checks performed by end-users in non-clinical settings, in addition to evaluation criteria for acceptable signal deterioration not being standardized. Apart from the development of sample-in–answer-out systems, recent studies exploring automated sequential reagent addition, increased liquid flow rates, and semi-quantitative signal readout approaches represent foundational concepts for achieving fully integrated μPAD versions for more complex assays in the future.Technologies such as SERS- and bioluminescence-based signal detection, CRISPR-based assay methods, and ML or DL-assisted image analysis and data interpretation have only more recently gained significant momentum in the field of μPADs. While SERS and bioluminescence still face major limitations in terms of the need of external Raman detecting modules and the instability of bioluminescent substrates, CRISPR-based assays and ML/DL approaches have already had significant impact in the field. Various CRISPR-based assays have been combined with μPADs due to the flexibility of their signal transduction mechanisms and are now actively contributing to the development of fully integrated NAT platforms. ML- and DL-assisted image analysis and data interpretation have enhanced μPAD performance without adding operational complexity, enabling shorter assay times without compromising sensitivity, minimizing sample matrix interference, and simplifying experimental optimization.
Finally, microchannel patterning methods on paper substrates are likely to become an increasingly critical issue for academic research. Currently, the wax printing method remains the most widely used technique for device prototyping in academic laboratories due to its simplicity, speed, and flexibility in design modification. However, with the discontinuation of commercially available wax printers, there is growing concern about the long-term sustainability of that approach. Although a range of alternative patterning methods has been reported,78,196–201 none have yet matched the ease of use and rapid turnaround time offered by wax printing.
Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included, and no new data were generated or analysed as part of this review.Supplementary information (SI): details of analytical performance of recent μPAD examples (2018–2025) in medical diagnostic applications (immunoassays, nucleic acid tests, etc.). See DOI: https://doi.org/10.1039/d5lc01085c.
Acknowledgements
Y. T. gratefully acknowledges funding from JST SPRING, Grant Number JPMJSP2123. During the preparation of this work the authors used ChatGPT to improve the readability and language of the manuscript. While using this tool, the authors reviewed and edited the text as needed and take full responsibility for the content of the published article.Notes and references
- J. P. Comer, Anal. Chem., 1956, 28, 1748–1750 CrossRef CAS.
- A. H. Free, E. C. Adams, M. L. Kercher, H. M. Free and M. H. Cook, Clin. Chem., 1957, 3, 163–168 CrossRef CAS.
- R. L. Campbell, D. B. Wagner and J. P. O'connell, US Pat., 4703017, 1987 Search PubMed.
- R. W. Rosenstein and T. G. Bloomster, US Pat., 4855240, 1989 Search PubMed.
- A. W. Martinez, S. T. Phillips, M. J. Butte and G. M. Whitesides, Angew. Chem., Int. Ed., 2007, 46, 1318–1320 CrossRef CAS PubMed.
- D. M. Cate, J. A. Adkins, J. Mettakoonpitak and C. S. Henry, Anal. Chem., 2015, 87, 19–41 CrossRef CAS PubMed.
- E. Noviana, T. Ozer, C. S. Carrell, J. S. Link, C. McMahon, I. Jang and C. S. Henry, Chem. Rev., 2021, 121, 11835–11885 Search PubMed.
- M.-N. Tsaloglou, D. C. Christodouleas, J. Milette, K. Milkey, I. C. Romine, J. Im, S. Lathwal, D. T. Selvam, H. D. Sikes and G. M. Whitesides, Lab Chip, 2025, 25, 741–751 RSC.
- E. Carrilho, A. W. Martinez and G. M. Whitesides, Anal. Chem., 2009, 81, 7091–7095 CrossRef CAS PubMed.
- K. Yamada, H. Shibata, K. Suzuki and D. Citterio, Lab Chip, 2017, 17, 1206–1249 Search PubMed.
- A. Sena-Torralba, R. Álvarez-Diduk, C. Parolo, A. Piper and A. Merkoçi, Chem. Rev., 2022, 122, 14881–14910 CrossRef CAS PubMed.
- J. Budd, B. S. Miller, N. E. Weckman, D. Cherkaoui, D. Huang, A. T. Decruz, N. Fongwen, G.-R. Han, M. Broto, C. S. Estcourt, J. Gibbs, D. Pillay, P. Sonnenberg, R. Meurant, M. R. Thomas, N. Keegan, M. M. Stevens, E. Nastouli, E. J. Topol, A. M. Johnson, M. Shahmanesh, A. Ozcan, J. J. Collins, M. Fernandez Suarez, B. Rodriguez, R. W. Peeling and R. A. McKendry, Nat. Rev. Bioeng., 2023, 1, 13–31 CrossRef CAS.
- J. L. Chen, D. I. Njoku, C. Tang, Y. Gao, J. Chen, Y.-K. Peng, H. Sun, G. Mao, M. Pan and N. F.-Y. Tam, Small Methods, 2024, 8, 2400155 CrossRef CAS PubMed.
- P. Aryal and C. S. Henry, Front. Lab Chip Technol., 2024, 3, 1467423 Search PubMed.
- P. M. Kalligosfyri and S. Cinti, ACS Mater. Lett., 2024, 6, 1184–1198 CrossRef CAS.
- J. D. Bishop, H. V. Hsieh, D. J. Gasperino and B. H. Weigl, Lab Chip, 2019, 19, 2486–2499 RSC.
- V.-T. Nguyen, S. Song, S. Park and C. Joo, Biosens. Bioelectron., 2020, 152, 112015 CrossRef CAS PubMed.
- J. R. Choi, K. W. Yong, R. Tang, Y. Gong, T. Wen, F. Li, B. Pingguan-Murphy, D. Bai and F. Xu, TrAC, Trends Anal. Chem., 2017, 93, 37–50 CrossRef CAS.
- Z. Li, Y. Bai, M. You, J. Hu, C. Yao, L. Cao and F. Xu, Biosens. Bioelectron., 2021, 177, 112952 CrossRef CAS PubMed.
- A. C. Mora and C. R. Mace, ACS Sens., 2025, 10, 3795–3805 CrossRef CAS PubMed.
- H. Fu, P. Song, Q. Wu, C. Zhao, P. Pan, X. Li, N. Y. K. Li-Jessen and X. Liu, Microsyst. Nanoeng., 2019, 5, 50 CrossRef.
- Y.-S. Chien, U.-I. Wu, Y.-H. Pan and C.-F. Chen, Biosens. Bioelectron., 2025, 287, 117724 CrossRef CAS PubMed.
- K. Tenda, B. Van Gerven, R. Arts, Y. Hiruta, M. Merkx and D. Citterio, Angew. Chem., Int. Ed., 2018, 57, 15369–15373 CrossRef CAS PubMed.
- D. Lee, C.-H. Chu and A. F. Sarioglu, ACS Sens., 2021, 6, 3204–3213 CrossRef CAS.
- P. Q. Nguyen, L. R. Soenksen, N. M. Donghia, N. M. Angenent-Mari, H. De Puig, A. Huang, R. Lee, S. Slomovic, T. Galbersanini, G. Lansberry, H. M. Sallum, E. M. Zhao, J. B. Niemi and J. J. Collins, Nat. Biotechnol., 2021, 39, 1366–1374 CrossRef CAS PubMed.
- T. Boonpoempoon, W. Wonsawat and T. Kaneta, Sci. Rep., 2019, 9, 12951 CrossRef PubMed.
- K. Inaba, T. Moriwaki, T. Maruyama and K. Morita, ACS Appl. Nano Mater., 2025, 8, 15572–15580 CrossRef CAS.
- T. Zhang, R. Deng, Y. Wang, C. Wu, K. Zhang, C. Wang, N. Gong, R. Ledesma-Amaro, X. Teng, C. Yang, T. Xue, Y. Zhang, Y. Hu, Q. He, W. Li and J. Li, Nat. Biomed. Eng., 2022, 6, 957–967 Search PubMed.
- S. K. Biswas, S. Chatterjee, S. Bandyopadhyay, S. Kar, N. K. Som, S. Saha and S. Chakraborty, ACS Sens., 2021, 6, 1077–1085 CrossRef CAS PubMed.
- C.-A. Chen, H. Yuan, C.-W. Chen, Y.-S. Chien, W.-H. Sheng and C.-F. Chen, Lab Chip, 2021, 21, 1908–1915 RSC.
- A. Yakoh, E. Mehmeti, K. Kalcher and S. Chaiyo, Anal. Chem., 2022, 94, 5893–5900 CrossRef CAS PubMed.
- S. Mansouri, S. Boulares, S. Chabchoub, Y. Alharbi and A. Alqahtani, Microchem. J., 2025, 209, 112670 CrossRef CAS.
- K. Yamada, K. Suzuki and D. Citterio, ACS Sens., 2017, 2, 1247–1254 CrossRef CAS.
- J. E. Butler, J. Immunoassay Immunochem., 2000, 21, 165–209 CrossRef CAS PubMed.
- E. Fu, T. Liang, P. Spicar-Mihalic, J. Houghtaling, S. Ramachandran and P. Yager, Anal. Chem., 2012, 84, 4574–4579 CrossRef CAS PubMed.
- A. Apilux, Y. Ukita, M. Chikae, O. Chailapakul and Y. Takamura, Lab Chip, 2013, 13, 126–135 RSC.
- B. Lutz, T. Liang, E. Fu, S. Ramachandran, P. Kauffman and P. Yager, Lab Chip, 2013, 13, 2840–2847 RSC.
- C. Srisomwat, A. Yakoh, N. Chuaypen, P. Tangkijvanich, T. Vilaivan and O. Chailapakul, Anal. Chem., 2021, 93, 2879–2887 CrossRef CAS PubMed.
- K. Yin, X. Ding, Z. Li, M. M. Sfeir, E. Ballesteros and C. Liu, Lab Chip, 2021, 21, 2730–2737 Search PubMed.
- J. Han, A. Qi, J. Zhou, G. Wang, B. Li and L. Chen, ACS Sens., 2018, 3, 1789–1794 CrossRef CAS PubMed.
- A. Yakoh, S. Chaiyo, W. Siangproh and O. Chailapakul, ACS Sens., 2019, 4, 1211–1221 CrossRef CAS PubMed.
- Y. Chen, Y. Hu and X. Lu, ACS Sens., 2023, 8, 2331–2339 Search PubMed.
- K. Maejima, Y. Hiruta and D. Citterio, Anal. Chem., 2020, 92, 4749–4754 Search PubMed.
- K. Khachornsakkul, W. Dungchai and N. Pamme, ACS Sens., 2022, 7, 2410–2419 CrossRef CAS PubMed.
- D. W. Bradbury, M. Azimi, A. J. Diaz, A. A. Pan, C. H. Falktoft, B. M. Wu and D. T. Kamei, Anal. Chem., 2019, 91, 12046–12054 CrossRef CAS PubMed.
- T. Akyazi, A. Tudor, D. Diamond, L. Basabe-Desmonts, L. Florea and F. Benito-Lopez, Sens. Actuators, B, 2018, 261, 372–378 Search PubMed.
- G.-R. Han, H. Ki and M.-G. Kim, ACS Appl. Mater. Interfaces, 2020, 12, 1885–1894 CrossRef CAS PubMed.
- R. Tang, H. Yang, J. R. Choi, Y. Gong, J. Hu, T. Wen, X. Li, B. Xu, Q. Mei and F. Xu, Microchim. Acta, 2017, 184, 2141–2150 CrossRef CAS.
- J. Park and J.-K. Park, Sens. Actuators, B, 2017, 246, 1049–1055 CrossRef CAS.
- E. B. Strong, C. Knutsen, J. T. Wells, A. R. Jangid, M. L. Mitchell, N. W. Martinez and A. W. Martinez, Inventions, 2019, 4, 20 CrossRef.
- D. Lee, T. Ozkaya-Ahmadov, C.-H. Chu, M. Boya, R. Liu and A. F. Sarioglu, Sci. Adv., 2021, 7, eabf9833 CrossRef CAS PubMed.
- C. Dai, H. Huang, Y. Zhang, Y. Liu, Y. Tao, T. Chen, Y. Zhang, C. Wan, S. Li, Z. Miao, Y. Li, P. Chen and B.-F. Liu, Biosens. Bioelectron., 2026, 292, 118083 Search PubMed.
- R. B. Channon, M. P. Nguyen, A. G. Scorzelli, E. M. Henry, J. Volckens, D. S. Dandy and C. S. Henry, Lab Chip, 2018, 18, 793–802 Search PubMed.
- N. Raj, V. Breedveld and D. W. Hess, Sens. Actuators, B, 2020, 320, 128606 CrossRef CAS.
- H. Fu, Z. Qin, X. Li, Y. Pan, H. Xu, P. Pan, P. Song and X. Liu, ACS Sens., 2023, 8, 3574–3584 CrossRef CAS PubMed.
- S. Yang, J. Zhu, L. Yang, H. Fa, Y. Wang, D. Huo, C. Hou, D. Zhong and M. Yang, ACS Sens., 2025, 10, 133–147 Search PubMed.
- H. Suzuki, G. Tong, P. Nath, Y. Hiruta and D. Citterio, ACS Sens., 2025, 10, 1811–1821 CrossRef CAS PubMed.
- P. Biswas, G. N. Mukunthan Sulochana, T. N. Banuprasad, P. Goyal, D. Modak, A. K. Ghosh and S. Chakraborty, ACS Sens., 2022, 7, 3720–3729 CrossRef CAS PubMed.
- S. Liu, T. Yu, L. Song, K. Kalantar-Zadeh and G. Liu, ACS Sens., 2025, 10, 4569–4579 Search PubMed.
- M. Wang, J. Qi, F. Li, Y. Liu, J. Xiang, B. Li, X. Zhuang, L. Chen and X. Fu, Sens. Actuators, B, 2026, 446, 138643 Search PubMed.
- E. W. Washburn, Phys. Rev., 1921, 17, 273–283 Search PubMed.
- D. M. Cate, W. Dungchai, J. C. Cunningham, J. Volckens and C. S. Henry, Lab Chip, 2013, 13, 2397–2404 RSC.
- J. E. Schonhorn, S. C. Fernandes, A. Rajaratnam, R. N. Deraney, J. P. Rolland and C. R. Mace, Lab Chip, 2014, 14, 4653–4658 Search PubMed.
- S. Jahanshahi-Anbuhi, P. Chavan, C. Sicard, V. Leung, S. M. Z. Hossain, R. Pelton, J. D. Brennan and C. D. M. Filipe, Lab Chip, 2012, 12, 5079–5085 RSC.
- C. K. Camplisson, K. M. Schilling, W. L. Pedrotti, H. A. Stone and A. W. Martinez, Lab Chip, 2015, 15, 4461–4466 RSC.
- J. A. Adkins, E. Noviana and C. S. Henry, Anal. Chem., 2016, 88, 10639–10647 Search PubMed.
- R. B. Channon, Y. Yang, K. M. Feibelman, B. J. Geiss, D. S. Dandy and C. S. Henry, Anal. Chem., 2018, 90, 7777–7783 CrossRef CAS PubMed.
- V. Soum, S. Park, A. I. Brilian, J.-Y. Choi, Y. Lee, W. Kim, O.-S. Kwon and K. Shin, Lab Chip, 2020, 20, 1601–1611 RSC.
- S. Boonkaew, A. Yakoh, N. Chuaypen, P. Tangkijvanich, S. Rengpipat, W. Siangproh and O. Chailapakul, Biosens. Bioelectron., 2021, 193, 113543 CrossRef CAS PubMed.
- K. Phoonsawat, T. Ozer, W. Dungchai and C. S. Henry, Analyst, 2022, 147, 4517–4524 Search PubMed.
- P. Aryal, A. W. Indrianingsih and C. S. Henry, Green Anal. Chem., 2024, 8, 100091 Search PubMed.
- C. Srisomwat, N. Bawornnithichaiyakul, S. Khonyoung, W. Tiyapongpattana, S. Butcha, N. Youngvises and O. Chailapakul, Talanta, 2024, 280, 126770 CrossRef CAS PubMed.
- X. Chen, H. Li, X. Li, M. Zhang, Q. He, J. Zhou, J. Zhong, H. Chen, Y. Shi, H. Chen, H. Zhang, L. T. Chu and W. Guo, Lab Chip, 2025, 25, 3839–3848 RSC.
- Y. B. Cho, D. C. Nguyen, S. H. Hua and Y. S. Kim, Microchem. J., 2024, 201, 110532 CrossRef CAS.
- P. Aryal, E. Brack, T. Alexander and C. S. Henry, Anal. Chem., 2023, 95, 5820–5827 Search PubMed.
- R. K. Arun, N. Priyadarshini, K. Chaudhury, N. Chanda, G. Biswas and S. Chakraborty, J. Micromech. Microeng., 2016, 26, 105008 CrossRef.
- H. Shigemori, K. Maejima, H. Shibata, Y. Hiruta and D. Citterio, Microchim. Acta, 2023, 190, 48 CrossRef CAS PubMed.
- C. Park, Y. D. Han, H. V. Kim, J. Lee, H. C. Yoon and S. Park, Lab Chip, 2018, 18, 1533–1538 RSC.
- A. Lomae, K. Teekayupak, P. Preechakasedkit, E. Pasomsub, T. Ozer, C. S. Henry, D. Citterio, T. Vilaivan, O. Chailapakul and N. Ruecha, Talanta, 2024, 279, 126613 CrossRef CAS PubMed.
- J. S. Link, J. O'Donnell-Sloan, S. Curdts, B. J. Geiss, D. S. Dandy and C. S. Henry, Anal. Chem., 2024, 96, 4111–4119 CrossRef CAS PubMed.
- C. Prakobdi, T. A. Baldo, P. Aryal, J. Link, P. Saetear and C. S. Henry, Anal. Methods, 2024, 16, 2489–2495 RSC.
- I. Macleod Briongos, Z. D. Call, C. S. Henry and D. L. Bark, Microfluid. Nanofluid., 2023, 27, 70 CrossRef PubMed.
- J. Yang, X. Feng, H. Li, P. Chang, X. Ren and X. Li, ACS Sens., 2025, 10, 5623–5632 CrossRef CAS PubMed.
- F. T. S. M. Ferreira, R. B. R. Mesquita and A. O. S. S. Rangel, Microchem. J., 2023, 193, 109102 Search PubMed.
- K. Khachornsakkul and W. Dungchai, ACS Sens., 2021, 6, 1339–1347 Search PubMed.
- Y. Li, L. Zhou, W. Ni, Q. Luo, C. Zhu and Y. Wu, Anal. Chem., 2019, 91, 14838–14841 CrossRef CAS PubMed.
- M. Celina, K. T. Gillen and R. A. Assink, Polym. Degrad. Stab., 2005, 90, 395–404 CrossRef CAS.
- S. Minnikanti, G. Diao, J. J. Pancrazio, X. Xie, L. Rieth, F. Solzbacher and N. Peixoto, Acta Biomater., 2014, 10, 960–967 CrossRef CAS PubMed.
- P. Panjan, V. Virtanen and A. M. Sesay, Talanta, 2017, 170, 331–336 CrossRef CAS PubMed.
- L. Hongtanee, S. Chaiyo, H. Wu and A. Yakoh, Adv. Healthcare Mater., 2025, 14, 2404320 CrossRef CAS PubMed.
- N. Thueankhum, L. Hongtanee, U. Boonyuen, A. Yakoh and K. Charoenkitamorn, Anal. Chem., 2025, 97, 20771–20779 CrossRef CAS PubMed.
- K. Fan, J. Zeng, C. Yang, G. Wang, K. Lian, X. Zhou, Y. Deng and G. Liu, ACS Sens., 2022, 7, 2049–2057 CrossRef CAS PubMed.
- A. Yamaguchi, S. Oyama, A. Ishida, T. Enomoto, N. Sanari, H. Miyaguchi and M. Tokeshi, ACS Appl. Bio Mater., 2025, 8, 1699–1706 CrossRef CAS PubMed.
- Y. Tanifuji, H. Suzuki, G. Tong, Y. Hiruta and D. Citterio, Anal. Methods, 2024, 16, 4143–4149 RSC.
- M. M. Calabretta, R. Álvarez-Diduk, E. Michelini, A. Roda and A. Merkoçi, Biosens. Bioelectron., 2020, 150, 111902 CrossRef CAS PubMed.
- I. Lewińska, M. Ścibisz and Ł. Tymecki, Anal. Chim. Acta, 2024, 1308, 342639 CrossRef PubMed.
- K. A. Ganaja, C. A. Chaplan, J. Zhang, N. W. Martinez and A. W. Martinez, Anal. Chem., 2017, 89, 5333–5341 CrossRef CAS PubMed.
- S. S. Nadar, P. D. Patil, M. S. Tiwari and D. J. Ahirrao, Crit. Rev. Biotechnol., 2021, 41, 1046–1080 CrossRef CAS PubMed.
- A. Baranwal, R. Shukla and V. Bansal, TrAC, Trends Anal. Chem., 2024, 172, 117573 CrossRef CAS.
- B. Das, J. L. Franco, N. Logan, P. Balasubramanian, M. I. Kim and C. Cao, Nano-Micro Lett., 2021, 13, 193 CrossRef CAS PubMed.
- K. Khachornsakkul, F. J. Rybicki and S. Sonkusale, Talanta, 2023, 260, 124538 CrossRef CAS PubMed.
- N. Kitchawengkul, A. Prakobkij, R. Saenmuangchin, D. Citterio, D. Nacapricha and P. Jarujamrus, Sens. Actuators, B, 2025, 435, 137671 CrossRef CAS.
- D. Cheng, J. Qin, Y. Feng and J. Wei, Biosensors, 2021, 11, 258 CrossRef CAS PubMed.
- L. Wang, Y. Chen, Y. Ji, L. Wang, X. Liu, F. Wang and C. Li, Anal. Chem., 2024, 96, 11353–11365 CrossRef CAS PubMed.
- S. E. Son, S. H. Cheon, W. Hur, H. B. Lee, D. H. Kim, C. H. Ha, S. J. Lee, D. K. Han and G. H. Seong, Biosens. Bioelectron., 2024, 243, 115752 CrossRef CAS PubMed.
- S. Yang, L. Zhao, X. Yang, L. Yang, H. Fa, Y. Wang, D. Huo, C. Hou, D. Zhong and M. Yang, Talanta, 2024, 278, 126523 CrossRef CAS PubMed.
- Y. Qin, Y. Qin, J. Sun, J. Qian, J. Yuan, J. Yao and M. Zhang, Chem. Eng. J., 2025, 514, 163033 Search PubMed.
- Q. H. Nguyen, D. H. Lee, P. T. Nguyen, P. G. Le and M. I. Kim, Chem. Eng. J., 2023, 454, 140541 CrossRef CAS.
- G. G. Lewis, M. J. DiTucci and S. T. Phillips, Angew. Chem., Int. Ed., 2012, 51, 12707–12710 CrossRef CAS PubMed.
- Y. Zhang, J. Fan, J. Nie, S. Le, W. Zhu, D. Gao, J. Yang, S. Zhang and J. Li, Biosens. Bioelectron., 2015, 73, 13–18 CrossRef CAS PubMed.
- M. Li, J. Tian, M. Al-Tamimi and W. Shen, Angew. Chem., Int. Ed., 2012, 51, 5497–5501 CrossRef CAS PubMed.
- S. Lee, J. Park and J.-K. Park, Sens. Actuators, B, 2018, 273, 322–327 CrossRef CAS.
- A. Teprek, V. Poetri Artono, W. Waiyawat, A. Limsakul, J. Shiowatana and A. Siripinyanond, Microchem. J., 2020, 158, 105284 CrossRef CAS.
- K. Khachornsakkul, D. Phuengkasem, K. Palkuntod, W. Sangkharoek, O. Jamjumrus and W. Dungchai, ACS Sens., 2022, 7, 2093–2101 Search PubMed.
- A. Katoh, K. Maejima, Y. Hiruta and D. Citterio, Analyst, 2020, 145, 6071–6078 RSC.
- M. Yang, W. Zhang, W. Zheng, F. Cao and X. Jiang, Lab Chip, 2017, 17, 3874–3882 RSC.
- Y. Manmana, S. Kinugasa, Y. Hiruta and D. Citterio, Anal. Chem., 2025, 97, 1500–1506 CrossRef CAS PubMed.
- A. Burklund, H. K. Saturley-Hall, F. A. Franchina, J. E. Hill and J. X. J. Zhang, Biosens. Bioelectron., 2019, 128, 97–103 CrossRef CAS PubMed.
- K. Yamada, D. Citterio and C. S. Henry, Lab Chip, 2018, 18, 1485–1493 RSC.
- S. Ohta, R. Hiraoka, Y. Hiruta and D. Citterio, Lab Chip, 2022, 22, 717–726 RSC.
- S. M. Russell, A. Alba-Patiño, M. Borges and R. de la Rica, ACS Sens., 2018, 3, 1712–1718 Search PubMed.
- Y.-Q. Yang, Y.-C. Yang, M.-H. Liu and Y.-H. Chan, Anal. Chem., 2020, 92, 1493–1501 CrossRef CAS PubMed.
- V. Oncescu, D. O'Dell and D. Erickson, Lab Chip, 2013, 13, 3232–3238 Search PubMed.
- A. K. Yetisen, J. L. Martinez-Hurtado, A. Garcia-Melendrez, F. da Cruz Vasconcellos and C. R. Lowe, Sens. Actuators, B, 2014, 196, 156–160 Search PubMed.
- G. Chen, C. Fang, H. H. Chai, Y. Zhou, W. Yun Li and L. Yu, Sens. Actuators, B, 2019, 281, 253–261 CrossRef CAS.
- S. Nie and S. R. Emory, Science, 1997, 275, 1102–1106 CrossRef CAS PubMed.
- K. Kneipp, Y. Wang, H. Kneipp, L. T. Perelman, I. Itzkan, R. R. Dasari and M. S. Feld, Phys. Rev. Lett., 1997, 78, 1667–1670 Search PubMed.
- A. Roda, M. Mirasoli, E. Michelini, M. Di Fusco, M. Zangheri, L. Cevenini, B. Roda and P. Simoni, Biosens. Bioelectron., 2016, 76, 164–179 CrossRef CAS PubMed.
- J. S. Gootenberg, O. O. Abudayyeh, J. W. Lee, P. Essletzbichler, A. J. Dy, J. Joung, V. Verdine, N. Donghia, N. M. Daringer and C. A. Freije, Science, 2017, 356, 438–442 CrossRef CAS PubMed.
- J. S. Chen, E. Ma, L. B. Harrington, M. Da Costa, X. Tian, J. M. Palefsky and J. A. Doudna, Science, 2018, 360, 436–439 CrossRef CAS PubMed.
- M. Bhaiyya, D. Panigrahi, P. Rewatkar and H. Haick, ACS Sens., 2024, 9, 4495–4519 CrossRef CAS PubMed.
- G.-R. Han, A. Goncharov, M. Eryilmaz, S. Ye, B. Palanisamy, R. Ghosh, F. Lisi, E. Rogers, D. Guzman, D. Yigci, S. Tasoglu, D. Di Carlo, K. Goda, R. A. McKendry and A. Ozcan, Nat. Commun., 2025, 16, 3165 CrossRef CAS PubMed.
- J. Langer, D. Jimenez de Aberasturi, J. Aizpurua, R. A. Alvarez-Puebla, B. Auguié, J. J. Baumberg, G. C. Bazan, S. E. J. Bell, A. Boisen, A. G. Brolo, J. Choo, D. Cialla-May, V. Deckert, L. Fabris, K. Faulds, F. J. García de Abajo, R. Goodacre, D. Graham, A. J. Haes, C. L. Haynes, C. Huck, T. Itoh, M. Käll, J. Kneipp, N. A. Kotov, H. Kuang, E. C. Le Ru, H. K. Lee, J.-F. Li, X. Y. Ling, S. A. Maier, T. Mayerhöfer, M. Moskovits, K. Murakoshi, J.-M. Nam, S. Nie, Y. Ozaki, I. Pastoriza-Santos, J. Perez-Juste, J. Popp, A. Pucci, S. Reich, B. Ren, G. C. Schatz, T. Shegai, S. Schlücker, L.-L. Tay, K. G. Thomas, Z.-Q. Tian, R. P. Van Duyne, T. Vo-Dinh, Y. Wang, K. A. Willets, C. Xu, H. Xu, Y. Xu, Y. S. Yamamoto, B. Zhao and L. M. Liz-Marzán, ACS Nano, 2020, 14, 28–117 CrossRef CAS PubMed.
- T. Itoh, M. Procházka, Z.-C. Dong, W. Ji, Y. S. Yamamoto, Y. Zhang and Y. Ozaki, Chem. Rev., 2023, 123, 1552–1634 CrossRef CAS PubMed.
- C. H. Lee, L. Tian and S. Singamaneni, ACS Appl. Mater. Interfaces, 2010, 2, 3429–3435 CrossRef CAS PubMed.
- W. W. Yu and I. M. White, Analyst, 2013, 138, 1020–1025 RSC.
- Y. Cao, Y. Sun, R.-J. Yu and Y.-T. Long, Microchim. Acta, 2023, 191, 8 CrossRef PubMed.
- V. Eskandari, H. Sahbafar, L. Zeinalizad, R. Marashipour and A. Hadi, J. Comput. Appl. Mech., 2022, 53, 142–156 Search PubMed.
- W. Y. Lim, C.-H. Goh, T. M. Thevarajah, B. T. Goh and S. M. Khor, Biosens. Bioelectron., 2020, 147, 111792 CrossRef CAS PubMed.
- S. Mabbott, S. C. Fernandes, M. Schechinger, G. L. Cote, K. Faulds, C. R. Mace and D. Graham, Analyst, 2020, 145, 983–991 RSC.
- S. Lee, H. Dang, J.-I. Moon, K. Kim, Y. Joung, S. Park, Q. Yu, J. Chen, M. Lu, L. Chen, S.-W. Joo and J. Choo, Chem. Soc. Rev., 2024, 53, 5394–5427 RSC.
- F. Zeng, W. Duan, B. Zhu, T. Mu, L. Zhu, J. Guo and X. Ma, Anal. Chem., 2019, 91, 1064–1070 Search PubMed.
- F. Zeng, T. Mou, C. Zhang, X. Huang, B. Wang, X. Ma and J. Guo, Analyst, 2019, 144, 137–142 Search PubMed.
- R. Griss, A. Schena, L. Reymond, L. Patiny, D. Werner, C. E. Tinberg, D. Baker and K. Johnsson, Nat. Chem. Biol., 2014, 10, 598–603 Search PubMed.
- L. Xue, Q. Yu, R. Griss, A. Schena and K. Johnsson, Angew. Chem., Int. Ed., 2017, 56, 7112–7116 Search PubMed.
- K. Tomimuro, K. Tenda, Y. Ni, Y. Hiruta, M. Merkx and D. Citterio, ACS Sens., 2020, 5, 1786–1794 Search PubMed.
- Y. Yang, A. Inoue, T. Yasuda, H. Ueda, B. Zhu and T. Kitaguchi, ACS Sens., 2024, 9, 5955–5965 Search PubMed.
- A. A. Shalaby, A. Salah, A. Ishida, M. Maeki and M. Tokeshi, Biosens. Bioelectron.: X, 2025, 24, 100621 Search PubMed.
- H. Martínez-Pérez-Cejuela, M. M. Calabretta, V. Bocci, M. D’Elia and E. Michelini, Biosensors, 2023, 13, 451 Search PubMed.
- J. P. Hunt, E. L. Zhao, T. J. Free, M. Soltani, C. A. Warr, A. B. Benedict, M. K. Takahashi, J. S. Griffitts, W. G. Pitt and B. C. Bundy, New Biotechnol., 2022, 66, 53–60 Search PubMed.
- A. Lopreside, L. Montali, B. Wang, A. Tassoni, M. Ferri, M. M. Calabretta and E. Michelini, Biosens. Bioelectron., 2021, 194, 113569 Search PubMed.
- F. Nazir, D. Gregucci, M. M. Calabretta, C. Cambrea, P. Vahidi, S. Lavrnić, A. Toscano and E. Michelini, Anal. Chem., 2025, 97, 18092–18100 Search PubMed.
- R. Kurita and R. Nishihara, Sens. Actuators, B, 2025, 423, 136700 CrossRef CAS.
- R. Shimazu, K. Tomimuro, Y. Ni, C. Malegori, V. Hamedpour, Y. Hiruta, P. Oliveri, M. Merkx and D. Citterio, Sens. Actuators, B, 2022, 352, 131002 Search PubMed.
- Y. Mizui, M. Eguchi, M. Tanaka, Y. Ikeda, H. Yoshimura, T. Ozawa, D. Citterio and Y. Hiruta, Org. Biomol. Chem., 2021, 19, 579–586 RSC.
- M. Jinek, K. Chylinski, I. Fonfara, M. Hauer, J. A. Doudna and E. Charpentier, Science, 2012, 337, 816–821 Search PubMed.
- Y. Dai, Y. Wu, G. Liu and J. J. Gooding, Angew. Chem., Int. Ed., 2020, 59, 20754–20766 Search PubMed.
- M. M. Kaminski, O. O. Abudayyeh, J. S. Gootenberg, F. Zhang and J. J. Collins, Nat. Biomed. Eng., 2021, 5, 643–656 Search PubMed.
- K. Pardee, A. A. Green, M. K. Takahashi, D. Braff, G. Lambert, J. W. Lee, T. Ferrante, D. Ma, N. Donghia and M. Fan, Cell, 2016, 165, 1255–1266 Search PubMed.
- J. S. Gootenberg, O. O. Abudayyeh, M. J. Kellner, J. Joung, J. J. Collins and F. Zhang, Science, 2018, 360, 439–444 CrossRef CAS PubMed.
- M. J. Kellner, J. G. Koob, J. S. Gootenberg, O. O. Abudayyeh and F. Zhang, Nat. Protoc., 2019, 14, 2986–3012 Search PubMed.
- J. P. Broughton, X. Deng, G. Yu, C. L. Fasching, V. Servellita, J. Singh, X. Miao, J. A. Streithorst, A. Granados, A. Sotomayor-Gonzalez, K. Zorn, A. Gopez, E. Hsu, W. Gu, S. Miller, C.-Y. Pan, H. Guevara, D. A. Wadford, J. S. Chen and C. Y. Chiu, Nat. Biotechnol., 2020, 38, 870–874 Search PubMed.
- V. Lapee-e, S. Nuanualsuwan, L. Hongtanee and A. Yakoh, Microchem. J., 2025, 211, 113055 Search PubMed.
- S. Dalgan and Q. Wei, Adv. Sens. Res., 2025, e00036 Search PubMed.
- D. Huang, D. Ni, M. Fang, Z. Shi and Z. Xu, Anal. Chem., 2021, 93, 16965–16973 Search PubMed.
- P. Li, S. Xu, Z. Dong, H. Liu, J. Huang, X. Deng, Y. Tao, H. Liu, Z. Lin and Z. Li, Biosens. Bioelectron., 2024, 271, 117087 Search PubMed.
- R. Yang, X. Guan, J. Zhang, J. Moon, C. Guo, Z. Jia, C. Hou, U. Ganbaatar, S. Tricarico, A. W. K. Ma and C. Liu, Biosens. Bioelectron., 2024, 271, 117054 CrossRef PubMed.
- X. Zhao, Y. He, S. Shao, Q. Ci, L. Chen, X. Lu, Q. Liu and J. Chen, ACS Sens., 2024, 9, 2413–2420 CrossRef CAS PubMed.
- X. Guan, R. Yang, J. Zhang, J. Moon, C. Hou, C. Guo, L. Avery, D. Scarola, D. S. Roberts, R. LaSala and C. Liu, Adv. Sci., 2025, 12, 2411021 Search PubMed.
- E. Hanson, N. Kalla, R. J. Tharu, M. M. Demir, B. H. Tok, M. A. Canbaz and M. V. Yigit, Small, 2025, 21, 2500784 Search PubMed.
- M. A. English, L. R. Soenksen, R. V. Gayet, H. de Puig, N. M. Angenent-Mari, A. S. Mao, P. Q. Nguyen and J. J. Collins, Science, 2019, 365, 780–785 CrossRef CAS PubMed.
- L. Li, G. Wang, Y. Rong, Y. Zhang, J. Huang, Y. Zhang, S. Ge, K. Cui, L. Zhang and J. Yu, Nano Energy, 2023, 116, 108768 Search PubMed.
- Y. Zhang, L. Gao, Z. Shi, Q. Wu and X. Miao, Bioelectrochemistry, 2025, 164, 108916 Search PubMed.
- Y. Zhang, K. Lv, Y. Zhang, Q. Mi, D. Ye, X. Chen and J. Zhang, Chem. Eng. J., 2025, 520, 165900 Search PubMed.
- A. Sen, M. Masetty, S. Weerakoon, C. Morris, J. S. Yadav, S. Apewokin, J. Trannguyen, M. Broom and A. Priye, Biosens. Bioelectron., 2024, 257, 116292 Search PubMed.
- Y. Joung, D. K. Han, H. Jang, T. Kang, L. Chen and J. Choo, ACS Sens., 2025, 10, 6253–6262 Search PubMed.
- Y. Tanifuji, G. Tong, Y. Hiruta and D. Citterio, Analyst, 2024, 149, 4932–4939 Search PubMed.
- H.-A. Joung, Z. S. Ballard, J. Wu, D. K. Tseng, H. Teshome, L. Zhang, E. J. Horn, P. M. Arnaboldi, R. J. Dattwyler, O. B. Garner, D. Di Carlo and A. Ozcan, ACS Nano, 2020, 14, 229–240 CrossRef CAS PubMed.
- Z. S. Ballard, H.-A. Joung, A. Goncharov, J. Liang, K. Nugroho, D. Di Carlo, O. B. Garner and A. Ozcan, npj Digit. Med., 2020, 3, 66 Search PubMed.
- Q. Bao, G. Li, W. Cheng, Z. Yang, Z. Qu, J. Wei and L. Lin, RSC Adv., 2023, 13, 23788–23795 RSC.
- L. Bi, H. Zhang, C. Mu, K. Sun, H. Chen, Z. Zhang and L. Chen, J. Hazard. Mater., 2025, 494, 138694 Search PubMed.
- A. Goncharov, H.-A. Joung, R. Ghosh, G.-R. Han, Z. S. Ballard, Q. Maloney, A. Bell, C. T. Z. Aung, O. B. Garner, D. D. Carlo and A. Ozcan, Small, 2023, 19, 2300617 Search PubMed.
- M. Eryilmaz, A. Goncharov, G.-R. Han, H.-A. Joung, Z. S. Ballard, R. Ghosh, Y. Zhang, D. Di Carlo and A. Ozcan, ACS Nano, 2024, 18, 16819–16831 CrossRef CAS PubMed.
- R. Ghosh, H.-A. Joung, A. Goncharov, B. Palanisamy, K. Ngo, K. Pejcinovic, N. Krockenberger, E. J. Horn, O. B. Garner, E. Ghazal, A. O'Kula, P. M. Arnaboldi, R. J. Dattwyler, A. Ozcan and D. Di Carlo, Nat. Commun., 2024, 15, 7124 CrossRef CAS PubMed.
- G.-R. Han, A. Goncharov, M. Eryilmaz, H.-A. Joung, R. Ghosh, G. Yim, N. Chang, M. Kim, K. Ngo, M. Veszpremi, K. Liao, O. B. Garner, D. Di Carlo and A. Ozcan, ACS Nano, 2024, 18, 27933–27948 CrossRef CAS PubMed.
- H. Sun, L. Xiong, Y. Huang, X. Chen, Y. Yu, S. Ye, H. Dong, Y. Jia and W. Zhang, Fundam. Res., 2022, 2, 476–486 CrossRef CAS PubMed.
- J. Zhu, K. Hoettges, Y. Wang, H. Ma, P. Song, Y. Hu, E. G. Lim and Q. Zhang, Anal. Chem., 2025, 97, 4515–4523 CrossRef CAS PubMed.
- Ö. B. Mercan, V. Kılıç and M. Şen, Sens. Actuators, B, 2021, 329, 129037 Search PubMed.
- Q. Ning, W. Zheng, H. Xu, A. Zhu, T. Li, Y. Cheng, S. Feng, L. Wang, D. Cui and K. Wang, Anal. Bioanal. Chem., 2022, 414, 3959–3970 Search PubMed.
- S. Duan, T. Cai, J. Zhu, X. Yang, E. G. Lim, K. Huang, K. Hoettges, Q. Zhang, H. Fu, Q. Guo, X. Liu, Z. Yang and P. Song, Anal. Chim. Acta, 2023, 1248, 340868 CrossRef CAS PubMed.
- J. S. Y. Low, T. M. Thevarajah, S. W. Chang and S. M. Khor, Sens. Actuators, B, 2023, 394, 134403 Search PubMed.
- H.-J. Jang, H.-A. Joung, A. Goncharov, A. G. Kanegusuku, C. W. Chan, K.-T. J. Yeo, W. Zhuang, A. Ozcan and J. Chen, ACS Nano, 2024, 18, 24792–24802 CrossRef CAS PubMed.
- G.-R. Han, A. Goncharov, M. Eryilmaz, S. Ye, H.-A. Joung, R. Ghosh, E. Ngo, A. Tomoeda, Y. Lee, K. Ngo, E. Melton, O. B. Garner, D. Di Carlo and A. Ozcan, Small, 2025, 21, 2411585 CrossRef CAS PubMed.
- D. M. Y. Tay, S. Kim, Y. Hao, E. H. Yee, H. Jia, S. M. Vleck, M. Chilekwa, J. Voldman and H. D. Sikes, Biosens. Bioelectron., 2023, 222, 114977 Search PubMed.
- K. Lv, Y. Zhang, K. Tang, W. Huang, F. Chen, M. Chen, Y. Wang and J. Zhang, ACS Sens., 2025, 10, 7002–7013 CrossRef CAS PubMed.
- A. Chauhan and B. J. Toley, Anal. Chem., 2021, 93, 8954–8961 Search PubMed.
- N. Raj, V. Breedveld and D. W. Hess, Microfluid. Nanofluid., 2021, 25, 47 Search PubMed.
- R. A. Ruiz, J. L. Gonzalez, M. Vazquez-Alvarado, N. W. Martinez and A. W. Martinez, Anal. Chem., 2022, 94, 8833–8837 Search PubMed.
- S. Seetasang and T. Kaneta, ACS Sens., 2022, 7, 1194–1200 Search PubMed.
- T. Monju, M. Hirakawa, S. Kuboyama, R. Saiki and A. Ishida, Sens. Actuators, B, 2023, 375, 132886 Search PubMed.
- L. Meng, D. Cao, J. O. Pedersen, G. Greczynski, V. Rogoz, W. Limbut and M. Eriksson, ACS Appl. Mater. Interfaces, 2025, 17, 39719–39731 Search PubMed.
- L. Gutiérrez-Gálvez, N. Seddaoui, L. Fiore, L. Fabiani, T. García-Mendiola, E. Lorenzo and F. Arduini, ACS Sens., 2024, 9, 4047–4057 Search PubMed.
- S. Kim, Y. Hao, E. A. Miller, D. M. Y. Tay, E. Yee, P. Kongsuphol, H. Jia, M. McBee, P. R. Preiser and H. D. Sikes, ACS Sens., 2021, 6, 1891–1898 Search PubMed.
- S. Liu, Y. Hou, Z. Li, C. Yang and G. Liu, ACS Sens., 2023, 8, 3520–3529 CrossRef CAS PubMed.
- S. Jackson, S. Lee and A. K. Badu-Tawiah, Anal. Chem., 2022, 94, 5132–5139 CrossRef CAS PubMed.
- B. S. Batule, Y. Seok and M.-G. Kim, Biosens. Bioelectron., 2020, 151, 111998 Search PubMed.
- B. P. Sullivan, A. T. Bender, D. N. Ngyuen, J. Y. Zhang and J. D. Posner, J. Chromatogr., B, 2021, 1163, 122494 Search PubMed.
- I. Choopara, A. Suea-Ngam, Y. Teethaisong, P. D. Howes, M. Schmelcher, A. Leelahavanichkul, S. Thunyaharn, D. Wongsawaeng, A. J. Demello, D. Dean and N. Somboonna, ACS Sens., 2021, 6, 742–751 CrossRef CAS PubMed.
- S. Cui, K. Wang, Y. Yang, X. Lv and X. Li, Anal. Bioanal. Chem., 2024, 417, 785–797 CrossRef PubMed.
- E. A. Phillips, T. J. Moehling, K. F. K. Ejendal, O. S. Hoilett, K. M. Byers, L. A. Basing, L. A. Jankowski, J. B. Bennett, L.-K. Lin, L. A. Stanciu and J. C. Linnes, Lab Chip, 2019, 19, 3375–3386 RSC.
- L. Bezinge, J. M. Lesinski, A. Suea-Ngam, D. A. Richards, A. J. deMello and C.-J. Shih, Adv. Mater., 2023, 35, 2302893 CrossRef CAS PubMed.
- R. Hiraoka, K. Kuwahara, Y.-C. Wen, T.-H. Yen, Y. Hiruta, C.-M. Cheng and D. Citterio, ACS Sens., 2020, 5, 1110–1118 Search PubMed.
- C. Park, H.-R. Kim, S.-K. Kim, I.-K. Jeong, J.-C. Pyun and S. Park, ACS Appl. Mater. Interfaces, 2019, 11, 36428–36434 Search PubMed.
- D. Kim, S. Kim and S. Kim, Analyst, 2020, 145, 5491–5499 RSC.
- J. Xu, H. Khan and L. Yang, Anal. Chem., 2021, 93, 14755–14763 Search PubMed.
- J. Cheng, Y. Fu, J. Guo and J. Guo, Sens. Actuators, B, 2023, 387, 133795 CrossRef CAS.
- M. M. P. Melo, A. O. S. S. Rangel and R. B. R. Mesquita, Talanta, 2026, 298, 128865 Search PubMed.
- J. Qi, B. Li, D. Lin, Z. Du, L. Fu, X. Wang, Z. Zhang, L. Luo and L. Chen, Anal. Chem., 2023, 95, 10486–10491 CrossRef CAS PubMed.
- S. Arif, M. Hassan, S. Rasheed, N. Ahmad, B. Fatima, S. G. Musharraf and D. Hussain, Biosens. Bioelectron., 2026, 291, 118025 CrossRef CAS PubMed.
- C. Chen, B. Ran, B. Liu, Y. Xiao, G. Tang, T. Shen, J. Liang and Z. Liang, Microchem. J., 2025, 210, 112966 CrossRef CAS.
- T. Komatsu, M. Maeki, A. Ishida, H. Tani and M. Tokeshi, ACS Sens., 2020, 5, 1287–1294 CrossRef CAS PubMed.
- M. Karimian, K. Dashtian, R. Zare-Dorabei and S. Norouzi, Anal. Chim. Acta, 2024, 1285, 342022 CrossRef CAS PubMed.
- K. Phoonsawat, K. Khachornsakkul, N. Ratnarathorn, C. S. Henry and W. Dungchai, ACS Sens., 2021, 6, 3047–3055 CrossRef CAS PubMed.
- D. Tu, A. Holderby, J. Dean, S. Mabbott and G. L. Coté, Anal. Chem., 2021, 93, 4497–4505 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |


