Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Size-based sorting of cancer cells reveals functional heterogeneity among subpopulations
Esra Yilmaz a,
Zhimeng Fanb,
Jason P. Beech
a,
Zhimeng Fanb,
Jason P. Beech a,
Vinay S. Swaminathan
a,
Vinay S. Swaminathan *b and
Jonas O. Tegenfeldt
*b and
Jonas O. Tegenfeldt *a
*a
aDepartment of Physics and NanoLund, Lund University, Lund, Sweden. E-mail: jonas.tegenfeldt@fysik.lu.se
bDepartment of Clinical Sciences, and The Wallenberg Centre for Molecular Medicine, Lund University, Lund, Sweden. E-mail: vinay.swaminathan@med.lu.se
First published on 25th March 2026
Abstract
Cancer cells display marked heterogeneity in size and morphology, traits long recognized in pathology as indicators of pleomorphism and poor prognosis. Yet, the functional significance and phenotypic consequences of these morphological variations within a single cancer cell population remain poorly understood. Here, we employed a deterministic lateral displacement (DLD) microfluidic device with a circular micro-pillar array to fractionate MDA-MB-231 breast cancer cells into distinct small and large subpopulations based on size. Following sorting, the isolated fractions were expanded in culture and subjected to assays to measure proliferation, migration and invasion. We found that these subpopulations maintained their characteristic sizes over several days and exhibited distinct functional behaviors that differentially contributed to the overall invasive phenotype of MDA-MB-231 cells. Notably, while the two subpopulations proliferated at the same rate, the small-cell fraction displayed enhanced migratory capacity and invasion both in 2D and 3D assays, thus representing the primary drivers of the invasive phenotype of MDA-MB-231 cells. Together, our findings demonstrate that label-free, size-based sorting reveals biologically meaningful functional heterogeneity within morphologically diverse cancer cell populations.
1 Introduction
Tumors consist of diverse subpopulations of cells that differ phenotypically, a heterogeneity linked to rapid growth, metastasis, plasticity, and therapy resistance.1–3 This heterogeneity arises from a number of different factors including genetic instability, non-genetic cellular plasticity, and influences from the tumor microenvironment such as nutrient availability and signaling. It is this heterogeneity that makes cancer cell profiling, diagnosis and treatment outcomes highly complex and challenging.4,5Besides genetic and cell state heterogeneity, cancer cells also exhibit morphological and biophysical variability. Among morphological features, pleomorphism, defined as variation in cell and nuclear size and shape, is central to tumor grading and prognosis6 and is routinely used in pathology as indicators of malignancy and correlated with poor clinical outcomes.7 Moreover, cell morphology, including cell size, is not only diagnostic but can also serve as a functional axis of biological diversity. For example, quantitative measures such as spread area, roundness, and irregularity have recently been shown to predict tumorigenicity and metastatic potential in breast cancer models more effectively than motility assays.8 Similarly, epithelial tissues exhibit cell-size pleomorphism that drives aberrant clone dispersal and influences tissue organization.9 In addition, changes in cell size and shape occur during the cell cycle and in response to local crowding, showing that morphological heterogeneity reflects dynamic and context-dependent cellular states.10 Interestingly, such heterogeneity is not limited to tumors in vivo; even established cancer cell lines, which are widely used as disease models, contain subpopulations that differ in morphology and behavior.11,12 However, whole population-level measurements of size often average out these differences, obscuring functionally distinct subsets withing a given population that may drive divergent biological responses and clinical outcomes.13,14
Given that morphological heterogeneity is biologically meaningful, it becomes important to isolate size-defined subpopulations and test whether they differ in functional traits such as proliferation, motility, and invasion. Traditional sorting methods like fluorescence-activated cell sorting (FACS) and magnetic-activated cell sorting (MACS) rely on antigen markers, which may perturb cells or fail to capture heterogeneity defined purely by morphology or other physical parameters.15,16
Label-free microfluidic methods exploit intrinsic biophysical properties—such as size, deformability, and stiffness—to sort cells without biochemical labels.17–20 These techniques can be classified into two groups, one in which external fields or forces are applied such as acoustophoresis, electrokinetics, and magnetophoresis, and another relying purely on hydrodynamic approaches (inertial focusing, deterministic lateral displacement, hydrodynamic filtration), both enabling scalable and reproducible isolation of functional subpopulations.20–26 Complementary tools like real-time deformability cytometry (RT-DC) extend this concept by rapidly quantifying mechanical phenotypes in flow.27
Among these, deterministic lateral displacement (DLD) is a well-established technique for separating particles and cells by size. DLD is characterized by a critical diameter, DC, such that particles with diameter D < DC are able to follow the flow in a zig-zag motion through a pillar array, while particles with D > DC are laterally displaced via interactions with the pillars, moving away from their original stream.28,29 DLD has been applied to blood cells, circulating tumor cells, and stem cells,30,31 demonstrating high throughput and viability retention. However, few studies have used DLD to sort solid tumor cells by size and then examined the biological significance of these subpopulations in the context of cancer cell heterogeneity and metastatic behavior.
In this study, we used DLD to separate the triple-negative breast cancer cell line MDA-MB-231 into fractions of small and large cells that together account for the size heterogeneity observed in the overall population. We found that the sorted fractions retained their respective size profiles after expansion for at least seven days, indicating that cell size was a stable morphological feature of cancer cells. We next examined whether these size-defined subpopulations represent distinct phenotypic and functional states. Functional assays revealed that, while proliferation rates between these 2 subgroups were comparable, smaller cells consistently exhibited greater migratory and invasive capacities in both two- and three-dimensional assays. These findings demonstrate that morphological diversity within a cancer cell population reflects stable and functionally distinct subpopulations.
2 Materials and methods
2.1 DLD device design
In the simplest description of DLD, particles are split into two size fractions around the critical size DC, with the two fractions following well defined trajectories that can be routed to separate device outlets. This simple behaviour is complicated by so-called dispersive or mixed mode trajectories which lie between zigzag and displacement mode trajectories and have been both predicted by simulations and observed in experiments.32–35 In addition, non-spherical and dynamically deformed particles have no well-defined single size, adding a random component to their trajectories. These dispersive trajectories are common for complex samples such as cells. They can lead to the routing of cells to the wrong outlets (with respect to size) and a considerable decrease in separation quality. We designed our devices with a third, intermediate outlet, between the zigzag and displacement outlets, which allows us to remove from downstream analysis, the majority of cells moving in dispersive, or mixed modes (for details see SI). This way we remove cells that lack a well-defined size and cells that follow anomalous trajectories due to the known possible artifacts of the DLD mechanism. While this was expected to lead to a drop in the number of recovered cells, it was expected to greatly improve purity of the two relevant fractions of cells moving in the zigzag (small) and displacement (large) modes.2.2 Fabrication and fluidics
The photomask for UV lithography was designed in L-Edit v.18.3 (Tanner Research, Monrovia, California, USA). A DLD device with nominal critical diameter 16 μm, gap size 48 μm and period, N = 20, with three inlets and three outlets was designed for cell sorting experiments. See Fig. 1. The photomask used for microfluidic chip fabrication was purchased from Delta Mask (Enschede, The Netherlands).A 4″ silicon wafer was baked for 5 minutes at 120 °C to dry. A 100 μm thick dry film resist (SUEX®, K100, DJ Microlaminates, Sudbury, Massachusetts, USA) was laminated onto the wafer using a laminating machine (Catena 35, Acco UK Ltd, Buckinghamshire, UK) at 65 °C. A pre-exposure bake at 85 °C on a hotplate (Model 1000-1 Precision Hot Plate, Electronic Micro Systems Ltd, West Middlands, UK) was performed to remove air bubbles and to help the film relax. Exposure at 365 nm was done in a contact mask aligner (Karl Suss MJB4 soft UV, Munich, Germany) for 34 s at a lamp power of 30 mW cm−2. Post exposure baking was done at 85 °C for 5 minutes. Development was done in Mr DEV 600 (Micro Resist Technology GmbH, Berlin, Germany) for 15 minutes plus 5 minutes in fresh developer followed by a rinse in flowing isopropanol and drying with nitrogen. A final bake was done in a convection oven at 200 °C for 2 hours. To reduce adhesion of PDMS to the master a layer of aluminium oxide (1 nm) followed by a monolayer of perfluordecyltriklorosilan (FDTS) was deposited in an ALD system (Fiji – Plasma Enhanced ALD, Veeco, NY, USA).
The microfluidic devices were fabricated through replica moulding using a silicon-based organic polymer, polydimethylsiloxane (PDMS), (Sylgard 184, Dow Corning, Midland, MI, USA). The monomer and curing agents were mixed in a ratio of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. The mixture was then degassed in a vacuum desiccator for 20 minutes to remove air bubbles. Subsequently, the PDMS mixture was carefully poured over the device molds. To solidify the polymer it was cured at 80 °C for 60 minutes. The resulting structured PDMS pieces were peeled from the molds, cut to shape, punched to create fluidic access and bonded to a glass slide via air plasma treatment (Zepto, Diener electronic GmbH & Co. KG, Ebhausen, Germany), glass slides (60 s) and PDMS (15 s). Each of the inlets in the PDMS device were filled with filtered MACS buffer. Silicon tubes were attached to the PDMS device.
1. The mixture was then degassed in a vacuum desiccator for 20 minutes to remove air bubbles. Subsequently, the PDMS mixture was carefully poured over the device molds. To solidify the polymer it was cured at 80 °C for 60 minutes. The resulting structured PDMS pieces were peeled from the molds, cut to shape, punched to create fluidic access and bonded to a glass slide via air plasma treatment (Zepto, Diener electronic GmbH & Co. KG, Ebhausen, Germany), glass slides (60 s) and PDMS (15 s). Each of the inlets in the PDMS device were filled with filtered MACS buffer. Silicon tubes were attached to the PDMS device.
2.3 Cell culture
MDA-MB-231 breast cancer cells (obtained from the American Type Culture Collection, ATCC, USA) were used for all the experiments presented in Fig. 2, 3 and 5. For the migration studies shown in Fig. 4, we used MDA-MB-231-Tomato-LUC cells, generated by stable transfection of the parental MDA-MB-231 line with a tdTomato-luciferase reporter construct. Both cell lines were cultured in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% FBS (Sigma-Aldrich) and 1% pen-strep (Sigma-Aldrich). Cells were cultured in an incubator at 37 °C, 5% CO2 with cell culture medium refreshed every 2–3 days, and the cells were passaged every four to six days. For experiments involving cell area quantification (day 1 and day 7), culture medium was replaced every three days to maintain appropriate nutrient levels and optimal growth conditions. For 2D migration assays, cells were imaged continuously over the defined experimental period without medium exchange to avoid mechanical disturbance that could interfere with trajectory analysis. For spheroid formation and subsequent Matrigel embedding experiments, medium was not exchanged during the defined experimental window to prevent disruption of spheroid integrity and matrix structure. For passaging, cells were trypsinized at 37 °C for 5 min and then collected after centrifugation at 700g for 5 min. For sorting experiments, MDA-MB-231 breast cancer cells were prepared in 10% Optiprep (to obtain neutral buoyancy), 4 mM EDTA and MACS buffer (Table S1).2.4 Sorting related imaging and image analysis
In the first part of the study, size distributions of the beads and cell subgroups (Fig. 1 and 2) were observed before and after sorting. Images of beads and cells plated on 35 mm petri dishes were acquired using an inverted epifluorescence microscope (Nikon Eclipse Ti, Nikon Corporation, Tokyo, Japan) equipped with a Plan Fluor ELWD 20×/0.45 NA objective (working distance: 7.4 mm) and a scientific Andor iXon EMCCD camera (Andor Technology, UK). Brightfield imaging was used to capture high-contrast images of cells and particles.Automated scans consisting of multiple adjacent fields of view (8 × 8) were captured for unsorted beads/cells and those collected from each outlet (small and large) across all sorting experiments. Images were saved in TIFF format and subsequently processed using a custom Python-based analysis pipeline. Automated image processing included grayscale normalization, contrast enhancement (CLAHE), and morphological segmentation to isolate single-cell regions. Individual objects were quantified using the regionprops module from scikit-image, extracting parameters such as area, perimeter, equivalent diameter, and centroid position. The segmentation analysis identified all objects within the field of view. Subsequently, objects were filtered based on circularity to exclude cell clusters and debris, ensuring that only well-defined cells were included in the analysis. A total of 418 unsorted cells were analyzed. Sorting experiments were performed in triplicate. For the small outlet, 314, 262, and 231 cells were analyzed in replicates 1–3, respectively. For the intermediate fraction, 598, 419, and 567 cells were analyzed, and for the large outlet, 476, 145, and 412 cells were analyzed across the three replicates. The resulting per-cell measurements were compiled into CSV datasets, which were later used for statistical analysis and visualization (e.g., plots shown in Fig. 2). Full details of the image segmentation workflow, parameter settings, and the full dataset of images together with the code and the csv dataset can be found in the SI (Fig. S2).
We define a critical diameter of the DLD device based on the experimental outcome of the sorting, as the size of a particle for which the probabilities to end up in the small and the large outlets are equal. We refer to these probabilities as the routing probabilities.36 We can formulate the problem using Bayes' rule with the routing probability of a particle of size di to end up in outlet k ∈ {small, intermediate, large} to be given by the conditional probability P(k|di). The critical diameter is thus defined as the size for which the routing probabilities for the small and the large outlets overlap, see eqn (1) and section 4 in SI. The approach was used to determine experimental DC for both microspheres (Fig. 1C) and cells (Fig. 2C). In this process a smooth behaviour of the routing probabilities close to DC due to large number of particles is important, while any noise for very small or very large particles is of less concern.
| P(k = small|di = DC) = P(k = large|di = DC) | (1) |
2.5 Cell-based assays
After sorting the cells, the small and the large cell fractions were studied phenotypically in three different assays as described below.For confocal microscopy, cells were seeded on glass-bottom 96-well plates (Mattek, USA) at a concentration of 2000 cells per well. Cells were then fixed in 4% paraformaldehyde, permeabilized in 0.2% Triton X-100, and blocked with 3% BSA prior to adding a mix of DAPI (Sigma, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400) and Phalloidin-647 (Sigma, 1
400) and Phalloidin-647 (Sigma, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400) for 2 hours. Samples were then washed three times with PBS and imaged. All images were acquired using a Nikon A1R HD confocal microscope (Nikon Corporation, Tokyo, Japan) equipped with Nikon Plan Apo λ 20×/0.75 NA objective and Nikon NIS-Elements software. Images were acquired using sequential scanning to minimize spectral overlap between fluorophores, with excitation lasers at 405 nm (DAPI), and 640 nm (Phalloidin-647). Fluorescence images were analyzed using ImageJ (National Institutes of Health, Bethesda, MD, USA).37
400) for 2 hours. Samples were then washed three times with PBS and imaged. All images were acquired using a Nikon A1R HD confocal microscope (Nikon Corporation, Tokyo, Japan) equipped with Nikon Plan Apo λ 20×/0.75 NA objective and Nikon NIS-Elements software. Images were acquired using sequential scanning to minimize spectral overlap between fluorophores, with excitation lasers at 405 nm (DAPI), and 640 nm (Phalloidin-647). Fluorescence images were analyzed using ImageJ (National Institutes of Health, Bethesda, MD, USA).37
It is important to note that the measurements presented in Fig. 2 were performed on cells in suspension. In contrast, the analyses shown in Fig. 3 were conducted after cell attachment under adherent culture conditions, where cells spread and adopt non-spherical morphologies. Thus, the cell area measurements at day 1 and day 7 represent projected adherent cell area. To ensure accurate quantification of cell area during dynamic spreading, the segmentation pipeline was designed without assumptions of circular cell geometry. Cell boundaries were identified using intensity-based thresholding and morphological processing, allowing detection of both compact, rounded cells and irregular, highly spread morphologies. Cell area was computed based on the total segmented pixel region for each cell. Representative examples demonstrating accurate boundary detection for both circular and non-circular cells are provided in Fig. S5.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 in cold serum-free medium, fibronectin (human, Merck) was prepared at a concentration of 10 μg mL−1 in PBS, and collagen I was prepared at 50 μg mL−1 in 0.02 N acetic acid. 50 μL of each coating solution was added to designated wells of the pre-chilled plates and incubated at 37 °C for one hour. Following incubation, the wells were gently washed with PBS to remove any unbound ECM components and then equilibrated with pre-warmed complete culture medium. After coating, unsorted cells and the sorted subpopulations were seeded into the prepared wells at a density of 2000 cells per well in 200 μL of complete medium. Cells were allowed to attach for 2 hours under standard culture conditions (37 °C, 5% CO2).
20 in cold serum-free medium, fibronectin (human, Merck) was prepared at a concentration of 10 μg mL−1 in PBS, and collagen I was prepared at 50 μg mL−1 in 0.02 N acetic acid. 50 μL of each coating solution was added to designated wells of the pre-chilled plates and incubated at 37 °C for one hour. Following incubation, the wells were gently washed with PBS to remove any unbound ECM components and then equilibrated with pre-warmed complete culture medium. After coating, unsorted cells and the sorted subpopulations were seeded into the prepared wells at a density of 2000 cells per well in 200 μL of complete medium. Cells were allowed to attach for 2 hours under standard culture conditions (37 °C, 5% CO2).Sorted MDA-MB-231-Tomato-LUC cells were labeled with the nuclear marker SiR-DNA kit (Spirochrome, Cat. No. CY-SC007). Images were acquired using an inverted Nikon Ti2-E wide-field fluorescence microscope with a Nikon Plan Apo λ 20×, 0.75 numerical aperture (NA) objective lens and perfect focus system (PFS) for maintenance of focus over time. Excitation and emission light were passed through TRITC (Exc. 543–566 nm, Em. 582–636 nm) and Cy5 (Exc. 590–645 nm, Em. 659–736 nm) filter cubes, all from Semrock. Samples were kept in a humidified atmosphere at (37 °C, 5% CO2) using an environmental chamber (Okolab). Images were acquired on a Nikon DS-Qi2 CMOS camera. The imaging of the samples was automated by generating stage positions covering the sample area using JOBS (NIS-Elements extension; Nikon) and a Nikon TI-S-ER motorized stage with an encoder. For time-lapse imaging, images were collected every 20 min for 20 hours. Single-cell migration analysis was performed using CellProfiler,38 with single-cell tracking specifically performed using the TrackObjects module. Cell trajectories were automatically generated using the established tracking algorithm without manual intervention. For each experimental condition, more than 20 high-quality cell trajectories were analyzed per biological replicate, with three independent replicates performed. To ensure reliable migration measurements, trajectories were filtered based on predefined criteria. Cells that remained non-motile throughout the imaging period, appeared only briefly in the field of view, were located at image boundaries, or corresponded to newly divided cells were excluded from analysis. These filtering steps were applied uniformly across all conditions to avoid analysis bias.
Trajectories were selected to represent cells whose migration behavior was close to the group average for each respective condition. Specifically, three representative cells per group and ECM condition were chosen from the full dataset based on their migration velocity being close to the population average enabling qualitative comparison (Fig. 4).
On day 7 when spheroids reached a uniform size, they were collected and gently transferred into pre-chilled 24-well plates containing a base layer of Matrigel® (Corning, Cat. No. 354234). Each spheroid was suspended in 25 μL of chilled growth factor-reduced Matrigel, placed centrally in the well, and allowed to solidify at 37 °C for 30–45 minutes. After gelation, 500 μL of complete medium was added on top of each well. Spheroid cultures were maintained for up to day 9. Brightfield images of spheroid formation and growth were acquired using ZOE Cell Imager (Bio-Rad, USA) equipped with a 10× objective lens. Embedded spheroids were imaged daily. Morphological changes (e.g., invasion, sprouting) were assessed and measured using ImageJ.37 The average spheroid size was determined based on the largest cross-sectional area (n = 15 spheroids per group). For each spheroid, two distinct thresholds were manually applied in ImageJ to delineate the spheroid core and the surrounding spread area.
3 Results and discussion
3.1 Performance characterization
To validate device performance prior to biological experiments, we introduced polystyrene calibration microspheres (10 μm CV (coefficient of variation) 13%, 16 μm CV 12%, 20 μm CV 9% and 27 μm CV 10%) through the sample inlet at 80 mbar using a pressure controller (MFCS-4C, Fluigent, Paris, France) (Fig. 1A) and analyzed the resulting composition of microspheres in each outlet. The device was designed with a critical diameter (DC ≈ 16 μm). To obtain a more rigorous estimate of the critical size based on actual separation results and from which purities and yields can be calculated, we used the Bayes' rule approach36 described above and in section 4 in the SI. Our results shed light on dispersive trajectories and show how designing devices to accommodate these improves separations. We observe that the distribution of routing probabilities P(intermediate) is broad, meaning that no particular subpopulation of particles is completely lost to the waste/intermediate outlet. However, the distribution does have a peak meaning that there is a size range of cells that are more likely to be lost. Since the intersection P(small) = P(large) is derived from measurements of the sizes of particles in the two outlets this point constitutes a good empirical definition of DC in our device. Because P(intermediate) peaks near this crossover we conclude that dispersive or mixed mode trajectories are most probable for particles close to the transition between zigzaging and displacement motion and that preferentially removing these particles increases the size separation between the other two fractions. Applying this method to our bead data (excluding clustered particles) yields an estimated critical size of DC = 19.4 μm (Fig. 1B).It is also possible to use routing probability analysis to estimate purity and yield. We define the purity in the small outlet as the proportion of particles smaller than DC found there, and in the large outlet as the proportion of particles larger than DC. We do this for raw, un-smoothed bin counts. For bead separations we reach 90% and 98% purities, respectively. Yields are defined as number of large particles in the large outlet divided by the total number of large particles, and correspondingly for the small particles. The yields are estimated by counting the particles in each outlet and comparing to the inlet population, giving us 58% and 28% for the large and small outlets, respectively. The remainder of the cells are routed to the intermediate outlet. Note: these values for yield are only valid under the assumption that we are sampling the cells from each outlet equally.
The DLD chip was assembled in a modular format to enhance reproducibility, simplify handling, and ensure compatibility with downstream cell culture assays. Together, these results confirm that the developed platform enables reliable size-based sorting, establishing its suitability for subsequent biological investigations.
3.2 Sorting of metastatic breast cancer cells into subpopulations
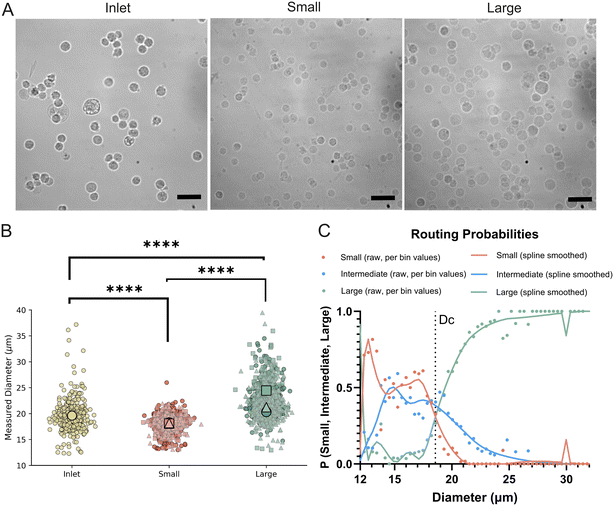
After establishing the robustness and performance of the DLD device, we next aimed to sort MDA-MB-231 cell line, a widely studied model of triple-negative breast cancer. These cells are characterized by high invasiveness, mesenchymal-like morphology, and substantial phenotypic heterogeneity, making them an ideal system to investigate size-based variation and have been extensively used to study migration and invasion in vitro,39–41 metastatic dissemination in vivo,42–44 and cellular mechanics such as deformability and stiffness,45–47 However, these studies typically report measurements as averages across the entire population, potentially masking subpopulation-level differences. By fractionating cells according to size, we sought to uncover whether distinct subsets within this commonly used model exhibit divergent behaviors that are otherwise obscured in bulk analyses.At an inlet pressure of 80 mbar, we measured cell diameters before and after sorting Fig. 2. The unsorted cell population displayed a mean diameter of 19.61 ± 2.80 μm, indicating a broad size distribution of MDA-MB-231 cells. Following sorting, the small-cell outlet yielded a mean diameter of 18.18 ± 1.55 μm, whereas the large-cell outlet was significantly enriched for bigger cells, with a mean diameter of 22.20 ± 3.10 μm, Fig. 2B.
Cell size was quantified from brightfield images one hour after seeding, ensuring that cells had adhered to the substrate but had undergone minimal spreading-induced changes in morphology and no cell division. This time point allowed us to capture the size characteristics of each subpopulation immediately following sorting, serving as a baseline for subsequent phenotypic comparisons.
As shown in the superplots of measured cell diameters, a clear and statistically significant difference was observed between the small and large cell subpopulations. The routing probability analysis further illustrates this size-dependent sorting behavior, with the intermediate outlet capturing a broad distribution that overlaps with both the small and large cells. As with the polystyrene sphere results, this demonstrates how the inclusion of a third, intermediate outlet improves separation results. From the crossover of the routing probability curves, the critical diameter (DC) was determined as 18.4 μm (Fig. 2C). Purity for small/large fractions was 58% and 93% respectively and yields 49% and 59%, under the assumption of equal sampling as described above for beads. Our findings establish that the DLD device robustly sorts breast cancer cells into distinct size-based subpopulations, providing a foundation for subsequent functional assays.
Although the nominal cut-off size DC of the DLD device is predicted to be ≈16 μm based on channel geometry and Davis' empirical equation,48 we find that DC, defined via routing probabilities, is ≈20.3 μm for polystyrene spheres and ≈18.6 μm for MDA-MB-231 cells. This apparent discrepancy arises from two key factors. Firstly, Davis' equation is a best fit model to experimental data across a broad array of DLD devices with non-negligible spread and should only be expected to give approximations of DC. Better approximations of DC require consideration of specific device and sample conditions. Secondly, there are considerable differences between rigid, polystyrene spheres and cells with inherent complexity and variation in deformability, morphology, and other properties that are very likely to influence behaviour during sorting.
Cells are also, with their greater complexity, inherently more difficult to image, segment and analyse than polystyrene spheres. While care has been taken to compensate for imaging errors and to filter the results of image segmentation to remove non-cells and clusters that would skew results (see SI for details), these effects may contribute to differences in observed DC. There is also a small probability that cells change size between the sorting and the imaging used for size analysis, but this time is too short (1 < hour) for cells to grow considerably and increases in apparent size due to interactions with the surface of the collection petri dishes (spreading) are also minimal.
3.3 Assessment of proliferation in sorted breast cancer cell populations
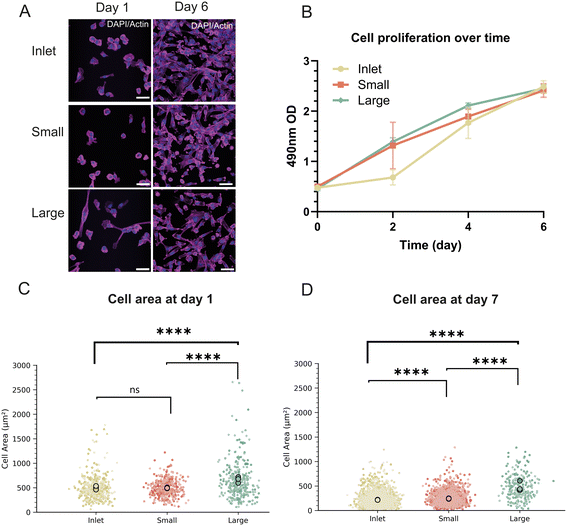
To determine if size-based subpopulations of MDA-MB-231 represents distinct phenotypic subsets, we first compared the proliferative behavior of the unsorted cells, small, and large subpopulations under standard 2D culture conditions. Proliferation was quantified using a CellTiter 96® assay performed on days 0, 2, 4, and 6 post-sorting (Fig. 3A). All three groups maintained robust growth across the six-day period, with surprisingly comparable proliferation rates (Fig. 3B). This shows that size-dependent subpopulations proliferate at the same rate and that exposing the cells to the DLD device does not impair cell viability and proliferative potential.Next, we assessed changes in cell size of the subpopulations after reattachment in culture over 7 days. (Fig. 3C and D). Measurement of cell spread area from F-actin stained images showed that the small-cell fractions consistently exhibited reduced adherent spread area, while large-cell fractions remained significantly larger, mirroring their diameters measured immediately after sorting in suspension. These stable differences persisted throughout the 7 days, demonstrating that DLD-defined size classes persist in both detached and adherent states over extended period of time and cell cycles. Additionally, cell area was compared between day 1 and day 7 for inlet, small, and large cell populations. All three groups exhibited a significant decrease in projected cell area over time (Fig. S6). Statistical analysis revealed highly significant differences between day 1 and day 7 for all subpopulations. These results indicate that cells in all groups reduce their spreading over prolonged adherent culture. Importantly, despite the overall decrease in size, the relative differences between small, inlet, and large populations were maintained.
Taken together, these results show that cell size is an inherent property of cells that persists over time and that these subpopulations proliferate at the same rate over 7 days.
3.4 ECM sensing and migration behavior of size-sorted breast cancer cells
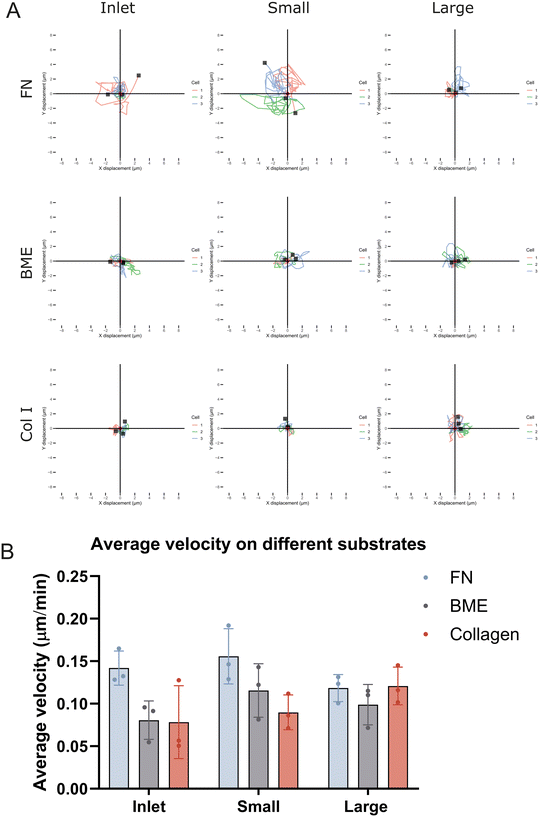
Since metastasis involves both proliferation as well as acquisition of a migratory phenotype which is strongly dictated by properties of the ECM, we next asked if size-sorted MDA-MB-231 cells differ in cell migration as well as sensitivity to different ECM cues during migration. Unsorted cells and cells from the small, and large subpopulations were thus seeded in wells coated with fibronectin (FN), basement membrane extract (BME), and collagen I (Col I), and imaged by live cell microscopy for 20 hours to measure morphology, and cell migration.In addition to migration trajectories, cell area, nuclei size, and total accumulated distance were quantified for all sorted subpopulations plus unsorted cells (inlet) across the different ECM substrates (FN, BME, and Col I) over the 20 hour period. Consistent with size based sorting results, here we found that large cells maintained significantly larger cell area and nuclei size compared to small, and unsorted (inlet) populations across all ECM substrates (Fig. S9A and B). Measurement of accumulated distance and average velocity however showed that overall small cells showed slightly higher migration velocities and accumulated distance on fibronectin and BME coated substrates while large cells were slightly faster on Col I (Fig. 4A, B and S9C). Most interestingly, we found that only the small cells show any sensitivity to changes in the ECM. The velocity of small cells was the highest on fibronectin followed by reduced velocities on BME and the slowest on Col I. In contrast, large cells showed no sensitivity across these three ECM coatings (Fig. 4B).
These observations are consistent with previous studies that highlight the importance of cell morphology in predicting metastatic behaviors. Notably, cell morphology, including size, aspect ratio, and roundness, has been shown to correlate with both in vitro migratory behavior and in vivo metastatic potential in TNBC models: smaller, rounder cell populations such as MDA-MB-231, BT549, and MDA-MB-468 exhibited significantly higher metastatic capability in murine models.8,49–51 In another study, it has been showed that a fast-moving subpopulation of MDA-MB-231 exhibited impaired tumor growth compared with the parental population, despite no significant differences in proliferation (Ki67) and decreased apoptosis in resected mammary tumors.8 Additionally, a recent study showed that analysis of subpopulations present at the invasive front suggests that a subpopulation of small deformable cells at the invasive fronts represent the most invasive subpopulation and may correspond to a subpopulation of CSCs, highlighting interplay between mechanical properties and size to regulate invasiveness.52
In our system small cells are more motile on the ECM protein fibronectin consistent with several previous studies that have highlighted how fibronectin-mediated adhesion and ECM-specific interactions influence both cell morphology and migratory behavior.53–55 This likely suggests different focal adhesion composition, dynamics and or signaling in the smaller subpopulation compared to large cells, which may lead to altered activation of mechanotransduction pathways that support faster migration. Taken together, these results underscore that cell size is a critical determinant of motility and potential predictors of clonal variability and metastatic potential.
3.5 3D invasion of size-sorted breast cancer cell populations
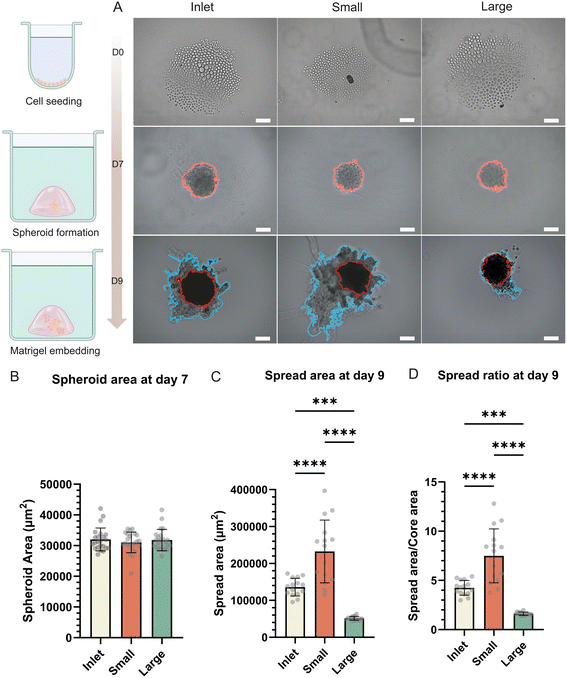
Since tumor invasion in vivo occurs within complex 3D environments rather than on 2D substrates, we next asked whether the functional differences between size-sorted subpopulations persist in complex 3D environments. To do so, we first generated spheroids from unsorted cells, and from small and large sorted MDA-MB-231 populations by culturing cells in non-adherent conditions for seven days (Fig. 5A). At this stage, spheroids from all groups were viable, compact, and comparable in size (mean areas: unsorted 0.032 ± 0.003 mm2, small 0.031 ± 0.003 mm2, large 0.031 ± 0.003 mm2; n = 23, p = n.s.), ensuring that initial spheroid area did not affect subsequent invasion analyses (Fig. 5B).To assess invasion, the spheroids were then embedded in Matrigel, a 3D basement membrane mimic and observed over the following days. Imaging of embedded spheroids revealed that spheroids derived from small cells exhibited significant invasive outgrowths, with irregular projections extending from the spheroid core. In contrast, large-cell spheroids largely retained compact morphology with well-defined boundaries, while spheroids based on unsorted cells showed intermediate behavior. Quantification of spheroid spread area over time confirmed that small-cell spheroids invaded significantly more than unsorted-cell and large-cell spheroids (day 9 mean areas: unsorted 0.136 ± 0.024 mm2, small 0.232 ± 0.084 mm2, large 0.051 ± 0.005 mm2) (Fig. 5C). To further characterize these differences, we calculated a spheroid spread ratio, defined as the projected spread area of the spheroid to its core area, which was 4.25 for the unsorted, 7.49 for the small, and 1.62 for the large (Fig. 5D). These results indicate that the small-cell subpopulation exhibited the highest spreading capacity, whereas spheroids generated from the large-cell fraction remained more compact, suggesting distinct adhesive and cytoskeletal behaviors between the size-sorted groups.
These findings demonstrate that, consistent with their enhanced 2D migration, small-cell subpopulations within MDA-MB-231 cell lines drive markedly greater invasion in 3D, whereas large-cell subpopulations remain restricted, highlighting stable, size-linked heterogeneity to invasive potential.
This approach, integrating microfluidic size-based sorting with 3D spheroid culture, provides a robust platform to investigate phenotypic differences in tumor cell invasiveness and growth dynamics within a single population of cells. Previous studies have demonstrated that 3D spheroid invasion assays better recapitulate tumor microenvironmental cues and invasion mechanisms than traditional 2D cultures.56–59 Moreover, size-dependent invasive behavior observed here is consistent with prior reports indicating that smaller breast cancer cells often possess increased motility, deformability, and stem-like characteristics that facilitate invasion and metastasis in primary tumors.60 The compact structure and limited spreading of large-cell spheroids may reflect a more differentiated, less invasive phenotype, consistent with tumor heterogeneity models emphasizing subpopulation specialization.61 Taken together, these results underscore the utility of combining microfluidic sorting and 3D spheroid invasion assays to dissect functional heterogeneity in breast cancer, providing insights relevant for therapeutic targeting of metastatic subpopulations.
4 Conclusions
Using deterministic lateral displacement (DLD), we successfully sorted metastatic breast cancer cells (MDA-MB-231) into two distinct subpopulations, small and large, based on their size. We show that dispersive trajectories are common close to the cutoff size (DC) in DLD devices of this kind when sorting both polystyrene spheres and cells and that this can be compensated for by careful design of device outlets to collect and remove errant cells from down stream analyses. This label-free, passive sorting approach enabled us to investigate functional heterogeneity among subpopulations within a single cell population. Comprehensive post-sorting analyses revealed that these morphologically distinct subpopulations also differ functionally. The small cell population within a single MDA-MB-231 population demonstrated higher motility and invasiveness, particularly in fibronectin-rich and 3D environments compared to large cells. This suggests to us that the aggressive phenotype observed in MDA-MB-231 cells is dominated by the small cell population, and this sorting enriches for this phenotype. In addition, size-based differences in morphology, adhesion dynamics, and invasion profiles suggest underlying mechanobiological heterogeneity within the metastatic population.Methodologically, this study establishes a microfluidics-based workflow for enriching functionally distinct cancer cell subsets and provides a foundation for further exploration of size, morphology and, because DLD has been shown to be sensitive to deformability,30,31,62 subsequently mechanobiology-associated traits in tumor progression. The approach not only provides a window into the relationship between cell physical state and metastatic behavior, but also offers a scalable, reproducible tool for downstream biochemical or biophysical assays.
While this study focuses on phenotypic characterization, further molecular investigation particularly of ECM-sensing and mechanotransduction pathways such as Hippo/YAP–TAZ signaling and focal adhesion dynamics would provide deeper mechanistic insight into the observed size-dependent heterogeneity and represents an important direction for future investigation. Future work will focus on quantifying deformability using DLD under varying flow regimes and extending this platform to circulating tumor cells (CTCs) from patient-derived samples. Applying this principle to CTCs may offer deeper insight into the mechanobiology of metastasis and facilitate the development of personalized diagnostics or therapeutic strategies targeting highly motile and invasive subpopulations.
Author contributions
Conceptualization, J. T., V. S., J. B.; methodology, J. T., V. S., J. B.; software, E. Y., Z. F. and J. B.; validation, E. Y., Z. F. and J. B.; formal analysis, E. Y., Z. F., J. B.; investigation, E. Y., Z. F., J. B.; resources, J. T; data curation, E. Y., Z. F. and J. B.; writing – original draft preparation, E. Y.; writing – review and editing, E. Y., Z. F., J. B., V. S., J. T; visualization, E. Y, Z. F. and J. B.; supervision, J. T., V. S., J. B.; project administration, J. T.; funding acquisition, J. T., V. S. All authors have read and agreed to the published version of the manuscript.Conflicts of interest
The authors declare no conflict of interest. The funders had no role in the design of the study, in the collection, analyses, or interpretation of data, in the writing of the manuscript, or in the decision to publish the results.Data availability
Data for this article, including spread sheets, videos and images are available at Harvard Dataverse. Fig. 1 raw data: https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/MI5RE8 Analyzed data: https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/MKC4QQ. Fig. 2 and S2–S4, Table 1 cell sorting data raw data: https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/NJZZKL and https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/9M5SGA. Analyzed data: https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/6BIKL2. Videos of cell sorting: raw data: https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/T1VNMP. Fig. 5 raw data: https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/JMYAVH.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5lc01042j.
Acknowledgements
This research was funded by the Swedish Research Council, grant number 2019-02355 and NanoLund, grant numbers p20-2019, staff01-2020, s01-2024. VS is also supported by the KAW foundation via the WCMM Lund program. All device processing was conducted within Lund Nano Lab. The European Union's Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement No 945378.References
- H. Maleki, A. R. Bahrami and M. M. Matin, Cancer cell cycle heterogeneity as a critical determinant of therapeutic resistance, Genes Dis., 2024, 11(1), 189–204, DOI:10.1016/j.gendis.2022.11.025.
- C. E. Meacham and S. J. Morrison, Tumour heterogeneity and cancer cell plasticity, Nature, 2013, 501(7467), 328–337, DOI:10.1038/nature12624.
- M. Tellez-Gabriel, B. Ory, F. Lamoureux, M.-F. Heymann and D. Heymann, Tumour heterogeneity: The key advantages of single-cell analysis, Int. J. Mol. Sci., 2016, 17(12), 2142, DOI:10.3390/ijms17122142.
- E. Melo, F. De Sousa, X. Wang, M. Jansen, E. Fessler, A. Trinh, L. P. de Rooij, J. H. de Jong, O. J. de Boer, R. van Leersum, M. F. Bijlsma, H. Rodermond, M. van der Heijden, C. J. van Noesel, J. B. Tuynman, E. Dekker, F. Markowetz, J. P. Medema and L. Vermeulen, Poor-prognosis colon cancer is defined by a molecularly distinct subtype and develops from serrated precursor lesions, Nat. Med., 2013, 19(5), 614–618, DOI:10.1038/nm.3174.
- R. Fisher, L. Pusztai and C. Swanton, Cancer heterogeneity: implications for targeted therapeutics, Br. J. Cancer, 2013, 108(3), 479–485, DOI:10.1038/bjc.2012.581.
- A. Haghofer, E. Parlak, A. Bartel, T. A. Donovan, C. A. Assenmacher, P. Bolfa, M. J. Dark, A. FuchsBaumgartinger, A. Klang, K. Jäger, R. Klopfleisch, S. Merz, B. Richter, F. Y. Schulman, H. Janout, J. Ganz, J. Scharinger, M. Aubreville, S. M. Winkler, M. Kiupel and C. A. Bertram, Nuclear pleomorphism in canine cutaneous mast cell tumors: Comparison of reproducibility and prognostic relevance between estimates, manual morphometry, and algorithmic morphometry, Vet. Pathol., 2025, 62(2), 161–177, DOI:10.1177/03009858241295399.
- I. Singh and T. P. Lele, Nuclear morphological abnormalities in cancer: A search for unifying mechanisms, Results Probl. Cell Differ., 2022, 70, 443–467, DOI:10.1007/978-3-031-06573-6_16.
- S. J. Conner, J. R. Guarin, T. T. Le, J. P. Fatherree, C. Kelley, S. L. Payne, S. R. Parker, H. Bloomer, C. Zhang, K. Salhany, R. A. McGinn, E. Henrich, A. Yui, D. Srinivasan, H. Borges and M. J. Oudin, Cell morphology best predicts tumorigenicity and metastasis in vivo across multiple tnbc cell lines of different metastatic potential, Breast Cancer Res., 2024, 26(1), 43, DOI:10.1186/s13058-024-01796-8.
- S. P. Ramanathan, M. Krajnc and M. C. Gibson, Cell-size pleomorphism drives aberrant clone dispersal in proliferating epithelia, Dev. Cell, 2019, 51(1), 49–61.e4, DOI:10.1016/j.devcel.2019.08.005.
- P.-H. Wu, M. J. Phillip, S. B. Khatau, W.-C. Chen, J. Stirman, S. Rosseel, K. Tschudi, J. Van Patten, M. Wong, S. Gupta, A. S. Baras, J. T. Leek, A. Maitra and D. Wirtz, Evolution of cellular morpho-phenotypes in cancer metastasis, Sci. Rep., 2015, 5(1), 18437, DOI:10.1038/srep18437.
- A. Krohn, T. Ahrens, A. Yalcin, T. Plönes, J. Wehrle, S. Taromi, S. Wollner, M. Follo, T. Brabletz, S. A. Mani, R. Claus, B. Hackanson and M. Burger, Tumor cell heterogeneity in small cell lung cancer (SCLC): phenotypical and functional differences associated with epithelial-mesenchymal transition (EMT) and DNA methylation changes, PLoS One, 2014, 9(6), e100249, DOI:10.1371/journal.pone.0100249.
- P. J. Keller, A. F. Lin, L. M. Arendt, I. Klebba, A. D. Jones, J. A. Rudnick, T. A. DiMeo, H. Gilmore, D. M. Jefferson, R. A. Graham, S. P. Naber, S. Schnitt and C. Kuperwasser, Mapping the cellular and molecular heterogeneity of normal and malignant breast tissues and cultured cell lines, Breast Cancer Res., 2010, 12(5), R87, DOI:10.1186/bcr2755.
- A. Gough, A. M. Stern, J. Maier, T. Lezon, T. Y. Shun, C. Chennubhotla, M. E. Schurdak, S. A. Haney and D. L. Taylor, Biologically relevant heterogeneity: Metrics and practical insights, SLAS Discovery, 2017, 22(3), 213–237, DOI:10.1177/2472555216682725.
- M. Proietto, M. Crippa, C. Damiani, V. Pasquale, E. Sacco, M. Vanoni and M. Gilardi, Tumor heterogeneity: preclinical models, emerging technologies, and future applications, Front. Oncol., 2023, 13, 1164535, DOI:10.3389/fonc.2023.1164535.
- D. J. Adams, U. Kim and H. T. Soh, Multitarget magnetic activated cell sorter, Proc. Natl. Acad. Sci. U. S. A., 2008, 105(47), 18165–18170, DOI:10.1073/pnas.0809795105.
- W. J. Tung, K. Heydari, R. Tirouvanziam, B. Sahaf, D. R. Parks, L. A. Herzenberg and L. A. Herzenberg, Modern flow cytometry: A practical approach, Clin. Lab. Med., 2007, 27(3), 453–468, DOI:10.1016/j.cll.2007.05.001.
- C. W. Shields IV, C. D. Reyes and G. P. López, Microfluidic cell sorting: a review of the advances in the separation of cells from debulking to rare cell isolation, Lab Chip, 2015, 15(5), 1230–1249, 10.1039/C4LC01246A.
- Y. Zhang, T. Zheng, L. Wang, L. Feng, M. Wang, Z. Zhang and H. Feng, From passive to active sorting in microfluidics: A review, Rev. Adv. Mater. Sci., 2021, 60(1), 313–324, DOI:10.1515/rams-2020-0044.
- J. Guck, S. Schinkinger, B. Lincoln, F. Wottawah, S. Ebert, M. Romeyke, D. Lenz, H. M. Erickson, R. Ananthakrishnan, D. Mitchell, J. Käs, S. Ulvick and C. Bilby, Optical deformability as an inherent cell marker for testing malignant transformation and metastatic competence, Biophys. J., 2005, 88(5), 3689–3698, DOI:10.1529/biophysj.104.045476.
- D. Di Carlo, Inertial microfluidics, Lab Chip, 2009, 9(21), 3038–3046, 10.1039/B912547G.
- E. U. Anand, C. Magnusson, A. Lenshof, Y. Ceder, H. Lilja and T. Laurell, Two-step acoustophoresis separation of live tumor cells from whole blood, Anal. Chem., 2021, 93(51), 17076–17085, DOI:10.1021/acs.analchem.1c04050.
- V. Varmazyari, H. Habibiyan, H. Ghafoorifard, M. Ebrahimi and S. Ghafouri-Fard, A dielectrophoresis-based microfluidic system having double-sided optimized 3D electrodes for labelfree cancer cell separation with preserving cell viability, Sci. Rep., 2022, 12(1), 12100, DOI:10.1038/s41598-022-16286-0.
- K. Xu, X.-L. Jiao, C.-Y. Chen, P.-P. Wang and C.-F. Chen, Isolation of circulating tumor cells based on magnetophoresis, Chin. J. Anal. Chem., 2022, 50(3), 100058, DOI:10.1016/j.cjac.2022.100058.
- L. Luo and Y. He, Magnetically driven microfluidics for isolation of circulating tumor cells, Cancer Med., 2020, 9(12), 4207–4231, DOI:10.1002/cam4.3077.
- L. R. Huang, E. C. Cox, R. H. Austin and J. C. Sturm, Continuous particle separation through deterministic lateral displacement, Science, 2004, 304(5673), 987–990, DOI:10.1126/science.1094567.
- M. Yamada and M. Seki, Hydrodynamic filtration for on-chip particle concentration and classification utilizing microfluidics, Lab Chip, 2005, 5(11), 1233–1239, 10.1039/B509386D.
- O. Otto, P. Rosendahl, A. Mietke, S. Golfier, C. Herold, D. Klaue, S. Girardo, S. Pagliara, A. Ekpenyong, A. Jacobi, M. Wobus, N. T. öpfner, U. F. Keyser, J. Mansfeld, E. Fischer-Friedrich and J. Guck, Real-time deformability cytometry: on-the-fly cell mechanical phenotyping, Nat. Methods, 2015, 12(3), 199–202, DOI:10.1038/nmeth.3281.
- A. Hochstetter, R. Vernekar, R. H. Austin, H. Becker, J. P. Beech, D. A. Fedosov, G. Gompper, S.-C. Kim, J. T. Smith, G. Stolovitzky, J. O. Tegenfeldt, B. H. Wunsch, K. K. Zeming, T. Krüger and D. W. Inglis, Deterministic lateral displacement: Challenges and perspectives, ACS Nano, 2020, 14(9), 10784–10795, DOI:10.1021/acsnano.0c05186.
- J. McGrath, M. Jimenez and H. Bridle, Deterministic lateral displacement for particle separation: a review, Lab Chip, 2014, 14(21), 4139–4158, 10.1039/C4LC00939H.
- J. P. Beech, S. H. Holm, K. Adolfsson and J. O. Tegenfeldt, Sorting cells by size, shape and deformability, Lab Chip, 2012, 12(6), 1048–1051, 10.1039/C2LC21083E.
- M. Xavier, S. H. Holm, J. P. Beech, D. Spencer, J. O. Tegenfeldt, R. O. C. Oreffo and H. Morgan, Label-free enrichment of primary human skeletal progenitor cells using deterministic lateral displacement, Lab Chip, 2019, 19(3), 513–523, 10.1039/C8LC01154K.
- T. Kulrattanarak, R. G. M. van der Sman, C. G. P. H. Schroën and R. M. Boom, Analysis of mixed motion in deterministic ratchets via experiment and particle simulation, Microfluid. Nanofluid., 2011, 10(4), 843–853, DOI:10.1007/s10404-010-0715-z.
- S.-C. Kim, B. H. Wunsch, H. Huan, J. T. Smith, R. H. Austin and G. Stolovitzky, Broken flow symmetry explains the dynamics of small particles in deterministic lateral displacement arrays, Proc. Natl. Acad. Sci. U. S. A., 2017, 114(26), E5034, DOI:10.1073/pnas.1706645114.
- E. Pariset, C. Pudda, F. Boizot, N. Verplanck, J. Berthier, A. Thuaire and V. Agache, Anticipating cutoff diameters in deterministic lateral displacement (DLD) microfluidic devices for an optimized particle separation, Small, 2017, 13(37), 11, DOI:10.1002/smll.201701901.
- R. Vernekar, T. Kruger, K. Loutherback, K. Morton and D. W. Inglis, Anisotropic permeability in deterministic lateral displacement arrays, Lab Chip, 2017, 17(19), 3318–3330, 10.1039/c7lc00785j.
- E. Akbari, E. Yilmaz, C. N. Prinz, J. P. Beech and J. O. Tegenfeldt, Using Bayes' Rule for Analysis of Microfluidic Particle and Cluster Sorting, Micromachines, 2026, 17(4), 396, DOI:10.3390/mi17040396.
- C. A. Schneider, W. S. Rasband and K. W. Eliceiri, Nih image to imagej: 25 years of image analysis, Nat. Methods, 2012, 9(7), 671–675, DOI:10.1038/nmeth.2089.
- C. McQuin, A. Goodman, V. Chernyshev, L. Kamentsky, B. A. Cimini, K. W. Karhohs, M. Doan, L. Ding, S. M. Rafelski, D. Thirstrup, W. Wiegraebe, S. Singh, T. Becker, J. C. Caicedo and A. E. Carpenter, Cellprofiler 3.0: Next-generation image processing for biology, PLoS Biol., 2018, 16(7), e2005970, DOI:10.1371/journal.pbio.2005970.
- N. Peela, F. S. Sam, W. Christenson, D. Truong, A. W. Watson, G. Mouneimne, R. Ros and M. Nikkhah, A three dimensional micropatterned tumor model for breast cancer cell migration studies, Biomaterials, 2016, 81, 72–83, DOI:10.1016/j.biomaterials.2015.11.039.
- X. Zhang, T. Chan and M. Mak, Morphodynamic signatures of mda-mb-231 single cells and cell doublets undergoing invasion in confined microenvironments, Sci. Rep., 2021, 11(1), 6529, DOI:10.1038/s41598-021-85640-5.
- O.-Y. Hong, H.-Y. Jang, Y.-R. Lee, S. H. Jung, H. J. Youn and J.-S. Kim, Inhibition of cell invasion and migration by targeting matrix metalloproteinase-9 expression via sirtuin 6 silencing in human breast cancer cells, Sci. Rep., 2022, 12(1), 12125, DOI:10.1038/s41598-022-16405-x.
- N. Asokan, S. Daetwyler, S. N. Bernas, C. Schmied, S. Vogler, K. Lambert, M. Wobus, M. Wermke, G. Kempermann, J. Huisken, M. Brand and M. Bornhäuser, Long-term in vivo imaging reveals tumor-specific dissemination and captures host tumor interaction in zebrafish xenografts, Sci. Rep., 2020, 10(1), 13254, DOI:10.1038/s41598-020-69956-2.
- M. A. Kane, K. G. Birmingham, B. Yeoman, N. Patel, H. Sperinde, T. G. Molley, P. Beri, J. Tuler, A. Kumar, S. Klein, S. Zare, A. Wallace, P. Katira and A. J. Engler, Adhesion strength of tumor cells predicts metastatic disease in vivo, Cell Rep., 2025, 44(3), 115359, DOI:10.1016/j.celrep.2025.115359.
- S. Javed, S. Soukhtehzari, N. Salmond, N. Fernandes and K. C. Williams, Development of an in vivo system to model breast cancer metastatic organotropism and evaluate treatment response using the chick embryo, iScience, 2023, 26(4), 106305, DOI:10.1016/j.isci.2023.106305.
- L. Shah, A. Latif, K. J. Williams and A. Tirella, Role of stiffness and physico-chemical properties of tumour microenvironment on breast cancer cell stemness, Acta Biomater., 2022, 152, 273–289, DOI:10.1016/j.actbio.2022.08.074.
- Z. Wang, F. Xu, D. Wu, W. Huang, Z. Chu and Y. Lin, Nuclear deformation and stiffness-dependent traction force generation dictate the migration of cells under confinement, ACS Appl. Mater. Interfaces, 2025, 17(16), 23731–23739, DOI:10.1021/acsami.5c03048.
- T. Fischer, A. Hayn and C. T. Mierke, Effect of nuclear stiffness on cell mechanics and migration of human breast cancer cells, Front. Cell Dev. Biol., 2020, 8, 393, DOI:10.3389/fcell.2020.00393.
- D. W. Inglis, J. A. Davis, R. H. Austin and J. C. Sturm, Critical particle size for fractionation by deterministic lateral displacement, Lab Chip, 2006, 6(5), 655–658, 10.1039/B515371A.
- P.-H. Wu, D. M. Gilkes, J. M. Phillip, A. Narkar, T. W.-T. Cheng, J. Marchand, M.-H. Lee, R. Li and D. Wirtz, Single-cell morphology encodes metastatic potential, Sci. Adv., 2020, 6(4), eaaw6938, DOI:10.1126/sciadv.aaw6938.
- S. M. Lyons, E. Alizadeh, J. Mannheimer, K. Schuamberg, J. Castle, B. Schroder, P. Turk, D. Thamm and A. Prasad, Changes in cell shape are correlated with metastatic potential in murine and human osteosarcomas, Biol. Open, 2016, 5(3), 289–299, DOI:10.1242/bio.013409.
- T. Kołodziej, A. Mielnicka, D. Dziob, A. K. Chojnacka, M. Rawski, J. Mazurkiewicz and Z. Rajfur, Morphomigrational description as a new approach connecting cell’s migration with its morphology, Sci. Rep., 2023, 13(1), 11006, DOI:10.1038/s41598-023-35827-9.
- Asadullah, S. Kumar, N. Saxena, M. Sarkar, A. Barai and S. Sen, Combined heterogeneity in cell size and deformability promotes cancer invasiveness, J. Cell Sci., 2021, 134(7), jcs250225, DOI:10.1242/jcs.250225.
- P. Kanchanawong and D. A. Calderwood, Organization, dynamics and mechanoregulation of integrin-mediated cell–ecm adhesions, Nat. Rev. Mol. Cell Biol., 2023, 24(2), 142–161, DOI:10.1038/s41580-022-00531-5.
- D. Missirlis, T. Haraszti, H. Kessler and J. P. Spatz, Fibronectin promotes directional persistence in fibroblast migration through interactions with both its cell-binding and heparin-binding domains, Sci. Rep., 2017, 7(1), 3711, DOI:10.1038/s41598-017-03701-0.
- M. C. Kim, D. M. Neal, R. D. Kamm and H. H. Asada, Dynamic modeling of cell migration and spreading behaviors on fibronectin coated planar substrates and micropatterned geometries, PLoS Comput. Biol., 2013, 9(2), e1002926, DOI:10.1371/journal.pcbi.1002926.
- L. E. Scott, D. B. Mair, J. D. Narang, K. Feleke and C. A. Lemmon, Fibronectin fibrillogenesis facilitates mechano-dependent cell spreading, force generation, and nuclear size in human embryonic fibroblasts, Integr. Biol., 2015, 7(11), 1454–1465, 10.1039/c5ib00217f.
- B. Pinto, A. C. Henriques, P. M. A. Silva and H. Bousbaa, Three-dimensional spheroids as in vitro preclinical models for cancer research, Pharmaceutics, 2020, 12(12), 1186, DOI:10.3390/pharmaceutics12121186.
- M. A. G. Barbosa, C. P. R. Xavier, R. F. Pereira, V. Petrikaitė and M. Helena Vasconcelos, 3D cell culture models as recapitulators of the tumor microenvironment for the screening of anti-cancer drugs, Cancers, 2022, 14(1), 190, DOI:10.3390/cancers14010190.
- J. Friedrich, C. Seidel, R. Ebner and L. A. Kunz-Schughart, Spheroid-based drug screen: considerations and practical approach, Nat. Protoc., 2009, 4(3), 309–324, DOI:10.1038/nprot.2008.226.
- P. A. Kenny, G. Y. Lee, C. A. Myers, R. M. Neve, J. R. Semeiks, P. T. Spellman, K. Lorenz, E. H. Lee, M. H. Barcellos-Hoff, O. W. Petersen, J. W. Gray and M. J. Bissell, The morphologies of breast cancer cell lines in three-dimensional assays correlate with their profiles of gene expression, Mol. Oncol., 2007, 1(1), 84–96, DOI:10.1016/j.molonc.2007.02.004.
- A. Marusyk, V. Almendro and K. Polyak, Intra-tumour heterogeneity: a looking glass for cancer?, Nat. Rev. Cancer, 2012, 12(5), 323–334, DOI:10.1038/nrc3261.
- K. K. Zeming, K. Y. Quek, W.-X. Sin, D. B. L. Teo, K.-W. Cheung, C. R. Goh, F. Kairi, E. Lee, F. L. W. I. Lim, M. S.-F. Seng, S. Y. Soh, M. E. Birnbaum and J. Han, Cell trajectory modulation: rapid microfluidic biophysical profiling of CAR T cell functional phenotypes, Nat. Commun., 2025, 16(1), 4775, DOI:10.1038/s41467-025-59789-w.
| This journal is © The Royal Society of Chemistry 2026 |