 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
20 years of microfluidic technology for advancing plant sciences
Louis D.
Cohen
 ,
Eleonora
Moratto
,
Eleonora
Moratto
 and
Claire E.
Stanley
and
Claire E.
Stanley
 *
*
Department of Bioengineering, Imperial College London, London, SW7 2AZ, UK. E-mail: claire.stanley@imperial.ac.uk
First published on 6th February 2026
Abstract
Understanding how plants respond to dynamic and spatially variable stimuli is a key goal in plant sciences. Traditional imaging methods often involve a trade-off between environmental control and spatial resolution, limiting their ability to capture real-time responses in high resolution. Microfluidic technology overcomes these limitations by facilitating precise control of environmental conditions and high-resolution live imaging. In the past two decades, microfluidic technology has increasingly been applied in plant sciences research. This review summarises current applications of microfluidic technology in plant sciences, including studies of root–rhizosphere interactions, tip-growing plant cells, plant protoplasts, and plant phenotyping. Emerging trends are explored, and key research gaps are highlighted.
1 Introduction
Plants must constantly react to environmental cues, requiring a high degree of phenotypic plasticity. How plants achieve this plasticity at different spatial and temporal scales remains an open question in plant sciences: how do plant roots respond to temporally and spatially heterogeneous chemical environments? How do plant roots interact with the soil microbiome at different scales? How do plant cells sense and respond to heterogeneous chemical environments, grow, and develop into complex multicellular organisms? Answering these questions requires precise control of plants' biotic and abiotic environments, combined with high-resolution spatial and temporal imaging of plants. Implementation of classical agar-based approaches1–3 to answer these questions is limited, due to coarse environmental control at the plate scale, and low-resolution imaging. Hence, microfluidic technology aims to fill this methodological gap. As a result, microfluidic technology has been increasingly utilised in the plant and wider biological sciences over the past two decades, having been adapted from its original use in the chemical sciences for the manipulation of very small fluid volumes (fL–nL) to perform chemical reactions with precise control and in a high-throughput manner.4Microfluidic devices are customised microenvironments that have at least one dimension less than one millimetre in size. They are often manufactured from transparent materials and exploit thin optical interfaces to facilitate high-resolution imaging of the enclosed samples. Inclusion of customised internal microstructures and control over fluid perfusion facilitate precise control and measurement of the internal microenvironment. This technology, sometimes referred to as lab-on-a-chip (LOC),5 “chip” technology, or micro-electromechanical (MEMS) systems,6 will be henceforth referred to as microfluidic technology, and associated microfluidic devices. Emerging from the use of microfluidic technology for the study of single cells, such as yeast7–9 and bacteria,7,10,11 the field of plant microfluidics was founded in 2006 with the development of a microfluidic device for trapping tobacco protoplasts12 and later expanded to the study of root-environment interactions with the “RootChip” devices from 2010 onwards.13 Today, the field of plant microfluidics encompasses many areas of plant biology, from plant–microbe interactions to high-throughput phenotyping of plant cells and seedlings.
To date, several reviews14–17 have been explored the application of microfluidic technology to plant sciences, but to date no single review has attempted to synthesise all existing research concerning the application of microfluidic technology to plant sciences. This review aims both to update the field, and to summarise all research that uses microfluidic technology to advance the field of plant sciences, encompassing research from 2006 to 2025. Microfluidic devices for plant sciences can be categorised by the type of biological question the microfluidic device has been designed to answer, which in turn determines device and experimental design. Therefore, we categorise existing research into microfluidic device-assisted: i) imaging of root–rhizosphere interactions, ii) imaging of tip-growing plant cells, iii) imaging and manipulation of single or small groups of plant cells, and iv) plant phenotyping. Current trends in the field and future directions of research are discussed, with a focus on novel microfluidic device designs and fabrication methods.
2 Microfluidic device-assisted imaging of root–rhizosphere interactions
Plant scientists have studied the relationship between plant roots and the rhizosphere for decades, and we now appreciate rhizospheric environments to be one of the most dynamic and heterogeneous environments on Earth.18 Plant roots are required to be phenotypically plastic to respond to this environmental dynamism and heterogeneity. Understanding phenotypic plasticity requires observation of plant phenotypes as a response to changing chemical, biological, and physical stimuli at multiple spatial and temporal scales. Classical approaches involve growing roots on agar plates and measuring the impact of the chemical, biological or physical agents on various root phenotypes such as root development, physiology, cytology, and gene expression.19 This approach has yielded valuable insights, such as the impact of nutrient deficiencies on Arabidopsis thaliana root plasticity,20 but there are several limitations to this approach. Firstly, high-resolution imaging of root development is not possible due to the thickness and opacity of agar. This limits the use of high numerical aperture objective lenses with short working distances, and roots grow out of focus due to gravitropic-induced bending. Secondly, classical approaches can achieve only coarse spatial control of chemical and biological agents, whereas plant developmental processes can be influenced on the micrometre scale. Microfluidic technology offers a way to overcome these limitations. Firstly, customisable microchannel heights facilitate focal confinement of the root, allowing high-resolution images to be taken over long time periods. Secondly, control of fluid perfusion combined with custom microchamber and microchannel geometries results in fine spatial and temporal control of the internal microenvironment. Finally, microfluidic technology can leverage other complementary technologies to probe further into root dynamics. For example, genetically encoded biosensors such as Förster resonance energy transfer (FRET) biosensors21 can provide information on intracellular metabolites and ions of individual plant cells in real time. In this section, the application of microfluidic technology for the investigation of root–rhizosphere interactions will be explored including: i) root–chemical, ii) root–microbe, and iii) root–physical interactions.2.1 Microfluidic technology facilitates high-resolution imaging of roots
Microfluidic devices are typically used to image plant roots by growing a primary root inside a microchannel with dimensions customised for the average diameter of the primary root of the plant being studied. This confines the growth of the primary root to a single focal plane, and aids root growth in a direction amenable to study. Seeds or seedlings (where seeds are germinated off-chip) are inoculated into microfluidic devices to image the growing root from a few days22 to over a month23 after germination. The first microfluidic devices developed to image primary roots consisted of a single straight microchannel to confine the primary root of a dicotyledonous plant such as A. thaliana (Fig. 1A–C). Device design progressed from confining the root apex,13 to the whole root or seedling22,24–28 inside the microchannel. This increased integration of the root into the microfluidic device facilitates longer experiments, as a smaller proportion of the root surface area is exposed to desiccation and nutrient depletion zones.29 Longer experiments are important for facilitating the study of later stages of plant development, and to study plants with longer lifecycles such as tree seedlings.23 For plants with root diameters less than 200 μm, such as A. thaliana, photolithography is an appropriate technique to fabricate the microchannels. However, it is difficult to use this technique for manufacturing channels heights larger than 200 μm. As most economically important crop plants possess roots with diameters more than 200 μm, other microfabrication techniques have been employed to focally confine larger root systems of plants such as rice (Oryza sativa),24Brachypodium distachyon30 and hemp (Cannabis sativa).31 For example, channel heights of 300 μm have been fabricated using Bosch etching,30 and channel heights of more than one millimetre have been fabricated using fused deposition modelling (FDM)24,31 and stereolithography (SLA)32 3D printing techniques (Box 1). Plants such as A. thaliana form a relatively large primary root that grows from the seed, which then branches laterally. This root architecture is amenable to a singular straight microchannel design. However, some plants develop fibrous root systems (Fig. 1D), which consist of multiple thinner roots growing from the seed. This makes the singular straight microchannel design unsuitable. Chai et al. overcame this issue using a multichambered “petaloid” design (Fig. 1E and F), consisting of five separate chambers for root observation, and showed the utility of the petaloid root growth microfluidic (PRGM) device in growing O. sativa (rice).24 In summary, microfluidic devices facilitate high resolution spatiotemporal imaging over long time periods through focal confinement of the plant root. Multiple microfabrication techniques have been employed for manufacturing microchannels of an appropriate size for focal confinement, which are dependent on the size and root architecture of the study species.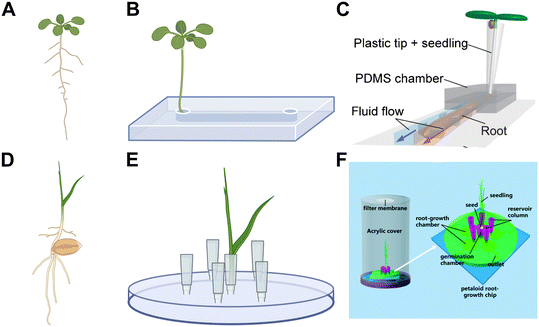 | ||
| Fig. 1 Plants require different microfluidic device designs due to differences in root architecture. (A–C) Plants that form a single primary root during the seedling stage such as A. thaliana (A) can be accommodated in a single straight microchannel device design (B), such as the dual-flow-RootChip (C). (D–F) Plants that form multiple such as rice and wheat often form fibrous root systems (D), which require more complicated device designs to accommodate multiple roots (E), such as the petaloid root growth microfluidic (PRGM) chip with 5 chambers to confine each root individually (F). Image in (C) produced with modification from ref. 28 with permission from John Wiley and Sons, copyright 2023. Image in (F) produced with modifications from ref. 24 with permission from the Royal Society of Chemistry, copyright 2018. Created in BioRender. Cohen, L. (2025) https://BioRender.com/hgd4y66. | ||
Box 1The microfabrication of microfluidic devices can be achieved through a variety of techniques. Photolithography is used to fabricate microfluidic devices with small feature sizes (microchannels, microchambers, etc.). Photolithography involves first spinning a photoresist onto a silicon wafer at a controlled thickness, determining the height of the structures. Subsequently, ultra-violet (UV) light is applied through a photomask to write structures onto the photoresist, determining the length and width of the structures. These patterned silicon wafers form the negative “master mould” of the microfluidic device. Photolithography reliably achieves feature size resolution as small as 1 μm and layer heights as small as 2 μm but typically requires access to specialised equipment and training. Moreover, the maximum height of structures produced by spin coating a single layer of commercially available photoresist is around 200 μm. While this channel height is suitable for A. thaliana primary roots, it is unsuitable for larger root systems. While it is possible to modify the photolithography process to produce thicker structures by replacing photoresist spin coating with constant volume injection,33 this has not yet been attempted for manufacturing large straight channels for roots, and layer heights have more variation than when spin coating. Therefore, alternative microfabrication technologies are required for large microchannel structures. 3D printing technologies have decreased in cost and increased in resolution over the past two decades, making them a viable alternative to photolithography for producing large channel designs (reviewed by Su et al.).34Fused deposition modelling (FDM) 3D printing involves melting a thermoplastic filament and depositing sequentially in layers to form a final master mould product. FDM 3D printing technologies have come down in cost drastically over the past two decades and are straightforward to use. Device designs can be iterated on very quickly, and the material cost of printing designs is low. However, FDM 3D printers are limited to minimum layer heights of several hundred micrometres, with minimum feature resolutions of 50 μm, making them unsuitable to fabricate the smaller layer heights needed for root hair, and protoplast trapping interaction studies. Moreover, issues with surface roughness (which are translated to the PDMS device) are an issue for achieving high-resolution imaging. Therefore, FDM 3D printing is a useful microfabrication technique for microfluidic device designs with simple, large microchamber geometries. Stereolithography (SLA) 3D printing involves using a light source to cure liquid resin into solid objects layer by layer. SLA printing can achieve feature resolutions of up to 10 μm (though this varies greatly depending on printer model) and offers a middle ground of resolution, price, and iterative speed, between FDM 3D printing and photolithography. SLA 3D printing technology can theoretically achieve layer heights as low as 15 μm, meaning it could be used for both small and large layer heights, making it amenable for root hair and protoplast trapping interaction studies. Initial surface roughness of the master mould can be post-processed to achieve low surface roughness. Hence, SLA 3D printing has been used for the manufacture of microfluidic devices used for root–chemical interactions32 and plant phenotyping35 (Fig. 13). By combining SLA 3D printing technology with transparent biocompatible resins, microfluidic devices can be manufactured directly, instead of producing a negative master mould and using PDMS. This makes scalable manufacturing of microfluidic devices possible, in use cases where many microfluidic devices are needed. As the cost of advanced SLA 3D printing technology continues to fall, and resolution continues to increase, this technology may rival the capabilities of photolithography and decrease the barrier to entry for microfluidic device manufacturing. Comparison of technologies for microfluidic device fabrication.
|
2.2 Microfluidic device-assisted imaging of root–chemical interactions
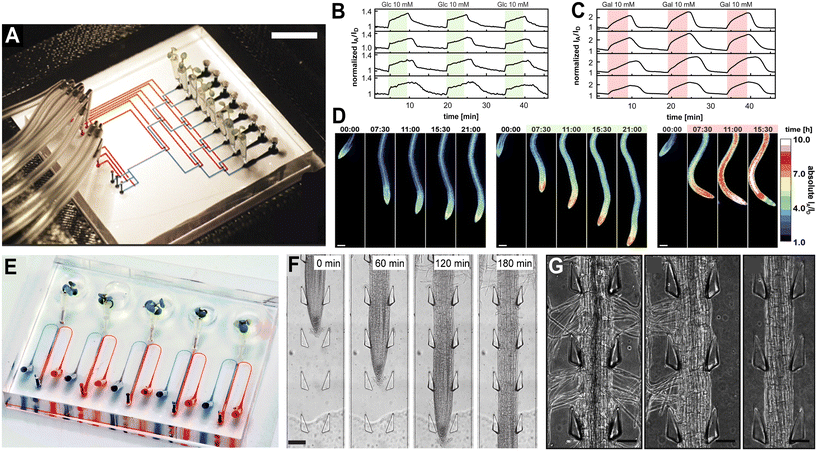 | ||
| Fig. 2 The RootChip and dual-flow-RootChip facilitate studies on root–chemical interactions. (A) The RootChip can accommodate eight mounted live A. thaliana plants, with eight control (blue) and flow (red) channels filled for illustration. Bar = 1 cm. (B and C) The RootChip was used to monitor intracellular glucose (B) and galactose (C) concentrations over time using a FRET biosensor. When exogenous glucose and galactose was applied, an initial increase and subsequent levelling off of both intracellular glucose and galactose concentrations was found, suggesting the presence of a uniport mechanism for hexoses. (D) The FRET intensity ratios of roots growing under the control (left), glucose (middle), and galactose (right) conditions over time was investigated. Images produced with modification from ref. 22 under a Creative Commons Attribution License CC-BY 4.0 (https://www.creativecommons.org/licenses/by/4.0/). (E) The dual-flow-RootChip can accommodate five mounted live A. thaliana plants. Two inlet channels serve each chamber, and red and blue dyes illustrate how two different reagents can be applied simultaneously to either side of the root. (F) A. thaliana roots are guided by PDMS support pillars. (G) Under symmetric osmotic stress conditions, where the medium contains 20% (w/v) PEG 8000 on both sides (right panel), no root hairs are observed. Under asymmetric osmotic stress conditions, with 20% (w/v) PEG 8000 applied only to the right side of the root (middle panel), root hairs are absent on the PEG-exposed side. In contrast, roots grown without PEG 8000 (left panel) show normal root hair development. Images produced with modification from ref. 28 with permission from RSC publishing, copyright 2018. | ||
Investigating root–chemical interactions of plants with fibrous root systems requires multiple root chambers to separate individual roots of fibrous root systems. In this instance, spatial manipulation of the chemical microenvironment can be achieved by differentially treating individual roots of the same root system. Chai et al. chemically treated individual roots of the fibrous root system of O. sativa with a range of concentrations of PEG.24 The authors showed that exposure to PEG in higher concentrations was correlated with roots showing delayed growth, and an increase in root hair density. In summary, manipulation of the chemical microenvironment can be achieved at different spatial scales – at the scale of a single root by separating which area of a single primary root is treated (proximal/distal, left–right axis), as well as at the scale of a root system by treating different roots of the same fibrous root system to different chemical treatments.
Aufrecht et al. took a different approach and designed the rhizosphere-on-a-chip microfluidic device to interface with liquid micro-junction surface sampling mass spectrometry (LMJ-SSP-MS) (Fig. 4).30 The authors achieved this by replacing the glass coverslip interface material with a polyester track-etched (PETE) membrane. This transparent membrane has a high density of 0.4 μm diameter pores, facilitating transport of small molecules such as amino acids from root exudates through the membrane for quantification, while preventing fluid leakage. The properties of this PETE membrane were leveraged to allow both an optical and chemical interface with the device, meaning the authors were able to image root growth at cellular resolution, as well as map the relationship between amino acid concentrations and artificial soil pore geometry at a coarser spatial scale. This membrane interface approach has the potential to provide both optical and chemical interfaces with the device but does not allow for plant tissue to be extracted, limiting analysis to specific analytical techniques compatible with membrane-interface sampling. Further development of both openable and membrane interface microfluidic devices is of great interest to the field of plant microfluidics. In summary, microfluidic devices can be designed to be amenable to analytical techniques to yield further insights into root–chemical interactions. Modifying microfluidic devices to enable compatibility with analytical techniques results in imaging trade-offs but has the potential to facilitate a broader range of data collection from microfluidic systems.
2.3 Microfluidic device-assisted imaging of root–microbe interactions
We have seen how microfluidic technology has contributed to an improved understanding of how plant roots respond to dynamic and spatially heterogeneous chemical environments. Biotic environments are another important factor contributing to rhizospheric heterogeneity and dynamism. Soil microbes interact with plant roots in pathogenic, commensal and mutualistic relationships, and have major impacts on plant health and ecosystem productivity.43 Classical plate, hydroponic, and pot-based assays for root–microbe interaction studies provide valuable insights into how microbes can alter plant phenotypes at coarse spatial scales, but understanding these interactions at a cellular level is difficult using classical techniques. Observing root–microbe interactions with cellular resolution allows the investigation of how microbes search space, forage for nutrients and disperse in rhizospheric environments. Microfluidic technology offers a way to construct a simplified rhizospheric ecosystem and facilitates high-resolution imaging of soil microbes, as well as their impact on plant roots. Observations of growth and space-searching behaviours of plant roots and soil microbes can be combined with fluorescently labelled microbes to characterise spatiotemporal variation in these interactions. Microfluidic devices used to investigate root–microbe interactions use single layer microchannel designs similar to devices used to investigate root–chemical interactions (Section 2.2). These microchannels facilitate focal confinement of the root, but not the smaller microbial partner, within the comparatively large plant root microchannel. Therefore, the growth and space-searching behaviour of the microbial partner cannot easily be observed in real time. However, biological inferences can be made using: i) observation of the impact of the microbial partner on root morphology, and ii) direct observation of the microbial partner using fluorescent microbes or image processing techniques.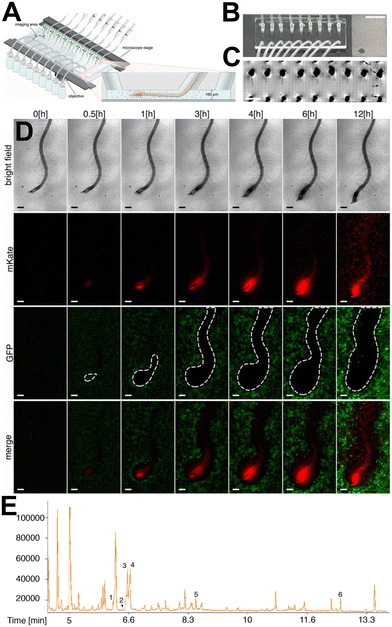 | ||
| Fig. 3 The Tracking Root Interaction System (TRIS) microfluidic device facilitates spatial root-bacterial interactions. (A) Schematic illustration of the TRIS device mounted on a microscope mount, including a side view of a single microfluidic root channel containing root and bacterial cells (insert, not to scale). (B) Top view of Arabidopsis thaliana plants growing inside plastic pipettes attached to the TRIS device. Scale bar, 1 cm. (C) Micrograph of nine A. thaliana roots growing inside the TRIS device captured with 10× magnification brightfield. Arrows: a, inlet port; b, tip inside a root-dedicated port with c, a root extending towards the outlet port; d, outlet port. Scale bar, 5 mm. (D) Select micrographs from four independent imaging experiments showing bacterial competition for the root surface. Wild-type (WT) A. thaliana (brightfield channel) was incubated with GFP-labelled Escherichia coli (green) and mKate-labelled Bacillus subtilis (red). White dashed area in GFP images indicate the exclusion zone of E. coli bacteria from the root by B. subtilis. Scale bars, 200 μm. (E) The TRIS device also facilitates single-root exudate metabolomics. Selected ion chromatogram showing a representative metabolite profile from a single-root exudate of a 10-day-old WT A. thaliana root grown in the TRIS device, analysed with gas chromatography coupled to time-of-flight mass spectrometry. The numbered peaks correspond to selected metabolites identified from authentic standards: (1) phosphoric acid, (2) isoleucine, (3) glycine, (4) succinic acid, (5) pyroglutamic acid, and (6) myo-inositol. Images produced with modification from ref. 45 with permission from Proceedings of the National Academy of Sciences of the United States of America, copyright 2017. | ||
In microbial partners where fluorescent labelling is more difficult, computational approaches can be used to improve the visualisation of the microbial partner in rhizospheric environments. Cohen et al. used software to track the movement of motile Phytophthora parasitica zoospores as they approached an A. thaliana root from the device outlet.47 The authors were able to show that compared to a non-root control, the presence of the root alters the speed distribution of zoospores, and that the movement speed and directedness of zoospores is only impacted at a distance of 300 μm or less from the root, suggesting a root-derived signal diffusible over short distances. Therefore, fluorescent labelling, whole-cell biosensors, and computational image processing are useful tools for investigating microbes in a large root channel to compensate for insufficient focal confinement.
2.4 Microfluidic device-assisted imaging of root–physical interactions
We have seen how microfluidic technology can uncover spatial interactions between roots and chemical agents, as well as roots and soil microbes, but just as important for understanding root–rhizosphere dynamics is the interplay between roots and the physical structure of the soil around them. Plant roots are influenced by and reshape the physical structure of soil, which impacts soil properties such as nutrient hotspots,48 contributing to the complexity of soil across a continuum of spatial scales. For example, it has been observed that bacterial strains that cannot coexist in a spatially homogeneous environment can coexist in a spatially heterogeneous environment.49 How the physical structure of soil generates these niches is a subject of great interest for soil scientists, but simplifying the soil system to investigate individual hypotheses is difficult. Microfluidic technology is uniquely suited to investigate this interplay as the complex structure of soil can be simplified to investigate individual hypotheses and therefore reduce the complexity of experiments. For example, Aufrecht et al. set out to investigate the impact of soil particle shape and size using the rhizosphere-on-a-chip device (Fig. 4).30 The authors constructed a natural sand-mimicking particle structure and introduced the model grass B. distachyon into this microchamber. The authors showed that this spatial heterogeneity resulted in hotpots of amino acid root exudates in comparison to an open geometry control, as measured using LMJ-SSP-MS, and profiled the variability in amino acids detected at several sampling sites (Fig. 4C). The growth of roots and root hairs into small interpore spaces (Fig. 4B) was observed alongside amino acid exudate data. A subsequent study using the same device but using Cannabis sativa (hemp) as the model organism, showed the utility of rhizosphere-on-a-chip devices for studying a range of plants with larger root systems.31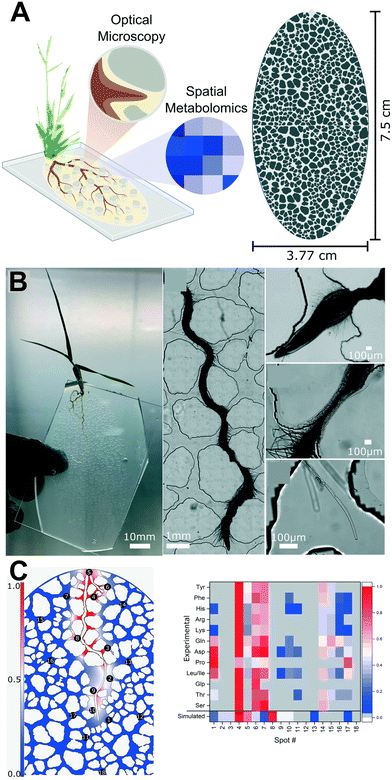 | ||
| Fig. 4 The rhizosphere-on-a-chip facilitates root–physical interaction studies. (A) The rhizosphere-on-a-chip mimics the pore structure of natural sands, and allows both optical microscopy and spatial metabolomics of the root's microenvironment through liquid micro-junction surface sampling mass spectrometry (LMJ-SSP-MS). (B) The rhizosphere-on-a-chip allows long-term culturing of Brachypodium distachyon roots (from left right); a seedling reaches developmental milestones two weeks post germination, roots are able to navigate small spaces, root hairs span inter-grain gaps. (C) Amino acid exudate hotposts are detected and spatially resolved within the microchannels. The left panel shows simulated root exudate concentrations after two hours of flux from the traced root boundaries of a biological replicate. Numbered circles represent spots where root exudates were sampled using LMJ-SSP-MS. The right panel shows a heat map comparing the amino acid profiles of simulated and experimental replicates from each spot sampled from a 12-day old B. distachyon root. Images produced with modification from ref. 30 under a Creative Commons Attribution License CC-BY 3.0 (https://creativecommons.org/licenses/by/3.0/). | ||
Microfluidic technology can also be utilised to investigate growth behaviour mechanics and quantify physical properties of plant. For example, Zhang et al. observed the impact of channel width and sharp corners on the rate of root growth in A. thaliana.50 Liu et al. used sodium chloride as an osmotic treatment to measure cell wall elasticity using the RootChip-8S.51 By applying media flow to growing A. thaliana roots, Agarwal et al. were able to induce and monitor thigmomorphogenesis inside a custom microfluidic device.52 Li et al. utilised a custom microfluidic device with a cantilevered sensing pillar to measure the protrusive force of a growing A. thaliana root.53 Lastly, intracellular dynamics upon root compression was monitored using a custom microfluidic device that applies mechanical stimulation to a growing A. thaliana root. Vinet et al. used a two-chamber microfluidic device design, with the top chamber exerting a compressive force on a growing A. thaliana root in the lower chamber. The authors were able to monitor changes in ROS and intracellular calcium using probes and GECIs respectively.54 In summary, microfluidic technology has recently given important insights into how roots interact with the physical structure of soil to create a spatially heterogeneous environment, but there are still many open questions. For example, integrating studies of physical structure with soil microbes, and looking at how physical structures generate unique chemical microenvironments, which in turn create biological niches for microbial colonisation.
3 Microfluidic device-assisted imaging of tip-growing plant cells
Tip-growing plant cells are an evolutionarily diverse group of plant cells, which share an ancestral mechanism of lateral extension. Unlike plant roots, which grow by meristematic division, tip-growing plant cells use turgor pressure and cell wall remodelling to expand and grow forwards.55 Three different types of tip-growing plant cells will be discussed: pollen tubes (tubular protrusions formed by the pollen grain of flowering plants in the angiosperm clade), moss protonemata (filamentous networks formed from haploid spores by mosses in the bryophyte clade), and root hairs (terminally differentiated lateral extensions of root epidermal cells (trichoblasts) and are formed by almost all land plants). Tip-growing plant cells are single cells, meaning their growth rate, direction, intracellular processes, and branching behaviour (where applicable) must be investigated at single-cell resolution. This makes tip-growing plant cells difficult to study in classical plate-based approaches due to insufficient focal confinement, and therefore the optical advantages afforded by microfluidic technology are particularly useful in studying tip-growing plant cells. As a result, the development of microfluidic technology for studying tip-growing plant cells has been an important research focus in the field of plant microfluidics over the past two decades. While these three types of tip-growing plant cells are morphologically and evolutionarily diverse, their shared method of growth means designing microfluidic devices to study their growth and development shares a common approach, with slight differences in device design depending on the biological question being asked. For example, a common device consideration is how to focally confine tip-growing plant cells. In pollen tubes and moss protonemata, this means specialised structures to trap pollen grains56 and moss spores or protoplasts, respectively.57 Whereas in root hairs, this takes the form of a multilayer device fabrication process, with separate layers to focally confine the root and root hairs.583.1 Microfluidic device-assisted studies on pollen tubes
Pollen tubes are tubular protrusions formed by the pollen grain when landing on a receptive stigma and deliver male gametes from the pollen grain to the female gametophyte in the flower ovule. The speed of pollen tube elongation directly influences fertilisation success, and therefore is strongly selected upon, resulting in pollen tube elongation rates up to 600 μm h−1.59 Pollen tube growth can be triggered in vitro with virtually identical morphology to in vivo,60 albeit with slower growth rates. The pollen tube must invade plant tissue inside the pistil of the receptive flower61 and grow through gaps between cells smaller than the diameter of the pollen tube.61,62 Though it is likely some digestive action by enzymes and programmed cell death of the pistil tissue facilitates pollen tube growth, it is unlikely that the substrate is liquefied completely.63 Therefore, the pollen tube likely exerts a penetrative force and must search space and navigate challenging environments using chemical, physical, and possibly electrical, cues. Microfluidic technology is a useful way to study pollen tube guidance as it allows high-resolution imaging of growing pollen tubes and can be designed to mimic the environment pollen tubes are exposed to in vivo.While these studies provided robust evidence for chemotropic growth on groups of pollen tubes, higher resolution spatial control of the chemical agent is required to investigate the chemotropic growth of single pollen tubes. This has been achieved through laminar flow, and through integrating microfluidic pumps. Nezhad et al. utilised laminar flow inside the pollen tube microchannel, allowing continuous exposure to two different chemical agents on either side of a C. japonica pollen tube72 (Fig. 5), in a similar way to the df-RootChip28 as seen previously (Fig. 2D–F). The utility of the device was demonstrated through two experiments. Firstly, asymmetric application of extracellular calcium resulted in the pollen tube navigating to an optimal calcium concentration, as opposed to navigating to one side or the other, as would be expected by a single molecular mechanism of chemotropism. This suggests multiple interacting regulatory mechanisms controlling calcium chemotropism. Secondly, the enzyme pectin methyl esterase, which acts to rigidify the pollen tube cell wall, was applied extracellularly on one side of the pollen tube. This resulted in the growth direction of the pollen tube to turn away from the agent, which supports existing hypotheses of the mechanism of directional growth in pollen tubes. Therefore, the separation of chemical agents either side of the pollen tube using laminar flow is a useful way to conduct targeted chemotropic assays. On the other hand, Yanagisawa et al. achieved spatial control using a microfluidic injector device, which can inject a small volume of sample into the growth chamber utilising an on-chip electro-osmotic pump.73 This allows for a small volume of reagent to be applied directly to growing A. thaliana pollen tubes. This technique offers an on-chip alternative to off-chip direct reagent application techniques, in which reagent is embedded onto a small bead, which in turn is placed on the growing pollen tube using a micromanipulator. This microfluidic injector device allows on-chip chemotropic assays without continuous flow, allowing chemotropic studies where flow is undesirable.
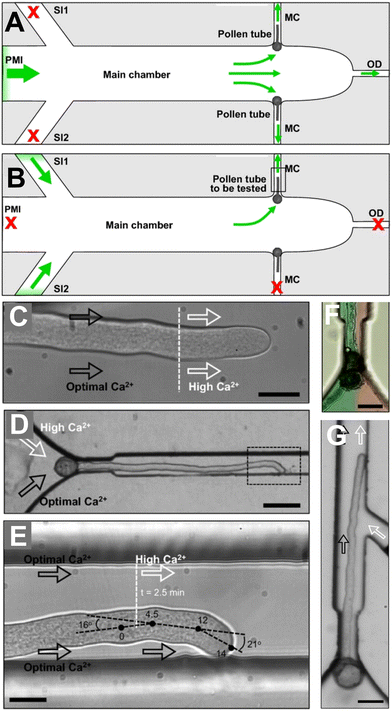 | ||
| Fig. 5 Microfluidic device facilitates asymmetric application of calcium to growing Camellia japonica pollen tubes. (A) Pollen grains are first deposited and orientated correctly during experimental setup. (B) The pollen medium inlet (PMI) and one of the media channels (MC) are blocked during the experiment. Medium is injected through two side inlets (SI1, SI2). Drawings not to scale. (C) Pollen tubes were found not to change orientation during a global change from optimal calcium (2.54 mM) to high calcium (5.08 mM) but were found to slow down by 20%. Scale bar 20 μm. (D) Pollen tubes were found to reorientate when subject to asymmetric calcium. Scale bar, 70 μm. (E) Medium with optimal calcium concentration (2.54 mM) was supplied through both SIs, then high calcium (5.08 mM) was supplied through one of the SIs. The pollen tube was found to changes its growth rate and turn twice. Scale bar, 20 μm. (F) Asymmetric application of calcium was achieved through laminar flow, exemplified using red and green food colouring. Scale bar, 50 μm. (G) Alternative design where the channel delivers calcium only to the apical region of the pollen tube, not the base. Scale bar, 50 μm. Images produced with modification from ref. 72 with permission from John Wiley and Sons, copyright 2014. | ||
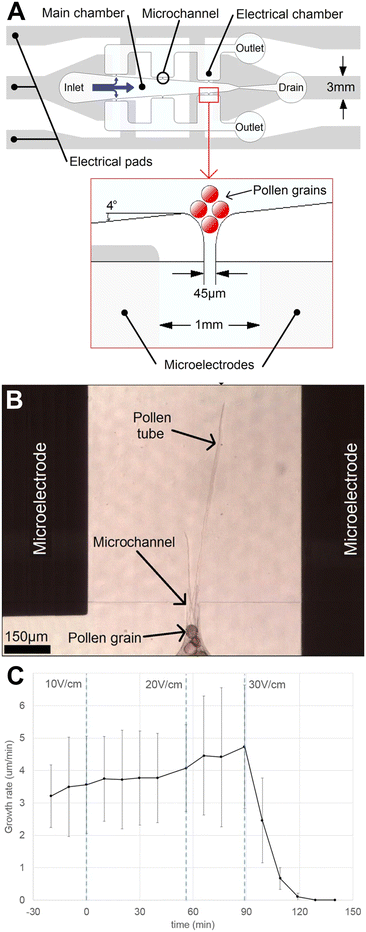 | ||
| Fig. 6 Single-cell electrical lab-on-chip (ELoC) reveals impact of electrical fields on pollen tube growth. (A) The single-cell ELoC schematic involves trapping pollen grains at the entrance to microchannels 45 μm wide. (B) Upon germination, the pollen tubes grow into a microchamber with microelectrodes on either side and are subjected to DC electric fields. (C) Exposure to weak DC electric fields was found to result in negligible impact on instantaneous growth rate, while exposure to DC electric fields above 30 V cm−1 was found to rapidly stop growth. Images produced with modification from ref. 74 under a Creative Commons Attribution License CC-BY 4.0 (http://creativecommons.org/licenses/by/4.0/). | ||
Microfluidic technology both facilitates focusing on the deformation of a single tip-growing cell at high magnification and manoeuvring cells into suitable positions to encourage turgor pressure to be exerted in a predefined function. Nezhad et al. used a modified version of the TipChip to measure the force exerted by growing pollen tubes when squeezing through gaps smaller than their diameter (Fig. 7).63 Changes in width were measured optically, and, in combination with optical measurements of PDMS deformation, the force exerted by the pollen was calculated using finite element modelling. Using another iteration of the TipChip, Nezhad et al. this time used the bending lab-on-chip (BLOC) to measure the Young's modulus (stiffness measurement) of the cell wall surrounding a pollen tube.78 The device traps a single pollen tube, and media flows perpendicular to the growth of the pollen tube. Optical measurements of the deformation of the pollen tube, combined with computational modelling, allowed the Young's modulus to be calculated. Burri et al. took a different approach and combined a microfluidic device with a force sensor, positioned at the output of one of the microchannels.80 This force sensor quantified the minimum force required for lily and Arabidopsis pollen tubes to sense and react to an obstacle. By combining the force readout data with microscopy data, the internal turgor pressure of growing pollen tubes was calculated. This force sensor readout has the potential to ameliorate some of the disadvantages of using deformation calculations to measure force exerted, but both methods give force readouts and turgor pressures with similar literature values to non-microfluidic measurements using probes. Therefore, microfluidic technology has improved upon established methods for chemosensing, electrosensing, and mechanosensing assays using high-resolution real-time imaging of single pollen tube cells.
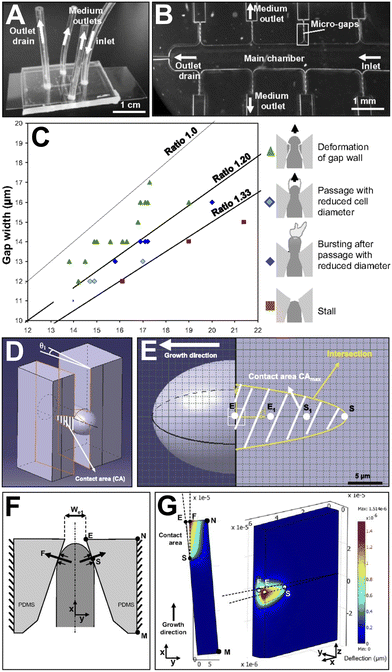 | ||
| Fig. 7 The TipChip facilitates studies on penetrative forces of pollen tubes using deformation from microgaps and finite element modelling. (A) The TipChip experimental setup illustrating locations of inlets and outlets. (B) The TipChip device design. (C) Variation in Camellia japonica pollen tube behaviour was found to depend on the ratio between gap and pollen tube width. Between ratios of 1.00 and 1.20, pollen tubes deform the microgap and pass through (green triangles). Between a ratio of 1.20 and 1.33, pollen tubes pass through with a reduced diameter (light blue diamond) and frequently burst after passing through (dark blue diamond). At a ratio of 1.33 and above, pollen tubes cannot pass through the microgap. (D) Finite element modelling simulating force exertion by the pollen tube on the walls of the microgap. (E) Three-dimensional model of the pollen tube and microgap when the pollen tube is in maximum contact with the microgap. The four points represent reference points for the contact area, starting with S upon initial contact, and finishing at E with exit from the microgap. (F) Top view of the modelled geometry is shown, along with relevant forces acing on the microgap. (G) The deflection of the poly(dimethylsiloxane) (PDMS) material is shown, with respect to the top view (left) and side view (right). The false colour scale represents the degree of sidewall deflection at an effective pressure of 0.15 MPa. Images produced with modification from ref. 63 with permission from Proceedings of the National Academy of Sciences of the United States of America, copyright 2013. | ||
3.2 Microfluidic-assisted studies on moss
Moss species, specifically the model moss Physcomitrella patens, have a simple body plan and efficient gene targeting via homologous recombination,81 which has resulted in interest in using P. patens as a model organism to study the evolution of developmental processes in land plants at a subcellular level.82 Moss protonema is the filamentous network formed from a haploid spore, with the apical cell within a filament dividing as the filaments grow radially outward. Within this network, buds form off protonematal filaments and eventually develop into gametophores. Non-microfluidic techniques have been used to study the development of moss protonema, but their utility is limited due to poor focal confinement. In one agar-based technique, protonemata are placed on an agar pad, sealed with a coverslip and imaged immediately. However, cells stop growing after a few hours (presumably due to insufficient gas exchange and limited nutrients). An alternative method involves culturing protonemata in a dish with a coverslip at the bottom covered with a thin film of agar for several days. The protonemata are given enough time to grow close enough to the coverslip and then imaged. The tissue grows well, however, few cells reach the surface of the coverslip, so only a small proportion of the sample can be imaged. Microfluidic technology has assisted in improving culturing and imaging of protonemata. Firstly, sufficient focal confinement means a majority of the sample can be imaged with high-resolution microscopy. Secondly, PDMS is highly air permeable,83 meaning P. patens is able to grow in these devices for several weeks.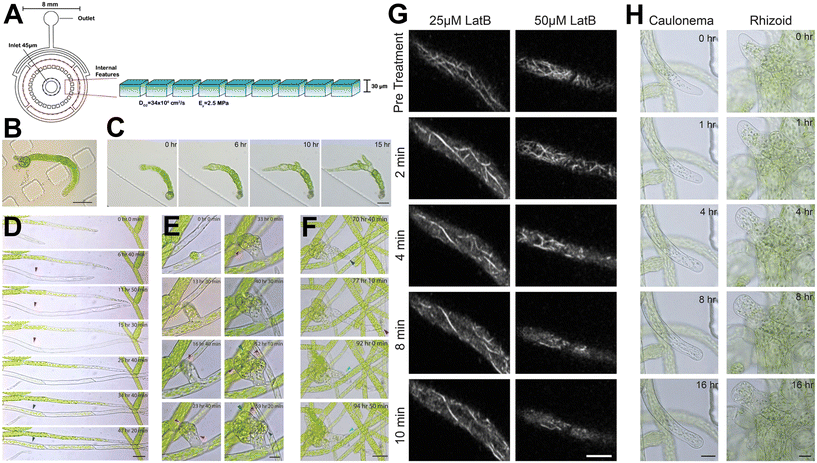 | ||
| Fig. 8 Microfluidic culture chamber facilitates the observation of developmental events, subcellular dynamics and pharmacological studies. (A) Schematic illustration of the microfluidic culture chamber, with a chamber height of 30 μm to focally confine Physcomitrella patens. (B) Protoplast regeneration was observed, as well as subsequent cell division events within 9 days. Scale bar, 50 μm. The initial protoplast is outlined with a black dashed line. (C) Multiple cell division events and branches were observed after protoplast regeneration over 15 hours. Scale bar, 50 μm. Developmental events of P. patens were observed inside microfluidic culture chambers. (D) A wild-type caulonemal subapical cell (red arrows) was observed turning into a chloroplast rich cell (blue arrows) after a day. Scale bar, 50 μm. (E) The development of a bud initial from a single cell was observed after 40 hours. Scale bar, 20 μm. A rhizoid initiated from the lateral basal cell (dark blue arrow). (F) Explosion of the rhizoid (purple arrow), with a new rhizoid tip reinitiated after 15 hours. Scale bar, 50 μm. (G) Two different concentrations of an actin-depolymerising drug, latrunculin B (LatB) were used on P. patens protonematal subapical cells expressing the actin fluorescent marker LifeAct-mRuby2. Rapid depolymerisation of most actin filaments in both concentrations was observed. (H) Tip growth was inhibited by exposure to the higher concentration of LatB. Images produced with modification from ref. 84 under a Creative Commons Attribution License CC-BY 4.0 (http://creativecommons.org/licenses/by/4.0/). | ||
3.3 Microfluidics-assisted studies on root hair growth dynamics
The root hair is a terminally differentiated lateral extension of root epidermal cells called trichoblasts. Root hairs display distinct cellular properties from trichoblasts, including cell and vacuole size.88 Root hair development occurs in three steps: (i) initiation via outgrowth from a trichoblast cell, (ii) elongation via tip growth, and (iii) maturation when the tip of the root hair cell ceases growing, and is well characterised. In A. thaliana, specific genes have been identified as important for root hair development, some of which have been mapped using the RootChip-8S microfluidic device.89,90 Root hairs are important for both uptaking water and nutrients from the soil, and to anchor the plant to the soil.91 They also play a crucial role in root–microbe interactions, such as nodulation, where root hairs are involved in the uptake of rhizobia through an infection thread.92 Generally, root hairs act as the first “point of contact” for any microbe initiating contact with the root surface, as root hair protrusions may cover the root, and extend up to several mm away from the root surface.88 Nuclear movement within root hairs may be associated with various environmental stimuli such as plant–microbe interactions, nutrient and water uptake, and light response.93 During root hair growth, nuclear position and movement are tightly regulated by actin and associated proteins in A. thaliana.94 However, due to the small diameter of root hairs (8–20 μm wide depending on the species), tracking the movement of root hair growth and organelles is difficult with classical plate-based techniques. Microfluidic technology, and fluorescent labelling, allow observation of root hair growth and subcellular dynamics in real time. Using the df-RootChip, Stanley et al. showed that root hairs can rapidly modulate their growth rate in response to phosphate availability in a cell-autonomous manner.28 Yanagisawa et al. looked at the ability and subsequent impact on the growth of root hairs to penetrate gaps smaller than the diameter of their cells.68 The authors found that 77% of A. thaliana root hairs were able to penetrate through 1 μm constrictions, suggesting the shape of the root hair apex can be deformed according to its environment. The authors next wanted to see whether root hair nuclei were able to cross the microgaps. The authors used genetically encoded fluorescent UBQ10pro::H2B-mClover line, in which the roots and root hair nuclei are fluorescently tagged. This allowed the authors to observe root hair nuclei passing through the 1 μm constrictions. Singh et al. investigated root hair nuclear dynamics further by analysing the shape and location of root hair nuclei during and after root hair extension.58 The authors used z-stacks to construct a three-dimensional model of nuclear shape through time. They found that the shape of root hair nuclei is highly dynamic and undergoes significant deformations in its three-dimensional shape. Moreover, the authors found that nuclear aspect ratio (ratio of width to height) increased when root hairs stopped growing (mature phase) compared to the elongation phase. In summary, it has been demonstrated that microfluidic technology can be used in combination with genetically encoded fluorescent labels to correlate organelle and root hair growth dynamics, contributing to a greater understanding of the genetic machinery that underpins root hair development.4 Microfluidic device-assisted plant protoplast, ovule, and embryo imaging and manipulation
Plant protoplasts are plant cells where the pecto-cellulosic cell wall has been removed, while keeping the cell membrane intact. They can originate from many parts of the plant (typically the mesophyll) and are useful as the removal of the cell wall allows easier genetic manipulation of plant cells using protoplast fusion. In suitable culturing conditions, plant protoplasts can dedifferentiate, divide, and subsequently re-differentiate to eventually reform the whole organism. Therefore, plant protoplasts are useful both as tools for genetic manipulation, and for plant development studies. Investigating plant development using protoplasts requires individual protoplasts to be trapped and observed throughout their developmental process in a high throughput manner, which is difficult in classical approaches using flasks. Microfluidic technology has been shown to offer a method to effectively culture, manipulate, and follow the development process of protoplasts over time. Plant embryos are the early stages of a plant that develops from a fertilised ovule. Studying the development of plant embryos and ovules underpins our understanding of plant development and cell fates. Classical approaches to observe plant ovules and embryos involve dissection, dispersion on a culture dish, and observation using an inverted microscope. Maintaining ovules and embryos in a fixed position over long time periods, as well as manipulating the orientation of ovules and embryos to observe all axes of development is challenging using this approach. Microfluidic technology therefore facilitates the trapping and manipulation of plant ovules and embryos for long-term studies in development.4.1 Microfluidic device-assisted imaging of plant protoplasts
Moreover, microfluidic technology has facilitated studies on protoplast fusion and cytoskeletal organisation. Typically, investigating which agents may or may not cause fusion of protoplasts is difficult, as timelapse microscopy of individual plant cells is required. Maintaining a suitable environment and keeping cells in focus is challenging in agar-based methods, especially if high-throughput investigation of different chemical agents is desired. Microfluidic technology is a useful way to both culture plant protoplasts and obtain optical data over long time periods. Wu et al. showed that microfluidic technology can be used for inducing and observing protoplast fusion events.95 The authors showed that PEG exposure resulted in chemically induced tobacco protoplast fusion events and were able to achieve a fusion efficiency of 28.8%, which is similar to the fusion efficiency achieved on classical agar plate-based approaches. Seidel et al. took this further, by delving deeper into the parameters that impact protoplast fusion efficiency, and by designing a microfluidic device to achieve targeted fusion of individual protoplasts,96 instead of the batch fusions performed by Wu et al.95 The authors were able to achieve this by trapping individual pairs of A. thaliana plant protoplasts in pillar trapping structures. These targeted protoplast fusions allowed Seidel et al. to observe the shape of the resulting cell body after fusion to check for abnormal cell shapes, as well as performing vitality staining for up to 10 hours after the suspected fusion event. This enabled the authors to be more confident in assays of fusion efficiency. The authors found that the fusion of protoplasts depended on the PEG concentration of the fusion buffer, the flow velocity inside the device, and the osmolarity of the fusion buffer. Moreover, Durand-Smet et al. used a microfluidic well device design to test the contribution of cell geometry of A. thaliana protoplasts with GFP-fused microtubules to the final organisation of the cytoskeleton.97 The authors trapped protoplasts in wells of different shapes, which facilitated long-term high-resolution imaging of the cytoskeleton using scanning confocal microscopy. The authors found that the cytoskeleton aligns with the long axis of the cell and developed a model for predicting the importance of microtubule severing to self-organising microtubules (Fig. 9). This study revealed that not only does the cytoskeleton impact the shape of the cell, but that inversely the shape of the cell impacts the cytoskeletal organisation Because the confined cells in the study were unable to elongate and their microtubules were predominantly aligned with the longitudinal axis, this study demonstrates that the transverse microtubule orientation characteristic of elongating plant cells is energy dependent and relaxes to a longitudinal configuration in the absence of elongation. Therefore, microfluidic technology has facilitated the targeted study of protoplast fusion and cytoskeletal structure, allowing individual fusion events and microtubule self-organisation to be observed and assessed in detail. This has contributed to a greater understanding of the parameters required for fusion, facilitating protoplast fusion to be used in a wider range of chemical environments.
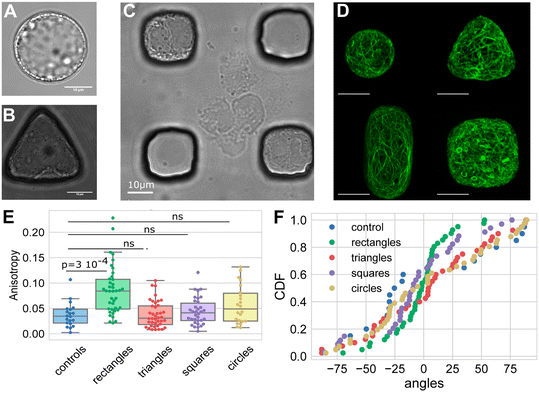 | ||
| Fig. 9 Custom microfluidic device microwell geometries facilitate studies on protoplast geometry–cytoskeleton interactions. (A–C) Description of the experimental setup. (A) Arabidopsis thaliana protoplasts are generated, and (B) trapped in micro-wells of different geometries forming (C) an array. (D) These different protoplast geometries were then used to investigate the impact of geometry on actin network organisation. Image shows protoplasts expressing FABD-GFP reporter confined in microwells of different geometries. Scale bars, 10 μm. (E) Boxplot of the anisotropy of the actin network for each geometry tested. (F) Graph showing the cumulative distribution of the average actin network angle across the different geometries tested. Images produced with modification from ref. 97 under a Creative Commons Attribution License CC-BY 4.0 (http://creativecommons.org/licenses/by/4.0/). | ||
4.2 Microfluidic device-assisted manipulation of plant cells
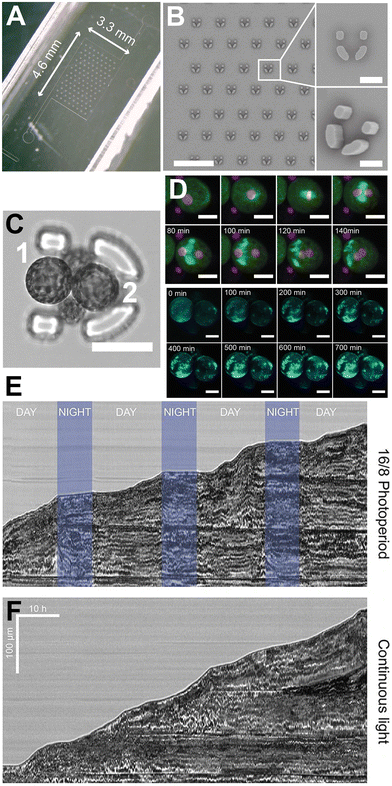 | ||
| Fig. 10 Microfluidic device for the trapping and observation of plant protoplasts. (A) Photograph of the microfluidic device. (B) SEM micrograph of the device. Scale bar, 500 μm (left), 40 μm (right). (C) Confocal image of a trap containing four H2B-mRFP/microtubules-GFP protoplasts. Scale bar, 40 μm. (D) Protoplasts 1 and 2 are in focus and labelled for subsequent observations. Confocal timelapse (top panel) of the first cell division event of protoplast number 2 (purple: H2B-mRFP, green: microtubules-GFP). The GFP channel is in maximum projection mode. Scale bar, 20 μm. Maximum projection confocal timelapse (bottom panel) of the cell wall regeneration around protoplasts 1 and 2 using the dye calcofluor white. Scale bar, 20 μm. Kymograph of the protonema growth under a 16–8 photoperiod (E) and under continuous light (F) using a custom microscope contained within a plant incubator. Images produced with modification ref. 57 under a Creative Commons Attribution License CC-BY 4.0 (http://creativecommons.org/licenses/by/4.0/). | ||
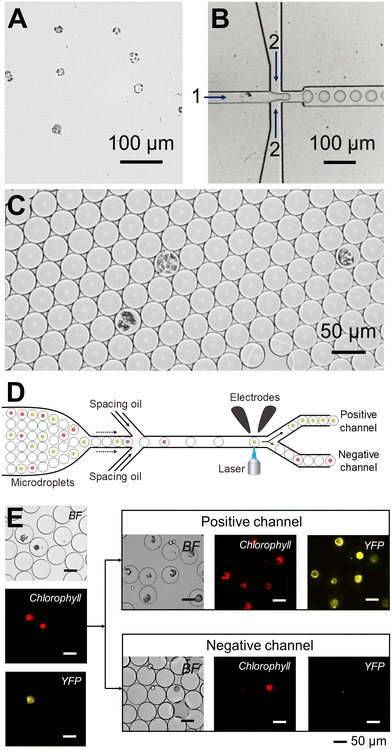 | ||
| Fig. 11 Droplet microfluidic technology facilitates precise manipulation of individual plant protoplasts. (A) Droplet microfluidic technology was used to encapsulate Marchantia polymorpha protoplasts using (B) a flow-focusing microfluidic device. (C) Image showing M. polymorpha protoplasts encapsulated into microdroplets. (D) Schematic of microfluidic device used to sort M. polymorpha protoplasts. (E) Representative brightfield and fluorescent micrographs used for sorting M. polymorpha protoplasts into positive and negative channels using their mVenus fluorescence intensity. Images produced with modification from ref. 104 under a Creative Commons Attribution License CC-BY 4.0 (http://creativecommons.org/licenses/by/4.0/). | ||
5 Microfluidic device-assisted plant screening
Optimising environmental conditions for plant growth and development is important for both plant and agricultural sciences. Due to the large number of environmental parameters plants are exposed to that influence growth and development such as temperature, nutrients, hormones, moisture and other chemical agents, high throughput phenotyping is necessary. Classical agar and soil mesocosm approaches for assessing the impact of a chemical agent on a plant phenotype involves preparing plates or pots of varying concentrations of the chemical agent, spotting seeds and observing plants over time, while incubating in a plant incubation chamber or growth room. This is space and time intensive, and not feasible for studies involving many different chemical agents. While more efficient methods of quantifying root traits using automated root analysis software105 has improved the efficiency of phenotyping some plant traits, a major limitation is the lack of multiplexing (the ability to test multiple chemical agents simultaneously) in a small environmentally controlled space. Efficient seed/seedling trapping and perfusion control makes microfluidic technology suitable to fill this methodological gap and facilitate high-throughput phenotyping of plant seedlings and organs.5.1 Microfluidic device-assisted high-throughput plant phenomics
Microfluidic technology facilitates high throughput imaging of seedling morphology at the organ and tissue level over a range of chemical conditions. High-throughput observation of changes in seedling morphology is used to link genotypes to phenotypes, potentially uncovering the mechanisms behind plant disease. Jiang et al. modified the design philosophy of early RootChip devices (Section 2) to allow for a vertically orientated device for gravitropism, and to allow seed germination to be monitored on chip (Fig. 12).26 The authors used hydrodynamic trapping to ensure a single Arabidopsis seed was trapped per well, and were able to phenotype different A. thaliana mutants, including cellular morphology using genetically fluorescent labels inside the device. The authors also show a variant of the device with an open top, that is shown to facilitate longer term growth studies of A. thaliana seedlings of up to 15 days, enabling later developmental stages to be investigated. This device is space efficient, but the device can only investigate one concentration of chemical agent (here tested using different media) at a time. To enable more precise control over the concentration of chemical agents, Jiang et al. modified the device to integrate a hormone concentration gradient generator (CGG).106 This was shown to offer fine control of concentration gradients through splitting, combining and mixing of two input concentrations, delivering 8 different output concentrations to the 8 seedlings in the plant chip. This technology is best suited for investigating the first developmental processes in small plants such as the model A. thaliana, where soil mesocosms are not required. Microfluidic technology is useful not only in high-throughput observation of plant phenotypes, but also in efficiently generating stimulus gradients. For example, Wang et al. used an inclined microfluidic device on top of a thermos-electric cooling plate, to generate a temperature gradient across an A. thaliana plant.107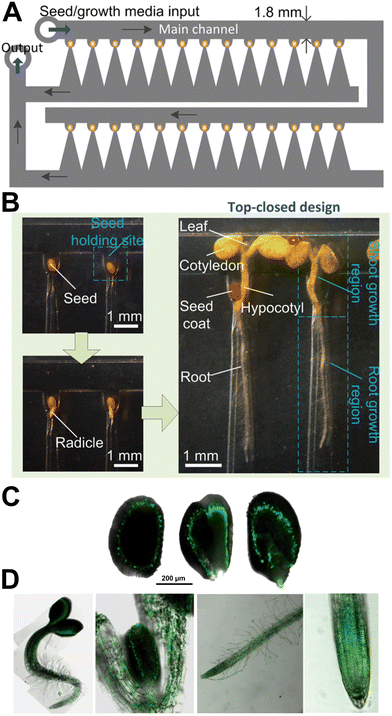 | ||
| Fig. 12 Microfluidic technology facilitates high-throughput phenotyping of plant germination and development. (A) Schematic illustration of the plant chip. (B) The device hydrodynamically traps Arabidopsis thaliana seeds in “holding sites”. Micrographs show in-chip seed germination and development, with major plant organs labelled. (C) IM GFP A. thaliana plants were grown inside the plant chip to follow IM expression, and confocal scanning laser microscopy was used to show seed germination over 18 hours post germination. (D) IM expression in several plant organs was also monitored. Left to right: 1 day old seedlings at a magnification of 10×, 5 day old leaves at 20×, 5 day old root at 10×, 5 day old root tip at 40×. Images produced with modification from ref. 26 with permission from the Royal Society of Chemistry, copyright 2014. | ||
5.2 Microfluidic device-assisted high-throughput imaging of gene expression
High-throughput plant phenotyping microfluidic devices can be combined with complementary techniques such as genetically encoded reporters and nucleic acid sequencing to leverage deeper insights into plant biology. Busch et al. introduced the RootArray, which can accommodate 64 A. thaliana seedlings, and has separate root and shoot compartments, which are supplied with liquid and gas respectively (Fig. 13).35 The authors used confocal microscopy to image 12 different transgenic reporter lines over several days and later reconstructed three-dimensional reconstructions from stitched high-resolution images. This allowed the authors to capture detailed expression maps of the 12 reporter constructs under different environmental conditions, allowing changes in gene expression to be linked to environmental conditions in a high-throughput manner.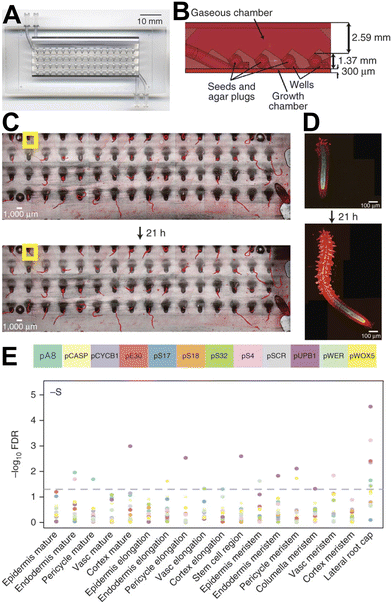 | ||
| Fig. 13 The RootArray microfluidic device facilitates high-throughput imaging of gene expression in Arabidopsis thaliana plants. (A) The RootArray, consisting of 64 individual wells. (B) Cross-section of the RootArray shows that each well connects the upper and lower chambers, enabling the roots to extend into the liquid chamber while the shoots grow upward into the gaseous chamber. The top and bottom boundaries of the illustration represent coverslips, which seal the RootArray device. Root growth can be imaged over multiple hours. (C) A representative low-resolution overview image. (D) A stitched high-resolution image of a single root captured at two different time points. (E) A gene expression map under sulphur-deficient medium obtained through microscopy acquisition of GFP reporter constructs and semi-automated image acquisition and analysis steps. Different tissue types are displayed along the x-axis, while the y-axis shows the −log10 false discovery rates (FDRs) representing changes in gene expression relative to expression in the standard medium for each color-coded reporter gene. The dashed horizontal line marks the 5% FDR significance threshold. “Vasc” denotes vasculature. Images produced with modification from ref. 35 under a Creative Commons Attribution License CC-BY 4.0 (http://creativecommons.org/licenses/by/4.0/). | ||
6 Outlooks and future research
6.1 Integration and iteration of microfluidic devices drives progress in plant microfluidics
The field of microfluidics has advanced plant sciences greatly in the past two decades, from single straight channel RootChip designs22 and single chamber devices for protoplast culturing,12 to microfluidic devices integrating on-chip chemical analysis,30 and force measurements.63 This trend of increased integration of previously off-chip experimental components into microfluidic devices has the benefit of increased miniaturisation of experimental components, finer spatial control of the internal microenvironment, and the facilitation of novel on-chip assays. Taking this further, microfluidic studies for plant sciences would benefit from the integration of a broader range of sensors for environmental measurements such as pH, temperature, and oxygen concentrations, as has been implemented in adjacent microfluidic fields108 and recently in root–rhizosphere interactions.109 However, increasing integration also increases the complexity of manufacturing and using microfluidic devices. Hence, the benefits of further integration of components into microfluidic devices must be weighed against the increased time investment, cost, and barrier to entry. Another factor contributing to progress in plant microfluidics concerns the reuse and iteration of existing microfluidic device designs, which has allowed useful device designs, such as the RootChip22 and the dual-flow-RootChip,28 to be reused or modified by other research groups.27,36,38–41,51,90,1106.2 Root–electric interaction studies
Interest in the field of root–electric interactions peaked between the 1960s and 1990s, with the discovery of a root electric signature.77,111 More recently, plant bioelectricity gained more attention with a variety of discoveries, including the importance of electric fields in pollination,112 electric field influence on pollen tube targeting, root electrotropic responses113 and the use of external electric fields to reduce root pathogen infection.114 Microfluidic technology has the potential to advance this area. Control and manipulation of microenvironments allow for more controlled electric field application and prevent media evaporation, while the use of very small amount of liquids significantly reduces Joule heating and can help stabilise ion currents.115 Microfluidic technology, together with the standardisation of the instruments and media used to measure electric fields and currents,77 could provide significant novel insights into plant bioelectricity.6.3 Rhizosphere-on-a-chip technology
Beyond bipartite root–microbe interactions, there is a need to develop generalised simplified rhizospheric ecosystems to study soil community dynamics.116 While a microfluidic device has successfully shown soil microbe community dynamics for soil and associated microbes,117 the inclusion of plant roots has not yet been achieved for reasons discussed above. Incorporating plant roots into simplified rhizospheric systems is an important goal to ensure a more representative soil ecosystem inside microfluidic devices, and to investigate the impact of root exudates on soil microbes. Soil structure is a key part of replicating simplified soil ecosystems on-chip. The field of transparent soil particles is a relatively new field that is advancing rapidly, where artificial soils are engineered with varying physiochemical properties to study specific aspects of soil structure.118 Super-absorbent polymers (SAPs) have been used to replicate the structural complexity of soil, while maintaining optically transparency for microscopy.119 While SAP-based systems have not yet been incorporated into microfluidic devices, the relationship between soil structure and the rhizosphere has been investigated using etched structures to replicate soil particles in plant devices,30,31 as well as photolithography for investigating bacterial accumulation in complex porous media.120 The next steps are to incorporate displaceable soil particles into microfluidic devices, which is more representative of edaphic environments. Therefore, further research is needed to build a picture of how the relationship between soil properties, root exudates, and soil microbes functions in simplified ecosystems. This research could uncover how unique biological niches are generated in soil through the relationship between roots, root exudates, and soil properties, helping to clarify global distributions of soil biodiversity.6.4 Root–hyphal interaction studies
Microfluidic technology is well-suited to follow the first few weeks of root–microbe interactions due to focal confinement the root, and potentially the microbial partner(s). Microfluidic technology for root–bacteria interactions23,45 and root–oomycete interactions39,44,47 has allowed researchers to delve deeper into the spatial relationships between roots and these two microbial taxa than classical approaches have allowed. Looking forward, there is a need to design microfluidic devices to study root–hyphal interactions by incorporating filamentous fungi and oomycetes into microfluidic devices.15,116 This would facilitate a new approach to study root–pathogen and root–mutualist interactions in real-time at cellular resolution and could have impacts on plant health and nutrition. While microfluidic devices have been successfully used to study the behaviour of fungal and oomycete spores and hyphae independently of plant roots,121,122 filamentous root interactors have not yet been incorporated into these simplified ecosystems due to plant roots requiring taller channel heights, which prevents focal confinement of smaller soil microbes. Very recently, new microfluidic technologies have been developed that allow the growth of individual hyphae of arbuscular mycorrhizal fungi (AMF) within confined microchannels for the first time.123,124 Progress has also been made in non-invasive monitoring of root–hyphal interactions in roots using transparent media,125 but non-invasive monitoring of the initiation of symbiosis between filamentous microbes and roots is not yet possible due to difficulties in imaging hyphal approach towards a root in real time. To bridge this technological challenge, progress in microfabrication techniques are needed to combine small (<10 μm) microchannels for focal confinement of hyphae with much larger (>200 μm) channels. While this combination of “thick and thin” microchannels has been achieved for focally confining root hairs of A. thaliana,58,68 microchannels for accommodating roots of relevant model plants such as Medicago truncatula and Nicotiana benthamiana require root microchannels significantly thicker than 200 μm. Future progress in microfabrication techniques to accommodate these larger model plants would also allow larger crop plants such as wheat and rice to more easily be incorporated into microfluidic devices. Studying crop plants directly instead of more established model plants, allows species-specific mechanisms to be investigated, and has direct applications to agriculture.6.5 Millifluidic devices
As explored in Box 1, there are various alternatives to standard photolithography for manufacturing microfluidic devices with layer heights taller than 200 μm, each with specific advantages and drawbacks. 3D printing is limited in manufacturing small feature sizes (less than 15 μm), but the cost of high-quality fused deposition modelling (FDM) and stereolithography (SLA) 3D printing machines has decreased rapidly in the last decade, reducing the barrier to entry for the adoption of microfluidic technology. This has resulted in the production of millifluidic devices (devices where the smallest dimension of the device is most easily measured in millimetres), which have facilitated studies into root–rhizosphere interactions. For example, the EcoFAB device,126 has allowed researchers to monitor root–rhizosphere interactions in a repeatable way by publishing guides on fabricating the EcoFAB device.126 This has resulted in modified EcoFAB devices such as the RootTRAPR device127 and Imaging EcoFAB. Therefore, while an exhaustive discussion of plant millifluidic technology is outside the scope of this review, many design principles and manufacturing techniques are applicable to microfluidic technology.6.6 Droplet and digital microfluidic technologies
Beyond microfluidic technology to study roots and soil, we have seen how microfluidic technology has contributed greatly to understanding how plant cells, particularly pollen tubes, function. The next frontier in this field involves translating advances from the fields of droplet and digital microfluidics to provide greater control and analytical ability to study plant cells. The former involves encapsulating (single) cells inside water-in-oil microdroplets, with the latter using electric fields to manipulate discrete microdroplets; both approaches allow precise control of droplets (i.e., fusion, splitting, dispensing etc.) in an automated way. Digital microfluidic platforms have already been used extensively for mammalian cell cultures,128 with only one example of this technique being applied to measure water permeability of isolated plant protoplasts.103 Droplet microfluidic technology has been applied in producing microfluidic bioreactors for the production of economically important plant-derived compounds.129 Automated manipulation and analysis of individual plant protoplasts could assist with high-throughput plant genotyping and phenotyping, paving the way to characterise the phenome of genetic models such as A. thaliana. Moreover, root and droplet microfluidic technology could be integrated, for example to encapsulate plant root exudates to provide a spatially explicit, time-resolved snapshot of the root secretome.6.7 Extracting samples from microfluidic devices
A common challenge in utilising microfluidic technology for the study of plants is the difficulty in extracting samples from inside microfluidic devices for further chemical analysis. Throughout, we have seen different approaches used to overcome this challenge. Chai et al. constructed an openable and resealable device, allowing samples to be extracted at coarse spatial resolution.42 Aufrecht et al. replaced a chemically impermeable glass coverslip with a selectively permeable PETE membrane, allowing on-device analysis of amino acid profiles.30 Maisch et al. separated a fluid extraction channel from the culture channel with a semi-permeable membrane, allowing cell exudates to diffuse through and be collected at specified time points.99 Finally, Dai et al. measured dissolved oxygen and pH inside a microfluidic device using microeletrode sensors, and at regularly spaced intervals collected soil pore water.109 All approaches have varying trade-offs with respect to the kind of data that can be extracted and impact on optical data. These tradeoffs should be carefully considered, and microfluidic devices should be designed from inception to interface with appropriate extraction techniques when required.6.8 Advancing image analysis
Another common challenge is extracting biological inferences from the imaging data collected during microfluidic experiments. Simple observations of behaviours are useful, but quantification of complex behaviours is difficult to achieve manually using classical analytical techniques such as the open source software package ImageJ.130 More advanced algorithms to remove background noise, track biological objects, and quantify network traits have had limited success with image data generated from microfluidic devices, as these specialised algorithms are trained on microscopy data not obtained from microfluidic devices, which can have different image parameters. More generalised artificial intelligence tools such as the Segment Anything Model131 may prove useful in increasing the throughput of image analysis pipelines and make quantitative measures of biological traits possible.6.9 Leaf-on-a-chip
Lastly, a notable absence in the field of microfluidic technology for plant sciences is the study of leaves. While leaves are more easily observed and analysed using classical approaches than their belowground root counterparts, microfluidic technology can and has started to provide useful benefits in studying these organs. For example, a microfluidic device was used to monitor electro-mechanical data across stomata in planta.132 PDMS imprints of leaves have also been used to investigate the effect of leaf topography on the microbiology of the leaf phyllosphere.133 Using microfluidic technology to further understand leaf functioning at the microscale could contributing to a greater understanding of the dynamics of photosynthesis, possibly resulting in efficiency gains in the leaves of genetically transformed crops. Generating a “leaf-on-a-chip” simplified leaf model could assist in selectively breeding for traits to improve leaf function, and aid areas outside of plant sciences by applying what makes leaves so efficient in their water and resource use to engineered products such as valves.1346.10 Accessibility in plant microfluidics
The field of plant microfluidics is now twenty years old. While significant progress has been made during this time, the field is still relatively new. Except for a handful of device designs discussed previously, most microfluidic device designs have been published once only, for a specific use case, without further adoption by the scientific community. As the field matures in the following twenty years of plant microfluidic research, a significant challenge remains of making plant microfluidics widely accessible to the broader plant sciences community through device standardisation. While at first glance, standardisation and customisation may seem at odds, further standardisation and customisation can be achieved in tandem. This could be achieved through further standardisation of existing successful device designs, targeted efforts to increase usability, and a larger focus on making device designs and manufacturing protocols open and accessible. Meanwhile, freedom of customisation can be pursued for novel experimental designs and capabilities. In this way, the next twenty years of plant microfluidic will drive many more advances in plant sciences.Author contributions
Conceptualisation: L. C., C. E. S.; funding acquisition: C. E. S.; supervision: C. E. S.; visualisation: L. C., C. E. S.; writing – original draft: L. C., E. M.; writing – review & editing: L. C., E. M., C. E. S.Conflicts of interest
None declared.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
We acknowledge financial support from the Leverhulme Centre for the Holobiont (RC-2021-058; funding awarded to C. E. S. to support L. C.'s PhD studentship), as well as from the Department of Bioengineering at Imperial College, London.References
- M. R. Appidi, A. N. Bible, D. L. Carper, S. S. Jawdy, R. J. Giannone, R. L. Hettich, J. Morrell-Falvey and P. E. Abraham, Mol. Plant-Microbe Interact., 2022, 35, 639–649 CrossRef CAS PubMed.
- M. Oliva and C. Dunand, New Phytol., 2007, 176, 37–43 CrossRef CAS PubMed.
- W. Xu, G. Ding, K. Yokawa, F. Baluška, Q.-F. Li, Y. Liu, W. Shi, J. Liang and J. Zhang, Sci. Rep., 2013, 3, 1273 CrossRef PubMed.
- D. J. Harrison, A. Manz, Z. Fan, H. Luedi and H. M. Widmer, Anal. Chem., 1992, 64, 1926–1932 CrossRef CAS.
- U. A. Gurkan, D. K. Wood, D. Carranza, L. H. Herbertson, S. L. Diamond, E. Du, S. Guha, J. Di Paola, P. C. Hines, I. Papautsky, S. S. Shevkoplyas, N. J. Sniadecki, V. K. Pamula, P. Sundd, A. Rizwan, P. Qasba and W. A. Lam, Lab Chip, 2024, 24, 1867–1874 RSC.
- C. Chircov and A. M. Grumezescu, Micromachines, 2022, 13, 164 CrossRef PubMed.
- A. Groisman, C. Lobo, H. Cho, J. K. Campbell, Y. S. Dufour, A. M. Stevens and A. Levchenko, Nat. Methods, 2005, 2, 685–689 CrossRef CAS PubMed.
- P. J. Lee, N. C. Helman, W. A. Lim and P. J. Hung, BioTechniques, 2008, 44, 91–95 CrossRef CAS PubMed.
- R. Davidsson, B. Johansson, V. Passoth, M. Bengtsson, T. Laurell and J. Emnéus, Lab Chip, 2004, 4, 488–494 RSC.
- J. Enger, M. Goksör, K. Ramser, P. Hagberg and D. Hanstorp, Lab Chip, 2004, 4, 196–200 RSC.
- C. Sakamoto, N. Yamaguchi and M. Nasu, Appl. Environ. Microbiol., 2005, 71, 1117–1121 CrossRef CAS PubMed.
- J.-M. Ko, J. Ju, S. Lee and H.-C. Cha, Protoplasma, 2006, 227, 237–240 CrossRef CAS PubMed.
- M. Meier, E. M. Lucchetta and R. F. Ismagilov, Lab Chip, 2010, 10, 2147–2153 RSC.
- M. Elitaş, M. Yüce and H. Budak, Analyst, 2017, 142, 835–848 RSC.
- C.-F. Kaiser, A. Perilli, G. Grossmann and Y. Meroz, J. Exp. Bot., 2023, 74, 3851–3863 CrossRef CAS PubMed.
- A. S. Nezhad, Lab Chip, 2014, 14, 3262–3274 RSC.
- N. Yanagisawa, E. Kozgunova, G. Grossmann, A. Geitmann and T. Higashiyama, Plant Cell Physiol., 2021, 62, 1239–1250 CrossRef CAS PubMed.
- L. Philippot, J. M. Raaijmakers, P. Lemanceau and W. H. van der Putten, Nat. Rev. Microbiol., 2013, 11, 789–799 Search PubMed.
- W. Xu, G. Ding, K. Yokawa, F. Baluška, Q.-F. Li, Y. Liu, W. Shi, J. Liang and J. Zhang, Sci. Rep., 2013, 3, 1273 CrossRef PubMed.
- B. D. Gruber, R. F. H. Giehl, S. Friedel and N. von Wiren, Plant Physiol., 2013, 163, 161–179 CrossRef CAS PubMed.
- S. Okumoto, Curr. Opin. Biotechnol., 2010, 21, 45–54 CrossRef CAS PubMed.
- G. Grossmann, W.-J. Guo, D. W. Ehrhardt, W. B. Frommer, R. V. Sit, S. R. Quake and M. Meier, Plant Cell, 2011, 23, 4234–4240 Search PubMed.
- M.-F. Noirot-Gros, S. V. Shinde, C. Akins, J. L. Johnson, S. Zerbs, R. Wilton, K. M. Kemner, P. Noirot and G. Babnigg, Front. Plant Sci., 2020, 11, 408 CrossRef PubMed.
- H. H. Chai, F. Chen, S. J. Zhang, Y. D. Li, Z. S. Lu, Y. J. Kang and L. Yu, Lab Chip, 2019, 19, 2383–2393 RSC.
- G. Dupouy, G. Singh, L. M. Schmidt-Speicher, E. Hoffmann, S. Baudrey, R. Ahrens, A. E. Guber, M. Ryckelynck, E. Herzog, M.-E. Chabouté and A. Berr, in Methods for Plant Nucleus and Chromatin Studies: Methods and Protocols, ed. C. Baroux and C. Tatout, Springer US, New York, NY, 2025, pp. 223–245 Search PubMed.
- H. Jiang, Z. Xu, M. Aluru and L. Dong, Lab Chip, 2014, 14, 1281–1293 RSC.
- V. Lanquar, G. Großmann, J. L. Vinkenborg, M. Merkx, S. Thomine and W. B. Frommer, New Phytol., 2014, 202, 198–208 CrossRef CAS PubMed.
- C. E. Stanley, J. Shrivastava, R. Brugman, E. Heinzelmann, D. van Swaay and G. Grossmann, New Phytol., 2018, 217, 1357–1369 CrossRef CAS PubMed.
- P. Bastian, A. Chavarría-Krauser, C. Engwer, W. Jäger, S. Marnach and M. Ptashnyk, J. Theor. Biol., 2008, 254, 99–109 Search PubMed.
- J. Aufrecht, M. Khalid, C. L. Walton, K. Tate, J. F. Cahill and S. T. Retterer, Lab Chip, 2022, 22, 954–963 RSC.
- I. A. Atoloye, D. Herrera, D. Veličković, C. S. Clendinen, K. Tate, A. Bhattacharjee, J. Aufrecht, T. Zeng, D. Rai and A. Bhowmik, Rhizosphere, 2025, 34, 101099 CrossRef.
- M. Moussus and M. Meier, Lab Chip, 2021, 21, 2557–2564 RSC.
- C.-H. Lin, G.-B. Lee, B.-W. Chang and G.-L. Chang, J. Micromech. Microeng., 2002, 12, 590 Search PubMed.
- R. Su, F. Wang and M. C. McAlpine, Lab Chip, 2023, 23, 1279–1299 RSC.
- W. Busch, B. T. Moore, B. Martsberger, D. L. Mace, R. W. Twigg, J. Jung, I. Pruteanu-Malinici, S. J. Kennedy, G. K. Fricke, R. L. Clark, U. Ohler and P. N. Benfey, Nat. Methods, 2012, 9, 1101–1106 CrossRef CAS PubMed.
- A. M. Jones, J. Å. Danielson, S. N. ManojKumar, V. Lanquar, G. Grossmann and W. B. Frommer, eLife, 2014, 3, e01741 CrossRef PubMed.
- M. Fendrych, M. Akhmanova, J. Merrin, M. Glanc, S. Hagihara, K. Takahashi, N. Uchida, K. U. Torii and J. Friml, Nat. Plants, 2018, 4, 453–459 CrossRef CAS PubMed.
- C. Allan, A. Tayagui, R. Hornung, V. Nock and C.-N. Meisrimler, Front. Plant Sci., 2023, 13, 1040117 CrossRef PubMed.
- C. Allan, Y. Sun, S. C. Whisson, M. Porter, P. C. Boevink, V. Nock and C.-N. Meisrimler, Lab Chip, 2024, 24, 5360–5373 RSC.
- N. F. Keinath, R. Waadt, R. Brugman, J. I. Schroeder, G. Grossmann, K. Schumacher and M. Krebs, Mol. Plant, 2015, 8, 1188–1200 CrossRef CAS PubMed.
- C. Brost, T. Studtrucker, R. Reimann, P. Denninger, J. Czekalla, M. Krebs, B. Fabry, K. Schumacher, G. Grossmann and P. Dietrich, Plant J., 2019, 99, 910–923 Search PubMed.
- H. Chai, C. Feng, K. Ning, W. Sun and L. Yu, Sens. Actuators, B, 2023, 392, 134107 Search PubMed.
- M. G. A. van der Heijden, R. D. Bardgett and N. M. van Straalen, Ecol. Lett., 2008, 11, 296–310 Search PubMed.
- A. Parashar and S. Pandey, Appl. Phys. Lett., 2011, 98, 263703 Search PubMed.
- H. Massalha, E. Korenblum, E. Korenblum, S. Malitsky, O. H. Shapiro and A. Aharoni, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 4549–4554 Search PubMed.
- J. M. Conway, P. J. Martinez, E. D. Wilson, N. M. Del Risco and J. L. Dangl, in Tissue Morphogenesis: Methods and Protocols, ed. C. M. Nelson, Springer US, New York, NY, 2024, pp. 213–228 Search PubMed.
- C. Cohen, F. X. Gauci, X. Noblin, E. Galiana, A. Attard and P. Thomen, Phys. Rev. E, 2025, 111, 024411 CrossRef CAS PubMed.
- M. Watt, W. K. Silk and J. B. Passioura, Ann. Bot., 2006, 97, 839–855 CrossRef PubMed.
- B. Borer, R. Tecon and D. Or, Nat. Commun., 2018, 9, 769 CrossRef PubMed.
- H. Zhang, X. Yan, M. Zhang, Y. Zhao, S. Jiang, Y. Jiang, Y. Wei, Y. Zhang and L. Sun, J. Agric. Food Chem., 2025, 73(48), 30895–30903 CrossRef CAS PubMed.
- S. Liu, S. Strauss, M. Adibi, G. Mosca, S. Yoshida, R. D. Ioio, A. Runions, T. G. Andersen, G. Grossmann, P. Huijser, R. S. Smith and M. Tsiantis, Curr. Biol., 2022, 32, 1974–1985.e3 CrossRef CAS PubMed.
- K. Agarwal, S. K. Mehta and P. K. Mondal, Lab Chip, 2024, 24, 3775–3789 Search PubMed.
- J. Li, M. Notaguchi, I. Kanno and H. Hida, IEEJ Trans. Electr. Electron. Eng., 2025, 20, 1134–1140 CrossRef.
- P. Vinet, V. Audemar, P. Durand-Smet, J.-M. Frachisse and S. Thomine, bioRxiv, 2025, preprint, DOI:10.1101/2025.10.22.683952.
- C. M. Rounds and M. Bezanilla, Annu. Rev. Plant Biol., 2013, 64, 243–265 Search PubMed.
- M. Ghanbari, A. S. Nezhad, C. G. Agudelo, M. Packirisamy and A. Geitmann, J. Biosci. Bioeng., 2014, 117, 504–511 CrossRef CAS PubMed.
- K. Sakai, F. Charlot, T. L. Saux, S. Bonhomme, F. Nogué, J. Palauqui and J. Fattaccioli, Plant Methods, 2019, 15, 79 Search PubMed.
- G. Singh, D. Pereira, S. Baudrey, E. Hoffmann, M. Ryckelynck, A. Asnacios and M.-E. Chabouté, Plant J., 2021, 108, 303–313 Search PubMed.
- J. H. Williams, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 11259–11263 CrossRef CAS PubMed.
- F. Bou Daher, Y. Chebli and A. Geitmann, Plant Cell Rep., 2009, 28, 347–357 CrossRef CAS PubMed.
- R. Palanivelu and T. Tsukamoto, WIREs Dev. Biol., 2012, 1, 96–113 Search PubMed.
- W. J. Uwate, J. Lin, K. Ryugo and V. Stallman, Can. J. Bot., 1982, 60, 98–104 Search PubMed.
- A. S. Nezhad, M. Naghavi, M. Packirisamy, R. B. Bhat and A. Geitmann, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 8093–8098 Search PubMed.
- C. G. Agudelo, A. Sanati, M. Ghanbari, M. Packirisamy and A. Geitmann, J. Micromech. Microeng., 2012, 22, 115009 CrossRef.
- C. G. Agudelo, A. S. Nezhad, A. S. Nezhad, M. Ghanbari, M. Naghavi, M. Packirisamy and A. Geitmann, Plant J., 2013, 73, 1057–1068 Search PubMed.
- A. Sanati Nezhad, M. Ghanbari, C. G. Agudelo, M. Naghavi, M. Packirisamy, R. B. Bhat and A. Geitmann, Biomed. Microdevices, 2014, 16, 23–33 Search PubMed.
- COMSOL Multiphysics® v. 6.4, https://www.comsol.com, COMSOL AB, Stockholm, Sweden Search PubMed.
- N. Yanagisawa, N. Sugimoto, H. Arata, T. Higashiyama and Y. Sato, Sci. Rep., 2017, 7, 1403 CrossRef PubMed.
- A. K. Yetisen, L. Jiang, J. R. Cooper, J. Cooper, Y. Qin, R. Palanivelu and Y. Zohar, J. Micromech. Microeng., 2011, 21, 054018 CrossRef.
- M. Horade, M. M. Kanaoka, M. Kuzuya, T. Higashiyama and N. Kaji, RSC Adv., 2013, 3, 22301–22307 RSC.
- N. Yanagisawa and T. Higashiyama, Biomicrofluidics, 2018, 12(2), 024113 Search PubMed.
- A. Nezhad, M. Packirisamy and A. Geitmann, Plant J., 2014, 80, 185–195 CrossRef PubMed.
- N. Yanagisawa, E. Kozgunova and T. Higashiyama, RSC Adv., 2021, 11, 27011–27018 RSC.
- C. Agudelo, M. Packirisamy and A. Geitmann, Sci. Rep., 2016, 6, 19812 Search PubMed.
- C. Hu, H. Vogler, M. Aellen, N. Shamsudhin, B. Jang, J. T. Burri, N. Läubli, U. Grossniklaus, S. Pané and B. J. Nelson, Lab Chip, 2017, 17, 671–680 Search PubMed.
- K. S. Elvira, X. C. i Solvas, R. C. R. Wootton and A. J. deMello, Nat. Chem., 2013, 5, 905–915 CrossRef CAS PubMed.
- E. Moratto and G. Sena, Bioelectricity, 2023, 5, 47–54 CrossRef.
- A. S. Nezhad, M. Naghavi, M. Packirisamy, R. B. Bhat and A. Geitmann, Lab Chip, 2013, 13, 2599–2608 Search PubMed.
- R. Benkert, G. Obermeyer and F.-W. Bentrup, Protoplasma, 1997, 198, 1–8 Search PubMed.
- J. T. Burri, H. Vogler, N. F. Läubli, C. Hu, U. Grossniklaus and B. J. Nelson, New Phytol., 2018, 220, 187–195 Search PubMed.
- W. Kammerer and D. J. Cove, Mol. Gen. Genet., 1996, 250, 380–382 Search PubMed.
- M. J. Prigge and M. Bezanilla, Development, 2010, 137, 3535–3543 Search PubMed.
- T. C. Merkel, V. I. Bondar, K. Nagai, B. D. Freeman and I. Pinnau, J. Polym. Sci., Part B: Polym. Phys., 2000, 38, 415–434 Search PubMed.
- C. S. Bascom, S.-Z. Wu, K. Nelson, J. Oakey and M. Bezanilla, Plant Physiol., 2016, 172, 28–37 Search PubMed.
- G. I. Jenkins and D. J. Cove, Planta, 1983, 157, 39–45 Search PubMed.
- F. Furt, K. Lemoi, E. Tüzel and L. Vidali, BMC Plant Biol., 2012, 12, 70 Search PubMed.
- E. Kozgunova and G. Goshima, Sci. Rep., 2019, 9, 15182 CrossRef PubMed.
- C. Grierson, E. Nielsen, T. Ketelaarc and J. Schiefelbein, Arabidopsis Book, 2014, vol. 12, DOI:10.1199/tab.0172.
- K. Yi, B. Menand, E. Bell and L. Dolan, Nat. Genet., 2010, 42, 264–267 CrossRef CAS PubMed.
- P. Denninger, A. Reichelt, V. A. F. Schmidt, D. G. Mehlhorn, L. Y. Asseck, C. E. Stanley, N. F. Keinath, J.-F. Evers, C. Grefen and G. Grossmann, Curr. Biol., 2019, 29, 1854–1865.e5 Search PubMed.
- S. Gilroy and D. L. Jones, Trends Plant Sci., 2000, 5, 56–60 CrossRef CAS PubMed.
- D. J. Gage, Microbiol. Mol. Biol. Rev., 2004, 68, 280–300 CrossRef CAS PubMed.
- A. H. N. Griffis, N. R. Groves, X. Zhou and I. Meier, Front. Plant Sci., 2014, 5, 129 Search PubMed.
- S. Zhang, J. Liu, X. Xue, K. Tan, C. Wang and H. Su, Biochem. Biophys. Res. Commun., 2019, 519, 783–789 Search PubMed.
- H. Wu, W. Liu, Q. Tu, N. Song, L. Li, J. Wang and J. Wang, Microfluid. Nanofluid., 2011, 10, 867–876 CrossRef CAS.
- T. Seidel, P. J. Artmann, I. Gkekas, F. Illies, A.-L. Baack and M. Viefhues, Plants, 2024, 13, 295 CrossRef CAS PubMed.
- P. Durand-Smet, T. A. Spelman, E. M. Meyerowitz and H. Jönsson, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 17399–17408 CrossRef CAS PubMed.
- S. Xu, Z. Sun, L. Liu, H. Liu, Y. Yang, S. Zhang, Y. Li, N. Bao, Y. Zhang and L. Sun, Processes, 2022, 10, 2507–2508 CrossRef CAS.
- J. Maisch, K. Kreppenhofer, S. Büchler, C. Merle, S. Sobich, B. Görling, B. Luy, R. Ahrens, A. E. Guber and P. Nick, J. Plant Physiol., 2016, 200, 28–34 Search PubMed.
- B. Zaban, W. Liu, X. Jiang and P. Nick, Sci. Rep., 2014, 4, 5852 Search PubMed.
- J. Park, D. Kurihara, T. Higashiyama and H. Arata, Sens. Actuators, B, 2014, 191, 178–185 CrossRef CAS.
- K. Gooh, M. Ueda, K. Aruga, J. Park, H. Arata, T. Higashiyama and D. Kurihara, Dev. Cell, 2015, 34, 242–251 CrossRef CAS PubMed.
- P. T. Kumar, F. Toffalini, D. Witters, S. Vermeir, F. Rolland, M. L. A. T. M. Hertog, B. M. Nicolaï, R. Puers, A. Geeraerd and J. Lammertyn, Sens. Actuators, B, 2014, 199, 479–487 CrossRef CAS.
- Z. Yu, C. R. Boehm, J. M. Hibberd, C. Abell, J. Haseloff, S. J. Burgess and I. Reyna-Llorens, PLoS One, 2018, 13, e0196810 CrossRef PubMed.
- T. Galkovskyi, Y. Mileyko, A. Bucksch, B. Moore, O. Symonova, C. A. Price, C. N. Topp, A. S. Iyer-Pascuzzi, P. R. Zurek, S. Fang, J. Harer, P. N. Benfey and J. S. Weitz, BMC Plant Biol., 2012, 12, 116 Search PubMed.
- H. Jiang, X. Wang, T. M. Nolan, Y. Yin, M. R. Aluru and L. Dong, in 2017 IEEE 12th International Conference on Nano/Micro Engineered and Molecular Systems (NEMS), 2017, pp. 756–760 Search PubMed.
- X. Wang, H. Jiang, Y. Wang and L. Dong, in 2017 IEEE 12th International Conference on Nano/Micro Engineered and Molecular Systems (NEMS), 2017, pp. 398–401 Search PubMed.
- M. A. Buttkewitz, C. Heuer and J. Bahnemann, Curr. Opin. Biotechnol., 2023, 83, 102978 Search PubMed.
- H. Dai, B. Wu, B. Chen, B. Ma and C. Chu, Environ. Sci. Technol., 2022, 56, 9075–9082 Search PubMed.
- A. Rizza, B. Tang, C. E. Stanley, G. Grossmann, M. R. Owen, L. R. Band and A. M. Jones, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e1921960118 Search PubMed.
- V. Ghildiyal, C. M. Altaner, B. Heffernan and M. C. Jarvis, Curr. For. Rep., 2025, 11, 7 Search PubMed.
- D. Clarke, H. Whitney, G. Sutton and D. Robert, Science, 2013, 340, 66–69 Search PubMed.
- M. Salvalaio, N. Oliver, D. Tiknaz, M. Schwarze, N. Kral, S.-J. Kim and G. Sena, Plant Physiol., 2022, 188, 1604–1616 CrossRef CAS PubMed.
- E. Moratto, Z. Tang, T. O. Bozkurt and G. Sena, Sci. Rep., 2024, 14, 19993 CrossRef CAS PubMed.
- Y.-S. Sun, Sensors, 2017, 17, 2048 CrossRef PubMed.
- C. E. Stanley, G. Grossmann, X. C. i Solvas and A. J. deMello, Lab Chip, 2016, 16, 228–241 RSC.
- P. M. Mafla-Endara, C. Arellano-Caicedo, K. Aleklett, M. Pucetaite, P. Ohlsson and E. C. Hammer, Commun. Biol., 2021, 4, 1–12 Search PubMed.
- B. Wang, H. Hou and Z. Zhu, Geotech. Res., 2021, 8, 130–138 CrossRef.
- L. Paré, C. Banchini, C. Hamel, L. Bernier and F. Stefani, Symbiosis, 2022, 88, 61–73 CrossRef.
- D. Scheidweiler, A. D. Bordoloi, W. Jiao, V. Sentchilo, M. Bollani, A. Chhun, P. Engel and P. de Anna, Nat. Commun., 2024, 15, 191 CrossRef CAS PubMed.
- A. J. Clark, E. Masters-Clark, E. Moratto, P. Junier and C. E. Stanley, Biomicrofluidics, 2024, 18, 054109 CrossRef PubMed.
- D. Sarkar, Y. Sun, A. Tayagui, R. Adams, A. Garrill and V. Nock, in 2022 IEEE 35th International Conference on Micro Electro Mechanical Systems Conference (MEMS), 2022, pp. 884–887 Search PubMed.
- E. C. Hammer, C. Arellano-Caicedo, P. M. Mafla-Endara, E. T. Kiers, T. Shimizu, P. Ohlsson and K. Aleklett, Fungal Ecol., 2024, 67, 101302 CrossRef.
- F. Richter, M. Calonne-Salmon, M. G. A. van der Heijden, S. Declerck and C. E. Stanley, Lab Chip, 2024, 24, 1930–1946 Search PubMed.
- J. McGaley, B. Schneider and U. Paszkowski, J. Microsc., 2025, 297, 289–303 CrossRef PubMed.
- J. Gao, J. Sasse, K. M. Lewald, K. Zhalnina, L. T. Cornmesser, T. A. Duncombe, Y. Yoshikuni, J. P. Vogel, M. K. Firestone and T. R. Northen, J. Visualized Exp., 2018, e57170 Search PubMed.
- P. Suwanchaikasem, A. Idnurm, J. Selby-Pham, R. Walker and B. A. Boughton, Plant Methods, 2022, 18, 46 CrossRef CAS PubMed.
- B. Gumuscu, Lab Chip, 2025, 25, 3297–3313 RSC.
- T. Finkbeiner, C. Manz, M. L. Raorane, C. Metzger, L. Schmidt-Speicher, N. Shen, R. Ahrens, J. Maisch, P. Nick and A. E. Guber, Protoplasma, 2022, 259, 173–186 CrossRef CAS PubMed.
- C. A. Schneider, W. S. Rasband and K. W. Eliceiri, Nat. Methods, 2012, 9, 671–675 CrossRef CAS PubMed.
- A. Kirillov, E. Mintun, N. Ravi, H. Mao, C. Rolland, L. Gustafson, T. Xiao, S. Whitehead, A. C. Berg, W.-Y. Lo, P. Dollár and R. Girshick, arXiv, 2023, preprint, arXiv:2304.02643, DOI:10.48550/arXiv.2304.02643.
- V. B. Koman, T. T. S. Lew, M. H. Wong, S.-Y. Kwak, J. P. Giraldo and M. S. Strano, Lab Chip, 2017, 17, 4015–4024 RSC.
- R. Soffe, M. Bernach, M. N. P. Remus-Emsermann and V. Nock, Sci. Rep., 2019, 9, 14420 CrossRef PubMed.
- L. Jingmin, L. Chong, X. Zheng, Z. Kaiping, K. Xue and W. Liding, PLoS One, 2012, 7, e50320 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |



