Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Point of care molecular cancer diagnostics
Seemesh Bhaskar *abc,
Saurabh Umrao
*abc,
Saurabh Umrao bcd,
Han Keun Lee
bcd,
Han Keun Lee ab,
Joseph Tibbs
ab,
Joseph Tibbs bd,
Amanda Bacon
bd,
Amanda Bacon bd,
Skye Shepherd
bd,
Skye Shepherd bd,
Takhmina Ayupova
bd,
Takhmina Ayupova bcd,
Fatma Uysal Ciloglu
bcd,
Fatma Uysal Ciloglu be,
Leyang Liu
be,
Leyang Liu ab,
Anqi Tan
ab,
Anqi Tan bd,
Wang-Chien Chen
bd,
Wang-Chien Chen ab,
My Thi Tra Nguyen
ab,
My Thi Tra Nguyen abp,
Maria Grace Scannell
abp,
Maria Grace Scannell ab,
Ugur Aygun
ab,
Ugur Aygun fg,
Ugur Parlatan
fg,
Ugur Parlatan f,
Catherine Zhang
f,
Catherine Zhang d,
Manish Kohli
d,
Manish Kohli h,
Guy R. Adami
h,
Guy R. Adami ij,
Wali Badar
ij,
Wali Badar k,
Ron C. Gaba
k,
Ron C. Gaba kl,
Aaron Mansfield
kl,
Aaron Mansfield m,
Joel Schwartz
m,
Joel Schwartz i,
Wang Xing
i,
Wang Xing bcdno,
Utkan Demirci
bcdno,
Utkan Demirci f and
Brian T. Cunningham
f and
Brian T. Cunningham *abcdno
*abcdno
aDepartment of Electrical and Computer Engineering, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA. E-mail: seemeshb@illinois.edu; bcunning@illinois.edu
bNick Holonyak Jr. Micro and Nanotechnology Laboratory, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA
cCarl R. Woese Institute for Genomic Biology, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA
dDepartment of Bioengineering, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA
eDepartment of Biomedical Engineering, Erciyes University, Kayseri, 38039, Türkiye
fBioAcoustic MEMS in Medicine BAMM Laboratory, Canary Center at Stanford for Cancer Early Detection, Stanford School of Medicine, Department of Radiology, Stanford University, Palo Alto, CA 94304, USA
gDepartment of Electrical and Electronics Engineering, Koç University, Istanbul, Turkiye
hDepartment of Internal Medicine and Huntsman Cancer Institute, University of Utah, Salt Lake City, UT 84112, USA
iDepartment of Oral Medicine and Diagnostics, College of Dentistry, University of Illinois Chicago, Chicago, IL 60612, USA
jUniversity of Illinois Cancer Center, University of Illinois Chicago, Chicago, IL 60612, USA
kDepartment of Radiology, University of Illinois at Chicago, 1740 West Taylor St, Chicago, IL, USA
lDepartment of Pathology, University of Illinois at Chicago, 1740 West Taylor St, Chicago, IL, USA
mMedical Oncology, Mayo Clinic, Rochester, MN 55905, USA
nDepartment of Chemistry, University of Illinois at Urbana-Champaign, Urbana, IL 61801, USA
oCancer Center at Illinois, Urbana, IL 61801, USA
pVinUni-Illinois Smart Health Center, VinUniversity, Hanoi 100000, Vietnam
First published on 10th February 2026
Abstract
Cancer remains one of the leading global health burdens, demanding swift, precise, and accessible diagnostic responses that can overcome the limitations of diagnostic tests performed in centralized laboratory settings. Point-of-care (POC) molecular diagnostics are emerging as transformative tools, enabling decentralized testing across hospitals, outpatient clinics, oncology practices, and dental practices, thereby accelerating clinical decision-making to improve patient outcomes. This critical review provides a comprehensive perspective on the clinical requirements that shape the design of POC diagnostics, where several case studies based on clinical scenarios serve as representative examples. We detail advancements in POC-compatible sample preparation, nucleic acid and protein biomarker detection, and imaging modalities, highlighting their translational value for real-world oncology applications. In addition, we analyze the global funding landscape, intellectual property, and regulatory frameworks that influence the transition of emerging technologies from academic research to commercial markets. Key enabling technologies, ranging from microfluidics and lab-on-a-chip platforms to nanomaterial-enhanced biosensors and smartphone-integrated devices, are critically evaluated alongside their limitations and challenges. Finally, we project future directions for POC molecular diagnostics, emphasizing interdisciplinary innovation, equitable access, and integration into precision medicine. By synthesizing clinical, technological, and regulatory perspectives, this review underscores the pivotal role of POC molecular diagnostics in shaping the future of cancer detection, therapy selection, therapy effectiveness monitoring, and personalized medicine.
1. Introduction
Prior to the rise of digital photography after 2010, taking pictures involved sending film to be developed in a remote laboratory. The limited number of exposures within a roll of film, combined with the need to personally deliver the film to a collection point, followed by waiting for up to several days to see the finished print had the natural consequence of limiting the number of photos we would take, while reducing the quality of the images due to the long time duration between clicking the shutter and seeing the outcome of our artistic vision. The costs of photography, both economic and in terms of effort, resulted from the friction and costs of the lab-based film processing paradigm. These costs were overcome through the availability of digital photography on mobile devices, allowing us, for the first time, to see our results immediately with nearly zero cost. The result was a greater degree of experimentation when there were few limits on the number of photos we could generate. A similar situation exists today in the realm of molecular diagnostic testing, which relies almost entirely upon gathering samples of bodily fluids that are subsequently shipped to a laboratory facility for processing.1,2 The facilities employ highly trained technicians who are capable of performing workflows that involve manual interactions of the sample, reagents, and instruments.3,4 The lab personnel strictly follow thoroughly documented sample preparation and testing protocols, where expensive and highly automated instruments have the capacity for performing many tests in parallel with high throughput.5,6 The laboratory-based approach is applied to several classes of molecular biomarkers for disease that include proteins, metabolites, and a variety of nucleic acids whose presence, concentration, and identity are used to inform clinicians about decisions that direct the path of patients through treatment options.7–9The accuracy and validity of medical diagnostic tests are extraordinarily important, as physicians rely on them for making a broad range of clinical decisions.10,11 Laboratory-based tests are highly trusted due to the quality control inherent in the regulatory approval process. Nonetheless, technology advances in microfluidic sample processing of clinical samples, automation of reagent handling within disposable cartridges, mass-manufacturable biosensing transducers, simplified molecular biology methods, and small/inexpensive detection instruments are advancing to the extent that the vision for performing molecular diagnostic tests at the “point of care” (POC) is becoming a reality.9,12–14 Particularly in the field of pathogen detection, POC tests can now claim several technologically and commercially successful examples, where detection of a pathogen-specific nucleic acid sequence, antigen, or antibody can be used to immediately prescribe an antibiotic, indicate an infectious state for the sake of quarantine, or determine if a person's immune system has responded to a prior exposure.15–18
Since the term “cancer liquid biopsy” was first introduced in 2010 to describe the analysis of circulating tumor cells (CTCs),19–21 the concept is now fully an aspect of mainstream clinical diagnostics that includes detection of a wide variety of nucleic acids, proteins, extracellular vesicles, cells, and metabolites.22 The attractive aspect of liquid biopsy compared to conventional needle biopsy is greatly reduced invasiveness for collecting a specimen.23,24 Blood plasma, urine, stool, saliva, oral rinse, tear fluid, and tissue swabs have all been found to contain cancer-relevant molecular biomarkers that have been released from tumor cells or that have been generated by other cells in response to the presence of tumor cells. As these sample types are much simpler to collect, compared to removing a volume of tissue from an internal organ, liquid biopsy offers the opportunity to repeat tests at frequent intervals to gain a deeper understanding of physiological processes occurring during and after treatment.25,26 Importantly, cancer treatment is most effective when it is diagnosed and treated at the earliest possible stage, when the size of tumors and the corresponding number of cancer cells are the lowest. As a result, cancer liquid biopsy is technically challenging because the molecular biomarkers found in easily-collected samples (such as circulating blood) are present at very low concentrations, that may represent only a few molecules in the test sample.27–30 With the advent of next-generation nucleic acid sequencing (NGS) and the steady reduction in the cost of sequencing, an accelerated understanding of the cancer genome is underway.31,32 Genomic studies that compare cancerous and healthy tissues are shedding light upon the complexity of cancer genomes, where single base mutations, copy number variants, methylation state, and genomic rearrangements that underly the cell's ability to evade the immune system, proliferate, and metastasize are being catalogued for every cancer type and stage.33–35 The genomic sequencing tools are being extended to not only catalog DNA, but also messenger RNA (mRNA), micro-RNA (miRNA), long noncoding RNA (lnc-RNA), pathogenic nucleic acids (such as from cancer-associated viruses) and proteins.36–39 A common pathway for discovering relevant biomarkers for liquid biopsy is to seek the presence of disease-relevant molecules in noninvasively obtained bodily fluids that are also present (or highly elevated) in cancerous tissue.40–42 The NGS technologies cast a broad net to characterize thousands of nucleic acid sequences and hundreds of miRNAs or proteins.27,43,44 However, in many circumstances a small subset of molecules is found to be most informative as disease biomarkers that, when considered as a group, can assist in predicting clinical outcomes or suggesting a specific treatment.45–48
An important well-known limitation of cancer liquid biopsy is that detection of a particular set of molecules in a bodily fluid does not indicate the location of a cancerous tumor, thus posing a major challenge for early cancer diagnosis, particularly for people with no symptoms.29,49 Thus, cancer molecular detection can be expected to be complemented by medical imaging.50–52 Cancer liquid biopsy is currently dominated by a laboratory-based testing paradigm. Detection of nucleic acid biomarkers using polymerase chain reaction (PCR), quantitative PCR (q-PCR), quantitative reverse transcriptase PCR (qRT-PCR), and droplet digital PCR (ddPCR) can identify the presence and concentration of a small number of nucleic acid target sequences, while NGS and various extensions (RNA-seq, miRNA-seq, ChIP-seq (Chromatin Immunoprecipitation Sequencing), ATAC-seq (Assay for Transposase-Accessible Chromatin Sequencing), CLIP-seq (Crosslinking Immunoprecipitation Sequencing)), and proximity probe methods for detecting proteins (O-Link, NULISA, SomaLogic) are available from laboratory testing services.42,53–55 Similarly, well established laboratory-based platforms are available for characterizing protein biomarkers that include highly automated Enzyme Linked Immunosorbent Assays (ELISA) and multiplexed assays using polymer beads, fluorescent tags, and chemiluminescent tags (for example, Luminex or MesoScale Discovery).56–58 Due to the complexity of the assay workflows (which may take up to several days to complete), high cost and complexity of the detection instruments, the need for precisely managing sample manipulations with reagents, and in some cases the need for large-scale data/computation resources with complex bioinformatics-based data interpretation, these approaches are not amenable for implementing in POC scenarios, such as an oncologist's office.59–62 The cost of laboratory-based diagnostic tests can be prohibitive.63–69
The focus of this critical review is to consider the needs and technological advances for POC cancer molecular diagnostics. We are focused upon tests that could be performed in a specialty physician's office (such as an oncologist, oral surgeon, gynecologist) or a health clinic, rather than tests that would be performed by a patient in their home. Since cancer is generally a disease that develops and advances gradually, one can legitimately ask why any POC cancer diagnostic tests would be necessary, as getting an answer within 30 minutes during a doctor's office visit would not be meaningful for treating a medical condition that is not acute in nature.66,70–72 In our review, we will describe several detailed representative clinically important scenarios in which a POC test is advantageous for the sake of efficient treatment and improving patient outcomes. For example, a patient may be diagnosed (through medical imaging or histopathology imaging of a needle biopsy) with cancer in a specific tissue, but selection of the most effective FDA-approved drug among many options can be guided by knowledge of the presence of a specific genomic variant. Similarly, genomic variants can be associated with more aggressive forms of cancer that can guide treatment towards immediate surgery rather than watchful waiting. Particularly in situations in which there is potential for a cancer to be aggressive, waiting for 4–8 weeks for comprehensive genomic testing may lead to increased risk, as the opportunity to begin effective therapy immediately can be performed before a tumor has an opportunity to reach an advanced stage or to metastasize. Once therapy is underway, a need exists for frequent monitoring of its effectiveness, as molecular biomarkers that decrease in concentration in response to therapy can indicate a promising approach, while the emergence of a new biomarker can indicate the onset of drug resistance and the need to change to a different therapy. Cancer survivors are concerned about recurrence, for whom frequent monitoring would be acceptable, even with some degree of false positive results.66,69,73 POC tests may also be effective for patients who present a visually observable lesion, where an accurate rule in/out test could be used to indicate the need for more comprehensive follow-up. An inherent characteristic of POC molecular diagnostic tests is that they will be performed one at a time, on demand, to obtain an answer in a short time period for the sake of making very specific clinical decisions.74–76 Thus, POC tests are likely to be focused upon detection of only a handful of biomarkers and considerations of cost (for the detection instrument and for single-use disposable microfluidic cartridges) and sample-to-answer time outweigh the need for large-scale multiplexing and assay throughput.77–79 Nonetheless, for broad adoption, POC tests must still match the detection limits, clinical sensitivity/selectivity,80 accuracy,81 and reproducibility82–84 of their laboratory-based counterparts to ensure that the results are valid and actionable.
While several prior reviews have provided comprehensive overviews of POC molecular diagnostic technologies,1,9,85–92 they have largely focused on assay formats, detection chemistries, or platform-level performance metrics in isolation. In contrast, the present review is explicitly structured around a clinically grounded, decision-driven framework tailored to oncology, in which representative cancer case studies are used to define the real-world constraints that shape POC design requirements. Rather than surveying technologies in the abstract, we begin with concrete clinical scenarios, such as early lesion triage, therapy selection based on actionable genomic variants, and longitudinal monitoring of treatment response, to identify the specific biomarker classes, sensitivity requirements, turnaround times, and workflow limitations that meaningfully influence patient management. A further distinguishing feature of this review is the deliberate integration of engineering considerations such as sample type, sample preparation burden, automation, assay complexity, and instrument cost, along with clinical decision points that occur at the level of the physician's office or specialty clinic. By explicitly linking these dimensions, we critically evaluate not only analytical performance, but also translational feasibility, scalability, and regulatory readiness in the context of oncology-specific use cases. In addition, this review places emphasis on pathways to clinical adoption, including regulatory precedents, intellectual property landscapes, funding mechanisms, and early commercialization efforts, thereby extending beyond laboratory demonstrations to assess what is realistically achievable at the POC. Collectively, this perspective positions the review as a bridge between cancer biology, microfluidic and biosensing technologies, and real-world clinical translation, complementing the existing literature while addressing gaps specific to POC cancer molecular diagnostics.
In addition to medically-relevant motivations to develop POC cancer diagnostic tests, there are several “human-centered” considerations. For example, patients experience considerable anxiety while waiting for test results to arrive.93–95 When a test is prescribed, many patients fail to make follow-up appointments for sample collection, potentially ignoring a problem or putting it on a low priority compared to other responsibilities. Large medical centers and research hospitals currently offer the most sophisticated testing and test analysis capabilities, requiring patients to take time from work for travel, which is especially a barrier for people living in rural areas. Many people lack health insurance or have insurance with large deductibles, bringing concerns about health care coverage for diagnostic tests costing hundreds to thousands of dollars.96,97 Fear of cancer and fear of unfamiliar doctors lead many patients to avoid taking extra steps for diagnostic tests. Overall, there are many societal, personal, and financial barriers that lead to healthcare disparities. These issues could be partially addressed through the availability of low cost, rapid, noninvasive, and actionable molecular diagnostic tests that can be performed and interpreted during a visit to a physician's office.
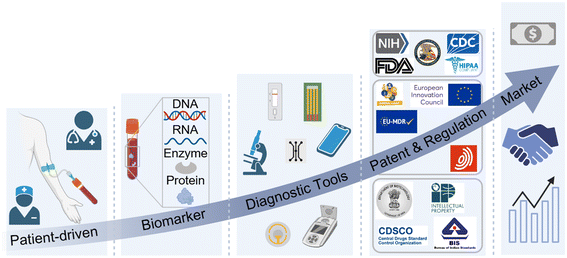
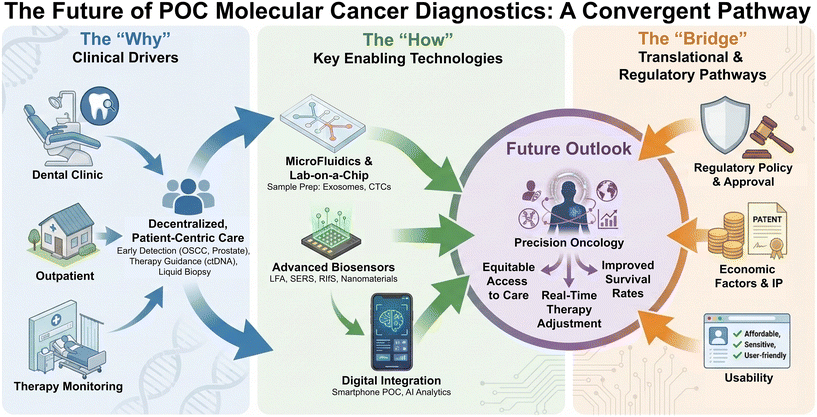
In light of these observations, here, we aim to provide a multidisciplinary perspective on the rapidly evolving field of cancer diagnostics, with a particular emphasis on approaches that are either already implemented at the POC or are advancing toward clinical translation. To anchor the discussion in real-world clinical needs, we first highlight several representative case studies to illustrate both the promise and the persistent gaps in early detection strategies (Fig. 1). Building upon this foundation, we present a structured literature survey that consolidates key developments in liquid biopsy, sample preparation, and biomarker targeting, supported by selected figures to illustrate technological advances and limitations. A dedicated section reviews technologies and patents that have achieved regulatory milestones or that have reached stages close to commercialization, offering concrete examples of translation beyond the laboratory. To complement this, we include a focused discussion on regulatory approvals and logistics, examining FDA, PMDA, and EMA case studies to provide readers with a clear view of the pathways and challenges involved in bringing POC diagnostic tools to market. We then turn to the role of funding mechanisms, presenting insights into grant proposals from both academic and industrial settings, while also drawing attention to the differing priorities and funding landscapes across countries. The review then summarizes current limitations and challenges, with comparative perspectives on technical, clinical, and systemic barriers, before turning the discussion to the future scope of the field. This forward-looking perspective, informed by inputs from clinical experts, identifies opportunities for innovation, integration, and broader accessibility of cancer diagnostics worldwide. Finally, we conclude with reflections on how collective efforts across science, medicine, and industry can accelerate the transition from discovery to clinical translation.
2. Clinical perspectives on cancer diagnostics
2.1. Unified clinical needs and constraints for point-of-care oncology diagnostics
Across diverse cancer types and clinical contexts, the motivation for POC molecular diagnostics arises from a common set of clinical constraints that limit the effectiveness of centralized laboratory testing. These shared limitations shape the functional requirements for POC systems and define the boundary conditions within which new technologies must operate to achieve clinical impact. While the specific biomarkers and decision points vary by disease, several unifying clinical needs recur across oncology.Turnaround time remains one of the most critical constraints in cancer diagnostics. In many clinical scenarios, including therapy selection, disease recurrence assessment, and longitudinal monitoring, results are required on the scale of hours to days rather than weeks. Delays associated with centralized molecular testing can postpone treatment initiation, complicate care coordination, and increase patient anxiety. Consequently, clinically meaningful POC diagnostics must deliver actionable molecular information within a timeframe aligned to real-world clinical decision windows, rather than merely achieving analytical sensitivity under ideal laboratory conditions. Invasiveness and patient burden further motivate the transition toward POC-compatible workflows. Tissue biopsies, while diagnostically informative, are often invasive, resource-intensive, and impractical for repeated sampling. This has driven increasing reliance on minimally invasive biofluids such as blood, saliva, urine, and other accessible matrices. However, the shift toward non-invasive sampling introduces additional challenges, including low analyte abundance, matrix interference, and biological variability, all of which impose stringent performance requirements on POC technologies.
Cost, accessibility, and infrastructure constraints play a decisive role in determining whether diagnostic innovations can be adopted beyond specialized centers. High capital costs, complex instrumentation, and reliance on centralized laboratory infrastructure limit access in community clinics, outpatient settings, and resource-constrained environments. POC diagnostics must therefore balance performance with affordability, portability, and robustness, enabling deployment in decentralized settings without sacrificing clinical reliability. Likewise, sample volume and quality limitations are particularly acute in POC contexts. Small-volume samples, variable collection conditions, and limited pre-analytical processing can significantly impact assay performance. Unlike centralized laboratories, where extensive sample preparation and quality control are routine, POC systems must accommodate variability while maintaining reproducibility. This necessitates integrated sample preparation, contamination control, and error-tolerant assay designs. Workflow complexity and operator dependence represent additional barriers to translation. Diagnostic workflows that require multiple manual steps, specialized training, or subjective interpretation are poorly suited for POC deployment. Clinically viable POC platforms must minimize hands-on time, reduce operator variability, and ideally integrate automation to ensure consistent performance across users and settings. Moreover, clinical actionability serves as a unifying criterion across all cancer types. High analytical sensitivity alone does not guarantee clinical utility unless results directly inform timely, actionable clinical decisions, including diagnostic classification, treatment selection, or disease monitoring.
Taken together, these shared clinical needs underscore that successful POC oncology diagnostics require more than miniaturized assays. They demand systems-level integration of sample handling, detection, workflow design, and decision relevance. Within this common framework, individual cancer types present distinct biomarker constraints and decision points, which are discussed in the following disease-specific sections on oral cancer, liver cancer prostate cancer, and lung cancer.
2.2. POC diagnostics for oral squamous cell carcinoma
Oral squamous cell carcinomas (OSCC) make up well over 90% of cancers of the tongue, floor of mouth, gums, and lips, and in Asian countries are one of the most common cancers.98 Despite occurring as chronic visible lesions in the oral cavity, OSCC is not diagnosed till late stages in about half of patients at which time it responds poorly to current treatments.99–102 This occurs despite the fact that 65% of adult Americans go to the dentist each year and having dental problems is a risk factor for oral cancer.103 The gold standard test, scalpel or punch biopsy followed by histopathology for evaluation of the tissue, is over 90% accurate in identifying early oral cancers when it is used104 but this protocol is one of several reasons for the dismal rate of early OSCC detection. Surgical biopsies require skill in knowing where to sample and, as a result, patients must be referred by their dentist to an oral surgeon. A second barrier is that often patients referred to an oral surgeon do not have their lesion surgically biopsied on their first consultation visit in part due to 3rd party requirements for reimbursement.105 To be reimbursed for a surgical biopsy with standard histopathology, the patient must come back for a second appointment for this expensive, labor-intensive procedure. The exception is if the oral surgeon is confident based on visual inspection that the lesion is cancerous, in which case a battery of tests are ordered at once including a surgical biopsy, CT scans and PET/CT scans to stage the cancer based on size, location, and presence of tumor in lymph nodes and distal body sites.101 An additional barrier to early diagnosis is that when offered the pain and discomfort of a surgical biopsy versus the “watch and wait” strategy of doing nothing, many patients prefer the latter. This sets up a situation where various barriers, some from the patient and some from the reimbursement system, make oral cancer detection difficult.A POC diagnostic for OSCC should leverage noninvasive sampling to allow broader clinical use, require minimal setup without anesthesia or surgery, and integrate smoothly into routine dental or clinic workflows. Next, the test and the instrument on which it was run would have to be simple and inexpensive to encourage reimbursement by third parties. This underscores the importance of developing biomarker panels and detection modalities that balance analytical sensitivity with simplicity, reproducibility, and rapid readout suitable for OSCC-specific clinical decision points. Rapid, POC-enabled diagnosis would allow timely referral for treatment, directly impacting the clinical decision window for OSCC management.99,100,102
A key goal of nonmetastatic cancer surgeries is to remove the entire tumor when possible. In the case of oral surgery, the clinician must often rely on visual and tactile clues to determine tumor borders and then to cut beyond them. When done properly this leaves a small tumor-free margin of normal tissue around the extirpated tumor and improves prognosis.106 During oral cancer surgery (>90% of which are OSCC) many oral surgeons in addition rely on intraoperation frozen section analysis of tumor.107 This allows the surgeon, while the patient is still under anesthesia and the surgical bed still open, to determine if there is a need to remove more tissue. The usage of intraoperation frozen section analysis during surgery requires that an oral pathologist be nearby to examine flash frozen and stained sections from surgically removed tissue margins at the time of surgery. This practice of having a pathologist immediately examine tissue takes anywhere from 15 to 30 minutes to complete and minimizes the chance of leaving residual tumor behind while also maximizing the preservation of normal tissue. A POC intraoperative instrument capable of assessing tumor margins in real time could guide surgical resection with improved precision, minimizing residual tumor while preserving healthy tissue. One group is working to develop an instrument with electrochemical sensors to measure hydroxyl radicals to precisely detect tumor and tumor-free tissue even when distorted physically during surgery,108,109 though additional rapid testing of a variety of marker molecules may be applicable to this task.
Saliva and saliva fractions, such as those enriched for extracellular vesicles, brushings from oral lesions, and even plasma can be analyzed for specific RNA, DNA, DNA methylation, and proteins that can be used to measure at least some component of tumor gene expression associated with OSCC.110–112 These approaches for marker measurement commonly use one of two protocols, detection of marker protein or RNA in fixed cells, or measurement of these or other markers in solubilized extracts. POC methods to detect OSCC that rely on measures of 2 to 3 proteins in fixed oral cells are one proposed POC technology, along with POC devices that rely on cell staining and AI-based analysis of cell morphology with additional antibody conjugated bead based immunoassay of one or more marker proteins.113,114 These biomarker-specific approaches highlight the constraints unique to OSCC POC testing, including the need to detect low-abundance nucleic acids or proteins from limited and heterogeneous oral samples, while maintaining rapid turnaround compatible with in-clinic decision-making,
2.3. Molecular and imaging perspectives in hepatocellular carcinoma
Hepatocellular carcinoma (HCC) represents a major healthcare burden, resulting in the third highest global cancer related mortality and a 5-year survival rate of only 18%.115,116 HCC is unique in that radiologic imaging can establish diagnosis in most cases, minimizing the need for invasive tissue biopsy except in select scenarios.117–119 Despite its diagnostic utility, imaging provides limited insight into tumor molecular subtypes, which are critical for predicting prognosis and guiding therapy. There are two major molecular subtypes of HCC, proliferative and non-proliferative, each encompassing approximately 50% of tumor prevalence. Proliferative HCCs—mediated by TP53 mutations—are poorly differentiated and portend a dismal prognosis, while non-proliferative HCCs—possessing CTNNB1 mutations—are well differentiated and show favorable outcomes.120,121 Each molecular subtype has unique immunohistochemical characteristics, genetic alterations, signaling pathways, and epigenetic features, all of which could have meaningful implications in therapeutic planning and treatment outcomes, either with targeted locoregional therapies (e.g., percutaneous ablation or transarterial chemoembolization/radioembolization) or systemic therapies like small molecule chemotherapy or immunotherapy.122–125 This highlights the HCC-specific challenge for POC diagnostics: the need to capture molecular subtype information, often from limited or non-invasive samples, to inform personalized treatment strategies. Current HCC treatment algorithms largely ignore molecular heterogeneity, presenting an unmet need for POC assays that can rapidly stratify patients by subtype to guide therapy. To this end, current HCC treatment algorithms do not account for inherent tumor biology and entail “one size fits all” therapeutic strategies unsupported by strong mechanistic foundations. For example, the widely applied Barcelona Clinic Liver Cancer (BCLC) system for HCC126 recommends treatment strategies based only on anatomic features of tumor size, focality, invasiveness, and metastatic spread, but does not account for tumor genetics. In theory, proliferative subtype HCC may require a more aggressive treatment regimen, such as combination therapy with both locoregional and systemic therapy using curated methods and agents, while monotherapy may be sufficient for non-proliferative HCC. However, in the current form, the BCLC scheme would designate identical therapy recommendations for comparably small, focal HCCs of differing genetic subtypes, which could influence clinical outcomes. To this end, the treatment response for proliferative versus non-proliferative HCC—at similar BCLC staging—to transarterial chemoembolization is drastically different (overall survival: 14.4 months versus 38.7 months).127 Because tissue biopsy carries procedural risks and is seldom performed, HCC POC strategies must rely on non-invasive samples such as circulating tumor DNA or microRNAs to assess molecular subtypes.128,129 Compounding the issue, even if tissue were readily available, existing molecular profiling technologies are laborious, and require whole exome sequencing, whole genome sequencing, RNA sequencing, or nestin staining.130 Circulating genetic material may serve as an alternative, non-invasive tool to investigate HCC molecular heterogeneity. Particularly, circulating tumor DNA and microRNA can serve as biomarkers for predicting molecular subtypes of HCC,131,132 which could then be correlated with clinical outcomes to advance rational therapeutic algorithms. Existing liquid biopsy workflows are limited by instrumentation cost, sample input requirements, and lengthy processing times, highlighting the HCC-specific challenge of achieving rapid molecular profiling in clinically actionable timeframes.133,134 Developing POC assays capable of rapid, minimally invasive molecular profiling would enable subtype-specific treatment decisions and facilitate precision oncology for HCC patients. Such information could be further supplemented by cross-sectional radiologic imaging with computed tomography (CT) or magnetic resonance imaging (MRI), which may serve to further enhance subtype prediction.125 Phenotypic imaging features such as tumor shape, architecture, texture, density or signal intensity, and contrast enhancement patterns can be used to develop radiomic models that can be combined with genomic markers to improve molecular profile predictability.127 In conclusion, advancement of POC molecular diagnostics—in combination with advanced non-invasive radiologic imaging and leveraging ever evolving artificial intelligence-based image analysis—can shepherd a new age of HCC treatment through non-invasive stratification of molecular subtypes and provide meaningful improvements in the clinical outcomes of this deadly disease. Integration of molecular biomarkers with imaging-derived radiomic features further defines the unique constraints for HCC POC tools: assays must be compatible with limited sample volumes, sensitive enough to detect subtype-defining signals, and rapid enough to inform treatment decisions within existing clinical workflows.2.4. Liquid biopsy-based copy number variation profiling for POC prostate cancer diagnostics
Prostate cancer will account for greater than 37![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 deaths in US males and over 325
000 deaths in US males and over 325![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 deaths worldwide in 2025.135 The management of metastatic prostate cancer has evolved rapidly, and while several intensified ADT-based drug combinations have been shown to slow the progression of metastatic hormone-sensitive prostate cancer (mHSPC),136 the emergence of drug resistance and progression to metastatic castrate resistant prostate cancer (mCRPC) occur inevitably. Most therapeutic regimens for mCRPC lack predictive biomarkers for treatment selection, emphasizing the need for molecular profiling to guide individualized therapy. Molecular biomarker development is needed to refine practice and identify mCRPC state patients with poor prognosis, for choosing from an increasing number of costly systemic treatments, for defining the optimal sequence of drug and drug combinations most likely to elicit responses and to track the emergence of primary and acquired drug resistance. POC approaches could ultimately enable rapid assessment of these biomarkers from minimally invasive blood samples, but assay sensitivity and specificity must accommodate low ctDNA abundance and variable CNV representation.
000 deaths worldwide in 2025.135 The management of metastatic prostate cancer has evolved rapidly, and while several intensified ADT-based drug combinations have been shown to slow the progression of metastatic hormone-sensitive prostate cancer (mHSPC),136 the emergence of drug resistance and progression to metastatic castrate resistant prostate cancer (mCRPC) occur inevitably. Most therapeutic regimens for mCRPC lack predictive biomarkers for treatment selection, emphasizing the need for molecular profiling to guide individualized therapy. Molecular biomarker development is needed to refine practice and identify mCRPC state patients with poor prognosis, for choosing from an increasing number of costly systemic treatments, for defining the optimal sequence of drug and drug combinations most likely to elicit responses and to track the emergence of primary and acquired drug resistance. POC approaches could ultimately enable rapid assessment of these biomarkers from minimally invasive blood samples, but assay sensitivity and specificity must accommodate low ctDNA abundance and variable CNV representation.
Tissue-based profiling in mCRPC is limited by insufficient material from metastatic lesions, particularly in bone, and is seldom feasible for routine monitoring.137 Clinical and molecular prognostic and predictive biomarkers previously investigated in blood include microRNAs (miRNAs or miRs),138 PSA, hemoglobin (HgB), lactate dehydrogenase (LDH) and alkaline phosphatase (ALP) levels,139 βIII-tubulin,140 circulating tumor cells (CTCs),141,142 AR-V7, AR-V9 splice variants,143–148 plasma circulating tumor DNA (ctDNA) alterations,149–157 and plasma cell-free DNA.158–160 This highlights a prostate cancer-specific constraint for POC molecular diagnostics: accurate detection of multiple ctDNA and protein biomarkers from limited plasma volumes, while maintaining quantitative fidelity suitable for clinical decision-making.
An emerging alternative is probing blood-based somatic alterations for identifying prognostic and predictive signatures for which consistent and accurate detection of the components of ctDNA remains a major challenge. Copy number variations (CNVs) are common molecular events in the mCRPC state161–163 and can be detected in plasma using low-pass next-generation sequencing (NGS) coverage. Concordance studies comparing CNVs detected in plasma/blood (also known as liquid biopsy) cell free DNA (cfDNA) with solid metastatic tissue biopsy approaches in the same patient demonstrate a high degree of agreement,164–166 thus permitting the use of less invasive cfDNA-based CNVs for clinical utility/application. The presence of specific CNV profiles and their evolution over time are potentially targetable therapeutically, unlike non-specific clinical laboratory prognostic measures. In this context multi-CNV risk scores have been developed that provide a tumor biology basis for targeting of high-risk patients with additional therapeutic interventions compared to the current standard of care.
The blood-based, ctDNA profiled, multi-gene CNV risk score derived from CNVs detected in plasma is based on detecting gains in AR, MYC, COL22A1, PIK3CA, PIK3CB, NOTCH1 and losses in TMPRSS2, NCOR1, ZPTB18, TP53, NKX3-1.167 This is a clear example of molecular profiling using blood samples that can be developed as a “lab on a chip” for enhancing care of patients. NGS-based CNV detection remains impractical for routine clinical use due to assay complexity, sample input requirements, and turnaround times, illustrating the technical constraints for prostate-specific POC solutions. Even with decreasing sequencing costs, multi-CNV profiling via NGS is limited by prolonged turnaround and the need for specialized interpretation.
As the detection limits for ctDNA can be as low as 0.1–5% of cfDNA, dependent upon the cancer stage, the lower detection limits and numerical precision of the sensing technology enable ctDNA fraction to be reduced and for small changes in CNV to be accurately measured. Using NGS and ddPCR, ctDNA is detectable when the tumor fraction is >3–5%, but the detectable ctDNA fraction is reduced as the CNV amplification factor increases. Traditional methods such as qRT-PCR for ctDNA have been considered as the gold standard for ctDNA quantification with femtomolar limits of detection.168 However, qRT-PCR requires complicated enzymatic amplification, and amplification bias impacts the accuracy of CNV quantitation.169 Thus, traditional methods for ctDNA detection do not meet current demands. PCR alternatives that include loop-mediated isothermal amplification (LAMP),170 strand displacement amplification (SDA),171 exponential amplification reaction (EXPAR),172,173 rolling circle amplification (RCA),174 and some enzyme-free amplifications, such as catalytic hairpin assembly (CHA),175 hybridization chain reaction (HCR),176,177 and entropy-driven catalysis,178 have been employed widely for the highly sensitive detection of DNA. Constrained by the detection limit and selectivity, none of these methods have been adopted for clinical use for ctDNA detection without enzymatic target amplification. Droplet digital PCR (ddPCR) has long been available for CNV detection,179 including in the context measuring genomic amplifications in the media of pure cancer cell lines and tumor models,180–182 and from tumor tissue183 although its complex workflow, cost/assay, and lack of multiplexing capability limit its utility for routine clinical monitoring of CNV panels in the background of cfDNA.
Emerging assays aim to enable sensitive, multiplexed CNV detection from plasma, with quantitative accuracy sufficient to inform clinical decisions for mCRPC patients. Developing ultrasensitive assays capable of accurately quantifying CNV biomarkers from limited plasma is critical for monitoring disease progression and guiding personalized therapy in mCRPC. For broad adoption in management of cancer, such a technology must have selectivity and quantitative accuracy for targeted genes.
2.5. Rapid ctDNA-based molecular testing in non-small cell lung cancer
Lung cancer is the leading cause of cancer-related mortality in the United States and worldwide. Among the histologic subtypes of lung cancer, non-small cell lung cancer (NSCLC), and more specifically adenocarcinoma, is the most prevalent. Advances in genomic profiling have resulted in increasingly refined molecular classifications of adenocarcinomas of the lung, as many of these molecular subsets of NSCLC can now be treated with targeted therapies. In fact, the FDA has approved more targeted therapies for lung cancer than any other cancer over the last decade. Many of these targeted therapies have been shown to be more effective than the prior standard of care of platinum doublet chemotherapy. Furthermore, immunotherapy is often less effective against many of the molecular subsets of NSCLC with targeted therapy options. Even with the recent FDA approvals of neo-adjuvant chemoimmunotherapy for patients with NSCLC that are surgical candidates, we still need to exclude the presence of EGFR and ALK alterations for patients to be eligible for this treatment approach. Rapid molecular testing in NSCLC is essential to identify actionable mutations, such as EGFR and ALK alterations, needed to guide targeted therapy and determine eligibility for chemoimmunotherapy.NGS profiling of tissue or ctDNA enables detection of actionable mutations in NSCLC, forming the basis for precision-targeted therapy decisions. Despite its utility, current NGS workflows can delay therapy initiation due to complex tissue acquisition and processing requirement. Tissue-based testing is challenged by procedural delays in anticoagulated patients, specimen transport between institutions, and frequent need for repeat biopsies due to insufficient or necrotic tissue, emphasizing the limitations of traditional workflows for NSCLC. Our real-world analysis of over 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 NSCLC patients demonstrates that standard molecular testing can take an average of 64 days from diagnosis to actionable results, highlighting the need for faster, disease-specific molecular assessment. ctDNA-based liquid biopsies have improved turnaround times to 7–10 days, yet further refinements in sensitivity, multiplexing, and workflow integration are needed to fully support NSCLC clinical decision-making. While ctDNA assays complement tissue testing, challenges remain in reliably detecting low-frequency mutations and integrating results into rapid treatment decisions for NSCLC patients.
000 NSCLC patients demonstrates that standard molecular testing can take an average of 64 days from diagnosis to actionable results, highlighting the need for faster, disease-specific molecular assessment. ctDNA-based liquid biopsies have improved turnaround times to 7–10 days, yet further refinements in sensitivity, multiplexing, and workflow integration are needed to fully support NSCLC clinical decision-making. While ctDNA assays complement tissue testing, challenges remain in reliably detecting low-frequency mutations and integrating results into rapid treatment decisions for NSCLC patients.
3. POC sample preparation
While POC cancer diagnostics have the potential to transform how and where we detect disease, one of the biggest hurdles is sample preparation. It is necessary to quickly and automatically isolate the target biomarkers from complex patient samples without relying on conventional lab equipment or complex protocols. Over the past five years, there has been progress in developing microfluidic cartridges that can process samples like blood, saliva, or urine for subsequent detection of molecular biomarkers. Liquid biopsy samples such as blood, saliva, urine, bile, ascites, sweat, and oral brushings can contain tumor-derived molecular markers, including circulating tumor DNA (ctDNA), various forms of RNA, proteins, and extracellular vesicles (EVs).184–190 We summarize these sample types and their molecular targets in Table 1.| Sample type | Tumor-derived molecular targets | POC considerations and examples | Reference |
|---|---|---|---|
| Blood (plasma, serum) | ctDNA (mutations, methylation), EVs/exosomes (oncoproteins, RNAs), CTCs, circulating proteins (antigens) | Most established liquid biopsy; many microfluidic systems integrate blood separation and on-chip extraction (e.g. cfDNA chips, CTC capture devices). Rapid processing needed to avoid cfDNA degradation | Javanmard et al.192 |
| Saliva | ctDNA (for oral or lung tumors), exosomal DNA/RNA (e.g. HPV DNA, microRNAs), proteins (IL-8, etc.) | Truly noninvasive self-collected sample. POC microfluidics like acoustofluidic exosome isolators have enhanced biomarker yields (15× exosomal RNA vs. centrifuge). Used for oral cancer and viral oncogene detection | Wang et al.197 |
| Urine | ctDNA (e.g. TERT or FGFR3 mutations in bladder cancer), exosomal RNA/proteins (PSA, PCA3 for prostate), tumor cells (rare) | Readily available; requires analyte concentration. Integrated chips perform size-based EV isolation and on-chip ELISA | Wang et al.205 |
| Centrifugal and paper-based microfluidics used for fast nucleic acid assays from urine | |||
| Bile | Whole tumor cells (cholangiocarcinoma cells), extracellular vesicles (bile-derived exosomes containing mutant DNA or ncRNA), and proteins (CA 19-9 in bile) | Collected during endoscopy. Viscous fluid with debris. New automated microfluidic systems achieve high sensitivity by magnetic bead capture of CCA cells. Integrating on-chip filtering (e.g. remove gallstones) is crucial | Liu et al.208 |
| Ascites/pleural effusion | Tumor cells (often abundant in ascites of ovarian, gastric cancer), EVs (carrying oncogenic miRNAs, proteins), ctDNA | High-volume fluids requiring high-throughput processing. Microfluidic CTC chips (e.g. spiral inertial sorters) can enrich cells from liters of fluid quickly. EVs can be isolated by immunoaffinity or acoustic methods similar to blood. On-site processing avoids cellular degradation post-drainage | Guo et al.191 |
| Sweat | Exosomal RNAs/proteins (e.g. dermcidin, PIP), small metabolites (lactate, etc.), cytokines (IL-1β) | Completely noninvasive and continuous sampling. Wearable microfluidic patches with built-in sensors can monitor sweat in real-time. Cancer use-case under exploration: one concept is a graphene-based immunosensor for tumor exosomes in sweat | Mirgh et al.184 |
| Oral brushings/swabs | DNA from captured cells (mutations, LOH), intact cells for cytology/CTC analysis, HPV RNA/DNA (for oropharyngeal cancer) | Minimally invasive “liquid biopsy of tissue”. Requires cell lysis or analysis on chip. Microfluidic platforms like SMILE allow on-chip cell processing and detection with minimal user steps. Useful for mass screening by dentists or in low-resource areas | Zoupanou et al.224 |
| Tear fluid | Proteins S100A8 and S100A9; peptides, lipids and small metabolites filtered from plasma that reflect systemic breast cancer changes | Noninvasive Schirmer-strip collection retrieves ≤5 μL; proteome-wide LC-MS/MS discovery panels can be translated to on-chip ELISA; logistic-regression-based cartridge assays achieve AUC 0.902 with 84.8% sensitivity and 86.4% specificity for breast cancer screening | Daily et al.220 |
| Exhaled breath condensate | Tumor DNA bearing TP53 and EGFR mutations; circulating miRNAs (miR-21, altered GATA6/NKX2-1 isoform ratios); metabolites (monoacylglycerols, fatty acid derivatives); volatile organic compound profiles | Ten-minute breath collection into a cooled condenser yields <1 mL; compatible with microfluidic PCR/NGS platforms and electronic-nose sensor arrays to capture tumor-specific “breathprints”; enables noninvasive point-of-care lung cancer screening and longitudinal molecular monitoring | Campanella et al.223 |
Recent POC-compatible methods focus on integrating sample preparation steps such as cell separation, lysis, filtration, and analyte purification onto microfluidic cartridges. This approach eliminates the need for conventional laboratory procedures like centrifugation or column extraction, which are impractical outside the laboratory. By automating these processes, such systems minimize the need for sample transport or preservation, allowing immediate analysis while the biomarkers are still intact.191 Here, we explore the latest advances in POC sample preparation for cancer diagnostics, focusing on how microfluidic platforms are being designed to be fast, user-friendly, and minimally invasive. We highlight representative developments from recent years (2019 to 2024), including integrated cartridges for blood-based ctDNA and EV isolation, saliva and urine processing devices, and novel approaches for bile, ascites, sweat and oral samples. Key features of each platform including their level of automation, processing time, and suitability for POC use are summarized in Table 2. Together, these advances illustrate a trend toward self-contained “sample-to-answer” systems that can bring molecular cancer diagnostics out of specialized labs and into clinics, rural settings, and primary care offices.
| Platform (year) & reference | Sample & target biomarkers | Key methodology and integration | Automation level/POC features | Processing time (approx.) |
|---|---|---|---|---|
| LabChip cfDNA processor (2019), Gwak et al.226 | Blood → ctDNA (plasma cell-free DNA) | Multi-layer microfluidic with on-chip plasma sep., solid-phase DNA extraction (surface-immobilized capture chemistry) | Fully automated (valves & pumps on chip) | ∼30 min to purified DNA (from 4 mL whole blood) |
| ∼2 user steps (load blood into cartridge → press start) | ||||
| One-button operation; no centrifuge needed | ||||
| Hand-powered ctDNA SERS chip (2024), Wu et al.227 | Blood (fingerprick) → ctDNA mutation | Size-based microfilter for rapid plasma separation; on-chip SERS nanoprobe for EGFR mutation (no PCR). Finger-driven vacuum for flow | Power-free operation (manual actuation) | 35 min total (blood-in to result) |
| ∼2–3 user steps (load fingerprick blood → actuate blister/finger vacuum → insert/read on portable Raman); suited for low-resource settings | ||||
| Acoustic EV separator (2020), Z. Wang et al.197 | Saliva → exosomes (HPV DNA, small RNA cargo) | Acoustofluidic (ultrasound) standing wave device; size-selective isolation of exosomes from raw saliva | Semi-automated with external actuator; ∼2–3 user steps (load saliva → start acoustic run → collect output for downstream PCR); requires small benchtop acoustic transducer (portable); no labels or reagents | <30 min (sample to exosomes ready for analysis); improved yield 15× vs. centrifuge |
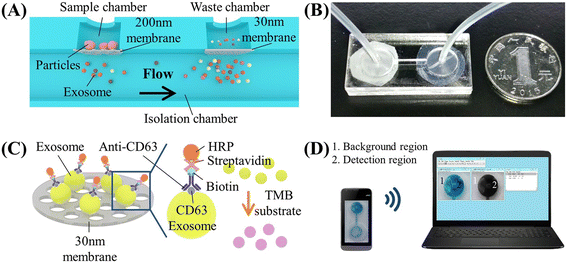
| Double-filtration exosome chip (2017), Liang et al.204 | Urine → EVs (exosomes) for bladder cancer | Stacked membrane filters (200 nm + 30 nm) in PMMA device isolate 30–200 nm EVs; on-chip direct ELISA (anti-CD63) with colorimetric readout | ∼4–6 user steps (load urine → run/flush through filters → add ELISA reagents/incubations as specified → add substrate → smartphone image/readout); self-contained: only syringe or pipette to load sample; color change imaged by smartphone. Low-cost disposable (<$5) | ∼1 hour (including incubation for ELISA) for result; hands-on steps minimal |
| “CAPTURE” bile CTC IMS (2024), Chang et al.206 | Bile → tumor cells (cholangiocarcinoma CTCs) | Centrifugal pretreatment (on-disc) + microfluidic capture module with aptamer/glycan-coated magnetic beads to bind CCA cells; on-chip fluorescence staining | Fully automated multi-module system; ∼2–3 user steps (load bile cartridge → place on centrifugal/IMS instrument and start → read fluorescence output); no manual intervention between steps; integrated “gallstone filter”. Requires bench-top centrifugal spinner (could be portable) | ∼2 hours total (including staining); high throughput (processes 1 mL bile). Validated 100% sensitivity in pilot study |
| SMILE Oral Cancer Chip (2021), Zoupanou et al.224 | Oral brushing → CTCs/cells (for oral SCC) | Two-chip system: one for on-chip sample prep (mixing, filtering to enrich CTCs), second for detection (electrochemical or optical analysis of cells). Made via low-cost 3D-printing and laser cutting | Semi-automated; ∼3–4 user steps (load brushing suspension → initiate on-chip enrichment/mixing → transfer or interface to detection module → read output); plug-and-play concept for non-specialists, but still requires module handling | ∼30–60 min for entire workflow; goal is rapid screening in clinic (currently at prototype stage, with ongoing sensitivity testing) |
| Immunomagnetic HER2 CTC Chip (2023), Parvin et al.264 | Blood → CTCs (breast cancer, HER2+) | Uses magnetic nanoparticles + microfluidic channel: blood incubated with anti-HER2 magnetic beads, then flowed through channel with magnet to pull out tagged CTCs | Low-complexity; manual incubation; ∼3–5 user steps (mix blood with magnetic nanoparticles/incubate → load into chip → apply magnet/flow → rinse if needed → microscope/on-chip stain readout); no external pumps (relies on capillary flow) | ∼20 min processing of 2 mL blood; designed for one-step CTC enrichment at POC (e.g. during chemo sessions) |
3.1. Sample types and target biomarkers
Different minimally invasive biofluids provide complementary windows into tumor biology, but their physical and chemical properties strongly constrain how POC devices must be engineered. Rather than only cataloging biomarkers, it is useful to view each sample type as a matrix with specific hurdles—viscosity, dilution, volume limits, enzymatic degradation, or cellular load—that dictate the necessary sample preparation strategies (Table 1). Below, we discuss how these matrix properties shape POC microfluidic design, highlighting representative platforms for each fluid.Peripheral blood is the most commonly used sample for liquid biopsy, but from an engineering standpoint it is a crowded, enzyme-rich colloid rather than a simple aqueous solution. Even a small draw (around 5–10 mL) can carry a wealth of tumor-related information from ctDNA, CTCs, and extracellular vesicles (EVs).192 Blood also contains tumor-associated proteins (such as PSA or CA-125), along with circulating microRNAs. However, ctDNA is typically present at very low levels, often less than 0.1% of total cell-free DNA, and is usually found as short fragments (about 50–150 base pairs), making efficient extraction a challenge.191 While blood carries large numbers of EVs (typically 109–1012 EVs mL−1), they are mixed with plasma proteins and lipoproteins, resulting in the need to separate them efficiently with low loss.193,194 Furthermore, high hematocrit and dense cellular content make blood non-ideal in microchannels, for instance red and white cells in blood increase apparent viscosity, cause non-uniform cell distributions and can clog constrictions, especially when flow is driven through narrow filters or sharp expansions. Pre-analytical studies have also shown that circulating DNA in blood is highly vulnerable to nuclease activity and pre-processing delays. Markus et al. demonstrated that plasma cell-free DNA (cfDNA) yield and fragment size are strongly affected by the tube type and processing protocol, using a multiplex ddPCR assay to show that certain tubes and centrifugation regimes preferentially preserve short fragments that are enriched for tumor-derived DNA.195 More recently, Peng et al. systematically dissected pre-analytical variables in both blood and urine and reported that tube anticoagulant, temperature, and delay time before plasma separation substantially alter measurable cfDNA concentrations and fragmentomics.196 These findings imply that POC systems which accept whole blood must integrate rapid plasma separation and extraction to minimize nuclease exposure and must be compatible with the specific chemistries of EDTA or cfDNA-stabilizing tubes that are often used in oncology practice. At the microscale, the high protein content and lipoprotein load of plasma promote nonspecific adsorption and sensor fouling, requiring antifouling coatings such as PEG or zwitterionic brushes on channel walls and electrodes to maintain analytical performance in repeated use. Rapid, closed, on-cartridge workflows that combine gentle plasma separation, controlled shear, and solid-phase or bead-based capture therefore emerge as a necessity rather than a convenience when blood is used as the input matrix.191
Saliva presents a nearly opposite problem. While it is easy to obtain noninvasively, it is rheologically complex and highly variable. Whole saliva is a mucin-rich, viscoelastic, shear-thinning fluid with pronounced extensional rheology, governed largely by the glycoproteins MUC5B and MUC7 and modulated by calcium, bicarbonate and pH.197 Vijay et al. quantified “spinnbarkeit” (filament-forming extensional viscosity) and showed strong correlations between extensional properties and ion/mucin content.198 Sjögren's and radiation-induced xerostomia further perturb this rheology, reducing both volume and elasticity of saliva.199 For microfluidic POC devices, this non-Newtonian behavior means that flow fronts can be unstable, bubbles are easily trapped, and mucus strands and cell clumps can bridge and occlude microstructures. Recent work using saliva as a diagnostic sample has found that simple microfiltration or clarification steps substantially improve the reproducibility of electrochemical and optical measurements by removing mucin strands and larger particulates before the analytical stage.200 Together, these observations dictate that saliva-based POC devices incorporate a “conditioning” stage, which is a coarse filtration or centrifugal clarification to remove food debris and clumps, controlled dilution with low-viscosity buffer to normalize rheology, and anti-fouling channel surfaces to mitigate persistent mucin adsorption. The acoustofluidic saliva chips used for label-free vesicle enrichment exploit bulk acoustic waves to size-focus particles without physical filters, precisely because traditional membrane-based approaches are vulnerable to clogging in this viscoelastic matrix.201
Urine, in contrast, is typically a low-viscosity, nearly Newtonian fluid that is physically easy to pump and fill into microchannels, but tumor-derived analytes in urine are often highly diluted. Hydration status and kidney function cause orders-of-magnitude variation in osmolality, ionic strength and pH, and cfDNA, exosomes, and tumor-associated proteins commonly appear at low-nanomolar or picomolar levels. Recent evaluations of urinary cfDNA workflows emphasize that attaining sufficient sensitivity for mutation detection often requires processing tens of milliliters of urine and that pre-analytical variables such as storage temperature and the presence of cell debris strongly influence recovery.196,202 Design of microfluidic POC systems for urine must therefore be high-throughput concentrators: they need large capture surface areas, parallelized filters, or high-surface-area beads to sweep rare targets out of a relatively large volume into a small elution or detection chamber.203 The double-filtration EV chips described here are a prime example of this logic: a 200 mm membrane excludes cells and coarse debris, while a 30 nm membrane retains 30–200 nm vesicles, effectively transforming tens of milliliters of dilute urine into microliter-scale, vesicle-enriched retentate that is compatible with on-chip immunoassays and smartphone-based imaging (Fig. 2).204 Physicochemically, the high ionic strength and urea content of urine can influence hybridization kinetics and sensor baselines; buffer exchange or controlled dilution on-chip thus becomes a second design requirement, aimed at regularizing ionic strength so that subsequent nucleic acid or electrochemical readouts are stable across patients and collection times.205
Bile is a more extreme example of a viscous, surfactant-rich matrix that nonetheless carries highly local information on biliary tract tumors. It is a multiphase emulsion containing bile salts, cholesterol, phospholipids, mucus, cellular debris and sometimes gallstones. These components make bile non-Newtonian and prone to forming mucus strands and aggregates that adhere to microchannel walls and obstruct narrow features. In addition, bile salts act as detergents that can denature immobilized antibodies or aptamers and disrupt hydrophobic/hydrophilic patterns that are commonly used to implement capillary valving or droplet formation. Chang et al. recently reported an integrated microfluidic system for cholangiocarcinoma cell detection from bile in which a centrifugal pretreatment module first removes mucus and traps stones, after which an immunomagnetic module uses an antibody/aptamer/glycosaminoglycan cocktail on magnetic beads to capture tumor cells.206 Briefly, the system includes a centrifugal pretreatment module (to remove mucus and block >1 mm gallstones by a micro-tier filter) and a cell capture module that uses functionalized magnetic beads. The beads are coated with a cocktail of affinity probes (aptamer HN16 and a glycosaminoglycan that bind CCA cells) to fish out cholangiocarcinoma cells from the bile. Captured cells are then automatically stained on-chip and identified. In a single-blind clinical study, the integrated bile-chip achieved 100% sensitivity and 83% specificity for CCA (26 patient samples), markedly improving on prior methods.207 This illustrates how complex liquid samples can be handled by combining mechanical (centrifugal) and microfluidic components in a POC device. Future bile diagnostics might integrate such sample preparation with on-chip molecular analysis such as mutation-specific DNA sequences from biliary EVs,208 enabling earlier cancer detection.
Ascites and pleural effusions represent large-volume, cell-rich fluids in which the dominant challenge is throughput rather than viscosity per se. These fluids can accumulate in volumes of hundreds of milliliters and contain mixtures of malignant cells, mesothelial cells, leukocytes and protein-rich exudate. Conventional cytology typically analyzes only small aliquots, which contributes to low diagnostic sensitivity. Inertial and viscoelastic microfluidic devices have been adapted to this matrix because their open, membrane-free channels are less prone to clogging at high cell densities. Spiral microchannels and elasto-inertial flows can continuously process diluted effusions at flow rates in the milliliters per minute range, separating larger, less deformable tumor cells and clusters from smaller hematopoietic cells based on size and mechanical properties.209 High cell density and fibrin can still precipitate clogging, so devices are typically operated with controlled pre-dilution and incorporate bypass branches and gradual expansions in channel cross-section to soften sudden pressure jumps.210 Proteolytic activity and inflammatory enzymes in ascites and pleural effusions can degrade vesicle surface proteins and soluble markers, so POC designs again benefit from integrated separation and immediate stabilization of the cell-free fraction (e.g., cooling, protease inhibitors, chaotropic buffers) inside the cartridge.211–213
Sweat illustrates the opposite corner of the design space: it is low in protein and nucleic acids, highly ionic, and produced locally at low flow rates, but it can be harvested continuously through the skin. Wearable microfluidic sensors exploit capillary and wicking effects in thin elastomeric channels to route sweat from the skin to sensing chambers before it can evaporate. Gao et al. reviewed recent flexible electrochemical sweat sensors and emphasized issues such as high background conductivity, sensor drift, and evaporation-driven concentration changes, which all complicate quantitative measurements in this matrix.214 Ramachandran et al. further highlighted that sweat analyte levels are strongly dependent on the local sweat rate and that microfluidic designs must strictly control the sampling area and back pressure to avoid inducing occlusion or backflow.215 For cancer applications that target sweat-derived vesicles or proteins, this means that POC devices will need extremely sensitive detection in tiny volume elements (often sub-microliter droplets), while also discriminating true biochemical changes from physical artifacts caused by variable sweat rate, evaporation, and skin-debris contamination. This drive designs toward closed, short channels, debris-excluding inlets, and either disposable microfluidic layers or robust antifouling surface chemistries, as well as multiplexed sensing (e.g., simultaneous measurement of sweat rate, conductivity and target biomarker) to enable on-device normalization.214,216,217
Oral brushings and swabs occupy an intermediate position between saliva and tissue biopsies. The primary sample is a dense slurry of epithelial cells, blood, mucus and microbiota, rather than a free-flowing liquid. Cell concentrations are high, and cells often form clusters embedded in mucus, producing a viscoelastic, heterogeneous suspension that sediments and aggregates during handling. Microfluidic systems that process these samples, such as the SMILE platform for oral cancer screening, therefore devote substantial design complexity to disaggregation and controlled resuspension, using micromixers, serpentine channels, or deterministic lateral displacement structures to break up clumps and achieve uniform cell distributions before enrichment or lysis. The mucus component introduces similar fouling and rheological problems seen in saliva, but with higher solids content, which can necessitate on-chip mucolytic treatment or dilution. Compared to blood-based CTC assays, oral brushing-based POC devices are thus fundamentally “cell-centric”: their front ends must be engineered to accept and homogenize dense, sticky cell suspensions and to maintain viability or nucleic acid integrity through mechanical perturbations that are more intense than those used for plasma-based workflows.
Tear fluid is nearly the opposite extreme: it is comparatively clean and protein-rich, but available volumes are minuscule and strongly dependent on sampling method. Schirmer strips, which wick tears onto cellulose paper, can collect on the order of 5–10 μL over several minutes but also induce reflex tearing and cause substantial adsorption of proteins to the strip matrix. Nättinen et al. compared capillary and Schirmer-strip sampling and found pronounced differences in tear proteome composition and relative protein abundances between methods, underscoring that the collection device acts as a strong pre-analytical filter.218 Recent analyses by Ngo et al. and others reinforce that protein recovery and total yield depend sensitively on elution conditions and that Schirmer strips, while convenient, may under-represent certain protein classes.219 Daily et al. showed that, even under these constraints, tear proteins such as S100A8/A9 measured from Schirmer-strip eluates can discriminate breast cancer from controls with an AUC of 0.902.220 For POC microfluidic design, these findings imply that the dominant constraints are volume and surface adsorption rather than viscosity: channels must have extremely low dead volume, internal surfaces must be engineered to minimize nonspecific adsorption of low-nanogram quantities of protein, and the cartridge should ideally accept both Schirmer eluates and capillary-collected tears with well-defined dilution factors. In practice, this leads to nanofluidic architectures and on-chip detection in nanoliter reaction chambers, rather than macroscopic dilution followed by off-chip assays.
Exhaled breath condensate rounds out the matrix landscape by adding volatility and environmental sensitivity to extreme dilution. EBC is obtained by cooling exhaled air for approximately ten minutes, generating sub-milliliter volumes of condensed airway lining fluid that contain dissolved gases, volatile organic compounds, small metabolites and trace amounts of macromolecules. The American Thoracic Society/European Respiratory Society recommendations and subsequent updates emphasize that the collection device geometry, condensation temperature and breathing pattern strongly influence EBC volume and composition and that standardization of these physical parameters is critical for comparability between studies.221,222 From a POC perspective, this means that the collection hardware and the analytical cartridge are essentially a single system: the microfluidic device must control cooling and flow, prevent re-equilibration with ambient air, and often perform pre-concentration steps (e.g., solid-phase extraction, controlled partial evaporation) to bring target analytes above detection limits. Moreover, because EBC is susceptible to contamination by oropharyngeal saliva and environmental aerosols, inlet designs and flow paths must physically segregate condensate associated with lower airway lining fluid from upstream droplets, for example by incorporating saliva traps, hydrophobic filters, or differential collection surfaces.223
3.2. Microfluidic and integrated platforms for sample preparation
Recent advances in POC technologies have harnessed microfluidics to carry out complex sample preparation steps—such as isolation, enrichment, and purification of cancer biomarkers—on a single, compact platform. In this section, we review the core technological strategies used in these devices, organized by biomarker target type. We also highlight representative systems that illustrate these approaches. For a comparative overview of their features and performance, please refer to Table 2.On-chip cfDNA extraction: silica column or magnetic bead DNA extraction kits are difficult to deploy outside laboratory settings, leading to the development of microfluidic analogs. For example, Lin et al. reported a microfluidic device (PIBEX) that uses a photopatterned polymer with –COOH groups to bind DNA, achieving efficient capture of ∼150 bp ctDNA fragments from plasma.225 On-chip solid-phase extraction can yield high recovery and purity at low cost, processing a few hundred microliters of plasma with minimal reagents. A fully automated cfDNA purifier by Gwak et al. integrates plasma separation, DNA binding, washing, and elution in one microfluidic cartridge.226 This device, controlled by electromagnetic valves, purifies cell-free DNA from 4 mL of whole blood in ∼30 minutes, delivering amplifiable DNA. Crucially, such rapid processing preserves the integrity of short-lived ctDNA and obviates the need for blood storage or shipping to a central lab. The extracted DNA can either be collected for analysis or analyzed on the same platform (by flowing the extracted material into an on-chip PCR chamber or sequencing library preparation module).
Hand-powered ctDNA chip: in resource-limited settings, electricity may be unavailable. Wu et al. demonstrated a finger-actuated microfluidic chip for mutation detection in ctDNA that requires no external power.227 In this design, a user presses a blister to create a vacuum that drives whole blood through a microscale filter trench, which rapidly removes blood cells and yields plasma. The plasma then enters a reaction zone with dried reagents for DNA detection by Surface Enhanced Raman Spectroscopy (SERS). The chip employs SERS probes that hybridize to a specific EGFR gene mutation (E746-A750 deletion common in lung cancer). If mutant ctDNA is present, a Raman-active signal is generated without the need for PCR amplification, and the result is read by a portable Raman instrument. This fully integrated device achieved mutation detection in 35 minutes from finger-stick blood, successfully identifying lung cancer patients, and its simplicity (finger pump + one reader) makes it suitable for use in clinics. While current SERS and optical readout instruments are still bench-top devices, progress in miniaturization suggests that handheld Raman readers or smartphone-based detectors could soon pair with such chips for field use.
Integrated nucleic acid analysis: beyond extraction, microfluidics can incorporate nucleic acid amplification and detection on-chip, creating sample-to-answer devices. One widely implemented concept is droplet digital PCR (ddPCR) on microfluidic arrays: after extraction, the DNA (or RNA after on-chip reverse transcription) is partitioned into thousands of nanoliter reactors on a chip, and target sequences are amplified and quantified by fluorescence.228 Another emerging approach is CRISPR-based nucleic acid detection on chip: Cas enzymes (like Cas13a) can be deployed to detect specific DNA/RNA sequences via their collateral cleavage of reporters, eliminating PCR entirely.229,230 A recent study integrated Cas13a reagents into a microfluidic electrochemical sensor and successfully detected a brain tumor-associated miRNA sequence in patient serum within 4 hours, using <1 μL sample.231 This marriage of microfluidics and CRISPR could potentially lead to amplification-free, single-step nucleic acid tests at the POC. Overall, through innovations like electric-field driven DNA capture, isotachophoresis pre-concentration, and miniaturized thermocyclers, microfluidic devices are making it feasible to perform complex genetic assays – from extraction to detection – in a self-contained cartridge the size of a credit card.
Transitional progress/near POC sample-to-answer: rapid ctDNA analysis at (or near) the POC requires efficient nucleic-acid extraction from a few milliliters of blood and fully integrated workflows. Cartridge PCR systems have been reported recently. The Biocartis Idylla system is an automated, sample-to-result platform used in decentralized labs that offers assays for ctDNA mutation detection.232–234 For example, the Idylla ctKRAS assay extracts and amplifies KRAS mutant DNA from just 1 mL of plasma in a single-use cartridge. Within ∼130 minutes, the instrument delivers a result, with all DNA extraction, PCR amplification, and result interpretation performed automatically. The system is currently employed only for research use. Such integration (with <1 minute of hands-on work) makes it feasible for hospital labs, or potentially near-patient settings, to rapidly genotype cancers (for example, identifying KRAS mutations in colorectal cancer) from blood. While many Idylla assays are currently research-use or CE-marked in Europe, they demonstrate clinical-grade performance comparable to centralized methods.235 Another example is Cepheid's GeneXpert platform, which has expanded beyond infectious disease into oncology. Xpert Bladder Cancer Detection is a cartridge test for hematuria patients that measures a panel of five mRNA biomarkers in urine.236 The urine is added to a self-contained GeneXpert cartridge, which lyses cells, extracts RNA, and performs RT-PCR, delivering results in ∼90 minutes. This test runs on the same GeneXpert machines widely used for TB and COVID-19 diagnostics, underscoring how a robust sample-preparation and detection cartridge system can be adapted to cancer biomarkers. Cepheid also markets an Xpert Breast Cancer STRAT4 (ref. 237) assay for rapid molecular subtyping of breast tumors (ER/PR/HER2/Ki67 mRNA) in <2 hours and quantitative Xpert BCR-ABL tests for leukemia monitoring,238 leveraging the platform's integrated sample prep and analysis. These illustrate the growing menu of “sample-in, answer-out” cartridges for cancer, enabling standardized molecular tests outside a centralized laboratory.
Immunoaffinity capture: many microfluidic platforms isolate EVs using immunoaffinity strategies, where microstructures such as microfluidic channels, pillar arrays, or magnetic beads are coated with antibodies that specifically bind to EV surface markers like CD63 or EpCAM. These affinity-based systems offer high specificity and purity. For instance, Chen et al. developed a device, which channels plasma through microchannels functionalized with anti-EpCAM antibodies, allowing selective capture of tumor-derived EVs expressing EpCAM, which can then be released for downstream analysis.240 Commercial platforms such as ExoChip use similar principles and have been tested at the patient bedside to rapidly enrich EVs for RNA profiling.241 Integration of immunocapture with on-chip enumeration has also been demonstrated, for example, a digital microfluidic ELISA system that captures EVs in microwell arrays and quantifies them using fluorescence-based detection.242,243 This form of “digital EV counting” enables quantitative, POC assessment of EV levels and could serve as a valuable tool for monitoring tumor burden in real time.
Size-based filtration and sorting: the double-filtration chip illustrated in Fig. 2 exemplifies size-exclusion-based isolation of EVs, a strategy that relies on physical characteristics rather than surface markers. Several microfluidic designs utilize either single membranes or nanoporous filters for this purpose. For example, a handheld device integrating a silicon nanowire filter was developed to trap EVs from serum based on pore size, enabling on-site RNA extraction.244 Label-free techniques such as inertial microfluidics (Dean flow) and deterministic lateral displacement (DLD) have also been adapted for EV isolation. These methods separate particles by size as they flow through specially designed channels. A recent DLD-based microarray chip used an array of micropillars to deflect larger particles (>150 nm) while allowing smaller EVs to pass through in a streamline. This platform achieved over 80% EV recovery in under 30 minutes.245 Acoustic separation also known as acoustofluidics is another promising label-free approach that uses sound waves to manipulate particles based on their size and density, directing them into distinct streamlines. This method has been applied successfully to isolate EVs from saliva, and similar acoustofluidic systems have demonstrated continuous, high-purity EV isolation from urine.197 These physical separation strategies offer distinct advantages: they avoid surface antigen bias, preserve the structural integrity of EVs, and are well-suited for downstream analyses such as RNA profiling or electron microscopy imaging.
Integrated EV analysis: an emerging trend in microfluidic EV platforms is the integration of isolation and detection into a single device to streamline POC workflow. For example, Xu et al. developed a two-stage microfluidic system that combined efficient EV capture and on-chip electrochemical detection.246 The first stage used a staggered Y-shaped micropillar array to mix the sample with magnetic beads conjugated with anti-CD63 antibodies, enabling selective binding of EVs. In the second stage, the bead-bound EVs were directed to an indium tin oxide (ITO) electrode, where they generated an electrochemical signal proportional to EV concentration. The fully automated process was completed in approximately one hour, demonstrating the feasibility of self-contained liquid biopsy cartridges. Another innovative approach combined on-chip exosome immunocapture with SERS.247 In this platform, EVs were first enriched via antibody-coated microchannels, then exposed to a SERS-active nanoprobe that produced a molecular fingerprint of their protein cargo. This allowed for sensitive, label-free tumor EV profiling without the need for off-chip assays.
Despite these advances, challenges in EV sample preparation remain. Achieving both high recovery and high throughput is difficult due to the low abundance of EVs in biofluids (often <108 particles per mL) and the risk of microchannel clogging. To address these limitations, researchers are introducing parallelized chip designs, featuring multiple channels to increase flow rates—and incorporating antifouling surface coatings to reduce blockages. As summarized in Table 2, several integrated EV platforms are now capable of processing 1–4 mL of biofluid in under an hour, representing a dramatic improvement over traditional ultracentrifugation workflows, which often take several hours. Importantly, these systems fit into portable form factors suitable for bedside or outpatient use. This opens the door to real-time cancer monitoring, for example, analyzing plasma EVs from lung cancer patients for EGFR mutations prior to each treatment cycle, directly in the clinic.
Translational progress: several translational platforms now enable fast EV isolation. For example, the ExoDisc (a lab-on-a-disc device) uses a spinning microfluidic disk with nanofilters to size-selectively capture exosomes.248–250 The ExoDisc can process a sample in ∼30 minutes (or ∼1 hour if coupled with on-disc ELISA detection) and yields high-purity EVs.251 Notably, both the original ExoDisc and an updated blood-compatible version (ExoDisc-B) have been commercialized, offering 10–40 minute EV prep from urine, plasma, or even whole blood.248–250 Other passive microfluidic devices include nanoporous membrane chips such as ExoTIC,252,253 which achieve high recovery rates from samples such as culture supernatant, plasma or urine, and nanostructured ciliated micropillars that enable high-specificity size-based capture. Recent work has also demonstrated biosensor-integrated recovery devices such as the biosensor integrated recovery device (BIRD),254 which uses a sequence of nanoporous membranes (450, 200, 100 nm) to rapidly enrich EVs and virions within ∼20 minutes, achieving ∼55% recovery in buffer and ∼10% in plasma while operating in a disposable, closed cartridge. Crucially, BIRD integrates directly with a portable photonic crystal-based label-free detection by a photonic resonator interferometric scattering microscopy (PRISM)254,255 reader for label-free imaging and digital counting of EVs, illustrating how preparation and detection can be combined in a single near-POC workflow. Active separation strategies are also advancing translation: acoustofluidic platforms (e.g., AcouTrap,256 now commercialized via AcouSort AB) use sound waves to gently enrich EVs from plasma, urine, and whole blood, while immunomagnetic chips (ExoSearch257) or dielectrophoretic devices258,259 isolate EVs by surface markers or electrical properties. Several companies are extending EV sample preparation to POC devices. For instance, BioFluidica's LiquidScan platform can immunocapture EVs (as well as cells and cfDNA) on a disposable plastic microfluidic chip, automating the enrichment of EVs from blood.260 These advances demonstrate how academic innovations in EV enrichment are rapidly translating into startup companies and commercially available platforms, bridging the gap toward clinically viable POC diagnostics.
Label-dependent CTC chips: many microfluidic platforms for CTC isolation261 rely on label-dependent capture strategies, using antibodies that target epithelial or tumor-specific surface markers, most notably EpCAM. These antibodies are immobilized on microchannel walls or conjugated to magnetic beads, enabling selective binding of CTCs while allowing other blood components to flow through.262 Systems such as the earlier-generation CTC-iChip and its successors have evolved toward greater simplicity, portability, and suitability for POC use.263 A recent innovation is an immunomagnetic separation chip that incorporates a handheld magnet for simple operation.264 In this device, blood is mixed with magnetic nanoparticles conjugated with anti-HER2 antibodies (targeting HER2-positive breast cancer cells), then passed through a microchannel with a zigzag geometry. A magnet positioned beneath the chip deflects and traps the tagged CTCs laterally, allowing other blood cells to pass by. This compact device, tested in a clinical pilot study, demonstrated high-purity isolation of HER2-positive CTCs and was designed for use in a physician's office with minimal supporting equipment. Another promising direction involves replacing antibodies with synthetic DNA or RNA aptamers that bind specific surface markers with high affinity.265 Aptamers offer enhanced thermal and chemical stability, allowing the devices to be stored for extended periods without refrigeration, which is a key advantage for decentralized or resource-limited settings. Aptamer-functionalized microfluidic chips have shown the ability to capture CTCs even after months of storage. When combined with on-chip staining or immunocytochemistry modules, these platforms offer an integrated solution for CTC isolation, identification, and characterization.266 Such tools could support personalized oncology, for example, the detection of HER2-positive CTCs could inform real-time decisions on HER2-targeted therapies.267
Label-free cell separation: label-free techniques such as size-based filtration, dielectrophoresis (DEP), and inertial focusing offer powerful alternatives for isolating CTCs without the need for molecular markers. This is especially valuable for cancers that undergo epithelial-to-mesenchymal transition (EMT), a process in which tumor cells downregulate surface markers like EpCAM, making antibody-based methods less effective.268 Among these, spiral microfluidic devices utilizing inertial focusing have shown particular promise.210,269 These systems consist of small plastic cartridges with curved microchannels that leverage inertial lift forces to separate larger, less deformable CTCs from smaller blood cells. Operating at flow rates of a few mL min−1, they enable continuous, high-throughput separation without moving parts. The simplicity and low manufacturing cost of these devices make them well-suited for disposable, POC use. Several clinical studies have deployed spiral chips in practice—for instance, isolating CTCs from ovarian cancer patient blood samples for downstream genomic analysis, all within 20 minutes of collection.270 Dielectrophoresis (DEP) is another label-free approach that separates cells based on their unique electrical properties.271 DEP chips generate alternating electric fields in a microchamber, pushing CTCs into separate flow paths distinct from leukocytes or other blood components.272,273 This method has even been successfully applied to pleural fluid samples, where tumor cells were isolated from white blood cells in lung cancer cases.209 One major advantage of label-free separation at the POC is that these techniques typically require little to no sample preparation; blood or fluid samples can be loaded directly into the device, and separation occurs in real time. However, label-free enrichment may co-isolate non-tumor cells.274 To improve specificity, a practical POC workflow might integrate these techniques with on-chip immunostaining or molecular assays to confirm cancer cell identity post-enrichment.
Fully integrated cell assays: the CAPTURE system for cholangiocarcinoma, discussed earlier, serves as a compelling example of integrating tumor cell isolation with downstream analysis on a single microfluidic platform. Building on this concept, recent developments have focused on devices that not only isolate CTCs but also enable their on-chip culture or molecular interrogation. One such innovation is the MyCTC chip, which captures CTCs directly from whole blood and maintains them in microscale culture chambers.226 This setup allows for short-term culture and drug susceptibility testing on patient-derived CTCs within a few days. While the culture component extends beyond the strict timeline of POC diagnostics, the initial CTC capture and seeding can be performed in near-patient settings. Afterward, the chip is simply incubated, enabling personalized therapy decisions without requiring conventional cell culture labs. Another integrated approach involves combining CTC capture with genetic analysis directly on the chip.262 In some systems, isolated CTCs are lysed in situ, and their DNA is channeled into on-chip polymerase chain reaction (PCR) modules for mutation screening or even into microfluidic units for sequencing library preparation.275 These platforms aim to deliver more than just cell enumeration—they facilitate functional and molecular profiling of tumor cells at or near the point of care. Though still in the experimental stage, these fully integrated systems represent the future of POC cell-based diagnostics. By enabling real-time testing of viable tumor cells—including drug response and mutation profiling—these technologies move beyond basic CTC counting and toward actionable, patient-specific cancer insights.
In summary, microfluidic engineering has enabled the development of a versatile toolkit of miniaturized sample preparation technologies—ranging from membranes and micromixers to magnetic bead systems and acoustic transducers—that can be tailored to meet diverse cancer diagnostic needs. Over the past five years, many of these innovations have advanced beyond proof-of-concept stages into functional prototypes that have been tested on clinical samples, often demonstrating performance on par with or even surpassing traditional laboratory methods. Table 2 highlights several representative integrated platforms, summarizing their technical features, target biomarkers, sample processing capabilities, and point-of-care readiness.
Fabrication strategies strongly influence scalability and cost per unit. High-throughput techniques such as injection molding, hot embossing, and roll-to-roll processing enable precise and reproducible replication of microchannel architectures with minimal per-device cost, making them more suitable for commercialization than photolithography- or soft-lithography-based approaches commonly used in laboratory prototypes.284–287 Injection molding of thermoplastics, in particular, supports tight dimensional tolerances, rapid cycle times, and seamless integration with downstream assembly processes, all of which are critical for clinical deployment.288,289 Paper-based and hybrid microfluidic platforms further extend scalability by reducing material and fabrication costs, enabling capillary-driven flow, and supporting disposable, single-use formats well suited for decentralized and low-resource settings.290–292 Emerging additive manufacturing approaches, including high-resolution stereolithography and digital light processing, offer unique advantages for rapid design iteration and complex architectures; however, challenges related to surface roughness, material biocompatibility, and production throughput currently limit their adoption for mass manufacturing. Bridging this gap will require optimization of printable materials and post-processing workflows that meet clinical and regulatory standards. Beyond materials and fabrication, true scalability requires system-level considerations, including integration of detection modules, fluidic interfaces, and user-friendly packaging. Reducing manual assembly steps, minimizing component count, and adopting standardized, modular architectures can substantially lower manufacturing complexity and facilitate quality control and regulatory approval.
As highlighted in Table 2, several integrated microfluidic platforms are already being adapted for scalable fabrication, demonstrating how early consideration of manufacturing challenges can accelerate the translation of laboratory prototypes to clinically deployable devices. Overall, addressing materials and fabrication scalability at the earliest stages of development is essential for bridging the gap between academic innovation and real-world deployment. For microfluidic POC cancer diagnostics to achieve clinical impact, analytical performance should be considered alongside manufacturability, reproducibility, and cost, ensuring that promising laboratory demonstrations can become robust, widely deployable diagnostic technologies.
4. Literature survey of emerging diagnostic technologies
4.1. Lateral flow assay (LFA), paper-based, and CRISPR technology-based diagnostics
One of the simplest, most affordable, and portable options for POC cancer diagnostics is the paper-based platform.300 Recent studies have applied new innovative strategies and designs to enhance the accuracy, sensitivity and simplicity of this platform for POC. Guo et al. developed an all-in-one pen-based paper chip (PPC) for detecting breast cancer-derived exosomal miRNA-21 (one of the most commonly upregulated miRNAs in cancers like lung, breast, and glioblastoma).298 This device integrates a lateral flow strip (LFS) with enzyme-free catalyzed hairpin assembly (CHA) amplification including two hairpin probes: FAM-labeled HP1 and biotinylated HP2 (Fig. 3a).298 In the absence of miR-21, HP1 and HP2 adopt stable stem-loop structures that prevent hybridization. After binding, miR-21 hybridizes with HP1, generating a single-stranded tail that promotes hybridization with HP2 to form double-stranded DNA.298 miR-21 is released from the HP1–HP2 complex, enabling effective recycling and signal amplification.298 The system had a limit of detection (LOD) of 25 fmol, showed high specificity against similar miRNAs, and successfully distinguished plasma from breast cancer patients and healthy controls, all by a compact pen-shaped, user-friendly format for a POCT device.298 Colorimetric assays also offer simple and widely used POC options. Zhang et al. introduced an innovative strategy for detecting methylation in miRNA by developing a DNAzyme-triggered rolling circle amplification (RCA) method for the identification of 5-methylcytosine-modified miRNA-21 (m5C-miRNA-21), a promising biomarker for colorectal cancer.299 In this study, the methylated miRNA is selectively cleaved by a DNAzyme, generating a fragment that serves as a primer for RCA (Fig. 3b).299 The resulting amplified products either induce gold nanoparticle (AuNP) aggregation, producing a visible red-to-blue color change with a LOD of 1 pM.299 This method can also be applied to a paper-based approach, by hybridizing with AuNP probes on a lateral flow dipstick for enhanced sensitivity, achieving an LOD of 0.1 pM.299
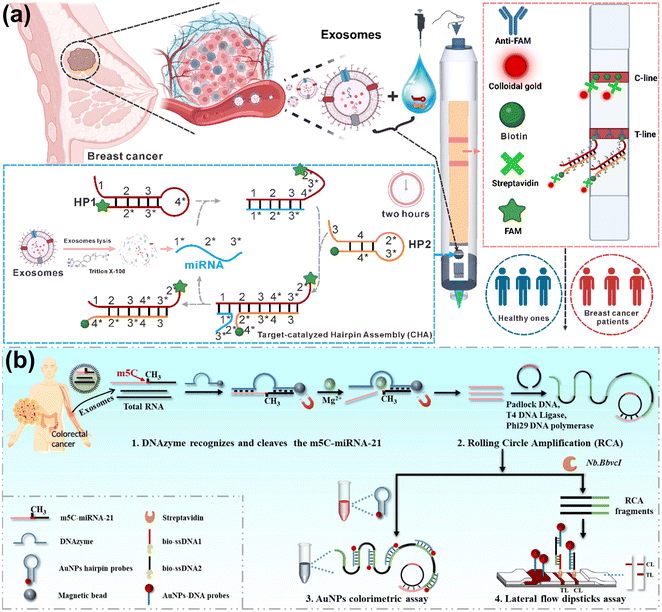 | ||
| Fig. 3 (a) Schematic illustration of the all-in-one pen-based paper chip (PPC) platform. Sample and reagents are added to the pen's reaction chamber, where enzyme-free catalyzed hairpin assembly (CHA) amplifies target miRNA-21. The target miRNA hybridizes with and opens the FAM-labeled hairpin probe 1 (HP1). The CHA reaction recycles the target miRNA, generating multiple HP1–HP2 duplexes for signal amplification. By pressing the pen, the reaction mixture is released onto the lateral flow strip, where the duplexes interact with gold nanoparticle–streptavidin conjugates to create a visible test line.298 Reproduced with permission from ref. 298. Copyright 2024, Royal Society of Chemistry. (b) Schematic illustration of the DNAzyme-RCA-based colorimetric and lateral flow dipstick assays for point-of-care detection of exosomal m5C-miRNA-21. m5C-miRNA-21 is recognized and cleaved by a DNAzyme. Cleaved fragments are primers to initiate padlock DNA cyclization, followed by rolling circle amplification (RCA). Detection step can be performed by colorimetric readout, where RCA products induce AuNP aggregation (red-to-blue color change), or by a lateral flow dipstick, where RCA fragments hybridize with AuNP–DNA probes to generate a visible test line. Reproduced with permission from ref. 299. Copyright 2024, Royal Society of Chemistry. | ||
CRISPR technology-based diagnostics are progressing rapidly in cancer mutation detection due to high specificity and sensitivity for identifying single-nucleotide or small insertion/deletion mutations.301,302 This method is now advancing toward two main trends: one-pot reactions and multiplex detection. One-pot systems perform CRISPR-based detection within a single reaction tube, minimizing handling steps, preventing contamination and reducing assay time. Li et al. developed a one-pot CRISPR-based platform for ROS1 fusions, which occur in approximately 2% of non-small cell lung cancer cases and are actionable with tyrosine kinase inhibitors.303 This system combines reverse transcription-recombinase polymerase amplification (RT-RPA) with Cas12a detection in a specially designed 3D-printed tube that physically separates the amplification and CRISPR reagents until amplification is complete, preventing premature cleavage (Fig. 4a).303 Detection is achieved via Cas12a trans-cleavage fluorescence at 40 °C, with a LOD of 5–10 copies per μL and high specificity against wild-type sequences.303 Clinical validation on FFPE samples (8 ROS1-positive, 12 ROS1-negative) demonstrated 87.5% sensitivity, 91.67% specificity, and an ROC AUC of 0.938.303 Multiplex platforms enable simultaneous detection of multiple targets or variant forms in a single assay, increasing throughput and clinical utility. The CoHIT platform, developed by Yin Liu and colleagues, integrates engineered Cas12a (enAsU-R) with enzymatic recombinase amplification (ERA) in a single-tube reaction for ultrasensitive detection of multiple same-site indels in the NPM1 gene.304 The enAsU-R nuclease exhibits high mismatch tolerance and broad PAM recognition (NNCV and TTTV), allowing a single crRNA to detect 14 different 4 bp insertions.304 In practice, patient genomic DNA is added to the reaction containing ERA reagents, primers, ssDNA fluorescent reporters, enAsU-R, and guide RNA.304 Amplification and Cas12a collateral cleavage occur at 39 °C, and the reaction can be completed within 30 minutes using a portable isothermal fluorescence detector (Fig. 4b).304 The system achieves LOD as low as 0.01% mutation frequency and successfully detects over 97% of known NPM1 4 bp insertions with a single crRNA.304
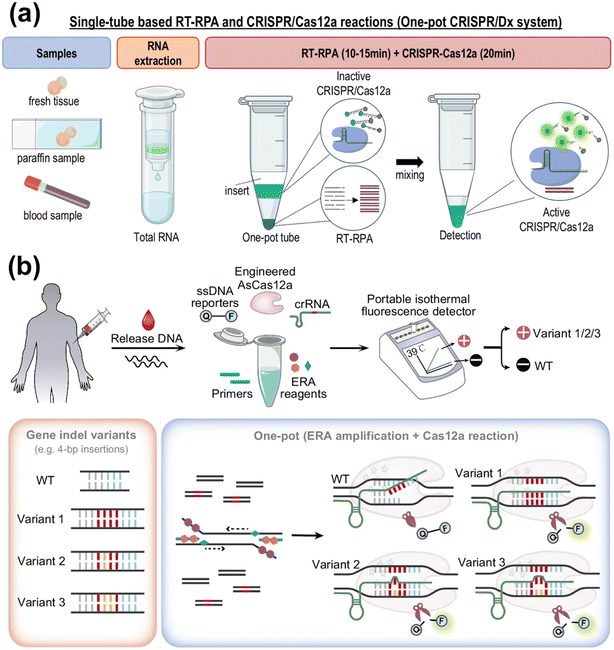 | ||
| Fig. 4 (a) Schematic illustration of the single-tube RT-RPA and CRISPR/Cas12a assay (one-pot CRISPR/Dx system) workflow. The RT-RPA reaction is initiated at the bottom of the tube, physically separated from CRISPR/Cas12a components. After amplification, the CRISPR/Cas12a mixture (initially contained in an insert) is released to the bottom of the tube by centrifugation, allowing amplicons to trigger Cas12a-mediated detection.303 Reproduced with permission from ref. 303. Copyright 2025, Royal Society of Chemistry. (b) Schematic illustration of the CoHIT system workflow. Genomic DNA was extracted from patient-derived cell samples and analyzed using a one-pot ERA–Cas12a reaction. The DNA is going under isothermal amplifon at 39 °C with ERA reagents, primers, FAM-ssDNA-BHQ1 reporters, engineered AsCas12a (enAsU-R), and a single crRNA. The crRNA binds variant 1–3 via sequence complementarity or mismatch tolerance, triggering Cas12a collateral cleavage and emitting green fluorescence, while wild type (WT) remain undetected. Detection completes within 30 min using a portable fluorescence detector.304 Reproduced with permission from ref. 304. Copyright 2024, Nature. | ||
Smartphone-assisted POC platforms are also developing due to their ability to combine portability with digital data capture. Yu et al. developed a split light-up aptamer assay for detecting methylation of the Septin9 gene (mSeptin9), a colorectal cancer biomarker.305 This system uses a methylation-sensitive restriction enzyme and isothermal amplification to produce single-stranded DNA that reassembles a split aptamer, which binds the dye auramine O to generate green fluorescence (Fig. 5).305 A smartphone camera captures the signal, achieving a detection limit of 0.769 nM within just 2–3 minutes (Fig. 5).305
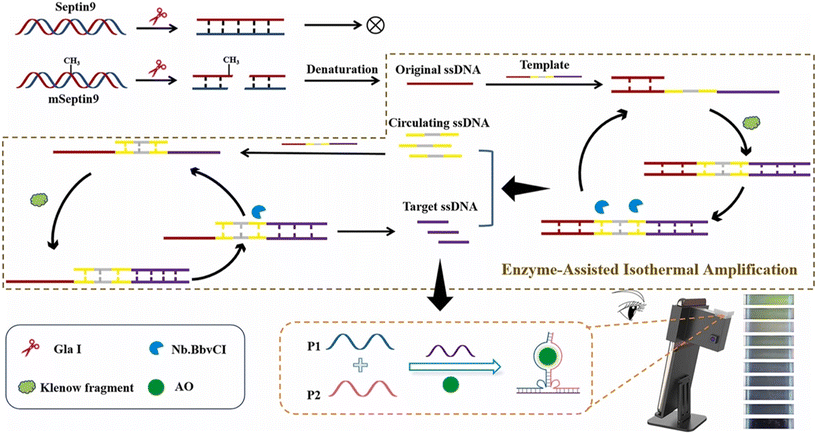 | ||
| Fig. 5 Schematic illustration of a smartphone-assisted POCT platform based on split light-up aptamer for mSeptin9 detection. mSeptin9 is cleaved by GlaI, generating primers for enzyme-assisted isothermal amplification. The resulting target ssDNA induces assembly of split DAP-10–42 aptamers, which activate auramine O fluorescence. The emitted signal is visualised under UV light and quantitatively analysed via smartphone imaging.305 Reproduced with permission from ref. 305. Copyright 2025, Elsevier. | ||
Biosensors are recommended to meet a variety of criteria for POC sensing. Many POC devices for viral detection follow ASSURED standards, which state that excellent POC diagnostics are ![[A with combining low line]](https://www.rsc.org/images/entities/char_0041_0332.gif) ffordable,
ffordable, ![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif) ensitive,
ensitive, ![[S with combining low line]](https://www.rsc.org/images/entities/char_0053_0332.gif) pecific,
pecific, ![[U with combining low line]](https://www.rsc.org/images/entities/char_0055_0332.gif) ser-friendly,
ser-friendly, ![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif) apid/Robust,
apid/Robust, ![[E with combining low line]](https://www.rsc.org/images/entities/char_0045_0332.gif) quipment-free and
quipment-free and ![[D with combining low line]](https://www.rsc.org/images/entities/char_0044_0332.gif) eliverable.322 REASSURED criteria also considers
eliverable.322 REASSURED criteria also considers ![[R with combining low line]](https://www.rsc.org/images/entities/char_0052_0332.gif) eal-time connectivity and
eal-time connectivity and ![[E with combining low line]](https://www.rsc.org/images/entities/char_0045_0332.gif) ase of specimen collection.323 For cancer diagnostics, prognostics, or screening applications, protein tests must be sensitive enough to detect low abundance in biofluids, particularly in early disease states, where concentrations might range from femtomolar to nanomolar.324 This is challenging for POC methods where the dynamic range may be limited. To increase the accuracy of cancer testing, screening technologies also require multiplexing to detect multiple proteins simultaneously, as a single biomarker often lacks predictive value.325,326 While few current assay methods for cancer diagnostics are able to meet all of these criteria, we will discuss novel methods for protein detection that have high potential for future POC applications in liquid biopsies.
ase of specimen collection.323 For cancer diagnostics, prognostics, or screening applications, protein tests must be sensitive enough to detect low abundance in biofluids, particularly in early disease states, where concentrations might range from femtomolar to nanomolar.324 This is challenging for POC methods where the dynamic range may be limited. To increase the accuracy of cancer testing, screening technologies also require multiplexing to detect multiple proteins simultaneously, as a single biomarker often lacks predictive value.325,326 While few current assay methods for cancer diagnostics are able to meet all of these criteria, we will discuss novel methods for protein detection that have high potential for future POC applications in liquid biopsies.
There are several lateral flow immunoassay methods in development to improve detection of proteins, which would make LFAs a more valuable tool for cancer diagnostics (Fig. 6). LFAs are rapid, low-cost, easy to use, and therefore are excellent options for point-of-care detection.327 When combined with strategies to enhance sensitivity and specificity,328 add quantitative analysis and multiplexing,329,330 and lower the LOD, LFAs can gain greater utility for cancer-related biosensing.331–333 Many of these strategies are covered in great detail in previous reviews.334–336 These can rely on enhanced labels for LFA assays, various imaging techniques, or may apply an isothermal amplification strategy before detection.337 Recently, Ye et al. used a gold-nanoparticle LFA paired with a portable reader to detect HER2-ECD protein, a biomarker for breast cancer, in serum.338 Nanozymes and other strategies such as nanoclusters have also been employed for the detection of EVs derived from breast cancer cells.339 Magnetic lateral flow biosensors are excellent at achieving higher sensitivity,340,341 such as in Ren et al., which reported detection of valosin-containing protein, a marker of cervical cancer, with limits as low as 25 fg mL−1.342,343 Ao et al. applied a near-infrared (NIR) fluorescence-based lateral flow assay with quantum dots to detect multiplexed proteins related to lung cancer.344 The numerous advancements in lateral flow immunoassays demonstrate high potential for future use in cancer diagnostics.
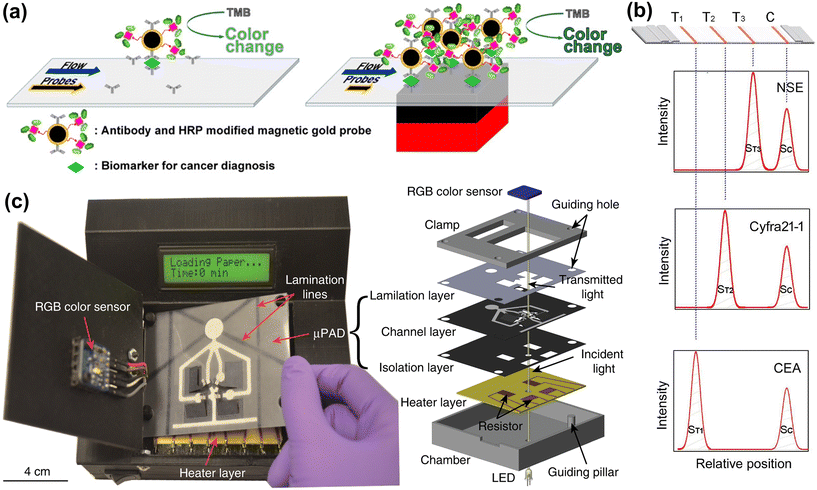 | ||
| Fig. 6 Novel paper-based protein detection methods. (a) Lateral flow assay utilizing magnetic enhancement. Reproduced with permission from ref. 342. Copyright 2019 American Chemical Society. (b) Multiplexed lateral flow immunoassay for lung cancer biomarkers with multiple test lines with quantum dots read out by a portable NIR fluorescence scanner. Reproduced with permission from ref. 344. Copyright 2022 Elsevier. (c) Microfluidic paper-based analytical device for multi-step immunoassays and colorimetric detection of tumor necrosis factor (TNF)-α. Reproduced with permission from ref. 345. Copyright 2019 Springer Nature. | ||
Another biosensing field with promising developments towards POC testing of proteins are microfluidic paper-based analytical devices (μPADs), which are low-cost and easy to operate. Many μPAD technologies have already been described for POC sensing and cancer diagnostics in other reviews.346,347 Unlike the absorbent pads of LFAs, μPADs use small microfluidic channels in paper to direct small liquid samples to different detection zones. These channels may be two-dimensional or three-dimensional throughout multiple layers of paper, allowing multi-step assay design. Detection of proteins through μPADs can be improved through different binding moieties, advanced imaging techniques, signal amplification before detection, and improvement of the fabrication process.345,348,349 A novel nano-ink was designed for μPAD using silver nanoparticles and graphene oxide for the detection of ovarian tumor protein CA 125.350 Paper combined with a PMMA paper-in-polymer pond was used to perform a sandwich immunoassay for multiplexed detection of protein cancer biomarkers.351 Wang et al. demonstrated a 3D paper analytical device for extracellular vesical profiling for PSMA, PSA, PTK-7, EpCAM and EGFR proteins to differentiate between healthy patients or patients with prostatitis and those with prostate cancer.352 Different binders such as aptamers or nanobodies have also been used for μPADs, increasing versatility and ease of labeling.353,354
There are many novel methods currently under development for protein detection that have the potential to achieve the sensitivity necessary for cancer diagnostics and screening tests while still meeting POC criteria. Some rely on “proximity” assay methods that convert protein targets to a nucleic acid signal before amplification,355,356 while other applications focus on amplifying the signal of individual proteins (Fig. 7).357–359 In a proximity assay, two antibody–oligonucleotide conjugates react with a target protein to transduce the signal into a nucleic acid that can be further amplified. While standard proximity assays rely on qPCR and thermocycling, making them less suitable for POC testing,360 variations have been developed using recombinase polymerase amplification (RPA), CRISPR enzymes,361,362 or catalytic hairpin assembly (CHA)363,364 for POC detection of proteins. In Tang et al. proximity-initiated transduction to a nucleic acid signal was combined with isothermal exponential amplification (EXPAR) and CHA for detection on a lateral flow assay in 5 minutes, with protein limits of detection down to 1 fM.365 Other assays rely on split enzymes such as a T7 RNA polymerase-linked immunosensing assay (TLISA), where the target antigen causes polymerase reassembly, and could reach nM limits of detection.366 These methods represent homogeneous assays that can be highly multiplexed and run at room temperature or low isothermal heating, making them applicable for ultrasensitive POC cancer diagnostics.
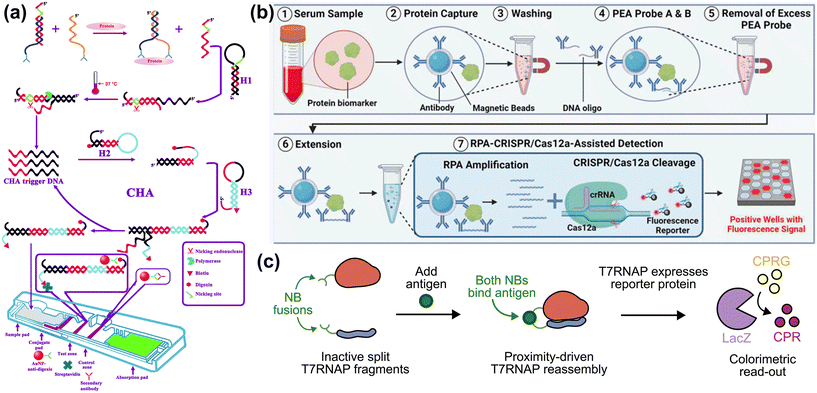 | ||
| Fig. 7 Current novel methods of protein detection for potential point-of-care applications. (a) Proximity-based catalytic hairpin amplification with detection on a lateral flow biosensor. Reproduced with permission from ref. 365. Copyright 2022, Royal Society of Chemistry. (b) A proximity assay using RPA and CRISPR/Cas12a cleavage for protein biomarker detection in serum, adapted from Shao et al. Reproduced with permission from ref. 361. Copyright 2024 IEEE Sensors. (c) Protein detection using split T7 RNA polymerase with colorimetric readout. Reproduced with permission from ref. 366. Copyright 2025 Science Advances. | ||
Emerging paper-based, lateral flow, and CRISPR-enabled platforms (section 4.1) directly intersect with the disease-specific constraints identified in section 2. These technologies address critical clinical needs such as rapid turnaround, minimal invasiveness, and limited sample requirements, which are particularly salient for early OSCC screening and NSCLC mutation profiling (sections 2.2 & 2.5). For example, an all-in-one pen-based paper chip for miRNA-21 leverages enzyme-free amplification to achieve femtomolar sensitivity in a compact format, illustrating how design innovation can bridge low-abundance biomarker detection with user-friendly operation. Similarly, one-pot CRISPR platforms achieving detection at single-digit copies per microliter demonstrate promising sensitivity for actionable mutations like ROS1 and NPM1 insertions relevant to targeted therapy decisions. Smartphone-assisted assays for methylated gene markers (e.g., mSeptin9) further exemplify how connectivity and portable imaging can support decentralized decision-making in real-world clinical workflows. These modalities align with broader trends in POC molecular diagnostics, including isothermal amplification and lab-on-chip miniaturization, that are driving market growth and encouraging integration of multiplex detection with simplified readouts. Collectively, these approaches suggest a feasible path toward clinical implementation for cancers where rapid, accurate molecular insights are needed at or near the point of care.
4.2. Electrochemical technologies for POC cancer diagnostics
Electrochemical biosensors have emerged as a promising platform for POC molecular diagnostics of cancer due to their high sensitivity, simplicity, low cost, and potential for miniaturization. These sensors convert biological interactions into measurable electrical signals, enabling the rapid and accurate detection of cancer-related biomarkers.367,368 Their compatibility with microfluidic systems and smartphone integration makes them especially attractive for resource-limited settings. The general working mechanism of an electrochemical sensor is demonstrated in Fig. 8a. There is a biological recognition element on the electrode surface, and the transduction signal can be generated with different mechanisms, such as the changes in the electron transfer rates, electrochemical potentials, charging/discharging of the capacitive elements, conductance, and impedance upon binding the target to a specifically modified surface.367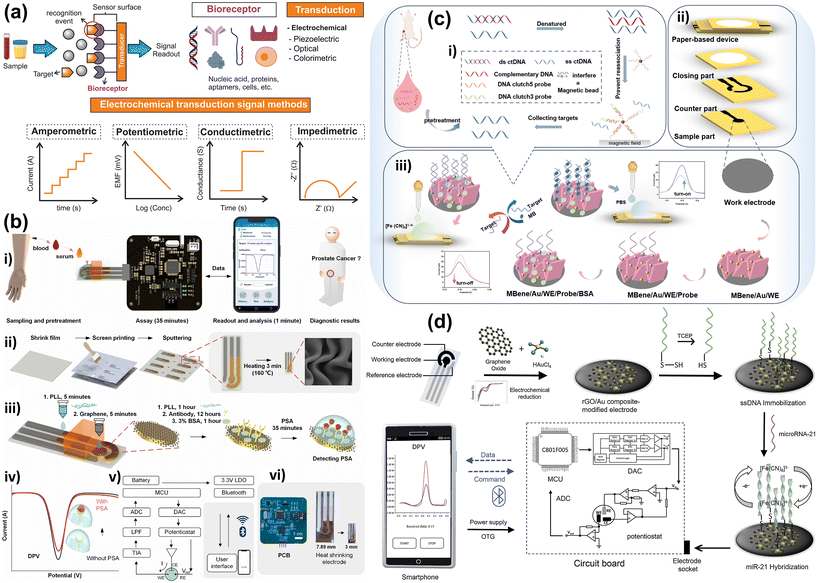 | ||
| Fig. 8 Different applications of electrochemical sensors for POC cancer diagnostic. (a) Schematic of electrochemical transduction signal methods and different types of electrochemical signals. Reproduced with permission from ref. 367. Copyright 2022, Elsevier. (b) PSA detection for prostate cancer diagnosis with the POC approach. The workflow for POC PSA detection (i), shrink polymer electrode manufacturing (ii), the modification of the sensor surface with recognition agents (iii), signal change depending on PSA binding status (iv), block diagram of the POC system (v), and photo of the printed circuit board and shrinking electrode (vi). Reproduced with permission from ref. 370. Copyright 2023, Elsevier. (c) The detection of ctDNA with an electrochemical paper-based analytical device. The sample pretreatment step (i), the electrochemical sensor (ii), and the detection procedure of ctDNA on the proposed sensor (iii). Reproduced with permission from ref. 372. Copyright 2024, Elsevier. (d) Schematic of the smartphone-based electrochemical biosensor. The biosensing system features a reduced graphene oxide/gold (rGO/Au) composite-modified electrode. A custom Android application on the smartphone allows for device control, data acquisition, and transmission. Reproduced with permission from ref. 373. Copyright 2020, Elsevier. | ||
In recent years, several electrochemical sensor-based devices have been developed that approach or reach clinical and commercial use, indicating their growing impact in the field of POC diagnostics for early cancer detection and monitoring. Chakraborty et al. used a flexible PET-based electrochemical aptasensor for carcinoembryonic antigen (CEA) detection, which is a glycoprotein in blood and might be used as an indicator for tumor screening.369 In this study, a dielectrophoresis-assisted graphene deposition followed by ZnO nanorod growth on a screen-printed electrode was performed. The graphene/ZnO nanorod architecture, functionalized with DNA aptamers via silanization, enabled highly specific and stable target recognition. A smartphone-controlled handheld potentiostat was integrated for pulse application, electrochemical impedance measurements, and on-site quantitative readout. This portable, low-cost platform demonstrates strong potential for POC CEA detection. However, the necessity for an external electric field and the complexity of electrode fabrication should be taken into consideration for mass production. Moreover, long-term aptamer stability under varying storage conditions requires further investigation before clinical adoption. Researchers have also explored electrochemical POC platforms for other cancer biomarkers. One representative example is prostate-specific antigen (PSA), a key indicator for prostate cancer. He et al. presented POC detection of prostate cancer by using PSA and an immunosensor integrated into a miniaturized electrochemical platform, as shown in Fig. 8b.370 A gold-coated shrinkable polymer electrode, forming nano–micro wrinkles upon shrinking, increased the binding surface area, while plasma treatment and graphene modification enhanced sensitivity. The portable system enabled detection in 20 μL of serum within 35 minutes, achieving a low detection limit of 0.38 fg mL−1. It showed reliable performance in clinical samples, comparable to commercial instruments. On the other hand, sputtering techniques and controlled thermal shrinking may present challenges during manufacturing. Also, the sensor's performance under different environmental and handling conditions should be evaluated carefully, which is of utmost importance for POC applications. While technical issues such as fabrication and environmental stability are critical, ensuring diagnostic value requires the incorporation of well-established biomarkers. Cancer antigen 15-3 (CA 15-3) is a glycoprotein produced by certain cells and can be used as a tumor biomarker found in body fluids for breast cancer. Pacheco et al. reported a molecularly imprinted polymer-based voltammetric sensor for the detection of CA 15-3. They directly imprinted CA 15-3 on a screen-printed gold electrode using electropolymerization.371 The peak current intensity of the redox probe and biomarker concentration show a linear relationship. The compatibility of screen-printed electrodes with compact electrochemical devices makes them particularly appealing for POC applications.
Nucleic acid-based biomarkers, including DNA and RNA, play a crucial role in cancer detection and response to treatment as well.374 ctDNAs which are released from tumor cells into the blood, carry important information about the mutations contained in their cell of origin. Liquid biopsy enables the non-invasive detection of ctDNA, providing valuable insights into tumor genetics.375 Following the approval of the first FDA-authorized liquid biopsy test, the cobas EGFR Mutation Test v2 (Roche Molecular Systems, Inc.), the number of studies focusing on ctDNA detection through liquid biopsy has increased substantially.376 Yang et al. developed an electrochemical paper-based device for the detection of ctDNA. In their study, they used a magnetic clutch probe to enrich ctDNA from tumor-carrying mice serum samples and MBene/Au for sensing, as depicted in Fig. 8c.372 MBene/Au not only shows a synergistic effect for signal enhancement but also provides more area for DNA probe immobilization. The electrochemical assay with turn-on and turn-off strategies shows a linear response in the concentration range of 1–50 pM, achieving limits of detection of 178 fM and 216 fM, respectively. There are also some studies that perform the detection of cancer by using alternative forms of circulating DNA beyond ctDNA. For instance, Keyvani et al. performed an integrated microfluidic electrochemical-based sensor for POC cervical cancer detection using human papilloma virus (hr-HPV) circulating DNA in whole-blood samples.377 The hr-HPV16 circulating DNA can be used as a marker of cervical cancer. The device is composed of two main parts: an automatic plasma separator to isolate plasma and a graphene oxide-based electrochemical sensor. The LOD was determined to be 0.48 μM (∼109 copies per mL). However, the concentration of hr-HPV16 circulating DNA in the plasma of cervical cancer patients has been reported to be approximately 109 copies per mL. Therefore, the system should be further improved by integrating DNA amplification or enhancing the sensitivity of the electrochemical sensor.
Several commercially available POC devices for molecular diagnostics have been developed for cancer. For example, Idylla®, developed by Biocartis, can detect mutations by using small tissue sections or plasma volumes with a simple workflow.378 Enabling clinical decision-making in oncology settings. Microribonucleic acids (miRNA) also play a key role in the molecular detection of cancer. Low et al. developed a portable, smartphone-based biosensor for the detection of miRNA-21, which is upregulated in non-small cell lung cancer in saliva.373 The device is composed of a reduced graphene oxide/gold (rGO/Au) composite and a circuit board (Fig. 8d). On the rGO/Au-modified electrode, hybridization of the miRNA-21 target with the ssDNA probe led to a concentration-dependent reduction in the peak current. Detection of miRNA-21 using the smartphone-based biosensing system covered concentrations between 1 × 10−4 M and 1 × 10−12 M, yielding results consistent with a commercial electrochemical workstation, as indicated by a high correlation coefficient (R2 = 0.99). Furthermore, the recovery rate is found to be 96.2–107.2% in spiked artificial saliva, and the detection limit is 1 pM. Beyond these examples, the literature includes related works as summarized in recent review articles.379,380
Electrochemical biosensing platforms (section 4.2) offer quantitative, high-sensitivity detection that maps directly onto several clinical constraints in section 2, especially for prostate cancer and NSCLC liquid biopsy contexts. Their ability to detect PSA, CEA, and circulating nucleic acids, including ctDNA, from microliter-scale samples responds to the challenge of limited sample volume and low analyte abundance in mCRPC and NSCLC (sections 2.4 & 2.5). The integration of electrochemical sensors with microfluidics and smartphone interfaces demonstrates how miniaturized systems can deliver actionable signals without the delays and infrastructure needs of centralized labs, addressing market demands for decentralized diagnostics highlighted in recent industry analyses. In particular, electrochemical paper-based devices that enrich ctDNA and generate fM-level analytic signals illustrate how biosensor innovation can achieve both sensitivity and usability. While fabrication complexity, environmental stability, and electrode longevity remain translational challenges, these technologies show how quantitative electrical transduction mechanisms can be adapted for diverse biomarker classes. This broad spectrum, from enzyme-free voltammetry to impedance spectroscopy for miRNA and protein targets, underscores the versatility of electrochemical platforms in fulfilling clinical use cases for screening, prognostication, and therapeutic stratification across OSCC, prostate cancer, and lung cancer populations.
4.3. Optical technologies for POC cancer diagnostics
Optical biosensing represents one of the most widely explored strategies for POC cancer diagnostics due to its high sensitivity, rapid signal transduction, and compatibility with portable instrumentation.77,381,382 These methods exploit diverse light–matter interactions, ranging from simple color changes visible to the naked eye to advanced nanophotonic phenomena capable of single-molecule detection.383–386 The versatility of optical biosensors allows detection of a broad spectrum of cancer biomarkers, including proteins, nucleic acids, EVs, and CTCs.387 Importantly, optical readouts can often be miniaturized and integrated with smartphone-based devices,388,389 making them highly attractive for decentralized cancer testing.390–393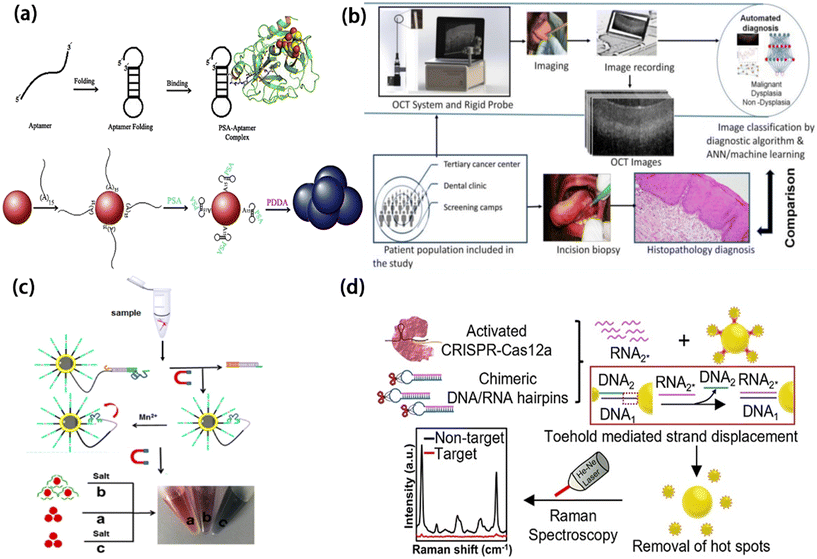 | ||
| Fig. 9 (a) Aptamer binding to prostate-specific antigen (PSA). The aptamer folds into a 3D structure under optimized conditions and interacts with PSA to form a stable complex. Reproduced with permission from ref. 396. Copyright 2020, Elsevier. (b) Combination of a magnetic 3D DNA walker with unmodified AuNPs for colorimetric detection of microRNA. Reproduced with permission from ref. 397. Copyright 2021, Elsevier. (c) Portable optical coherence tomography (OCT) system for imaging oral mucosal lesions; OCT images were analyzed by automated algorithms and compared with histological diagnosis. Reproduced with permission from ref. 399. Copyright 2021, MDPI. (d) CRISPR–Cas12a-assisted SERS biosensor employing chimeric DNA/RNA hairpins (displacers). Target-induced Cas12a activation releases RNA that triggers toehold-mediated strand displacement, leading to disassembly of core–satellite AuNP nanoclusters with Raman tags and a sharp decrease in SERS intensity. Reproduced with permission from ref. 403. Copyright 2021, Ivyspring International Publisher. | ||
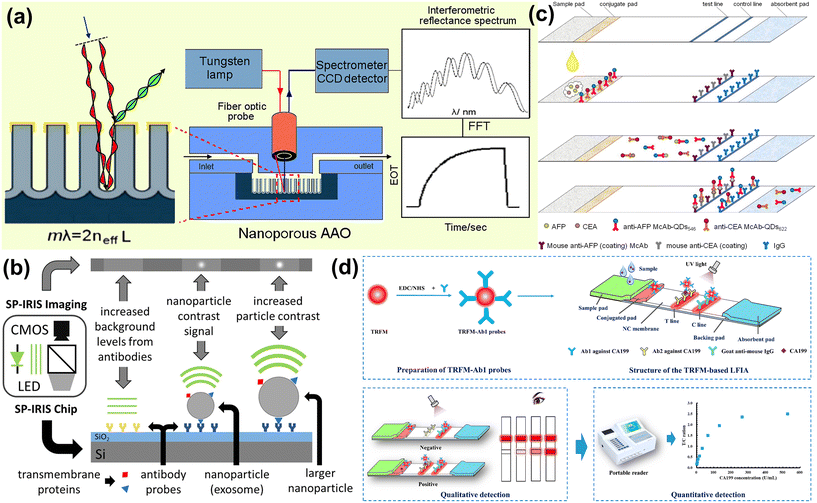 | ||
| Fig. 10 (a) RIfS microchip biosensor comprising an anodic aluminum oxide (AAO) nanopore integrated with a microfluidic chip; binding-induced shifts in Fabry–Perot interference fringes were analyzed by FFT to yield effective optical thickness (EOT) as the detection signal. Reproduced with permission from ref. 406. Copyright 2012, Elsevier. (b) Principle of SP-IRIS: monochromatic LED illumination of the sensor surface generates interferometrically enhanced nanoparticle scattering signatures captured on a CMOS camera. Reproduced with permission from ref. 407. Copyright 2022, MDPI. (c) Multicolor quantum dot (QD)-based ICTS for simultaneous detection of AFP and CEA; QD–Ab conjugates bind targets and produce distinct fluorescence signals at the test lines. Reproduced with permission from ref. 408. Copyright 2015, Elsevier. (d) Time-resolved fluorescence microsphere (TRFM)-based LFIA for rapid qualitative and quantitative detection of CA 19-9. Reproduced with permission from ref. 409. Copyright 2023, Springer Nature. | ||
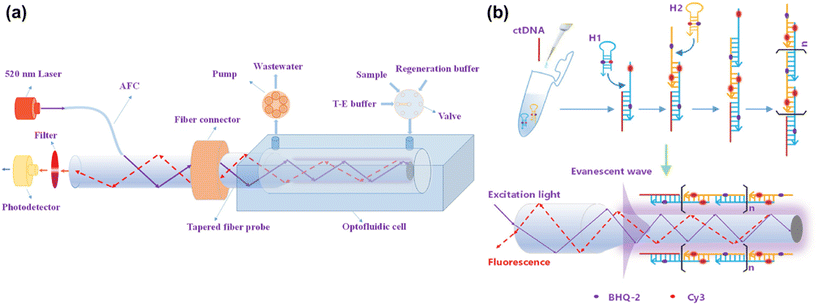 | ||
| Fig. 11 Design and operation of the HCR–LOFD system. (a) Schematic of the lab-on-fiber platform, comprising a 520 nm laser, asymmetric Y-branch fiber coupler (AFC), optofluidic unit, and photodetector. (b) Target ctDNA hybridizes with hairpin H1, triggering HCR with hairpin H2 to form nicked DNA helices. This structural change separates fluorophore–quencher pairs, reducing FRET and restoring Cy3 fluorescence within the evanescent field. Emitted fluorescence is coupled back through the AFC to a photodiode for end-point or real-time detection. The system enables quantitative ctDNA detection within 6 min at room temperature, directly from diluted whole blood.414 Reproduced with permission from ref. 414. Copyright 2023, Elsevier. | ||
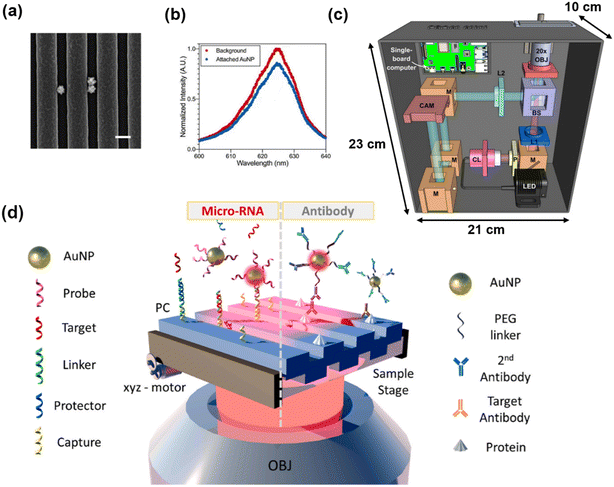 | ||
| Fig. 12 Photonic resonator absorption microscopy for point-of-care biosensing. (a) Principles of PRAM operation using photonic crystals (SEM scale bar 200 nm) and (b) nanoparticles coupled with a red LED. Reproduced with permission from ref. 427. Copyright 2022, Elsevier. (c) Miniaturized PRAM-mini setup and dimensions for point-of-care detection, adapted from Khemtonglang et al. Reproduced with permission from ref. 424. Copyright 2024, Optica Publishing Group. (d) PRAM assays for testing microRNA and proteins, adapted from Liu et al. Reproduced with permission from ref. 423. Copyright 2024, Elsevier. | ||
Representative examples of optical biosensing modalities (section 4.3) capture a wide variety of methods such as colorimetric, OCT, SERS, RIfS, fluorescence, and photonic crystal-based detection methodologies that collectively address different facets of the clinical gaps identified in section 2. The simplicity and visual interpretability of colorimetric assays create low-barrier tools for early OSCC screening (section 2.2), while label-free, high-resolution imaging such as portable OCT offers non-invasive structural insights complementary to molecular data in tumor margin assessment. Advanced optical techniques like SERS and fluorescence provide ultra-high sensitivity needed to detect rare ctDNA variants or EV signatures crucial for HCC and NSCLC subgroup stratification (sections 2.3 & 2.4). Photonic crystal-enabled digital counting methods, as shown in Fig. 12, illustrate how nanophotonic resonance can be harnessed for multiplexed detection of miRNAs and proteins with digital resolution, pointing toward solutions for multiplexed identification and detection in heterogeneous cancers like PCa and HCC. These optical platforms are often compatible with portable, camera-based readouts and map onto broader clinical trends toward decentralized, rapid phenotypic and molecular characterization. While challenges remain in substrate reproducibility and user-independent interpretation, the integration of optical biosensing with computational analytics offers a compelling strategy to meet the high analytical and operational standards demanded for real-world POC oncology diagnostics.
While a wide array of emerging POC cancer detection technologies exists, their translation into routine clinical practice remains limited. The comparative table highlights key modalities, from lateral flow assays, CRISPR-based nucleic acid detection, and electrochemical sensors to advanced optical platforms such as OCT, SERS, and photonic crystal-based digital detection, along with their POC advantages, implementation barriers, and clinical status. Despite strong analytical performance in controlled settings, challenges such as low biomarker abundance, sample complexity, assay standardization, multiplexing limitations, and regulatory hurdles hinder widespread adoption. Centralized laboratory methods continue to dominate due to established sensitivity, reproducibility, and clinical reliability. Table 3 is intended as a representative, critical comparison rather than an exhaustive catalog. Across modalities, real clinical sensitivity and specificity remain foremost challenges; many POC platforms perform well in controlled labs but falter with complex matrices like blood or saliva. Pre-analytical complexity, reagent stability, workflow standardization, multi-analyte multiplexing, clinical trust, and regulatory requirements further limit decentralized adoption. These realities explain why many promising technologies remain in proof-of-concept or early validation phases rather than routine clinical use.
| Sl. no. | Technology | Advantages | Challenges/barriers | Clinical translation status |
|---|---|---|---|---|
| 1 | Lateral flow/paper-based | Very low cost, rapid, easy to use, highly portable; scalable for resource-limited settings; compatible with simple amplification strategies | Intrinsically limited sensitivity and dynamic range for low-abundance cancer biomarkers; difficult to achieve robust multiplexing; user interpretation variability; often proof-of-concept only despite enhancements (nanoparticles, coatings) | Many pilots; few broad cancer applications in clinical use |
| 2 | CRISPR-based NA detection | Very high specificity and potential for amplification-free workflows; suited for multiplexed detection; programmable and adaptable to new targets | Complex sample prep and reagent stability issues; cold-chain requirements; multi-step reactions; limited standardized protocols; real-world robustness unclear | Early clinical validations; translation ongoing |
| 3 | Electrochemical sensors | High sensitivity, quantitative output, smartphone compatibility; integrates well with microfluidics; amenable to miniaturization | Fabrication complexity; environmental noise and calibration issues; robust analytical performance in real biofluids still limited; need large-scale clinical validation | Promising prototypes; few widespread commercial products |
| 4 | Colorimetric/simple optical sensors | Intuitive visual readout; minimal instruments needed; rapid results; scalable and low-cost | Limited dynamic range for low-abundance targets; often can't reach clinical sensitivity needed for early cancer detection | Some niche use; broader translation limited by performance |
| 5 | Optical coherence tomography (OCT) | Real-time, label-free imaging; excellent for structural/lesion characterization; portable devices emerging | High equipment cost; specialized training required; not molecularly specific; often complements, not replaces, centralized diagnostics | Clinically used mainly for structural imaging |
| 6 | Surface-Enhanced Raman Spectroscopy (SERS) | Ultra-high analytical sensitivity; can detect ctDNA/EVs at very low levels; multiplexable; label-free options | Substrate reproducibility and biological fluid complexity challenge clinical consistency; specialized interpretation needed | Active research; limited translation |
| 7 | Photonic crystal-based digital detection | Highly sensitive digital counting; low sample volumes; compatible with multiplexed POC detection | Fabrication and alignment complexity; limited commercial availability; requires readout instrumentation | Proof-of-concept; early translational studies |
In summary, the technologies surveyed in section 4, spanning paper-based assays, CRISPR-enhanced nucleic acid detection, electrochemical sensors, and advanced optical modalities, collectively demonstrate how modern biosensing platforms are converging toward solutions for the specific clinical constraints identified in section 2. Many paper-based and LFA systems achieve rapid, minimally invasive readouts suitable for early screening scenarios such as OSCC or NSCLC mutation triage. CRISPR-integrated workflows demonstrate the potential for high specificity and low copy number detection, a capability that directly addresses low-abundance biomarkers in ctDNA and microRNAs, critical for precision stratification in prostate cancer and lung cancer therapy decisions. Electrochemical platforms balance quantitative performance and small sample volumes, aligning with the needs for serial liquid biopsies and frequent monitoring in metastatic prostate cancer and molecular subtype surveillance in HCC. Optical techniques, from colorimetric assays that offer simple visual cues to SERS and photonic crystal technologies that push analytical sensitivity to the attomolar range, further expand the toolkit for detecting diverse analytes including proteins, exosomes, and nucleic acids. Many of these technologies already demonstrate components of ASSURED/REASSURED criteria (e.g., rapidity, affordability, user friendliness), and smartphone or camera-linked readouts promise seamless integration with clinical workflows for decentralized deployment.
It is important to acknowledge that the landscape of POC cancer diagnostics extends beyond the specific platforms reviewed here. High-sensitivity sequencing-based liquid biopsy methods such as CAPP-Seq, immune-cell-based biomarker assays (e.g., EDIM), plasmonic and SPR enhancements (including graphene-augmented SPR), magnetic modulation biosensing, high-resolution ultrasound imaging, AI-informed decisions and sophisticated lab-on-chip microfluidic systems are among powerful approaches reported in the literature but not covered in this review.252,379,430–440 These techniques illustrate alternative paradigms (e.g., label-free, cascade amplification, real-time imaging) that have strong translational promise and may complement or synergize with the platforms presented here. While it is beyond the scope of this review to exhaustively survey all existing technologies, the representative examples selected collectively highlight both the current state of the art and the trajectory toward clinically impactful, integrated POC solutions capable of meaningfully addressing the disease-specific diagnostic needs described in section 2.
5. Challenges for regulatory approval of POC diagnostics
Moving any diagnostic technology used to make health care decisions from concept to clinical use requires a regulatory strategy that defines not only whether a technology platform has sound physics and chemistry principles, but also the manufacturability, quality control, documentation, reproducibility, and quantitative accuracy to ensure that the information provided is valid and reliable enough to be incorporated into medical practice.441–443 For platforms built on unconventional physical principles or designed to detect emerging molecular targets, regulatory planning is integrated into product development at an early stage to ensure that it will function as expected in real-world settings.444–446The U.S. Food and Drug Administration (FDA) approve devices for commercial use. The FDA defines how credibility is established, how risk is assessed, and how innovation is translated into clinical relevance. For molecular diagnostic technologies, regulatory classification sets a standard for evidence generation that requires substantial financial resources. Classification hinges on a device's potential risk and the degree to which it introduces new technology.447,448 Those deemed low risk are designated as class I and are often allowed to bypass premarket evaluation under the framework of general controls.449,450 Most diagnostics, however, are classified as class II and follow the 510(k) pathway, which requires demonstrating substantial equivalence to a legally marketed device in both intended use and core technological characteristics.449,451 While this framework supports iterative innovation, it poses clear constraints: molecular diagnostics technologies, such as POC approaches utilizing detection methods and assay protocols that differ fundamentally from well-established laboratory methods may not qualify.
For platforms that deviate from conventional electrochemical, enzymatic, or immunologic formats, a predicate device may not exist. In those cases, equivalence becomes a regulatory dead end. For novel, low- to moderate-risk technologies, the De Novo pathway provides a more appropriate mechanism for FDA review.452,453 For diagnostic technologies that fall outside existing regulatory templates where no predicate device captures the mechanism, form factor, or intended use, the De Novo pathway provides a means to establish legitimacy from first principles.454 The De Novo pathway to regulatory approval enables the FDA to assess novel technologies based on their individual risk profile, rather than through forced comparisons, provided that the device's safety can be reasonably assured through well-defined engineering and procedural controls. A successful De Novo submission also establishes a new regulatory pathway for future devices.455–457 When supported by updated clinical evidence or real-world performance data, developers may petition the FDA for risk class reclassification, creating a more predictable and expedited route for similar devices that follow. For developers introducing first-in-class systems, this route offers a balance that requires the same level of analytical validation expected in a 510(k), but adds further scrutiny around risk mitigation, labeling, and system-level quality assurance. Since De Novo review is based on an independent assessment of risk rather than alignment with a predicate, it often allows developers to advance without the scale of clinical evidence typically demanded for high-risk devices under the premarket approval (PMA) process.452,458–460
In contrast, when a device is intended to inform high-stakes medical decisions where the consequences of failure are serious or irreversible, the FDA assigns it to the class III category and evaluates it through the PMA process, representing the most demanding and data-intensive route.454,461 For technologies that aim to detect unestablished biomarkers with significant clinical consequences, particularly in complex fields such as cancer and neurodevelopment, the PMA route remains the principal option. PMA demands thorough testing and clinical evidence that confirms the device will serve patients without introducing unacceptable risk. When clinical data are needed to support a regulatory submission, particularly for novel or high-risk diagnostics, developers must first secure an investigational device exemption (IDE).462 The IDE enables investigators to collect human clinical data using a device that has not yet received FDA approval, provided the study is governed by a rigorously designed protocol, reviewed and approved by an institutional review board, and executed under Good Clinical Practice standards, such as subject protection and data credibility, which are central to IDE approval. While resource-intensive, IDE studies can yield the type of prospective, multicenter evidence that strengthens FDA submissions and builds early momentum for payer engagement and clinical adoption.463,464
Recognizing the potential impact of transformative technologies, the FDA has introduced expedited mechanisms to support a path to market. Among them, the Breakthrough Devices Program offers a formalized track for technologies that address unmet clinical needs, creating a structure for early and continuous dialogue with reviewers and enabling faster review timelines for devices that offer measurable improvements over current options.465,466 Biomolecular diagnostic technologies granted this designation gain early input on study design, streamlined feedback on submission strategy, and direct access to regulatory guidance – all of which help reduce uncertainty and improve investor confidence. Although the program does not relax evidentiary standards, it fosters a more agile and transparent process.
The strength of the data package remains constant across all regulatory routes. Validation studies must show not only sensitivity and specificity but also consistent performance across a range of operators, sample types, and environmental contexts encountered in clinical settings.467 For POC platforms, usability testing plays a central role in confirming that the device can be operated safely and effectively by its intended users across a range of clinical environments. For POC diagnostic technologies that are multiplexed or quantitative, especially those targeting small RNAs or other low-abundance biomarkers, additional rigor is required around normalization strategies and assay linearity.468 Clinical validation must demonstrate that the technology performs reliably within the target population and intended clinical setting. The most persuasive data emerge from prospective, blinded studies embedded in real diagnostic workflows, where device performance is evaluated directly against recognized clinical benchmarks.
As the POC diagnostic field moves steadily toward individualized and anticipatory care, robust evidence becomes indispensable not only for FDA approval but also for convincing clinicians and payers of the technology's value. Alongside these submission-specific demands, all devices are subject to a foundational set of requirements referred to as general controls.469,470 These include establishment registration, product listing, accurate labeling, and compliance with the FDA's Quality System Regulation outlined in 21 CFR Part 820.471 Diagnostic developers, even at the prototype stage, must plan for quality management systems that include design controls, risk management, document traceability, and change control.
For academic teams considering commercialization, this often requires establishing a commercial entity that can take over the responsibility for building quality control infrastructure from the ground up or working with manufacturing partners who can navigate both engineering constraints and regulatory expectations. These tasks are important, and not generally conducted by graduate students who are focused upon completing coursework and evaluating scientific hypotheses that are hallmark of PhD theses. Overlooking this foundation not only complicates scale-up but risks noncompliance during FDA inspections, an avoidable setback that can stall momentum just as a technology is ready to launch. Importantly, regulatory responsibility does not end at clearance. Post-market oversight is a critical extension of the FDA's regulatory role, particularly for diagnostic technologies intended for widespread or decentralized use. Once a device is cleared and in clinical use, manufacturers must remain engaged by tracking performance in real-world settings, reporting adverse events through the FDA's Medical Device Reporting system, and initiating corrective actions – such as updates to labeling, safety communications, or product recalls – if new evidence suggests patient risk.471–473 In molecular diagnostic markets characterized by rapid iteration and real-time updates, post-market systems must be built to both report and learn from field experiences, ideally using structured performance analytics that inform next-generation improvements.
Early engagement with the FDA through the Q-Submission (Q-Sub) program can provide streamline the approval process.473 These pre-submission meetings provide POC diagnostic developers with an opportunity to present their device concept, seek clarification on specific regulatory questions, and align with the FDA on evidentiary expectations before initiating resource-intensive validation studies.
Securing FDA clearance does not complete the regulatory process; additional steps are required. For diagnostic platforms intended for use in decentralized settings or certified laboratories, developers must determine whether additional regulatory mechanisms—such as the Clinical Laboratory Improvement Amendments (CLIA) waiver or Laboratory Developed Test (LDT)—are required to support real-world deployment.474,475 A CLIA waiver enables diagnostic use outside of high-complexity labs, but only when a device is shown to be simple to operate, analytically robust, and minimally susceptible to user error.473,476 Meeting this standard requires studies that assess reproducibility across untrained users, operational variability, and comparison with validated laboratory methods. These expectations influence both system design and validation strategy from the outset and should be treated as integral to the product development process.
In contrast, some diagnostic technology developers may consider offering their assay as an LDT, designed, manufactured, and used within a single high-complexity clinical laboratory certified under the CLIA framework.477,478 The FDA historically exempted LDTs from premarket review under enforcement discretion, provided they remained within a single certified laboratory. That policy has now shifted.479 In April 2024, the FDA finalized its long-anticipated rule withdrawing enforcement discretion for LDTs. Under the new policy, most LDTs are required to follow the same regulatory controls as commercial diagnostics, in response to increasing use of advanced methods with direct patient impact.480 These include premarket review (unless exempt), compliance with quality systems, and post-market responsibilities. Though the rollout will accommodate legacy and low-risk tests, the LDT pathway now demands the same regulatory rigor as commercial assays.
Designing POC cancer diagnostic tests for clinical translation also requires confronting the geographic disparities that shape healthcare delivery. Many technologies originate in high-resource research environments, but their most meaningful applications often emerge in areas where diagnostic infrastructure is limited. For platforms intended for global health or deployment in underserved regions, the regulatory strategy must anticipate constraints in workforce training, environmental stability, and supply chain resilience.481 This includes building systems that maintain functional stability without refrigeration, can be used accurately by operators with limited training, and integrate with data reporting mechanisms outside centralized electronic systems.
In the European Union (EU), in vitro diagnostics are governed by the In Vitro Diagnostic Regulation (IVDR 2017/746), which came into force in 2022.482 Authorization relies on Conformité Européenne (CE) marking, supported by conformity assessment carried out by Notified Bodies.483 What is examined during review depends in part on device risk but usually includes analytical performance, clinical evidence, and whether the submitted evidence is sufficient to support the intended use. Where diagnostics are classified as higher risk, the process is further structured through the use of Common Specifications, which establish shared technical reference points across the system and serve a function broadly comparable to that of FDA guidance documents. Approval does not conclude regulatory involvement, and manufacturers are expected to maintain post-market performance follow-up and vigilance activities within an established quality management system as part of ongoing oversight.
Regulatory approaches across Asia differ in their formal structures, though many have aligned with international norms and shared expectations. In Japan, regulatory review conducted by the Pharmaceuticals and Medical Devices Agency follows an approach that is broadly consistent with FDA practice, especially in how technical documentation and clinical validation are used to support diagnostics intended for clinical decision-making.484 Under the National Medical Products Administration, devices in China are approved through a registration pathway in which technical expectations and clinical evaluation are assessed through a layered review.485 In India, regulation is carried out by the Central Drugs Standard Control Organization under the Medical Device Rules of 2017, which use risk-based classification to define both clinical evidence requirements and expectations for quality system documentation.486 Taken together, these systems point toward increasing regulatory attention to clinical evidence generation, traceability, and oversight that extends across the full lifecycle of diagnostic products.
6. Limitations and challenges
While the preceding sections of this critical review highlighted the developments and promise of POC cancer diagnostics with topics ranging from clinical perspectives on diverse cancers, emerging optical, electrochemical, and CRISPR-based detection platforms, to the global funding landscape and the rise of AI-enabled biosensing technologies, it is equally pertinent to critically examine the challenges that remain in translating these technologies from bench to bedside. Despite remarkable advances in POC sample preparation, microfluidic integration, and real-time molecular analysis, significant barriers exist in terms of cost-performance trade-offs, automation and system integration, clinical usability, and regulatory approval pathways. Moreover, issues associated with sample handling, preservation, and data management, especially when incorporating AI-based analytics, continue to limit large-scale clinical deployment and patient accessibility. In this section, we discuss these limitations and challenges in detail, focusing on practical, technological, and translational aspects that currently hinder the widespread adoption of POC cancer diagnostics. We begin by addressing the clinical and logistical barriers to implementing chairside diagnostic devices in oral healthcare settings, where both technical and socioeconomic factors play decisive roles in determining adoption and impact.Chairside device development in oral care scenarios needs to consider clinical requirements to accelerate acquisition and utilization of the device by the clinical community. In general, clinical needs include improvement of clinical diagnosis, evaluation, monitoring of treatment with enhancement of clinical outcomes while improving quality of life. However, for the oral health professional there are additional hurdles compared to their medical counterpart. The oral health professionals, like many medical professionals, are small business entrepreneurs. Health professionals in private clinics as well as in academic or public health clinics must justify any new expense, incorporation of any new device or sampling protocol into the flow of patients in practice. Delays in diagnosis and treatment care can damage patient relationships with doctors and reduce time for procedures which also affect profitability of the practice. In addition, reimbursement for costs and time allocated to procedures often is covered through insurance. However, for oral health professionals this can be more difficult than for medical professionals. Oral health services are limited by any coverage through Medicaid or Medicare, and thus dental professionals must learn to use medical insurance codes to obtain reimbursement. There are codes for oral brush cytology, and oral microbiome but the oral health professional needs to provide the correct justification based on the guidelines provided by the insurance coverage and for medical professional with more experience this can be provided more readily. In addition, oral health professionals are increasingly aware of genomic, immunologic and microbiome-based platforms of data sets, that require processing of information to (1) enhance early diagnosis, (2) modify or support on-going monitoring of oral disease or (3) initiate modification in treatment. However, adaptation procedures and protocols will be needed to encourage and reinforce initiation of testing in the clinic or dental office. We expect that implementation of chairside POC tests can be useful to improve communication between clinicians and patients which can support clinical practice development and enhance long-term follow up emphasizing low cost, ease of use, and implementation of continuous screening at a high comfort level for clinicians and patient.487–492
It is important to realize that detection of oral disease: infection, inflammation and/or neoplasia is not exclusive to oral biology and health conditions. There is increasing awareness that there are multiple oral to non-oral networks of bidirectional integrated pathways. For this reason, use of non-invasive, repeatable approaches using oral brush, saliva gathered in a tube, or harvest of gingival crevicular fluid using a paper strip is rapidly becoming an important avenue for early detection of oral and non-oral cancers. Non-invasive approaches provide samples that characterize the patient environmental exposome experience by harvesting the oral microbiome. The oral microbiome and the gut microbiome are bidirectionally interactive with oral and gut mucosa while both tissues are in bidirectionally contact with brain, heart, kidneys, and skin tissues. Various platforms discussed in this review will provide devices and quantitative detection for early diagnosis, and treatment monitoring of oral and non-oral systemic disease pathogenesis and when used with support from additional data set analyses and artificial intelligence predictive modeling opening a new era for intervening in systemic disease pathogenesis through characterizing oral fluid and biology.
It is assumed that the development of a chairside device will be achieved by incorporating into the device a “lab-on-a-chip” technology after non-invasive harvesting (e.g., brush, swab, paper) of fluids, proteins, cells or other biologic information vehicles (e.g., exosomes, multilamellar vehicles). The selection of this technology should incorporate a variety of assessments. For example, these can include an analysis of macro-environment composition (e.g., pollutant-particulate irritants, nutrients-diet, toxins).493 Another set of technologies should include evaluation of different omics that characterize macro and microenvironments, and the microbiome as a bridging and mediating system regulating a variety of human cell types. These include microbiome-viromics with speciation identification; metagenomics characterizing metabolic derivatives and metabolite interactive products; describing microbiome and human oral cells and tissues integration.494,495 Additional human cell and tissue normal or abnormality levels are evaluated using genomics and proteomics technologies.496,497 For example, characterizing DNA and RNA structural changes and stabilities, metabolic biochemistry, cellular energy utilization and oxidative stresses. Any chairside device should provide a dynamic view of cell and tissue viability related to cell and tissue maintenance and/or stability of genomic structures while identifying biomolecular condensate products validating functional significance to markers.498–500 Using these rich sources for an array of markers of various types, we can attain a clinical protocol to bolster clinical judgement within an in-depth rapid comprehensive overview of the drivers for a specific disease at multiple time points during pathogenesis. This approach can also be further improved in accuracy determined by depressing false positive and negatives by transferring lab chip results to an AI data analysis system. Integration to improve predictive value for diagnosis, and achievable treatment monitoring for proper treatments achieving best outcome.
6.1. Trade-off between the performance and cost
Qualitative and quantitative estimation of cancer biomarkers with high sensitivity as well as specificity is extremely important when it comes to cancer POC diagnostics. This is extremely relevant in cancer detection as identification of biomarkers in bodily fluids has the most life-saving potential when the molecules can be detected at low concentrations, before cancer has reached an advanced stage. Thus, there is an inherent challenge for POC cancer molecular diagnostics that seek to provide ultrasensitive detection of low-concentration analytes, but at low cost, rapid sample-to-answer time, and minimal workflow complexity. Due to the combined requirements, POC molecular diagnostic technologies for cancer represent a scientific and molecular frontier that requires a multidisciplinary combination of innovative solutions. For instance, while the paper-based diagnostic tests are user-friendly and cost-effective, they are often not regarded as suitable for cancer diagnostics due sensitivity limitations of the platform. Nonetheless, paper-based assay formats can utilize novel nanoparticle reporters and detection instruments that can, in some cases, provide sensitivity superior to standard laboratory-based ELISA.501–504A further challenge for all molecular diagnostic technologies used in cancer is the heterogeneity that is inherent in human biology, resulting in broad ranges of concentration for any biomarker across the population, and the difficulty of establishing a threshold that clearly separates healthy from disease states, particularly when only one biomarker is considered. As highlighted in several of the technologies highlighted in this review, false negatives and positives can be reduced through multiplexed co-detection of several biomarkers.
As highlighted in the review, POC cancer molecular diagnostics is an inherently multidisciplinary field that brings together hardware engineers, software engineering, physicians, biologists, AI experts, nano-scientists, chemists, and bio-physicists.
In spite of numerous advancements in the broad domain of technology development for precise cancer biomarker detection and its development via various clinical trials, patenting, start-up to company launch, to being a successful product in the market the challenges that still remain are associated with the reproducibility and reliability of the devices when subjected to new settings and user groups. Calibration/quantitation issues, user error, local environmental/handling errors, manufacturing reproducibility, reagent storage/aging, and many other issues serve to threaten the validity of test results that will be used to inform clinical decisions. Further complications arise from cross-reactivity in complex matrices like saliva, urine, sweat, which are often less understood than blood serum or plasma, especially when the patient is taking medications, exposed to various environments, and experiencing other health conditions besides cancer. Biological assays often demand minimal fluctuations in the environmental conditions such as temperature and humidity, which if not considered, can compromise the accuracy of tests performed in relatively uncontrolled POC conditions. POC technologies face additional challenges compared to their laboratory-based counterparts due to the complex scenarios and diverse field settings where the tests will be performed.
6.2. Automation and integration
A clear trend in microfluidic-based cancer diagnostics at the POC is the shift from single-function components toward fully integrated platforms capable of performing sample preparation steps.505 Early academic research prototypes often addressed just one element of the workflow, for example, a chip that enriches EVs for offline analysis, or a device that performs cell lysis but requires off-chip DNA extraction. Recently there has been progress towards “sample-to-answer” systems that integrate lysis, purification, enrichment, and detection into a unified workflow. This level of integration enhances operational efficiency, reduces user error, and minimizes the need for manual handling or laboratory equipment. For instance, the bile-CAPTURE system integrates cell isolation and on-chip immunostaining, allowing users to initiate complex diagnostic protocols simply by loading the sample and pressing “start”.206 Similarly, a hand-powered ctDNA chip combines plasma separation and mutation detection into a single disposable device, accepting raw blood as input and delivering a binary diagnostic readout (mutation present or absent) without requiring external instrumentation or trained personnel.227 Such closed-loop platforms are essential for reliable POC use, especially in decentralized or resource-limited environments.Sample processing speed is another critical consideration. Many emerging POC microfluidic devices can complete sample processing in under an hour—and some in minutes—compared to traditional methods like ultracentrifugation (>5 hours for EVs) or standard DNA extraction (∼2 hours with incubation). Rapid turnaround not only improves workflow efficiency but can be clinically impactful, enabling real-time decisions in intraoperative settings or acute oncology care, where a patient may receive an answer and treatment plan during their visit to a health provider. For example, a fast ctDNA test at the bedside could indicate whether a patient is responding to targeted therapy, allowing same-day treatment adjustments. Microfluidics also excels at processing small sample volumes, often requiring only microliters. While this can limit sensitivity for extremely dilute biomarkers, it enables minimally invasive sampling from finger-stick blood or small volumes of saliva or urine, facilitating more frequent and patient-friendly testing. Innovative designs have begun to address volume constraints: acoustic exosome chips process several milliliters of biofluid through continuous-flow operation, while centrifugal microfluidics divide larger volumes into multiple chambers for parallelized processing.205 Together, these advances highlight how automation, integration, and speed are redefining the capabilities of microfluidic platforms for POC cancer diagnostics, moving them closer to real-world clinical deployment.
6.3. Avoiding sample preservation and transport
One of the key advantages of POC sample preparation is the ability to process analytes immediately after collection, minimizing degradation and preserving molecular integrity. Many tumor-derived biomarkers are inherently unstable: circulating free DNA (cfDNA) is rapidly degraded by nucleases, exosomal RNAs can decay if vesicles rupture, and CTCs may die or lyse within hours in standard collection tubes. Conventional workflows often require specialized blood collection tubes with preservatives, cold-chain transport, and tightly controlled timelines—adding cost, complexity, and risk of failure if protocols are not strictly followed. POC devices eliminate many of these issues by transforming the clinic into the laboratory. For example, microfluidic cfDNA platforms are engineered to process fresh blood and immobilize nucleic acids onto a solid phase within minutes. This immediate capture effectively “locks in” the molecular state, protecting cfDNA from enzymatic degradation and removing the need for plasma preservation. Similarly, microfluidic devices that capture EVs directly onto chip surfaces soon after collection help preserve their RNA content. Immobilization allows for gentle washing steps that remove proteases and RNases, stabilizing the vesicles and their cargo. A notable example is the acoustofluidic saliva exosome device developed by the Wang group.197 Saliva contains high levels of digestive enzymes that can degrade DNA and RNA. However, the platform isolates EVs in real-time as the saliva is being collected, significantly improving RNA preservation compared to storing or shipping samples to a distant lab.Avoiding transport also broadens access to advanced diagnostics. Patients in remote or resource-limited settings can still benefit from molecular testing if the assays are performed on-site. Instead of sending a blood sample to a centralized lab and waiting days for results, a nurse or clinician can run a microfluidic cartridge and receive results within the same appointment. This immediacy can transform clinical decision-making. For instance, a POC ctDNA test could be used to detect emerging drug resistance mutations during an oncology visit, enabling same-day adjustments in therapy rather than requiring a follow-up once lab results become available.
6.4. Clinical readiness, usability and regulation
The platforms discussed vary widely in their maturity and practical deployment. Instrumented cartridge systems such as GeneXpert and Idylla are used in clinical labs or trials worldwide.506,507 They come with vetted quality controls, standard operating procedures, and in some cases regulatory approvals. For instance, GeneXpert's oncology tests (Bladder Cancer, BCR-ABL, etc.) and Idylla assays have CE-IVD status in Europe. On the other hand, newer entrants such as ExoDisc and LiquidScan are at various stages of validation. A critical aspect of clinical readiness is usability: POC devices must be operable by non-specialists and have fail-safes. Many platforms achieve this with automation and self-contained reagents. For example, the Xpert cartridges have internal controls and self-guiding interfaces, making them easy to implement even in modest labs.5086.5. AI-based POC diagnostics: limitations and road ahead
Artificial intelligence (AI) increasingly plays the central role in point-of-care (POC) molecular diagnostics, particularly where rapid interpretation of complex signals is required.509,510 Unlike lab-based environments, POC settings demand speed, robustness, and interpretability under constraints of cost and operator expertise. AI's role is therefore not only computational but translational: it enables cloud- or smartphone-based readers, simplifies device hardware, and provides clinical decision support at the time and place of care. Below are listed some recent applications of AI in POC devices.Panels of biomarkers are much more informative than just one biomarker for cancer, whether the molecules are ctDNA, miRNA, EVs, or proteins. It is possible to use AI to combine molecular biomarkers of multiple types with other health information from medical imaging and patient characteristics to develop AI-driven models that have prognostic and predictive value to gauge cancer aggressiveness, the treatment option most likely to succeed, and survival. One potential value of POC cancer molecular diagnostics is the ability to perform the same test many times to develop longitudinal profiles that reveal trends that would otherwise be missed by expensive and invasive single-use tests. The ability to measure longitudinal trends across several molecular biomarkers provides another type of signature that can inform AI models.
AI-based POC diagnostics face numerous obstacles on the path towards clinical acceptance. Reported advantages are not generalizable beyond the local site of improvement when data are split at the measurement rather than patient level, indicating the need for multi-site validation and pre-registered analysis protocol. Smartphone readers remain vendor-sensitive, optics-sensitive, and light-sensitive, requiring device-invariant algorithms and calibrated standardization. Assay stability is also a concern: LFAs, spectroscopy, and CRISPR chemistries all have lot-to-lot drift and reagent instability that needs traceable calibration, on-card controls, and stable lyophilized reagents. For nucleic-acid analysis, contamination and carryover provide sources of false positives unless closed cartridges and enzymatic protection are implemented. Cross-reactivity and signal distortion threaten multiplex panels if orthogonal targets and internal standards are not used. Threshold drift and imprecision make quantitative performance more difficult, which necessitates calibrated reference materials and predetermined decision thresholds.
Real-world deployment adds user error, data governance, and model lifecycle hazards. Simplified procedures, QC notifications, encryption, versioning, and post-market drift monitoring are required safeguards. Equity also matters: devices must be validated in heterogeneous populations and settings with commoditized consumables and sturdy supply chains. In short, technologically promising AI-driven POC tests depend on their usefulness, which rests on formal validation, standard chemistries, stable reagents, and subsequent trials demonstrating clinical utility.
AI's translational promise lies in augmenting, not replacing molecular assays. Cloud-based AI interpretation via smartphones enables use in dental clinics, rural health centers, and low-resource settings where central labs are inaccessible. AI can also integrate ctDNA, miRNA, protein, and EV signatures into compact diagnostic scores that improve accuracy in therapy selection and monitoring. Importantly, low-cost AI-enhanced POC tools offer a path to democratizing cancer diagnostics globally, provided external validation and careful regulatory design are prioritized. In conclusion, AI allows for rapid and prolonged deployment of POC cancer tests but risks overfitting, bias, and regulatory overload. Most important are the optimal practices, which are algorithmic-independent ones: partition at the patient level, externally validate, pre-register analytic strategies, and replicate results with orthogonal assays (e.g., targeted MS, qPCR). This ensures that AI-driven POC tests are scientifically sound, clinically beneficial, and universally accessible.
7. Future scope and perspectives
7.1. Convergent framework for POC cancer diagnostics: clinical drivers, technologies, and translation
Fig. 13 provides a convergent framework illustrating how future point-of-care (POC) molecular cancer diagnostics can be realized through the alignment of clinical drivers, enabling technologies, and translational pathways. The left panel (“why”) highlights the clinical forces motivating POC adoption, including the shift toward decentralized, patient-centric care across outpatient settings, dental clinics, and therapy-monitoring contexts. These use cases underscore the need for early detection, real-time therapy guidance, and longitudinal disease monitoring using minimally invasive samples such as liquid biopsies. Collectively, these clinical scenarios define stringent requirements for rapid turnaround, operational simplicity, and clinically actionable readouts at or near the point of care.The central panel (“how”) captures the technological convergence enabling these clinical objectives, emphasizing advances in microfluidics and lab-on-a-chip platforms for integrated sample preparation, enrichment, and handling of complex analytes such as circulating tumor DNA, exosomes, and circulating tumor cells. These platforms are complemented by advanced biosensing strategies, including plasmonic, Raman, and fluorescence-based approaches, often enhanced through nanomaterials to improve sensitivity and multiplexing. Digital integration, particularly via smartphone-based POC interfaces and AI-driven analytics, further supports real-time decision-making and data connectivity. The right panel (“bridge”) highlights the translational and regulatory pathways required to move these technologies into clinical practice, including regulatory approval, economic viability, intellectual property considerations, and usability. At the center, the future outlook emphasizes precision oncology outcomes which are equitable access to care, real-time therapy adjustment, and improved survival, thereby reinforcing that successful POC deployment requires coordinated progress across clinical, technological, and translational domains.
While this framework outlines what is needed for future POC cancer diagnostics, the pace and direction of progress are strongly shaped by funding priorities, policy initiatives, and translational incentives. An examination of global funding trends, intellectual property activity, and clinical adoption provides insight into how these convergent technologies are being translated toward real-world impact.
7.2. Funding, policy, and translational ecosystem shaping POC innovation
The US National Institutes of Health (NIH) is the largest source of governmental funding for cancer research, and the National Cancer Institute was responsible for awarding over $4 billion (USD) of the NIH's overall $32 billion grant budget for the 2024 fiscal year.511 By using the NIH RePORTer database, over two million abstracts for funded grants were assessed using keywords in the title and abstract to categorize projects that mentioned one or more type of cancer. Fig. 14 presents an analysis of NIH-funded awards from 1989 to 2025, subsequently classified into six thematic fields: (i) assays and screening methods for cancer diagnostics, (ii) therapeutics for the direct treatment of cancer, (iii) mechanistic studies of cancer biology at the molecular or cellular level, (iv) strategies for cancer prevention, (v) devices, including POC technology development for cancer research or clinical care, and (vi) computational tools used in the study of cancer. Keywords were selected to minimize category overlap while still capturing the breadth of research trends and are available in the SI.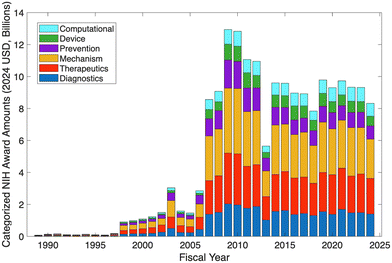 | ||
| Fig. 14 Analysis of NIH-funded awards from 1989 to 2024, categorized using thematic keywords, with the total award amounts plotted annually. Normalized keyword searches in award abstracts were used to determine the category of each award, and monetary amounts were converted to 2024 USD by using yearly CPI data.513 This is our original analysis; no permission from other publishers is required. | ||
Compared with funding in 2000, the amount awarded to cancer-focused grants has increased dramatically, and when adjusted for inflation the peak occurred in 2009 at almost $13 billion (normalized to 2024 USD). For the last ten years, grant amounts have remained more steady between $8 and $10 billion after a sharp downturn caused by the fiscal cliff in 2013.512 What was not constant, however, were the number of abstracts that claimed to develop point-of-care diagnostics or screening. These rose steadily from grant amounts of less than $1 million in 2000 to almost $40 million in 2024, proportionally outpacing the growth of other cancer funding. Although this specific category still accounts for a relatively small proportion of the overall cancer funding budget (rising to about 0.5% of cancer-related grant funding in 2024), this consistent increase indicates that the field is becoming a focus for more researchers and that granting agencies are willing to fund promising POC approaches in cancer diagnostics. Among this pool of 802 grants, nearly one quarter mentioned cervical cancer, making it the most-referenced cancer type, with an emphasis on protein biomarker detection. The second most-referenced pathology, breast cancer, accounted for 11% of the grants, and these primarily proposed a genetic subtyping or nucleic acid profiling method as the subject of the grant.
However, the clinical research funded by the NIH is only one facet of the funding story. In the United States, the National Science Foundation (NSF) is charged with advancing basic science research, and by examining historical grant awards we can extract trends relevant to POC cancer diagnostics. A search using the keyword “cancer” identified 4567 awards; each year since 2003, $30–80 million USD (adjusted for inflation) have been allocated to some fundamental study of the nature of cancer or oncology. Although a broad range of grant targets are present in the data, one notable shift in language is the appearance of terms such as “artificial intelligence” and “machine learning”, which became prevalent in abstracts starting in 2017. Computational tools have always been prevalent in the study of cancer, but the new focus on AI has dominated the awards landscape in recent years, with 25% of grants from the NSF prominently featuring those terms in 2025. Since 2015, aided by advances in other scientific domains, researchers have increasingly focused on developing multidisciplinary systems, shifting from single-substrate oriented studies toward integrated platforms capable of delivering sensitive, scalable, and label-free diagnostic solutions and therapy guidance. These systems have leveraged breakthroughs in materials science, microfluidics, wearable technologies, and biophysical modeling to create versatile, patient-centered tools. This trend is typified by the Cancer Moonshot Initiative, with fiscal year 2016 resulting in the highest single-year number of NSF cancer-related awards in the dataset and 2017 containing the highest real monetary spending on cancer-related awards by the agency.514 However, in subsequent years, the number of cancer-associated NSF grants has stagnated, with a modest downward trend. This suggests a potential pivot in funding priorities, increased competition from other federal initiatives, or a strategic shift toward translational and commercialization pathways outside the traditional NSF funding mechanisms. While the NSF is responsible for improvements in foundational understanding necessary to advance science as a whole, clinical and translational research are not a focus of that agency.
Outside of the United States, significant funding for POC cancer diagnostics research is available through the European Commission's Horizon Europe grant program, which establishes seven-year grant periods to fund strategic goals related to research and innovation. The Horizon Europe website lists five “missions”, one of which is cancer prevention and treatment, with a stated goal of saving 3 million lives by 2030.515 The total yearly cancer-related funding rose to approximately €500 million in 2022, which is an order of magnitude smaller than the NIH cancer budget. However, of these awards 1.8% of allocated funds were for grants mentioning the keyword “point-of-care” in their objectives,516 representing a significant emphasis on innovation in these technologies when compared with the less than 0.5% monetary share that point-of-care NIH cancer grants exhibited in the same time period. With that in mind, we may contextualize the statements made by Berns et al. in 2020 in their recommendations for the Horizon program. They emphasized that any advances in early detection and screening must also consider which technologies will best reduce the risks associated with increased testing,517 a challenge which POC devices must seek to overcome. Finally, in 2025, the proposed budget for the 2028–2034 Horizon Europe funding cycle would double total grant funding compared to the previous Horizon program, which is indicative of an emphasis on R&D in the next decade.518 Although this analysis only considers a subset of available funding worldwide, the trend for POC research seems to be positive, driven by proposals that build on the molecules and mechanisms discovered through basic research and aimed at improving outcomes for patients in clinics around the world.
Beyond public research funding, translational readiness is also reflected in intellectual property activity, clinical trial engagement, and early-stage commercialization efforts. Together, these indicators provide a complementary view of how POC cancer diagnostics are progressing from concept to deployment.
7.3. Commercialization and clinical translation: patents, trials, and SBIR activity
The patent landscape for cancer or oncology POC diagnostics demonstrates a marked trend of increasing innovation and commercial interest over the period from 2010 to 2024. As shown in Fig. 15a, the patent applications (blue bars) exhibited a consistent upward trajectory, peaking in 2021 with over 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 new filings, indicating robust research and development in the field. Similarly, issued US patents (green bars) also demonstrated significant growth, reaching their highest level in 2019 with approximately 13
500 new filings, indicating robust research and development in the field. Similarly, issued US patents (green bars) also demonstrated significant growth, reaching their highest level in 2019 with approximately 13![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 patents granted, suggesting a consistently high rate of innovation and commercialization potential. The data for 2023–2024 shows an apparent decline in both applications and grants due to the inherent time lag (typically 1–3 years) between an application's filing and its official public disclosure and eventual grant in intellectual property databases.519,520 Overall, the trend from 2010 through 2022 reveals a highly active and productive phase for intellectual property development in cancer POC diagnostics.
000 patents granted, suggesting a consistently high rate of innovation and commercialization potential. The data for 2023–2024 shows an apparent decline in both applications and grants due to the inherent time lag (typically 1–3 years) between an application's filing and its official public disclosure and eventual grant in intellectual property databases.519,520 Overall, the trend from 2010 through 2022 reveals a highly active and productive phase for intellectual property development in cancer POC diagnostics.
Since 2010, the number of clinic trials and NIH project trends have displayed positive growth for the cancer POC field. As seen in Fig. 15b, the number of funded NIH grants (blue bars) displays a consistent positive trajectory with a spike in 2018. NIH grants typically span 3–5 years, thus the number of funded grants in a year indicates commitments into future years.521,522 Meanwhile, the number of clinical trials related to POC cancer diagnostics increased at a linear rate.
To track the development of projects devoted towards commercialization, the Small Business Innovation Research (SBIR) database of federal grants was used to track trends in overall cancer diagnostics business development. Cancer-related SBIR grants comprised approximately 4.7% of all SBIR grants awarded in the past ten years. This relatively small yet significant portion highlights the selective but growing emphasis on translational cancer research within the broader innovation ecosystem (that is specific to this review). The data suggest that while cancer research represents only a fraction of total SBIR activity, it continues to attract sustained federal investment, reflecting its high societal and clinical impact. A closer examination of the cancer-focused SBIR portfolio reveals a distinct distribution of priorities within the funded projects. Among all cancer-related SBIR grants, therapeutics-oriented projects accounted for 16.2%, emphasizing the enduring interest in drug discovery, targeted delivery systems, and precision medicine strategies. These projects often bridge fundamental discoveries in molecular oncology with the development of clinically actionable therapeutic modalities which are indeed areas traditionally dominant in cancer research funding.
In contrast, projects centered on POC cancer diagnostics represented only 2.9% of the cancer-related SBIR grants. Despite this modest fraction, these POC efforts are strategically significant, as they focus on technologies that can decentralize cancer diagnostics, shorten turnaround times, and facilitate early detection in resource-limited or outpatient settings. The relatively low percentage may reflect both the technical challenges of translating advanced biosensing platforms into robust, regulatory-compliant clinical devices and the comparatively higher costs and risks associated with therapeutic development that tend to attract more established funding streams.
Overall, these distributions underscore the current landscape of innovation supported by SBIR programs, one in which cancer research occupies a specialized but impactful niche, with most investments directed toward therapeutic solutions and a smaller, yet promising, subset devoted to diagnostic and POC advancements. These trends collectively highlight where translational innovation is concentrated and where opportunities remain to enhance diagnostic technology development. In summary, these ecosystem-level trends suggest growing translational momentum for POC cancer diagnostics. At the technology level, continued progress will depend on how sample preparation, integration, sensitivity, and usability evolve to meet real-world clinical constraints.
7.4. Technology-level future directions for POC cancer diagnostics
POC-compatible sample preparation methods for cancer diagnostics have rapidly advanced, largely driven by advances in microfluidic engineering. These platforms can now process a wide range of biofluids from a drop of blood to saliva, urine, ascitic fluid, or oral swabs. By focusing on minimally and noninvasive samples (Table 1) and integrating multiple steps on-chip (Table 2), these systems address real-world clinical needs. They are faster, more compact, and easier to operate than conventional lab-based methods yet maintain, and often exceed, the sensitivity and specificity of traditional assays. Recent successes, such as integrated EV isolation chips and automated cfDNA extractors, demonstrate that complex sample preparation tasks like isolating nanoscale vesicles or purifying fragmented nucleic acids can be performed at the POC without compromising performance. In fact, many microfluidic devices deliver biomarker yields equal to or better than gold-standard methods (e.g., acoustic EV isolation outperforming ultracentrifugation, microfluidic solid-phase DNA capture matching or exceeding column-based protocols). These systems' efficiency, coupled with their low sample volume requirements and rapid turnaround times, makes them ideally suited for decentralized screening, early cancer detection, and longitudinal patient monitoring.Looking forward, we anticipate several trends and developments:
• Further integration and automation: while current devices integrate many steps, full sample-to-answer systems with built-in detection (optical/electrical readout) will become more common. The ultimate goal is a cartridge for the user to input a raw sample, and after inserting the cartridge into an automated instrument, the outputs a result (e.g., “mutation X detected” or “EV count above threshold”) on a desktop instrument, handheld reader or smartphone. Achieving this vision will require robust fluidic control (valves, pumps) that are inexpensively mass-manufactures, as well as calibration-free detection methods (using internal controls and calibration standards). Advances in 3D-printing and microfluidic manufacturing are helping produce more complex chips at lower cost, which supports integration.
• Clinical validation and standardization: to transition from a lab prototype to clinical tool, POC methods must be validated in larger patient cohorts and standardized. This includes establishing performance metrics (sensitivity, specificity, LOD) under real-world conditions. Some biomarkers lack standard reference materials, making comparisons difficult. For example, different EV isolation chips might yield slightly different subpopulations of vesicles. As the field matures, we need standard protocols or reference samples to fairly evaluate technologies. Regulatory approval pathways (e.g. FDA) will require demonstration that an integrated POC test is as reliable as the centralized lab test it aims to replace. Encouragingly, a few microfluidic POC cancer tests are already in clinical trials or nearing regulatory submission, especially in the liquid biopsy space for lung and colon cancer.
• Improving sensitivity for early detection: a key promise of liquid biopsy is earlier cancer detection. POC tests will play a major role in population screening only if they can detect extremely low biomarker levels (e.g. one mutated DNA fragment among 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 normal ones). Additionally, combining multiple analytes, for instance, detecting a panel of, say, 5 different cancer-related mutations or proteins on the same chip can improve overall diagnostic accuracy. Microfluidics excel at multiplexing, so we expect future cartridges will concurrently isolate and test several classes of biomarkers (perhaps CTCs + ctDNA + EVs together) to build a more comprehensive picture of disease.
000 normal ones). Additionally, combining multiple analytes, for instance, detecting a panel of, say, 5 different cancer-related mutations or proteins on the same chip can improve overall diagnostic accuracy. Microfluidics excel at multiplexing, so we expect future cartridges will concurrently isolate and test several classes of biomarkers (perhaps CTCs + ctDNA + EVs together) to build a more comprehensive picture of disease.
• User-centered design: to truly bring these technologies to clinics and remote areas, device designers are focusing on usability. The end-user might be an oncologist, a nurse, a dentist, or even the patient. Interfaces like simple push-button starts, clear sample loading steps, and smartphone integration for displaying results are being refined. Training requirements must be minimal – ideally the device operation should be almost as simple as a home glucose test. Some teams are developing AI-powered smartphone apps to guide the user (by recognizing cassette orientation, volume sufficiency, etc.) and interpret faint results, adding another layer of reliability at POC.
In conclusion, the convergence of microfluidics, nanotechnology, and biomedical engineering in the past five years has brought us to the brink of a new era in cancer diagnostics, where high-quality molecular testing can be done anywhere, anytime. By focusing on minimally invasive samples and integrating sample preparation into portable devices, these advances uphold the promise of liquid biopsy – safer, more accessible cancer diagnosis and monitoring – and extend it to the point of need. With continued progress, we foresee that routine check-ups could include on-the-spot cancer biomarker panels, and patients in all global regions could benefit from early detection through a simple chip-based test. The momentum in research and the early clinical demonstrations reviewed here strongly suggest that POC cancer diagnostics will become a practical reality in the near future, fundamentally improving how we detect and treat cancer.
8. Conclusions and outlook
POC cancer diagnostics represent a transformative shift in oncology, enabling rapid, accessible, and patient-centered detection of disease outside traditional laboratory settings. This critical review has traced the pathway from fundamental research to clinical translation, highlighting the importance of reliable sample preparation, biomarker selection, and integrated detection strategies. Advances in biosensors, microfluidics, miniaturized detection instruments, novel biochemistry methods, genomic/proteomics-informed selection of informative biomarker panels, and high-sensitivity assays have expanded the analytical capabilities of POC platforms, while emerging technologies such as multi-omics integration and AI-driven interpretation promise to enhance precision and early detection. However, translational success depends not only on technological innovation but also on thoughtful implementation strategies, including regulatory compliance, funding mechanisms, and academic–industry collaboration. Addressing these factors is essential to ensure that POC diagnostics are robust, reproducible, and suitable for deployment in diverse clinical and resource-limited settings.Looking forward, the field must balance innovation with accessibility and equity. Overcoming challenges such as device standardization, operational limitations, and cost-effectiveness will be critical to broader adoption. Integration of digital health tools, real-time data analysis, and patient-centered design can further enhance clinical utility and longitudinal monitoring. By fostering collaborations across scientific, clinical, and commercial domains, POC cancer diagnostics have the potential to revolutionize early detection and improve outcomes globally. A holistic, multidimensional approach, combining technological rigor with translational insight and equitable deployment, will be key to realizing the full promise of this rapidly advancing field.
Author contributions
Seemesh Bhaskar: conceptualization, data curation, formal analysis, funding acquisition, investigation, methodology, project administration, software, supervision, validation, visualization, writing – original draft, writing – review & editing; Saurabh Umrao: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Han Keun Lee: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Joseph Tibbs: data curation, formal analysis, investigation, methodology, software, validation, visualization, writing – original draft; Amanda Bacon: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Skye Shepherd: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Takhmina Ayupova: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Fatma Uysal Ciloglu: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Leyang Liu: data curation, formal analysis, investigation, methodology, software, validation, visualization, writing – original draft; Anqi Tan: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Wang-Chien Chen: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; My Thi Tra Nguyen: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Maria Grace Scannell: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Ugur Aygun: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Ugur Parlatan: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Catherine Zhang: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Manish Kohli: data curation, formal analysis, investigation, methodology, software, supervision, validation, visualization, writing – original draft; Guy R. Adami: data curation, formal analysis, investigation, methodology, software, supervision, validation, visualization, writing – original draft, writing – review & editing; Wali Badar: data curation, formal analysis, investigation, methodology, software, validation, writing – original draft; Ron C. Gaba: data curation, formal analysis, investigation, methodology, software, supervision, validation, visualization, writing – original draft, writing – review & editing; Aaron Mansfield: data curation, formal analysis, investigation, methodology, software, supervision, validation, visualization, writing – original draft, writing – review & editing; Joel Schwartz: data curation, formal analysis, investigation, methodology, software, supervision, validation, visualization, writing – original draft, writing – review & editing; Xing Wang: data curation, formal analysis, investigation, methodology, software, supervision, validation, visualization, writing – original draft, writing – review & editing; Utkan Demirci: data curation, formal analysis, investigation, methodology, software, supervision, validation, visualization, writing – original draft, writing – review & editing; Brian T. Cunningham: conceptualization, formal analysis, funding acquisition, investigation, methodology, project administration, resources, supervision, validation, visualization, writing – original draft, writing – review & editing.Conflicts of interest
Utkan Demirci is a co-founder of and has an equity interest in: (i) Vetmotl Inc., (ii) LevitasBio, (iii) Hermes Biosciences, (iv) Enabridge. Utkan Demirci's interests were reviewed and managed in accordance with his institutional conflict-of-interest policies. Brian T. Cunningham is a co-founder of PhotonicDx. All authors have no other conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5lc01014d.
Acknowledgements
Seemesh Bhaskar (S. B.) acknowledges funding from the Carl R. Woese Institute for Genomic Biology (IGB). Ugur Aygun (U. A.) acknowledges funding from the European Union’s Horizon Europe research and innovation programme under the Marie Skłodowska-Curie grant agreement No. 101066038. Fatma Uysal Ciloglu (F. U. C.) is supported by the Scientific and Technological Research Council of Türkiye (TUBITAK) during her post-doctoral research.References
- V. Gubala, L. F. Harris, A. J. Ricco, M. X. Tan and D. E. Williams, Anal. Chem., 2012, 84, 487–515 CrossRef CAS PubMed.
- S. Bhaskar, W. Wang, H. Lee, L. Liu, S. Umrao, W. Liu, A. Bacon, J. Tibbs, K. Khemtonglang, A. Tan, T. Ayupova, W.-C. Chen, X. Wang and B. T. Cunningham, Chem. Rev., 2025, 125(14), 6435–6540 CrossRef CAS PubMed.
- C. Che, N. Li, K. D. Long, M. Á. Aguirre, T. D. Canady, Q. Huang, U. Demirci and B. T. Cunningham, Lab Chip, 2019, 19, 3943–3953 RSC.
- L. Liu, J. Tibbs, N. Li, A. Bacon, S. Shepherd, H. Lee, N. Chauhan, U. Demirci, X. Wang and B. T. Cunningham, Biosens. Bioelectron., 2023, 228, 115197 CrossRef CAS PubMed.
- S. Nayak, N. R. Blumenfeld, T. Laksanasopin and S. K. Sia, Anal. Chem., 2017, 89, 102–123 CrossRef CAS PubMed.
- C. Wang, M. Liu, Z. Wang, S. Li, Y. Deng and N. He, Nano Today, 2021, 37, 101092 CrossRef CAS PubMed.
- H. Yuan, Z. Miao, C. Wan, J. Wang, J. Liu, Y. Li, Y. Xiao, P. Chen and B.-F. Liu, Lab Chip, 2025, 25, 1015–1046 RSC.
- C. P. Y. Chan, W. C. Mak, K. Y. Cheung, K. K. Sin, C. M. Yu, T. H. Rainer and R. Renneberg, Annu. Rev. Anal. Chem., 2013, 6, 191–211 Search PubMed.
- A. M. Foudeh, T. Fatanat Didar, T. Veres and M. Tabrizian, Lab Chip, 2012, 12, 3249–3266 RSC.
- A. M. Caliendo, D. N. Gilbert, C. C. Ginocchio, K. E. Hanson, L. May, T. C. Quinn, F. C. Tenover, D. Alland, A. J. Blaschke, R. A. Bonomo, K. C. Carroll, M. J. Ferraro, L. R. Hirschhorn, W. P. Joseph, T. Karchmer, A. T. MacIntyre, L. B. Reller, A. F. Jackson and f. t. I. D. S. o. America, Clin. Infect. Dis., 2013, 57, S139–S170 Search PubMed.
- L. Váradi, J. L. Luo, D. E. Hibbs, J. D. Perry, R. J. Anderson, S. Orenga and P. W. Groundwater, Chem. Soc. Rev., 2017, 46, 4818–4832 RSC.
- A. E. Ongaro, Z. Ndlovu, E. Sollier, C. Otieno, P. Ondoa, A. Street and M. Kersaudy-Kerhoas, Lab Chip, 2022, 22, 3122–3137 RSC.
- Y. Zhang and Y. Liu, Sens. Diagn., 2022, 1, 648–672 RSC.
- A. K. Yetisen, M. S. Akram and C. R. Lowe, Lab Chip, 2013, 13, 2210–2251 RSC.
- M. Parihar, N. W. N. Sahana, R. Biswas, B. Dehury and N. Mazumder, RSC Adv., 2025, 15, 29267–29283 RSC.
- M. Sharafeldin and J. J. Davis, Anal. Chem., 2021, 93, 184–197 CrossRef CAS PubMed.
- R. Franco-Duarte, L. Černáková, S. Kadam, K. S. Kaushik, B. Salehi, A. Bevilacqua, M. R. Corbo, H. Antolak, K. Dybka-Stępień, M. Leszczewicz, S. Relison Tintino, V. C. Alexandrino de Souza, J. Sharifi-Rad, H. D. M. Coutinho, N. Martins and C. F. Rodrigues, Microorganisms, 2019, 7, E130 CrossRef PubMed.
- Y. Joung, K. Kim, J. E. An, S. Park, Q. Yu, M. Lu, J. Chen, S.-W. Joo and J. Choo, Trends Biotechnol., 2025, 43, 1048–1061 CrossRef CAS PubMed.
- H. Yin, M. Zhang, Y. Zhang, X. Zhang, X. Zhang and B. Zhang, Mol. Biomed., 2025, 6, 18 CrossRef CAS PubMed.
- A. Tadimety, A. Closson, C. Li, S. Yi, T. Shen and J. X. J. Zhang, Crit. Rev. Clin. Lab. Sci., 2018, 55, 140–162 CrossRef CAS PubMed.
- C. Alix-Panabières and K. Pantel, Mol. Oncol., 2021, 15, 1617–1621 CrossRef PubMed.
- K. Kurma, Z. Eslami-S, C. Alix-Panabières and L. Cayrefourcq, Cell Adhes. Migr., 2024, 18, 1–26 CrossRef CAS PubMed.
- S. Perakis and M. R. Speicher, BMC Med., 2017, 15, 75 CrossRef PubMed.
- M. Ilié and P. Hofman, Transl. Lung Cancer Res., 2016, 5, 420 CrossRef PubMed.
- M. Mino-Kenudson, Transl. Lung Cancer Res., 2016, 5, 424 CrossRef PubMed.
- F. Castro-Giner, S. Gkountela, C. Donato, I. Alborelli, L. Quagliata, C. K. Ng, S. Piscuoglio and N. Aceto, Diagnostics, 2018, 8, 31 CrossRef PubMed.
- G. Siravegna, S. Marsoni, S. Siena and A. Bardelli, Nat. Rev. Clin. Oncol., 2017, 14, 531–548 CrossRef CAS PubMed.
- C. Alix-Panabieres, Nature, 2020, 579, S9 CrossRef CAS PubMed.
- L. Ma, H. Guo, Y. Zhao, Z. Liu, C. Wang, J. Bu, T. Sun and J. Wei, Signal Transduction Targeted Ther., 2024, 9, 336 CrossRef PubMed.
- F. Fenizia, A. De Luca, R. Pasquale, A. Sacco, L. Forgione, M. Lambiase, A. Iannaccone, N. Chicchinelli, R. Franco and A. Rossi, Future Oncol., 2015, 11, 1611–1623 CrossRef CAS PubMed.
- U. Malapelle, P. Pisapia, D. Rocco, R. Smeraglio, M. di Spirito, C. Bellevicine and G. Troncone, Transl. Lung Cancer Res., 2016, 5, 505 CrossRef CAS PubMed.
- M. Chen and H. Zhao, Hum. Genomics, 2019, 13, 34 CrossRef PubMed.
- T.-M. Wu, J.-B. Liu, Y. Liu, Y. Shi, W. Li, G.-R. Wang, Y.-S. Ma and D. Fu, Cancer Control, 2020, 27, 1073274820934805 CrossRef PubMed.
- E. Zulato, V. Tosello, G. Nardo, L. Bonanno, P. Del Bianco and S. Indraccolo, Diagnostics, 2021, 11, 1468 CrossRef CAS PubMed.
- C. Lin, X. Liu, B. Zheng, R. Ke and C.-M. Tzeng, Life, 2021, 11, 890 CrossRef CAS PubMed.
- G. K. Bhatti, N. Khullar, I. S. Sidhu, U. S. Navik, A. P. Reddy, P. H. Reddy and J. S. Bhatti, Metab. Brain Dis., 2021, 36, 1119–1134 CrossRef CAS PubMed.
- R. Hamam, D. Hamam, K. A. Alsaleh, M. Kassem, W. Zaher, M. Alfayez, A. Aldahmash and N. M. Alajez, Cell Death Dis., 2017, 8, e3045 CrossRef PubMed.
- X. Qi, D.-H. Zhang, N. Wu, J.-H. Xiao, X. Wang and W. Ma, J. Med. Genet., 2015, 52, 710–718 CrossRef PubMed.
- R. A. Boon, N. Jaé, L. Holdt and S. Dimmeler, J. Am. Coll. Cardiol., 2016, 67, 1214–1226 CrossRef CAS PubMed.
- D. Xiang, S. Shigdar, G. Qiao, T. Wang, A. Z. Kouzani, S.-F. Zhou, L. Kong, Y. Li, C. Pu and W. Duan, Theranostics, 2015, 5, 23 CrossRef PubMed.
- J. D. Cohen, A. A. Javed, C. Thoburn, F. Wong, J. Tie, P. Gibbs, C. M. Schmidt, M. T. Yip-Schneider, P. J. Allen and M. Schattner, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 10202–10207 CrossRef CAS PubMed.
- E. Heitzer, I. S. Haque, C. E. Roberts and M. R. Speicher, Nat. Rev. Genet., 2019, 20, 71–88 CrossRef CAS PubMed.
- E. Kilgour, D. G. Rothwell, G. Brady and C. Dive, Cancer Cell, 2020, 37, 485–495 CrossRef CAS PubMed.
- D. W. Cescon, S. V. Bratman, S. M. Chan and L. L. Siu, Nat. Cancer, 2020, 1, 276–290 CrossRef CAS PubMed.
- G. Cheng, Adv. Drug Delivery Rev., 2015, 81, 75–93 CrossRef CAS PubMed.
- A. R. Thierry, S. El Messaoudi, P. Gahan, P. Anker and M. Stroun, Cancer Metastasis Rev., 2016, 35, 347–376 CrossRef CAS PubMed.
- K. Zen and C. Y. Zhang, Med. Res. Rev., 2012, 32, 326–348 CrossRef PubMed.
- N. Kosaka, H. Iguchi and T. Ochiya, Cancer Sci., 2010, 101, 2087–2092 CrossRef CAS PubMed.
- S. N. Lone, S. Nisar, T. Masoodi, M. Singh, A. Rizwan, S. Hashem, W. El-Rifai, D. Bedognetti, S. K. Batra and M. Haris, Mol. Cancer, 2022, 21, 79 CrossRef CAS PubMed.
- A. Pulumati, A. Pulumati, B. S. Dwarakanath, A. Verma and R. V. Papineni, Cancer Rep., 2023, 6, e1764 Search PubMed.
- T. Hussain and Q. T. Nguyen, Adv. Drug Delivery Rev., 2014, 66, 90–100 CrossRef CAS PubMed.
- X. Jiang, Z. Hu, S. Wang and Y. Zhang, Cancers, 2023, 15, 3608 CrossRef PubMed.
- M. Antunes-Ferreira, D. Koppers-Lalic and T. Würdinger, Mol. Oncol., 2021, 15, 1727–1743 CrossRef CAS PubMed.
- H. Husain and V. E. Velculescu, JAMA, J. Am. Med. Assoc., 2017, 318, 1272–1274 CrossRef PubMed.
- E. Heitzer, P. Ulz and J. B. Geigl, Clin. Chem., 2015, 61, 112–123 CrossRef CAS PubMed.
- V. M. Corman, V. C. Haage, T. Bleicker, M. L. Schmidt, B. Mühlemann, M. Zuchowski, W. K. Jo, P. Tscheak, E. Möncke-Buchner, M. A. Müller, A. Krumbholz, J. F. Drexler and C. Drosten, Lancet Microbe, 2021, 2, e311–e319 CrossRef CAS PubMed.
- L. Cohen and D. R. Walt, Chem. Rev., 2019, 119, 293–321 CrossRef CAS PubMed.
- M. Platchek, Q. Lu, H. Tran and W. Xie, SLAS Discovery, 2020, 25, 1197–1213 CrossRef CAS PubMed.
- F. Chowdhury, A. Williams and P. Johnson, J. Immunol. Methods, 2009, 340, 55–64 CrossRef CAS PubMed.
- P. J. Tighe, R. R. Ryder, I. Todd and L. C. Fairclough, Proteomics: Clin. Appl., 2015, 9, 406–422 CAS.
- H. Ahsan, Comp. Clin. Pathol., 2022, 31, 333–345 CrossRef CAS PubMed.
- E. C. Breen, S. M. Reynolds, C. Cox, L. P. Jacobson, L. Magpantay, C. B. Mulder, O. Dibben, J. B. Margolick, J. H. Bream and E. Sambrano, Clin. Vaccine Immunol., 2011, 18, 1229–1242 CrossRef CAS PubMed.
- H. Päkkilä and T. Soukka, Bioanal. Rev., 2011, 3, 27–40 CrossRef.
- K. Simeon, M. Sharma, J. Dorward, J. Naidoo, N. Dlamini, P. Moodley, N. Samsunder, R. V. Barnabas, N. Garrett and P. K. Drain, PLoS One, 2019, 14, e0223669 CrossRef CAS PubMed.
- A. Mouseli, M. Barouni, M. Amiresmaili, S. M. Samiee and L. Vali, Electron. Physician, 2017, 9, 4077 CrossRef PubMed.
- R. C. Fitzgerald, A. C. Antoniou, L. Fruk and N. Rosenfeld, Nat. Med., 2022, 28, 666–677 CrossRef CAS PubMed.
- A. A. Fryer and W. S. A. Smellie, J. Clin. Pathol., 2013, 66, 62–72 CrossRef PubMed.
- R. Sullivan, J. Peppercorn, K. Sikora, J. Zalcberg, N. J. Meropol, E. Amir, D. Khayat, P. Boyle, P. Autier and I. F. Tannock, Lancet Oncol., 2011, 12, 933–980 CrossRef PubMed.
- S. Srivastava, E. J. Koay, A. D. Borowsky, A. M. De Marzo, S. Ghosh, P. D. Wagner and B. S. Kramer, Nat. Rev. Cancer, 2019, 19, 349–358 CrossRef CAS PubMed.
- K. Payne, S. P. Gavan, S. J. Wright and A. J. Thompson, Nat. Rev. Genet., 2018, 19, 235–246 CrossRef CAS PubMed.
- A. George, D. Riddell, S. Seal, S. Talukdar, S. Mahamdallie, E. Ruark, V. Cloke, I. Slade, Z. Kemp and M. Gore, Sci. Rep., 2016, 6, 29506 CrossRef PubMed.
- X. Chen, J. Gole, A. Gore, Q. He, M. Lu, J. Min, Z. Yuan, X. Yang, Y. Jiang and T. Zhang, Nat. Commun., 2020, 11, 3475 CrossRef CAS PubMed.
- J. A. Todd, Nat. Genet., 2006, 38, 731–733 CrossRef CAS PubMed.
- J. D. Schiffman, P. G. Fisher and P. Gibbs, American Society of Clinical Oncology Educational Book, 2015, vol. 35, pp. 57–65 Search PubMed.
- D. Crosby, S. Bhatia, K. M. Brindle, L. M. Coussens, C. Dive, M. Emberton, S. Esener, R. C. Fitzgerald, S. S. Gambhir and P. Kuhn, Science, 2022, 375, eaay9040 CrossRef CAS PubMed.
- L. Necula, L. Matei, D. Dragu, A. I. Neagu, C. Mambet, S. Nedeianu, C. Bleotu, C. C. Diaconu and M. Chivu-Economescu, World J. Gastroenterol., 2019, 25, 2029 CrossRef CAS PubMed.
- B. Kaur, S. Kumar and B. K. Kaushik, Biosens. Bioelectron., 2022, 197, 113805 CrossRef CAS PubMed.
- J. A. Kim, D. J. Wales and G.-Z. Yang, Prog. Biomed. Eng., 2020, 2, 042001 CrossRef.
- J. Wang, Biosens. Bioelectron., 2006, 21, 1887–1892 CrossRef CAS PubMed.
- W. J. Peveler, M. Yazdani and V. M. Rotello, ACS Sens., 2016, 1, 1282–1285 CrossRef CAS PubMed.
- E. C. Cauberg, D. M. de Bruin, D. J. Faber, T. G. van Leeuwen, J. J. de la Rosette and T. M. De Reijke, Eur. Urol., 2009, 56, 287–297 CrossRef PubMed.
- P. Khosravi, T. J. Fuchs and D. J. Ho, Cancer Res., 2025, 85, 2356–2367 CrossRef CAS PubMed.
- G. Cserni, S. Bianchi, W. Boecker, T. Decker, M. Lacerda, F. Rank, C. A. Wells and E. W. G. f. B. S. Pathology, Cancer, 2005, 103, 358–367 CrossRef PubMed.
- K. Roper, A. Abdel-Rehim, S. Hubbard, M. Carpenter, A. Rzhetsky, L. Soldatova and R. D. King, J. R. Soc. Interface, 2022, 19, 20210821 CrossRef CAS PubMed.
- C. A. Holland and F. L. Kiechle, Curr. Opin. Microbiol., 2005, 8, 504–509 CrossRef PubMed.
- X. Wang, X.-Z. Hong, Y.-W. Li, Y. Li, J. Wang, P. Chen and B.-F. Liu, Mil. Med. Res., 2022, 9, 11 Search PubMed.
- M. Zarei, Biosens. Bioelectron., 2017, 98, 494–506 CrossRef CAS PubMed.
- Y. Xiao, M. Zhou, C. Liu, S. Gao, C. Wan, S. Li, C. Dai, W. Du, X. Feng, Y. Li, P. Chen and B.-F. Liu, Biosens. Bioelectron., 2024, 255, 116240 CrossRef CAS PubMed.
- Y. Song, Y.-Y. Huang, X. Liu, X. Zhang, M. Ferrari and L. Qin, Trends Biotechnol., 2014, 32, 132–139 CrossRef CAS PubMed.
- H. Yuan, P. Chen, C. Wan, Y. Li and B.-F. Liu, TrAC, Trends Anal. Chem., 2022, 157, 116814 CrossRef CAS PubMed.
- A. Malekjahani, S. Sindhwani, A. M. Syed and W. C. Chan, Acc. Chem. Res., 2019, 52, 2406–2414 CrossRef CAS PubMed.
- P. K. Drain, E. P. Hyle, F. Noubary, K. A. Freedberg, D. Wilson, W. R. Bishai, W. Rodriguez and I. V. Bassett, Lancet Infect. Dis., 2014, 14, 239–249 CrossRef PubMed.
- P. N. Butow, J. N. Kazemi, L. J. Beeney, A. M. Griffin, S. M. Dunn and M. H. Tattersall, Cancer, 1996, 77, 2630–2637 CrossRef CAS PubMed.
- S. S. Raab and D. M. Grzybicki, Ca-Cancer J. Clin., 2010, 60, 139–165 CrossRef PubMed.
- A. Winterbottom and D. Harcourt, J. Adv. Nurs., 2004, 48, 226–233 CrossRef PubMed.
- J. Zhao, X. Han, L. Nogueira, S. A. Fedewa, A. Jemal, M. T. Halpern and K. R. Yabroff, Ca-Cancer J. Clin., 2022, 72, 542–560 CrossRef PubMed.
- M. T. Halpern, E. M. Ward, A. L. Pavluck, N. M. Schrag, J. Bian and A. Y. Chen, Lancet Oncol., 2008, 9, 222–231 CrossRef PubMed.
- R. L. Siegel, K. D. Miller, H. E. Fuchs and A. Jemal, Ca-Cancer J. Clin., 2021, 71, 7–33 CrossRef PubMed.
- D. Lee, K. U. Jung, H. O. Kim, H. Kim and H.-K. Chun, Medicine, 2018, 97, e12244 CrossRef PubMed.
- J. H. Kim, Y. H. Choi, S.-Y. An, H. Y. Son, C. Choi, S. Kim, J. Chung and H. S. Na, Int. J. Oral Biol., 2018, 43, 13–21 CrossRef.
- M. W. Lingen, E. Abt, N. Agrawal, A. K. Chaturvedi, E. Cohen, G. D'Souza, J. Gurenlian, J. R. Kalmar, A. R. Kerr, P. M. Lambert, L. L. Patton, T. P. Sollecito, E. Truelove, M. P. Tampi, O. Urquhart, L. Banfield and A. Carrasco-Labra, J. Am. Dent. Assoc., JADA, 2017, 148, 712–727.e710 CrossRef PubMed.
- A. S. Ho, S. Kim, M. Tighiouart, A. Mita, K. S. Scher, J. B. Epstein, A. Laury, R. Prasad, N. Ali and C. Patio, Cancer, 2018, 124, 3154–3162 CrossRef PubMed.
- E. A. K. Clemencia, M. Vargas and J. A. Yellowitz, CENTERS FOR DISEASE CONTROL AND PREVENTION National Center for Health Statistics, 2000, https://www.cdc.gov/nchs/data/ahcd/agingtrends/03oral.pdf.
- S. Chen, M. Forman, P. M. Sadow and M. August, Kouqiang Hemian Waike Zazhi, 2016, 74, 959–964 Search PubMed.
- N. Osazuwa-Peters, J. F. Zeitouni, L. E. Simon and M. A. Varvares, JAMA Oncol., 2025, 11(9), 959–960 CrossRef PubMed.
- P. A. Brennan, F. Dylgjeri, R. D. Coletta, G. Arakeri and A. M. Goodson, J. Oral Pathol. Med., 2022, 51, 311–314 CrossRef PubMed.
- M. L. Urken, J. Yun, M. P. Saturno, L. A. Greenberg, R. L. Chai, K. Sharif and M. Brandwein-Weber, Oral Oncol., 2023, 143, 106445 CrossRef PubMed.
- H. Ghaedamini, S. Duanghathaipornsuk, P. Onusko, A. M. Binsheheween and D. S. Kim, Biosensors, 2023, 13(2), 254 CrossRef CAS PubMed.
- A. A.-R. Dong-Shik Kim, R. Chakravarti, N. Dominiak and P. Chaudhuri, Award Abstract # 2141183 PFI TT, 2022–2025, https://www.nsf.gov/awardsearch/showAward?AWD_ID=2141183.
- Ó. Rapado-González, S. Salta, R. López-López, R. Henrique, M. M. Suárez-Cunqueiro and C. Jerónimo, Clin. Epigenet., 2024, 16, 105 CrossRef PubMed.
- A. Balapure, S. K. Dubey, A. Javed, S. Chattopadhyay and S. Goel, Anal. Methods, 2024, 16, 6098–6118 RSC.
- L. Arantes, A. C. De Carvalho, M. E. Melendez and A. Lopes Carvalho, Expert Rev. Mol. Diagn., 2018, 18, 85–112 CrossRef CAS PubMed.
- M. P. McRae, K. S. Rajsri, T. M. Alcorn and J. T. McDevitt, Sensors, 2022, 22(17), 6355 CrossRef PubMed.
- S. K. Ghosh, Y. Man, A. Fraiwan, C. Waters, C. McKenzie, C. Lu, D. Pfau, H. Kawsar, N. Bhaskaran, P. Pandiyan, G. Jin, F. Briggs, C. C. Zender, R. Rezaee, F. Panagakos, J. E. Thuener, J. Wasman, A. Tang, H. Qari, T. Wise-Draper, T. S. McCormick, A. Madabhushi, U. A. Gurkan and A. Weinberg, Cell Rep. Med., 2024, 5, 101447 CrossRef CAS PubMed.
- S. F. Altekruse, K. A. McGlynn and M. E. Reichman, J. Clin. Oncol., 2009, 27, 1485–1491 CrossRef PubMed.
- A. Villanueva, N. Engl. J. Med., 2019, 380, 1450–1462 CrossRef CAS PubMed.
- C. B. van der Pol, C. S. Lim, C. B. Sirlin, T. A. McGrath, J. P. Salameh, M. R. Bashir, A. Tang, A. G. Singal, A. F. Costa, K. Fowler and M. D. F. McInnes, Gastroenterology, 2019, 156, 976–986 CrossRef PubMed.
- Y. T. Lee, J. J. Wang, Y. Zhu, V. G. Agopian, H. R. Tseng and J. D. Yang, Clin. Liver Dis., 2021, 17, 409–413 CrossRef PubMed.
- A. Furlan, N. Dasyam, C. Buros, C. P. Thompson, M. I. Minervini and A. S. Kierans, Curr. Probl. Diagn. Radiol., 2024, 53, 235–238 CrossRef PubMed.
- A. Villanueva and Y. Hoshida, J. Hepatol., 2011, 55, 724–725 CrossRef PubMed.
- C. Xu, Z. Xu, Y. Zhang, M. Evert, D. F. Calvisi and X. Chen, J. Clin. Invest., 2022, 132(4), e154515 CrossRef CAS PubMed.
- J. Calderaro, M. Ziol, V. Paradis and J. Zucman-Rossi, J. Hepatol., 2019, 71, 616–630 CrossRef CAS PubMed.
- S. Rebouissou and J. C. Nault, J. Hepatol., 2020, 72, 215–229 CrossRef CAS PubMed.
- A. X. Zhu, A. R. Abbas, M. R. de Galarreta, Y. Guan, S. Lu, H. Koeppen, W. Zhang, C. H. Hsu, A. R. He, B. Y. Ryoo, T. Yau, A. O. Kaseb, A. M. Burgoyne, F. Dayyani, J. Spahn, W. Verret, R. S. Finn, H. C. Toh, A. Lujambio and Y. Wang, Nat. Med., 2022, 28, 1599–1611 CrossRef CAS PubMed.
- D. Kadi, M. F. Yamamoto, E. C. Lerner, H. Jiang, K. J. Fowler and M. R. Bashir, J. Liver Cancer, 2023, 23, 284–299 CrossRef PubMed.
- M. Reig, A. Forner, J. Rimola, J. Ferrer-Fabrega, M. Burrel, A. Garcia-Criado, R. K. Kelley, P. R. Galle, V. Mazzaferro, R. Salem, B. Sangro, A. G. Singal, A. Vogel, J. Fuster, C. Ayuso and J. Bruix, J. Hepatol., 2022, 76, 681–693 CrossRef PubMed.
- Y. Bao, J. X. Li, P. Zhou, Y. Tong, L. Z. Wang, D. H. Chang, W. W. Cai, L. Wen, J. Liu and Y. D. Xiao, Radiology, 2023, 308, e230457 CrossRef PubMed.
- D. B. McGill, J. Rakela, A. R. Zinsmeister and B. J. Ott, Gastroenterology, 1990, 99, 1396–1400 CrossRef CAS PubMed.
- M. A. Silva, B. Hegab, C. Hyde, B. Guo, J. A. Buckels and D. F. Mirza, Gut, 2008, 57, 1592–1596 CrossRef CAS PubMed.
- R. Xue, L. Chen, C. Zhang, M. Fujita, R. Li, S. M. Yan, C. K. Ong, X. Liao, Q. Gao, S. Sasagawa, Y. Li, J. Wang, H. Guo, Q. T. Huang, Q. Zhong, J. Tan, L. Qi, W. Gong, Z. Hong, M. Li, J. Zhao, T. Peng, Y. Lu, K. H. T. Lim, A. Boot, A. Ono, K. Chayama, Z. Zhang, S. G. Rozen, B. T. Teh, X. W. Wang, H. Nakagawa, M. S. Zeng, F. Bai and N. Zhang, Cancer Cell, 2019, 35(6), 932–947 CrossRef CAS PubMed.
- X. Xu, Y. Tao, L. Shan, R. Chen, H. Jiang, Z. Qian, F. Cai, L. Ma and Y. Yu, J. Cancer, 2018, 9, 3557–3569 CrossRef PubMed.
- C. Campani, S. Imbeaud, G. Couchy, M. Ziol, T. Z. Hirsch, S. Rebouissou, B. Noblet, P. Nahon, K. Hormigos, S. Sidali, O. Seror, V. Taly, N. Ganne Carrie, P. Laurent-Puig, J. Zucman-Rossi and J. C. Nault, Gut, 2024, 73, 1870–1882 CrossRef CAS PubMed.
- D. A. Ezeife, E. Spackman, R. A. Juergens, J. J. Laskin, J. S. Agulnik, D. Hao, S. A. Laurie, J. H. Law, L. W. Le, L. A. Kiedrowski, B. Melosky, F. A. Shepherd, V. Cohen, P. Wheatley-Price, R. Vandermeer, J. J. Li, R. Fernandes, A. Shokoohi, R. B. Lanman and N. B. Leighl, Ther. Adv. Med. Oncol., 2022, 14, 17588359221112696 CrossRef PubMed.
- S. Connal, J. M. Cameron, A. Sala, P. M. Brennan, D. S. Palmer, J. D. Palmer, H. Perlow and M. J. Baker, J. Transl. Med., 2023, 21, 118 CrossRef PubMed.
- R. L. Siegel, T. B. Kratzer, A. N. Giaquinto, H. Sung and A. Jemal, Ca-Cancer J. Clin., 2025, 75, 10–45 CrossRef PubMed.
- S. Gillessen, A. Bossi, I. D. Davis, J. de Bono, K. Fizazi, N. D. James, N. Mottet, N. Shore, E. Small, M. Smith, C. J. Sweeney, B. Tombal, E. S. Antonarakis, A. M. Aparicio, A. J. Armstrong, G. Attard, T. M. Beer, H. Beltran, A. Bjartell, P. Blanchard, A. Briganti, R. G. Bristow, M. Bulbul, O. Caffo, D. Castellano, E. Castro, H. H. Cheng, K. N. Chi, S. Chowdhury, C. S. Clarke, N. Clarke, G. Daugaard, M. De Santis, I. Duran, R. Eeles, E. Efstathiou, J. Efstathiou, O. N. Ekeke, C. P. Evans, S. Fanti, F. Y. Feng, V. Fonteyne, N. Fossati, M. Frydenberg, D. George, M. Gleave, G. Gravis, S. Halabi, D. Heinrich, K. Herrmann, C. Higano, M. S. Hofman, L. G. Horvath, M. Hussain, B. A. Jereczek-Fossa, R. Jones, R. Kanesvaran, P. L. Kellokumpu-Lehtinen, R. B. Khauli, L. Klotz, G. Kramer, R. Leibowitz, C. Logothetis, B. Mahal, F. Maluf, J. Mateo, D. Matheson, N. Mehra, A. Merseburger, A. K. Morgans, M. J. Morris, H. Mrabti, D. Mukherji, D. G. Murphy, V. Murthy, P. L. Nguyen, W. K. Oh, P. Ost, J. M. O'Sullivan, A. R. Padhani, C. J. Pezaro, D. M. C. Poon, C. C. Pritchard, D. M. Rabah, D. Rathkopf, R. E. Reiter, M. A. Rubin, C. J. Ryan, F. Saad, J. P. Sade, O. Sartor, H. I. Scher, N. Sharifi, I. Skoneczna, H. Soule, D. E. Spratt, S. Srinivas, C. N. Sternberg, T. Steuber, H. Suzuki, M. R. Sydes, M. E. Taplin, D. Tilki, L. Turkeri, F. Turco, H. Uemura, H. Uemura, Y. Urun, C. L. Vale, I. van Oort, N. Vapiwala, J. Walz, K. Yamoah, D. Ye, E. Y. Yu, A. Zapatero, T. Zilli and A. Omlin, Eur. J. Cancer, 2023, 185, 178–215 CrossRef PubMed.
- R. E. Jimenez, T. D. Atwell, H. Sicotte, B. Eckloff, L. Wang, P. Barman, J. P. Sinnwell, P. W. Eiken, B. P. McMenomy, W. Tan, L. Wang, R. E. Carlson and M. Kohli, Mayo Clin. Proc. Innov. Qual. Outcomes, 2019, 3, 14–22 CrossRef PubMed.
- J. Larson, M. O. Ozen, M. Kohli, D. Akin and U. Demirci, Adv. Biol., 2023, 7, e2200327 CrossRef PubMed.
- R. Iacovelli, C. Ciccarese, G. Schinzari, E. Rossi, B. A. Maiorano, S. Astore, T. D'Angelo, A. Cannella, C. Pirozzoli, M. A. Teberino, F. Pierconti, M. Martini and G. Tortora, Expert Rev. Mol. Diagn., 2020, 20, 195–205 CrossRef CAS PubMed.
- G. Ploussard, S. Terry, P. Maille, Y. Allory, N. Sirab, L. Kheuang, P. Soyeux, N. Nicolaiew, E. Coppolani, B. Paule, L. Salomon, S. Culine, R. Buttyan, F. Vacherot and A. de la Taille, Cancer Res., 2010, 70, 9253–9264 CrossRef CAS PubMed.
- J. S. de Bono, H. I. Scher, R. B. Montgomery, C. Parker, M. C. Miller, H. Tissing, G. V. Doyle, L. W. Terstappen, K. J. Pienta and D. Raghavan, Clin. Cancer Res., 2008, 14, 6302–6309 CrossRef CAS PubMed.
- A. Goldkorn, B. Ely, D. I. Quinn, C. M. Tangen, L. M. Fink, T. Xu, P. Twardowski, P. J. Van Veldhuizen, N. Agarwal, M. A. Carducci, J. P. Monk, 3rd, R. H. Datar, M. Garzotto, P. C. Mack, P. Lara Jr., C. S. Higano, M. Hussain, I. M. Thompson Jr., R. J. Cote and N. J. Vogelzang, J. Clin. Oncol., 2014, 32, 1136–1142 CrossRef CAS PubMed.
- E. S. Antonarakis, C. Lu, B. Luber, H. Wang, Y. Chen, M. Nakazawa, R. Nadal, C. J. Paller, S. R. Denmeade, M. A. Carducci, M. A. Eisenberger and J. Luo, JAMA Oncol., 2015, 1, 582–591 CrossRef PubMed.
- M. Nakazawa, C. Lu, Y. Chen, C. J. Paller, M. A. Carducci, M. A. Eisenberger, J. Luo and E. S. Antonarakis, Ann. Oncol., 2015, 26, 1859–1865 CrossRef CAS PubMed.
- W. Onstenk, A. M. Sieuwerts, J. Kraan, M. Van, A. J. Nieuweboer, R. H. Mathijssen, P. Hamberg, H. J. Meulenbeld, B. De Laere, L. Y. Dirix, R. J. van Soest, M. P. Lolkema, J. W. Martens, W. M. van Weerden, G. W. Jenster, J. A. Foekens, R. de Wit and S. Sleijfer, Eur. Urol., 2015, 68, 939–945 CrossRef CAS PubMed.
- H. I. Scher, R. P. Graf, N. A. Schreiber, B. McLaughlin, D. Lu, J. Louw, D. C. Danila, L. Dugan, A. Johnson, G. Heller, M. Fleisher and R. Dittamore, Eur. Urol., 2017, 71, 874–882 CrossRef CAS PubMed.
- H. I. Scher, D. Lu, N. A. Schreiber, J. Louw, R. P. Graf, H. A. Vargas, A. Johnson, A. Jendrisak, R. Bambury, D. Danila, B. McLaughlin, J. Wahl, S. B. Greene, G. Heller, D. Marrinucci, M. Fleisher and R. Dittamore, JAMA Oncol., 2016, 2, 1441–1449 CrossRef PubMed.
- S. T. Tagawa, E. S. Antonarakis, A. Gjyrezi, G. Galletti, S. Kim, D. Worroll, J. Stewart, A. Zaher, T. P. Szatrowski, K. V. Ballman, K. Kita, S. Tasaki, Y. Bai, L. Portella, B. J. Kirby, F. Saad, M. A. Eisenberger, D. M. Nanus and P. Giannakakou, Clin. Cancer Res., 2019, 25, 1880–1888 CrossRef CAS PubMed.
- V. Conteduca, A. Jayaram, N. Romero-Laorden, D. Wetterskog, S. Salvi, G. Gurioli, E. Scarpi, E. Castro, M. Marin-Aguilera, C. Lolli, G. Schepisi, A. Maugeri, A. Wingate, A. Farolfi, V. Casadio, A. Medina, J. Puente, M. J. M. Vidal, R. Morales-Barrera, J. C. Villa-Guzman, S. Hernando, A. Rodriguez-Vida, A. Gonzalez-Del-Alba, B. Mellado, E. Gonzalez-Billalabeitia, D. Olmos, G. Attard and U. De Giorgi, Eur. Urol., 2019, 75, 368–373 CrossRef CAS PubMed.
- V. Conteduca, D. Wetterskog, M. T. A. Sharabiani, E. Grande, M. P. Fernandez-Perez, A. Jayaram, S. Salvi, D. Castellano, A. Romanel, C. Lolli, V. Casadio, G. Gurioli, D. Amadori, A. Font, S. Vazquez-Estevez, A. Gonzalez Del Alba, B. Mellado, O. Fernandez-Calvo, M. J. Mendez-Vidal, M. A. Climent, I. Duran, E. Gallardo, A. Rodriguez, C. Santander, M. I. Saez, J. Puente, D. Gasi Tandefelt, A. Wingate, D. Dearnaley, P. Collaborators, G. Spanish Oncology Genitourinary, F. Demichelis, U. De Giorgi, E. Gonzalez-Billalabeitia and G. Attard, Ann. Oncol., 2017, 28, 1508–1516 CrossRef CAS PubMed.
- M. Du, Y. Tian, W. Tan, L. Wang, L. Wang, D. Kilari, C. C. Huang, L. Wang and M. Kohli, Prostate Cancer Prostatic Dis., 2020, 23, 705–713 CrossRef CAS PubMed.
- A. Romanel, D. Gasi Tandefelt, V. Conteduca, A. Jayaram, N. Casiraghi, D. Wetterskog, S. Salvi, D. Amadori, Z. Zafeiriou, P. Rescigno, D. Bianchini, G. Gurioli, V. Casadio, S. Carreira, J. Goodall, A. Wingate, R. Ferraldeschi, N. Tunariu, P. Flohr, U. De Giorgi, J. S. de Bono, F. Demichelis and G. Attard, Sci. Transl. Med., 2015, 7, 312re310 Search PubMed.
- S. Salvi, V. Casadio, V. Conteduca, S. L. Burgio, C. Menna, E. Bianchi, L. Rossi, E. Carretta, C. Masini, D. Amadori, D. Calistri, G. Attard and U. De Giorgi, Br. J. Cancer, 2015, 112, 1717–1724 CrossRef CAS PubMed.
- S. Salvi, V. Casadio, V. Conteduca, C. Lolli, G. Gurioli, F. Martignano, G. Schepisi, S. Testoni, E. Scarpi, D. Amadori, D. Calistri, G. Attard and U. De Giorgi, Oncotarget, 2016, 7, 37839–37845 CrossRef PubMed.
- W. Abida, J. Cyrta, G. Heller, D. Prandi, J. Armenia, I. Coleman, M. Cieslik, M. Benelli, D. Robinson, E. M. Van Allen, A. Sboner, T. Fedrizzi, J. M. Mosquera, B. D. Robinson, N. De Sarkar, L. P. Kunju, S. Tomlins, Y. M. Wu, D. Nava Rodrigues, M. Loda, A. Gopalan, V. E. Reuter, C. C. Pritchard, J. Mateo, D. Bianchini, S. Miranda, S. Carreira, P. Rescigno, J. Filipenko, J. Vinson, R. B. Montgomery, H. Beltran, E. I. Heath, H. I. Scher, P. W. Kantoff, M. E. Taplin, N. Schultz, J. S. deBono, F. Demichelis, P. S. Nelson, M. A. Rubin, A. M. Chinnaiyan and C. L. Sawyers, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 11428–11436 CrossRef CAS PubMed.
- M. Annala, G. Vandekerkhove, D. Khalaf, S. Taavitsainen, K. Beja, E. W. Warner, K. Sunderland, C. Kollmannsberger, B. J. Eigl, D. Finch, C. D. Oja, J. Vergidis, M. Zulfiqar, A. A. Azad, M. Nykter, M. E. Gleave, A. W. Wyatt and K. N. Chi, Cancer Discovery, 2018, 8, 444–457 CrossRef CAS PubMed.
- E. Gonzalez-Billalabeitia, V. Conteduca, D. Wetterskog, A. Jayaram and G. Attard, Prostate Cancer Prostatic Dis., 2019, 22, 195–205 CrossRef CAS PubMed.
- A. Kienel, D. Porres, A. Heidenreich and D. Pfister, J. Urol., 2015, 194, 966–971 CrossRef CAS PubMed.
- S. Kwee, M. A. Song, I. Cheng, L. Loo and M. Tiirikainen, Clin. Transl. Sci., 2012, 5, 65–70 CrossRef PubMed.
- N. Mehra, D. Dolling, S. Sumanasuriya, R. Christova, L. Pope, S. Carreira, G. Seed, W. Yuan, J. Goodall, E. Hall, P. Flohr, G. Boysen, D. Bianchini, O. Sartor, M. A. Eisenberger, K. Fizazi, S. Oudard, M. Chadjaa, S. Mace and J. S. de Bono, Eur. Urol., 2018, 74, 283–291 CrossRef CAS PubMed.
- H. Sicotte, K. R. Kalari, S. Qin, S. M. Dehm, V. Bhargava, M. Gormley, W. Tan, J. P. Sinnwell, D. W. Hillman, Y. Li, P. T. Vedell, R. E. Carlson, A. H. Bryce, R. E. Jimenez, R. M. Weinshilboum, M. Kohli and L. Wang, Mol. Cancer Res., 2022, 20, 1739–1750 CrossRef CAS PubMed.
- L. Wang, S. M. Dehm, D. W. Hillman, H. Sicotte, W. Tan, M. Gormley, V. Bhargava, R. Jimenez, F. Xie, P. Yin, S. Qin, F. Quevedo, B. A. Costello, H. C. Pitot, T. Ho, A. H. Bryce, Z. Ye, Y. Li, P. Eiken, P. T. Vedell, P. Barman, B. P. McMenomy, T. D. Atwell, R. E. Carlson, M. Ellingson, B. W. Eckloff, R. Qin, F. Ou, S. N. Hart, H. Huang, J. Jen, E. D. Wieben, K. R. Kalari, R. M. Weinshilboum, L. Wang and M. Kohli, Ann. Oncol., 2018, 29, 352–360 CrossRef CAS PubMed.
- D. Robinson, E. M. Van Allen, Y. M. Wu, N. Schultz, R. J. Lonigro, J. M. Mosquera, B. Montgomery, M. E. Taplin, C. C. Pritchard, G. Attard, H. Beltran, W. Abida, R. K. Bradley, J. Vinson, X. Cao, P. Vats, L. P. Kunju, M. Hussain, F. Y. Feng, S. A. Tomlins, K. A. Cooney, D. C. Smith, C. Brennan, J. Siddiqui, R. Mehra, Y. Chen, D. E. Rathkopf, M. J. Morris, S. B. Solomon, J. C. Durack, V. E. Reuter, A. Gopalan, J. Gao, M. Loda, R. T. Lis, M. Bowden, S. P. Balk, G. Gaviola, C. Sougnez, M. Gupta, E. Y. Yu, E. A. Mostaghel, H. H. Cheng, H. Mulcahy, L. D. True, S. R. Plymate, H. Dvinge, R. Ferraldeschi, P. Flohr, S. Miranda, Z. Zafeiriou, N. Tunariu, J. Mateo, R. Perez-Lopez, F. Demichelis, B. D. Robinson, M. Schiffman, D. M. Nanus, S. T. Tagawa, A. Sigaras, K. W. Eng, O. Elemento, A. Sboner, E. I. Heath, H. I. Scher, K. J. Pienta, P. Kantoff, J. S. de Bono, M. A. Rubin, P. S. Nelson, L. A. Garraway, C. L. Sawyers and A. M. Chinnaiyan, Cell, 2015, 161, 1215–1228 CrossRef CAS PubMed.
- V. A. Adalsteinsson, G. Ha, S. S. Freeman, A. D. Choudhury, D. G. Stover, H. A. Parsons, G. Gydush, S. C. Reed, D. Rotem, J. Rhoades, D. Loginov, D. Livitz, D. Rosebrock, I. Leshchiner, J. Kim, C. Stewart, M. Rosenberg, J. M. Francis, C. Z. Zhang, O. Cohen, C. Oh, H. Ding, P. Polak, M. Lloyd, S. Mahmud, K. Helvie, M. S. Merrill, R. A. Santiago, E. P. O'Connor, S. H. Jeong, R. Leeson, R. M. Barry, J. F. Kramkowski, Z. Zhang, L. Polacek, J. G. Lohr, M. Schleicher, E. Lipscomb, A. Saltzman, N. M. Oliver, L. Marini, A. G. Waks, L. C. Harshman, S. M. Tolaney, E. M. Van Allen, E. P. Winer, N. U. Lin, M. Nakabayashi, M. E. Taplin, C. M. Johannessen, L. A. Garraway, T. R. Golub, J. S. Boehm, N. Wagle, G. Getz, J. C. Love and M. Meyerson, Nat. Commun., 2017, 8, 1324 CrossRef PubMed.
- D. H. Hovelson, C. J. Liu, Y. Wang, Q. Kang, J. Henderson, A. Gursky, S. Brockman, N. Ramnath, J. C. Krauss, M. Talpaz, M. Kandarpa, R. Chugh, M. Tuck, K. Herman, C. S. Grasso, M. J. Quist, F. Y. Feng, C. Haakenson, J. Langmore, E. Kamberov, T. Tesmer, H. Husain, R. J. Lonigro, D. Robinson, D. C. Smith, A. S. Alva, M. H. Hussain, A. M. Chinnaiyan, M. Tewari, R. E. Mills, T. M. Morgan and S. A. Tomlins, Oncotarget, 2017, 8, 89848–89866 CrossRef PubMed.
- A. W. Wyatt, M. Annala, R. Aggarwal, K. Beja, F. Feng, J. Youngren, A. Foye, P. Lloyd, M. Nykter, T. M. Beer, J. J. Alumkal, G. V. Thomas, R. E. Reiter, M. B. Rettig, C. P. Evans, A. C. Gao, K. N. Chi, E. J. Small and M. E. Gleave, J. Natl. Cancer Inst., 2017, 109(12), djx118 CrossRef PubMed.
- J. Huang, M. Du, A. Soupir, L. Wang, W. Tan, K. R. Kalari, D. Kilari, J. Park, C. C. Huang, M. Kohli and L. Wang, Cancers, 2022, 14(19), 4714 CrossRef CAS PubMed.
- F. Causa, A. Aliberti, A. M. Cusano, E. Battista and P. A. Netti, J. Am. Chem. Soc., 2015, 137, 1758–1761 CrossRef CAS PubMed.
- F. Moltzahn, A. B. Olshen, L. Baehner, A. Peek, L. Fong, H. Stöppler, J. Simko, J. F. Hilton, P. Carroll and R. Blelloch, Cancer Res., 2011, 71, 550–560 CrossRef CAS PubMed.
- C. Li, Z. Li, H. Jia and J. Yan, Chem. Commun., 2011, 47, 2595–2597 RSC.
- G. T. Walker, M. S. Fraiser, J. L. Schram, M. C. Little, J. G. Nadeau and D. P. Malinowski, Nucleic Acids Res., 1992, 20, 1691–1696 CrossRef CAS PubMed.
- L. Yang, J. Fang, J. Li, X. Ou, L. Zhang, Y. Wang, Z. Weng and G. Xie, Anal. Chim. Acta, 2021, 1143, 157–165 CrossRef CAS PubMed.
- R.-D. Li, B.-C. Yin and B.-C. Ye, Biosens. Bioelectron., 2016, 86, 1011–1016 CrossRef CAS PubMed.
- M. M. Ali, F. Li, Z. Zhang, K. Zhang, D.-K. Kang, J. A. Ankrum, X. C. Le and W. Zhao, Chem. Soc. Rev., 2014, 43, 3324–3341 RSC.
- J. Bao, C. Hou, Y. Zhao, X. Geng, M. Samalo, H. Yang, M. Bian and D. Huo, Talanta, 2019, 196, 329–336 CrossRef CAS PubMed.
- Z. Wu, G.-Q. Liu, X.-L. Yang and J.-H. Jiang, J. Am. Chem. Soc., 2015, 137(21), 6829–6836 CrossRef CAS PubMed.
- Y. Nie, X. Yuan, P. Zhang, Y. Q. Chai and R. Yuan, Anal. Chem., 2019, 91, 3452–3458 CrossRef CAS PubMed.
- C.-P. Liang, P.-Q. Ma, H. Liu, X. Guo, B.-C. Yin and B.-C. Ye, Angew. Chem., Int. Ed., 2017, 56, 9077–9081 CrossRef CAS PubMed.
- P. H. McCord, Euphytica, 2016, 209, 439–448 CrossRef CAS.
- Y. Zhang, E.-T. Tang and Z. Du, PLoS One, 2016, 11, e0146784 CrossRef PubMed.
- M. Lodrini, A. Sprüssel, K. Astrahantseff, D. Tiburtius, R. Konschak, H. N. Lode, M. Fischer, U. Keilholz, A. Eggert and H. E. Deubzer, Oncotarget, 2017, 8, 85234–85251 CrossRef PubMed.
- A. S. Whale, J. F. Huggett, S. Cowen, V. Speirs, J. Shaw, S. Ellison, C. A. Foy and D. J. Scott, Nucleic Acids Res., 2012, 40, e82 CrossRef CAS PubMed.
- N. J. Heredia, P. Belgrader, S. Wang, R. Koehler, J. Regan, A. M. Cosman, S. Saxonov, B. Hindson, S. C. Tanner, A. S. Brown and G. Karlin-Neumann, Methods, 2013, 59, S20–S23 CrossRef CAS PubMed.
- D. Mirgh, A. Krishnan and S. Gorai, J. Liq. Biopsy, 2023, 2, 100122 CrossRef PubMed.
- Z. Pos, O. Pos, J. Styk, A. Mocova, L. Strieskova, J. Budis, L. Kadasi, J. Radvanszky and T. Szemes, Int. J. Mol. Sci., 2020, 21(22), 8634 CrossRef CAS PubMed.
- Y. Naito and K. Honda, J. Pers. Med., 2023, 13(2), 303 CrossRef PubMed.
- B. Michela, Diagnostics, 2021, 11(8), 1391 CrossRef CAS PubMed.
- L. L. Leung, X. Qu, B. Chen and J. Y. Chan, Extracell. Vesicles Circ. Nucleic Acids, 2024, 5, 639–659 CAS.
- D. He, B. Cui, H. Lv, S. Lu, Y. Zhu, Y. Cheng, L. Dang and H. Zhang, Biomolecules, 2024, 14(7), 847 CrossRef CAS PubMed.
- D. Yu, Y. Li, M. Wang, J. Gu, W. Xu, H. Cai, X. Fang and X. Zhang, Mol. Cancer, 2022, 21, 56 CrossRef PubMed.
- Q. R. Guo, L. L. Zhang, J. F. Liu, Z. Li, J. J. Li, W. M. Zhou, H. Wang, J. Q. Li, D. Y. Liu, X. Y. Yu and J. Y. Zhang, Nanotheranostics, 2021, 5, 73–89 CrossRef PubMed.
- S. Haghjooy Javanmard, L. Rafiee, M. Bahri Najafi, D. Khorsandi, A. Hasan, G. Vaseghi and P. Makvandi, Environ. Res., 2023, 238, 117083 CrossRef CAS PubMed.
- H. Kupcova Skalnikova, B. Bohuslavova, K. Turnovcova, J. Juhasova, S. Juhas, M. Rodinova and P. Vodicka, Proteomes, 2019, 7, 17 CrossRef PubMed.
- K. B. Johnsen, J. M. Gudbergsson, T. L. Andresen and J. B. Simonsen, Biochim. Biophys. Acta, Rev. Cancer, 2019, 1871, 109–116 CrossRef CAS PubMed.
- H. Markus, T. Contente-Cuomo, M. Farooq, W. S. Liang, M. J. Borad, S. Sivakumar, S. Gollins, N. L. Tran, H. D. Dhruv, M. E. Berens, A. Bryce, A. Sekulic, A. Ribas, J. M. Trent, P. M. LoRusso and M. Murtaza, Sci. Rep., 2018, 8, 7375 CrossRef PubMed.
- H. Peng, M. Pan, Z. Zhou, C. Chen, X. Xing, S. Cheng, S. Zhang, H. Zheng and K. Qian, Front. Cell Dev. Biol., 2024, 12, 1385041 CrossRef PubMed.
- Z. Wang, F. Li, J. Rufo, C. Chen, S. Yang, L. Li, J. Zhang, J. Cheng, Y. Kim, M. Wu, E. Abemayor, M. Tu, D. Chia, R. Spruce, N. Batis, H. Mehanna, D. T. W. Wong and T. J. Huang, J. Mol. Diagn., 2020, 22, 50–59 CrossRef CAS PubMed.
- A. Vijay, T. Inui, M. Dodds, G. Proctor and G. Carpenter, PLoS One, 2015, 10, e0135792 CrossRef PubMed.
- N. M. Chaudhury, G. B. Proctor, N. G. Karlsson, G. H. Carpenter and S. A. Flowers, Mol. Cell. Proteomics, 2016, 15, 1048–1059 CrossRef CAS PubMed.
- C. Cafolla, J. Philpott-Robson, A. Elbourne and K. Voïtchovsky, ACS Appl. Mater. Interfaces, 2024, 16, 44–53 CrossRef CAS PubMed.
- S. Bakhshandeh, Nat. Rev. Bioeng., 2025, 3, 359–359 CrossRef CAS.
- A. Eberhard, T. Moser, L. Ziegler, G. Vlachos, M. Loibner, T. Bauernhofer, M. Balic, A. Gerger, N. Dandachi, C. Beichler, L. Glawitsch, M. Moser, R. Graf, P. M. Abuja, M. Schmitz, T. Krenz, T. Voss, D. Mancarella and E. Heitzer, iScience, 2025, 28, 113632 CrossRef CAS PubMed.
- N. Lee, A. Canagasingham, M. Bajaj, R. Shanmugasundaram, A. Hutton, J. Bucci, P. Graham, J. Thompson and J. Ni, Front. Oncol., 2022, 12, 1019391 CrossRef CAS PubMed.
- L. G. Liang, M. Q. Kong, S. Zhou, Y. F. Sheng, P. Wang, T. Yu, F. Inci, W. P. Kuo, L. J. Li, U. Demirci and S. Wang, Sci. Rep., 2017, 7, 46224 CrossRef PubMed.
- Y. Wang, Y. Gao and Y. Song, ChemMedChem, 2022, 17, e202200422 CrossRef CAS PubMed.
- J. L. Chang, C. J. Huang, Y. C. Tsai, N. J. Chiang, Y. S. Huang, S. C. Hung, Y. S. Shan and G. B. Lee, Lab Chip, 2024, 24, 375–382 RSC.
- L. Y. Hung, N. J. Chiang, W. C. Tsai, C. Y. Fu, Y. C. Wang, Y. S. Shan and G. B. Lee, Sci. Rep., 2017, 7, 4248 CrossRef PubMed.
- F. Liu, X. Hao, B. Liu, S. Liu and Y. Yuan, Clin. Chim. Acta, 2023, 551, 117593 CrossRef CAS PubMed.
- Y. Zhang, J. Zhao, H. Yu, P. Li, W. Liang, Z. Liu, G. B. Lee, L. Liu, W. J. Li and Z. Wang, Sci. Adv., 2020, 6, eaba9628 CrossRef CAS PubMed.
- M. A. Mansor, C. Yang, K. L. Chong, M. A. Jamrus, K. Liu, M. Yu, M. R. Ahmad and X. Ren, ACS Omega, 2025, 10, 6306–6322 CrossRef CAS PubMed.
- S. Cheng, Y. Li, H. Yan, Y. Wen, X. Zhou, L. Friedman and Y. Zeng, Lab Chip, 2021, 21, 3219–3243 RSC.
- J. M. Nikoloff, M. A. Saucedo-Espinosa and P. S. Dittrich, Anal. Chem., 2023, 95, 1933–1939 CrossRef CAS PubMed.
- Y. Meng, M. Asghari, M. K. Aslan, A. Yilmaz, B. Mateescu, S. Stavrakis and A. J. deMello, Chem. Eng. J., 2021, 404, 126110 CrossRef CAS.
- F. Gao, C. Liu, L. Zhang, T. Liu, Z. Wang, Z. Song, H. Cai, Z. Fang, J. Chen, J. Wang, M. Han, J. Wang, K. Lin, R. Wang, M. Li, Q. Mei, X. Ma, S. Liang, G. Gou and N. Xue, Microsyst. Nanoeng., 2023, 9, 1 CrossRef PubMed.
- B. Ramachandran and Y. C. Liao, Biomicrofluidics, 2022, 16, 051501 CrossRef CAS PubMed.
- S. Li and Q. Ma, Biosens. Bioelectron., 2022, 214, 114554 CrossRef CAS PubMed.
- H. Chi, L. Shi, S. Gan, G. Fan and Y. Dong, Biosensors, 2025, 15(4), 199 CrossRef CAS PubMed.
- J. Nättinen, U. Aapola, A. Jylhä, A. Vaajanen and H. Uusitalo, Transl. Vis. Sci. Technol., 2020, 9, 16 CrossRef PubMed.
- G. H. Ngo, M. Chadha, Y. J. Sun, G. Yu, S. H. Lee, T. C. Yeh, D. R. P. Almeida, A. G. Bassuk, P. Mruthyunjaya, A. Dufour and V. B. Mahajan, STAR Protoc., 2025, 6, 104146 CrossRef CAS PubMed.
- A. Daily, P. Ravishankar, S. Harms and V. S. Klimberg, PLoS One, 2022, 17, e0267676 CrossRef CAS PubMed.
- I. Horváth, J. Hunt, P. J. Barnes, K. Alving, A. Antczak, E. Baraldi, G. Becher, W. J. van Beurden, M. Corradi, R. Dekhuijzen, R. A. Dweik, T. Dwyer, R. Effros, S. Erzurum, B. Gaston, C. Gessner, A. Greening, L. P. Ho, J. Hohlfeld, Q. Jöbsis, D. Laskowski, S. Loukides, D. Marlin, P. Montuschi, A. C. Olin, A. E. Redington, P. Reinhold, E. L. van Rensen, I. Rubinstein, P. Silkoff, K. Toren, G. Vass, C. Vogelberg and H. Wirtz, Eur. Respir. J., 2005, 26, 523–548 CrossRef PubMed.
- B. R. Winters, J. D. Pleil, M. M. Angrish, M. A. Stiegel, T. H. Risby and M. C. Madden, J. Breath Res., 2017, 11, 047107 CrossRef PubMed.
- A. Campanella, S. De Summa and S. Tommasi, J. Breath Res., 2019, 13, 044002 CrossRef CAS PubMed.
- S. Zoupanou, A. Volpe, E. Primiceri, C. Gaudiuso, A. Ancona, F. Ferrara and M. S. Chiriaco, Micromachines, 2021, 12(8), 885 CrossRef PubMed.
- H. Lee, C. Park, W. Na, K. H. Park and S. Shin, npj Precis. Oncol., 2020, 4, 3 CrossRef CAS PubMed.
- H. Gwak, J. Kim, S. Cha, Y. P. Cheon, S. I. Kim, B. Kwak, K. A. Hyun and H. I. Jung, Biomicrofluidics, 2019, 13, 024113 CrossRef PubMed.
- L. Wu, X. Liu, Y. Zhang, Z. Yang, L. Chen, S. Zong, J. Li, Y. Cui and Z. Wang, Sens. Actuators, B, 2024, 401, 135081 CrossRef CAS.
- C. Bayona, T. Ranđelović, C. Olaizola-Rodrigo and I. Ochoa, Bioeng. Transl. Med., 2025, e70032 CrossRef.
- Z. Huang, J. Fang, M. Zhou, Z. Gong and T. Xiang, Front. Microbiol., 2022, 13, 1011399 CrossRef PubMed.
- J. Zhou, Z. Li, J. Seun Olajide and G. Wang, Heliyon, 2024, 10, e26179 CrossRef CAS PubMed.
- S. N. Zakiyyah, K. S. P. Sima, C. Y. Kharismasari, M. Xi, S. Gaffar, M. Ozsoz, F. Paolucci, G. Valenti and Y. W. Hartati, Sens. Actuat. Rep., 2025, 100358 Search PubMed.
- P. Gilson, C. Saurel, J. Salleron, M. Husson, J. Demange, J. L. Merlin and A. Harle, Sci. Rep., 2021, 11, 10470 CrossRef CAS PubMed.
- H. Huang, S. Springborn, K. Haug, K. Bartow, H. Samra, S. Menon and A. C. Mackinnon, J. Mol. Diagn., 2019, 21, 862–872 CrossRef CAS PubMed.
- M. W. Linder, M. E. Egger, T. Van Meter, S. N. Rai, R. Valdes Jr., M. B. Hall, X. Wu, N. Alghamdi and J. A. Chesney, Mol. Diagn. Ther., 2021, 25, 361–371 CrossRef CAS PubMed.
- A. Leone, L. A. Muscarella, P. Graziano, A. Tornese, L. R. Grillo, A. Di Lorenzo, M. Bronzini, S. Scarpino, A. Sparaneo and G. Rossi, Cancers, 2023, 15(1), 292 CrossRef CAS PubMed.
- C. Schmitz-Drager, P. J. Goebell, E. Paxinos, E. Bismarck, J. Chen, P. Balakrishnan, M. Bates, T. Ebert, B. J. Schmitz-Drager and N. Benderska-Soder, Bladder Cancer, 2024, 10, 25–33 CrossRef PubMed.
- P. C. C. Borges, H. B. Spencer, C. Barbosa, V. Costa, A. Furtado, M. C. Leal, C. Lopes, D. Ferreira, A. L. Carvalho, I. Dos-Santos-Silva and L. L. Santos, Ecancermedicalscience, 2023, 17, 1530 CrossRef PubMed.
- O. Sala-Torra, L. Beppu, Q. Wu, E. Welch, E. Berthier, J. P. Radich and V. G. Oehler, Leukemia, 2024, 38, 1822–1824 CrossRef CAS PubMed.
- S. M. Mousavi, S. M. Amin Mahdian, M. S. Ebrahimi, M. Taghizadieh, M. Vosough, J. Sadri Nahand, S. Hosseindoost, N. Vousooghi, H. A. Javar, B. Larijani, M. R. Hadjighassem, N. Rahimian, M. R. Hamblin and H. Mirzaei, Mol. Ther.--Nucleic Acids, 2022, 28, 758–791 CrossRef CAS PubMed.
- C. Chen, J. Skog, C. H. Hsu, R. T. Lessard, L. Balaj, T. Wurdinger, B. S. Carter, X. O. Breakefield, M. Toner and D. Irimia, Lab Chip, 2010, 10, 505–511 RSC.
- S. S. Kanwar, C. J. Dunlay, D. M. Simeone and S. Nagrath, Lab Chip, 2014, 14, 1891–1900 RSC.
- S. Surappa, P. Multani, U. Parlatan, P. D. Sinawang, J. Kaifi, D. Akin and U. Demirci, Lab Chip, 2023, 23, 2942–2958 RSC.
- M. He, J. Crow, M. Roth, Y. Zeng and A. K. Godwin, Lab Chip, 2014, 14, 3773–3780 RSC.
- H. Im, H. Shao, Y. I. Park, V. M. Peterson, C. M. Castro, R. Weissleder and H. Lee, Nat. Biotechnol., 2014, 32, 490–495 CrossRef CAS PubMed.
- B. H. Wunsch, J. T. Smith, S. M. Gifford, C. Wang, M. Brink, R. L. Bruce, R. H. Austin, G. Stolovitzky and Y. Astier, Nat. Nanotechnol., 2016, 11, 936–940 CrossRef CAS PubMed.
- H. Xu, C. Liao, P. Zuo, Z. Liu and B. C. Ye, Anal. Chem., 2018, 90, 13451–13458 CrossRef CAS PubMed.
- J. Li, Y. Li, P. Li, Y. Zhang, L. Du, Y. Wang, C. Zhang and C. Wang, Acta Biomater., 2022, 144, 1–14 CrossRef CAS PubMed.
- V. Sunkara, C. J. Kim, J. Park, H. K. Woo, D. Kim, H. K. Ha, M. H. Kim, Y. Son, J. R. Kim and Y. K. Cho, Theranostics, 2019, 9, 1851–1863 CrossRef CAS PubMed.
- H. K. Woo, V. Sunkara, J. Park, T. H. Kim, J. R. Han, C. J. Kim, H. I. Choi, Y. K. Kim and Y. K. Cho, ACS Nano, 2017, 11, 1360–1370 CrossRef CAS PubMed.
- L. Leggio, G. Paterno, S. Vivarelli, A. Bonasera, B. Pignataro, N. Iraci and G. Arrabito, iScience, 2023, 26, 108105 CrossRef CAS PubMed.
- M. Havers, A. Broman, A. Lenshof and T. Laurell, Anal. Bioanal. Chem., 2023, 415, 1265–1285 CrossRef CAS PubMed.
- F. Liu, O. Vermesh, V. Mani, T. J. Ge, S. J. Madsen, A. Sabour, E. C. Hsu, G. Gowrishankar, M. Kanada, J. V. Jokerst, R. G. Sierra, E. Chang, K. Lau, K. Sridhar, A. Bermudez, S. J. Pitteri, T. Stoyanova, R. Sinclair, V. S. Nair, S. S. Gambhir and U. Demirci, ACS Nano, 2017, 11, 10712–10723 CrossRef CAS PubMed.
- P. D. Sinawang, P. Multani, M. O. Ozen, J. Wong, D. Akin, C. Hanson, M. Larsen, E. Ampaw, M. T. Rondina, N. D. Tolley, L. Wang, B. T. Cunningham, M. Kohli and U. Demirci, J. Extracell. Biol., 2025, 4, e70071 CrossRef CAS PubMed.
- L. Liu, T. Ayupova, S. Umrao, L. D. Akin, H. K. Lee, J. Tibbs, X. Wang, U. Demirci and B. T. Cunningham, Lab Chip, 2025, 25, 2073–2084 RSC.
- L. Liu, J. Tibbs, N. Li, A. Bacon, S. Shepherd, H. Lee, N. Chauhan, U. Demirci, X. Wang and B. T. Cunningham, Biosens. Bioelectron., 2023, 228, 115197 CrossRef CAS PubMed.
- A. Ku, H. C. Lim, M. Evander, H. Lilja, T. Laurell, S. Scheding and Y. Ceder, Anal. Chem., 2018, 90, 8011–8019 CrossRef CAS PubMed.
- Z. Zhao, Y. Yang, Y. Zeng and M. He, Lab Chip, 2016, 16, 489–496 RSC.
- M. Sharma, M. Sheth, H. M. Poling, D. Kuhnell, S. M. Langevin and L. Esfandiari, Sci. Rep., 2023, 13, 18293 CrossRef CAS PubMed.
- L. Shi, D. Kuhnell, V. J. Borra, S. M. Langevin, T. Nakamura and L. Esfandiari, Lab Chip, 2019, 19, 3726–3734 RSC.
- M. Hu, V. Brown, J. M. Jackson, H. Wijerathne, H. Pathak, D. C. Koestler, E. Nissen, M. L. Hupert, R. Muller, A. K. Godwin, M. A. Witek and S. A. Soper, Anal. Chem., 2023, 95, 7665–7675 CrossRef CAS PubMed.
- L. Descamps, D. Le Roy and A. L. Deman, Int. J. Mol. Sci., 2022, 23(4), 1981 CrossRef CAS PubMed.
- S. M. Abusamra, R. Barber, M. Sharafeldin, C. M. Edwards and J. J. Davis, Sens. Diagn., 2024, 3, 562–584 RSC.
- A. Mishra, S. B. Huang, T. Dubash, R. Burr, J. F. Edd, B. S. Wittner, Q. E. Cunneely, V. R. Putaturo, A. Deshpande, E. Antmen, K. A. Gopinathan, K. Otani, Y. Miyazawa, J. E. Kwak, S. Y. Guay, J. Kelly, J. Walsh, L. T. Nieman, I. Galler, P. Chan, M. S. Lawrence, R. J. Sullivan, A. Bardia, D. S. Micalizzi, L. V. Sequist, R. J. Lee, J. W. Franses, D. T. Ting, P. A. R. Brunker, S. Maheswaran, D. T. Miyamoto, D. A. Haber and M. Toner, Nat. Commun., 2025, 16, 32 CrossRef PubMed.
- D. Parvin, Z. S. Hashemi, F. Shokati, Z. Mohammadpour and V. Bazargan, ACS Omega, 2023, 8, 21745–21754 CrossRef CAS PubMed.
- A. D. Ellington and J. W. Szostak, Nature, 1990, 346, 818–822 CrossRef CAS PubMed.
- A. A. Kajani, L. Rafiee, M. Samandari, M. A. Mehrgardi, B. Zarrin and S. H. Javanmard, RSC Adv., 2022, 12, 32834–32843 RSC.
- D. Sun, Y. Ma, M. Wu, Z. Chen, L. Zhang and J. Lu, J. Pharm. Anal., 2023, 13, 340–354 CrossRef PubMed.
- C. Alix-Panabieres and K. Pantel, Nat. Rev. Cancer, 2014, 14, 623–631 CrossRef CAS PubMed.
- N. Herrmann, P. Neubauer and M. Birkholz, Biomicrofluidics, 2019, 13, 061501 CrossRef CAS PubMed.
- C. Magnusson, P. Augustsson, E. Undvall Anand, A. Lenshof, A. Josefsson, K. Welen, A. Bjartell, Y. Ceder, H. Lilja and T. Laurell, Anal. Chem., 2024, 96, 6914–6921 CrossRef CAS PubMed.
- Z. Tian, X. Wang and J. Chen, Microsyst. Nanoeng., 2024, 10, 117 CrossRef CAS PubMed.
- V. Varmazyari, H. Habibiyan, H. Ghafoorifard, M. Ebrahimi and S. Ghafouri-Fard, Sci. Rep., 2022, 12, 12100 CrossRef CAS PubMed.
- I. Najafipour, P. Sadeh, A. M. Amani, H. Kamyab, S. Chelliapan, S. Rajendran, A. B. Peñaherrera-Pazmiño and S. Jamalpour, Sens. Actuat. Rep., 2025, 100304 Search PubMed.
- Z. Guo and W. Xia, Med-X, 2024, 2, 28 CrossRef CAS.
- A. Abdulla, T. Zhang, S. Li, W. Guo, A. R. Warden, Y. Xin, N. Maboyi, J. Lou, H. Xie and X. Ding, Microsyst. Nanoeng., 2022, 8, 13 CrossRef CAS PubMed.
- E. Berthier, E. W. Young and D. Beebe, Lab Chip, 2012, 12, 1224–1237 RSC.
- C. Chen, B. T. Mehl, A. S. Munshi, A. D. Townsend, D. M. Spence and R. S. Martin, Anal. Methods, 2016, 8, 6005–6012 RSC.
- K. Raj M and S. Chakraborty, J. Appl. Polym. Sci., 2020, 137, 48958 CrossRef CAS.
- P. S. Nunes, P. D. Ohlsson, O. Ordeig and J. P. Kutter, Microfluid. Nanofluid., 2010, 9, 145–161 CrossRef CAS.
- C.-W. Tsao, Micromachines, 2016, 7, 225 CrossRef PubMed.
- K. Childers, I. M. Freed, M. L. Hupert, B. Shaw, N. Larsen, P. Herring, J. H. Norton, F. Shiri, J. Vun and K. J. August, Lab Chip, 2024, 24, 4422–4439 RSC.
- W. S. R. Lago, C. Aymes-Chodur, A. P. Ahoussou and N. Yagoubi, J. Mater. Sci., 2017, 52, 6879–6904 CrossRef CAS.
- S. Venkatesan, J. Jerald, P. Asokan and R. Prabakaran, Recent Advances in Mechanical Engineering: Select Proceedings of NCAME 2019, 2020, pp. 235–245 Search PubMed.
- G. M. Whitesides, E. Ostuni, S. Takayama, X. Jiang and D. E. Ingber, Annu. Rev. Biomed. Eng., 2001, 3, 335–373 CrossRef CAS PubMed.
- Y. Xia and G. M. Whitesides, Angew. Chem., Int. Ed., 1998, 37, 550–575 CrossRef CAS PubMed.
- B. K. Gale, A. R. Jafek, C. J. Lambert, B. L. Goenner, H. Moghimifam, U. C. Nze and S. K. Kamarapu, Inventions, 2018, 3, 60 CrossRef.
- L. Dhakar, S. Gudla, X. Shan, Z. Wang, F. E. H. Tay, C.-H. Heng and C. Lee, Sci. Rep., 2016, 6, 22253 CrossRef CAS PubMed.
- M. Heckele and W. Schomburg, J. Micromech. Microeng., 2003, 14, R1 CrossRef.
- J. Giboz, T. Copponnex and P. Mélé, J. Micromech. Microeng., 2007, 17, R96 CrossRef CAS.
- L. S. A. Busa, S. Mohammadi, M. Maeki, A. Ishida, H. Tani and M. Tokeshi, Micromachines, 2016, 7, 86 CrossRef PubMed.
- H. A. Silva-Neto, I. V. Arantes, A. L. Ferreira, G. H. do Nascimento, G. N. Meloni, W. R. de Araujo, T. R. Paixão and W. K. Coltro, TrAC, Trends Anal. Chem., 2023, 158, 116893 CrossRef CAS.
- W. Zhou, M. Dou, S. S. Timilsina, F. Xu and X. Li, Lab Chip, 2021, 21, 2658–2683 RSC.
- W. Lou, L. Zhang and J. Wang, Int. Immunopharmacol., 2024, 142, 113157 CrossRef CAS PubMed.
- D. Hanahan, Cancer Discovery, 2022, 12, 31–46 CrossRef CAS PubMed.
- L. Falzone, S. Salomone and M. Libra, Front. Pharmacol., 2018, 9, 1300 CrossRef CAS PubMed.
- L. Syedmoradi, M. L. Norton and K. Omidfar, Talanta, 2021, 225, 122002 CrossRef CAS PubMed.
- Y. Zu, H. Chang and Z. Cui, Nexus, 2025 Search PubMed.
- S. Guo, H. Xie, X. Zhao, H. He, X. Feng, Y. Li, B.-F. Liu and P. Chen, Analyst, 2024, 149, 1250–1261 RSC.
- H. Zhang, Y. Tang, Y. Zhou, Y. Wang, H. Si, L. Li and B. Tang, Chem. Sci., 2024, 15, 9345–9352 RSC.
- M. Mahmoodpour, B. A. Kiasari, M. Karimi, A. Abroshan, D. Shamshirian, H. Hosseinalizadeh, A. Delavari and H. Mirzei, Front. Oncol., 2023, 13, 1131435 CrossRef CAS PubMed.
- H. Wang, R. Liu, K. Dong, L. Zhang, J. Zhang, X. Zhang, J. Zhang, X. Xiao, W. Zhang and X. Wang, Anal. Chim. Acta, 2023, 1246, 340886 CrossRef CAS PubMed.
- H. Yin, W. Xue and D. G. Anderson, Nat. Rev. Clin. Oncol., 2019, 16, 281–295 CrossRef CAS PubMed.
- J. Li, C.-H. Lau, J. Wang, W. Wu, Z. Huang, X. Chen, J. Li, Y. Huang, T. Wang and Y. Li, Anal. Methods, 2025, 17, 6304–6316 RSC.
- Y. Liu, X. Liu, D. Wei, L. Dang, X. Xu, S. Huang, L. Li, S. Wu, J. Wu and X. Liu, Nat. Commun., 2024, 15, 5014 CrossRef CAS PubMed.
- S. Yu, T. Liu, K. Zhang, Y. Wu and L. Ding, Sens. Actuators, B, 2025, 138303 CrossRef CAS.
- G. S. Omenn, Proteomics, 2006, 6, 5662–5673 CrossRef CAS PubMed.
- V. Urbiola-Salvador, D. Miroszewska, A. Jabłońska, T. Qureshi and Z. Chen, Biochim. Biophys. Acta, Mol. Cell Res., 2022, 1869, 119266 CrossRef CAS PubMed.
- Y. Zhou, L. Tao, J. Qiu, J. Xu, X. Yang, Y. Zhang, X. Tian, X. Guan, X. Cen and Y. Zhao, Signal Transduction Targeted Ther., 2024, 9, 132 CrossRef PubMed.
- S. Zafar, A. Hafeez, H. Shah, I. Mutiullah, A. Ali, K. Khan, G. Figueroa-González, O. D. Reyes-Hernández, L. I. Quintas-Granados, S. I. Peña-Corona, L. N. Kiyekbayeva, M. Butnariu, C.-E. Tota, A. Caunii, D. Büsselberg, J. Sharifi-Rad and G. Leyva-Gómez, Eur. J. Med. Res., 2025, 30, 760 CrossRef PubMed.
- A. D. Barker, M. M. Alba, P. Mallick, D. B. Agus and J. S. H. Lee, Mol. Cell. Proteomics, 2023, 22(7), 100569 CrossRef CAS PubMed.
- J. Zhang, Q. Wei, D. Dong and L. Ren, Clin. Chim. Acta, 2021, 523, 19–25 CrossRef CAS PubMed.
- Z. Wei, Y. Zhang, H. Lu, J. Ying, H. Zhao and J. Cai, Transl. Cancer Res., 2022, 11, 669–677 CrossRef CAS PubMed.
- S. Das, M. K. Dey, R. Devireddy and M. R. Gartia, Sensors, 2024, 24, 37 CrossRef CAS PubMed.
- L. Cohen and D. R. Walt, Chem. Rev., 2019, 119, 293–321 CrossRef CAS PubMed.
- A. K. Füzéry, J. Levin, M. M. Chan and D. W. Chan, Clin. Proteomics, 2013, 10, 13 CrossRef PubMed.
- S. Sundarraj, B. Sundaramahalingam, G. Rajagopal, M. Sundar and R. Thangam, in Protein Detection, ed. Y. Tutar and L. Tutar, IntechOpen, London, 2022, DOI:10.5772/intechopen.101050.
- N. C. Japp, J. J. Souchek, A. R. Sasson, M. A. Hollingsworth, S. K. Batra and W. M. Junker, J. Immunol. Res., 2021, 2021, 9942605 Search PubMed.
- S. Kakkar, P. Gupta, S. P. Singh Yadav, D. Raj, G. Singh, S. Chauhan, M. K. Mishra, E. Martín-Ortega, S. Chiussi and K. Kant, Mater. Today Bio, 2024, 28, 101188 CrossRef CAS PubMed.
- Y. Zhang, Y. Chai, Z. Hu, Z. Xu, M. Li, X. Chen, C. Yang and J. Liu, Front. Bioeng. Biotechnol., 2022, 10, 866368 CrossRef PubMed.
- U. Landegren and M. Hammond, Mol. Oncol., 2021, 15, 1715–1726 CrossRef PubMed.
- Y. Zhou, T. M. Lih, J. Pan, N. Höti, M. Dong, L. Cao, Y. Hu, K.-C. Cho, S.-Y. Chen, R. V. Eguez, E. Gabrielson, D. W. Chan, H. Zhang and Q. K. Li, J. Hematol. Oncol., 2020, 13, 170 CrossRef CAS PubMed.
- M. Naseri, Z. M. Ziora, G. P. Simon and W. Batchelor, Rev. Med. Virol., 2022, 32, e2263 CrossRef CAS.
- K. J. Land, D. I. Boeras, X.-S. Chen, A. R. Ramsay and R. W. Peeling, Nat. Microbiol., 2019, 4, 46–54 CrossRef CAS PubMed.
- R. Bhawal, A. L. Oberg, S. Zhang and M. Kohli, Cancers, 2020, 12(9), 2428 CrossRef CAS PubMed.
- J. F. Rusling, C. V. Kumar, J. S. Gutkind and V. Patel, Analyst, 2010, 135, 2496–2511 RSC.
- D. Wenk, C. Zuo, T. Kislinger and L. Sepiashvili, Clin. Proteomics, 2024, 21, 6 CrossRef CAS PubMed.
- J. Hu, S. Wang, L. Wang, F. Li, B. Pingguan-Murphy, T. J. Lu and F. Xu, Biosens. Bioelectron., 2014, 54, 585–597 CrossRef CAS PubMed.
- H. Yuan, R. Yong, W. Yuan, Q. Zhang, E. G. Lim, Y. Wang, F. Niu and P. Song, Microsyst. Nanoeng., 2025, 11, 101 CrossRef CAS PubMed.
- S. Liu, Y. Liao, R. Shu, J. Sun, D. Zhang, W. Zhang and J. Wang, ACS Nano, 2024, 18, 27167–27205 CrossRef CAS PubMed.
- E. Lamprou, P. M. Kalligosfyri and D. P. Kalogianni, Biosensors, 2025, 15, 68 CrossRef CAS PubMed.
- Y. Zhang, Q. Zhang, Z. Li, X. Zhou, C. Liu, H. Zha, X. Zhang, H. Zhao, G. Lü, J. Li and X. Li, J. Mater. Chem. B, 2025, 13, 5746–5761 RSC.
- D. M. Kinyua, D. M. Memeu, C. N. Mugo Mwenda, B. D. Ventura and R. Velotta, Sensors, 2025, 25, 5414 CrossRef CAS PubMed.
- C. Dincer, R. Bruch, A. Kling, P. S. Dittrich and G. A. Urban, Trends Biotechnol., 2017, 35, 728–742 CrossRef CAS PubMed.
- A. Sena-Torralba, R. Álvarez-Diduk, C. Parolo, A. Piper and A. Merkoçi, Chem. Rev., 2022, 122, 14881–14910 CrossRef CAS PubMed.
- Y. Liu, L. Zhan, Z. Qin, J. Sackrison and J. C. Bischof, ACS Nano, 2021, 15, 3593–3611 CrossRef CAS PubMed.
- X. Nan, L. Yang and Y. Cui, Clin. Chim. Acta, 2023, 544, 117337 CrossRef CAS PubMed.
- C. Parolo, A. de la Escosura-Muñiz and A. Merkoçi, Biosens. Bioelectron., 2013, 40, 412–416 CrossRef CAS PubMed.
- L. Ye, X. Xu, A. Qu, L. Liu, C. Xu and H. Kuang, Nano Res., 2024, 17, 5452–5460 CrossRef CAS.
- X. Lin, P. Zhou, Q. Li and Y. Pang, Anal. Chem., 2024, 96, 10686–10695 CrossRef CAS PubMed.
- A. Moyano, E. Serrano-Pertierra, J. M. Duque, V. Ramos, E. Teruel-Barandiarán, M. T. Fernández-Sánchez, M. Salvador, J. C. Martínez-García, L. Sánchez, L. García-Flórez, M. Rivas and M. d. C. Blanco-López, Sensors, 2021, 21, 3756 CrossRef CAS PubMed.
- C. Wang, R. Xiao, S. Wang, X. Yang, Z. Bai, X. Li, Z. Rong, B. Shen and S. Wang, Biosens. Bioelectron., 2019, 146, 111754 CrossRef CAS PubMed.
- W. Ren, S. I. Mohammed, S. Wereley and J. Irudayaraj, Anal. Chem., 2019, 91, 2876–2884 CrossRef CAS PubMed.
- W. Ren, I.-H. Cho, Z. Zhou and J. Irudayaraj, Chem. Commun., 2016, 52, 4930–4933 RSC.
- L. Ao, T. Liao, L. Huang, S. Lin, K. Xu, J. Ma, S. Qiu, X. Wang and Q. Zhang, Chem. Eng. J., 2022, 436, 135204 CrossRef CAS.
- H. Fu, P. Song, Q. Wu, C. Zhao, P. Pan, X. Li, N. Y. K. Li-Jessen and X. Liu, Microsyst. Nanoeng., 2019, 5, 50 CrossRef PubMed.
- A. A. Shalaby, C.-W. Tsao, A. Ishida, M. Maeki and M. Tokeshi, Sens. Actuators, B, 2023, 379, 133243 CrossRef CAS.
- N. Yonet-Tanyeri, B. Z. Ahlmark and S. R. Little, Adv. Mater. Technol., 2021, 6, 2170045 CrossRef.
- T. Lam, J. P. Devadhasan, R. Howse and J. Kim, Sci. Rep., 2017, 7, 1188 CrossRef PubMed.
- E. Noviana, T. Ozer, C. S. Carrell, J. S. Link, C. McMahon, I. Jang and C. S. Henry, Chem. Rev., 2021, 121, 11835–11885 CrossRef CAS PubMed.
- F. Bahavarnia, A. Saadati, S. Hassanpour, M. Hasanzadeh, N. Shadjou and A. Hassanzadeh, Int. J. Biol. Macromol., 2019, 138, 744–754 CrossRef CAS PubMed.
- S. S. Timilsina and X. Li, Lab Chip, 2024, 24, 4962–4973 RSC.
- Y. Wang, C. Wu, G. Yu and S. Yu, Biosens. Bioelectron., 2025, 287, 117695 CrossRef CAS PubMed.
- Y.-X.-Y. Gong, X.-F. Huang, J. Liang, J. Xie and L.-S. Qing, Anal. Bioanal. Chem., 2025, 417, 3529–3550 CrossRef CAS PubMed.
- S. Boonkaew, K. Szot-Karpińska, J. Niedziółka-Jönsson, A. de Marco and M. Jönsson-Niedziółka, ACS Sens., 2024, 9, 3066–3074 CrossRef CAS PubMed.
- M. Lundberg, A. Eriksson, B. Tran, E. Assarsson and S. Fredriksson, Nucleic Acids Res., 2011, 39, e102 CrossRef CAS PubMed.
- M. Lundberg, S. B. Thorsen, E. Assarsson, A. Villablanca, B. Tran, N. Gee, M. Knowles, B. S. Nielsen, E. González Couto, R. Martin, O. Nilsson, C. Fermer, J. Schlingemann, I. J. Christensen, H. J. Nielsen, B. Ekström, C. Andersson, M. Gustafsson, N. Brunner, J. Stenvang and S. Fredriksson, Mol. Cell. Proteomics, 2011, 10, M110.004978 CrossRef PubMed.
- R. Gupta, P. Gupta, S. Wang, A. Melnykov, Q. Jiang, A. Seth, Z. Wang, J. J. Morrissey, I. George, S. Gandra, P. Sinha, G. A. Storch, B. A. Parikh, G. M. Genin and S. Singamaneni, Nat. Biomed. Eng., 2023, 7, 1556–1570 CrossRef CAS PubMed.
- P. Barya, Y. Xiong, S. Shepherd, R. Gupta, L. D. Akin, J. Tibbs, H. Lee, S. Singamaneni and B. T. Cunningham, Small, 2023, 19, 2207239 CrossRef CAS PubMed.
- A. Seth, Y. Liu, R. Gupta, Z. Wang, E. Mittal, S. Kolla, P. Rathi, P. Gupta, B. A. Parikh, G. M. Genin, S. Gandra, G. A. Storch, J. A. Philips, I. A. George and S. Singamaneni, Nano Lett., 2024, 24, 229–237 CrossRef CAS PubMed.
- Y. S. Park, S. Choi, H. J. Jang and T. H. Yoo, Front. Bioeng. Biotechnol., 2023, 11, 1188313 CrossRef PubMed.
- F. Shao, et al., Digital Magnetic Proximity Extension RPA-CRISPR/Cas12a-Assisted Immunoassay with Attomolar Sensitivity, 2024 IEEE Sensors, Kobe, Japan, 2024, pp. 1–4, DOI:10.1109/SENSORS60989.2024.10784846.
- Y. Tang, T. Song, L. Gao, S. Yin, M. Ma, Y. Tan, L. Wu, Y. Yang, Y. Wang, T. Lin and F. Li, Nat. Commun., 2022, 13, 4667 CrossRef CAS PubMed.
- Y. Tang, Y. Lin, X. Yang, Z. Wang, X. C. Le and F. Li, Anal. Chem., 2015, 87, 8063–8066 CrossRef CAS PubMed.
- J. L. Liu, D. Zabetakis, L. C. Shriver-Lake, G. P. Anderson, E. R. Goldman and H. Bui, Adv. Sens. Res., 2024, 3, 2300111 CrossRef.
- Z. Tang, W. Zhao, Y. Deng, Y. Sun, C. Qiu, B. Wu, J. Bao, Z. Chen and L. Yu, Analyst, 2022, 147, 1709–1715 RSC.
- M. A. McSweeney, A. T. Patterson, K. Loeffler, R. Cuellar Lelo de Larrea, M. P. McNerney, R. S. Kane and M. P. Styczynski, Sci. Adv., 2025, 11, eado6280 CrossRef CAS PubMed.
- L. C. Lopes, A. Santos and P. R. Bueno, Sens. Actuat. Rep., 2022, 4, 100087 Search PubMed.
- M. Mohammadniaei, H. V. Nguyen, M. V. Tieu and M.-H. Lee, Micromachines, 2019, 10, 662 CrossRef PubMed.
- B. Chakraborty, A. Das, N. Mandal, N. Samanta, N. Das and C. R. Chaudhuri, Sci. Rep., 2021, 11, 2962 CrossRef CAS PubMed.
- W. He, L. Liu, Z. Cao, Y. Lin, Y. Tian, Q. Zhang, C. Zhou, X. Ye and T. Cui, Biosens. Bioelectron., 2023, 228, 115193 CrossRef CAS PubMed.
- J. G. Pacheco, M. S. Silva, M. Freitas, H. P. Nouws and C. Delerue-Matos, Sens. Actuators, B, 2018, 256, 905–912 CrossRef CAS.
- S. Yang, L. Zhao, X. Yang, L. Yang, H. Fa, Y. Wang, D. Huo, C. Hou, D. Zhong and M. Yang, Chem. Eng. J., 2024, 498, 155345 CrossRef CAS.
- S. S. Low, Y. Pan, D. Ji, Y. Li, Y. Lu, Y. He, Q. Chen and Q. Liu, Sens. Actuators, B, 2020, 308, 127718 CrossRef.
- J. Zhao, K. Xia, P. He, G. Wei, X. Zhou and X. Zhang, Coord. Chem. Rev., 2023, 497, 215456 CrossRef CAS.
- F. Cheng, L. Su and C. Qian, Oncotarget, 2016, 7, 48832 CrossRef PubMed.
- cobas EGFR Mutation Test v2, https://www.fda.gov/drugs/resources-information-approved-drugs/cobas-egfr-mutation-test-v2, Content current as of: 06/02/2016.
- F. Keyvani, N. Debnath, M. A. Saleh and M. Poudineh, Nanoscale, 2022, 14, 6761–6770 RSC.
- Idylla, (https://biocartis.com), https://www.biocartis.com/en-US.
- S. Singh, P. S. Podder, M. Russo, C. Henry and S. Cinti, Lab Chip, 2023, 23, 44–61 RSC.
- R. Laocharoensuk, Electroanalysis, 2016, 28, 1716–1729 CrossRef CAS.
- S. Mariani and M. Minunni, Anal. Bioanal. Chem., 2014, 406, 2303–2323 CrossRef CAS PubMed.
- C. Zong, M. Xu, L.-J. Xu, T. Wei, X. Ma, X.-S. Zheng, R. Hu and B. Ren, Chem. Rev., 2018, 118, 4946–4980 CrossRef CAS PubMed.
- S. Bhaskar, S. M. Lis S, S. Kanvah, S. Bhaktha B. N and S. S. Ramamurthy, ACS Appl. Opt. Mater., 2023, 1, 159–172 CrossRef CAS.
- S. Bhaskar, A. K. Singh, P. Das, P. Jana, S. Kanvah, S. Bhaktha B N and S. S. Ramamurthy, ACS Appl. Mater. Interfaces, 2020, 12, 34323–34336 CrossRef CAS PubMed.
- A. Rai, S. Bhaskar, K. M. Ganesh and S. S. Ramamurthy, ACS Appl. Nano Mater., 2022, 5, 12245–12264 CrossRef CAS.
- D. Thacharakkal, S. Bhaskar, T. Sharma, G. Rajaraman, S. Sathish Ramamurthy and C. Subramaniam, Chem. Eng. J., 2023, 148166, DOI:10.1016/j.cej.2023.148166.
- Y. J. Kim, W.-Y. Rho, S.-M. Park and B.-H. Jun, J. Hematol. Oncol., 2024, 17, 10 CrossRef PubMed.
- M. Zhang, X. Cui and N. Li, Mater. Today Bio, 2022, 14, 100254 CrossRef PubMed.
- H. Zong, Y. Zhang, X. Liu, Z. Xu, J. Ye, S. Lu, X. Guo, Z. Yang, X. Zhang, M. Chai, M. Fan, Y. Liao, W. Yang, Y. Wu and D. Zhang, View, 2023, 4, 20220062 CrossRef CAS.
- S. Bhaskar, Micromachines, 2023, 14, 574 CrossRef PubMed.
- S. Bhaskar, N. C. S. S. Kowshik, S. P. Chandran and S. S. Ramamurthy, Langmuir, 2020, 36, 2865–2876 CrossRef CAS PubMed.
- A. Rai, S. Bhaskar and S. S. Ramamurthy, ACS Appl. Nano Mater., 2021, 4, 5940–5953 CrossRef CAS.
- V. S. K. Cheerala, K. M. Ganesh, S. Bhaskar, S. S. Ramamurthy and S. C. Neelakantan, Langmuir, 2023, 39, 7939–7957 CrossRef CAS PubMed.
- R. Elghanian, J. J. Storhoff, R. C. Mucic, R. L. Letsinger and C. A. Mirkin, Science, 1997, 277, 1078–1081 CrossRef CAS PubMed.
- N. Xia, D. Deng, Y. Wang, C. Fang and S.-J. Li, Int. J. Nanomed., 2018, 13, 2521–2530 CrossRef CAS PubMed.
- O. H. Shayesteh and R. Ghavami, Spectrochim. Acta, Part A, 2020, 226, 117644 CrossRef CAS PubMed.
- L. Wang, Z.-J. Liu, H.-X. Cao and G.-X. Liang, Sens. Actuators, B, 2021, 337, 129813 CrossRef CAS.
- H. Duan, Y. Zhao, X. Hu, M. Liang, X. Yang, L. Yu, B. T. Oranj, V. Romanovski, P. Li and Z. Zhang, Foods, 2024, 13, 3188 CrossRef CAS PubMed.
- B. L. James, S. P. Sunny, A. E. Heidari, R. D. Ramanjinappa, T. Lam, A. V. Tran, S. Kankanala, S. Sil, V. Tiwari, S. Patrick, V. Pillai, V. Shetty, N. Hedne, D. Shah, N. Shah, Z.-P. Chen, U. Kandasarma, S. A. Raghavan, S. Gurudath, P. B. Nagaraj, P. Wilder-Smith, A. Suresh and M. A. Kuriakose, Cancers, 2021, 13, 3583 CrossRef PubMed.
- Y.-Q. Xiong, Y. Mo, Y.-Q. Wen, M.-J. Cheng, S.-T. Huo, X.-J. Chen and Q. Chen, J. Biomed. Opt., 2018, 23, 1 Search PubMed.
- S. S. Akella, J. Lee, J. R. May, C. Puyana, S. Kravets, V. Dimitropolous, M. Tsoukas, R. Manwar and K. Avanaki, Sci. Rep., 2024, 14, 8900 CrossRef CAS PubMed.
- I. Amygdalos, E. Hachgenei, L. Burkl, D. Vargas, P. Goßmann, L. I. Wolff, M. Druzenko, M. Frye, N. König, R. H. Schmitt, A. Chrysos, K. Jöchle, T. F. Ulmer, A. Lambertz, R. Knüchel-Clarke, U. P. Neumann and S. A. Lang, J. Cancer Res. Clin. Oncol., 2023, 149, 3575–3586 CrossRef PubMed.
- B. Yin, Q. Zhang, X. Xia, C. Li, W. K. H. Ho, J. Yan, Y. Huang, H. Wu, P. Wang, C. Yi, J. Hao, J. Wang, H. Chen, S. H. D. Wong and M. Yang, Theranostics, 2022, 12, 5914–5930 CrossRef CAS PubMed.
- A. Ramola, A. K. Shakya and A. Bergman, Biosensors, 2025, 15, 292 CrossRef CAS PubMed.
- M. Bally, M. Halter, J. Vörös and H. M. Grandin, Surf. Interface Anal., 2006, 38, 1442–1458 CrossRef CAS.
- T. Kumeria, M. D. Kurkuri, K. R. Diener, L. Parkinson and D. Losic, Biosens. Bioelectron., 2012, 35, 167–173 CrossRef CAS PubMed.
- X. Jiao, T. Peng, Z. Liang, Y. Hu, B. Meng, Y. Zhao, J. Xie, X. Gong, Y. Jiang, X. Fang, X. Yu and X. Dai, Int. J. Mol. Sci., 2022, 23, 9991 CrossRef CAS PubMed.
- C. Wang, F. Hou and Y. Ma, Biosens. Bioelectron., 2015, 68, 156–162 CrossRef CAS PubMed.
- K. Brosamer, K. Kourentzi, R. C. Willson and B. V. Vu, Commun. Eng., 2023, 2, 31 CrossRef CAS PubMed.
- G. G. Daaboul, P. Gagni, L. Benussi, P. Bettotti, M. Ciani, M. Cretich, D. S. Freedman, R. Ghidoni, A. Y. Ozkumur, C. Piotto, D. Prosperi, B. Santini, M. S. Ünlü and M. Chiari, Sci. Rep., 2016, 6, 37246 CrossRef CAS PubMed.
- L. Zhang, Y. Mazouzi, M. Salmain, B. Liedberg and S. Boujday, Biosens. Bioelectron., 2020, 165, 112370 CrossRef CAS PubMed.
- A. I. Barbosa, P. Gehlot, K. Sidapra, A. D. Edwards and N. M. Reis, Biosens. Bioelectron., 2015, 70, 5–14 CrossRef CAS PubMed.
- M. J. Mickert, Z. Farka, U. Kostiv, A. Hlaváček, D. Horák, P. Skládal and H. H. Gorris, Anal. Chem., 2019, 91, 9435–9441 CrossRef CAS PubMed.
- J. Xu, X. Han, W. Xu, J. Liu, A. Zhu, D. Song and F. Long, Talanta, 2023, 259, 124475 CrossRef CAS PubMed.
- W. Liu, S. Li, E. Chow, S. Bhaskar, Y. Fang and B. T. Cunningham, Adv. Mater. Technol., 2025, 10, 2401837 CrossRef CAS PubMed.
- S. Bhaskar, W. Liu, S. Shepherd, D. Wijewardena, J. Tibbs and B. T. Cunningham, Adv. Funct. Mater., 2026, 36(5), e12295 CrossRef CAS.
- L. Zhou, Y. Xiong, A. Dwivedy, M. Zheng, L. Cooper, S. Shepherd, T. Song, W. Hong, L. T. P. Le, X. Chen, S. Umrao, L. Rong, T. Wang, B. T. Cunningham and X. Wang, Sci. Robot., 2024, 9, eadi2084 CrossRef PubMed.
- N. Li, T. D. Canady, Q. Huang, X. Wang, G. A. Fried and B. T. Cunningham, Nat. Commun., 2021, 12, 1744 CrossRef CAS PubMed.
- S. Shepherd, W. Liu, S. Bhaskar and B. T. Cunningham, Anal. Bioanal. Chem., 2025, 417, 6781–6796 CrossRef CAS PubMed.
- S. Bhaskar, Y. Xiong, S. Shepherd, J. Tibbs, A. K. Bacon, W. Liu, L. D. Akin, T. Ayupova, W. Wang, H. Lee, L. Liu, A. Tan, K. Khemtonglang, X. Wang and B. T. Cunningham, in Nano-Engineering at Functional Interfaces for Multi-Disciplinary Applications, ed. S. S. Ramamurthy, S. Bhaskar and N. Reddy, Elsevier, 2025, pp. 123–156, DOI:10.1016/B978-0-443-21691-6.00007-X.
- Y. Xiong, S. Shepherd, J. Tibbs, A. Bacon, W. Liu, L. D. Akin, T. Ayupova, S. Bhaskar and B. T. Cunningham, Micromachines, 2023, 14, 668 CrossRef PubMed.
- S. Ghosh, N. Li, Y. Xiong, Y. G. Ju, M. P. Rathslag, E. G. Onal, E. Falkiewicz, M. Kohli and B. T. Cunningham, Biomed. Opt. Express, 2021, 12, 4637–4650 CrossRef PubMed.
- W. Liu, T. Ayupova, W. Wang, S. Shepherd, X. Wang, L. D. Akin, M. Kohli, U. Demirci and B. T. Cunningham, Biosens. Bioelectron., 2024, 264, 116643 CrossRef CAS PubMed.
- K. Khemtonglang, W. Liu, H. Lee, W. Wang, S. Li, Z. Y. Li, S. Shepherd, Y. Yang, D. G. Diel, Y. Fang and B. T. Cunningham, Biomed. Opt. Express, 2024, 15, 5691–5705 CrossRef CAS PubMed.
- T. D. Canady, N. Li, L. D. Smith, Y. Lu, M. Kohli, A. M. Smith and B. T. Cunningham, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 19362–19367 CrossRef CAS PubMed.
- X. Wang, S. Shepherd, N. Li, C. Che, T. Song, Y. Xiong, I. R. Palm, B. Zhao, M. Kohli, U. Demirci, Y. Lu and B. T. Cunningham, Angew. Chem., Int. Ed., 2023, 62, e202217932 CrossRef CAS PubMed.
- B. Zhao, W. Wang, N. Li, T. Garcia-Lezana, C. Che, X. Wang, B. Losic, A. Villanueva and B. T. Cunningham, Talanta, 2022, 241, 123256 CrossRef CAS PubMed.
- S. Li, W. Wang, W. Liu, C. Chen, S. Shepherd, F. Yuan, J. M. Reinhart, D. G. Diel, B. T. Cunningham and Y. Fang, IEEE Sens. J., 2025, 25(14), 26599–26607 Search PubMed.
- H. Lee, S. Li, L. Liu, W. Wang, T. Ayupova, J. Tibbs, C. Kim, Y. Fang, M. N. Do and B. T. Cunningham, Biosens. Bioelectron., 2025, 281, 117455 CrossRef CAS PubMed.
- N. Soda, K. Clack and M. J. Shiddiky, Sens. Diagn., 2022, 1, 343–375 RSC.
- B. Tomasik, M. Skrzypski, M. Bieńkowski, R. Dziadziuszko and J. Jassem, Transl. Lung Cancer Res., 2023, 12, 594 CrossRef CAS PubMed.
- L. Gorgannezhad, M. Umer, M. N. Islam, N.-T. Nguyen and M. J. Shiddiky, Lab Chip, 2018, 18, 1174–1196 RSC.
- A. J. Bronkhorst and S. Holdenrieder, Med. Genet., 2023, 35, 201–235 CAS.
- H. Lee, T.-H. Shin, J. Cheon and R. Weissleder, Chem. Rev., 2015, 115, 10690–10724 CrossRef CAS PubMed.
- K. M. Koo, N. Soda and M. J. Shiddiky, Curr. Opin. Electrochem., 2021, 25, 100645 CrossRef CAS.
- Y. Xianyu, Q. Wang and Y. Chen, TrAC, Trends Anal. Chem., 2018, 106, 213–224 CrossRef CAS.
- N. J. Greenidge, B. Calmé, A. C. Moldovan, B. Abaravicius, J. W. Martin, N. Marahrens, J. Woolfrey, B. Scaglioni, D. S. Chathuranga and S. Mitra, Sci. Robot., 2025, 10, eadq4198 CrossRef PubMed.
- J. M. Lavista Ferres, E. K. Fishman, E. Catmull, B. Vogelstein and J. T. Vogelstein, Cancer Discovery, 2025, 15, 1982–1984 CrossRef PubMed.
- C.-W. Ju, R. Lyu, H. Li, J. Wei, A. J. Parra Vitela, U. Dougherty, A. Kwesi, A. Luna, X. Zhu, S. Shen, Y. Liu, L. Wang, X. Cui, Y. Xu, B. Jiang, Y. Ji, P. Xia, D. C. West-Szymanski, C. Sun, Y. Zhong, C. Ye, A. Moran, C. Lehmann, E. Pamer, W. Zhang, M. Bissonnette, L.-S. Zhang and C. He, Nat. Biotechnol., 2025 DOI:10.1038/s41587-025-02731-8.
- M. O. Oyowvi, K. H. Babawale, A. D. Atere and B. Ben-Azu, J. Ovarian Res., 2025, 18, 96 CrossRef PubMed.
- L. G. Trubek, T. R. Oliver, C.-M. Liang, M. Mokrohisky and T. C. Campbell, Journal of Health Care Law and Policy, 2010, 14, 1–21 Search PubMed.
- L. S. Gold, G. Klein, L. Carr, L. Kessler and S. D. Sullivan, Cancer Imaging, 2012, 12, 13 CrossRef PubMed.
- K. Offit, C. M. Sharkey, D. Green, X. Wu, M. Trottier, J. G. Hamilton, M. F. Walsh, S. Dandiker, S. Belhadj and S. M. Lipkin, J. Clin. Oncol., 2023, 41, 11–21 CrossRef PubMed.
- V. Akpe and I. E. Cock, Journal of Nanotheranostics, 2025, 6(4), 29 CrossRef.
- V. Pawnikar and M. Patel, Ann. Pharm. Fr., 2025, 83(4), 637–648 CrossRef CAS PubMed.
- A. Sadana, Fractal Binding and Dissociation Kinetics for Different Biosensor Applications, Elsevier Science, Netherlands, 2007, p. 265 Search PubMed.
- Overview of Device Regulation, https://www.fda.gov/medical-devices/device-advice-comprehensive-regulatory-assistance/overview-device-regulation, Content current as of: 01/31/2024.
- Classify Your Medical Device, https://www.fda.gov/medical-devices/overview-device-regulation/classify-your-medical-device, Content current as of: 02/07/2020.
- Overview of IVD Regulation, https://www.fda.gov/medical-devices/ivd-regulatory-assistance/overview-ivd-regulation, accessed Content current as of: 12/20/2024.
- Regulatory Controls, https://www.fda.gov/medical-devices/overview-device-regulation/regulatory-controls, Content current as of: 03/27/2018.
- Premarket Notification 510(k), https://www.fda.gov/medical-devices/premarket-submissions-selecting-and-preparing-correct-submission/premarket-notification-510k, Content current as of: 08/22/2024.
- M. Aboy, C. Crespo and A. Stern, npj Digit. Med., 2024, 7, 29 CrossRef PubMed.
- J. L. Johnston, S. S. Dhruva, J. S. Ross and V. K. Rathi, JAMA Intern. Med., 2020, 180, 1701–1703 CrossRef PubMed.
- De Novo Classification Request, https://www.fda.gov/medical-devices/premarket-submissions-selecting-and-preparing-correct-submission/de-novo-classification-request, Content current as of: 12/20/2024.
- S. Al-Jawadi, P. Capasso and M. Sharma, Pharm. Dev. Technol., 2018, 23, 953–963 CrossRef CAS PubMed.
- G. A. Van Norman, JACC Basic Transl. Sci, 2016, 1, 277–287 CrossRef PubMed.
- M. L. Buenafe, FDLI Update, 2017, p. 10, https://www.fdli.org/2017/04/factors-consider-submitting-de-novo-request/.
- J. Naganuma, in Innovation in Anesthesiology, Elsevier, 2024, pp. 275–292 Search PubMed.
- E. I. Kreydin, P. Kohli and P. Gad, in Translational Urology, Elsevier, 2025, pp. 47–50 Search PubMed.
- J. J. Darrow, J. Avorn and A. S. Kesselheim, JAMA, J. Am. Med. Assoc., 2021, 326, 420–432 CrossRef PubMed.
- Premarket Approval (PMA), https://www.fda.gov/medical-devices/premarket-submissions-selecting-and-preparing-correct-submission/premarket-approval-pma, Content current as of: 05/16/2019.
- Investigational Device Exemption (IDE), https://www.fda.gov/medical-devices/premarket-submissions-selecting-and-preparing-correct-submission/investigational-device-exemption-ide, Content current as of: 08/22/2024.
- J. B. Carter and A. W. Beck, JAMA Surg., 2025, 160, 161–162 CrossRef PubMed.
- M. J. Rossi and J. Fatima, Seminars in Vascular Surgery, 2022, 35(3), 374–379 CrossRef PubMed.
- Breakthrough Devices Program, https://www.fda.gov/medical-devices/how-study-and-market-your-device/breakthrough-devices-program, Content current as of: 08/20/2025.
- R. Conroy, Cancer Network, 2023 Search PubMed , https://www.cancernetwork.com/authors/russ-conroy.
- GUIDANCE DOCUMENT: Policy for Coronavirus Disease-2019 Tests (Revised): Guidance for Developers and Food and Drug Administration Staff, https://www.fda.gov/regulatory-information/search-fda-guidance-documents/policy-coronavirus-disease-2019-tests-revised, Content current as of: 01/12/2023.
- GUIDANCE DOCUMENT: Highly Multiplexed Microbiological/Medical Countermeasure In Vitro Nucleic Acid Based Diagnostic Devices; Guidance for Industry and Food and Drug Administration Staff, https://www.fda.gov/regulatory-information/search-fda-guidance-documents/highly-multiplexed-microbiologicalmedical-countermeasure-in-vitro-nucleic-acid-based-diagnostic-devices, Content current as of: 02/08/2018.
- General Controls for Medical Devices, https://www.fda.gov/medical-devices/regulatory-controls/general-controls-medical-devices, accessed Content current as of: 12/15/2023.
- Classify Your Medical Device, https://www.fda.gov/medical-devices/overview-device-regulation/classify-your-medical-device, Content current as of: 02/07/2020.
- Quality System (QS) Regulation/Medical Device Current Good Manufacturing Practices (CGMP), https://www.fda.gov/medical-devices/postmarket-requirements-devices/quality-system-qs-regulationmedical-device-current-good-manufacturing-practices-cgmp, Content current as of: 01/31/2024.
- Medical Device Reporting (MDR): How to Report Medical Device Problems, https://www.fda.gov/medical-devices/medical-device-safety/medical-device-reporting-mdr-how-report-medical-device-problems, Content current as of: 03/27/2025.
- MDR Data Files, https://www.fda.gov/medical-devices/medical-device-reporting-mdr-how-report-medical-device-problems/mdr-data-files, Content current as of: 09/09/2025.
- C.-L. Han, C.-T. Lai, A. J. Reyes, H.-C. Yang, J.-Y. Lu, S.-R. Shih, K.-Y. Chen, A. N. Hoofnagle, S.-L. Yu and W. Bocik, Clin. Proteomics, 2024, 21, 12 CrossRef CAS PubMed.
- J. E. Szalados, in The Medical-Legal Aspects of Acute Care Medicine: A Resource for Clinicians, Administrators, and Risk Managers, Springer, 2021, pp. 463–477 Search PubMed.
- CLIA Waiver by Application, https://www.fda.gov/medical-devices/ivd-regulatory-assistance/clia-waiver-application, Content current as of: 02/29/2024.
- J. T. Jørgensen, Expert Rev. Mol. Diagn., 2024, 24, 245–247 CrossRef PubMed.
- K. C. Graden, S. A. Bennett, S. R. Delaney, H. E. Gill and M. A. Willrich, Lab. Med., 2021, 52, 315–328 CrossRef PubMed.
- Laboratory Developed Tests, https://www.fda.gov/medical-devices/in-vitro-diagnostics/laboratory-developed-tests, Content current as of: 09/19/2025.
- Reuters, US FDA tightens scrutiny of lab-developed tests with new rule, https://www.reuters.com/business/healthcare-pharmaceuticals/us-fda-publishes-final-rule-laboratory-developed-tests-2024-04-29/, Updated April 29, 2024.
- GUIDANCE DOCUMENT: Recommendations for Clinical Laboratory Improvement Amendments of 1988 (CLIA) Waiver Applications for Manufacturers of In Vitro Diagnostic Devices, https://www.fda.gov/regulatory-information/search-fda-guidance-documents/recommendations-clinical-laboratory-improvement-amendments-1988-clia-waiver-applications.
- O. J. o. t. E. Union, REGULATION (EU) 2017/746 OF THE EUROPEAN PARLIAMENT AND OF THE COUNCIL, https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:32017R0746&utm_.
- Notified bodies for medical devices, https://health.ec.europa.eu/medical-devices-topics-interest/notified-bodies-medical-devices_en?utm_.
- Pmda - Pharmaceuticals and Medical Devices Agency, https://www.pmda.go.jp/english/review-services/index.html.
- 2026 National Medical Products Administration, China, https://english.nmpa.gov.cn/Regulations.html?utm_.
- D. S. P. Agarwal and D. T. A. B. Chairman, Central drugs standard control organization, New Delhi, 2017, https://www.vmgmc.edu.in/pdfs/Guidelines_For_Good_Clinical_Practices.pdf Search PubMed.
- S. L. Tracy, Dent. Mater., 2014, 30, 1–15 CrossRef PubMed.
- A. T. Schroen, G. R. Petroni, H. Wang, M. J. Thielen, D. Sargent, J. K. Benedetti, W. M. Cronin, D. L. Wickerham, X. F. Wang and R. Gray, Clin. Trials, 2011, 8, 591–600 CrossRef PubMed.
- B. R. Johnson, A. Schwartz, J. Goldberg and A. Koerber, J. Dent. Educ., 2006, 70, 133–141 CrossRef PubMed.
- R. Byakodi, S. Byakodi, S. Hiremath, J. Byakodi, S. Adaki, K. Marathe and P. Mahind, J. Community Health, 2012, 37, 316–319 CrossRef PubMed.
- F. Farina and N. Cirillo, Cancers, 2024, 16, 4110 CrossRef CAS PubMed.
- W. C. Wood, Ann. Surg. Oncol., 2001, 8, 471 CrossRef CAS PubMed.
- J. Lau, O. Guru, S. Warnakulasuriya, R. Balasubramaniam, A. Frydrych and O. Kujan, Japanese Dental Science Review, 2024, 60, 53–72 CrossRef PubMed.
- N. Shimpi, I. Glurich, R. Rostami, H. Hegde, B. Olson and A. Acharya, J. Pers. Med., 2022, 12(4), 614 CrossRef PubMed.
- A. Romano, D. Di Stasio, M. Petruzzi, F. Fiori, C. Lajolo, A. Santarelli, A. Lucchese, R. Serpico and M. Contaldo, Cancers, 2021, 13(12), 2864 CrossRef CAS PubMed.
- K. J. Fouillen, C. Clément, E. Aous, D. Le Goff, J. Y. Le Reste and M. Guillou-Landreat, Drug Alcohol Rev., 2025, 44, 667–680 CrossRef PubMed.
- J. A. Alfaro, A. Sinha, T. Kislinger and P. C. Boutros, Nat. Methods, 2014, 11, 1107–1113 CrossRef CAS PubMed.
- B. L. Greenberg, M. Glick, J. Frantsve-Hawley and M. L. Kantor, J. Am. Dent. Assoc., JADA, 2010, 141, 52–62 CrossRef PubMed.
- N. Bostanci and G. N. Belibasakis, Clin. Oral Investig., 2023, 27, 971–978 CrossRef PubMed.
- B. Yılmaz and G. Emingil, Expert Rev. Proteomics, 2024, 21, 417–429 CrossRef PubMed.
- J. Luan, A. Seth, R. Gupta, Z. Wang, P. Rathi, S. Cao, H. Gholami Derami, R. Tang, B. Xu, S. Achilefu, J. J. Morrissey and S. Singamaneni, Nat. Biomed. Eng., 2020, 4, 518–530 CrossRef CAS PubMed.
- J. Luan, J. J. Morrissey, Z. Wang, H. G. Derami, K.-K. Liu, S. Cao, Q. Jiang, C. Wang, E. D. Kharasch, R. R. Naik and S. Singamaneni, Light: Sci. Appl., 2018, 7, 29 CrossRef PubMed.
- P. Barya, Y. Xiong, S. Shepherd, R. Gupta, L. D. Akin, J. Tibbs, H. Lee, S. Singamaneni and B. T. Cunningham, Small, 2023, 19, 2207239 CrossRef CAS PubMed.
- A. Seth, Y. Liu, R. Gupta, Z. Wang, E. Mittal, S. Kolla, P. Rathi, P. Gupta, B. A. Parikh and G. M. Genin, Nano Lett., 2023, 24, 229–237 CrossRef PubMed.
- D. Strohmaier-Nguyen, C. Horn and A. J. Baeumner, Biosens. Bioelectron., 2025, 267, 116795 CrossRef CAS PubMed.
- Y. Makutani, K. Sakai, M. Yamada, T. Wada, T. Chikugo, T. Satou, Y. Iwasa, H. Yamamoto, M. A. de Velasco, K. Nishio and J. Kawamura, Int. J. Clin. Oncol., 2022, 27, 1180–1187 CrossRef CAS PubMed.
- L. Denny, R. Saidu, R. Boa, N. Mbatani, D. Castor, J. Moodley and L. Kuhn, Sci. Rep., 2023, 13, 16182 CrossRef CAS PubMed.
- S. A. Yerim, Y. C. Khazraji, R. Bekkali, M. Bennai, N. Bahra, I. Chaoui, F. Z. Chellat, Z. Gaizi, N. Tachfouti, A. Benabdellah, B. Belkadi, M. Attaleb, M. A. Berraho and M. El Mzibri, Tumour Virus Res., 2025, 19, 200318 CrossRef CAS PubMed.
- F. Haghayegh, A. Norouziazad, E. Haghani, A. A. Feygin, R. H. Rahimi, H. A. Ghavamabadi, D. Sadighbayan, F. Madhoun, M. Papagelis, T. Felfeli and R. Salahandish, Adv. Sci., 2024, 11, 2400595 CrossRef CAS PubMed.
- G.-R. Han, A. Goncharov, M. Eryilmaz, S. Ye, B. Palanisamy, R. Ghosh, F. Lisi, E. Rogers, D. Guzman, D. Yigci, S. Tasoglu, D. Di Carlo, K. Goda, R. A. McKendry and A. Ozcan, Nat. Commun., 2025, 16, 3165 CrossRef CAS PubMed.
- P. A. Williams, S. K. Zaidi and R. Sengupta, Clin. Cancer Res., 2024, 30(19), 4296–4298 CrossRef PubMed.
- N. Bompey, InsideHealthPolicy.com's FDA Week, 2013, vol. 19, pp. 1–8.
- U. S. B. o. L. Statistics, CPI Inflation Calculator, https://www.bls.gov/data/inflation_calculator.htm, (accessed December 18, 2025).
- S. Barlas, Pharmacy and Therapeutics, 2016, 41(5), 290–295 Search PubMed.
- J. Celis and U. Ringborg, Mol. Oncol., 2024, 18, 785–792 CrossRef PubMed.
- F. Morea, A. Soraci and D. De Stefano, Open Res. Eur., 2025, 4, 268 Search PubMed.
- A. Berns, U. Ringborg, J. E. Celis, M. Heitor, N. K. Aaronson, N. Abou-Zeid, H. O. Adami, K. Apostolidis, M. Baumann, A. Bardelli, R. Bernards, Y. Brandberg, C. Caldas, F. Calvo, C. Dive, A. Eggert, A. Eggermont, C. Espina, F. Falkenburg and E. Voest, Mol. Oncol., 2020, 14, 1589–1615 CrossRef PubMed.
- S. Marti and A. M. Peneoasu, 2025 SURVEY–REPORT II, 2026, https://eua.eu/images/publications/Publication_PDFs/CDE_Doctoral_Education_Survey_Part_2_FIN_Online.pdf Search PubMed.
- N. Dass, V. Nanda and S. C. Xiao, J. Corp. Finance, 2017, 44, 353–374 CrossRef.
- J. Lerner and A. Seru, Rev. Financ. Stud., 2022, 35, 2667–2704 CrossRef.
- National Institutes of Health, NIH grants policy statement, US Department of Health and Human Services, Public Health Service, National Institutes of Health, 1998 Search PubMed.
- New Procedures and Instructions for Submitting Annual Progress Reports for Multi-Year Funded (MYF) Awards, (accessed October 6, 2010).
| This journal is © The Royal Society of Chemistry 2026 |