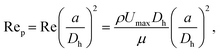Open Access Article
Open Access ArticleEstimating single-cell elastic modulus in a serial microfluidic cytometer from time-of-flight and fluorescence signals analysis
Graylen R.
Chickering
 ab,
Leroy L.
Jia
ab,
Leroy L.
Jia
 b,
Matthew
DiSalvo
b,
Matthew
DiSalvo
 b,
Megan A.
Catterton
b,
Paul N.
Patrone
b,
Eric M.
Darling
b,
Megan A.
Catterton
b,
Paul N.
Patrone
b,
Eric M.
Darling
 *a and
Gregory A.
Cooksey
*a and
Gregory A.
Cooksey
 *b
*b
aBrown University, Box G-B397, Providence, Rhode Island 02912, USA. E-mail: eric_darling@brown.edu
bNational Institute of Standards and Technology, 100 Bureau Drive, Gaithersburg, Maryland 20899, USA. E-mail: gregory.cooksey@nist.gov
First published on 21st April 2026
Abstract
Cellular state, function, and disease all contribute to whole-cell mechanical properties. Investigating these relationships is often difficult due to low measurement throughput, inability to draw one-to-one connections between mechanical and biochemical properties, and significant or unknown measurement uncertainty. To address these needs, we demonstrate that a serial microfluidic cytometer can realize high-throughput estimates of elastic modulus and size from fluorescence signals and time-of-flight (TOF) measurements of cell-like particles in flow. To analyze the resulting data, we leverage a combined spectral time-series analysis (STA) of fluorescence measurements and a mechanics-based Gaussian-process regression model. Critically, the former yields independent estimates of the particle size, whereas the latter characterizes the relationship between size, elasticity, and TOF, thereby allowing us to decouple such effects and extract estimates of elastic modulus. We calibrate the model using cell-like polyacrylamide microparticles with a range of known sizes (8.9 μm to 23 μm diameter) and stiffnesses (0.1 kPa to 9.1 kPa). The calibrated model is then applied to estimate the per-particle size and elastic modulus of live MG-63 osteosarcoma cells. Cell viability through the device was high (>90%), and the median diameter of 16.3 μm and elastic modulus of 0.9 kPa for MG-63s were consistent with light microscopy and AFM measurements. Thus, our novel device and model have the potential to expand mechanophenotyping capabilities by enabling high-throughput, single-cell measurements with uncertainty quantification. Furthermore, this emerging flow cytometry technique is directly compatible with fluorescence measurements of biochemical composition.
Introduction
Cellular elasticity, or Young's modulus, which primarily reflects changes in a cell's cytoskeletal structure, is an important metric in both physiological and pathological processes.1–16 In health, elastic modulus can be linked to cell state, activation, and differentiation. Each immune cell type has a typical elastic modulus range appropriate for its function, and changes in elastic modulus can reflect activation in response to a pathogen or therapy.17,18 Stem cells exhibit elastic modulus changes as they differentiate, enabling researchers to monitor and direct lineage-specific development by measuring elastic modulus.19,20 Additionally, elastic modulus changes are observed during mitosis, indicating a useful parameter for understanding cell proliferation and tissue regeneration.21,22 Cell elastic modulus can be a critical biomarker to understand disease mechanisms, to diagnose conditions, and to provide an accurate prognosis. This has been thoroughly documented in cancer, where reduced cell elastic modulus is correlated with increased metastatic potential.23,24 As a result, measurements of the Young's modulus can aid in the classification of tumor aggressiveness and disease prognosis. Similarly, blood diseases, like malaria or sickle cell, involve stiffening of affected red blood cells.25–29 Mechanical changes on a cellular level are also seen in neurodegenerative diseases, cardiovascular disease, and chronic inflammatory illnesses. In many instances, monitoring cell stiffening could lead to more accurate prognostics and better patient outcomes by identifying disease progression before downstream symptoms have developed. Because of these connections between cell mechanics and various states of health and disease, high-throughput platforms capable of measuring elastic modulus are advantageous in tracking disease progression, monitoring health, and screening drug candidates for novel therapies.30,31 Moreover, many of these cell types of interest may be rare (fewer than one circulating tumor cell per milliliter of blood from a patient with metastatic cancer) or part of a heterogeneous sample.32 Ultimately, robust, high-throughput tools to measure cell mechanophenotype in conjunction with validation of cell identity and function through staining with biochemical markers will advance both fundamental biology and translational research.Currently, however, existing methods for mechanophenotyping are limited. Commercial methods include 1) optical tweezers/stretchers, in which a highly focused laser beam is used to trap and deform cells with a given force, 2) micropipette aspiration, where deformability is measured by applying a suction force to the cell's membrane, and 3) atomic force microscopy (AFM), where a small cantilever applies a force to the cell and deformations are measured by a laser. AFM is the gold standard for mechanophenotyping, but there remain many shortcomings.33–35 First, cells must be adhered to a surface, which can result in cellular responses to the elastic modulus of the substrate.36–39 Results can also vary significantly based on the cantilever tip and calibration, as well as where on the cell the cantilever makes contact.40–42 Additionally, error can accumulate after calibration due to environmental fluctuations and drift or leveling, resulting in uncertainties of 20% or more.43 The processing of data can also affect results depending on the models and assumptions used.44 Finally, a critical limitation of AFM is its low throughput, at just (1 to 5) s−1 for single-cell measurements.45 Alternative commercial approaches are either similarly low throughput or are prone to destroying samples.46,47 To remedy throughput issues, many researchers have developed microfluidic deformability- or imaging-based devices for mechanophenotyping at higher throughput as compared to commercial methods. Some of these approaches include measurement of deformation in the direction of flow,48 observing changes in time of flight (TOF) or particle shape following compression of cells through an orifice,49–55 stretching particles in extensional flows perpendicular to their entry path,56 acoustics,57–59 and inertially displacing cells to their varied dynamic equilibrium points.60 These designs are capable of throughputs on the order of (102 to 103) s−1, much closer to the 104 s−1 capabilities of fluorescence-based cell sorting.55,56
To our knowledge, only two published devices exist with capabilities to match label-free deformation measurements with fluorescence biomarker abundance for cell delineation. The real-time 1D-imaging fluorescence and deformability cytometry (RT-DC)61 is capable of making single-event measurements of both cell mechanics and fluorescence markers at high throughput and has been successfully applied in a variety of clinical conditions.62–64 Additionally, fluorescence imaging deformability cytometry (FI-DC) combines fluorescence imaging with shear-flow deformation to image cell shape and size.65 As with any device, though, these approaches have limitations. The FI-DC relies on high-speed imaging for both deformation and fluorescence information. To our knowledge, the deformations have not been translated to elastic modulus. FI-DC and RT-DC are both constrained by requiring a high-viscosity buffer to measurably deform the cells in flow. Existing versions of these devices also rely on a single measurement of cells, such that there can be no quantification of uncertainty, a limitation shared by almost all existing flow-based cytometers.
We recently developed an optical cytometer capable of per-particle uncertainty quantification by measuring each particle multiple times.66,67 Measurement error in the system is further reduced by using a 3-dimensional hydrodynamic and inertial focusing strategy to uniformly place particles in the same channel location upstream of the serial interrogation regions. By pre-focusing particles to a stable dynamic equilibrium position, we found that per-particle TOF for similarly sized rigid particles could be controlled to within less than 0.2% coefficient of variation (CV; standard deviation divided by population mean). Moreover, the serial cytometer used here stores the entire time-resolved fluorescence signal, unlike standard flow cytometers that discard this information. Deeper analysis of these signal shapes further permits estimation of particle size and velocity as well as identification and deconstruction of doublet signals into the most likely singlets.68,69 This device has been well-characterized for its fluorescence measurement capabilities, and a companion manuscript details how size and velocity impact interpreted area.70 Studies and models of particles in rectangular flow channels have shown that particles inertially sort based on size and elastic modulus, with larger and more deformable particles migrating toward the center of the channel where velocity streamlines are faster.71–77
The goal of this study was to address the common shortcomings of mechanophenotyping cytometers by adapting the high-throughput serial microfluidic cytometer for simultaneous mechanophenotyping and fluorescence measurements of particles and cells. First, the serial cytometer was calibrated using well-characterized cell-like particles that mimic the physiologic size and elastic modulus of cells. Through high-precision measurements of TOFs using the serial cytometer, we were able to generate a relationship between TOF, size, and elasticity. The TOF measurements are also used to assess the accuracy of estimates of particle velocity from signal shapes. Estimates of particle size from signals analysis were also compared to distributions of particle size determined from microscopy. Finally, we demonstrated the feasibility of rapid, single-event mechanical characterization using the calibrated models to estimate size and elastic modulus of living cells. Importantly, the serial cytometer did not significantly alter cell viability and allowed us to measure only active cells using a fluorescent viability stain.
Methods
Cell-like microparticle generation and characterization
Polyacrylamide microparticles (PA MPs) with cell-like size and elasticity were made using a microfluidic droplet generator as previously described.4 The continuous phase used in the droplet generator was 2.5% by weight Hypermer SP6 (Croda Industrial; see Disclaimer) surfactant in 1-octadecene (MilliporeSigma), which was mixed for at least 20 min and 0.22 μm filtered before degassing in a syringe. The dispersed phase consisted of acrylamide (Bio-Rad) and bis-acrylamide (Bio-Rad) in different ratios targeting elastic/Young's moduli ranging from 0.1 kPa to 10 kPa, along with ammonium persulfate (APS, MilliporeSigma), lithium phenyl-2,4,6-trimethylbenzoylphosphinate (LAP, MilliporeSigma), liquid fluorescent ink (Sharpie), and de-ionized (DI) water. This solution was prepared, 0.22 μm filtered, degassed for at least 20 min, and then transferred to a syringe. After droplet generation and ultraviolet (UV) curing, MPs were washed three times with filtered 0.1% (by volume) Triton X-100 (MilliporeSigma) in DI water, followed by three washes with filtered DI water, and set aside to hydrate for at least 72 h before characterization. Particle size was determined from fluorescent microscopy images. Particles were dispensed on a glass coverslip and left to settle undisturbed before imaging. Images were then analyzed in ImageJ to calculate diameter from the cross-sectional area of the particles using the calibrated image pixel size. Elastic moduli were measured with atomic force microscopy (AFM) indentation. Particles were dispensed onto plasma-treated coverslips and allowed to settle and adhere for 15 minutes before indenting (1 to 2) μm with a 0.03 N m−1 cantilever tipped with a 5 μm diameter polystyrene (PS) bead. Elastic moduli were calculated from the resulting force vs. indentation curves using a modified Hertz model.78,79Cell culture, staining, and characterization
MG-63 osteosarcoma cells (ATCC) were grown in Minimum Essential Medium Eagle (MEM, Cellgro) supplemented with 10% (by volume) fetal bovine serum (Zen-Bio) and 1% penicillin–streptomycin (Hyclone). Living cells were stained with 0.1 μmol L−1 calcein AM (MilliporeSigma) for 15 min at room temperature prior to cytometry experiments. Cell viability before and after processing was determined by staining cells with trypan blue (Gibco) and counting on a hemocytometer before and after cytometer runs.Mechanophenotyping fluorescence cytometer (MFC) fabrication
Device fabrication followed previously described procedures.68 In brief, templates for both the top and bottom layers of device features were created using photolithography on silicon wafers as previously described. These molds were then cast in 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 (crosslinker
10 (crosslinker![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) base by mass) poly(dimethylsiloxane) (PDMS) (Sylgard 184, Dow Corning), and the top and bottom layers were cut out, punched with 1 mm inlets, and joined together following transfer of a thin spin-coated layer of PDMS crosslinker onto one layer. Features were aligned under a microscope and then cured for 1 h at 70 °C. To mitigate light leakage, black PDMS (Sylgard 170, Dow Corning) was filled into channels alongside the waveguides and microfluidic channels of the MFC. Waveguides were filled with optical adhesive (Norland 88, Edmund Optics), followed by insertion of optical fibers (0.1 numerical aperture [NA], 105 μm core size for excitation and 0.22 NA, 105 μm core size for detection) and curing in place with 1 h of UV exposure by lamp (365 nm). Finally, waveguides were cured with 4 min of UV exposure by laser (375 nm, 30 mW power) directed through each fiber. Syringes were filled with sample and sheath fluids, mounted in pumps, and coupled to the MFC with fluorinated ethylene–propylene tubing (0.762 mm inner diameter [ID], 1.58 mm outer diameter [OD]) inside of a flexible tubing sheath (1.58 mm ID, 3.175 mm OD) to reduce the effects of external vibrations on flow and performance. The tubing was fitted with 21-gauge blunt-tip needles that were inserted into the device inlets. The design iteration reported on here consisted of four sheath fluid inlets (to volumetrically partition sheath fluid “above”, “below”, “left”, and “right” of the sample core) and one sample inlet that converge into a 40 μm × 80 μm rectangular main channel. Channel dimensions were chosen to accommodate typical cell sizes of 5 to 20 μm and to reflect a 2 to 1 aspect ratio for effective inertial focusing.68,80 The main channel passes through four interrogation regions, each with waveguides to deliver laser excitation or to collect fluorescence emission and forward-scattered signals (Fig. 1A). Additionally, to facilitate imaging of particle streamlines, the device contained two expansion regions with a 4× increase in channel width, located after the first interrogation region and just prior to the device outlet. While this design incorporated two different excitation beam-shaping strategies for the laser, the study here focuses on regions 3 and 4 of the device, which feature a waveguide design for a step-function profile (Fig. 1B).
base by mass) poly(dimethylsiloxane) (PDMS) (Sylgard 184, Dow Corning), and the top and bottom layers were cut out, punched with 1 mm inlets, and joined together following transfer of a thin spin-coated layer of PDMS crosslinker onto one layer. Features were aligned under a microscope and then cured for 1 h at 70 °C. To mitigate light leakage, black PDMS (Sylgard 170, Dow Corning) was filled into channels alongside the waveguides and microfluidic channels of the MFC. Waveguides were filled with optical adhesive (Norland 88, Edmund Optics), followed by insertion of optical fibers (0.1 numerical aperture [NA], 105 μm core size for excitation and 0.22 NA, 105 μm core size for detection) and curing in place with 1 h of UV exposure by lamp (365 nm). Finally, waveguides were cured with 4 min of UV exposure by laser (375 nm, 30 mW power) directed through each fiber. Syringes were filled with sample and sheath fluids, mounted in pumps, and coupled to the MFC with fluorinated ethylene–propylene tubing (0.762 mm inner diameter [ID], 1.58 mm outer diameter [OD]) inside of a flexible tubing sheath (1.58 mm ID, 3.175 mm OD) to reduce the effects of external vibrations on flow and performance. The tubing was fitted with 21-gauge blunt-tip needles that were inserted into the device inlets. The design iteration reported on here consisted of four sheath fluid inlets (to volumetrically partition sheath fluid “above”, “below”, “left”, and “right” of the sample core) and one sample inlet that converge into a 40 μm × 80 μm rectangular main channel. Channel dimensions were chosen to accommodate typical cell sizes of 5 to 20 μm and to reflect a 2 to 1 aspect ratio for effective inertial focusing.68,80 The main channel passes through four interrogation regions, each with waveguides to deliver laser excitation or to collect fluorescence emission and forward-scattered signals (Fig. 1A). Additionally, to facilitate imaging of particle streamlines, the device contained two expansion regions with a 4× increase in channel width, located after the first interrogation region and just prior to the device outlet. While this design incorporated two different excitation beam-shaping strategies for the laser, the study here focuses on regions 3 and 4 of the device, which feature a waveguide design for a step-function profile (Fig. 1B).
Sample and sheath solution preparation
For non-biological samples, sheath fluid was prepared as 0.1% (by volume) Triton X-100 (MilliporeSigma) in DI water. Sample solutions with PS and/or PA MPs were prepared in 0.1% (by volume) Triton X-100 in DI water and adjusted to or near neutral buoyancy using OptiPrep Density Gradient Medium (MilliporeSigma). Biological samples were prepared in Hank's Buffered Saline Solution (Cytiva) with 18% (by volume) OptiPrep. All samples were spiked with 15.25 μm Dragon Green fluorescent PS microspheres (Bangs Labs) as a reference control and filtered through a 40 μm cell strainer before loading into a 1 mL glass syringe. The four-way sheath fluid was used to hydrodynamically focus the sample into a tight core. The lateral position of the core was set off-center of the channel to align with a stable inertial node of the PS MP control, determined by adjusting the ratio of left to right sheath flow rates until real-time metrics were optimized.68 For data presented here, a 3× to 25× ratio of left to right flow rates was used, with the sample flow rate at 2×, and the above and below sheath at 5×. Total flow rates of either 40 μL min−1 or 80 μL min−1 were used for mechanophenotyping experiments. The strength of inertial effects on particles can be characterized by the particle-based Reynolds numbers (Rep), which takes the ratio of inertial effects (from particle size and particle velocity relative to the fluid) to the viscous effects. Specifically, we define Rep aswhere a is the particle diameter, Dh is the hydraulic diameter, ρ is the density of the fluid and μ is the dynamic viscosity, and Umax is the maximum velocity.68 Here, Reps are 1.9 and 3.9 for the lower and higher flow rates, respectively. Samples were prepared at concentrations that targeted event rates between (1 and 30) s−1 in the MFC to reduce particle–particle interactions in flow.
Particle position imaging and analysis
For particle streamline measurements, we collected images in a region of the chip with an approximate 7.5-fold expansion in channel width, which causes total flow to slow down and particle streamlines to separate, allowing for better visualization. Particles used in this portion of the study were all approximately the same size, nominally 20 μm, and were not used for MFC data collection or calibration because they were fabricated without 488 nm excitable fluorophores. Populations of different elastic moduli were each run independently in the device. Flowing particles were illuminated under phase contrast microscopy, and videos (approximately 110 s−1 frame rate; pixel size of 0.69 μm) were recorded by a complementary metal-oxide-semiconductor camera. Images were background subtracted by a temporal median. The background was further suppressed by morphological processing with a structuring element diameter of 30 μm, facilitating the segmentation of the particles by manual intensity thresholding. Particle centroids were measured from circles fitted to each segmentation.Signal acquisition
A 488 nm, 200 mW diode laser was used to illuminate particles or cells. Fluorescence emissions were optically filtered with a (500 to 540) nm bandpass filter and then collected by photomultiplier tubes, and transmitted light was filtered and collected by silicon photodetectors. These signals were then digitized by a data acquisition card at 2 × 106 s−1 and processed by signals analysis routines, as described previously (Fig. 1C).68,69 Importantly, data containing detected events were extracted, processed, and saved in their entirety, and segments of the data stream without any triggered events were discarded. Additionally, TOF and per-particle fluorescence and scatter imprecision were calculated after serial events were matched using the previously described forward-projection time subdivision method, which predicts particles' TOF from one region to the next, matching the signal from an upstream region to the signal that falls within the particle's projected arrival timeframe at a subsequent region.68Mechanophenotyping
Following data collection, each population (PS MP control and experimental population) was gated manually by fluorescence signal height and area using a generous gating approach in which likely outliers were included, but obvious doublet populations were excluded. For the TOF approach, TOF was determined as the difference in timestamps of matched same-particle events from one serial interrogation region to the next. To control for run-to-run variation, the median TOF between regions of the control population was used as the reference TOF for that experimental run. The percent increase or decrease in TOF of each experimental particle was made relative to this reference and referred to as TOFrel. For analysis of the fluorescence signals themselves, post hoc analysis described previously was performed on the data sets to extract the instantaneous velocity and relative diameter of particles as they passed through each laser interrogation region.69 Once again, the control population was used as a reference for this analysis. While instantaneous velocity and TOF are both affected by particle mechanophenotype, TOF is an average over time, whereas instantaneous velocity is more likely to be affected by random error, like any momentary disruptions of particle equilibrium. Thus, we found that TOF was a more accurate, sensitive, and robust metric. Analysis of instantaneous velocity data can be found in the SI section. Variation between measurements of all particles in a population and variation within the repeat measurements of single particles were quantified. Relationships between measured TOF and particle size and elastic modulus were calculated through regression analyses of PA MP data. While this study used a 4-region device, we will be focusing on data collected after region 2 of the device, which includes two sets of signals analysis data (from regions 3 and 4) and two sets of TOF data (region 2 to 3 and region 3 to 4). Particles are assumed to have finished migrating to their equilibrium positions by these later regions. Additionally, this design includes a channel expansion between regions 1 and 2, which affects flight path and, in concentrated samples, increases the risk of particle interactions, therefore impacting TOF. Data from regions 1 and 2 are included in the SI section.Data analysis
Cytometer data was collected in at least three runs for each experimental MP lot, with each collection containing thousands of events per region. Events that were not matched to one particle across all four regions were discarded along with outliers, which were identified as values within the relative population data that fell outside three median absolute deviations in the TOF and velocity data. Most of these outliers were particles with improbable values, likely due to event matching errors or collisions. For each region of TOF data, outliers accounted for less than 2% of the total data. Because particles with outlier data in any given region were discarded for all 4 regions, this totaled about 7% of events. Data were then analyzed on both the population level from region to region and within the repeat measurements of individual particles. Any reported upper adjacent values are equivalent to the upper whisker of the box-and-whisker; in other words, this is the largest data point that does not exceed the value of the third quartile plus 1.5 times the inter-quartile range (IQR). Population metrics are reported as mean ± standard deviation throughout. Reported p-values are from paired t-tests.To test the hypothesis that TOF can be used to estimate elasticity, we fit the mean time-of-flight data to a model of the form
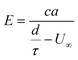 | (1) |
We anticipate that eqn (1) does not completely capture the TOF-elasticity relationship, although it should provide a semi-quantitative approximation that describes the data to within reasonable accuracy. Development of an improved model is reserved for a subsequent manuscript. To better account for any systematic model-form error associated with eqn (1), we also fit the residuals using Gaussian process regression (GPR),82 which leads to a modification of eqn (1) having the form
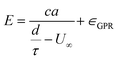 | (2) |
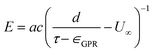 | (3) |
Results
PA MP population metrics
Microparticle lots were created across a range of sizes and elastic moduli. For MFC data collection and calibration, we focused on 4 lots of similar elastic modulus but varying sizes and 5 lots of similar size but different elastic moduli. Lots were chosen based on physiological relevance. The lots with similar elastic modulus all had AFM-characterized elastic moduli between 1 kPa and 2 kPa, with mean diameters of (8.9 ± 0.8) μm for the smallest lot up to (23.2 ± 3.0) μm for the largest lot. For lots of similar size, mean particle diameters ranged from 12 μm to 13 μm, with the softest lot at elastic modulus (0.13 ± 0.04) kPa and the stiffest lot at elastic modulus of (9.1 ± 0.3) kPa (Fig. 2). While reported CVs for elastic moduli are influenced by the inherent error and variability of AFM measurements, CVs for the 9 different lots ranged from 4% to 39% for elastic moduli and were less than or equal to 15% for diameter except for the 7 kPa lot, which had a CV of 26% for diameter. | ||
| Fig. 2 Distribution of microparticle lots as characterized by atomic force microscopy (elastic modulus) and microscopy (size). | ||
Mechanophenotyping: particle drift
As a proof of concept to observe streamline drift, we measured positions of particles of similar size (nominal 20 μm diameter) but different elastic moduli (as characterized by AFM) across the width of the channel (Fig. 3A). While the expansion poses an increased risk of particle–particle interactions at the expansion inlet, images generally captured only one particle entering and passing through the expansion region at a time with our prepared sample concentrations. Rigid polystyrene particles remained tightly aligned with the focus streamline (mean distance of (0 ± 2) μm from the focus), whereas highly deformable MPs ((0.7 ± 0.1) kPa) had migrated to streamlines closer to the channel center, (57 ± 24) μm from the focus (Fig. 3B). Of the two intermediate particle populations, the higher modulus population had a much greater variability in modulus ((5.2 ± 1.9) kPa) compared to the more elastic population ((2.5 ± 0.2) kPa). Although the drifts of these populations were not significantly different at limited image resolution and sample size, we found that the variability in population drift reflected the trend in elastic modulus variability, at (8 ± 7) μm and (9 ± 6) μm, for the 5.2 kPa and 2.5 kPa populations, respectively (Fig. 3B). All other differences in drifts were significant at a significance level of 5%. Given that the population under 1 kPa had a drift that was much greater than that of particles 2.5 kPa or higher, the sensitivity of this method may be better at lower—and physiologically more common—elastic moduli. Across the entire chip, the increased flight pathway required for particles to drift is minimal relative to the overall length of the straight, unexpanded device channel, meaning that greater drift toward higher velocity, more central streamlines should result in reduced TOF along the chip. Barring differences in AFM- and microscopy-characterized population variability, real-time inter-region velocity CV during these experiments was higher for softer particle lots, a sign that the inertial node of those particles was farther from the chosen focus streamline of the rigid PS MPs (elastic modulus on the order of 1 GPa). This observation is discussed further below.Mechanophenotyping: time-of-flight
After visually confirming that with decreasing stiffness, particles move to faster streamlines that are closer to the center of the channel, we compared time-of-flight measurements for characterized particle lots.After confirming that the instrument-induced variations in velocity variation were well controlled, the TOFs for the elastic MP formulations were analyzed. Comparing runs of the same particle lot, the largest difference in median percent decrease TOF was 0.72% (1.7% median versus 0.98% median), indicating good reproducibility.
Grouping the same particle lot runs together, we found that, as expected, the change in TOF relative to the control population was dependent on particle lot size and elastic modulus (Fig. 4A). Population TOF CVs were between 0.6% to 2.5%, and the CVs were correlated (R2 = 0.80 to 0.82) with the variation in elastic modulus within the particle lots themselves. For example, the lot with the most elastic modulus variation at (0.34 ± 0.13) kPa (39% CV) had the most TOF measurement variation (2.5% CV). Likewise, the lot with the least stiffness variation at (9.1 ± 0.3) kPa had the least TOF measurement variation (0.8% CV). In contrast, particle lot diameter CVs were not correlated to the TOF CVs (R2 = 0.04 to 0.13). The separation of TOF results for particles of similar size but different elastic modulus, or particles of similar elastic modulus but different size, was significant (p ≪ 0.0001) for almost all lots at all regions, excluding 1.8 kPa vs. 1.3 kPa lots from region 2 to 3, which were not significantly different from one another. The mean 1.3 kPa lot, though softer than the mean 1.8 kPa lot, was also smaller, with an 8.9 μm mean diameter compared to 12.2 μm diameter, respectively. Therefore, the effect of the elastic modulus difference between these two lots may have been negated by the effect of the diameter difference. This will be addressed further in the next section.
As indicated by the measurements, the precision of the TOF method correlated strongly with the CVs on single particle measurements across the three sequential measurements of TOF. For the control particles, across all runs and all regions (including region 1 to 2 TOF), the per-particle TOF CVs were under 2.5%, with the upper adjacent value being just under 1.4% CV, showing great precision. The experimental lots had more variation between single-particle measurements, although all median per-particle precisions were still 4.1% or less (Fig. 4B). This is less than half the variation we recorded in repeated AFM testing of single particles (Fig. S24). Particles that were more similar in size or elasticity to the focused control population tended to have lower per-particle CVs. For the softest particle lot with the highest variation in per-particle measurements, the upper adjacent was less than 6.4% CV per-particle. Excluding outliers, where individual TOF measurements were not physically possible and likely resulted from matching errors or particle–particle interactions and accounted for about 2% of data points per region, no particle had a CV greater than 7.9%. Out of nearly 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 particles, fewer than 1000 particles (<2%) had a per-particle CV over 5.0%.
000 particles, fewer than 1000 particles (<2%) had a per-particle CV over 5.0%.
Mechanophenotyping: spectral time-series analysis (STA)
We used the raw signal shape collected at each of the four interrogation regions to estimate instantaneous velocity and relative particle size.69,70 In the device iteration used in this study, the waveguides for focusing the laser differed between the first two and last two regions. The first two regions had a waveguide and lens design that produced a light beam with a Gaussian profile; in contrast, the last two regions had a waveguide design whose resulting beam had a step function profile. We found that the Fourier space analyses of the raw signals from the first two regions were less accurate at estimating instantaneous particle size (when compared to microscopy) and velocity (when compared to TOF). For this reason, STA data reported here will focus on regions 3 and 4. The same clusters that were used in the TOF analysis were used for each experiment in the spectral time-series analysis. As previously mentioned, TOF, being robust to random error, is more representative of equilibrium particle speed than instantaneous velocity, so we will focus on STA of size rather than velocity here.Combined methods and deconvolving size
To deconvolve the effects of size and elastic modulus, we coupled the relative size outputs from STA with TOFrel values. The physics-based GPR described above was trained on mean calibration lot values of elasticity, particle diameter, and TOFrel to yield a general relationship for individual particle elasticity as a function of particle TOF and diameter values (Fig. 5A). Here, we coupled STA of size at regions 3 and 4 with the TOFrel values from region 2 to 3 and region 3 to 4. GPR was also used to calculate confidence intervals for predicted particle elasticity (Fig. 5B). While this approach was also attempted using region 1 to 2 TOF data, we saw strong inconsistencies for lower elastic modulus data as expected from our hypothesis that the channel expansion thwarted accurate equilibrium velocity measurements or particle migrations were incomplete at that stage. In general, there is excellent agreement between cytometer measurements of size/modulus of particles and their externally measured values. In general, there is excellent agreement between cytometer measurements of size/modulus of particles and their externally measured values. The discrepancies that did exist were primarily in particles that would have the greatest deviation from the rigid PS MP control, at the lowest range of modulus and the highest range of size. For example, particles that measured larger by microscopy appeared smaller by STA (red and pink in Fig. 5B). Overall, we can estimate both size and stiffness on a per-particle basis in reasonable agreement with orthogonal measurement approaches of AFM and microscopy.Human cells
To validate the method with a living sample, live calcein-AM-stained MG-63 osteosarcoma cells were tested. Viability of the cells before and after testing was over 90%, indicating good cell health under the flow conditions and making downstream analyses/applications possible in the future. Fluorescing viable cells were gated for analysis. TOF results were normally distributed with a 2% to 3% CV for the population. The size estimation from the STA was fairly accurate, with (16.3 ± 3.0) μm from microscopy compared to an estimated (16.6 ± 7.9) μm at region 4. Given a larger sample size and greater possibility for outliers and doublets, the median ± IQR from this data was an even more accurate match at (16.3 ± 5.4) μm. The 2nd and 3rd quartiles of the estimated diameters also matched well with one standard deviation of the characterized size. Using these estimated sizes as inputs to the regressions from cell-like particle testing, the resulting elastic modulus estimations using the GPR were (1.8 ± 1.8) kPa compared to AFM characterization of the cells at (0.9 ± 0.6) kPa (Fig. 5C). However, an important aspect of this model is its quantified uncertainty. When model error is considered, the results do fall within 1 standard deviation of the AFM measurements. It is worth noting that a preliminary model using a log regression rather than the physics-based equation estimated the cell population at (0.9 ± 0.5) kPa. Although this model was replaced with the GPR model because of better alignment across the wide range of physiological values, this simple log regression may prove more accurate for mid-range physiological data until further adjustments are made to the GPR model.Discussion
In this study, we demonstrated that this serial cytometer can measure particle size and elastic modulus in a physiologically relevant range using a combination of time-of-flight and analysis of the fluorescence signal. Additionally, because each measurement is repeated, we can quantify the uncertainty of each measurement on a per-particle basis. Inertial focusing gives a stable TOF for particles of target characteristics, but differences in size and elasticity lead to drift. Thus, while we expect very low CV for the focused rigid control particle, a change in CV is expected for experimental particles. We found the TOF per-particle measurement CVs to be 8% or better for all particles, with over 98% of particles having under 5% per-particle CV. Reproducibility run-to-run was also very high when using rigid polystyrene particles as a control and reference (CV < 3%). We were able to accurately estimate particle size from the raw fluorescence signals and then use that size estimate in a model fit to calibration particles to deconvolve the effects of particle mechanics on TOF, thereby providing a measurement of particle elastic modulus. Measurements in the device aligned well with standard orthogonal approaches and were made for particles that covered a range of physiological values.We were able to apply this approach to the MG-63 cell line, measuring the fluorescence of the calcein AM viability stain along with size measurements from our device that aligned almost exactly with measurements obtained using current traditional approaches, with the added benefit of repeated measurements, increased throughput, and the ability to simultaneously measure a fluorescent biomarker. While the GPR model results for elastic modulus in this cell population were not as well matched to the AFM and microscopy estimates, our preliminary log-based model showed general agreement with standard approaches using reference particles. This information will direct future work to refine our model for live samples. Additionally, we confirmed cells remained viable, indicating limited damage to the cells during processing and enabling downstream analysis of cells in the future. Overall, we have independently determined size and stiffness with error quantification on a per-particle basis at a high degree of resolution, precision, and reproducibility, and at improved throughput as compared to AFM.
This work builds on previous reports with the serial cytometer to characterize and reduce measurement variability and to retrieve high-resolution particle information, including size and elastic modulus, from fluorescence signals.68–70 The approach taken here also combines methods supported by other microfluidics models and research. Inertial focusing of particles based on size or elastic modulus in a rectangular flow channel has been modeled and shown to be an effective sorting method in microfluidic devices.71–75 Equilibrium velocity of particles in microflows, measured using methods like spatially modulated emission,76 has been shown to relate to both particle size and elastic modulus.77 In those studies, larger and more deformable particles are consistently shown to migrate onto faster velocity streamlines toward the channel center.83 However, most mechanophenotyping devices, whether using inertial migration or direct imaging and measurement of particle deformation, fail to support fluorescence measurements of biochemical markers. As discussed earlier, the only other devices, to our knowledge, to combine size and elastic modulus in conjunction with fluorescence measurements are limited by a dependence on high-speed imaging and high-viscosity fluids and by a lack of error quantification.61,65 In future studies, we plan to expand on the fluorescence capabilities of the device to compare mechanophenotypes among cells of different types and states.
While the prospect of high-throughput, flow cytometry-based mechanophenotyping is promising, the proposed method in this study was not without limitations. This study did not examine a small fraction of particles whose events could not be matched at all regions of the chip or exhibited outlying TOF or instantaneous velocity. Such particles could be rare mismatches from the particle matching algorithm, which we have previously shown encounters ambiguity in particle matching at high velocities and particle event rates.68 Alternatively, the outlier events may be particles being dynamically influenced by inertial particle–particle and particle–fluid interactions, which would result in disruptions to the equilibration of velocity. Future studies aiming to account for these resolving particle dynamics may add insight into new optimizations to our mechanophenotyping approach.
Initially in our microfluidic system, the variables of particle size and deformability are completely convolved together in TOF. By combining known calibration values and STA size estimates, we can begin to decouple the impact of these variables on TOF to create our model. Ideally, fully validating this system requires precise, per-particle ground truth measurements for each of the particle properties. However, AFM deformability and microscopy size measurements could only be used as population-wide characterizations, which were further subject to sizable uncertainties. Discrepancies between estimated size and modulus in our model compared to standard measurements tended to occur at the lowest range of elastic modulus and the highest range of size, where our model underestimated size or overestimated stiffness. Errors in size and velocity estimates—whether due to instrumentation and design limits, like particle size relative to excitation beam width, or random error, such as tumbling/deformations altering the perceived particle size—will also lead to a less accurate modulus estimation. Additionally, our model of modulus may be skewed in the fitting process, for example, due to data from higher elastic modulus particles where modulus appeared insensitive to TOF. Cells and particles, despite being of similar size and stiffness, may behave differently in flow due to internal structure and may require separate models for the greatest accuracy. Novel methods for per-particle characterization, including ultra-high-speed imaging, may aid future refinement of the models we propose here. Furthermore, future models of modulus may be refined by weighting the model fit according to the uncertainty in the dependent factors.
The device used in this study consisted of 4 separate interrogation regions, each one collecting spectral time-series traces of both fluorescence and transmitted light, but experimental aspects of the design limited our analysis here to only some of the replicate measurements. We now know that the step function profile in the latter 2 regions of the chip is preferred to the Gaussian profile when conducting STA. Additionally, several small changes could greatly improve TOF measurement. Change in TOF could be made more sensitive with a longer distance between regions that increases the flight path of particles. TOF in all regions could be used if particles are provided a greater migration equilibration time prior to the first measurement and if channel expansions that complicate TOF with non-linear streamlines are removed.
While established serial cytometry methods provide access to time-averaged particle velocities over centimeter-long trajectories, the STA we applied in this work offers instantaneous velocities within less than a millimeter of travel. We chose to focus on time-averaged (TOF) over instantaneous velocity to measure elastic modulus because of the wider spread in instantaneous velocity data within particle lots and inaccuracies due to added uncertainty by the simultaneous estimation of particle metrics in STA. However, future work to rectify or combine both types of velocities may improve mechanophenotyping precision.
An advantage of STA was the ability to measure relative particle size for particles such as microgels with insufficient light scattering for conventional sizing via forward scattering; however, fluorescence data was required. In cells, the size estimate may be improved by including scattering data for these analyses, and size and elasticity estimates could even be made in a label-free sample. Further refinements to both the STA and the chip design, like optimizing signal windowing and reference signal selection, or rendering all four regions usable for the size estimation by altering the excitation beam profile, should improve the robustness and precision of particle size estimates.
Conclusion
The device and method described here increase the precision, confidence, and consistency of single-cell or -particle mechanics measurements while vastly improving throughput as compared to the standard methods of AFM and microscopy. We have overcome a major limitation of similar experimental devices that require high-speed cameras to decouple size and elastic modulus effects or use high-viscosity fluids and harsh flow conditions. In fact, this device uses fluids like buffered saline and requires no imaging to decouple cell size and elastic modulus. This approach does have a limit on throughput due to the possibility of particle–particle interactions in more concentrated samples, but this limit is three orders of magnitude greater than that of mechanophenotyping with AFM. Further investigations covering a broader range of physical properties will be conducted to improve the estimation of particle size from raw signals, an essential step in accurate calculations of elastic modulus. While fluorescence was only used here to generically label particles, we plan to utilize fluorescence for cell-type- and -state-specific biomarker measurements in heterogeneous clinical samples. This will allow connections to be made between cell elasticity and cell biomarkers on a single-cell basis, an as-yet challenging task, shedding light on the understanding of cell health and disease.Author contributions
Conceptualization, GRC, MD, GAC, EMD; methodology, GRC, LLJ, MD, GAC; software, MD, PNP, GRC, LLJ, MAC; validation, GRC; formal analysis, GRC, LLJ, PNP; investigation, GRC; resources, GAC, PNP, EMD; writing – original draft, GRC, LLJ; writing – review & editing, GRC, LLJ, MD, GAC, EMD, PNP; visualization, GRC; supervision, GAC, EMD; funding acquisition, GAC, EMD.Conflicts of interest
EMD is an inventor on a patent for the use of hyper-compliant polymer particles for calibration of flow-based cytometers (WO 2018/102480). MD, GAC, and PNP are coinventors on a patent on time-resolved optical profilometry (US 12541051 B2). GAC and PNP are coinventors on a patent for serial flow cytometry (US 12429412 B2).Data availability
Models and algorithms used here are described in supplementary information (SI) or Patrone et al., 2023 (DOI: https://doi.org/10.1371/journal.pone.0295502). Data table provided in SI.Supplementary information: the SI includes figures showing modeling results, TOF measurements for various particle types, estimates for instantaneous velocities and particle diameter from STA, comparison of these estimates to elasticity measurements from AFM, and technical replicates demonstrating AFM measurement variability. See DOI: https://doi.org/10.1039/d5lc00930h.
Disclaimer
Certain equipment, instruments, software, or materials are identified in this paper in order to specify the experimental procedure adequately. Such identification is not intended to imply recommendation or endorsement of any product or service by NIST, nor is it intended to imply that the materials or equipment identified are necessarily the best available for the purpose.Acknowledgements
Financial support for this project was provided in part by the National Science Foundation (CMMI 2054193, EMD). GRC acknowledges support of a Graduate Student Measurement Science and Engineering (GMSE) Fellowship from NIST. Research performed in part at the NIST Center for Nanoscale Science and Technology (CNST).References
- C. Zhu, G. Bao and N. Wang, Cell mechanics: mechanical response, cell adhesion, and molecular deformation, Annu. Rev. Biomed. Eng., 2000, 2, 189–226, DOI:10.1146/annurev.bioeng.2.1.189 , PubMed PMID: 11701511.
- M. E. Dempsey, G. R. Chickering, R. D. Gonzalez-Cruz, V. C. Fonseca and E. M. Darling, Discovery of surface biomarkers for cell mechanophenotype via an intracellular protein-based enrichment strategy, Cell. Mol. Life Sci., 2022, 79(6), 320, DOI:10.1007/s00018-022-04351-w , Epub 20220527, PubMed PMID: 35622146; PMCID: PMC9239330.
- R. A. Gutierrez, W. Fang, H. Kesari and E. M. Darling, Force sensors for measuring microenvironmental forces during mesenchymal condensation, Biomaterials, 2021, 270, 120684, DOI:10.1016/j.biomaterials.2021.120684 , Epub 20210120, PubMed PMID: 33535143; PMCID: PMC7906959.
- R. Dubay, J. Fiering and E. M. Darling, Effect of elastic modulus on inertial displacement of cell-like particles in microchannels, Biomicrofluidics, 2020, 14(4), 044110, DOI:10.1063/5.0017770 , Epub 20200803, PubMed PMID: 32774585; PMCID: PMC7402708.
- R. D. Gonzalez-Cruz, K. N. Dahl and E. M. Darling, The Emerging Role of Lamin C as an Important LMNA Isoform in Mechanophenotype, Front. Cell Dev. Biol., 2018, 6, 151, DOI:10.3389/fcell.2018.00151 , Epub 20181120, PubMed PMID: 30450357; PMCID: PMC6224339.
- N. R. Labriola, J. S. Sadick, J. R. Morgan, E. Mathiowitz and E. M. Darling, Cell Mimicking Microparticles Influence the Organization, Growth, and Mechanophenotype of Stem Cell Spheroids, Ann. Biomed. Eng., 2018, 46(8), 1146–1159, DOI:10.1007/s10439-018-2028-4 , Epub 20180418, PubMed PMID: 29671154; PMCID: PMC6039261.
- R. D. Gonzalez-Cruz, J. S. Sadick, V. C. Fonseca and E. M. Darling, Nuclear Lamin Protein C Is Linked to Lineage-Specific, Whole-Cell Mechanical Properties, Cell. Mol. Bioeng., 2018, 11(2), 131–142 CrossRef CAS PubMed , Epub 20180515, PubMed PMID: 29755599; PMCID: PMC5943047.
- M. K. Shah, I. H. Garcia-Pak and E. M. Darling, Influence of Inherent Mechanophenotype on Competitive Cellular Adherence, Ann. Biomed. Eng., 2017, 45(8), 2036–2047, DOI:10.1007/s10439-017-1841-5 , Epub 20170428, PubMed PMID: 28447179; PMCID: PMC5529242.
- Y. Xia, E. M. Darling and W. Herzog, Functional properties of chondrocytes and articular cartilage using optical imaging to scanning probe microscopy, J. Orthop. Res., 2018, 36(2), 620–631, DOI:10.1002/jor.23757 , Epub 20171122, PubMed PMID: 28975657; PMCID: PMC5839958.
- N. R. Labriola and E. M. Darling, Temporal heterogeneity in single-cell gene expression and mechanical properties during adipogenic differentiation, J. Biomech., 2015, 48(6), 1058–1066, DOI:10.1016/j.jbiomech.2015.01.033 , Epub 20150217, PubMed PMID: 25683518; PMCID: PMC4380682.
- E. M. Darling, M. Topel, S. Zauscher, T. P. Vail and F. Guilak, Viscoelastic properties of human mesenchymally-derived stem cells and primary osteoblasts, chondrocytes, and adipocytes, J. Biomech., 2008, 41(2), 454–464, DOI:10.1016/j.jbiomech.2007.06.019 , Epub 20070906, PubMed PMID: 17825308; PMCID: PMC2897251.
- E. K. Yim, E. M. Darling, K. Kulangara, F. Guilak and K. W. Leong, Nanotopography-induced changes in focal adhesions, cytoskeletal organization, and mechanical properties of human mesenchymal stem cells, Biomaterials, 2010, 31(6), 1299–1306, DOI:10.1016/j.biomaterials.2009.10.037 , Epub 20091103, PubMed PMID: 19879643; PMCID: PMC2813896.
- E. M. Darling, Force scanning: a rapid, high-resolution approach for spatial mechanical property mapping, Nanotechnology, 2011, 22(17), 175707, DOI:10.1088/0957-4484/22/17/175707 , Epub 20110318, PubMed PMID: 21411911; PMCID: PMC3150532.
- M. Kanthilal and E. M. Darling, Characterization of mechanical and regenerative properties of human, adipose stromal cells, Cell. Mol. Bioeng., 2014, 7(4), 585–597, DOI:10.1007/s12195-014-0350-y , PubMed PMID: 25484990; PMCID: PMC4255916.
- R. D. Gonzalez-Cruz and E. M. Darling, Adipose-derived stem cell fate is predicted by cellular mechanical properties, Adipocyte, 2013, 2(2), 87–91, DOI:10.4161/adip.23015 , Epub 20130628, PubMed PMID: 23805404; PMCID: PMC3661107.
- R. D. Gonzalez-Cruz, V. C. Fonseca and E. M. Darling, Cellular mechanical properties reflect the differentiation potential of adipose-derived mesenchymal stem cells, Proc. Natl. Acad. Sci. U. S. A., 2012, 109(24), E1523–E1529, DOI:10.1073/pnas.1120349109 , Epub 20120523, PubMed PMID: 22615348; PMCID: PMC3386052.
- N. Bufi, M. Saitakis, S. Dogniaux, O. Buschinger, A. Bohineust, A. Richert, M. Maurin, C. Hivroz and A. Asnacios, Human Primary Immune Cells Exhibit Distinct Mechanical Properties that Are Modified by Inflammation, Biophys. J., 2015, 108(9), 2181–2190, DOI:10.1016/j.bpj.2015.03.047 , PubMed PMID: 25954876; PMCID: PMC4423053.
- P. Jung, X. Zhou, S. Iden, M. Bischoff and B. Qu, T cell stiffness is enhanced upon formation of immunological synapse, eLife, 2021, 10, e66643, DOI:10.7554/eLife.66643 , Epub 20210727, PubMed PMID: 34313220; PMCID: PMC8360652.
- Y. Morita, T. Mukai, Y. Ju and S. Watanabe, Evaluation of stem cell-to-tenocyte differentiation by atomic force microscopy to measure cellular elastic moduli, Cell Biochem. Biophys., 2013, 66(1), 73–80, DOI:10.1007/s12013-012-9455-x , PubMed PMID: 23090789.
- G. Yourek, M. A. Hussain and J. J. Mao, Cytoskeletal changes of mesenchymal stem cells during differentiation, ASAIO J., 2007, 53(2), 219–228, DOI:10.1097/MAT.0b013e31802deb2d , PubMed PMID: 17413564; PMCID: PMC4035052.
- P. Chugh, A. G. Clark, M. B. Smith, D. A. D. Cassani, K. Dierkes, A. Ragab, P. P. Roux, G. Charras, G. Salbreux and E. K. Paluch, Actin cortex architecture regulates cell surface tension, Nat. Cell Biol., 2017, 19(6), 689–697, DOI:10.1038/ncb3525 , Epub 20170522, PubMed PMID: 28530659; PMCID: PMC5536221.
- Y. Fujii, W. C. Koizumi, T. Imai, M. Yokobori, T. Matsuo, K. Oka, K. Hotta and T. Okajima, Spatiotemporal dynamics of single cell stiffness in the early developing ascidian chordate embryo, Commun. Biol., 2021, 4(1), 341, DOI:10.1038/s42003-021-01869-w , Epub 20210316, PubMed PMID: 33727646; PMCID: PMC7966737.
- V. Swaminathan, K. Mythreye, E. T. O'Brien, A. Berchuck, G. C. Blobe and R. Superfine, Mechanical stiffness grades metastatic potential in patient tumor cells and in cancer cell lines, Cancer Res., 2011, 71(15), 5075–5080, DOI:10.1158/0008-5472.CAN-11-0247 , Epub 20110603, PubMed PMID: 21642375; PMCID: PMC3220953.
- M. Lekka, Discrimination Between Normal and Cancerous Cells Using AFM, BioNanoScience, 2016, 6, 65–80, DOI:10.1007/s12668-016-0191-3 , Epub 20160130, PubMed PMID: 27014560; PMCID: PMC4778153.
- B. E. Barber, B. Russell, M. J. Grigg, R. Zhang, T. William, A. Amir, Y. L. Lau, M. D. Chatfield, A. M. Dondorp, N. M. Anstey and T. W. Yeo, Reduced red blood cell deformability in Plasmodium knowlesi malaria, Blood Adv., 2018, 2(4), 433–443, DOI:10.1182/bloodadvances.2017013730 , PubMed PMID: 29487058; PMCID: PMC5858481.
- E. Evans, N. Mohandas and A. Leung, Static and dynamic rigidities of normal and sickle erythrocytes. Major influence of cell hemoglobin concentration, J. Clin. Invest., 1984, 73(2), 477–488, DOI:10.1172/JCI111234 , PubMed PMID: 6699172; PMCID: PMC425039.
- G. B. Nash, C. S. Johnson and H. J. Meiselman, Mechanical properties of oxygenated red blood cells in sickle cell (HbSS) disease, Blood, 1984, 63(1), 73–82 CrossRef CAS PubMed , PubMed PMID: 6689955.
- J. T. T. Rathnam, M. J. Grigg, A. M. Dondorp, T. William, M. Rajasekhar, G. Rajahram, J. A. Simpson, B. E. Barber and N. M. Anstey, Reduced red blood cell deformability in vivax malaria, J. Infect. Dis., 2024, 231(3), e566–e569, DOI:10.1093/infdis/jiae490 , Epub 20241007, PubMed PMID: 39374370.
- E. Muhammed, J. Cooper, D. Devito, R. Mushi, M. Del Pilar Aguinaga, D. Erenso and H. Crogman, Elastic property of sickle cell anemia and sickle cell trait red blood cells, J. Biomed. Opt., 2021, 26(9), 096502, DOI:10.1117/1.JBO.26.9.096502 , PubMed PMID: 34590447; PMCID: PMC8479689.
- M. Kozminsky and L. L. Sohn, The promise of single-cell mechanophenotyping for clinical applications, Biomicrofluidics, 2020, 14(3), 031301, DOI:10.1063/5.0010800 , Epub 20200609, PubMed PMID: 32566069; PMCID: PMC7286698.
- E. M. Darling and D. Di Carlo, High-Throughput Assessment of Cellular Mechanical Properties, Annu. Rev. Biomed. Eng., 2015, 17, 35–62, DOI:10.1146/annurev-bioeng-071114-040545 , PubMed PMID: 26194428.
- M. J. Alvarez Cubero, J. A. Lorente, I. Robles-Fernandez, A. Rodriguez-Martinez, J. L. Puche and M. J. Serrano, Circulating Tumor Cells: Markers and Methodologies for Enrichment and Detection, Methods Mol. Biol., 2017, 1634, 283–303, DOI:10.1007/978-1-4939-7144-2_24 , PubMed PMID: 28819860.
- D. Delgado, S. Desroches, G. Kang, Y. Aldabii and A. R. Harris, An open-source combined atomic force microscope and optical microscope for mechanobiology studies, Heliyon, 2024, 10(19), e38214, DOI:10.1016/j.heliyon.2024.e38214 , Epub 20240920, PubMed PMID: 39386786; PMCID: PMC11462344.
- K. Molnar and J.-B. Manneville, Emerging mechanobiology techniques to probe intracellular mechanics, npj Biol. Phys. Mech., 2025, 2(1), 12, DOI:10.1038/s44341-025-00016-4.
- C. M. White, M. A. Haidekker and W. S. Kisaalita, Ratiometric Nanoviscometers: Applications for Measuring Cellular Physical Properties in 3D Cultures, SLAS Technol., 2020, 25(3), 234–246, DOI:10.1177/2472630319901262 , Epub 20200130, PubMed PMID: 31997709.
- R. G. Wells, The role of matrix stiffness in regulating cell behavior, Hepatology, 2008, 47(4), 1394–1400, DOI:10.1002/hep.22193 , PubMed PMID: 18307210.
- C. Zhang, Y. Tan, J. Feng, C. Huang, B. Liu, Z. Fan, B. Xu and T. Lu, Exploration of the Effects of Substrate Stiffness on Biological Responses of Neural Cells and Their Mechanisms, ACS Omega, 2020, 5(48), 31115–31125, DOI:10.1021/acsomega.0c04279 , Epub 20201129, PubMed PMID: 33324820; PMCID: PMC7726759.
- B. Yi, Q. Xu and W. Liu, An overview of substrate stiffness guided cellular response and its applications in tissue regeneration, Bioact. Mater., 2022, 15, 82–102, DOI:10.1016/j.bioactmat.2021.12.005 , Epub 20211225, PubMed PMID: 35386347; PMCID: PMC8940767.
- S. Y. Tee, J. Fu, C. S. Chen and P. A. Janmey, Cell shape and substrate rigidity both regulate cell stiffness, Biophys. J., 2011, 100(5), L25–L27, DOI:10.1016/j.bpj.2010.12.3744 , PubMed PMID: 21354386; PMCID: PMC3043219.
- S. V. Kontomaris, A. Stylianou, G. Chliveros and A. Malamou, Overcoming Challenges and Limitations Regarding the Atomic Force Microscopy Imaging and Mechanical Characterization of Nanofibers, Fibers, 2023, 11(10), 83, DOI:10.3390/fib11100083.
- C. Pimenta-Lopes, C. Suay-Corredera, D. Velázquez-Carreras, D. Sánchez-Ortiz and J. Alegre-Cebollada, Concurrent atomic force spectroscopy, Commun. Phys., 2019, 2, 1, DOI:10.1038/s42005-019-0192-y.
- A. J. Whitehead, N. J. Kirkland and A. J. Engler, Atomic Force Microscopy for Live-Cell and Hydrogel Measurement, Methods Mol. Biol., 2021, 2299, 217–226, DOI:10.1007/978-1-0716-1382-5_16 , PubMed PMID: 34028746; PMCID: PMC9074798.
- F. Marinello, S. Carmignato, A. Voltan, E. Savio and L. De Chiffre, Error Sources in Atomic Force Microscopy for Dimensional Measurements: Taxonomy and Modeling, J. Manuf. Sci. Eng., 2010, 132(3), 030903, DOI:10.1115/1.4001242.
- B. Feng, Y. Li, R. Li and H. Li, Error analysis in calculation and interpretation of AFM tip-surface interaction forces, Adv. Colloid Interface Sci., 2022, 306, 102710, DOI:10.1016/j.cis.2022.102710 , Epub 20220602, PubMed PMID: 35691096.
- Z. Wang, L. Liu, Y. Wang, N. Xi, Z. Dong, M. Li and S. Yuan, A fully automated system for measuring cellular mechanical properties, J. Lab. Autom., 2012, 17(6), 443–448, DOI:10.1177/2211068212460236 , Epub 20120926, PubMed PMID: 23015516.
- K. C. Neuman and A. Nagy, Single-molecule force spectroscopy: optical tweezers, magnetic tweezers and atomic force microscopy, Nat. Methods, 2008, 5(6), 491–505, DOI:10.1038/nmeth.1218 , PubMed PMID: 18511917; PMCID: PMC3397402.
- R. M. Hochmuth, Micropipette aspiration of living cells, J. Biomech., 2000, 33(1), 15–22, DOI:10.1016/s0021-9290(99)00175-x , PubMed PMID: 10609514.
- O. Otto, P. Rosendahl, A. Mietke, S. Golfier, C. Herold, D. Klaue, S. Girardo, S. Pagliara, A. Ekpenyong, A. Jacobi, M. Wobus, N. Topfner, U. F. Keyser, J. Mansfeld, E. Fischer-Friedrich and J. Guck, Real-time deformability cytometry: on-the-fly cell mechanical phenotyping, Nat. Methods, 2015, 12(3), 199–202, DOI:10.1038/nmeth.3281 , Epub 20150202, PubMed PMID: 25643151.
- S. Byun, S. Son, D. Amodei, N. Cermak, J. Shaw, J. H. Kang, V. C. Hecht, M. M. Winslow, T. Jacks, P. Mallick and S. R. Manalis, Characterizing deformability and surface friction of cancer cells, Proc. Natl. Acad. Sci. U. S. A., 2013, 110(19), 7580–7585, DOI:10.1073/pnas.1218806110 , Epub 20130422, PubMed PMID: 23610435; PMCID: PMC3651488.
- M. J. Rosenbluth, W. A. Lam and D. A. Fletcher, Analyzing cell mechanics in hematologic diseases with microfluidic biophysical flow cytometry, Lab Chip, 2008, 8(7), 1062–1070, 10.1039/b802931h , Epub 20080605, PubMed PMID: 18584080; PMCID: PMC7931849.
- J. R. Lange, J. Steinwachs, T. Kolb, L. A. Lautscham, I. Harder, G. Whyte and B. Fabry, Microconstriction arrays for high-throughput quantitative measurements of cell mechanical properties, Biophys. J., 2015, 109(1), 26–34, DOI:10.1016/j.bpj.2015.05.029 , PubMed PMID: 26153699; PMCID: PMC4571004.
- K. D. Nyberg, K. H. Hu, S. H. Kleinman, D. B. Khismatullin, M. J. Butte and A. C. Rowat, Quantitative Deformability Cytometry: Rapid, Calibrated Measurements of Cell Mechanical Properties, Biophys. J., 2017, 113(7), 1574–1584, DOI:10.1016/j.bpj.2017.06.073 , PubMed PMID: 28978449; PMCID: PMC5627151.
- A. Adamo, A. Sharei, L. Adamo, B. Lee, S. Mao and K. F. Jensen, Microfluidics-based assessment of cell deformability, Anal. Chem., 2012, 84(15), 6438–6443, DOI:10.1021/ac300264v , Epub 20120710, PubMed PMID: 22746217; PMCID: PMC3418411.
- A. C. Rowat, D. E. Jaalouk, M. Zwerger, W. L. Ung, I. A. Eydelnant, D. E. Olins, A. L. Olins, H. Herrmann, D. A. Weitz and J. Lammerding, Nuclear Envelope Composition Determines the Ability of Neutrophil-type Cells to Passage through Micron-scale Constrictions, J. Biol. Chem., 2013, 288(12), 8610–8618, DOI:10.1074/jbc.M112.441535 , PubMed PMID: WOS:000316564500061.
- X. Yang, Z. Chen, J. Miao, L. Cui and W. Guan, High-throughput and label-free parasitemia quantification and stage differentiation for malaria-infected red blood cells, Biosens. Bioelectron., 2017, 98, 408–414, DOI:10.1016/j.bios.2017.07.019 , Epub 20170708, PubMed PMID: 28711027; PMCID: PMC5558593.
- D. R. Gossett, H. T. Tse, S. A. Lee, Y. Ying, A. G. Lindgren, O. O. Yang, J. Rao, A. T. Clark and D. Di Carlo, Hydrodynamic stretching of single cells for large population mechanical phenotyping, Proc. Natl. Acad. Sci. U. S. A., 2012, 109(20), 7630–7635, DOI:10.1073/pnas.1200107109 , Epub 20120430, PubMed PMID: 22547795; PMCID: PMC3356639.
- P. Augustsson, J. T. Karlsen, H. W. Su, H. Bruus and J. Voldman, Iso-acoustic focusing of cells for size-insensitive acousto-mechanical phenotyping, Nat. Commun., 2016, 7, 11556, DOI:10.1038/ncomms11556 , Epub 20160516, PubMed PMID: 27180912; PMCID: PMC4873643.
- H. Wang, Z. Liu, D. M. Shin, Z. G. Chen, Y. Cho, Y. J. Kim and A. Han, A continuous-flow acoustofluidic cytometer for single-cell mechanotyping, Lab Chip, 2019, 19(3), 387–393, 10.1039/c8lc00711j , PubMed PMID: 30648172.
- J. Y. Hwang, J. Kim, J. M. Park, C. Lee, H. Jung, J. Lee and K. K. Shung, Cell Deformation by Single-beam Acoustic Trapping: A Promising Tool for Measurements of Cell Mechanics, Sci. Rep., 2016, 6, 27238, DOI:10.1038/srep27238 , Epub 20160608, PubMed PMID: 27273365; PMCID: PMC4897707.
- S. C. Hur, N. K. Henderson-MacLennan, E. R. B. McCabe and D. Di Carlo, Deformability-based cell classification and enrichment using inertial microfluidics, Lab Chip, 2011, 11(5), 912–920, 10.1039/c0lc00595a , PubMed PMID: WOS:000287409600019.
- P. Rosendahl, K. Plak, A. Jacobi, M. Kraeter, N. Toepfner, O. Otto, C. Herold, M. Winzi, M. Herbig, Y. Ge, S. Girardo, K. Wagner, B. Baum and J. Guck, Real-time fluorescence and deformability cytometry, Nat. Methods, 2018, 15(5), 355–358, DOI:10.1038/nmeth.4639 , Epub 20180402, PubMed PMID: 29608556.
- K. R. Bashant, A. Vassallo, C. Herold, R. Berner, L. Menschner, J. Subburayalu, M. J. Kaplan, C. Summers, J. Guck, E. R. Chilvers and N. Toepfner, Real-time deformability cytometry reveals sequential contraction and expansion during neutrophil priming, J. Leukocyte Biol., 2019, 105(6), 1143–1153, DOI:10.1002/JLB.MA0718-295RR , Epub 20190305, PubMed PMID: 30835869; PMCID: PMC7587463.
- M. Herbig, A. Jacobi, M. Wobus, H. Weidner, A. Mies, M. Krater, O. Otto, C. Thiede, M. T. Weickert, K. S. Gotze, M. Rauner, L. C. Hofbauer, M. Bornhauser, J. Guck, M. Ader, U. Platzbecker and E. Balaian, Machine learning assisted real-time deformability cytometry of CD34+ cells allows to identify patients with myelodysplastic syndromes, Sci. Rep., 2022, 12(1), 870, DOI:10.1038/s41598-022-04939-z , Epub 20220118, PubMed PMID: 35042906; PMCID: PMC8766444.
- A. Walther, A. Mackens-Kiani, J. Eder, M. Herbig, C. Herold, C. Kirschbaum, J. Guck, L. D. Wittwer, K. Beesdo-Baum and M. Krater, Depressive disorders are associated with increased peripheral blood cell deformability: a cross-sectional case-control study (Mood-Morph), Transl. Psychiatry, 2022, 12(1), 150, DOI:10.1038/s41398-022-01911-3 , Epub 20220408, PubMed PMID: 35396373; PMCID: PMC8990596.
- H. E. Muñoz, J. Lin, B. G. Yeh, T. Biswas and D. Di Carlo, Fluorescence imaging deformability cytometry: integrating nuclear structure with mechanical phenotyping, Med-X, 2023, 1, 10, DOI:10.1007/s44258-023-00008-w.
- M. E. Piyasena and S. W. Graves, The intersection of flow cytometry with microfluidics and microfabrication, Lab Chip, 2014, 14(6), 1044–1059, 10.1039/c3lc51152a , PubMed PMID: 24488050; PMCID: PMC4077616.
- H. Daguerre, M. Solsona, J. Cottet, M. Gauthier, P. Renaud and A. Bolopion, Positional dependence of particles and cells in microfluidic electrical impedance flow cytometry: origin, challenges and opportunities, Lab Chip, 2020, 20(20), 3665–3689, 10.1039/d0lc00616e , Epub 20200911, PubMed PMID: 32914827.
- M. DiSalvo, P. N. Patrone, A. J. Kearsley and G. A. Cooksey, Serial flow cytometry in an inertial focusing optofluidic microchip for direct assessment of measurement variations, Lab Chip, 2022, 22, 3217–3228, 10.1039/d1lc01169c , Epub 20220720, PubMed PMID: 35856829.
- P. N. Patrone, M. DiSalvo, A. J. Kearsley, G. B. McFadden and G. A. Cooksey, Reproducibility in cytometry: Signals analysis and its connection to uncertainty quantification, PLoS One, 2023, 18(12), e0295502, DOI:10.1371/journal.pone.0295502 , Epub 20231222, PubMed PMID: 38134031; PMCID: PMC10745152.
- M. A. Catterton, M. DiSalvo, P. N. Patrone and G. A. Cooksey, Estimating Particle Size and Velocity from Fluorescence Pulses: A Practical Validation Study of Flow Cytometry Signals Analysis, PLoS One, 2026, accepted Search PubMed.
- S. C. Hur, N. K. Henderson-MacLennan, E. R. McCabe and D. Di Carlo, Deformability-based cell classification and enrichment using inertial microfluidics, Lab Chip, 2011, 11(5), 912–920, 10.1039/c0lc00595a , Epub 20110127, PubMed PMID: 21271000.
- C. Liu, G. Hu, X. Jiang and J. Sun, Inertial focusing of spherical particles in rectangular microchannels over a wide range of Reynolds numbers, Lab Chip, 2015, 15(4), 1168–1177, 10.1039/c4lc01216j , PubMed PMID: 25563524.
- Y. Zhou, Z. Ma and Y. Ai, Sheathless inertial cell focusing and sorting with serial reverse wavy channel structures, Microsyst. Nanoeng., 2018, 4, 5, DOI:10.1038/s41378-018-0005-6 , Epub 20180507, PubMed PMID: 31057895; PMCID: PMC6220157.
- S. Tanriverdi, J. Cruz, S. Habibi, K. Amini, M. Costa, F. Lundell, G. Martensson, L. Brandt, O. Tammisola and A. Russom, Elasto-inertial focusing and particle migration in high aspect ratio microchannels for high-throughput separation, Microsyst. Nanoeng., 2024, 10, 87, DOI:10.1038/s41378-024-00724-2 , Epub 20240625, PubMed PMID: 38919163; PMCID: PMC11196675.
- K. Hood, S. Kahkeshani, D. Di Carlo and M. Roper, Direct measurement of particle inertial migration in rectangular microchannels, Lab Chip, 2016, 16(15), 2840–2850, 10.1039/c6lc00314a , Epub 20160628, PubMed PMID: 27349748.
- N. Heinß, S. Alebrand, J. Wittek and M. Baßler, Equilibrium transport velocity of deformable cells and rigid spheres in micro-channels under laminar flow conditions, Microfluid. Nanofluid., 2019, 24, 3, DOI:10.1007/s10404-019-2305-z.
- L. Schott, C. Sommer, J. Wittek, K. Myagmar, T. Walther and M. Baßler, Cell Size Discrimination Based on the Measurement of the Equilibrium Velocity in Rectangular Microchannels, Micromachines, 2015, 6(5), 634–647, DOI:10.3390/mi6050634.
- N. R. Labriola, E. Mathiowitz and E. M. Darling, Fabricating polyacrylamide microbeads by inverse emulsification to mimic the size and elasticity of living cells, Biomater. Sci., 2016, 5(1), 41–45, 10.1039/c6bm00692b , PubMed PMID: 27935612; PMCID: PMC5201106.
- E. M. Darling, S. Zauscher and F. Guilak, Viscoelastic properties of zonal articular chondrocytes measured by atomic force microscopy, Osteoarthr. Cartil., 2006, 14(6), 571–579, DOI:10.1016/j.joca.2005.12.003 , Epub 2006/02/16, PubMed PMID: 16478668.
- D. Di Carlo, Inertial microfluidics, Lab Chip, 2009, 9(21), 3038–3046, 10.1039/b912547g , Epub 20090922, PubMed PMID: 19823716.
- C. K. W. Tam and W. A. Hyman, Transverse motion of an elastic sphere in a shear field, J. Fluid Mech., 2006, 59(1), 177–185, DOI:10.1017/s0022112073001497.
- C. E. Rasmussen and C. K. I. Williams, Gaussian Processes for Machine Learning, The MIT Press, 2005 Search PubMed.
- Y. L. Chen, Inertia- and deformation-driven migration of a soft particle in confined shear and Poiseuille flow, RSC Adv., 2014, 4(34), 17908–17916, 10.1039/c4ra00837e , PubMed PMID: WOS:000335227700064.
| This journal is © The Royal Society of Chemistry 2026 |