 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
The evolution of nanopore measurements: from biological out-of-plane pores to plastic in-plane pores
Khurshed
Akabirov†
 ab,
Hanna
Nguyen†
ab,
Shakila
Peli Thanthri
ab,
Hanna
Nguyen†
ab,
Shakila
Peli Thanthri
 ab,
Sheila
M. Barros
ab,
Sheila
M. Barros
 ab,
Maximillian
Chibuike
ab,
Maximillian
Chibuike
 ab,
Sunggook
Park
ab,
Sunggook
Park
 bc and
Steven A.
Soper
bc and
Steven A.
Soper
 *abcdefg
*abcdefg
aDepartment of Chemistry, The University of Kansas, Lawrence, KS 66045, USA
bCenter of BioModular Multiscale Systems for Precision Medicine, USA
cDepartment of Industrial and Mechanical Engineering, Louisiana State University, Baton Rouge, LA 70803, USA
dDepartment of Mechanical Engineering, The University of Kansas, Lawrence, KS 66045, USA
eBioengineering Program, The University of Kansas, Lawrence, KS 66045, USA
fKU Cancer Center, University of Kansas Medical Center, Kansas City, KS 66160, USA. E-mail: ssoper@ku.edu
gKansas Institute of Precision Medicine, Kansas City, KS 66160, USA
First published on 28th January 2026
Abstract
Nanopore sensing provides an ideal strategy for the label-free detection of single molecules in a variety of application scenarios. Working under the principle of resistive pulse sensing (RPS), nanopores consist of constrictions with sub-100 nm dimensions to enable single-molecule resolution by matching pore size to target dimensions (scaling); the optimal signal-to-noise ratio (SNR) results when the electrically biased pore is comparable in size to the molecule to be analyzed. When single molecules are electrokinetically transported through such remarkably small pores, they temporarily disturb the flux of ions moving through them, generating unique signals. These signals vary based upon the molecules' shape, size, orientation, and other physicochemical properties. Nanopores are generally divided into two main categories owing to their fabrication approach and material: biological and solid state. While biological nanopores have been the dominant sensor format due to their exceptionally small size, solid-state nanopores can demonstrate high performance characteristics attributed to their rigidity, stability, and high versatility in shape, material, and configuration. This review will explore the state-of-the-art in biological and solid-state nanopores and their abilities to detect and identify single biomolecules in a label-free manner. We will also review two topographical configurations of nanopore sensors; in-plane and out-of-plane sensors. The evolution of nanopore sensing will be reviewed, starting with out-of-plane biological sensors and progressing to in-plane sensors fabricated in plastics via replication technologies.
Introduction
Resistive pulse sensing (RPS) uses an electrically biased nano-constriction (i.e., nanopore) with the monitoring of ionic current as individual entities translocate through the nanoscale pore. This technology traces its roots back to the pioneering work of Wallace H. Coulter, who developed the first “hole-based” sensing technique in the late 1940s and was later patented in 1953 as a method for detecting and counting micrometer-sized blood cells suspended in liquid (i.e., Coulter counter).1 According to the Coulter principle,2 a particle moving through a constriction of similar size that separates two electrolyte-filled chambers produces a change in constriction resistance that is proportional to the volume of the particle translocating through the pore. While its initial development arose from the need for counting blood cells, DeBlois et al.3 adapted the technology about two decades later for the detection of colloidal particles and bacteriophages using sub-micrometer sized pores. Reducing the constriction size improved resolution and enabled the detection of individual biomolecules, thereby paving the way for new applications with major advancements occurring in the 1990s through efforts in single-molecule DNA sequencing.4–7 Since then, nanopore sensing applications have expanded due to their single molecule sensitivity, label-free capability, remarkable versatility, and enhanced insights into nanoscale phenomena.8 These insights resulted from an amalgamation of multiple disciplines including chemistry, biology, physics, engineering, and materials science. Nanopore sensing applications now encompass a vast array of molecule types and particles such as mono-9 and oligonucleotides,10 proteins,11–15 carbohydrates,16 viruses,17–21 extracellular vesicles,22–24 and others.Single entity occupation within a nanopore modulates the flow of ions either by physically obstructing the pore, which leads to a decrease in ionic current (current blockade), or by introducing a local enrichment of charge carriers such as counterions associated with highly charged entities, which can increase the ionic current. Because detection is dependent on ionic current blockade or enhancement, the translocation dynamics (i.e., current change magnitude, duration, and signal shape) may be correlated to unique characteristics and behavior of the entity moving through the pore under the influence of an electric field. These include properties such as size,19,25–27 conformation,11 mechanical stability/deformation,21,28 surface charge,29 and/or electrophoretic mobility.14,18,23,25 The diameter of classical nanopore sensors range from 1–100 nm while those of extended nanopore sensors range from 100–1000 nm.30 The sensor's physical (size, shape, orientation, conductivity) and chemical (composition and surface functional groups) properties as well as run buffer conditions may be tailored to fit the application need.
In this manuscript, we will provide a timely review of the evolution of RPS nanopore sensors using different materials and configurations for the sensing of single molecules. We will also discuss the basic operating principles of nanopore sensing, including ionic current modulation, voltage-driven analyte translocation, and signal interpretation based on molecular properties. This will give the reader a fundamental basis from which performance comparisons can be made for pores of different configurations and/or materials.
General operating considerations for RPS
The high surface area-to-volume ratio associated with nanopores elicit a significant contribution of surface properties on the transport dynamics of single molecules through the pores and their RPS signals,8,32 which are typically negligible at the microscale. These phenomena include, but are not limited to, electrical double layer (EDL) overlap, ion permselectivity, and ion concentration polarization. Because electrostatic forces are a major contributor to these phenomena, factors such as electrolyte type and concentration, pH, and voltage must be considered as well. Finally, pore size and shape can affect the resultant signal observed in RPS. These effects will be discussed briefly in the sections that follow.Surface charge effects
Many surfaces acquire an electric charge when in contact with an electrolyte solution. The nature and density of this charge depend on a combination of material properties (surface functional groups), solution characteristics (pH and ionic species), and certain interactions (e.g., adsorption/desorption at solid/liquid interfaces). This surface charge affects the distribution of hydrated counterions (ions of opposite charge to the surface charge) and co-ions in solution forming two distinct layers of ions known collectively as the EDL. Due to electrostatic interactions, a single layer of hydrated counterions is positioned adjacent to the surface forming the Stern layer (Fig. 1a). This fixed layer of counterions does not sufficiently screen the surface charge, so additional hydrated counterions accumulate next to the Stern layer.31 Because of their greater distance from the surface, these counterions are mobile due to Brownian motion. To maintain electrical neutrality, hydrated co-ions are also present. Together, these mobile counterions and co-ions form the diffuse layer where the concentration of counterions exceeds that of co-ions until it gradually approaches equilibrium with the bulk solution. | ||
| Fig. 1 (a) The formation of electric double layer (EDL) on solid/liquid interface also known as Gouy–Chapman–Stern model. Negatively charged surface attracts/adsorbs non-hydrated anions, partially hydrated cations and fully hydrated cations forming inner Helmholtz (Stern layer) and outer Helmholtz (diffuse layer or Gouy–Chapman) planes. Diffuse layer mainly consists of hydrated co-ions. Reproduced with permission from Schoch et al.31 Copyright 2005 American Institute of Physics. (b) Ion enrichment (cathodic side) and depletion zones (anodic side) formed at a micro- to nanostructure interface. | ||
The interface between the Stern and diffuse layers is known as the slip or shear plane. The potential difference at this interface is called the zeta (ζ) potential. The ζ potential decays exponentially in the diffuse layer and the characteristic length over which this decay occurs is known as the Debye length (λD). The Debye length is represented by;8
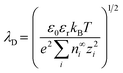 | (1) |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 electrolyte solution at 25 °C, eqn (1) simplifies to;8
1 electrolyte solution at 25 °C, eqn (1) simplifies to;8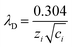 | (2) |
Under a bias voltage, charged entities migrate electrophoretically towards electrodes of opposite charge. Neutral particles move with the bulk fluid flow, which is towards the cathode for negatively charged surfaces. This bulk flow, known as the electroosmotic flow (EOF), is generated from the viscous drag exerted by cations in the diffuse layer as they migrate towards the cathode. EOF can either reduce or increase the driving force on the molecule/particle depending on the direction of the electrophoretic mobility (μep). While particle charge density influences its μep, surface charge density and solution characteristics influence the electroosmotic mobility (μeof), which can be described by eqn (3);8
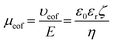 | (3) |
When the dimension of the nanopore is comparable to that of the EDL, EDL overlap can occur transforming the classical plug-like EOF profile into a more parabolic flow profile.33 EDL overlap also causes the nanopore to become charge-selective, leading to ion current rectification (ICR) and concentration polarization.32,34–36 With EDL overlap, the ion flux must pass through the diffuse layer, which has a non-uniform distribution of ions despite being electrically neutral. This non-uniformity creates a potential gradient that favors the transport of counterions while hindering the movement of co-ions. When there is asymmetric surface charge and/or asymmetric nanopore geometry, ICR can occur stemming from the unequal migration of cations and anions that generate a higher ion current in one direction.34,35,37,38 This is observed as a deviation from ohmic behavior in the current–voltage (I–V) curve. The degree of rectification is determined from the rectification ratio (the ratio of current at voltages of opposite polarity but equal magnitude).37 Lower ionic strength solutions typically produce greater rectification because surface charge contributes more significantly to the total current. Because analyte-induced changes on the nanopore surface can modulate the I–V response, ICR can affect the accuracy of applications that rely on an Ohmic response.
Concentration polarization, while related to ICR, differs in its manifestation. It describes the spatial distribution of ions within and around a nanopore and can be induced by EDL overlap or by an electric field without requiring surface charge asymmetry.39 Ion transport involves both diffusion and electrokinetics. For negatively charged surfaces, when diffusive transport is slower than electrokinetic transport (i.e., sufficiently high applied voltage), freely migrating cations concentrate on the cathodic side (enrichment zone) at a micro- to nanostructure interface leaving the anodic side depleted of cations (depletion zone); see Fig. 1b.33,34,36 Co-ions follow a similar enrichment and depletion pattern to preserve charge balance.
Effects of voltage, salt, surface charge, and pore dimensions on RPS
Despite its well-established name, RPS is only a partial descriptor for the operational principles of the method because it monitors transient current changes rather than direct resistance. Current changes can result from blockades (resistive pulses), enhancements (conductive pulses),40–42 or a combination of both (biphasic pulses)43,44 depending on analyte and pore surface charge, buffer composition, and applied voltage. The baseline or open pore current (I0) is described by eqn (4),45 where V is the bias voltage, μ+ and μ− represent the electrophoretic mobilities of cations and anions, respectively, n± is the number density of the salt ions, e is the elementary charge, heff is the effective pore length or membrane thickness of the pore, and d is the diameter of the pore.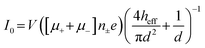 | (4) |
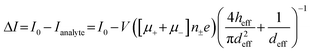 | (5) |
Salt type influences ΔI through its conductivity (e.g., K+ > Na+ > Li+); Fig. 2. While salt type weakly affects ΔI, it strongly impacts pulse duration via charge screening by counterions.46 Smaller, more strongly hydrated ions (Li+ < Na+ < K+) screen the surface charge more effectively, thereby reducing effective charge and increasing td. Higher salt concentrations similarly affect pulse characteristics as they enhance charge screening. If charge reduction is such that μep < μeof, the translocation would be EOF-dominant.15 For proteins or protein-containing analytes, different salt types exert chaotropic (salting-out) or kosmotropic (salting-in) effects in accordance with the Hofmeister series.15 Asymmetric salt conditions, where the salt concentration is lower in the cis (analyte-loading) chamber relative to that in the trans (analyte-receiving) chamber, have been shown to increase event rates and pulse durations for negatively charged analytes.47,48 Wanunu et al.47 noted that the increased pumping of cations into the cis chamber creates a polarized environment near the pore. This polarization increases the potential well adjacent to the pore funneling analytes towards the entrance and enhancing their capture rate. For negatively charged analytes and surfaces, a higher trans-side salt concentration increases pulse durations due to an increased counterion flux to the cis side, which enhances EOF and reduces μapp. Additionally, a lower salt concentration results in a higher electrical resistance leading to a greater voltage drop outside the pore (cis side), and consequently a lower voltage drop inside the pore compared to symmetric salt conditions. This reduced effective voltage results in a weaker electric field slowing down analyte translocation through the nanopore.47
 | ||
| Fig. 2 Effects of salt type on the RPS signal characteristics for both ssDNA and dsDNA analytes. (a) Representative current traces for 48.5 kbp λ-dsDNA translocations recorded at 5 kHz with different monovalent salts. (b) Similar recordings in 1 M, 2 M, and 4 M LiCl solutions. (c) Translocation traces for heat-denatured M13mp18 ssDNA in the presence of 8 M urea and 1 M monovalent salts, recorded at 30 kHz. (d–f) Corresponding histograms of translocation times for the conditions in (a–c). Reproduced with permission from Kowalczyk et al.,46 Copyright 2012 American Chemical Society. | ||
The non-specific nature of charge screening also affects the surface charge of the pore reducing co-ion exclusion and increasing event rates. At low salt concentrations, surface charge dominates contributing significantly to the total ion current. Conversely, at higher salt concentrations pore conductance is comparable to that of the bulk solution.40 Charge screening of analyte surface charge diminishes ΔI by adding counterions to the ion flux reducing the number of net displaced ions (i.e., volume exclusion effect). Sufficiently enhanced analyte conductance, facilitated by a counterion cloud, can lead to observable conductive pulses.40–42
In sensors exhibiting concentration polarization, biphasic pulses may occur.43,49 In such cases, analyte migration through ion-rich regions results in resistive pulses, because current change is primarily due to volume exclusion of ions. Migration through ion-depleted regions produces conductive pulses when screening counterions sufficiently counteract the volume exclusion effect.43,44 For applications in which controlling surface charge is crucial, surfaces can be engineered to have the desired charge density via material selection, surface activation or modification, and/or buffer optimization.
RPS signal, noise, and bandwidth characteristics
In RPS, changes in ionic current due to a translocation event are transduced by the appropriate electronics. Application of a bias voltage with Ag/AgCl electrodes (reversible and non-polarizable)50 across an electrolyte solution on either side of the nanopore generates a baseline ionic current (I0), typically in the nanoampere (nA) range, with transient current changes as small as picoamperes (pA).18,23,51–53 Current amplifier circuity is able to amplify these changes to a measurable level and convert the RPS events to a voltage signal. High-frequency noise is then removed from the signal using a low-pass filter (LPF), which, in conjunction with the electronic amplifier circuitry and the RC time constant of the nanopore, establishes the system bandwidth (BW). A common challenge in RPS is that the timescale of translocation events (as short as tens of microseconds) can be shorter than the system's temporal resolution (temporal resolution ∼1/BW). A higher BW improves temporal resolution but also increases system noise, creating a trade-off in signal fidelity (see Fig. 3a and below for further discussion of BW).54–57 | ||
Fig. 3 (a) Comparison of four current transient amplitudes obtained via electrokinetic translocation of icosahedral virus particle through conical nanopore collected at four different BWs. (b) Four different noise associated with RPS and their characteristic frequency response range. (c–h) Theoretical illustration of four different noise: (-----) thermal noise, (-·-··) headstage and amplifier noise, (-··-) dielectric noise, (⋯⋯) quantization noise, (![[thick line, graph caption]](https://www.rsc.org/images/entities/char_e117.gif) ) total noise generated by the glass and PET substrates at 1, 10 and 40 kHz frequencies. (a) and (c–h) Reproduced with permission from Uram et al.,54 Copyright 2008 American Chemical Society. (b) Reproduced with permission from Fragasso et al.,55 Copyright 2020 American Chemical Society. ) total noise generated by the glass and PET substrates at 1, 10 and 40 kHz frequencies. (a) and (c–h) Reproduced with permission from Uram et al.,54 Copyright 2008 American Chemical Society. (b) Reproduced with permission from Fragasso et al.,55 Copyright 2020 American Chemical Society. | ||
Noise and BW
Noise and BW are essential parameters to consider in any RPS measurement. Complete resolution of an RPS event requires a specific BW. Operating at a low BW can lead to signal attenuation (i.e., smaller amplitude) and pulse broadening (i.e., wider pulse width),15,54,58 as illustrated in Fig. 3a; the signal amplitude is comparable for a system BW of 40 and 10 kHz, but the current trace at 40 kHz has higher system noise. It is standard practice to set a detection threshold based on the root-mean-square (RMS) noise in I0. Higher RMS noise necessitates a higher detection threshold, potentially excluding low-amplitude translocation events and lowering the overall event rate. Understanding the sources and characteristics of noise can aid in noise reduction in current traces. Increasing the BW improves the system's temporal resolution but also may permit higher-frequency noise to pass reducing the SNR.Each component in the measurement system (i.e., nanopore, headstage, amplifier, LPF, and digitizer) has a BW and the component with the lowest BW typically sets the upper limit on the operating temporal resolution.54 Nanopore substrates are typically dielectric materials (i.e., SiO2, SiNx, glass, polymer), which have both resistive and capacitive components.59 The BW can be estimated from;54,59,60
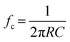 | (6) |
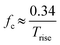 | (7) |
A nanopore behaves as a resistor within a sensor network and noise arising from a resistive component is thermal noise (also known as Johnson–Nyquist noise). Thermal noise results from the agitation of charge carriers inside the sensor.63 This type of thermal noise is white, meaning its power density is independent of frequency.55,64 Another component of white noise is shot noise and is due to the quantized nature of charge and arises from charge carriers crossing a potential barrier65 (i.e., interface between electrode and buffer). White noise typically dominants at higher frequencies (1 kHz to 1 kHz or more) compared to 1/f noise.
At high frequencies (>1 kHz), dielectric materials used to fabricate the nanopore exhibit energy dissipation in the form of thermal energy contributing to a specific type of noise known as dielectric noise. This noise arises as a consequence of the insulating material not being able to perfectly respond to the rapidly alternating electric fields. Additionally, the capacitance of the substrate material can also generate capacitive noise, especially in high-impedance environments.55
An analog signal is converted to a digital signal using an A/D converter. Quantization is the mapping of the continuous values of an analog signal to discrete values representing the digital signal by the A/D converter. Each value to the nearest integer multiple of the A/D conversion is the smallest change in signal that the digitizer can distinguish.63 This approximation introduces a type of error known as quantization noise, which is the difference between the actual analog signal and the digitized signal. The conversion resolution is dependent on the bit depth of the digitizer. The higher the bit depth the smaller the quantizing step (also called the least significant bit, or LSB) and thus, lower quantization noise. In addition to quantization noise, the headstage and amplifier also can generate high-frequency capacitive noise (>10 kHz) from electrode wiring and coaxial cables, amplifier input, and feedback elements.63,66
Uram et al.54 determined the theoretical noise levels in a nanopore measurement and compared them to experimentally measured noise using an Axopatch 200B amplifier at 1 kHz, 10 kHz, and 40 kHz for glass, and polyethylene terephthalate (PET) substrates. While theoretical quantization noise was minimal across all conditions likely due to high digitizer resolution and system filtering. Thermal noise dominated at lower pore resistances (≤30 MΩ) due to increased current levels. At a BW of 10 kHz, thermal noise dominated up to resistances ∼400 MΩ (Fig. 3c–h). PET exhibited higher dielectric noise compared to glass, which was attributed to PET's higher capacitance and dissipation factors. At higher applied voltages, noise increased significantly and was attributed to 1/f noise. Patil et al.67 used molecular dynamic simulations to predict noise in solid-state nanopores (SSNPs) concluding that low-frequency (<100 Hz) 1/f noise was due to surface charge density aligning with the number fluctuation theory. Due to these noise sources, conventional patch-clamp amplifiers are limited to a BW of ∼100 kHz. However, most experiments are performed at ∼10 kHz to reduce overall noise levels and ensure signal fidelity.
Noise suppression and temporal resolution for RPS measurements
In addition to employing appropriate high-pass filters (HPF) and LPF, several strategies can be incorporated to minimize noise in RPS measurements: (i) increasing pore resistance by reducing pore diameter or increasing pore length to reduce thermal noise; (ii) increasing surface hydrophilicity to reduce nanobubble formation54 or increase surface charge density leading to surface conduction dominating over bulk conduction all of which reduce flicker noise;67 (iii) selecting substrate materials with low dielectric loss to lower dielectric noise; (iv) use of extremely small area capacitor plates and shortening the wiring length between the amplifier and sensor to reduce capacitive noise; and (v) shielding the system with a Faraday cage to help isolate the RPS experiment from external electromagnetic interferences.Digital post processing with bandpass filters are routinary used to reduce the RMS in I0.68,69 Bessel filters are commonly used as LPF, but can also attenuate blockade signal amplitude especially for fast translocation events. Even in the case for events that are scored properly, high-frequency components in the RPS signal can be lost as well when employing LBF.70
Wavelet and deep learning have also been proposed to reduce noise for RPS measurements. For the wavelet method, the signal and background noise are separated in the wavelet domain with a breakdown of the signal that shows the time and frequency of the events allowing large-magnitude wavelet components to remain, while small-magnitude components are eliminated.69 In the case of deep learning, the Noise-2-Noise technique uses pairs of noisy nanopore signals to denoise them without requiring a clean signal and then employing convolutional neural networks to learn the mapping from noisy data followed by a deconvolution or reconstruction of the data revealing features in the signals that otherwise would be hidden.71
Rosenstein et al.66 developed a custom-built preamplifier using complementary metal-oxide semiconductor (CMOS) technology that was tested by integrating it to a SSNP embedded within a thin SiNx membrane. The amplifier could operate at a BW of 1 MHz and produced a SNR >5 for RPS events at a temporal resolution of 1 μs. The preamplifier circuitry, positioned directly within the fluid chamber, reduced parasitic capacitance from the SiNx and electrode (Ag/AgCl electrodes fabricated directly on the preamplifier surface), thereby lowering high-frequency capacitance noise. Shekar et al.72 later implemented a similar CMOS setup with a 10 MHz BW that was able to achieve a temporal resolution approaching 100 ns with a SNR of 10 at 5 MHz BW. Hartel et al.73 utilized a 500 kHz BW with a temporal resolution of 2 μs to perform a beta distribution analysis to observe events as fast as 35 ns. For a more extensive discussion on high BW approaches in nanopore recordings, readers are directed to this comprehensive review.74
Nanopore sensors can be modeled as a resistor–capacitor (RC) circuit that will determine the speed of the ionic current modulation measurements. In this context, in-plane pores fabricated in a polymer or glass exhibit distinct RC characteristics due to their unique material properties, pore size, geometry, and surface chemistry. Combined with the polymer capacitance and its unique dielectric properties, this will result in characteristic resistance–capacitance filtering.75 Conversely, out-of-plane pores create a RC circuit resulting in high-capacitance limiting time resolution.55
Sampling rate and signal aliasing in RPS
Sampling rate (Hz) is the number of data points recorded per unit time and the appropriate rate is dependent on the type of analysis to be performed. For time-domain analyses (e.g., pulse analysis), sufficient measurements must be recorded to ensure sampling intervals are shorter than translocation events to prevent signal distortion or missed events. According to the Nyquist theorem,76 the sampling frequency should be ≥2× the system BW. However, in nanopore RPS applications a rate of ≥10× the BW is often used.9,15,21,24 For frequency-domain analyses (e.g., noise analysis), it is essential to prevent aliasing, a phenomenon where high frequency components (above the Nyquist frequency) fold back into the lower frequency range distorting signals in the frequency range of interest. To prevent this, a low-pass anti-aliasing filter (a filter with a sharp attenuation) can be placed before signal digitization. Butterworth and elliptic filters are commonly used in frequency-domain analyses due to their sharp attenuation. In contrast, Bessel filters are common for time-domain analyses due to their lower overshoot, a phenomenon where the recorded signal exceeds the expected value during a transient response.63 Oversampling is less common in frequency-domain measurements not because it causes aliasing, but because it may unnecessarily increase data volume without any improvement in frequency information.In the sections that follow, different pore configurations (in-plane versus out-of-plane) and the use of different materials for nanopore measurements will be discussed. Regardless of the pore configuration or material, noise contributions to the RPS data remain relevant as discussed above, even though their magnitude and impact may vary depending on the pore configuration and/or material used for the nanopore. Also, the underlying physics of RPS signal generation are similar in all cases. We will highlight these common principles and variations as we explore each of the pore materials and configurations discussed in the following sections.
Pore configurations: out-of-plane vs. in-plane pores
Accurate detection and identification of analytes are not only determined by the driving voltage, buffer composition, and electronic parameters (see eqn (4) and (5)), but also the associated nanopore shape and configuration. While the pore size is important in terms of determining the SNR in any RPS measurement, the pore configuration including its orientation with respect to the translocation direction (out-of-plane vs. in-plane), surface chemistry, and shape can have a profound impact on the measured response as well as noise contributions to the measurement. For example, adjusting pore geometry can enhance the detection accuracy and sensitivity.11,77 Additionally, modifying surface properties such as charge density or functional groups11,23,78 can affect potential analyte–nanopore wall interactions, concentration polarization and/or ICR.Nanofluidic sensors in general contain micro- and nanoscale structure interfaces. Such interfaces may introduce entropic barriers during the voltage-driven translocations of large biomolecules, that are not observed in microscale structures.8,33,79–82 Proper configuration of the micro-nanostructure interface can lower the entropic barrier providing smooth entry of macro-molecules into the nanopore, thereby increasing event capture rate.83 Additionally, surface charge effects of the pore must be carefully considered as they can determine the amount of concentration polarization, ICR, and/or translocation time and signal amplitude, all of which affects overall sensor performance.
Nanopore geometries broadly speaking can consist of the out-of-plane11,12,21,84 or the in-plane configuration.24,85 In this review, we define out-of-plane nanopore sensors as those in which the analyte translocates through the pore orthogonally to the sensor's substrate, while for in-plane pores the analyte translocates parallel to the sensor substrate. In-plane and in some cases, out-of-plane nanopores can be designed as a single pore, multiple pores in series,51,86 or multiple pores in parallel,17 which can increase the information content of nanopore RPS measurements (serial configurations) or the measurement throughput (parallel configurations), respectively.
Out-of-plane nanopores
Out-of-plane biological nanopores
Over the past few decades, many researchers have widely used biological out-of-plane nanopores in a broad range of applications including DNA/RNA sequencing, and protein analysis to name a few. They can be engineered by various techniques preserving their physical and chemical stability across different devices.87 Examples of different biological nanopores includes α-HL,88,89 MspA,87,90 and φ29.91Table 1 summarizes their important operational parameters. The α-HL nanopore from Staphylococcus aureus possesses a molecular mass of ∼33 kDa and contains three distinct regions; the cis entrance, central constriction, and trans exit.88,92 The α-HL pore has been utilized for DNA sequencing as it allows the translocation of single stranded (ss) DNA through the nanopore with each nucleotide generating distinct RPS signals due to their structural variations (Table 1a).88 The MspA porin nanopore, on the other hand, is extracted from the Mycobacterium chelone and contains a more cylindrical shape (Table 1b).90 Its octameric structure can make it an ideal candidate for nanopore sequencing because of the ability to generate distinctive RPS signals for A, T, C and G.93 The third nanopore type is φ29 from Bacillus subtilis and has a propeller-like shape (Table 1c).91| Nanopore type | Structural view | Pore diameter | References | |
|---|---|---|---|---|
| a Reproduced with permission from Tanaka et al.,88 Copyright 2011 The Protein Society. b Reproduced from Faller et al.90 with permission from American Association for the Advancement of Science. c Reproduced with permission from Guasch et al.,91 Copyright 2002 Academic Press. | ||||
| α-HL |

|
1.4 nm | Tanaka et al.88 | |
| Ying et al.89 | ||||
| MspA |

|
1.2 nm | Faller et al.90 | |
| φ29 |

|
3.5 nm | Guasch et al.91 | |
Stoddart et al. compared the RPS behavior of wild-type (WT) and E11N/K147N modified α-HL nanopores by shuttling homopolymer and heteropolymer ssDNAs through the pores and observing current drop amplitudes.94 They immobilized oligonucleotides at the pore vicinity via biotin tags at their 3′ termini. Upon application of a bias voltage, the oligonucleotides were captured by the nanopore and held there over a period of time, while residual current generated by each nucleotide were detected and recorded.
The specific nucleotide sequences used in these experiments consisted of 5′CCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCNCCCCCCCCBtn-3′ (Fig. 4a), 5′CCCCCCCCCCCCCCCCCCCCCCCCCCNCCCCCCCCCCCCCBtn-3′ (Fig. 4b) and 5′CCCCCCCCCCCCCCCCCCCCCCNCCCCCCCCCCCCCCCCCBtn-3′ (Fig. 4c), where N was either G, A, T, or C. Both nanopores were used to distinguish the nucleotide at the “N” position. The E11N/K147N modified α-HL nanopore provided superior performance on the identification of nucleotide-specific current levels over its WT counterpart by comparing their Ires (residual current) distribution. Clarke et al. mutated α-HL nanopore to contain an am6amPDP1βCD adapter with a cysteine substitution at position 119, 121, 123, 135 or 137 amino acid residues. Electrokinetic translocation of single dNMP molecules through the nanopore produced the current trace shown in Fig. 4d.95 The modified nanopore established discrimination of all four nucleotides at the 99% confidence level (Fig. 4e). They further investigated the RPS performance of this mutated nanopore by translocating ssDNA mixed with Exo-1 and detected RPS signals produced by the enzyme-assisted DNA digestion of DNA under asymmetric salt conditions (see Fig. 4f).95
 | ||
| Fig. 4 Molecule translocation through biological nanopores. Histogram plot (top) for Ires and a table (bottom) highlighting the average Ires for DNA/α-HL pore interaction where oligonucleotides have a sequence of (a) 5′-C31NC7Btn-3′, (b) 5′C26N13Btn-3′, and (c) 5′C22N17Btn-3′. Reproduced from Stoddart et al.94 with permission from Proceedings of the National Academy of Sciences. (d) Current trace generated by translocation of dNMPs through modified α-HL nanopore and (e) Gaussian fit histogram plot of Ires for in the presence of four nucleotides. (f) Schematic image of ssDNA/Exo-1 interaction in the cis chamber and produced mononucleotide translocation through modified α-HL nanopore. Reproduced from Clarke et al.95 with permission from Proceedings of Nature Nanotechnology. | ||
While biological nanopores are known for their exceptionally small size and relatively low noise, there are several challenges associated with them.87 The process of designing, fabrication, and modification of biological nanopores are often time-consuming and complex. Additionally, laboratory-purified proteins are available in limited quantities, which makes large-scale nanopore analyses difficult.87 Also, the fragility of the lipid bilayer can be problematic, especially when an electric field is applied across the nanopore limiting its operational lifespan.96 However, efforts have been made to stabilize the supporting biological membrane such as encapsulation of free-standing planar lipid membranes in a polymer hydrogel via in situ photopolymerization of poly(ethylene glycol)dimethacrylate (PEGDMA).97
Out-of-plane solid-state nanopores (SSNPs)
Out-of-plane SSNPs are typically fabricated in free-standing membranes such as silicon nitride (SiNx), silicon dioxide (SiO2),98 glass,99 or ultrathin 2D materials such as molybdenum disulfide100 or boron nitride.101 These membranes are placed between two fluidic reservoirs made out of materials such as Teflon,21 poly(dimethylsiloxane),99 polymethylmethacrylate,102 or polycarbonate.103 The membrane-embedded out-of-plane pores can be fabricated using transmission electron microscopy (TEM),104 where an electron beam is tightly focused to a desired spot on the membrane and used to open the membrane to crease a nanopore. While TEM offers high precision, it requires expensive instrumentation and has low throughput; typically producing ∼1 device per hour. Focused ion beam (FIB) milling using Ga+ (ref. 105 and 106) or He+ ions107 improves device production throughput allowing the fabrication of 20–30 devices per hour. Controlled dielectric breakdown (CDB)108,109 has emerged recently wherein an applied voltage is used to form a pore at a membrane defect. This approach does not require sophisticated equipment and in principle, allows for parallelized fabrication to increase production. However, CDB has limitations. For example, pore formation in CDB is stochastic, leading to variability in pore size and location. It is also limited to producing single-pore devices and thus, challenges associated with increasing parallelization. Moreover, the nanopores created by CDB tend to grow over time limiting their long-term stability and utility.Inorganic membrane pores
Table 2 summarizes various membrane-based nanopore types. The pore developing process within the membrane requires precise alignment of the ion beam on the membrane surface and controlling the exposure time to provide the most desirable pore shape and diameter. When the dosage of the pore-forming beam is carefully controlled, the size of a nanopore can be adjusted to <5 nm.110| Pore type | Pore height | Pore diameter | Pore fabrication | Schematic view of experimental set up (left) and SEM/AFM images of nanopore size | Reference |
|---|---|---|---|---|---|
| a Reproduced with permission from Xia et al.,119 Copyright 2022 American Chemical Society. b Reproduced from Kennedy et al.121 with permission from Proceedings of Nature Nanotechnology. c Reproduced from Liu et al.,100 with permission from American Chemical Society. d Reproduced from Wang et al.127 with permission from Proceedings of Nature Nanotechnology. e Reproduced from Traversi et al.126 with permission from Proceedings of Nature Nanotechnology. | |||||
| SiNx | >20 nm | >4.8 nm | HF etching |

|
Xia et al.119 |
| SiNx | ∼10 nm | <1 nm | Electron bean-induced sputtering |

|
Kennedy et al.121 |
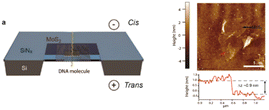
| MoS2/SiNx | 0.7 nm (MoS2)/20 nm (SiNx) | >5 nm | TEM |
|
Liu et al.100 |
| MoS2 | ∼10 nm | ∼0.8 nm | CVD |

|
Wang et al.127 |
| SiNx/GNR | 0.335 nm (GNR)/20 nm (SiNx) | ∼10 nm | TEM |
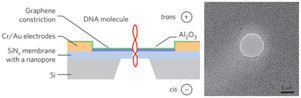
|
Traversi et al.126 |
Membrane-based nanopore fabrication and implementation was first reported by Li J. et al. in 2001,111 who used an argon (Ar+) ion beam to generate 5 nm pores on the surface of a bowl-shaped SiNx membrane.111 When testing the nanopore, a constant ionic current (I0) of 1.66 nA resulted at 120 mV applied voltage. When a 500 bp dsDNA was electrokinetically translocated through the nanopore, the results indicated current blockage dwell times in the millisecond regime. Current reductions (ΔI) were 88% lower compared to the open-pore current value.111
Silicon nitride nanopores
SiNx membranes are made of materials that exhibit excellent mechanical and chemical stability. Particularly, the nanopore size and shape are more adjustable in comparison to biological nanopores.112 The fabrication process of the pores within the SiNx membrane window involves: (1) deposition of SiNx film on the Si substrate; (2) photolithography and reactive ion etching to open a window on the membrane; and (3) nanopore formation using FIB milling,111,113 electron beam lithography (TEM),114,115 or laser assisted drilling.116,117 Alternatively, chemical etching methods can be employed for nanopore formation.118,119 Xia et al. fabricated 20–100 nm thick low-stress SiNx membranes on a glass substrate (Table 2a and b).119 Then, nanopores with diameters ranging from sub-nm to 100 nm were chemically etched into the membrane using hydrofluoric acid (HF).119 Zvuloni et al. employed laser drilling (LD) to fabricate pores in the 2.5–5 nm range.116 These nanopores were subsequently used for analyzing single protein molecules. Unlike biological nanopores, SiNx can be integrated with other techniques to enhance signal sensitivity. For example, Dela Torre et al. integrated TiO2 coated SiNx nanopore arrays (6 × 6) with total internal reflection fluorescence (TIRF) spectroscopy to analyze 1 kbp dsDNAs, which were covalently attached to streptavidin/quantum dots.120 Non-propagating evanescence radiation was able to excite the fluorophores only when the DNA-dot duplexes were electrokinetically directed towards the nanopore offering a unique and highly selective DNA detection technique.120 Timp's group121 reported a sub-nanometer SiNx pore for protein sequencing, which used a quadromer (four amino acid residue) appendage to resolve single residues with volume differences of ∼0.07 nm3.Aluminum oxide nanopores
Aluminum oxide (Al2O3) nanopores are gaining interest due to their optical and electrochemical properties, pore size adjustability, and high thermal stability.122 Compared to SiNx-based nanopores, Al2O3 nanopores offer lower noise, which is beneficial for high-sensitivity detection.123 A study by Venkatesan et al. demonstrated that DNA translocation was significantly slower using an Al2O3 nanopore compared to SiNx nanopores.124 The primary reason for this observation was ascribed to electrostatic attraction between the negatively charged DNA molecules and the positively charged Al2O3 nanopore walls, which partially reduced the speed of DNA during translocation.124Two-dimensional nanopores
Monolayer or 2D nanopores are exceptionally thin, which result in increased RPS signal amplitudes, but consequently short dwell times, which makes the system BW considerations critical. Commonly used materials for this platform are MoS2 or graphene.125 Liu et al. fabricated MoS2 a nanopore sensor with 0.7 nm thickness and compared its performance to SiNx by translocating λ-DNA through the 2D nanopores (Table 2c and d).100 The current transient data revealed a 5-fold increase in signal amplitude and a 10-fold improvement in SNR.100 Traversi et al. was able to deposit a 0.335 nm thin graphene layer on a SiNx membrane surface by chemical vapor deposition (CVD) (Table 2e).126 They integrated graphene nanoribbons (GNR) across the membrane and electrokinetically translocated DNA molecules through the nanopore employing two detection modes; current-based and electrical-based modes. The current-based mode was initiated by RPS criteria, and the electrical-based mode was achieved via inducing the transmembrane potential perpendicular to the DNA translocation direction.126The 2D pores have recently been demonstrated to detect individual amino acids, post-translational modifications,127 single nucleotides128 and DNA molecules.100 2D nanopores have also been used in studies of neurodegenerative disease with results associated with the characterization of oligomers of amyloid-forming proteins demonstrating high resolution when compared to a similar characterization using TEM or mass photometry.129
There are challenges with 2D nanopores such as short dwell times, which often exceeds the operational BW of the measurement system.112 Also, there are challenges associated with membrane current leakage introduced during film growth and membrane fabrication, damage of the 2D material due to electrical discharges, and the fact that the thin membrane nanopores are prone to widening and damage when in solution. These and other challenges as well as some alternative solutions are discussed in detail by Graf et al.130
Another important consideration is that because the ionic current through a nanopore is inversely proportional to its length and the translocation of a molecule blocks a large fraction of the ions occupying the pore volume, 2D nanopores have the advantage of generating high ΔI values.130 However, the thin MoS2 membranes come with a consequence of low mechanical stability as well as charge fluctuation of its suspended membrane, which increases 1/f noise. Thus, the signal to noise ratio (SNR) is ∼10,131,132 and consequently not so different from the values observed in pores constructed in SiNx membranes.133 Another challenge with graphene nanopores arises from clogging caused by hydrophobic interactions between DNA and graphene requiring a coating such as a self-assembled pyrenediol monolayer.134
Another interesting membrane-based out-of-plane pore system is MXene consisting of early transition metal carbides and/or nitrides of chemical formula Mn+1XnTx, where M is an early transition metal, x denotes carbon and/or nitrogen, and Tx represents surface functional groups such as –O–, –OH, –F, or –Cl.135 Mojtabavi et al.136 reported the development of a liquid–liquid interface assembly of Ti2CTx and Ti3C2Tx MXene sheets with an ∼60% process yield rate for producing the nanometer-sized pores in free-standing MXene membranes. The MXene pores exhibited lower noise characteristics when compared to the other 2D systems, such as graphene or MoS2. However, the MXene pores show challenges as do other 2D membrane pores such as poor structural stability, oxidation, and high-cost production.135,137
Track etched nanopores
Track etched nanopores are a class of out-of-plane SSNP fabricated by ion irradiation followed by chemical etching of polymer membranes, such as polycarbonate (PC) or polyethylene terephthalate (PET) (Fig. 5). This fabrication strategy enables production of cylindrical or conical shaped pores with tunable dimensions.138,139 In the standard fabrication process, high energy heavy ions (e.g., Kr, Xe, Au) are accelerated and bombarded onto a polymer film creating latent tracks, which form along the path the heavy ions travel. Subsequent chemical etching, usually done with a strong base such as NaOH, selectively removes damaged regions, which then form nanopores with size and shape that can be tuned by adjusting the etching time, temperature, and chemical concentration.140 | ||
| Fig. 5 (a) SEM image of a multi-porous (108 nanopores per cm2) PET membrane etched for 120 min (dbase 232 nm). Nanopore geometries: (b) cylindrical, (c) conical, (d) bullet-like, (e) double-conical (hourglass), (f) cigar-shaped track etched nanopores. Reproduced from Kaya et al.139 with permission from The Electrochemical Society. (g) Experimental setup for DNA (A10/T10, A40/T40 and T40) sensing through track-etched PET conical nanopore. (h) Examples of current traces and zoomed-in current blockade traces recorded for A10/T10 at (i) 250 mV and (ii) 500 mV; A40/T40 at (iii) 250 mV and (iv) 500 mV; T40 at (v) 250 mV and (vi) 500 mV. The symbol * identifies examples of current blockade signals zoomed below each trace. The current traces were obtained using pore with tip diameter = 3 nm and base diameter = 200 nm. Reproduced from Meyer et al.143 with permission from the Multidisciplinary Digital Publishing Institute. (i) Schematic illustration of BSA transport through a chiral porous membrane (i). The driving voltage was +2 V (anode was in the permeate side). (ii) Time-dependent fluorescence change when BSA was allowed to be transported through bare nanopores and L/D-Cys immobilized nanopores. (j) Time-dependent current changes during BSA translocation in L- (i) and D-Cys (ii) modified single nanopores under the same conditions. Reproduced with permission from Zhang et al.,142 Copyright 2017 Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim. | ||
Track-etched nanopores can be made into various shapes as shown in Fig. 5a–e. The shape of the pore directly affects the electrical signal, translocation speed, and rectification behavior of ions and molecules passing through them. Cylindrically shaped pores (Fig. 5b) have a constant diameter throughout the pore length and are formed by symmetric etching from both sides of the membrane. These pores are commonly used for size-based sensing and transport studies of single molecules. Conically shaped pores (Fig. 5c) have a narrow tip at one end of and a wider base at the opposite end and are generated by asymmetric etching (etching one side more aggressively than the other). These pores exhibit current rectification and are useful for directional transport and gating applications. The double conically shaped pores (Fig. 5d) provide symmetrical electric fields and can enhance analyte capture rate. These pores are created by etching both sides of the membrane in a controlled manner to form two conical shapes that meet at the center of the membrane. Bullet-like track-etched pores (Fig. 5e) refer to a pore geometry that combines a steep taper near the tip with a gradual widening toward the base resembling the shape of a bullet. During the fabrication, one side of the membrane is etched more aggressively often using a high-concentration of NaOH and etching is stopped quickly after breakthrough limiting enlargement of the tip. These pores have the ability for electric field focusing, translocation control, and improved directional selectivity. The cigar shaped pores (Fig. 5f) are a variant of asymmetric conical pores, where the pore gradually tapers from a wider base at the middle to a narrower tip at both ends and resembles the rounded profile of a cigar. This shape is fabricated through controlled etching. The gradual tapering of the shape produces a spatially varying electric field, which can help slow down the translocation of biomolecules like DNA or proteins. The smoother geometry also reduces fluctuations in the ionic current, which leads to increased SNR of RPS signals.
As an alternative approach for producing conical pores in a plastic, Choi et al. utilized UV nanoimprint lithography (UV-NIL) of a SU-8 resin to produce an array of pores.141 In this process, the Si master consisted of an array of needles with a tip diameter of ∼25 nm. Following UV-NIL, the SU-8 membrane could be heated to near its glass transition temperature to induce polymer reflow to allow shrinkage of the pores at a rate of 2.7 nm min−1 that could produce polymer pores of <10 nm. The device was used to detect the translocation of λ-DNA through the pores.
A study done by Meyer et al. demonstrated the use of track-etched nanopore in PET membranes for ssDNA and dsDNA translocation as shown in Fig. 5f and g.143 Analysis using current blockade amplitudes and dwell times resulted in poor resolution due to overlapping signals among different DNA types (A10/T10, A40/T40 and T40). Machine learning significantly enhanced the discriminatory power of a conical track-etched nanopore for detecting short DNAs with a classification accuracy from ∼50% (using two features) to as high as 82% (using all five features), especially at lower voltages and smaller pore sizes (see Fig. 5h(i–vi)).
The ability to modify chemically the surface of track-etched nanopores is an advantage of these out-of-plane pores. Surface modification of the pore with charged or biological molecules can improve the selectivity of the measurement or control analyte translocation speed. For example, Thangaraj et al. demonstrated ssDNA (40 mer) and dsDNA (10 mer and 40 mer) translocation through PET track-etched nanopores coated with Al2O3 by atomic later deposition.144 Modified pores with Al2O3 (isoelectric point ∼5) generated dwell times of 0.85 ms, 1.07 ms, and 0.65 ms for ssDNA, 10 mer dsDNA and 40 mer, respectively.
In another example, Zhang et al. studied selective protein (BSA; bovine serum albumin) translocation through chiral-modified conical shaped nanopores fabricated in a polyimide (PI) membrane.142 The experimental setup is shown in Fig. 5i(i). When the BSA was labeled with a fluorescent dye, they could use fluorescence detection to monitor how much protein was able to translocate through the nanopore membrane; D-Cysteine modified pores provided higher efficiency transport through the pore compared to a bare pore and a pore containing L-cysteine (see Fig. 5i(ii)). L-Cysteine modified (L-Cys) nanopores improved the protein capture rate compared to pores modified with D-cysteine (12.9 events per s for L-Cys pores and 1.6 for D-Cys pores; see Fig. 5j(i) and (ii)). This phenomenon was caused by BSA drawn into the modified nanopores using chiral gate modulation.142
As a final example, Ma et al. investigated the translocation of hyaluronic acid, HA, through glass conical nanopores both with and without immobilized enzymes.145 Without enzymes, the HA transit through different pore diameters and polymer concentrations followed de Gennes' scaling laws. When hyaluronidase was grafted at the base side of the pore, HA chains were enzymatically cleaved at the entrance, and degradation products were detected downstream; grafting at the tip side led to prolonged dwell times, reflecting HA–enzyme binding interactions before degradation.
Multi-pore sensors using in-series or in-parallel configurations
Out-of-plane nanopores can be arranged with a dual in-series or parallel configuration. For the in-series configuration, a layer-by-layer fabrication strategy is typically employed. For example, Chou et al.146 reported the use of a bilayer-coupled in-series nanopore sensor consisting of a vertically stacked architecture The sensor contained a top two-dimensional (2D) layer with a single pore ∼2 nm in diameter and a bottom SiNx layer with one or more pores ranging from 10–31 nm in diameter depending on the fabrication method (Fig. 6a). The dual-pore in-series sensor was used for the guiding, tracking, and detection of single molecules. | ||
| Fig. 6 (a) Fabrication schematic involving electron beam lithography (EBL), etching, transfer of two-dimensional materials, and alternating current scanning transmission electron microscopy sculpting (AC-STEM). SiN nanopores serve as a guiding and reusable (GURU) platform. The pore thicknesses are denoted as tSiN and t2D, the diameters as dSiN and d2D, and the overall thickness is represented as tGURU. Device configurations are represented as [N, M], for NSiN pores and M 2D pores. σSiN and σ2D represent the surface charge densities of silicon nitride (SiN) and 2D nanopores, respectively. Experimentally interlayer separation, L, can be manipulated by adjusting the time of reactive ion etching (RIE). Reproduced with permission from Chou et al.,146 Copyright with permission from Proceedings of Nature Nanotechnology. (b) A diagram showing a 3D architecture for single-cell lysis to DNA detection by ionic current measurement. A voltage, Vb, applied across the vertically stacked two nanopore membranes and the corresponding ionic current measured monitored with a pair of Ag/AgCl electrodes in phosphate buffered saline (PBS). (c) A photographic view of the integrated nanopore emphasing the top multipore sheet. The 50 nm-thick SiNx nanopore membrane in dark square area. (d) A 3D model illustrating the cross-sectional architecture of the integrated nanopore. (e and f) False-colored scanning electron micrographs of the SiO2 multipore and SiNx nanopore. Reproduced with permission from Tsutsui et al.,147 Copyright 2021 Wiley-VCH GmbH. (g) Schematic depiction of the experimental configuration illustrating a twin barrel nanopore. Reproduced from Cadinu et al.,148 with permission from American Chemical Society. | ||
Tsutsui and coworkers described a 3D-integrated nanopore device for the detection of single DNA molecules.147 The device consisted of a SiO2 multi-pore sheet stacked vertically on a SiNx nanopore membrane with all layers integrated onto a Si wafer. RPS was used to reveal signatures indicative of the folding states of polynucleotides. The authors demonstrated the device's ability to detect E. coli-derived DNA without amplification (Fig. 6b–f).
In a parallel dual out-of-plane configuration, Cadinu et al. reported a double-barrel nanopore design for regulating the molecular transport of single molecules.148 The two pores were ∼20 nm apart and located at the tip of a double-barrel quartz nanopipette (Fig. 6g).
In-plane nanopores
A unique feature of in-plane nanopore sensors is in their architectural flexibility. In contrast to out-of-plane sensor configurations, where the nanopore lies within a membrane separating two vertically aligned electrolyte-filled reservoirs, in-plane sensors have nanostructures that connect input and output microscale channels parallel to the surface. These micro/nanochannel interfaces use either a direct interface (i.e., blunt interface) or engineered structures to create a size-gradient (tapering geometry) of nanochannels that can enhance sampling efficiency. Also, unlike out-of-plane sensors, in-plane sensors enable simple integration with planner microfluidics facilitating sample pre-processing prior to the RPS measurement. In-plane nanopore sensors can also contain a single nanopore, multiple nanopores in series (to sequentially detect analyte properties), or arrays of parallel nanopores (to enhance throughput), depending on the measurement needs. The use of multiple nanopores enable unique measurements such as the time-of-flight (ToF) that are not achievable in single nanopore systems.As shown in Fig. 3, the noise in any RPS measurement arises from various sources including 1/f noise, white noise, and high-frequency noise typically resulting from capacitive and dielectric characteristics of the sensor. Beyond noise in any nanopore measurement, another key factor must be considered is the mass detection limit, which is related to the sampling efficiency of the sensing system. Although nanopore sensors can detect single molecules, they suffer from poor sampling efficiency. As a result, large mass loads of sample are required for the sensor to achieve reasonable detection rates (i.e., high capture rates). While increasing the applied voltage can increase the frequency of RPS events, it also increases the analyte velocity, which can detrimentally affect the event rate due to BW issues.
Glass (inorganic) in-plane nanopores
In-plane nanopores can be fabricated in glass or quartz substrates via e-beam lithography149 or focused ion beam milling, which are considered top-down fabrication strategies.150,151 The attractive nature of this fabrication strategy is that multiple pores (≥2) can be placed in series to offer some unique measurement opportunities as discussed above. For example, each nanopore in-series serves as an independent sensing element allowing transduction of the analyte more than once. Also, as an analyte moves sequentially from one pore to the next, the analyte's μapp can be calculated based on the distance between the two nanopores, the electric field strength, and the ToF between the pores. It should be noted that μapp is not nanopore-dependent and this can produce an RPS-independent and molecular-dependent characteristic that can aid in molecule or nanoparticle identification.Zhang et al. used FIB milling to fabricate two circular in-plane nanopores in a glass substrate (Fig. 7ai–vi).85 Two distinct FIB milling strategies were employed for nanopore fabrication. The first design used setting the FIB stage at 54° and drilling a lamella wall on the glass surface at 0° followed by changing the stage alignment to 0° and milling the nanopores at an incident angle of 36° (Fig. 7ai–iii).151 For the second design, the stage was tilted by 28° while the incident angle was set at 26° to construct a tilted lamella design (Fig. 7aiv–vi). The incident beam manipulation strategy and substrate orientation offered control over the final nanopore geometry. The vertical lamella design produced a conical shaped pore, while the tilted lamella formed a cylindrically shaped nanopore (Fig. 7aiii and vi).151
 | ||
| Fig. 7 Glass in-plane nanopore devices. (a) Schematic view of vertical (i) and tilted lamella (ii) fabricated by adjusting the stage and the incident beams angles (ii), SEM images of vertical lamella (iii) and tilted lamella (iv) holding two in-plane circular nanopores directly milled on the lamella walls (scale bar: 1 μm). SEM images of conical (v) and cylindrical (vi) nanopore shapes initiated due to lamella wall alignments (scale bar: 200 nm). Reproduced from Zhang et al.85 with permission from American Chemical Society. (b) SEM image of three in-plane nanopores milled by FIB instrument. (c) Schematic diagram of the entire device containing 4 reservoirs, 2 V-shape microchannels, nanofilters, nanochannels and nanopores. (d) A visual illustration of the EV particles translocating through the nanopores and generating current transient amplitudes. (e) Current transient amplitudes of 67, 82 and 115 nm EV particles. Reproduced from Young et al.23 with permission from American Chemical Society. | ||
The same group later developed an in-plane nanopore sensor featuring three in-plane nanopores in-series, each with dimensions of 200 nm width, 200 nm depth, and 600 nm length as illustrated by SEM images in Fig. 7b.152 Similar to the previous design, this device integrated microscale regions fabricated via photolithography and wet chemical etching with nanoscale features fabricated via FIB milling. However, instead of creating a circular nanopore into a lamella wall, they applied a vertical incident beam at 90° perpendicular to the substrate surface to create nanofilters, nanochannels, and nanopores (Fig. 7c).152 Moreover, this sensor design added a 3rd nanopore to improve detection accuracy. As shown in Fig. 7d and e, translocation of one biological nanoparticle through the three in-plane nanopore sensor generated three consecutive current blockades.152 Each particle size was inversely proportional to the signal amplitude as predicted by eqn (5).115,153,154 The main limitation associated with these glass-based devices is the low device production rate; each device requires multiple techniques for micro/nanostructure fabrication including photolithography, etching, and FIB milling.
Harms et al.155 utilized a dual-nanopore configuration (two nanopores in series) spaced 2 μm apart and each nanopore was 50 × 50 × 40 nm in width, depth, and length (W × D × L) to analyze hepatitis B virus (HBV) capsids. The μapp was calculated from the time difference (Δt) between two consecutive RPS translocation events and reflects the vector sum of the EOF and the electrophoretic mobility (μep) of the particle. In a follow up study, this same group employed a similar sensor design but increased the nanopore length to 400 nm for size resolution enhancement. This modification allowed size-based discrimination of HBV capsids by analyzing their distinct ionic current blockade signatures.18
Kondylis et al.51 used in-plane nanopore devices containing two, four, or eight pores in series, spaced 500 nm apart to study the assembly of HBV capsids. The use of multiple pores supported enhanced precision, allowing the researchers to detect subtle differences in μapp of capsids. Additionally, they were able to monitor the real-time changes in particle size and their relative abundances during the first hour of in vitro capsid assembly. In another capsid assembly study, Zhou et al.19 improved measurement precision via multicycle RPS in which each capsid was electrophoretically moved back and forth through four in-plane pores in series by alternating the polarity of the applied voltage. This allowed collecting ∼1000 individual RPS measurements, which lead to discriminating particles by size. They utilized the same method to investigate HBV capsid disassembly at varying concentrations of a chaotropic agent (guanidine hydrochloride).156 Furthermore, Li et al.157 employed four in-plane pores in series to determine that RNA length and structure, which effected capsid geometry during the assembly of Simian Virus 40 (SV40).
Plastic (organic) in-plane nanopores and their surface modification
Over the past several years, plastics, in particular thermoplastics, have been utilized to make nanopores offering significant advantages in terms of fabrication and engineering surface chemistry. For example, in-plane nanopores can be fabricated using low cost and high throughput replication techniques, such as nano-injection molding.158 Plastics are robust materials that can tolerate different temperatures, pH, and salt conditions. Additionally, plastics exhibit diverse surface chemistries displaying high susceptibility to various surface activation, modification, and immobilization techniques.159Thermoplastics, are a class of polymers that consist of long chains of covalently linked repeating units with the chains held together by weak intermolecular forces. They are characterized by their ability to repeatedly undergo a phase transition either into a malleable form (T > Tg; glass transition temperature) or a melt form (T > Tm; melting temperature) and reshaped without undergoing significant chemical degradation. This unique property, stemming from their molecular structure, makes them susceptible to various manufacturing processes even at the nanoscale. Each type of thermoplastic has unique chemical, mechanical, and thermal properties (see Fig. 8a), which are determined by the chemical structure of the thermoplastic (Fig. 8c–e). However, these properties can be systematically altered using a copolymer, such as COC, which consists of ethylene and norbornene monomer units. As seen in Fig. 8b, the Tg of COC depends heavily on the mole fraction of norbornene in the copolymer.
 | ||
| Fig. 8 (a) Chemical, mechanical and thermal properties of various thermoplastics. CTE – coefficient of thermal expansion. (b) The glass transition temperature (Tg) of the copolymer, COC, which consists of ethylene and norbornene units. The Tg depends on the mole fraction of norbornene in the copolymer. Chemical structures of different thermoplastics including PC (c), COC and COP (d) and PET (e). (f) Photo-oxidation of PMMA using either UV/O3 or O2 plasma to convert the methyl ester into surface-confined carboxylic acid groups. Reproduced from Chan et al.,160 with permission from Elsevier B.V. (g and h) Superresolution images of COC and PMMA, respectively, following UV/O3 photo-oxidation at different irradiation times. Reproduced from O'Neil et al.161 with permission from American Chemical Society. | ||
Thermoplastics can be modified upon exposure to UV/O3 or an O2 plasma, which induces a photo-oxidation reaction that alters the surface chemistry of the plastic (see Fig. 8f).160 The primary modification is the formation of oxygen-containing groups including surface carboxyl groups that affects surface charge and wettability of the plastic. Characterization of these activated surfaces has been demonstrated.162,163 The surface charge and wettability can be controlled by dosing the plastic with different irradiation times. For example, using superresolution microscopy (STORM), which provides sub-diffraction limited spatial resolution, the distribution of –COOH groups on COC and PMMA were determined (see Fig. 8g and h).161 As seen from these superresolution images, the surface charge density was dependent on the dose of the activating radiation. Fig. 8g shows increases in –COOH density as COC substrates were exposed from 1 to 15 min of UV/O3 radiation after which a decrease in –COOH density was observed. Different effects were observed for PMMA (Fig. 8h), where the –COOH surface density was significantly less than COC and showed an abrupt decrease after 1 min of UV/O3 irradiation.
In terms of nanopore sensing, plastics do offer some unique advantages including the ability to manufacture in-plane pores using replication-based techniques that can improve the widespread use of RPS label-free detection, dielectric properties that can reduce high frequency noise, and the ability to easily modify their surfaces. However, there are some challenges including their high surface hydrophobicity in their native state (water contact angle >90° in some cases) that requires surface activation to make the nanostructures they contain more hydrophilic. Additionally, their amorphous nature creates non-homogeneous surface charge densities following activation (see Fig. 8g and h) that can give rise to nanobubble formation as well as create surface ionizable groups both of which can increase the 1/f noise in the RPS measurement.
Thermoplastic in-plane nanopore sensors fabricated using replication technology
One of the advantages of using thermoplastics is their ability to be molded into mixed-scale structures (nm → mm) using replication technologies such as nanoimprint lithography (NIL), hot embossing, or injection molding.9,86,158,159,165 The advantage of these approaches for nanopore fabrication is that Si masters can be prepared with the appropriate topology and then used as a master to generate molding tools for any of the aforementioned production processes. For example, in the case of nanopore sensors, a fluidic network can be generated in the Si master using conventional photolithography followed by FIB milling to produce the necessary nanostructures (hybrid lithography). Then, the Si master can be used to make resin stamps for prototyping or Ni molding tools for high-scale production (see Fig. 9a–c). There are several advantages of this approach: (i) any substrate material (i.e., thermoplastic) can be used at the replication stage to suite the application need; (ii) different replication methods can be employed to produce the mixed-scale structures; (iii) no dependency on FIB or photolithography to make each device reducing cost and increasing production rate; (iv) ability to make mixed-scale structures to improve device figures-of-merit; and (v) integration of sample processing capabilities to the nanofluidic chip prior to making the RPS measurement. | ||
| Fig. 9 Diagram showing the general fabrication and assembly steps for making in-plane nanopore sensors in plastics. (a) Fabrication of the Si master mold, which utilized a combination of conventional photolithography to make the necessary microstructures followed by focused ion beam milling to produce the nanostructures, which included nanochannels and nanopores. (b) Fabrication of a resin stamp using UV-NIL with the Si master serving as the template. (c) Replication of the final device into a plastic using either thermal NIL or nano-injection molding. Reproduced from Shiri et al.158 with permission from The Royal Society of Chemistry. (d) Hybrid thermal fusion bonding of a low Tg cover plate to a higher Tg substrate containing the micro- and nanostructures. This step serves to enclose the fluidic channels and cover the in-plane nanopore. Reproduced from Uba et al.164 The Royal Society of Chemistry. The in-plane nanopores fabricated in the plastic could be reduced in size in a controllable manner during the thermal fusion bonding step, as shown in (e and f) in terms of the pore depth and overall cross-sectional area, respectively. (g) RPS peak amplitude for the translocation of λ-DNA (48.5 kbp) through plastic in-plane pores made in PMMA with a COC 8007 cover plate for three different devices assembled at different thermal fusion bonding pressures, 110, 170, and 200 psi. Reproduced from Athapattu et al.52 with permission from American Chemical Society. | ||
Nanoimprint lithography (NIL) has been used to achieve high quality nanoscale imprints.166–168 This instrument is capable of imprinting sub-10 nm structures by accurately controlling the heat and pressure during device fabrication.169 Recently, nano-injection molding has been reported to produce high quality nanostructures.158 Nano-injection molding of nanosensors was demonstrated using resin stamps as a prototyping tool.158 The primary advantage of injection molding over NIL is that the production rate is higher and at a lower unit cost.170
A challenge associated with making nanofluidic devices using top-down strategies as outlined above is the need for enclosing the fluidic network with a cover plate. While there are several methods to bond a plastic cover plate to the substrate containing the fluidic network, the approach adopted must not damage the underlying nanostructures. The choice of thermoplastic as the cover plate depends on the type of experiment being conducted. If detection and identification of biomolecules involve spectroscopic analyses where the light should penetrate into the device and excite the targets followed by microscopic detection of the reporters, it's crucial to select a cover plate that exhibits excellent optical transparency. However, this is not an issue for RPS measurements where only electrical signals are used to analyze the target.
To minimize nanostructure deformation, a hybrid thermal fusion bonding technique was reported (see Fig. 9d),164 which consists of using a Tg cover plate thermal fusion bonded to a higher Tg substrate. This hybrid method has resulted in making nanopore sensor devices with a process yield rate >90%.
Recently, Athapattu et al. reported a technique to tailor the size of in-plane plastic nanopores as part of the thermal fusion bonding step;52 the goal was to increase the SNR for the RPS by reducing the size of in-plane nanopores made via replication. This was accomplished by increasing the bonding pressure while holding the temperature constant at 70 °C (i.e., close to the Tg of a COC cover plate but below the Tg of the substrate).52 Increasing the pressure from 110 to 200 psi led to a reduction in the apparent diameter of the nanopore from 22.3 ± 1.4 nm to 10.2 ± 1.5 nm (inter-device standard deviations), an approximate 60% reduction (see Fig. 9e and f). The low standard deviations in these values indicated the high reproducibility of the fabrication method as well as the pore shrinkage process during thermal fusion bonding.
To demonstrate the ability to improve the SNR of the RPS measurements using this pore shrinking method, double-stranded DNA (λ-DNA) was electrically translocated through in-plane plastic nanopores (see Fig. 9g). The average peak amplitude of λ-DNA RPS events for devices bonded at 110 psi was 130 pA for λ-DNA corresponding to a pore depth of ∼22 nm as measured from conductance plots of 1 M KCl. Devices bonded at 170 psi had a pore depth of ∼13 nm and generated a RPS amplitude of 280 pA for λ-DNA and devices bonded at 200 psi possessed a depth of ∼10 nm and yielded a RPS amplitude of 437 pA for λ-DNA.
Fig. 10 shows the results for an injection molded plastic nanofluidic device containing in-plane nanopore sensors integrated within a nanofluidic circuit allowing on-chip sample processing prior to performing the RPS measurement.158Fig. 10a displays the nanofluidic network fabricated in Si using both photolithography to define microstructures and FIB milling to construct the necessary nanostructures.158 The device structural dimensions ranged from ∼10 μm (microchannels) down to ∼30 nm (in-plane pores). AFM images of a solid-phase bioreactor (Fig. 10b) and an in-plane nanopore (Fig. 10c) are shown.158 The effective diameter of the in-plane nanopores in the assembled nanofluidic device, estimated using open pore current measurements, was ∼12 nm.83,158 Assembled COP/COC dual in-plane nanopore devices were tested with a rCMP solution (10 nM) in 1× NEBuffer 3 (pH 7.9) under an applied voltage of 2.5 V and RPS detection was carried out. Fig. 10d–f shows representative RPS data along with histograms of td and ΔI/I0.
 | ||
| Fig. 10 (a) Si master mold for preparing resin stamps for nano-injection molding. The nanofluidic circuit contains several functional components to allow for inputting single molecules into an immobilized nanoscale enzymatic reactor (INER) and monitoring products generated using the products in real-time using a dual in-plane nanopore sensor equipped with a ToF nanotube and two in-plane pores. (b) Atomic force microscopy (AFM) image of the INER. (c) AFM image of an in-plane pore (input pore; Fig. 5a). Both AFM images are of the nano-injection molded structures. (d) RPS data for a 100 nM solution of rCMPs in 1× NEBuffer 3 using the dual in-plane nanopore sensor. (e and f) Histograms of the dwell time (td) and the normalized RPS peak amplitude (ΔI/I0), respectively. Reproduced from Shiri et al.158 with permission from The Royal Society of Chemistry. | ||
The current blockade amplitude (ΔI) for a single-pore system is described by eqn (5) and demonstrated that ΔI is influenced by multiple parameters including the applied voltage, salt (electrolyte) type and concentration, pore size, and analyte size. ΔI scales proportionally with the hydrodynamic cross-section of the analyte and inversely proportional to pore volume indicating that smaller pores and larger analytes yield more prominent current blockades. For a dual in-plane nanopore sensor, the current modulation is more complex than the single-pore system. For such systems, ΔI can be given by eqn (8), which accounts for both nanopores and their contributions to the overall current modulation;165
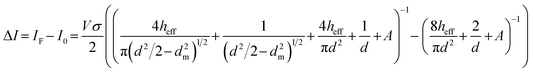 | (8) |
Thermoplastic nanopore sensors can be engineered to contain more than one sensing pore within a single device (see Fig. 10).168 The multi-pore format offers several advantages for not only single biomolecule detection, but can: (i) increase the accuracy of deducing RPS parameters;86,171 and (ii) inclusion of an RPS-independent parameter called the time-of-flight (ToF), which correlates with the μapp of the molecule.158,168 Because the ToF is independent of the RPS signal, a new set of parameters can be used to alter its value without affecting the RPS dependent variables such as td and ΔI/I0. The parameter space to alter the ToF is large because it is an electrophoretic and/or chromatographic effect that can be altered by changing the wall chemistry in the flight tube, the length of the flight tube, applied voltage, or carrier electrolyte.
Fig. 11 shows the comparison of noise characteristics and electric field distribution of a dual in-plane nanopore sensor with those of traditional out-of-plane nanopore sensors. There are notable differences between a single out-of-plane nanopore and the dual in-plane nanopore system, particularly in their power spectral density profiles (see Fig. 11a and b). At higher frequencies, the noise observed for the dual out-of-plane system is mainly caused by capacitive and dielectric effects, whereas the in-plane configuration has reduced high-frequency noise.56 The use of thick and highly insulating dielectric substrates, such as a plastic, can minimize thermal voltage noise arising from the pore material's parasitic capacitance and dielectric loss tangent. Lee et al. were able to generate low RMS noise levels at sub-pA in I0 for PET pores of ∼1.5 nm diameter.172
 | ||
| Fig. 11 Noise characteristics from different types of nanopores. (a) Power spectral density functions for three different out-of-plane nanopores made in a SiNx membrane (SSNP), PDMS (SSNP), and alpha hemolysin (biological NP). Taken from Tabard-Cossa et al.56 (b) Power spectral density of a COP/COP in-plane pore. Reproduced with permission from Shivanka et al.,165 copyright 2021 American Chemical Society. (c) Extension of electric field from an out-of-plane pore into the adjoining cis-chamber. Reproduced from Wanunu et al.47 with permission from Nature Nanotechnology. (d) COMSOL simulation showing extension of electric field across the pore traversing into the sampling microchannel. (e) Electric potential drop through a dual in-plane nanopore sensor, which is comprised of two in-plane pores in series and flanking a nanometer flight tube. (f) An in-plane nanopore showing the relative voltage drop at each longitudinal distance along the pore axis. Cs = capacitance of the polymer substrate (cross-hashed area; dot-dashed line shows capacitor plates, d = effective distance between plates), RNC = nanochannel flight tube resistance, and RMC = microchannel resistance. e and f Reproduced from Rathnayaka, et al.86 with permission from The Royal Society of Chemistry. | ||
Additional factors were proposed to explain the anomalous data shown in Fig. 11a and b: (i) the extended pore length; the in-plane pores are longer than biological pores like α-hemolysin resulting in longer dwell times. This phenomenon reduces the signal distortion during analyte translocation caused by BW limitations. (ii) The use of a low ionic strength carrier electrolyte. Typically, nanopore measurements are made in the presence of 1 M KCl; Smeets et al. reported lower ionic strength electrolytes (at low salt concentrations) can yield better RPS SNR for relatively larger pore sizes.173 (iii) The absence of surface protonation/deprotonation noise; plastic in-plane pores can be operated at a pH > pKa of the surface carboxyl groups (pKa ∼ 5.0), which can avoid protonation/deprotonation contributions to 1/f noise.174 (iv) The high mechanical stability; plastic in-plane pores offer greater mechanical stability compared to pores suspended on thin membranes that reduces 1/f noise.
The voltage drop for out-of-plane pores is almost exclusively across the pores while that is not the case for in-plane pores that are embedded within a nanofluidic network (Fig. 11c and d).47 This architecture allows in-plane pores to achieve higher sampling efficiency than their out-of-plane counterparts. This enhanced performance is due to the dual in-plane nanopore sensor placed within a mixed-scale fluidic network. For example, the dual in-plane nanopore sensor shown in Fig. 10 was able to detect as low as 3.9 fg of input rNMPs, while out-of-plane pores typically require about 1 μg of material to produce reasonable capture rates. This is predicated on the fact that the sampling zone (r*), where the electric field is sufficiently strong to electrokinetically drive molecules into the nanopore. The size of this zone can be calculated from eqn (9);47
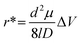 | (9) |
Interestingly, despite the larger pore volume of the sensing in-plane pores relative to out-of-plane sensors, RPS detection of single nucleotides was successfully demonstrated using plastic in-plane pores while for an out-of-plane sensor required α-hemolysin pores modified with a β-cyclodextrin moiety to achieve similar detection efficiency. The in-of-plane pore volumes were 320 nm3vs. out-of-plane α-hemolysin pore volume that is ∼18.2 nm3.176 Considering only volume displacement effects (i.e., Coulter principle), the detection of single nucleotides should be challenging using the in-plane nanopore system. However, the mixed-scale fluidic circuit, which consisted of nanochannels, and two in-plane pores placed in series, introduces additional physical effects which contribute to the resultant RPS signals. For example, Lee et al. demonstrated that ΔI for single-molecule RPS events can be significantly enhanced using a guiding nanochannel in series with the sensing nanopore due to compartmental limitations on ion transport as well as the drag forces exerted on the translocating molecules induced by the EOF in the guiding nanochannel.177 Furthermore, as depicted in Fig. 11e and f, the electric field strength in the dual in-plane nanopores are larger than the flight tube, but there is still an appreciable electric field drop within the nanochannel flight tube. This helps to suppress diffusional drift of the analytes and thus, minimize the variation in the ToF values.86
Choi et al. developed three different dual in-plane nanopore devices using PEGDA, each containing flight tubes with lengths ranging from 0.5–5.0 μm and were designed to improve nucleotide identification by enhancing differences in the ToF values (Fig. 12a–c).9 SEM images of the nanochannel flight tubes incorporated into the dual in-plane nanopore sensor are shown in Fig. 12d–i. After translocating individual deoxynucleotide monophosphates (dNMPs) through each device, the study revealed that identification accuracy for the four dNMPs improved from 55 to 94% with increasing flight tube length. The reason behind this improvement was ascribed to the combined effects of electrophoresis and chromatography in the nanometer sized flight tube (nano-EC). Because chromatographic separation resolution is column length dependent, longer flight tubes provide better separation in the ToF values.9Fig. 12k displays three pairs of current transients resulting from a single dNMP passing through the dual in-plane nanopore sensor.
 | ||
| Fig. 12 Thermoplastic-based fluidic device. (a) Schematically drawn set-up image of dual in-plane nanopore sensor, which consists of two in-plane nanopores flanking a nanometer-scale flight tube to allow carrying out nanoscale electrochromatography from which the ToF is deduced. The data includes the traditional RPS parameters (event amplitude and dwell time) and the ToF parameter. (b) Assembled sensor made from PEGDA. (c) Fabrication protocol used to make the device, which consisted of using UV-NIL from a Si master. (d–f) SEMs of the Si master mold including the dual in-plane pores flanking the flight tube, flight tube, and in-plane pores. (g–i) SEMs of NIL imprinted device made from PEGDA including the entire sensor, flight tube, and in-plane pores. Panels (a–i and k) were reproduced with permission from Choi et al.,9 copyright 2021 Wiley-VCH GmbH. (j) Schematic of sensor showing the two phases of the measurement including translocation through both pores and then the electrokinetic motion through the nano-EC flight tube. Within the flight tube, electrochromatography occurs and yields molecular-dependent ToF values due to differences in μapp (apparent electrophoretic mobility). (k) RPS trace data for dCMP showing pairs of RPS events. (i) Histogram distributions of μapp for the canonical dNMPs and m-dCMP collected by PMMA/COC (PMMA as substrate and COC as cover plate) device containing 100 μm nanochannel. Reproduced with permission from O'Neil et al.,178 copyright 2018 Elsevier B.V. | ||
The nano-EC of nucleotides within a nanofluidic channel was investigated by O'Neil et al.178 using laser-induced fluorescence detection. They electrokinetically transported dye-labeled dNMPs through 100 μm long nanochannels fabricated in PMMA/COC thermoplastics. As observed in Fig. 12i, the dNMP nano-EC migration order of nucleotides was identical to the results obtained by Choi et al.,9 who used RPS detection. The surface charges generated due to chemical groups (e.g., carboxyl groups) on plastic walls either by O2 plasma or UV/O3 activation can influence ion migration, especially because diffusional distances are smaller in nanometer columns compared to microscale columns; thus, nano-EC effects are predominate (Fig. 12j).30 Additionally, surface activation increases the hydrophilicity and thus the wettability of the plastic surface, which is crucial for maintaining consistent EOF.179 When an electric potential is applied across a nanochannel, the diffuse layer near the surface charges drags the bulk fluid towards the cathode generating the EOF.180 The net μapp of the analyte is a combination of the electrophoretic mobility (μep) and the mobility due to EOF (μeof), which is further modified by solute–wall interactions.181
Altering O2 plasma or UV/O3 activation time affects μeof due to increases in the surface charge density within the nanochannel and can also act as a functional scaffold for surface modification using standard coupling chemistries. For example, Uba et al. chemically modified the surface of activated PMMA/COC nanochannels with ethylenediamine to the surface carboxy groups using EDC/NHS coupling chemistry.182 As a result, an inversion in the EOF was seen; 1.02 ± 0.017 × 10−4 cm2 V−1 s−1 for the carboxylated surface to −0.75 ± 0.021 × 10−4 cm2 V−1 s−1 for the diamine surface.182
Other molecule types can be detected and identified as well using the dual in-plane nanopore sensor. For example, Fig. 13 shows data using the dual in-plane nanopore sensor for detecting and identifying a series of peptides with the use of both the RPS and ToF parameters along with machine learning.183Fig. 13a–c display current transient signals collected at time intervals of 200 ms, 50 ms and 100 μs.183 These signals were generated by the peptide metenkephalin electrokinetically shuttled through the sensor using 2.5 V. Here, the dual in-plane nanopore chip was made from PMMA and contained a COC cover plate. The red dashed line represents the amplitude threshold used by the authors, which was set to keep the false positive rate to 0 in the blank. Inspection of Fig. 13d shows the apparent mobilities of four peptides (metenkephalin, bradykinin, C natriuretic and C 3-33 peptides) as they were electrokinetically transported through a microchannel, nanochannel, and nanopore.183 The authors observed that as the channel size was reduced, μapp became smaller due to increased wall interactions.183 Smaller column lengths also reduced the differences in the ToF values. Although, the peptides did display differences in their ToF values (Fig. 13e), significant cluster overlap was observed when peptides were compared solely by their dwell times (Fig. 13f) and ΔI/I0 (Fig. 13g).183 Therefore, the authors implemented a neural network machine learning algorithm to improve the identification accuracy. Specifically, they used five input values (dwell time 1 and 2, ΔI/I0 1 and 2 and ToF) to train the model and performed a validation test to determine the classification accuracy (Fig. 13h). As a result, they acquired 95.6% and 100% accuracy during training and validation tests, respectively.
 | ||
| Fig. 13 RPS data results acquired using series of peptides using dual in-plane nanopore sensor. (a) Current trace containing RPS events acquired by translocating metenkephalin peptide. (b) Paired events showing ToF values of the metenkephalin peptide. (c) Single event showing ΔI/I0 and peak dwell time. (d) The μapp of four peptides measured specifically in the regions of microchannel, nanochannel and nanopore. Violin plot showing the distribution of ToF (e), dwell time (f), and ΔI/I0 (g). Neural network machine learning analysis of four peptides (h). Reproduced with permission from Chibuike et al.,183 copyright 2025 American Chemical Society. | ||
Comparison between in-plane versus out-of-plane nanopore sensors
Table 3 lists the performance metrics for both in-plane and out-of-plane nanopore sensors. Generally speaking there are advantages and disadvantages for both nanosensor platforms, but irrespective of configuration both offer single-molecule detection capabilities in a label-free manner and with the signal intensity inversely dependent on the size of the pore with respect to the target analyzed. The in-plane pores can easily be fine-tune in terms of the pore size and shape during ion beam milling or electron beam lithography. In addition, both in-plane and out-of-plane nanopore sensors can be made from organic or in-organic substrates. However, the out-of-plane pores can use biological nanopores while this becomes difficult for the in-plane format.| Single molecule sensing by RPS | |||
|---|---|---|---|
| LOD | <1 amol due to high e-field extension into sampling microchannel | >10 nmol due to abrupt size reduction (sampling reservoir to nanopore) | In-planes allow the use of gradient structure transitions while out-of-plane use a step transition |
| Bandwidth | High signal bandwidth 73/130 MHz (COP/PMMA) | Good signal bandwidth 16/32 MHz (SiN/PET) | In-plane have lower surface area capacitor plates |
| Signal integrity | Highly reproducible – depends on fabrication reproducibility | Highly reproducible for biological and modest for SSNP | For in-plane SSNP, milling conditions and pore closing are challenging |
| Operational time | Long if cover plate remains sealed to substrate | Modest due to instability of lipid bilayer for biological pores | Will see baseline drift (i.e., changes in I0) over time for both systems |
| Noise characteristics | Low noise at high frequency due to low capacitance and dielectric of plastics | High noise at high frequency due to capacitance and dielectric considerations | Material selection and small capacitor plates for in-plane pores; out-of-plane pores are suspended within membranes |
| Multi-detection | Very simple due to planar nature of pore and fluidics | Challenging due to the non-transparency of membrane and alignment issues with vertical pore | Optical alignment issues become less problematic for in-plane pores, but must be careful with material selection (i.e., optical transparency) |
| Manufacturing | |||
|---|---|---|---|
| Multiple pores (series or in-parallel) | Many pores can be placed in series or parallel (lithography dependent) | Pores in series determined by layer-by-layer fabrication; pores in parallel as well | In-plane pores can produce many (>8) in series or even in-parallel, but limited for out-of-plane pores for in-series |
| Material selection | High – organics (polymers), inorganics (Si-based) | High – biological, inorganic (SiN, SiO2, Al2O3), 2D materials (hBN, MoS2), organic (track-etched polymers) | No reported papers on using biological pores for in-plane configuration |
| A variety of polymers are available for in-plane configuration | |||
| Surface modification | Activating the substrate using UV/O3 for plastics followed by covalent modifications | Site-directed mutagenesis for biological pores and UV/O3 for plastics | Plastic in-plane and out-of-plane pores can undergo similar activation and modification strategies |
| Manufacturing scalability | High-scale production at low cost when using thermoplastic as substrate | Low-scale production at high cost; CMOS production line can increase production rate | Top-down fabrication for in-plane pores and ion/electron beam milling for out-of-plane pores |
| Integration with sample pre-processing | Simple due to chip-format of in-plane pores | Difficult due to non-chip format of out-of-plane pores | Microchip-to-nanochip interfaces simple for in-plane pores |
| Multiple pores (series or in-parallel) | Many pores can be placed in series or parallel (lithography dependent) | Pores in series determined by layer-by-layer fabrication; pores in parallel as well | In-plane pores can produce many (>8) in series or even in-parallel, but limited for out-of-plane pores for in-series |
In terms of noise characteristics, the in-plane pores can display higher mechanical stability compared to out-of-plane nanopore sensors, which are embedded within thin membranes. Also, capacitive noise for the in-plane pores is lower because the capacitor plates are smaller for these pore configurations. When the in-plane or even out-of-plane pores are made in a thermoplastic, this can make the dielectric noise smaller compared to SiNx membranes.
When seeking to build multiple pores in series, this becomes a simple and more flexible process for in-plane pores compared to out-of-plane pores because the fluidic circuit can be laid out on a single plane and then fabricated using, for example, ion beam milling. In addition, the spacing between the pores is easily adjusted to allow for securing molecular characteristics of the target through the apparent electrophoretic mobility that can aid in molecule identification.
The particularly attractive operational characteristic of the in-plane pores is their ability to provide high sampling efficiency that can generate favorable mass LODs. This results from the size gradients in pore/channel dimensions that can produce high electric fields extending into micro-sampling channels as opposed to relatively large sample reservoirs that flank either side of a membrane containing the nanopore as is the case for out-of-plane nanopore sensors.
Conclusions
Nanopore-based systems coupled with RPS have emerged as a powerful technique for the detection and identification of single molecules across a wide range of applications due to their ability to operate in a label-free manner and requiring rather simple instrumentation. Biologically derived nanopores such as α-hemolysin or MspA have significantly contributed to the progress of single molecule RPS sensing, but integrating them into broad user communities with long operational times is challenged due to the fragile nature of the lipid bilayer they are suspended in. This limitation has spawned the renaissance of SSNPs, which offer greater mechanical stability, improved fabrication versatility and broader flexibility in material selection, ranging from inorganic materials (Si and glass) to organic polymers (PET). In many cases, the nanopore systems employ a single pore configured in an out-of-plane format, where the analyte translocation is orthogonal with respect to the plane of the sensor. In these configurations, molecular identification primarily relies on conventional RPS parameters, namely the normalized current drop (ΔI/I0) and dwell time (td). While attractive results have emanated from this approach, for example in the case of DNA/RNA sequencing, emerging applications increasingly demand more complex and information-rich datasets requiring additional parameters for single molecule identification. This is particularly true for the case of epitranscriptomic RNA modifications (>170 different modifications), and post-translational modifications (PTMs) in proteins, where subtle structural variations are challenging to resolve only with RPS parameters. Another limitation of out-of-plane nanopore sensors are their low mass loading efficiency. While amplification strategies can be envisioned to improve the sensitivity of nucleic acid detection, the amplification process can potentially mask epigenetic and epitranscriptomic modifications. The absence of amplification strategies for proteins further limits the detection sensitivity and characterization.In-plane RPS nanopore sensors have recently emerged as a promising advancement. This platform offers double-sensing, where two spatially separated nanopores are connected via an intervening nanochannel. This geometry not only provides single-molecule detection in a label-free manner, but also facilitates ToF analysis, offering an additional single-molecule discriminatory parameter. The ToF is drawn from a combination of electrophoresis and chromatography with each molecule possessing a unique ToF value depending on its size and charge as well as its propensity to interact with the nanochannel wall.
Although glass-based in-plane nanopore sensors have been used to detect and identify a range of single molecule types, their low-throughput production and labor-intensive fabrication processes remain a challenge. In contrast, thermoplastic-based nanopore systems address these limitations by enabling low-cost and scalable fabrication, flexibility in device material selection, and highly sensitive RPS measurements. In addition, simple activation and surface modification strategies are available to further expand the application portfolio for these sensors. These advancements make thermoplastic nanopores a compelling alternative for next-generation molecular diagnostics and bioanalytical applications.
A main challenge with nanopore sensors is their wide-scale dissemination is their fabrication issues. Production of sub-100 nm or even sub-nm dimensions for sensing small objects such as the amino acids,127 require a combination of conventional photolithography and electron beam lithography or ion beam milling to make solid-state nanopores (in-plane or out-of-plane). While biological nanopores can ameliorate this issue, other pendant problems arise including mechanical stability and the limited operational lifetime of the sensor due to issues with the supporting lipid bilayer.97
Possible venues for increasing the access of academic labs to SSNPs (in-plane or out-of-plane) for research is through commercialization, soliciting support for small-scale production from foundries, or access to in-house fabrication resources. Shown in Table 4 is a cost-of-good SNR.
| Step | Lithography & FIBa | CMOSb | Injection molding | ||
|---|---|---|---|---|---|
| Cost/chip ($) | Cost/chip ($) | Step | |||
| Nano | Nano | Micro | Nano | ||
a Costs for lithography/FIB steps taken from typical user fees for various foundries.
b Same processing steps as for (1), but production scaleup included.
c Tooling for producing a Ni mold insert using photolithography only for micromolding and photolithography and FIB for nano-molding.
d Amortization for 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 chips.
e Includes equipment usage ($0.40 per chip) and labor ($0.10 per chip).
f Labor for cleaning chip.
g Includes labor, cover plate material, equipment amortization for fusion bonding.
h Structure replication used as limiting step.
i Yield rate set by success of fusion bonding. 000 chips.
e Includes equipment usage ($0.40 per chip) and labor ($0.10 per chip).
f Labor for cleaning chip.
g Includes labor, cover plate material, equipment amortization for fusion bonding.
h Structure replication used as limiting step.
i Yield rate set by success of fusion bonding.
|
|||||
| Optical mask | $0.10 (4″–8 per wafer) | 0.05 | 0.50 | Mold insert | |
| Photolithography | 2.0 | 0.50 | 0.50 | 0.50 | Molding |
| Etching (DRIE) | 10 | 0.50 | 0.50 | 0.50 | Post-processing |
| Material | Silicon | 0.25 | 0.25 | Polymer | |
| Sputter coating, FIB milling, and assembly | 75 | 0.75 | 0.75 | 0.75 | UV exposure and assembly |
| Chemicals | 0.75 | 0.25 | 0.25 | 0.25 | Chemicals |
| Labor | 2.25 | 0.75 | 0.75 | 0.75 | Labor |
| Total cost/chip | 90.10 | 87.10 | 3.05 | 3.50 | Total cost/chip |
| Production rate | 8 per day | 1000 per day | 960 per day | Production rate | |
| Yield rate | 80% | 70% | 100% | 90% | Yield rate , |
Conflicts of interest
The authors declare no competing interests.Abbreviations
| A/D/(ADC) | Converter analog-to-digital converter |
| AFM | Atomic force microscopy |
| BW | Bandwidth (range of frequencies the modulated signal occupies) |
| COP | Cyclic olefin polymer |
| COC | Cyclic olefin copolymer |
| CTE | Coefficient of thermal expansion |
| DNA | Deoxyribonucleic acid |
| ssDNA | Single stranded DNA |
| dsDNA | Double stranded DNA |
| λ D | Debye length (λD) measure of the electric double layer thickness |
| EDL | Electrical double layer |
| EOF | Electroosmotic flow |
| EBL | Electron beam lithography |
| PTMs | Post-translational modifications |
| EV | Extracellular vesicles |
| EDC/NHS | 1-Ethyl-3-(3-dimethylaminopropyl)carbodiimide/N-hydroxysuccinimide |
| FIB | Focused ion beam |
| GNR | Graphene nanoribbon |
| HBV | Hepatitis B virus |
| ICR | Ion current rectification |
| HA | Hyaluronic acid |
| ICP | Ion concentration polarization |
| I–V curve | Plot of current response to applied voltage |
| LPF | Low-pass filter |
| MspA porin | Mycobacterial porins (cell membrane pore formed by beta-barrel protein of Mycobacterium smegmatis) |
| WT | Wild type (gene or phenotype as it occurs in nature) |
| NIL | Nanoimprint lithography (method of fabrication of nanometer scale that uses a mold for patterning of structures under controlled pressure and temperature) |
| NMPs | Nucleoside monophosphate |
| dNMPs | Deoxyribonucleoside monophosphate (building blocks of DNA) |
| rNMPs | Ribonucleoside monophosphates (building blocks of RNA) physical mold to stamp a pattern onto a material) |
| PC | Polycarbonate |
| PDMS | Polydimethylsiloxane |
| PET | Polyethylene terephthalate |
| PMMA | Poly(methyl methacrylate) |
| RNA | Ribonucleic acid |
| RPS | Resistive pulse sensing |
| SNR | Signal-to-noise ratio |
| SSNPs | Solid-state nanopores |
| STORM | Stochastic Optical Reconstruction Microscopy |
| TEM | Transmission electron microscopy |
| T g | Glass transition temperature (temperature where a material transition between glassy/rigid to soft/soft/leathery state) |
| T m | Melting temperature (temperature at which one half of a complex is disrupted, is a measure of thermal stability) |
| ToF | Time-of-flight (in the context of nanopore sensors, the term is used to define the time between the translocation of pore 1 and pore 2 in dual in-plane nanopore systems) |
| UV/O3 | Ultraviolet light/ozone treatment (process based on ultraviolet light used to generate ozone molecules, which then oxidizes and cleans surfaces by breaking down organic contaminants or to generate oxygen functional groups in material surfaces such as polymers) |
| Zeta (ζ) potential | Defined as the electric potential at the shear surface of solid surfaces and colloidal particles at the particle–liquid interface |
| α-HL | Staphylococcal α-hemolysin (α-HL) a β-barrel pore-forming toxin |
| ΔI | Change in ionic current |
| I 0 | Baseline ionic current (commonly referred as background current or open pore current) |
| t d | Dwell time (residence time of an analyte inside or interacting with the nanopore) |
| μ ep | Electrophoretic mobility (movement of the charge particles under applied electric field) |
| μ eof | Electroosmotic mobility (movement of the bulk solution) |
| μ app | Apparent mobility which is the sum of electroosmotic mobility and electrophoretic mobility |
| η | Viscosity of buffer solution |
| ε o | Permittivity of free space (F m−1) |
| ε r | Relative permittivity |
| ΔV | Voltage drop |
| Cis side | In nanopore sensor context refers to the side of the membrane where the analyte is added |
| Trans side | Opposite side of the membrane pore where the analyte translocates to |
Data availability
All data associated with this manuscript is detailed in the form of figures and tables included in the manuscript. The data presented in all figures and tables can be made available to the readership upon request to the corresponding author of this manuscript.References
- W. H. Coulter, US Pat., 2656508, 1953 Search PubMed.
- M. Don, J. Lab. Autom., 2016, 8, 72–81 CrossRef.
- R. W. DeBlois and C. P. Bean, Rev. Sci. Instrum., 1970, 41, 909–916 Search PubMed.
- J. Kasianowicz, B. Walker, M. Krishnasastry and H. Bayley, MRS Online Proc. Libr., 1993, 330, 217–223 CrossRef.
- J. J. Kasianowicz, E. Brandin, D. Branton and D. W. Deamer, Proc. Natl. Acad. Sci. U. S. A., 1996, 93, 13770–13773 Search PubMed.
- H. Bayley and P. S. Cremer, Nature, 2001, 413, 226–230 Search PubMed.
- D. Deamer, M. Akeson and D. Branton, Nat. Biotechnol., 2016, 34, 518–524 CrossRef CAS PubMed.
- R. B. Schoch, J. Han and P. Renaud, Rev. Mod. Phys., 2008, 80, 839–883 CrossRef CAS.
- J. Choi, Z. Jia, R. Riahipour, C. J. McKinney, C. A. Amarasekara, K. M. Weerakoon-Ratnayake, S. A. Soper and S. Park, Small, 2021, 17, e2102567 CrossRef PubMed.
- J. Wang, M. Y. Li, J. Yang, Y. Q. Wang, X. Y. Wu, J. Huang, Y. L. Ying and Y. T. Long, ACS Cent. Sci., 2020, 6, 76–82 CrossRef CAS PubMed.
- J. Houghtaling, C. Ying, O. M. Eggenberger, A. Fennouri, S. Nandivada, M. Acharjee, J. Li, A. R. Hall and M. Mayer, ACS Nano, 2019, 13, 5231–5242 CrossRef CAS PubMed.
- C. Plesa, S. W. Kowalczyk, R. Zinsmeester, A. Y. Grosberg, Y. Rabin and C. Dekker, Nano Lett., 2013, 13, 658–663 CrossRef CAS PubMed.
- M. O'Donohue, J. Saharia, N. Bandara, G. Alexandrakis and M. J. Kim, Electrophoresis, 2023, 44, 349–359 CrossRef PubMed.
- J. Saharia, Y. Bandara, G. Goyal, J. S. Lee, B. I. Karawdeniya and M. J. Kim, ACS Nano, 2019, 13, 4246–4254 CrossRef CAS PubMed.
- J. Saharia, Y. Bandara, B. I. Karawdeniya, C. Hammond, G. Alexandrakis and M. J. Kim, RSC Adv., 2021, 11, 24398–24409 RSC.
- F. Rivas, P. L. DeAngelis, E. Rahbar and A. R. Hall, Sci. Rep., 2022, 12, 4469 CrossRef CAS PubMed.
- S. Vaidyanathan, H. Wijerathne, S. S. T. Gamage, F. Shiri, Z. Zhao, J. Choi, S. Park, M. A. Witek, C. McKinney, M. Verber, A. R. Hall, K. Childers, T. McNickle, S. Mog, E. Yeh, A. K. Godwin and S. A. Soper, Anal. Chem., 2023, 95, 9892–9900 CrossRef CAS PubMed.
- Z. D. Harms, D. G. Haywood, A. R. Kneller, L. Selzer, A. Zlotnick and S. C. Jacobson, Anal. Chem., 2015, 87, 699–705 CrossRef CAS PubMed.
- J. Zhou, P. Kondylis, D. G. Haywood, Z. D. Harms, L. S. Lee, A. Zlotnick and S. C. Jacobson, Anal. Chem., 2018, 90, 7267–7274 CrossRef CAS PubMed.
- A. Arima, M. Tsutsui, I. H. Harlisa, T. Yoshida, M. Tanaka, K. Yokota, W. Tonomura, M. Taniguchi, M. Okochi, T. Washio and T. Kawai, Sci. Rep., 2018, 8, 16305 CrossRef PubMed.
- B. I. Karawdeniya, Y. Bandara, A. I. Khan, W. T. Chen, H. A. Vu, A. Morshed, J. Suh, P. Dutta and M. J. Kim, Nanoscale, 2020, 12, 23721–23731 RSC.
- M. L. Pleet, S. Cook, V. A. Tang, E. Stack, V. J. Ford, J. Lannigan, N. Do, E. Wenger, J. L. Fraikin, S. Jacobson, J. C. Jones and J. A. Welsh, Nano Lett., 2023, 23, 9195–9202 CrossRef PubMed.
- T. W. Young, M. P. Kappler, N. M. Hockaden, R. L. Carpenter and S. C. Jacobson, Anal. Chem., 2023, 95, 16710–16716 Search PubMed.
- Z. Zhao, S. Vaidyanathan, P. Bhanja, S. Gamage, S. Saha, C. McKinney, J. Choi, S. Park, T. Pahattuge, H. Wijerathne, J. M. Jackson, M. L. Huppert, M. A. Witek and S. A. Soper, Electroanalysis, 2022, 34, 1961–1975 Search PubMed.
- R. W. DeBlois, C. P. Bean and R. K. A. Wesley, J. Colloid Interface Sci., 1977, 61, 323–335 CrossRef CAS.
- R. W. DeBlois and R. K. Wesley, J. Virol., 1977, 23, 227–233 Search PubMed.
- P. Kondylis, C. J. Schlicksup, N. E. Brunk, J. Zhou, A. Zlotnick and S. C. Jacobson, J. Am. Chem. Soc., 2019, 141, 1251–1260 Search PubMed.
- A. Darvish, G. Goyal, R. Aneja, R. V. Sundaram, K. Lee, C. W. Ahn, K. B. Kim, P. M. Vlahovska and M. J. Kim, Nanoscale, 2016, 8, 14420–14431 Search PubMed.
- J. A. Eldridge, G. R. Willmott, W. Anderson and R. Vogel, J. Colloid Interface Sci., 2014, 429, 45–52 Search PubMed.
- C. Rathnayaka, C. A. Amarasekara, K. Akabirov, M. C. Murphy, S. Park, M. A. Witek and S. A. Soper, J. Chromatogr. A, 2022, 1683, 463539 CrossRef CAS PubMed.
- R. B. Schoch, H. Van Lintel and P. Renaud, Phys. Fluids, 2005, 17, 100604 CrossRef.
- A. Plecis, R. B. Schoch and P. Renaud, Nano Lett., 2005, 5, 1147–1155 CrossRef CAS PubMed.
- A. Holtzel and U. Tallarek, J. Sep. Sci., 2007, 30, 1398–1419 CrossRef PubMed.
- K. Zhou, J. M. Perry and S. C. Jacobson, Annu. Rev. Anal. Chem., 2011, 4, 321–341 Search PubMed.
- C. Wei, A. J. Bard and S. W. Feldberg, Anal. Chem., 1997, 69, 4627–4633 Search PubMed.
- Q. Pu, J. Yun, H. Temkin and S. Liu, Nano Lett., 2004, 4, 1099–1103 CrossRef CAS.
- M. L. Kovarik, K. Zhou and S. C. Jacobson, J. Phys. Chem. B, 2009, 113, 15960–15966 Search PubMed.
- Y. Zhou, X. Liao, J. Han, T. Chen and C. Wang, Chin. Chem. Lett., 2020, 31, 2414–2422 Search PubMed.
- K. Zhou, M. L. Kovarik and S. C. Jacobson, J. Am. Chem. Soc., 2008, 130, 8614–8616 Search PubMed.
- R. M. M. Smeets, U. F. Keyser, D. Krapf, M.-Y. Wu, N. H. Dekker and C. Dekker, Nano Lett., 2006, 6, 89–95 CrossRef CAS PubMed.
- G. Goyal, K. J. Freedman and M. J. Kim, Anal. Chem., 2013, 85, 8180–8187 CrossRef CAS PubMed.
- L. S. Lastra, Y. Bandara, M. Nguyen, N. Farajpour and K. J. Freedman, Nat. Commun., 2022, 13, 2186 Search PubMed.
- E. Weatherall and G. R. Willmott, J. Phys. Chem. B, 2015, 119, 5328–5335 CrossRef CAS PubMed.
- W.-J. Lan, C. Kubeil, J.-W. Xiong, A. Bund and H. S. White, J. Phys. Chem. C, 2014, 118, 2726–2734 CrossRef CAS.
- M. Wanunu, T. Dadosh, V. Ray, J. Jin, L. McReynolds and M. Drndic, Nat. Nanotechnol., 2010, 5, 807–814 CrossRef CAS PubMed.
- S. W. Kowalczyk, D. B. Wells, A. Aksimentiev and C. Dekker, Nano Lett., 2012, 12, 1038–1044 CrossRef CAS PubMed.
- M. Wanunu, W. Morrison, Y. Rabin, A. Y. Grosberg and A. Meller, Nat. Nanotechnol., 2010, 5, 160–165 CrossRef CAS PubMed.
- I. W. Leong, M. Tsutsui, K. Yokota and M. Taniguchi, Anal. Chem., 2021, 93, 16700–16708 Search PubMed.
- J. Menestrina, C. Yang, M. Schiel, I. Vlassiouk and Z. S. Siwy, J. Phys. Chem. C, 2014, 118, 2391–2398 CrossRef CAS.
- I. W. Leong, S. Kishimoto, M. Tsutsui and M. Taniguchi, iScience, 2022, 25, 105073 CrossRef CAS PubMed.
- P. Kondylis, J. Zhou, Z. D. Harms, A. R. Kneller, L. S. Lee, A. Zlotnick and S. C. Jacobson, Anal. Chem., 2017, 89, 4855–4862 CrossRef CAS PubMed.
- U. S. Athapattu, C. Rathnayaka, S. Vaidyanathan, S. S. T. Gamage, J. Choi, R. Riahipour, A. Manoharan, A. R. Hall, S. Park and S. A. Soper, ACS Sens., 2021, 6, 3133–3143 Search PubMed.
- J. Choi, C. C. Lee and S. Park, Microsyst. Nanoeng., 2019, 5, 12 CrossRef PubMed.
- J. D. Uram, K. Ke and M. Mayer, ACS Nano, 2008, 2, 857–872 CrossRef CAS PubMed.
- A. Fragasso, S. Schmid and C. Dekker, ACS Nano, 2020, 14, 1338–1349 CrossRef CAS PubMed.
- V. Tabard-Cossa, D. Trivedi, M. Wiggin, N. N. Jetha and A. Marziali, Nanotechnology, 2007, 18, 305505 CrossRef.
- R. M. M. Smeets, U. F. Keyser, N. H. Dekker and C. Dekker, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 417–421 CrossRef CAS PubMed.
- D. Pedone, M. Firnkes and U. Rant, Anal. Chem., 2009, 81, 9689–9694 CrossRef CAS PubMed.
- R. A. Levis and J. L. Rae, in Methods in Enzymology, Academic Press, 1998, vol. 293, pp. 218–266 Search PubMed.
- F. J. Sigworth, in Single-Channel Recording, ed. B. Sakmann and E. Neher, Springer US, Boston, MA, 1995, pp. 95–127, DOI:10.1007/978-1-4419-1229-9_4.
- D. Colquhoun and F. J. Sigworth, Single-Channel Recording, 1995, ch. 19, pp. 483–587, DOI:10.1007/978-1-4419-1229-9_19.
- F. N. Hooge, Phys. Lett. A, 1969, 29, 139–140 CrossRef.
- R. Sherman-Gold, The Axon Guide, Axon Instruments, Inc., 1993 Search PubMed.
- D. P. Hoogerheide, S. Garaj and J. A. Golovchenko, Phys. Rev. Lett., 2009, 102, 256804 Search PubMed.
- Y. M. Blanter and M. Büttiker, Phys. Rep., 2000, 336, 1–166 Search PubMed.
- J. K. Rosenstein, M. Wanunu, C. A. Merchant, M. Drndic and K. L. Shepard, Nat. Methods, 2012, 9, 487–492 CrossRef CAS PubMed.
- O. Patil, D. Manikandan and V. V. R. Nandigana, Mol. Simul., 2020, 46, 1011–1016 CrossRef CAS.
- Z.-X. Wei, Y.-L. Ying, M.-Y. Li, J. Yang, J.-L. Zhou, H.-F. Wang, B.-Y. Yan and Y.-T. Long, Anal. Chem., 2019, 91, 10033–10039 Search PubMed.
- S. Shekar, C.-C. Chien, A. Hartel, P. Ong, O. B. Clarke, A. Marks, M. Drndic and K. L. Shepard, Nano Lett., 2019, 19, 1090–1097 CrossRef CAS PubMed.
- C. Wen, D. Dematties and S.-L. Zhang, ACS Sens., 2021, 6, 3536–3555 CrossRef CAS PubMed.
- M. Tsutsui, T. Takaai, K. Yokota, T. Kawai and T. Washio, Small Methods, 2021, 5, 2100191 CrossRef CAS PubMed.
- S. Shekar, D. J. Niedzwiecki, C. C. Chien, P. Ong, D. A. Fleischer, J. Lin, J. K. Rosenstein, M. Drndic and K. L. Shepard, Nano Lett., 2016, 16, 4483–4489 CrossRef CAS PubMed.
- A. J. W. Hartel, P. Ong, I. Schroeder, M. H. Giese, S. Shekar, O. B. Clarke, R. Zalk, A. R. Marks, W. A. Hendrickson and K. L. Shepard, Proc. Natl. Acad. Sci. U. S. A., 2018, 115, E1789–E1798 CrossRef CAS PubMed.
- A. J. W. Hartel, S. Shekar, P. Ong, I. Schroeder, G. Thiel and K. L. Shepard, Anal. Chim. Acta, 2019, 1061, 13–27 CrossRef CAS PubMed.
- C. Rathnayaka, I. A. Chandrosoma, J. Choi, K. Childers, M. Chibuike, K. Akabirov, F. Shiri, A. R. Hall, M. Lee and C. McKinney, Lab Chip, 2024, 24, 2721–2735 RSC.
- H. Austerlitz, in Data Acquisition Techniques Using PCs (Second Edition), ed. H. Austerlitz, Academic Press, San Diego, 2003, pp. 51–77, DOI:10.1016/B978-012068377-2/50004-8.
- M. Davenport, K. Healy, M. Pevarnik, N. Teslich, S. Cabrini, A. P. Morrison, Z. S. Siwy and S. E. Létant, ACS Nano, 2012, 6, 8366–8380 Search PubMed.
- K. Zhou, L. Li, Z. Tan, A. Zlotnick and S. C. Jacobson, J. Am. Chem. Soc., 2011, 133, 1618–1621 CrossRef CAS PubMed.
- Y. Green, R. Eshel, S. Park and G. Yossifon, Nano Lett., 2016, 16, 2744–2748 CrossRef CAS PubMed.
- M. L. Kovarik and S. C. Jacobson, Anal. Chem., 2008, 80, 657–664 CrossRef CAS PubMed.
- A. Alizadeh, W. L. Hsu, M. Wang and H. Daiguji, Electrophoresis, 2021, 42, 834–868 CrossRef CAS PubMed.
- Y. Green, R. Abu-Rjal and R. Eshel, Phys. Rev. Appl., 2020, 14, 014075 Search PubMed.
- J. Wu, J. Choi, F. I. Uba, S. A. Soper and S. Park, Micro Nano Eng., 2023, 21, 100230 CrossRef CAS PubMed.
- Q. Chen and Z. Liu, Sensors, 2019, 19, 1886 CrossRef CAS PubMed.
- M. Zhang, Z. D. Harms, T. Greibe, C. A. Starr, A. Zlotnick and S. C. Jacobson, ACS Nano, 2022, 16, 7352–7360 CrossRef CAS PubMed.
- C. Rathnayaka, I. A. Chandrosoma, J. Choi, K. Childers, M. Chibuike, K. Akabirov, F. Shiri, A. R. Hall, M. Lee, C. McKinney, M. Verber, S. Park and S. A. Soper, Lab Chip, 2024, 24, 2721–2735 RSC.
- S. Akhtarian, S. Miri, A. Doostmohammadi, S. K. Brar and P. Rezai, Bioengineered, 2021, 12, 9189–9215 CrossRef CAS PubMed.
- Y. Tanaka, N. Hirano, J. Kaneko, Y. Kamio, M. Yao and I. Tanaka, Protein Sci., 2011, 20, 448–456 CrossRef CAS PubMed.
- Y.-L. Ying, C. Cao and Y.-T. Long, Analyst, 2014, 139, 3826–3835 RSC.
- M. Faller, M. Niederweis and G. E. Schulz, Science, 2004, 303, 1189–1192 CrossRef CAS PubMed.
- A. Guasch, J. Pous, B. Ibarra, F. X. Gomis-Rüth, J. M. a. Valpuesta, N. Sousa, J. L. Carrascosa and M. Coll, J. Mol. Biol., 2002, 315, 663–676 CrossRef CAS PubMed.
- H. Bayley, Clin. Chem., 2015, 61, 25–31 CrossRef CAS PubMed.
- I. M. Derrington, T. Z. Butler, M. D. Collins, E. Manrao, M. Pavlenok, M. Niederweis and J. H. Gundlach, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 16060–16065 CrossRef PubMed.
- D. Stoddart, A. J. Heron, E. Mikhailova, G. Maglia and H. Bayley, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 7702–7707 CrossRef CAS PubMed.
- J. Clarke, H.-C. Wu, L. Jayasinghe, A. Patel, S. Reid and H. Bayley, Nat. Nanotechnol., 2009, 4, 265–270 CrossRef CAS PubMed.
- A. Balijepalli, J. Ettedgui, A. T. Cornio, J. W. Robertson, K. P. Cheung, J. J. Kasianowicz and C. Vaz, ACS Nano, 2014, 8, 1547–1553 CrossRef CAS PubMed.
- T.-J. Jeon, N. Malmstadt and J. J. Schmidt, J. Am. Chem. Soc., 2006, 128, 42–43 CrossRef CAS PubMed.
- A. J. Storm, J. H. Chen, X. S. Ling, H. W. Zandbergen and C. Dekker, Nat. Mater., 2003, 2, 537–540 CrossRef CAS PubMed.
- J. D. Uram, K. Ke, A. J. Hunt and M. Mayer, Small, 2006, 2, 967–972 CrossRef CAS PubMed.
- K. Liu, J. Feng, A. Kis and A. Radenovic, ACS Nano, 2014, 8, 2504–2511 CrossRef CAS PubMed.
- S. Liu, B. Lu, Q. Zhao, J. Li, T. Gao, Y. Chen, Y. Zhang, Z. Liu, Z. Fan, F. Yang, L. You and D. Yu, Adv. Mater., 2013, 25, 4549–4554 CrossRef CAS PubMed.
- Y. Zhang, D. Ma, Z. Gu, L. Zhan and J. Sha, Nanomaterials, 2021, 11, 2450 CrossRef CAS PubMed.
- A. Darvish, J. S. Lee, B. Peng, J. Saharia, R. VenkatKalyana Sundaram, G. Goyal, N. Bandara, C. W. Ahn, J. Kim, P. Dutta, I. Chaiken and M. J. Kim, Electrophoresis, 2019, 40, 776–783 CrossRef CAS PubMed.
- A. J. Storm, J. H. Chen, X. S. Ling, H. W. Zandbergen and C. Dekker, Nat. Mater., 2003, 2, 537–540 CrossRef CAS PubMed.
- V. J. Gadgil, H. D. Tong, Y. Cesa and M. L. Bennink, Surf. Coat. Technol., 2009, 203, 2436–2441 CrossRef CAS.
- C. J. Lo, T. Aref and A. Bezryadin, Nanotechnology, 2006, 17, 3264 CrossRef CAS.
- J. Yang, D. C. Ferranti, L. A. Stern, C. A. Sanford, J. Huang, Z. Ren, L. C. Qin and A. R. Hall, Nanotechnology, 2011, 22, 285310 CrossRef PubMed.
- H. Kwok, K. Briggs and V. Tabard-Cossa, PLoS One, 2014, 9, e92880 CrossRef PubMed.
- M. Waugh, K. Briggs, D. Gunn, M. Gibeault, S. King, Q. Ingram, A. M. Jimenez, S. Berryman, D. Lomovtsev, L. Andrzejewski and V. Tabard-Cossa, Nat. Protoc., 2020, 15, 122–143 CrossRef CAS PubMed.
- J. P. Fried, J. L. Swett, B. P. Nadappuram, J. A. Mol, J. B. Edel, A. P. Ivanov and J. R. Yates, Chem. Soc. Rev., 2021, 50, 4974–4992 RSC.
- J. Li, D. Stein, C. McMullan, D. Branton, M. J. Aziz and J. A. Golovchenko, Nature, 2001, 412, 166–169 CrossRef CAS PubMed.
- H. Qiu, W. Zhou and W. Guo, ACS Nano, 2021, 15, 18848–18864 CrossRef CAS PubMed.
- H. D. Tong, H. V. Jansen, V. J. Gadgil, C. G. Bostan, E. Berenschot, C. J. van Rijn and M. Elwenspoek, Nano Lett., 2004, 4, 283–287 CrossRef CAS.
- M. D. Fischbein and M. Drndić, Nano Lett., 2007, 7, 1329–1337 CrossRef CAS PubMed.
- M. Wanunu, T. Dadosh, V. Ray, J. Jin, L. McReynolds and M. Drndić, Nat. Nanotechnol., 2010, 5, 807–814 CrossRef CAS PubMed.
- E. Zvuloni, A. Zrehen, T. Gilboa and A. Meller, ACS Nano, 2021, 15, 12189–12200 CrossRef CAS PubMed.
- S. Tan, L. Wang, J. Yu, C. Hou, R. Jiang, Y. Li and Q. Liu, Nanoscale Res. Lett., 2015, 10, 1–10 Search PubMed.
- S. Dutt, B. I. Karawdeniya, Y. N. D. Bandara, N. Afrin and P. Kluth, Anal. Chem., 2023, 95, 5754–5763 Search PubMed.
- Z. Xia, A. Scott, R. Keneipp, J. Chen, D. J. Niedzwiecki, B. DiPaolo and M. Drndic, ACS Nano, 2022, 16, 18648–18657 Search PubMed.
- R. Dela Torre, J. Larkin, A. Singer and A. Meller, Nanotechnology, 2012, 23, 385308 Search PubMed.
- E. Kennedy, Z. Dong, C. Tennant and G. Timp, Nat. Nanotechnol., 2016, 11, 968–976 CrossRef CAS PubMed.
- S. Liu, J. Tian and W. Zhang, Nanotechnology, 2021, 32, 222001 CrossRef CAS PubMed.
- B. M. Venkatesan, B. Dorvel, S. Yemenicioglu, N. Watkins, I. Petrov and R. Bashir, Adv. Mater., 2009, 21, 2771–2776 CrossRef CAS PubMed.
- B. M. Venkatesan, A. B. Shah, J. M. Zuo and R. Bashir, Adv. Funct. Mater., 2010, 20, 1266–1275 Search PubMed.
- Y. Feng, Y. Zhang, C. Ying, D. Wang and C. Du, Genomics, Proteomics Bioinf., 2015, 13, 4–16 CrossRef CAS PubMed.
- F. Traversi, C. Raillon, S. Benameur, K. Liu, S. Khlybov, M. Tosun, D. Krasnozhon, A. Kis and A. Radenovic, Nat. Nanotechnol., 2013, 8, 939–945 CrossRef CAS PubMed.
- F. Wang, C. Zhao, P. Zhao, F. Chen, D. Qiao and J. Feng, Nat. Commun., 2023, 14, 2895 CrossRef CAS PubMed.
- J. Feng, K. Liu, R. D. Bulushev, S. Khlybov, D. Dumcenco, A. Kis and A. Radenovic, Nat. Nanotechnol., 2015, 10, 1070–1076 CrossRef CAS PubMed.
- S. Awasthi, C. Ying, J. Li and M. Mayer, ACS Nano, 2023, 17, 12325–12335 CrossRef CAS PubMed.
- M. Graf, M. Lihter, M. Thakur, V. Georgiou, J. Topolancik, B. R. Ilic, K. Liu, J. Feng, Y. Astier and A. Radenovic, Nat. Protoc., 2019, 14, 1130–1168 CrossRef CAS PubMed.
- S. Garaj, S. Liu, J. A. Golovchenko and D. Branton, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 12192–12196 CrossRef CAS PubMed.
- H. Arjmandi-Tash, L. A. Belyaeva and G. F. Schneider, Chem. Soc. Rev., 2016, 45, 476–493 RSC.
- M.-H. Lee, A. Kumar, K.-B. Park, S.-Y. Cho, H.-M. Kim, M.-C. Lim, Y.-R. Kim and K.-B. Kim, Sci. Rep., 2014, 4, 7448 CrossRef PubMed.
- G. F. Schneider, Q. Xu, S. Hage, S. Luik, J. N. Spoor, S. Malladi, H. Zandbergen and C. Dekker, Nat. Commun., 2013, 4, 2619 CrossRef PubMed.
- S. Jangra, B. Kumar, J. Sharma, S. Sengupta, S. Das, R. Brajpuriya, A. Ohlan, Y. K. Mishra and M. Goyat, J. Energy Storage, 2024, 101, 113810 CrossRef.
- M. Mojtabavi, A. VahidMohammadi, W. Liang, M. Beidaghi and M. Wanunu, ACS Nano, 2019, 13, 3042–3053 CrossRef CAS PubMed.
- M. T. Hossaın, M. R. Repon, M. A. Shahid, A. Ali and T. Islam, ChemElectroChem, 2024, 11, e202400008 CrossRef.
- P. Apel, Radiat. Meas., 2001, 34, 559–566 CrossRef CAS.
- D. Kaya and K. Keçeci, J. Electrochem. Soc., 2020, 167, 037543 CrossRef CAS.
- Z. Siwy and A. Fuliński, Phys. Rev. Lett., 2002, 89, 198103 CrossRef CAS PubMed.
- J. Choi, C. C. Lee and S. Park, Microsyst. Nanoeng., 2019, 5, 12 CrossRef PubMed.
- F. Zhang, Y. Sun, D. Tian and H. Li, Angew. Chem., Int. Ed., 2017, 56, 7186–7190 CrossRef CAS PubMed.
- N. Meyer, J.-M. Janot, M. Lepoitevin, M. Smietana, J.-J. Vasseur, J. Torrent and S. Balme, Biosensors, 2020, 10, 140 CrossRef CAS PubMed.
- V. Thangaraj, M. Lepoitevin, M. Smietana, E. Balanzat, M. Bechelany, J.-M. Janot, J.-J. Vasseur, S. Subramanian and S. Balme, Microchim. Acta, 2016, 183, 1011–1017 CrossRef CAS.
- T. Ma, J.-M. Janot and S. Balme, Nanoscale, 2020, 12, 7231–7239 RSC.
- Y.-C. Chou, C.-Y. Lin, A. Castan, J. Chen, R. Keneipp, P. Yasini, D. Monos and M. Drndić, Nat. Nanotechnol., 2024, 19, 1686–1692 CrossRef CAS PubMed.
- M. Tsutsui, K. Yokota, A. Arima, T. Washio, Y. Baba and T. Kawai, Small Methods, 2021, 5, 2100542 CrossRef CAS PubMed.
- P. Cadinu, G. Campolo, S. Pud, W. Yang, J. B. Edel, C. Dekker and A. P. Ivanov, Nano Lett., 2018, 18, 2738–2745 CrossRef CAS PubMed.
- Z. D. Harms, K. B. Mogensen, P. S. Nunes, K. Zhou, B. W. Hildenbrand, I. Mitra, Z. Tan, A. Zlotnick, J. P. Kutter and S. C. Jacobson, Anal. Chem., 2011, 83, 9573–9578 CrossRef CAS PubMed.
- Z. D. Harms, D. G. Haywood, A. R. Kneller, L. Selzer, A. Zlotnick and S. C. Jacobson, Anal. Chem., 2015, 87, 699–705 CrossRef CAS PubMed.
- M. Zhang, Z. D. Harms, T. Greibe, C. A. Starr, A. Zlotnick and S. C. Jacobson, ACS Nano, 2022, 16, 7352–7360 CrossRef CAS PubMed.
- T. W. Young, M. P. Kappler, N. M. Hockaden, R. L. Carpenter and S. C. Jacobson, Anal. Chem., 2023, 95, 16710–16716 CrossRef CAS PubMed.
- R. M. Smeets, U. F. Keyser, D. Krapf, M.-Y. Wu, N. H. Dekker and C. Dekker, Nano Lett., 2006, 6, 89–95 CrossRef CAS PubMed.
- O. K. Zahid, F. Rivas, F. Wang, K. Sethi, K. Reiss, S. Bearden and A. R. Hall, Nanomed.: Nanotechnol., Biol. Med., 2021, 35, 102407 CrossRef CAS PubMed.
- Z. D. Harms, K. B. Mogensen, P. S. Nunes, K. Zhou, B. W. Hildenbrand, I. Mitra, Z. Tan, A. Zlotnick, J. P. Kutter and S. C. Jacobson, Anal. Chem., 2011, 83, 9573–9578 CrossRef CAS PubMed.
- J. Zhou, A. Zlotnick and S. C. Jacobson, Anal. Chem., 2022, 94, 985–992 CrossRef CAS PubMed.
- C. Li, A. R. Kneller, S. C. Jacobson and A. Zlotnick, ACS Chem. Biol., 2017, 12, 1327–1334 CrossRef CAS PubMed.
- F. Shiri, J. Choi, C. Vietz, C. Rathnayaka, A. Manoharan, S. Shivanka, G. Li, C. Yu, M. C. Murphy and S. A. Soper, Lab Chip, 2023, 23, 4876–4887 RSC.
- C. A. Amarasekara, U. S. Athapattu, C. Rathnayaka, J. Choi, S. Park and S. A. Soper, Electrophoresis, 2020, 41, 1627–1640 CrossRef CAS PubMed.
- C. M. Chan, T. M. Ko and H. Hiraoka, Surf. Sci. Rep., 1996, 24, 1–54 CrossRef CAS.
- C. E. O'Neil, J. M. Jackson, S. H. Shim and S. A. Soper, Anal. Chem., 2016, 88, 3686–3696 CrossRef PubMed.
- J. M. Jackson, M. A. Witek, M. L. Hupert, C. Brady, S. Pullagurla, J. Kamande, R. D. Aufforth, C. J. Tignanelli, R. J. Torphy, J. J. Yeh and S. A. Soper, Lab Chip, 2014, 14, 106–117 RSC.
- S. Wei, B. Vaidya, A. B. Patel, S. A. Soper and R. L. McCarley, J. Phys. Chem. B, 2005, 109, 16988–16996 CrossRef CAS PubMed.
- F. I. Uba, B. Hu, K. Weerakoon-Ratnayake, N. Oliver-Calixte and S. A. Soper, Lab Chip, 2015, 15, 1038–1049 RSC.
- S. Shivanka, F. Shiri, M. Chibuike, C. McKinney, M. Verber, J. Choi, S. Park, A. R. Hall and S. A. Soper, Sci. Rep., 2025, 15, 13742 CrossRef CAS PubMed.
- S. Vaidyanathan, K. M. Weerakoon-Ratnayake, F. I. Uba, B. Hu, D. Kaufman, J. Choi, S. Park and S. A. Soper, Lab Chip, 2021, 21, 1579–1589 RSC.
- C. A. Amarasekara, C. Rathnayaka, U. S. Athapattu, L. Zhang, J. Choi, S. Park, A. C. Nagel and S. A. Soper, J. Chromatogr. A, 2021, 1638, 461892 CrossRef CAS PubMed.
- J. Choi, Z. Jia, R. Riahipour, C. J. McKinney, C. A. Amarasekara, K. M. Weerakoon-Ratnayake, S. A. Soper and S. Park, Small, 2021, 17, 2102567 CrossRef CAS PubMed.
- K. M. Weerakoon-Ratnayake, F. I. Uba, N. J. Oliver-Calixte and S. A. Soper, Anal. Chem., 2016, 88, 3569–3577 CrossRef CAS PubMed.
- H. Becker, Lab Chip, 2009, 9, 2759–2762 RSC.
- S. Seth and A. Bhattacharya, Sci. Rep., 2021, 11, 9799 CrossRef CAS PubMed.
- M.-H. Lee, A. Kumar, K.-B. Park, S.-Y. Cho, H.-M. Kim, M.-C. Lim, Y.-R. Kim and K.-B. Kim, Sci. Rep., 2014, 4, 1–7 Search PubMed.
- R. M. M. Smeets, U. F. Keyser, N. H. Dekker and C. Dekker, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 417–421 CrossRef CAS PubMed.
- R. L. McCarley, B. Vaidya, S. Y. Wei, A. F. Smith, A. B. Patel, J. Feng, M. C. Murphy and S. A. Soper, J. Am. Chem. Soc., 2005, 127, 842–843 CrossRef CAS PubMed.
- M. Wanunu, W. Morrison, Y. Rabin, A. Y. Grosberg and A. Meller, Nat. Nanotechnol., 2010, 5, 160–165 CrossRef CAS PubMed.
- M. Ayub, S. W. Hardwick, B. F. Luisi and H. Bayley, Nano Lett., 2013, 13, 6144–6150 CrossRef CAS PubMed.
- K. Lee, H. Lee, S.-H. Lee, H.-M. Kim, K.-B. Kim and S. J. Kim, Nanoscale, 2017, 9, 18012–18021 RSC.
- C. O'Neil, C. A. Amarasekara, K. M. Weerakoon-Ratnayake, B. Gross, Z. Jia, V. Singh, S. Park and S. A. Soper, Anal. Chim. Acta, 2018, 1027, 67–75 CrossRef PubMed.
- K. M. Weerakoon-Ratnayake, C. E. O'Neil, F. I. Uba and S. A. Soper, Lab Chip, 2017, 17, 362–381 Search PubMed.
- G. W. Slater, F. Tessier and K. Kopecka, Microb. Biotechnol., 2010, 121–134 CAS.
- J. Zhong and X. Wang, Evaluation technologies for food quality, Woodhead Publishing, 2019 Search PubMed.
- F. I. Uba, S. R. Pullagurla, N. Sirasunthorn, J. Wu, S. Park, R. Chantiwas, Y.-K. Cho, H. Shin and S. A. Soper, Analyst, 2015, 140, 113–126 RSC.
- M. Chibuike, C. Rathnayaka, S. Shivanka, J. Choi, M. Verber, S. Park and S. A. Soper, Anal. Chem., 2024, 97, 427–435 Search PubMed.
Footnote |
| † These authors contributed equally to this manuscript. |
| This journal is © The Royal Society of Chemistry 2026 |
