Open Access Article
Open Access ArticleAn automated and portable platform for rapid cell-free DNA isolation and its application in microbial DNA metagenomic sequencing from human blood samples
Linda Marriotta,
Ana Martinez-Lopez ab,
Antonio Ligaa,
Kazuhiro Horiba
ab,
Antonio Ligaa,
Kazuhiro Horiba c,
Amanda Warrb,
Jacob N. Phulusad,
Radhe Shantha Kumar
c,
Amanda Warrb,
Jacob N. Phulusad,
Radhe Shantha Kumar e,
Laura Carey
e,
Laura Carey f,
Yoshinori Ito
f,
Yoshinori Ito g,
Benjamin J. Parcellh,
Nicholas R. Lesliea,
Nicholas A. Feasey
g,
Benjamin J. Parcellh,
Nicholas R. Lesliea,
Nicholas A. Feasey i,
Shevin T. Jacobf,
Jamie Rylance
i,
Shevin T. Jacobf,
Jamie Rylance f and
Maïwenn Kersaudy-Kerhoas
f and
Maïwenn Kersaudy-Kerhoas *a
*a
aSchool of Engineering, Heriot-Watt University, UK. E-mail: m.kersaudy-kerhoas@hw.ac.uk
bThe Roslin Institute, University of Edinburgh, UK
cPathogen Genomics Center, National Institute of Infectious Diseases, Japan
dMalawi-Liverpool Wellcome Clinical Research Programme, Kamuzu University of Health Sciences, Blantyre, Malawi
eQueens' college, University of Cambridge, UK
fDepartment of Clinical Sciences, Liverpool School of Tropical Medicine, UK
gDepartment of Pediatrics, Aichi Medical University, Japan
hSchool of Medicine, University of Dundee, UK
iSchool of Medicine, University of St Andrews, UK
First published on 9th March 2026
Abstract
The prompt identification of pathogens in human circulation in a clinically deployable format remains an unmet clinical need. The established test for infection diagnostics remains blood culture, which typically takes 2–4 days and is positive in less than 15% of cases, with many prevalent pathogens difficult or impossible to culture. While microbial cfDNA in blood could facilitate the diagnosis of sepsis, febrile and infectious conditions, sample preparation for cell-free DNA (cfDNA) analysis in decentralised settings presents challenges due to its complexity and the low concentration and fragmented nature of cfDNA in blood plasma. We developed a portable and automated platform and a consumable (CNASafe) for cfDNA isolation from human plasma samples. The platform-device performance was evaluated by comparing relative cfDNA yield against a reference (QIAGEN QIAamp Circulating Nucleic Acid Kit). cfDNA eluates from ten non-cultured blood samples from hospital patients were sequenced on a nanopore sequencer, and results compared to blood cultures. Extraction of cfDNA using the CNASafe device was completed in 40 minutes, compared to the 1 hour 15 min reference protocol. The device achieved an average relative cfDNA recovery of 100.5% over 333 unique extractions encompassing all parameter variations, demonstrating a performance equivalent to the reference kit. From the patient samples, a sufficient quantity of microbial cfDNA was extracted to either identify pathogens missed by blood cultures or confirm negative cultures. The CNASafe platform and real-time nanopore sequencing offer a promising solution for the rapid deployment of metagenomic diagnostics, enabling pathogen identification within a few hours in decentralised clinical environments.
Introduction
Over the past few decades, circulating nucleic acids (CNAs) in blood plasma have emerged as important biomarkers for several medical applications. CNAs have been particularly well-established in oncology diagnostics,1,2 serving as an attractive alternative to conventional tissue biopsy. Liquid biopsies are less invasive and enable repeated testing for longitudinal assessments. Liquid biopsy has also been identified as a valuable oncological predictive and prognostic tool,3 as well as an effective way to monitor treatment4 by detecting minimal residual disease and predicting recurrence.5 Beyond personalized oncology, the potential of cell-free DNA (cfDNA) has been demonstrated across various applications including, non-invasive prenatal testing through the detection of fetal cfDNA in maternal blood,6 organ transplantation monitoring via circulating donor DNA,7,8 and toxicology applications through circulating microRNA (miRNA) detection.9,10 More recently, cfDNA has been used to identify infectious pathogens through circulating microbial DNA detection.11–13In the latter case, microbial cfDNA enables the identification of disease-causing pathogens and provides accurate, rapid, and clinically actionable diagnosis of bloodstream infections. Bloodstream infections and sepsis lead to high mortality rates, prolonged hospitalizations, and pose a significant socio-economic burden globally.14 Current diagnostic procedures rely on blood cultures, which lack specificity and sensitivity, and often have a time-to-result exceeding 72 hours in clinical practice, typically after broad-spectrum antibiotics have been administered. The incorrect use of antibiotics has been associated with an increased risk of antimicrobial resistance (AMR) and mortality.15 Additionally, parasites and viruses cannot be cultured, and fungi require specific and often challenging culture conditions, limiting the utility of blood culture in infectious disease diagnosis. Protein biomarkers, such as procalcitonin (PCT) and C-reactive protein (CRP), are useful for establishing if there is an infection present but do not inform the presence of specific pathogens. PCR, including multiplex PCR panels can be effective, but have several limitations: their multiplex power is limited and, for a specific pathogen, PCR methods only probe one or a few selected genomic sites. Microbial cfDNA released from lysed pathogens in the infected human circulation has been shown to have high sensitivity for the detection of sepsis-causing pathogens, and its use could result in improvement of turnaround times (TATs) for results and improved antimicrobial stewardship.11 This is particularly valuable in conditions where the aetiology is difficult or impossible to determine through conventional methods such as blood cultures.
Whatever the clinical application considered, the hurdles are similar for liquid biopsies to reach their promise. Samples must be prepared consistently and in a way that maximises the nucleic acid yield while maintaining quality and avoiding background genomic DNA interference. CNAs are circulating in a complex matrix, comprising white blood cells, red blood cells, proteins, and metabolites, that would inhibit their detection if not removed. The current solution for isolating circulating cell-free nucleic acids from blood is a skilled, labour- and resource-intensive process. Most of the commercial methods for isolating CNAs from plasma are based on the properties of solid substrate, usually a silica membrane within a spin column to reversibly bind CNAs in presence of a binding buffer.16 QIAamp Circulating Nucleic Acid Kit (Qiagen) is a market reference. However, other commercial solid phase extraction methods include the use of magnetic beads as substrate for CNA adsorption,17 such as the MagMAX cfDNA Isolation Kit from Thermo Fisher Scientific, EZ1&2 ccfDNA Kit from Qiagen, and Maxwell® RSC ccfDNA Plasma Kit from Promega. Both filtration capture columns and magnetic silica beads rely on either a manual workflow or bulky, costly, and semi-automated platforms operated by skilled staff; these approaches require a centralised lab and are not compatible with blood processing in a decentralised manner.18
Microfluidics has emerged as a possible toolbox to miniaturise and facilitate some challenging steps of liquid biopsy analysis. The potential advantages of microfluidic platforms for point-of-care applications have been understood for many years and include full automation, self-contained reaction volumes, integration of multi-step processes, increased speed, portability, and ease of disposal.19 In the context of CNAs, some researchers focused on improving the substrate for nucleic acids adsorption.20–23 While most microfluidic applications and above examples are only suitable for small volume (microlitres) samples,24 larger volume applications have been developed as well, typically approaching cfDNA recovery similar to manual or robotic magnetic bead kits, on 0.5 to under 4 mL of plasma in 15 min to 2 hours, but never combining all of optimised features in a single device, and never compared to the gold standard silica membrane columns.25–29
Here, to address these gaps, we have developed CNASafe, a novel automated microfluidic cfDNA extraction platform with an injection-moulded consumable. We demonstrate its use directly in iSEP-SEQ, a simple and versatile cell-free DNA metagenomic workflow to identify pathogens in non-cultured blood samples.
Material and methods
Overall concept
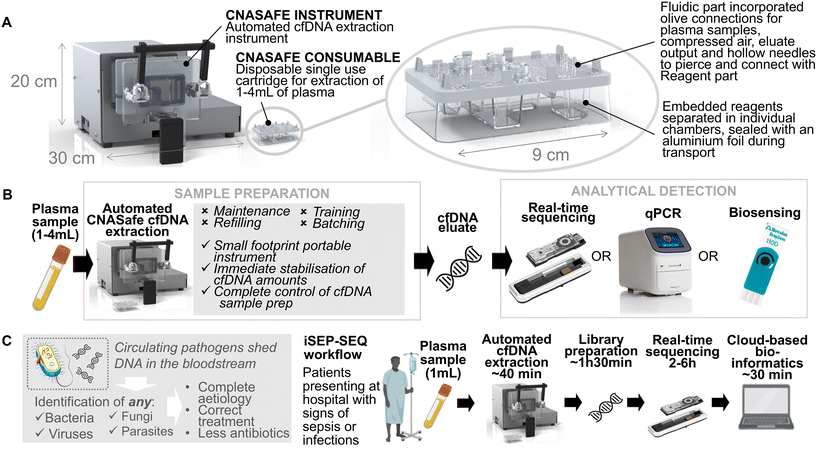
The CNASafe platform comprises a moulded consumable cartridge and an automated instrument (Fig. 1A) and was designed to deliver convenient and immediate sample preparation. It requires no manual intervention in the field, such as reagent pipetting and mixing. The CNASafe platform produces CNA eluates compatible with all major analytical platforms for example quantitative PCR, sequencing workflows or biosensors (Fig. 1B). CNASafe was integrated to iSEP-SEQ, an unbiased cell-free DNA (cfDNA) metagenomic workflow able to identify any DNA-based pathogen (DNA viruses, bacteria, as well as fungi and various parasites) at species level, directly from a plasma sample (Fig. 1C). iSEP-SEQ includes a rapid sequencing assay, delivering a clinically actionable result in the form of a list of top pathogens (viruses, bacteria, as well as fungi and various parasites) at species level.Cartridge fabrication and assembly
Both sides of the CNASafe cartridge were injection moulded in COC TOPAS 8007S-04 by third parties upon specified design. The use of injection moulding ensures the best feature repeatability over other laboratory prototyping methods.30,31 Further details including cartridge filling are described in Method S1. A toggle-clamp mechanism in the instrument allows the combination of the two sides of the cartridge once placed inside the unit. Once ready to use, the sample tube is attached to the cartridge via a tube and an automated instrument. The instrument includes stepper motors to actuate the turning valves, a membrane air pump (Schwarzer precision SP270EC), and an electric heater, and is controlled by a Labview application accessible through a laptop. On completion of the workflow, the eluate obtained was stored at −20 °C until use. Video documentation of the cartridge's internal operation was not possible due to the platform's opaque construction.Healthy donors and patient human samples
For the CNASafe cartridge performance experiments, fresh healthy human blood samples were obtained from the Scottish National Blood Transfusion Services, contract reference 18–20 with Ethics Approval from Office for Research Ethics Committees Northern Ireland (ORECNI) (reference: 18/NI/0148) or Cambridge Biosciences. Cambridge Biosciences operates in conjunction with Research Donors Ltd, which has an HTA license (Human Tissue Authority) and ethical approval from the HRA (Health Research Authority) to operate as a Research Tissue Bank (RTB). Informed consents were obtained from all human participants in this study. Samples were used for consented purpose only and stored accordingly to the Human Tissue Act. Blood samples were spun at 3270 g for 10 min upon reception and the plasma was removed. The plasma was then spun at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 g for 10 min, removed from any pelleted material, and pooled. Aliquots of pooled plasma were stored at −20 °C until further use.
000 g for 10 min, removed from any pelleted material, and pooled. Aliquots of pooled plasma were stored at −20 °C until further use.
For the iSEP-SEQ workflow validation using patient samples, frozen plasma samples from patients hospitalised with sepsis and enrolled in the Cardiovascular Responses In Septic Patients (CRISP) study were obtained through the Malawi-Liverpool-Wellcome Programme. These blood samples were obtained between October 2018 and January 2020 for blood cultures. Plasma was removed from any remaining blood samples, frozen and stored at the Queen Elizabeth Central Hospital at −80 °C. Ten samples were shipped on dry-ice to Heriot-Watt University in October 2021. Upon receipt, samples were stored at −20 °C until cfDNA extraction.
The CRISP study was approved by the National Health Sciences Research Committee in Malawi (reference: 2157), and Liverpool School of Tropical Medicine Research Ethics Committee (reference: 18-062). Approval for use in the iSEP-SEQ workflow was granted by Heriot-Watt University Research Ethics Committee (reference: 1285-iSEP-SEQ_ARCS/320361).
Blood cultures
Blood cultures were processed on an automated BacT/ALERT system (bioMérieux, France). Enterobacteriaceae and oxidase-positive Gram-negative bacilli were identified by API (BioMérieux, France). The blood culture service at the Queen Elizabeth Central Hospital is ISO15189 compliant and accredited the UK National External Quality Assessment Service (UK NEQAS). Bacteria that form part of the normal skin or oral flora, including diphtheroids, bacilli, micrococci, coagulase-negative staphylococci, and α-haemolytic streptococci (other than S pneumoniae), were considered to be contaminants.cfDNA extraction
Manual extractions were performed for benchmark references following the QIAamp Circulating Nucleic Acid Kit (Qiagen) protocol for the purification of circulating nucleic acids from serum or plasma (QIAamp Circulating Nucleic Acid Handbook 10/2019). cfDNA was extracted from 1–4 mL plasma, with cfDNA eluted in 65 μL Buffer AVE. Details corresponding to the CNASafe extractions in the result section are in Method S2.DNA quantification
For cartridge performance experiments, real-time quantitative PCR was performed using 2× Power SYBR® Green PCR Master Mix (Thermo Fisher Scientific) to amplify a 90 bp target with LINE (long interspersed nuclear element) primers (final concentration 200 nM): forward 5′-TGC CGC AAT AAA CAT ACG TG-3′ and reverse 5′-GAC CCA GCC ATC CCA TTA C-3′. 1 μL of DNA template was added to a total reaction volume of 12.5 μL. Thermal cycling conditions involved a 10 minute cycle at 95 °C followed by 40 cycles with 15 seconds at 95 °C and 60 seconds at 60 °C. Samples were amplified in triplicates using Applied Biosystems StepOnePlus™ Real-Time PCR System (Applied Biosystems). Quality controls are discussed in Method S3. DNA yield was expressed as relative recovery compared to the commercial benchmark (Qiagen QIAamp cfDNA Kit). For each sample, identical volumes of the same plasma were processed in parallel on both the microfluidic device and the benchmark kit.Nanopore sequencing and bioinformatics
For all workflow validation samples, library preparation was carried out over 120 min using the Ligation Sequencing Kit (SQK-LSK110, ONT) with no modification to the manufacturer's protocol, beyond the use of SPRIselect beads rather than AmpureXP beads. For sequencing optimisation, libraries were prepared from 50 μL of extraction eluate regardless of the DNA concentration. For sequencing of clinical samples, sequencing libraries were prepared from 45 μL of extraction eluate regardless of the DNA concentration measured using the Qubit. The molarity of the loaded libraries varied between 7.6 to 112.5 fmol. Libraries were sequenced on a MinION Mk1C, Minknow version 21.02.2 or greater, using single-use Flongle Flow Cells (FLO-FLG001, ONT). Sequencing was performed for 6 h with flow cell pore counts ranging from 51–95 at the start of sequencing. High accuracy basecalling was performed using Guppy version 4.0.11 or greater. Qscores were set below 1. Pathogen identification was carried out using the WIMP (v2021.02.10) workflow in EPI2ME (ONT) or a bioinformatic pipeline PATHDET v6.0 or greater.32 All sequencing reads generated were analysed in both the WIMP and PATHDET workflows; no trimming of adapter sequences took place before analysis. Absolute total number of reads were recorded, as well as total number of pathogens reads and number of reads for any pathogens with more than three reads. Pathogen reads were normalised as reads per million reads of total sequencing output (RPM). The compatibility of the CNASafe technology to the nanopore sequencing workflow is presented in Method S4 and Fig. S4.Statistical analysis
Conditions were analysed using GraphPad Prism software. Statistical significance was determined by unpaired parametric Student's t-test or ANOVA. Where unequal sample sizes were present, statistical testing was conducted using Welch's t-test. Significance threshold was set at a p-value less than or equal to 0.05.Results
Cartridge and platform design and operation
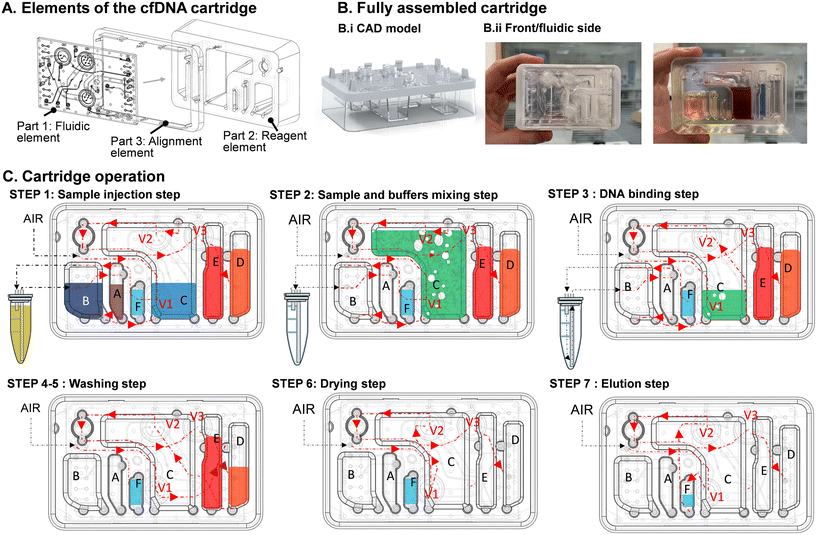
The CNASafe cartridge includes all necessary reagents and relevant fluidics to allow a fully self-contained extraction (Fig. 2A–C): one part containing all the fluidic components (part 1), a second part storing all reagents for the solid phase extraction (part 2), and a third part (part 3) which acts as a transport guard and an alignment feature. The design builds on a previous lab-prototyped version that explored a vertical approach and gravity- and bubble-based mixing.33 We have applied Design for Manufacturing principle, to produce this new design and enable the use of injection moulding to produce the devices. Injection Moulding is a high-throughput manufacturing method which allows highly reproducible features30,31 There are several unique features to this cartridge arrangement: (i) the three-dimensional nature of the cartridge creates enough space for reagent storage for extraction of millilitre volumes compatible with typical clinical sample volumes for CNA applications; (ii) the reagents are embedded into double-use ‘blisters’ (i.e., needles are used to pierce and access their contents as in a traditional blister, but the rigid nature of the blisters mean that they can be used for further steps, such as mixing); and (iii) the configuration of the inlet and outlet coming into and leaving the mixing chamber enables the effective mixing of reagents followed by siphoning onto the silica membrane. The cartridge design contains an embedded silica membrane. In this work, we prioritised a cartridge connected to an external silica membrane.The CNASafe platform prioritises operational simplicity with automated fluid handling, pre-filled cartridges, and a minimal user interface, to enable deployment in decentralised or resource-limited environments. These design choices reduce operator variability and allow extraction to be performed by non-specialists, but it inevitably limits the degree to which the workflow can be tuned or adapted. Fixed reagent volumes and set experimental sequences make the system easy to use but constrain protocol optimisation, which bring some limitations for certain applications, although if fully productised, different versions of the cartridge and automation programs could be supplied.
Optimisation of CNA extractions on healthy donor samples
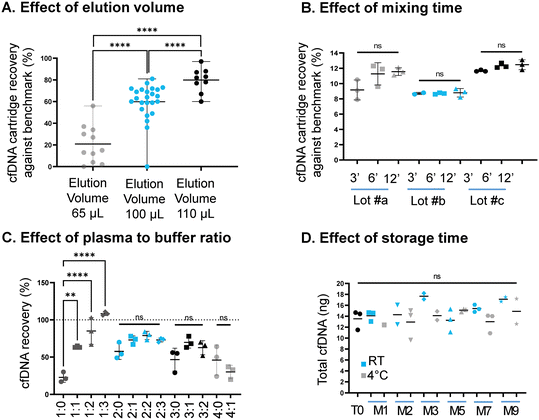
The optimal conditions should provide the highest cfDNA recovery, reduce extraction time and allow storage. Both the mean cartridge eluate volume (Fig. S1) and total cfDNA obtained (Fig. 3A) were significantly increased with a higher elution buffer volume. The maximum eluate volume achieved was 63 μL for 110 μL of elution buffer, for a cfDNA recovery of 67%. We did not push further the volume of elution buffer due to limitation in the maximum volume allowed in the consumable and selected 110 μL for the following experiments. The variability primarily in the dataset reflects biological differences in plasma composition (including for example specific composition, hemolysis) cfDNA binding efficiency, and possible artefacts in the cartridges themselves, which although moulded and thus less subject to variability as a laboratory-made device are still assembled in a research laboratory. When comparing the total amount of cfDNA obtained from the extractions (not taking into account the mixing time), we observed that the column lot number had a significant impact on the result, with significant difference in the total amount of cfDNA recovered between 3 silica membrane lots (Fig. 3B, one-way ANOVA for all three lots, for 3, 6 and 12 min, where respectively p = 0.0018; p = 0.0008; p < 0.0001). Although the reason behind these differences has yet to be elucidated, these results highlight the importance of the silica membrane in cfDNA recovery and call for more research into the differences observed between lot numbers, and the possibility to tune the design of silica membranes to improve cfDNA recovery. The total cfDNA obtained from each mixing time was compared for each column batch: there was no significant difference seen between mixing times for lots #b and #c (cfDNA amount in Fig. S2). Six minutes was selected as the mixing time for the following experiments unless otherwise stated.The starting sample volume is an important consideration in cfDNA analyses. Most analyses use a starting volume of plasma between 1 to 4 mL. Here we tested 1–4 mL of plasma in increments of 1 mL, with, or without topping volume of DPBS. The highest cfDNA recovery was achieved when the total starting sample volume was equal to 4 mL (Fig. 3C). No additional increase in the recovery was observed when the starting volume exceeded 4 mL, indicating that increasing the starting volume to 5 mL may have a diluting effect on the reagents. The variation of viscosity of the starting samples and its effects explain the increase of cfDNA recovery for 1 mL plasma with increasing volume of DPBS (27%, 64%, 85% and 108%, respectively for no DPBS added, and 1 mL, 2 mL and 3 mL of topping DPBS volume) and why the combination 1 mL plasma + 3 mL DPBS was the optimal experimental condition in terms of cfDNA recovery.
Inter-cartridge variability
To evaluate the consistency and robustness of the consumable devices, we examined whether cfDNA amount retrieved varied across different manufacturing lots of extraction cartridges. A unique pooled sample was used for this set of experiment, enabling a direct comparison. Like all other experiments, the cartridge device was used only once which enables the evaluation of inter-cartridge variability. Fig. S3 shows cfDNA amount across same sample in triplicate for several mixing time and membrane lot number (same experiment as shown in Fig. 3B). cfDNA amount did not show significant difference within membrane lot number, and the mean coefficient of variations across all subsets was below 5.5%, which is acceptable for devices that combines high-throughput injection moulding and manual laboratory assembly.Stability over time and temperature storage
Prefilled cartridges need to withstand a certain shelf-life in order to be stocked and used when required. There was no significant difference in the total amount of cfDNA obtained from the cartridges, stored at RT or 4 °C, over the 9 months storage period (Fig. 3D). This confirms that the extraction reagents are stable for up to 9 months when stored in the CNASafe cartridge at room temperature or 4 °C.Large scale dataset results
During the development and utilisation of the CNASafe cfDNA extraction cartridge, a total of 474 extractions were performed. The complete dataset is provided with this manuscript (https://doi.org/10.6084/m9.figshare.28075193.v1). Of these, 358 extractions were conducted after transitioning the sealing process in-house, yielding 333 successful eluates. The average cfDNA recovery from these 333 eluates, encompassing all parameter variations, was 100.5%, demonstrating a performance equivalent to the industry gold standard for cfDNA yield.iSEP-SEQ workflow validation with patient clinical samples
Following the optimisation of the sequencing workflow, the application of the iSEP-SEQ workflow to the identification of pathogens was investigated using banked plasma samples from ten patients with sepsis as described in methods. The clinical presentation of the patients is summarised in Table 1.| Age | Sex | HIV/ART status | CD4 cell count (cells per uL) | Antibiotics, in the last 4 weeks | Fever | Cough | Chest pain | Headache | Vomiting | T (°C) | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| CRISP 1 | 30–40 | Male | HIV+, ART | 69 | Amoxycillin | Yes | Yes | No | No | Yes | 39.2 |
| CRISP 2 | 20–30 | Female | HIV+, ART | 240 | Erythromycin metronidazole | Yes | No | Yes | Yes | No | 38.1 |
| CRISP 3 | 30–40 | Male | HIV+, ART | 50 | Positive, but unknown prescription | No | No | No | No | No | 38.5 |
| CRISP 4 | 30–40 | Female | HIV+, ART | 611 | Positive, but unknown prescription | No | Yes | Yes | No | No | 36 |
| CRISP 5 | 30–40 | Female | HIV+, ART | 45 | Ceftriaxone ciprofloxacin | Yes | Yes | Yes | No | No | 37.6 |
| CRISP 6 | 40–50 | Female | HIV+, ART | 15 | Ciprofloxacin | Yes | Yes | No | No | Yes | 36.1 |
| CRISP 7 | 30–40 | Male | HIV+, ART | 126 | Amoxycillin | Yes | Yes | Yes | Yes | No | 39.1 |
| CRISP 8 | 20–30 | Male | Negative | NA | Doxycycline | Yes | Yes | Yes | No | No | 38.1 |
| CRISP 9 | 20–30 | Female | HIV+, ART | 440 | Ceftriaxone | Yes | Yes | No | No | Yes | 34.3 |
| CRISP 10 | 40–50 | Male | HIV+, ART | 166 | Metronidazole, Bactrim | Yes | Yes | Yes | No | Yes | NA |
The samples were passed through the full iSEP-SEQ workflow. cfDNA was extracted from banked plasma samples ranging from 300–1000 μL using the CNASafe cartridges. All samples produced an eluate. One CNASafe extraction did not run to completion (sample 6), but an eluate from this sample was still obtained after the column was removed from the cartridge and the remaining steps (wash, dry and elution) performed on the bench. The elution volume range was 52.4 ± 2.8 μL, excluding the sample manually recovered from the cartridge. All samples were successfully sequenced, but one set of sample files (sample 3) was lost during transfer due to an IT error. The samples were passed through PATHDET, a proprietary bioinformatic pipeline hosted by the University of Tokyo. In addition, EPI2ME, the bioinformatic platform from Oxford Nanopore Technologies was also used to compare with PATHDET. Summary data from the processed samples are presented in Table 2, and an extended statistical data table is available in Table S1. EPI2ME statistics are provided in Table S2 as a secondary verification of the PATHDET results and for additional statistics. The total number of reads varied between 47![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 078 and 873
078 and 873![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 892 and the percentage unclassified ranged between 5 to 12.6% (EPI2ME data).
892 and the percentage unclassified ranged between 5 to 12.6% (EPI2ME data).
| Case no. | iSEP-SEQ mNGS results (with PATHDET bioinformatic workflow) | Blood culture result | Concordance and comparison | |||
|---|---|---|---|---|---|---|
| iSEP-SEQ mNGS quantitative result (identified organisms (species, 3 reads or more)) | Number of reads | RPM | iSEP-SEQ mNGS qualitative result | |||
| CRISP 1 | None | — | — | Negative | Negative | Negative concordance |
| CRISP 2 | Human gammaherpesvirus 8 | 3 | 13 | Small signal for human gammaherpesvirus | Negative | iSEP-SEQ identification of HHV8 |
| CRISP 4 | Human gammaherpesvirus 8 | 5 | 59 | Small signal for human gammaherpesvirus 8 and Arcobacter cryaerophilus | Positive, ‘gpc seen’, ‘CoNS’ | iSEP-SEQ identfication of HHV8 and A. cryaerophilus below threshold, but plausible |
| Arcobacter cryaerophilus | 5 | 59 | ||||
| Rhizobacter gummiphilus | 3 | 35 | ||||
| Cutibacterium acnes | 3 | 35 | ||||
| CRISP 5 | None | — | — | Negative | Positive ‘gnb seen’ ‘K. pneumoniae’ | iSEP-SEQ false negative due to small total number of reads |
| CRISP 6 | Methylobacterium sp. XJLW | 366 | 1743 | Strong signal for Methylobacterium | Positive ‘gpc seen’ ‘CoNS’ | iSEPSEQ identification of Methylobacterium plausible given the strong signal |
| Methylobacterium phyllosphaerae | 202 | 962 | ||||
| Methylobacterium oryzae | 55 | 261 | ||||
| Human betaherpesvirus 5 | 3 | 14 | ||||
| CRISP7 | Cutibacterium acnes | 4 | 13.7 | Negative (contamination) | Negative | Negative concordance |
| CRISP 8 | Streptococcus pneumoniae | 124 | 385 | Strong signal for Streptococcus pneumoniae | Positive – ‘gnb seen’ ‘No growth but gpc seen’ | iSEP-SEQ identification of S. pneumoniae which matches the clinical context |
| Methylobacterium sp. XJLW | 6 | 18 | ||||
| Methylobacterium phyllosphaerae | 4 | 12 | ||||
| CRISP 9 | Methylobacterium sp. XJLW | 4 | 18 | Negative (contamination) | Negative | Negative concordance |
| CRISP 10 | Methylobacterium sp. XJLW | 33 | 85 | Negative (contamination) | Negative | Negative concordance |
| Methylobacterium phyllosphaerae | 17 | 44 | ||||
| Methylobacterium oryzae CBMB20 | 3 | 7 | ||||
In six out of nine cases, iSEP-SEQ was consistent with blood culture results. These include CRISP 1, 2, 7, 9 and 10, where the blood cultures were found negative, and where iSEP-SEQ identified bacteria in low numbers, below the threshold of for positive identification of bacteria.11,12
In CRISP 4, iSEP-SEQ identified Arcobacter cryaerophilus. While previously described as a pathogen in certain patients,34,35 the signal is below the 100 RPM for positive identification of bacteria.36 Human gammaherpesvirus 8 (HHV-8) was found at a level of 59 RPM, above the threshold generally set for viruses.37 This virus would not have been identified in the blood culture. Although not a sepsis-causing pathogen, HHV viruses can cause community-acquired pneumonia (CAP) in compromised hosts38 and are a major cause of morbidity in patients with AIDS and other immunosuppressive disorders.39 The patient's severe dyspnoea, chest pain and cough with mucus support this possibility.
In CRISP 5, cultures identified K. pneumoniae. Due to a lower number of reads in that particular sample, it is likely to have been missed by iSEP-SEQ. This could be prevented in the future by processing larger volumes of plasma.
In CRISP 6, iSEP-SEQ identified three Methylobacterium species which cultures failed to detect, which instead identified coagulase-negative staphylococci, considered to be a contaminant from the patient's skin. Methylobacterium, which exists in soil, dust, water, and sewage, rarely cause human disease, but have been previously identified as responsible for bacteremia in immunocompromised patients (patient had a CD4 count of 15 cells per μL), such as patients with HIV included in this analysis.40,41 Methylobacterium is a fastidious, slow-growing bacterium that can be easily missed during microbiological surveillance.42 It is also a known contaminant of DNA extraction kits.43 However, the high RPM value (2966 RPM at Methylobacterium genus level), 100 times above typical threshold values and the fact that another sample was extracted after CRISP 6 using reagents from the same potentially contaminated batch number and showed no evidence of such heavy Methylobacterium contamination, point to this being a true positive.
To rule out contamination, we implement multiple controls throughout the workflow. Water blanks are routinely processed through the CNASafe apparatus. Exemplar resulting negative-control data (included in Table S3) show no detectable contamination from reagents, plasticware, or the microfluidic cassettes. In addition, the negative healthy controls in the preliminary data on microbial mix (‘Compatibility of CNASafe technology to nanopore sequencing workflow’ Method S4 and Fig. S4) also show that the on-chip cfDNA extraction does not introduce contaminants. Together, these data explain why the observed unusually high-RPM Methylobacterium signal is unlikely to originate from reagent contamination and instead reflects true sample biology.
In CRISP 8, iSEP-SEQ workflow identified a strong Streptococcus pneumoniae signal (124 reads corresponding to 8% of total reads, 385 RPM and covering 9% of the organism's genome). This was not identified in blood culture, which only stated that Gram-positive cocci were observed. Streptococcus pneumoniae is a bacteria responsible for severe lung infection, a result highly consistent with the presentation of the patient (coughing sputum, fever, chest pain and difficulty breathing).
While these results would benefit from confirmation by PCR, it highlights the potential of the microfluidic-enabled portable workflow for the culture-free and fast identification of sepsis-causing pathogens.
Discussion
While cfDNA has been shown to be a powerful biomarker, cfDNA extraction remains a challenge. Skilled staff are employed to process samples for cfDNA manually or using high-throughput robots. The former is subject to human performance and suffers from variability from one person to the next. The latter requires high-capital investment in centralised facilities, extensive staff training, high-maintenance and long turnaround time due to the need to batch processes. Thus, complex sample processing is a factor that restricts the clinical availability and time-to-result of cfDNA-based assays. This is especially relevant to a context like infection diagnosis where time is critical and where diagnosis may be needed in remote settings without high-capital infrastructure.We have developed an integrated single-use cartridge produced by high-volume injection moulding and containing pre-filled reagents. Together with its companion automation platform, the device transforms a sample of plasma into an eluate of cfDNA in less than 45 minutes, with scope to further reduce the extraction time. In total, over 400 human plasma samples were used in the investigation of various parameters, including the elution buffer volume, mixing time and silica membrane lot number, as well as plasma and DPBS sample volume. A subset of these results has been presented here, and the complete dataset is available. Following preliminary optimisation, the average recovery of CNASafe cartridge was found to be 100.5% relatively to the gold standard in the field of cfDNA extraction (QIAamp CNA Kit, manual extraction), highlighting the performance of the proposed platform for rapid, automated, and highly efficient extraction of cfDNA from human plasma samples.
Our microfluidic CNASafe platform demonstrates competitive performance with existing cfDNA extraction technologies while offering advantages in automation and validation rigour. The system processes 1–4 mL of plasma with full automation in 40 minutes, positioning it among higher-capacity platforms in the field. This volume exceeds most semi-automated systems,23,27,44–48 which typically process ≤1 mL, and is comparable to fully automated platforms like Kim et al.26 and Lee et al.,28 which handle >3 mL and 0.5–1 mL respectively in 30 and 15 minutes.
While our 40 minute processing time is longer than some fully automated systems, it substantially outperforms manual approaches like Campos et al.,23 which would require over 8 hours for 1 mL at the maximum stated flow rate. The fully automated workflow eliminates the manual intervention steps required by semi-automated or manual platforms.
A key strength is our validation methodology. We characterised relative cfDNA recovery to a gold standard using >400 healthy donor samples processed identically on both our device and the Qiagen QIAamp benchmark, achieving 100% relative recovery by quantitative qPCR. This contrasts with most studies, which report recovery rates of 50–92% using synthetic DNA spikes, often at unspecified concentrations or with non- or semi-quantitative methods, making direct comparisons difficult. This literature comparison, which is available in a separate SI file, highlights the necessity for adequate validation standards in the field. Such validation standards could be developed in partnership with clinicians, and organisations like BloodPAC (https://www.bloodpac.org/). CNASafe provides an automated solution for the extraction of cfDNA in an environment where speed is critical, and samples cannot be batched for delayed extraction.
While nanopore sequencing has been widely used for genomic surveillance of emerging viruses, its clinical metagenomic applications for pathogen detection remain limited, in part because of the complex sample preparation and the need for skilled staff to operate complex batched workflows. Here we demonstrated that the performance of an integrated cartridge (CNASafe) to isolate cfDNA and reduce the time to identification of circulating microbial cfDNA without the need to batch samples. The sample preparation is pathogen-agnostic and does not introduce biases to organisms, unlike several methods that enrich pathogen-specific reads and focus on bacteria and fungi. Most metagenomic studies rely on long sequencing sequencing run times exceeding 12 h, although there is a tendency to reduce sequencing time in recent years.12 In contrast, our sequencing run times did not exceed 6 h on the low capital investment and portable ONT platform, and 2 h sequencing may be sufficient to detect pathogens in circulation and responsible for diseases and infections.12,32 If reduced to 2 h, the whole iSEP-SEQ workflow could be performed in under 5 h, providing a field-based, fast, sequencing-based identification of blood-borne pathogens. We have demonstrated that the CNASafe cfDNA eluate is compatible with ONT sequencing, known to be sensitive to contaminants, and applied the complete workflow on ten clinical samples, uncovering several organisms which were not identified by blood cultures. Our validation with nine sepsis patients also distinguishes this work from the predominantly oncology-focused applications in the literature, establishing the platform as a robust solution for infectious disease diagnostics where rapid pathogen identification from cfDNA is critical.
The proposed iSEP-SEQ metagenomic workflow would enable prompt aetiological investigations in decentralised settings or clinics without well-equipped centralised laboratories, leading to more consistent, more accurate and faster results than current solutions, alleviating the need for additional invasive procedures (e.g., bronchoscopy or bronchoalveolar lavage), reducing the time spent in hospital and associated costs, as well as reducing adverse effects from broad-spectrum antibiotics, thus leading to better outcome for patients.
Author contributions
The corresponding author takes full responsibility that all authors on this publication have met the following required criteria of eligibility for authorship: (a) significant contributions to the conception and design, acquisition of data, or analysis and interpretation of data; (b) drafting or revising the article for intellectual content; (c) final approval of the published article; and (d) agreement to be accountable for all aspects of the article thus ensuring that questions related to the accuracy or integrity of any part of the article are appropriately investigated and resolved. Nobody who qualifies for authorship has been omitted from the list.Conceptualisation: Linda Marriott, Ana Martinez-Lopez, Antonio Liga, Nicholas Feasey, Shevin T. Jacob, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas. Data curation: Linda Marriott, Ana Martinez-Lopez, Nicholas Feasey, Shevin T. Jacob, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas. Formal analysis: Linda Marriott, Ana Martinez Lopez, Radhe Shantha Kumar, Benjamin J. Parcell, Nicholas R. Leslie, Nicholas Feasey, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas. Funding acquisition: Nicholas Feasey, Shevin T. Jacob, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas. Investigation: Linda Marriott, Ana Martinez-Lopez, Antonio Liga, Shevin T. Jacob, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas. Methodology: Linda Marriott, Ana Martinez-Lopez, Antonio Liga, Kazuhiro Horiba, Amanda Warr, Shevin T. Jacob, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas. Project administration: Linda Marriott, Antonio Liga, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas. Resources: Jacob N. Phulusa, Laura Carey, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas. Software: Linda Marriott, Ana Martinez-Lopez, Antonio Liga, Kazuhiro Horiba, and Yoshinori Ito. Supervision: Yoshinori Ito, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas. Validation: Linda Marriott, Ana Martinez-Lopez, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas. Visualisation: Linda Marriott, Antonio Liga, and Maïwenn Kersaudy-Kerhoas. Writing – original draft: Linda Marriott, Antonio Liga, and Maïwenn Kersaudy-Kerhoas. Writing – review & editing: Linda Marriott, Ana Martinez-Lopez, Antonio Liga, Kazuhiro Horiba, Amanda Warr, Jacob N. Phulusa, Radhe Shantha Kumar, Laura Carey, Yoshinori Ito, Benjamin J. Parcell, Nicholas R. Leslie, Nicholas Feasey, Shevin T. Jacob, Jamie Rylance, and Maïwenn Kersaudy-Kerhoas.
Conflicts of interest
AL, MKK and KH have filed and been granted patents related to this work.Data availability
Data regarding the details of all 400+ extractions performed are available at FigShare at https://figshare.com/articles/dataset/CNASafe_Dataset/28075193?file=51336866.Supplementary information (SI): Supplemental Method 1: CNASafe cartridge assembly. Supplemental method 2: cfDNA extractions on CNASafe device. Supplemental Method 3: qPCR quality controls. Supplemental Method 4: Compatibility of CNASafe technology to nanopore sequencing workflow. Supplemental Figure 1: Impact of on-chip elution volume on eluate volume. Supplemental Figure 2: Impact of lot number and mixing time on amount of cfDNA (ng). Supplemental Figure 3: (A) Total amount cfDNA (ng) for different lot number and mixing time and (B) corresponding coefficient of variations. Supplemental Figure 4: Characterisation of sequencing workflow on cartridge. Supplementary Table S1: Extended sequencing data for clinical patient samples in the CRISP iSEP-SEQ study. Supplementary Table S2: Statistics for EPI2ME bioinformatic workflow. Supplementary Table S3: Epi2me raw data and statistics for blank water sample. Supplemental Table S4: CfDNA extraction comparison table. See DOI: https://doi.org/10.1039/d5lc00876j.
Acknowledgements
We acknowledge Engineering and Physical Science Research Council EP/R00398X/1, Royal Academy of Engineering Frontiers of Development scheme, Heriot-Watt University, Scottish Enterprise High Growth Spin Out funding, and Santander Universities Santander X award for funding. MKK thanks E. S. for her suggestions towards the structure of this manuscript and some of the data presentation.References
- H. Schwarzenbach, D. S. B. Hoon and K. Pantel, Cell-free nucleic acids as biomarkers in cancer patients, Nat. Rev. Cancer, 2011, 11, 426–437, DOI:10.1038/nrc3066.
- S. Volik, M. Alcaide, R. D. Morin and C. Collins, Cell-free DNA (cfDNA): Clinical Significance and Utility in Cancer Shaped By Emerging Technologies Early Discovery and Applications of cfDNA, Mol. Cancer Res., 2016, 14(10), 898–908, DOI:10.1158/1541-7786.MCR-16-0044.
- D. Fernandez-Garcia, A. Hills, K. Page, R. K. Hastings, B. Toghill and K. S. Goddard, et al., Plasma cell-free DNA (cfDNA) as a predictive and prognostic marker in patients with metastatic breast cancer, Breast Cancer Res., 2019, 21, 149, DOI:10.1186/s13058-019-1235-8.
- M. Oellerich, E. Schütz, J. Beck, P. Kanzow, P. N. Plowman and G. J. Weiss, et al., Using circulating cell-free DNA to monitor personalized cancer therapy, Crit. Rev. Clin. Lab. Sci., 2017, 54, 205–218, DOI:10.1080/10408363.2017.1299683.
- J. Tie, Y. Wang, C. Tomasetti, L. Li, S. Springer and I. Kinde, et al., Circulating tumor DNA analysis detects minimal residual disease and predicts recurrence in patients with stage II colon cancer, Sci. Transl. Med., 2016, 8(346), 346ra92, DOI:10.1126/scitranslmed.aaf6219.
- Y. M. D. Lo, N. Corbetta, P. F. Chamberlain, V. Rai, I. L. Sargent and C. W. G. Redman, et al., Presence of fetal DNA in maternal plasma and serum, Lancet, 1997, 350, 485–487, DOI:10.1016/S0140-6736(97)02174-0.
- J. Beck, S. Bierau, S. Balzer, R. Andag, P. Kanzow and J. Schmitz, et al., Digital droplet PCR for rapid quantification of donor DNA in the circulation of transplant recipients as a potential universal biomarker of graft injury, Clin. Chem., 2013, 59, 1732–1741, DOI:10.1373/clinchem.2013.210328.
- E. Schütz, A. Fischer, J. Beck, M. Harden, M. Koch and T. Wuensch, et al., Graft-derived cell-free DNA, a noninvasive early rejection and graft damage marker in liver transplantation: A prospective, observational, multicenter cohort study, PLoS Med., 2017, 14(4), e1002286, DOI:10.1371/journal.pmed.1002286.
- K. Wang, S. Zhang, B. Marzolf, P. Troisch, A. Brightman and Z. Hu, et al., Circulating microRNAs, potential biomarkers for drug-induced liver injury, Proc. Natl. Acad. Sci. U. S. A., 2009, 106(11), 4402–4407, DOI:10.1073/pnas.0813371106.
- A. D. B. Vliegenthart, C. Berends, C. M. J. Potter, M. Kersaudy-Kerhoas and J. W. Dear, MicroRNA-122 can be measured in capillary blood which facilitates point-of-care testing for drug-induced liver injury, Br. J. Clin. Pharmacol., 2017, 83(9), 2027–2033, DOI:10.1111/bcp.13282.
- T. A. Blauwkamp, S. Thair, M. J. Rosen, L. Blair, M. S. Lindner and I. D. Vilfan, et al., Analytical and clinical validation of a microbial cell-free DNA sequencing test for infectious disease, Nat. Microbiol., 2019, 4, 663–674, DOI:10.1038/s41564-018-0349-6.
- C. Grumaz, A. Hoffmann, Y. Vainshtein, M. Kopp, S. Grumaz and P. Stevens, et al., Rapid Next-Generation Sequencing–Based Diagnostics of Bacteremia in Septic Patients, J. Mol. Diagn., 2020, 22, 405–418, DOI:10.1016/j.jmoldx.2019.12.006.
- K. Horiba, J. I. Kawada, Y. Okuno, N. Tetsuka, T. Suzuki and S. Ando, et al., Comprehensive detection of pathogens in immunocompromised children with bloodstream infections by next-generation sequencing, Sci. Rep., 2018, 8, 1–9, DOI:10.1038/s41598-018-22133-y.
- S. Dugani, J. Veillard and N. Kissoon, Reducing the global burden of sepsis. CMAJ, Can. Med. Assoc. J., 2017, 189(1), E2–E3, DOI:10.1503/cmaj.160798.
- F. Prestinaci, P. Pezzotti and A. Pantosti, Antimicrobial resistance: a global multifaceted phenomenon, Pathog. Global Health, 2015, 109, 309, DOI:10.1179/2047773215Y.0000000030.
- R. Boom, J. A. Sol, M. M. M. Salimans, C. L. Jansen, P. M. E. Wertheim-Van Dillen and J. Van Der Noordaa, Rapid and Simple Method for Purification of Nucleic Acids, J. Clin. Microbiol., 1990, 28(3), 495–503, DOI:10.1128/jcm.28.3.495-503.1990.
- M. A. M. Gijs, F. Lacharme and U. Lehmann, Microfluidic applications of magnetic particles for biological analysis and catalysis, Chem. Rev., 2010, 110, 1518–1563, DOI:10.1021/cr9001929.
- S. J. Reinholt and A. J. Baeumner, Microfluidic isolation of nucleic acids, Angew. Chem., Int. Ed., 2014, 53, 13988–14001, DOI:10.1002/anie.201309580.
- A. M. Streets and Y. Huang, Chip in a lab: Microfluidics for next generation life science research, Biomicrofluidics, 2013, 7(1), 011302, DOI:10.1063/1.4789751.
- N. C. Cady, S. Stelick and C. A. Batt, Nucleic acid purification using microfabricated silicon structures, Biosens. Bioelectron., 2003, 19(1), 59–66, DOI:10.1016/s0956-5663(03)00123-4.
- M. C. Breadmore, K. A. Wolfe, I. G. Arcibal, W. K. Leung, D. Dickson and B. C. Giordano, et al., Microchip-Based Purification of DNA from Biological Samples, Anal. Chem., 2003, 75, 1880–1886, DOI:10.1021/ac0204855.
- K. Perez-Toralla, I. Pereiro, S. Garrigou, F. Di Federico, C. Proudhon and F. C. Bidard, et al., Microfluidic extraction and digital quantification of circulating cell-free DNA from serum, Sens. Actuators, B, 2019, 286, 533–539, DOI:10.1016/j.snb.2019.01.159.
- C. D. M. Campos, S. S. T. Gamage, J. M. Jackson, M. A. Witek, D. S. Park and M. C. Murphy, et al., Microfluidic-based solid phase extraction of cell free DNA, Lab Chip, 2018, 18, 3459–3470, 10.1039/c8lc00716k.
- C. W. Price, D. C. Leslie and J. P. Landers, Nucleic acid extraction techniques and application to the microchip, Lab Chip, 2009, 9, 2484–2494, 10.1039/B907652M.
- M. Amasia and M. Madou, Microfluidic device for whole blood sample preparation [Internet], Google Patents, 2015, Available from: https://www.google.com/patents/US9186672.
- C.-J. Kim, J. Park, V. Sunkara, T.-H. Kim, Y. Lee and K. Lee, et al., Fully automated, on-site isolation of cfDNA from whole blood for cancer therapy monitoring, Lab Chip, 2018, 18, 1320–1329, 10.1039/C8LC00165K.
- F. Hu, J. Li, N. Peng, Z. Li, Z. Zhang and S. Zhao, et al., Rapid isolation of cfDNA from large-volume whole blood on a centrifugal microfluidic chip based on immiscible phase filtration, Analyst, 2019, 144, 4162–4174, 10.1039/C9AN00493A.
- H. Lee, C. Park, W. Na, K. H. Park and S. Shin, Precision cell-free DNA extraction for liquid biopsy by integrated microfluidics, npj Precis. Oncol., 2020, 4, 1–10, DOI:10.1038/s41698-019-0107-0.
- F. Schlenker, P. Juelg, J. Lüddecke, N. Paust, R. Zengerle and T. Hutzenlaub, Nanobead handling on a centrifugal microfluidic LabDisk for automated extraction of cell-free circulating DNA with high recovery rates, Analyst, 2023, 148, 932–941, 10.1039/D2AN01820A.
- R. Filion, A. St-Georges-Robillard, H. Chaudoir, S. Ritch, M. Ndongo and X. Y. Chua, et al., Mass fabrication of PDMS microfluidic devices by injection molding and applications in sensitive 3D spheroid and explant culture, Sci. Rep., 2025, 15, 33218, DOI:10.1038/s41598-025-16863-z.
- U. N. Lee, X. Su, D. J. Guckenberger, A. M. Dostie, T. Zhang and E. Berthier, et al., Fundamentals of rapid injection molding for microfluidic cell-based assays, Lab Chip, 2018, 18, 496–504, 10.1039/C7LC01052D.
- K. Horiba, Y. Torii, Y. Aizawa, M. Yamaguchi, K. Haruta and T. Okumura, et al., Performance of Nanopore and Illumina Metagenomic Sequencing for Pathogen Detection and Transcriptome Analysis in Infantile Central Nervous System Infections, Open Forum Infect. Dis., 2022, 9(10), ofac504, DOI:10.1093/ofid/ofac504.
- M. Kersaudy-Kerhoas, A. Liga, A. Roychoudhury, M. Stamouli, R. Grant and D. S. Carrera, et al., Microfluidic system for near-patient extraction and detection of miR-122 microRNA biomarker for drug-induced liver injury diagnostics, Biomicrofluidics, 2022, 16(2), 024108, DOI:10.1063/5.0085078.
- P. R. Hsueh, L. J. Teng, P. C. Yang, S. K. Wang, S. C. Chang and S. W. Ho, et al., Bacteremia caused by Arcobacter cryaerophilus 1B, J. Clin. Microbiol., 1997, 35, 489, DOI:10.1128/jcm.35.2.489-491.1997.
- T. P. Ramees, K. Dhama, K. Karthik, R. S. Rathore, A. Kumar and M. Saminathan, et al., Arcobacter: an emerging food-borne zoonotic pathogen, its public health concerns and advances in diagnosis and control – a comprehensive review, Vet. Q., 2017, 37, 136–161, DOI:10.1080/01652176.2017.1323355.
- W. Gu, X. Deng, M. Lee, Y. D. Sucu, S. Arevalo and D. Stryke, et al., Rapid pathogen detection by metagenomic next-generation sequencing of infected body fluids, Nat. Med., 2020, 27, 115–124, DOI:10.1038/s41591-020-1105-z.
- Y. Wang, T. Chen, S. Zhang, L. Zhang, Q. Li and Q. Lv, et al., Clinical evaluation of metagenomic next-generation sequencing in unbiased pathogen diagnosis of urinary tract infection, J. Transl. Med., 2023, 21, 762, DOI:10.1186/s12967-023-04562-0.
- B. A. Cunha, Cytomegalovirus Pneumonia: Community-Acquired Pneumonia in Immunocompetent Hosts, Infect. Dis. Clin. North Am., 2010, 24, 147, DOI:10.1016/j.idc.2009.10.008.
- N. Salomon and D. C. Perlman, Cytomegalovirus pneumonia, Semin. Respir. Infect., 1999, 14, 353–358, DOI:10.1016/S0012-3692(16)58747-9.
- A. L. Truant, R. Gulati, O. Giger, V. Satishchandran and J. G. Caya, Methylobacterium bacteremia in AIDS, Clin. Microbiol. Infect., 1998, 4, 112–113, DOI:10.1111/j.1469-0691.1998.tb00369.x.
- J. W. Sanders, J. W. Martin, M. Hooke and J. Hooke, Methylobacterium mesophilicum Infection: Case Report and Literature Review of an Unusual Opportunistic Pathogen, Clin. Infect. Dis., 2000, 30, 936–938, DOI:10.1086/313815.
- K. J. Szwetkowski and J. O. Falkinham, Methylobacterium spp. as Emerging Opportunistic Premise Plumbing Pathogens, Pathogens, 2020, 9, 149, DOI:10.3390/pathogens9020149.
- A. Glassing, S. E. Dowd, S. Galandiuk, B. Davis and R. J. Chiodini, Inherent bacterial DNA contamination of extraction and sequencing reagents may affect interpretation of microbiota in low bacterial biomass samples, Gut Pathog., 2016, 8, 1–12, DOI:10.1186/s13099-016-0103-7.
- J. Yang, P. R. Selvaganapathy, T. J. Gould, D. J. Dwivedi, D. Liu and A. E. Fox-Robichaud, et al., A microfluidic device for rapid quantification of cell-free DNA in patients with severe sepsis, Lab Chip, 2015, 15, 3925–3933, 10.1039/C5LC00681C.
- H. G. Kye, C. D. Ahrberg, B. S. Park, J. M. Lee and B. G. Chung, Separation, Purification, and Detection of cfDNA in a Microfluidic Device, BioChip J., 2020, 14, 195–203, DOI:10.1007/s13206-020-4208-1.
- H. Gwak, J. Kim, S. Cha, Y. P. Cheon, S. I. Kim and B. Kwak, et al., On-chip isolation and enrichment of circulating cell-free DNA using microfluidic device, Biomicrofluidics, 2019, 13, 024113, DOI:10.1063/1.5100009.
- L. Schneider, T. Usherwood and A. Tripathi, A microfluidic platform for high-purity cell free DNA extraction from plasma for non-invasive prenatal testing, Prenatal Diagn., 2022, 42, 240–253, DOI:10.1002/pd.6092.
- C. E. Jin, B. Koo, T. Y. Lee, K. Han, S. B. Lim and I. J. Park, et al., Simple and Low-Cost Sampling of Cell-Free Nucleic Acids from Blood Plasma for Rapid and Sensitive Detection of Circulating Tumor DNA, Adv. Sci., 2018, 5, 1800614, DOI:10.1002/advs.201800614.
| This journal is © The Royal Society of Chemistry 2026 |