Modeling amyotrophic lateral sclerosis (ALS) in vitro: from mechanistic studies to translatable drug discovery
Kathryn G.
Maskell
 *a,
Anthony L.
Cook
*a,
Anthony L.
Cook
 b,
Anna E.
King
b,
Anna E.
King
 b,
Tracey C.
Dickson†
b,
Tracey C.
Dickson†
 a and
Catherine A.
Blizzard†
a and
Catherine A.
Blizzard†
 *a
*a
aMenzies Institute for Medical Research, Tasmania, Australia. E-mail: kathryn.maskell@utas.edu.au; Catherine.blizzard@utas.edu.au
bWicking Dementia Research and Education Centre, Tasmania, Australia
First published on 7th January 2026
Abstract
Amyotrophic lateral sclerosis is a rapidly progressing, fatal neurodegenerative disease that causes selective degeneration of the corticomotor system. Currently, ALS remains incurable, and the available treatment options offer little in the way of extending life or improving quality of life. This is due, at least in part, to a lack of representative disease models. In vitro modeling offers rapid, experimentally accessible platforms for mechanistic discovery research and drug screening, but modeling the complexity of ALS – a multicellular, multisystem disease – in a dish, is not without its challenges. Here, we review the current landscape of in vitro pre-clinical ALS research, with emphasis on the development of compartmentalised culture and the promise this holds for translatable modeling of ALS.
1 Introduction
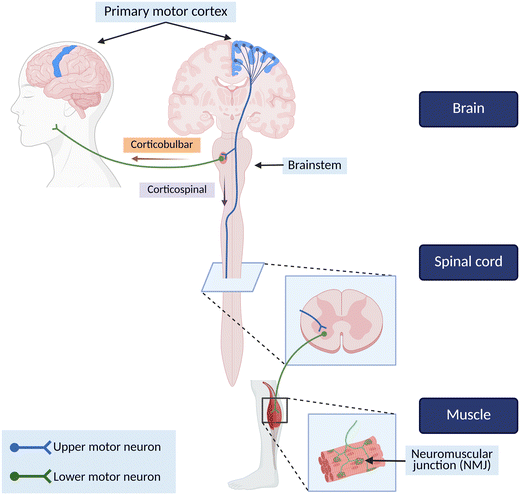
Motor neuron diseases cause degeneration of the corticomotor system, which consists of the motor cortex in the brain, the brain stem, the ventral horn of the spinal cord and the muscles, together facilitating controlled, complex body movements (Fig. 1). The most common form of motor neuron disease is amyotrophic lateral sclerosis (ALS). There is significant heterogeneity in disease onset, disease progression and responses to treatment in ALS, but symptom progression generally entails a relentless loss of the ability to move, eat, speak, and, ultimately, breathe. On average, patients have a life expectancy of 2–5 years, with a median survival time of 2.5 years after symptom onset.1,2 Research into ALS aetiology has revealed over 50 causative genetic mutations that can promote a familial form of ALS (fALS). Pathogenic variants in the superoxide dismutase (SOD1), fused in sarcoma (FUS), TAR DNA-binding protein (TARDBP), and chromosome 9 open reading frame 72 (C9orf72) genes are the most common cause of fALS, accounting for roughly 70% of fALS cases.3 However, 90% of all ALS cases arise with no known familial association and are thus currently referred to as sporadic ALS (sALS).4 It is not yet known what causes sALS, but it is likely that interactions between genetic polymorphisms and environmental and lifestyle factors, including ageing-related factors, contribute.5,6ALS is characterised by involvement of upper motor neurons (UMNs) in the brain, lower motor neurons (LMNs) in the spinal cord, and skeletal and smooth muscle in the periphery (Fig. 1). We still do not understand where the disease begins, how it spreads, or why it selectively attacks UMNs and LMNs. Key pathological processes include circuit dysfunction such as synaptic loss, changes in excitability, and neuroinflammatory processes, as well as cell-centric disturbances such as mitochondrial dysfunction, protein aggregation, impaired RNA processing, and defective autophagy.7–12 However, it remains unclear which of these is a primary driver versus a compensatory mechanism. The involvement of the corticomotor system in its entirety renders a uniquely complex setting for understanding neurodegeneration in ALS. Indeed, dysfunction in this disease can spread through the central nervous system and into the periphery, or vice versa, affecting cells in discrete biological compartments.13 For example, the central nervous system operates in cerebrospinal fluid (CSF), a tightly regulated, immune-privileged environment that is kept separate from systemic circulation by the blood brain barrier. In comparison, muscle tissue resides in an immunologically active environment directly exposed to systemic circulation. In addition, the regenerative capacity of muscle tissue exceeds that of nervous tissue, meaning that it is likely that mechanisms of dysfunction may vary between environments and cell types. Understanding where disease starts, how it spreads and how to treat it in such a complex setting will require accurate preclinical models which best replicate this complexity.14
The preclinical landscape has recently changed, with the Food and Drug Administration (FDA) modernisation act 2.0 passing the U.S. senate in late 2022, permitting the FDA to accept in vitro evidence as sufficient for progression of novel therapeutics to clinical trials.15,16 More recently, the NIH made a further decision that grant proposals solely using animal models would no longer be funded.17 These changes emphasise the value of in vitro modeling and create an incentive to further advance our in vitro tools and technology in the race against ALS. For a model to be valid for ALS research, be that mechanistic or for drug testing, it must replicate the disease as closely as possible, utilizing cells that are suited to the research question. This review will cover the considerations, regarding cell type and environment, that are necessary to create a model of ALS in a dish. Further, we will reflect on the limitations that persist and the highlights of research breakthroughs that have already come from in vitro models.
2 Sourcing selectively vulnerable cell types for modeling ALS
Selective vulnerability of specific neuronal cell types is observed across many neurodegenerative conditions.18 ALS is characterized by the progressive degeneration of both upper and lower motor neurons, making these the primary cell types of interest for in vitro modeling of the disease (Fig. 1). Both these cell types possess structural and functional characteristics that are integral to their identity and likely play a significant role in their selective vulnerability in ALS. UMNs are glutamatergic, multipolar neurons characterised by a large pyramidal cell body that resides in layer V of the motor cortex. They have an apical dendrite extending to higher cortical layers with many branching basolateral dendrites, each covered in dendritic spines, and an axon projecting to the brain stem or spinal cord which synapses onto LMNs, either directly or via a spinal interneuron. LMNs are cholinergic multipolar neurons characterised by a large cell body residing in the ventral horn of the spinal cord and a long axon projecting into the periphery to synapse onto smooth or skeletal muscle via the highly specialised neuromuscular junction (NMJ). It is becoming apparent that upper and lower motor neurons experience unique types of dysfunctions in ALS and may even be differentially vulnerable to disease mechanisms.19–21 For example, LMNs with nuclear depletion of the RNA/DNA-binding protein TDP-43 exhibit cytoplasmic aggregates of phosphorylated TDP-43, whereas UMN nuclear loss of TDP-43 does not coincide with any cytoplasmic aggregate formation in human tissue.19 As such, it is important that in vitro models of ALS involving upper and lower motor neurons recapitulate key features of their identity to serve as a useful platform for screening new therapies or understanding disease mechanisms.2.1 Primary neurons
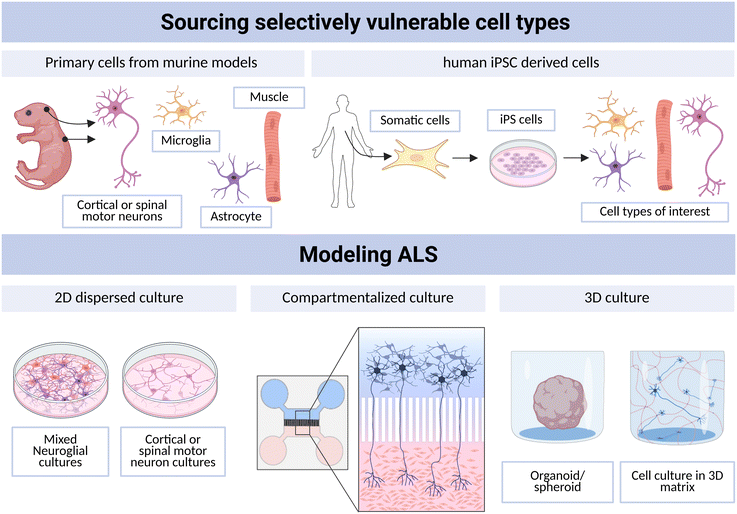
ALS has been studied in vitro using primary cells derived from animal models, such as rats or mice (Fig. 2). While primary cells can be derived from human tissues too, this is impracticable for neuronal tissues. Therefore, the term ‘primary neurons’ hereafter refers specifically to those obtained from rodent sources. Primary neurons undergo substantial differentiation in vivo before becoming post-mitotic, and therefore carry signatures of their unique identities into the dish. LMNs can be partially isolated from dissociated spinal cord tissue via centrifugation due to their large size in comparison to the other cells in the dissociated tissue, enabling production of cultures that are 90% pure for LMNs, as identified by expression of choline acetyl transferase (ChAT) and Islet1 (ISL1).22–24To drive ALS-associated pathological phenotypes in primary neurons, they can be sourced from transgenic animals, transfected with plasmids harbouring pathological genes, or treated with substances that drive ALS-associated pathologies. Primary culture models have been used to investigate the role of TDP-43 pathology in neurons.25,26 For example, Pisciottani et al. generated a model of TDP-43 proteinopathy in mouse cortical neurons by transfecting them with TDP-43 constructs. They found that transfected neurons exhibited characteristics such as cytoplasmic aggregation of TDP-43 (colocalised with stress granules), decreased axonal protein synthesis, oxidative stress, decreased spontaneous bursting activity, decreased synaptic vesical formation and decreased intracellular calcium levels.27 Importantly, these characteristics reflect those observed in animal models, human cell culture models and human postmortem tissue, suggesting that primary cell culture models can exhibit disease-relevant phenotypes.
A significant limitation of primary culture is the presence of species differences between rodents and humans that may impact the physiological relevance of these models to human disease. In addition, while LMNs can be partially isolated from spinal cord suspensions, some studies have identified that the survival of LMNs in high purity cultures is very low, given the necessity of non-neuronal cells, particularly muscle cells and astrocytes, for LMN identity and survival.28,29 Regarding isolation of primary UMNs, there are currently no reliable UMN specific markers, so there is no way to identify the UMNs from the other neuronal populations dissociated from brain tissue. This is a significant limitation given the specific vulnerability of UMNs to ALS pathology, and this problem continues to hinder our ability to establish authentic UMNs from stems cells. Neuronal-like cell lines, such as N2a, MN1 and NSC-34, offer the advantage of high throughput, cost effectiveness and lack of immediate ethical impact. However, as an ALS disease model, they possess inherent limitations related to their transformed nature and genetic instability driven by indefinite division. Alternatively, human embryonic stem cells (hESCs) can be differentiated into neurons and other relevant cell types, and further genetically modified to model ALS-associated pathologies or incorporated into complex 3D models of nervous tissue.30,31 However, their significant ethical implications and limited resourcing renders this option less attractive.32
2.2 Induced pluripotent stem cells
The development of induced pluripotent stem cells (iPSCs) was a pivotal breakthrough for in vitro studies, improving physiological relevance of cell models and reducing the dependence on animal models.33,34 In the context of neurodegenerative diseases, iPSC technology allows differentiation of cells from patients with neurodegenerative conditions into disease-relevant cell types (such as UMNs and LMNs) and subsequent comparison to those of unaffected individuals or, ideally, gene-corrected controls (Fig. 2). This allows significant insights into cellular pathological changes that arise from the unique genetic signature carried by individual patients.Despite being a pivotal addition to the in vitro toolkit, some limitations remain in iPSC technology. Importantly, our current inability to generate authentic iUMNs limits our ability to investigate UMN-specific contributions to disease progression. In addition, variation in iLMN differentiation protocols affects the yield, purity and phenotype of the resultant culture.60–62 ALS research has also revealed that specific subtypes of LMNs are more vulnerable to degeneration.63 Indeed, oculomotor neurons are resistant to degeneration, despite ubiquitous expression of pathogenic genetic variants, emphasizing the need to be able to effectively differentiate and identify subtypes of LMNs.64 In addition, some caveats exist that may be important to consider when interpreting iPSC data for translation. Namely, the retainment and erasure of different epigenetic markers in iPSCs may cause information to be lost regarding the contribution of non-genetic factors to neurodegenerative disease pathogenesis. While some studies show that epigenetic memory can be retained by specific methods of differentiation,65,66 the extent to which other epigenetic alterations are retained is not yet known. Additionally, a significant issue associated with the application of iPSCs to studying neurodegenerative diseases in general is the cellular rejuvenation that occurs during the iPSC reprogramming process, meaning that the resultant cells are not replicative of the age-affected cells that are so vulnerable in neurodegenerative diseases.67 Additional cell stressors and environmental factors can be employed to drive some forms of age-associated cellular damage.68,69 Recently, it has been shown that inhibition of epigenetic modifiers in neural progenitors drives rapid maturation of iPSC-derived neurons, perhaps paving the way to generating more physiologically relevant models for neurodegenerative diseases.70
3 Modelling ALS in vitro
Accurately modeling complex diseases requires more than the inclusion of relevant cell types, as the interplay between the cell types and their spatial arrangement may play a crucial role in disease pathology. A critical challenge lies in organising the cells into a physiologically relevant and experimentally compatible conformation. Here, we explain the principals of 2D compartmentalised culture and 3D culture systems and then discuss the experimental readouts compatible with these methods. Finally, we highlight recent findings from in vitro ALS models that demonstrate their utility in advancing our understanding of the disease, and point out current limitations.3.1 Compartmentalised 2D cultures
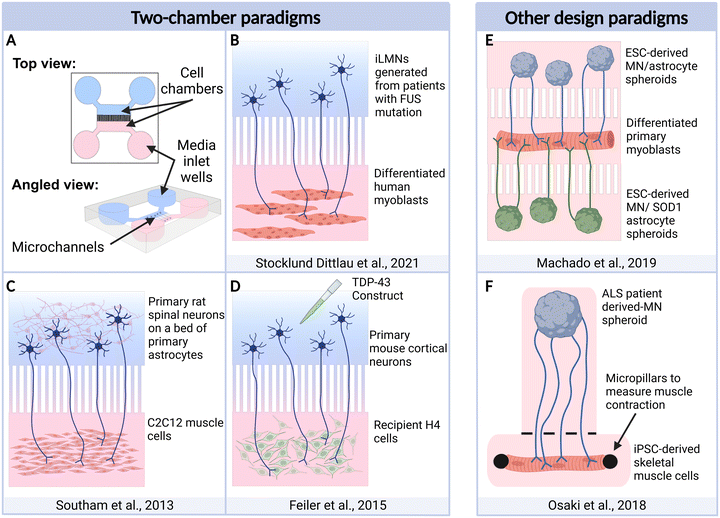
2-Dimensional (2D) culture systems consist of cells grown in a dispersed monolayer and is the simplest and most cost-effective form of in vitro research (Fig. 2). Historically, 2D culture has been the principal method of studying cellular mechanisms of disease as the conditions are largely controlled and cells can be examined in an isolated manner. 2D culture has led to significant advances in understanding pathobiological mechanisms of neurodegeneration but has limits to its translatability in complex diseases like ALS. Human neuroanatomy is highly organised, and cellular function and health is likely to be highly dependent on these organised connections. This is particularly true for studies of ALS, in which multiple regions of the nervous system are affected and there appears to be directional spread of pathology through circuits.71 As such, it is possible that 2D monocultures may provide false positives or false negatives when used to screen novel therapeutic agents. The development of microfluidic platforms for in vitro studies has offered immense progress to overcoming this limitation, providing a model for questions pertaining to network dysfunction and spread of disease. Whilst dispersed 2D culture methods are still valid, the explosion of microfluidic culture paradigms and 3D cultures present new opportunities to model ALS in a dish and interrogate disease progression and therapeutic options, with the intention of translation.For application to neuronal studies, a microfluidic device generally contains some configuration of 3 main elements: inlet and outlet wells where media/treatments are added, chambers where the cells reside, and microchannels connecting the chambers (Fig. 3A). Cells are deposited into the inlet wells, and capillary action pulls them into the chambers. An important feature of microfluidic models is the ability to promote fluid flow and/or a diffusion gradient through the fluid. Fluid flow can be achieved by manipulation of the volume at the inlets and outlets of the device, which results in passive pumping of fluid by capillary action and hydrostatic pressure, or by active fluid pumping through the device.79 This continuous fluid perfusion enables circulation of nutrients and removes waste, replicating the role of the fluid flow of cerebrospinal fluid (CSF) in vivo.80 In the context of ALS, circulation of CSF may be important in understanding how pathology and pathological cues travel through the central nervous system, as well as being an important aspect of consideration in therapeutic delivery.81 In addition, concentration gradients can be established in microfluidic systems by adding a substance (growth factor, solute, drug) at a high concentration to a particular region of the device, causing a diffusion gradient to passively establish through the device. For example, Chennampally et al. used a diffusion gradient of rapamycin across a culture of iPSC-derived motor neurons carrying the ALS-associated A315T mutation in the TARDBP gene. This enabled them to identify a therapeutic window: rapamycin concentrations between 0.4–1 μM rescued 40.44% of neurons, while lower concentrations were ineffective and higher concentrations were toxic.82 Low dose rapamycin has now been found to be safe for patients with ALS in a phase 2 clinical trial.83
Microfluidic models are a promising culture platform for ALS research and have shown great value already (see applications and outcomes section). However, there is currently significant variability in microfluidic models of ALS, in the specific design of the model, the fabrication process and the cell handling within the model (such as surface pre-treatment, seeding density, the culture medium used for multi-cell type models, and control of media circulation). While flexibility in design is a key strength of microfluidic modelling (particularly for mechanistic studies), a model of ALS for the purpose of therapeutic screening will need to be more consistent to ensure reproducibility. PDMS is also known to absorb small molecules, which is an important consideration for drug screening applications, and solutions for this issue, as explained by Gomez-Sjoberg et al., are yet to be broadly applied.77 In addition., with the added physiological relevance of compartmentalised cultures, there are some introduced stressors, such as physical sheer stress and altered soluble gas availability, that make culturing cells in microfluidic devices more challenging than in traditional cultureware.84 Specifically, seeding density, cell survival, media changing schedules and media components will all need to be reconsidered when translating from dispersed 2D culture to microfluidic culturing. Accordingly, microfluidic use can be manually arduous and require technical expertise, as much of the process cannot be, or is yet to be, automated in an academic laboratory setting.
3.2 Modeling ALS in 3D
The most recent advancement made to in vitro modeling has been the development of 3-dimensional (3D) culture systems. 3D culture systems include those that are 3D with regards to the cellular environment or 3D with regard to the arrangement of the cells themselves. Cell culture in a 3D environment refers to the seeding of cells into a 3D extracellular matrix-like scaffolding, within which the cells can move freely and interact with each other and the matrix. 3D tissue culture involves the co-culture of immature cells and depends on their directed self-assembly into 3D clusters (Fig. 2).85 These clusters (often referred to as organoids or spheroids) can be grown in a matrix or free-floating in culture media.Currently, there are some limitations on the use of 3D organoids and spheroids for the study of neurodegenerative disease. Importantly, recent single-cell transcriptomic characterisation of cerebral organoids suggests that even after being grown for 12 months, only 15% of the excitatory neurons show signatures of early post-natal stages, with most of the nuclei in the organoid aligning with early foetal signatures of human brains.100 This suggests that cerebral organoids are useful for modeling the developing brain, and thus clearly suited to the study of neurodevelopmental disorders, but their utility for modeling neurodegenerative disease may need further development. In addition, 3D culture methods (including cells grown in 3D matrices) are complex in nature, more expensive than 2D methods, and require technical expertise and large numbers of replicates to account for culture–culture variability.101 Regardless, organoids introduce an aspect of dimensionality to disease modeling that is otherwise unachievable, giving them the potential to significantly improve translational outcomes.
4 Readouts and experimental compatibility of microfluidic and 3D models of ALS
One of the key strengths of cell culture-based models compared to animal models is their high compatibility with diverse experimental manipulations and analytical readouts. This is essential for validating the models, probing mechanistic pathways and assessing therapeutic outcomes. Importantly, in culture-based modeling, the cellular environment is readily accessible for addition of experimental manipulations or therapeutics. In compartmentalised on-the-chip models, these treatments can be added in a site-specific manner by establishing fluidic isolation of one region from others. This is achieved through precise control of the volume of media feeding into the two chambers connected by microchannels.102–104 Further, concentration gradients and continuous fluidic perfusion can be established. Following experimental manipulation, readout methods such as electrophysiology, live/timelapse microscopy, calcium imaging, region-specific-omics studies, reporter assays, optogenetics and immunocytochemistry can all be adapted for microfluidic studies (Table S1).Electrophysiological outcomes are a central measure of neuronal and circuit health, and it is thus essential that electrophysiological monitoring is compatible with 2D and 3D cell-models. Traditional patch clamp electrophysiology remains a significant challenge in on-the-chip models, because while the media is accessible, the cells generally are not. However, microelectrode array technology has been successfully integrated into microfluidic models, enabling compartment specific measurements of spontaneous and evoked potentials, synaptic function and network function, and is compatible with 2D and 3D cell-models.75,105,106 In addition, the ease of accessibility means that treatments can be added to modify neuronal activity or mimic excitotoxicity, such as tetrodotoxin or kainic acid, respectively, and the cellular and network electrophysiological outcomes can be monitored in real time.107–109
Imaging-based readouts are also compatible with on-the-chip models. For 2D on-the-chip models, live cell microscopy can be used to observe axonal growth and cellular degeneration in real time, and calcium imaging can be used to observe intracellular calcium flux and thus indicate neuronal excitability and network function.110 Following cell fixation, immunocytochemistry can proceed as usual, allowing for identification of cell types, cell health markers and protein aggregation labelling. For 3D models, live imaging is often used to record the growth of organoids over time.106 At the end point of an experiment, organoids can be sectioned and fixed, making them compatible with immunohistochemistry. Prior to fixation, the sections, or even the whole organoid, can also be used for calcium imaging or patch clamp electrophysiology.111
Compartmentalised on-the-chip models also allow compartment-specific omics studies, allowing comparative omics investigations of the neuronal somatodendritic compartment versus axonal compartment, or cortical compartment versus spinal compartment.112 This provides a deep exposition on the gene and protein expression profiles in response to disease processes, potentially revealing mechanistic and compensatory mechanisms of specific cells in the circuit. 3D organoids can also be dissociated into a cell suspension and then used for omics studies.106
A shortcoming of many of these options are that these observation methods are, in their nature, end-point observations of the culture. Biosensor integration into on-the-chip models is revolutionising our ability to continuously monitor live culture analytics in real-time.113 Biosensors, by definition, are analytical tools that use biological molecules (enzymes or antibodies) to detect the presence of chemicals. Biosensor integration, however, more broadly refers to integration of readout methods into microfluidic models to capture complex biological or physiological readouts. For example, incorporation of MEAs into microfluidic devices is an example of biosensor integration. Biosensors have been developed to report (optically, magnetically or electrically) on culture properties such as levels of dissolved gases, pH, nutrients, metabolites and biomarkers, as well as cell-centric read outs such as cell growth and survival, electrical and mechanical properties, nucleocytoplasmic translocation, autophagy, and mitochondrial health.113–115 In models of ALS, biosensor integration can provide continuous, non-invasive readouts of physiological and pathological changes in the model, providing a dynamic window into ALS progression and therapeutic responses. As an example of biosensor integration into ALS microfluidic models, Uzel et al. tailored their on the chip model to have micropillars in the muscle compartment, between which the myofibers formed a muscle bundle, and the pillars were able to record the force of muscle contraction, providing a live readout of circuit function.116
Importantly, although many chemical biosensor assays exist, few are useful for long-term, continuous monitoring given their dependence on laser excitation, which causes photobleaching over time. As such, biosensor development that is not dependent on laser excitation for detection is currently an important area of research. For example, Zhang et al. developed a reactive oxygen species (ROS)-responsive probe that undergoes a structural change in the presence of ROS, resulting in a significantly enhanced Raman scattering signal detectable via surface-enhanced Raman spectroscopy (SERS).117 This approach demonstrated high specificity and sensitivity for ROS detection in vivo and in vitro, and its design is well-suited for adaptation to microfluidic platforms, where real-time, spatially resolved monitoring of oxidative stress could greatly enhance mechanistic studies and therapeutic screening.
The readout methods discussed here, while they are not exhaustive, have facilitated highly informative research in the ALS research space.
5 Applications and outcomes of ALS in vitro studies
The development of microfluidic techniques has revolutionised our capacity to investigate how pathology spreads through corticofugal tracts in ALS. For example, spread of pathological TAR DNA-binding protein 43 (TDP-43) appears to be an accurate predictor of symptom progression, but whether the spread of TDP-43 pathology is due to direct transmission of protein intercellularly or a consequence of increasing neuronal dysfunction remains unknown.71,118 2D dispersed culture investigations into TDP-43 pathology revealed that cytoplasmic expression of TDP-43 (be that WT or harbouring an fALS associated mutation A315T) is toxic to neurons, and that the amount of TDP-43 in the cytoplasm (but not in the nucleus) is a strong predictor of cell death.119,120 This provides an example of how simple in vitro paradigms can demystify cellular pathological phenomena observed in clinics or in vivo settings, but does not allow investigation of spread of pathology, given the disordered connections and lack of directional resolution. It has not been clear, for example, whether spread of TDP-43 requires contact between cells or if its transmission occurs predominantly via secretion of extracellular vesicles. To answer this question, Feiler et al. cultured primary mouse cortical neurons into one compartment of a microfluidic device and transfected them with a luciferase-tagged TDP-43 construct, or just the luciferase construct, and recipient H4 cells into the other compartment (Fig. 3D).121 This experiment revealed increased luciferase activity in the cultures treated with the TDP-43 construct compared to the control, demonstrating that TDP-43 can undergo directed anterograde, contact-mediated spread between cells via axons. By reversing the paradigm and adding TDP-43 to the dendritic compartment, the authors also identified retrograde spread of TDP-43, indicating bidirectional transmission of TDP-43. This platform of in vitro studies presents exciting opportunities to study how ALS pathology spreads through the corticomotor system, how different pathological mechanisms affect different cell types and what mechanisms protect some cell types from degeneration.Compartmentalised microfluidic platforms are particularly useful for studying neuromuscular junction (NMJ) pathology. The NMJ is a specialised synapse between motor neurons and muscle fibres. An action potential in the LMN leads to release of acetylcholine at the neuromuscular junction, which initiates a post-synaptic current in the muscle that ultimately results in muscle fibre contraction. The NMJ is highly vulnerable in ALS, and it is widely accepted that the disassembly of the NMJ is pivotal to symptom onset and progression.122 Early attempts at generating an in vitro model of the NMJ utilised human iLMNs with C2C12 mouse myoblasts in a 2D culture dish, and demonstrated functional NMJ formation through electrophysiological stimulation and post-synaptic current record.123 However, this approach lacks the spatial separation and distinct environments that LMNs and muscle cells naturally inhabit, limiting its physiological relevance. We have previously overcome these limitations by generating a microfluidic-based motor unit model by culturing embryonic rat LMNs on a bed of astrocytes in one compartment and rat myoblasts in another124 (Fig. 3C). With this protocol, motor neurons extended distally and formed neuromuscular junctions with myocytes as evidenced by clustered presynaptic synaptophysin and postsynaptic acetylcholine receptors.125 Using this model, we showed that rodent LMNs degenerate following somatodendritic, but not axonal, exposure to kainic acid, supporting the die-forward hypothesis of pathological spread in ALS. The addition of iPSC technology to microfluidic devices has improved the translatability of microfluidic motor unit models, combining the benefits of human cells and compartmentalised culture. For example, in 2021, Stoklund Dittlau et al. generated a microfluidic model of the human motor unit using iLMNs and human myoblasts (Fig. 3B). NMJ-like connections in the myoblast compartment were quantified in cultures grown from patients harbouring FUS mutations and CRISPR-corrected control cultures, and revealed that FUS mutant neurons, compared to controls, formed fewer NMJs and those that did form were less likely to have multiple contact points, indicative of defective maturation.126
As an example of the value of non-compartmentalised 3D culture, a recent study by Faustino Martins et al. showed the generation of a hESC-derived 3D, functional neuromuscular organoid that self-organised into neural and muscular regions. Additionally, the organoid formed functional NMJs that, upon stimulation of the neural region, were able to confer observable contractions in the muscle cells.106 The group also establish a neuromuscular organoid from iPSCs derived from a patient with spinal muscular atrophy, and the organoid exhibited reduced NMJ formation and contractility.127 In an effort to integrate a cortical region into a neuromuscular organoid, Andersen et al. generated organoids resembling the cerebral cortex, the spinal cord/hindbrain and skeletal muscle from human iPSCs and cultured them together to generate a corticomotor assembloid.128 Optogenetic stimulation of the cortical organoid was able to translate through the miniaturised circuit and cause recordable muscle contraction in the muscle organoid. Neuromuscular organoids and cerebrospinal-muscular assembloids formed from patient-derived iPSCs could form an invaluable platform to study motor dysfunction in ALS and screen novel therapies. However, these models do not exhibit the compartmentalised nature of the motor system as the neuronal and muscular cell types are co-cultured in the same media with no physical boundary and limited axonal distance between them.
In contrast, Machado et al. designed a compartmentalised device capable of housing spheroids (Fig. 3E).31 This device included three open-top compartments, where spheroids containing ESC-derived motor neurons (MNs) and astrocytes were placed in the outer compartments, while primary myofibers were positioned in the central compartment. The spheroids projected axons through the microchannels connecting the compartments and formed NMJs onto the myofiber. This format allowed the authors to assess the NMJ formation and functionality from different populations – and potentially genotypes – of neural/astrocyte spheroids, onto the same myofiber. They found that when the astrocytes in the MN/astrocyte spheroid were affected by a mutation in the SOD1 gene, the resultant NMJs exhibited reduced myofiber contractions. In 2018, Osaki et al. generated an elaborate human ALS on-a-chip model.95 iPSCs derived from a patient with sALS were differentiated into 3D skeletal muscle bundles as well as iLMNs and cultured into ECM scaffolded microfluidic devices, forming functional motor units (Fig. 3F). By optogenetically stimulating the iLMNs, the authors observed that the sALS-derived motor unit model generated fewer muscle contractions than controls, as well as aggregation of TDP-43 in the cytoplasm. Additionally, the LMN degeneration and contraction deficits were rescued with a combination of treatments, bosutinib and rapamycin, which have now been assessed in phase one and phase two clinical trials respectively.83,96
These examples present convincing evidence that in vitro models of ALS are valuable for questions ranging from simple, single cell mechanistic studies to complex, multicellular 3D modelling of pathological spread. In fact, of the three currently published examples of drug screening in iPSC culture translating into a clinical trial (bosutinib, ropinirole and retigabine), all utilised 2D monoculture of iPSC-derived motor neurons from ALS patients42,47,129 – confirming that simple monoculture systems can be informative for drug screening trials. Furthermore, these examples provide evidence that they are relevant, in that they reflect complex pathological signatures observed in humans and animal models, and that their translational potential is vast.49,83,96,130
6 Future directions and conclusion
In the future, in vitro models of ALS are likely to increase in complexity, model currently unknown aspects of disease, and identify additional drugs with translational potential. For example, not only will they have the necessary cell populations in the dish, but each in their own relevant extracellular matrix with associated supporting tissues such as vasculature, glial populations and immune cells. To support data collection from increasingly complex models, multiplexed biosensors—targeting aspects such as protein aggregation, oxidative and mitochondrial stress, and electrophysiological dysfunction—will need to be integrated into microfluidic devices. This integration will enable continuous monitoring of cell health, pathological spread, and therapeutic responses throughout the system. However, this should not be misinterpreted—complex, multi-cellular or multi-tissue models are not necessary for every ALS research question. There is substantial value in cell-autonomous studies, particularly for mechanistic insights. Nonetheless, for translational research, the use of more complex models becomes increasingly important. In the present, it is clear that the issues of clarifying cell identity in iPSC studies, our inability to generate bona fide iUMNs, and the limited cellular maturity/ageing associated with iPSC-derived cells continue to impede our ability to faithfully model ALS in a dish. Regarding translatability, the miniaturisation of model systems can lead to challenges in understanding dosage, toxicology, drug availability and delivery upon translation into the clinic. Specifically, models currently do not incorporate the role of the blood brain barrier or liver filtration in therapeutic function, meaning that direct translation of novel therapeutics (rather than repurposed therapeutics) from the dish to the clinic need to be carefully considered.Here, in the context of the FDA no longer necessitating animal model experiments to progress a drug to clinical trials, we have discussed the ways that in vitro models are being used to model and to study ALS. The purpose of this reflection is to identify the strengths and the limitations of our current in vitro models that may hinder their ability to be standalone preclinical platforms of drug discovery and mechanistic research. Promisingly, microfluidic devices and iPSC technology have revolutionised our ability to model human diseases in a compartmentalised way that is reflective of anatomical organisation. This has been particularly useful in the study of ALS given that it is a multi-compartment, multicellular disease that affects a very specific circuit. Furthermore, microfluidic models are inherently scalable, allowing researchers to run high-throughput, parallel experiments with precise control over cell types and conditions, making them ideal for drug screening, mechanistic studies, and personalised medicine. Indeed, new advances in in vitro technologies have enabled significant mechanistic discoveries and multiple instances of iPSC-based drug screens that have progressed to clinical trials. In contrast to the resource-intensive and ethically constrained nature of animal studies, in vitro models provide a relevant, scalable and ethically favourable alternative that is well-positioned to meet the demands of mechanistic and translational research.
Conflicts of interest
There are no conflicts of interest to declare.List of common abbreviations
| ALS | Amyotrophic lateral sclerosis |
| fALS | Familial amyotrophic lateral sclerosis |
| sALS | Sporadic amyotrophic lateral sclerosis |
| UMN | Upper motor neuron |
| LMN | Lower motor neuron |
| ESC | Embryonic stem cell |
| iPSC | Induced pluripotent stem cell |
| iLMN | Induced lower motor neuron |
| iUMN | Induced upper motor neuron |
| TDP-43 | TAR DNA binding protein 43 |
Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review. Supplementary information (SI): supplementary Table 1 details the studies mentioned in section 4 and the readout methods applied. See DOI: https://doi.org/10.1039/d5lc00577a.Acknowledgements
Tasmanian Masonic Medical Research Foundation. FightMND Mid-career Fellowship. The Massey Charitable Trust Grant.References
- K. Traxinger, C. Kelly, B. A. Johnson, R. H. Lyles and J. D. Glass, Prognosis and epidemiology of amyotrophic lateral sclerosis: Analysis of a clinic population, 1997-2011, Neurol. Clin. Pract., 2013, 3(4), 313–320 CrossRef.
- P. Mehta, D. K. Horton, E. J. Kasarskis, E. Tessaro, M. S. Eisenberg, S. Laird and J. Iskander, CDC Grand Rounds: National Amyotrophic Lateral Sclerosis (ALS) Registry Impact, Challenges, and Future Directions, MMWR Morb. Mortal. Wkly. Rep., 2017, 66(50), 1379–1382 CrossRef.
- E. P. McCann, K. L. Williams, J. A. Fifita, I. S. Tarr, J. O'Connor and D. B. Rowe, et al., The genotype-phenotype landscape of familial amyotrophic lateral sclerosis in Australia, Clin. Genet., 2017, 92(3), 259–266 CrossRef PubMed.
- G. Kim, O. Gautier, E. Tassoni-Tsuchida, X. R. Ma and A. D. Gitler, ALS Genetics: Gains, Losses, and Implications for Future Therapies, Neuron, 2020, 108(5), 822–842 CrossRef PubMed.
- R. Vasta, R. Chia, B. J. Traynor and A. Chiò, Unraveling the complex interplay between genes, environment, and climate in ALS, EBioMedicine, 2022, 75, 103795 CrossRef PubMed.
- A. Al-Chalabi, A. Calvo, A. Chio, S. Colville, C. M. Ellis and O. Hardiman, et al., Analysis of amyotrophic lateral sclerosis as a multistep process: a population-based modelling study, Lancet Neurol., 2014, 13(11), 1108–1113 CrossRef PubMed.
- O. Aousji, S. Feldengut, S. Antonucci, M. Schon, T. M. Boeckers and J. Matschke, et al., Patterns of synaptic loss in human amyotrophic lateral sclerosis spinal cord: a clinicopathological study, Acta Neuropathol. Commun., 2023, 11(1), 120 CrossRef CAS PubMed.
- P. Menon, M. Higashihara, M. van den Bos, N. Geevasinga, M. C. Kiernan and S. Vucic, Cortical hyperexcitability evolves with disease progression in ALS, Ann. Clin. Transl. Neurol., 2020, 7(5), 733–741 CrossRef PubMed.
- A. D. Calma, N. Pavey, P. Menon and S. Vucic, Neuroinflammation in amyotrophic lateral sclerosis: pathogenic insights and therapeutic implications, Curr. Opin. Neurol., 2024, 37(5), 585–592 CrossRef CAS PubMed.
- E. F. Smith, P. J. Shaw and K. J. De Vos, The role of mitochondria in amyotrophic lateral sclerosis, Neurosci. Lett., 2019, 710, 132933 CrossRef PubMed.
- S. J. Barmada and R. N. A. Linking, Dysfunction and Neurodegeneration in Amyotrophic Lateral Sclerosis, Neurotherapeutics, 2015, 12(2), 340–351 CrossRef CAS PubMed.
- A. Amin, N. D. Perera, P. M. Beart, B. J. Turner and F. Shabanpoor, Amyotrophic Lateral Sclerosis and Autophagy: Dysfunction and Therapeutic Targeting, Cell, 2020, 9(11), 2413 CrossRef CAS.
- P. Menon, N. Geevasinga, M. van den Bos, C. Yiannikas, M. C. Kiernan and S. Vucic, Cortical hyperexcitability and disease spread in amyotrophic lateral sclerosis, Eur. J. Neurol., 2017, 24(6), 816–824 CrossRef CAS PubMed.
- P. Van Damme, W. Robberecht and L. Van Den Bosch, Modelling amyotrophic lateral sclerosis: progress and possibilities, Dis. Models Mech., 2017, 10(5), 537–549 CrossRef CAS.
- Consolidated Appropriations Act, S., 2023, 3209.
- E. Y. Adashi, D. P. O'Mahony and I. G. Cohen, The FDA Modernization Act 2.0: Drug Testing in Animals is Rendered Optional, Am. J. Med., 2023, 136(9), 853–854 CrossRef CAS.
- NIH, NIH to prioritize human-based research technologies, 2025, Available from: https://nihrecord.nih.gov/2025/07/04/nih-prioritize-human-based-research-technologies.
- M. Kampmann, Molecular and cellular mechanisms of selective vulnerability in neurodegenerative diseases, Nat. Rev. Neurosci., 2024, 25(5), 351–371 CrossRef CAS.
- H. Braak, A. C. Ludolph, M. Neumann, J. Ravits and K. Del Tredici, Pathological TDP-43 changes in Betz cells differ from those in bulbar and spinal alpha-motoneurons in sporadic amyotrophic lateral sclerosis, Acta Neuropathol., 2017, 133(1), 79–90 CrossRef CAS.
- L. A. Reale, M. S. Dyer, S. E. Perry, K. M. Young, T. C. Dickson and A. Woodhouse, et al., Pathologically mislocalised TDP-43 in upper motor neurons causes a die-forward spread of ALS-like pathogenic changes throughout the mouse corticomotor system, Prog. Neurobiol., 2023, 226, 102449 CrossRef CAS PubMed.
- A. E. King, T. C. Dickson, C. A. Blizzard, S. S. Foster, R. S. Chung and A. K. West, et al., Excitotoxicity mediated by non-NMDA receptors causes distal axonopathy in long-term cultured spinal motor neurons, Eur. J. Neurosci., 2007, 26(8), 2151–2159 CrossRef CAS.
- M. J. Beaudet, Q. Yang, S. Cadau, M. Blais, S. Bellenfant and F. Gros-Louis, et al., High yield extraction of pure spinal motor neurons, astrocytes and microglia from single embryo and adult mouse spinal cord, Sci. Rep., 2015, 5, 16763 CrossRef CAS.
- H. M. Zhang, N. Robinson, I. Gomez-Curet, W. Wang and M. A. Harrington, Neuronal and network activity in networks of cultured spinal motor neurons, NeuroReport, 2009, 20(9), 849–854 CrossRef PubMed.
- D. J. Graber and B. T. Harris, Purification and culture of spinal motor neurons from rat embryos, Cold Spring Harb. Protoc., 2013, 2013(4), 319–326 Search PubMed.
- S. J. Barmada, G. Skibinski, E. Korb, E. J. Rao, J. Y. Wu and S. Finkbeiner, Cytoplasmic Mislocalization of TDP-43 Is Toxic to Neurons and Enhanced by a Mutation Associated with Familial Amyotrophic Lateral Sclerosis, J. Neurosci., 2010, 30(2), 639 CrossRef CAS.
- R. A. K. Atkinson, H. L. Fair, R. Wilson, J. C. Vickers and A. E. King, Effects of TDP-43 overexpression on neuron proteome and morphology in vitro, Mol. Cell. Neurosci., 2021, 114, 103627 CrossRef CAS.
- A. Pisciottani, L. Croci, F. Lauria, C. Marullo, E. Savino and A. Ambrosi, et al., Neuronal models of TDP-43 proteinopathy display reduced axonal translation, increased oxidative stress, and defective exocytosis, Front. Cell. Neurosci., 2023, 17, 1253543 CrossRef CAS PubMed.
- M. Bucchia, S. J. Merwin, D. B. Re and S. Kariya, Limitations and Challenges in Modeling Diseases Involving Spinal Motor Neuron Degeneration in Vitro, Front. Cell. Neurosci., 2018, 12, 61 CrossRef.
- I. Vargova, J. Kriska, J. C. F. Kwok, J. W. Fawcett and P. Jendelova, Long-Term Cultures of Spinal Cord Interneurons, Front. Cell. Neurosci., 2022, 16, 827628 CrossRef CAS PubMed.
- W. Guo, M. Naujock, L. Fumagalli, T. Vandoorne, P. Baatsen and R. Boon, et al., HDAC6 inhibition reverses axonal transport defects in motor neurons derived from FUS-ALS patients, Nat. Commun., 2017, 8(1), 861 CrossRef PubMed.
- C. B. Machado, P. Pluchon, P. Harley, M. Rigby, V. G. Sabater and D. C. Stevenson, et al., In Vitro Modeling of Nerve–Muscle Connectivity in a Compartmentalized Tissue Culture Device, Adv. Biosyst., 2019, 3(7), 1800307 CrossRef.
- W. Zakrzewski, M. Dobrzynski, M. Szymonowicz and Z. Rybak, Stem cells: past, present, and future, Stem Cell Res. Ther., 2019, 10(1), 68 CrossRef CAS PubMed.
- K. Takahashi and S. Yamanaka, Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors, Cell, 2006, 126(4), 663–676 CrossRef CAS PubMed.
- K. Takahashi, K. Tanabe, M. Ohnuki, M. Narita, T. Ichisaka and K. Tomoda, et al., Induction of pluripotent stem cells from adult human fibroblasts by defined factors, Cell, 2007, 131(5), 861–872 CrossRef CAS.
- M. A. Saporta, V. Dang, D. Volfson, B. Zou, X. S. Xie and A. Adebola, et al., Axonal Charcot-Marie-Tooth disease patient-derived motor neurons demonstrate disease-specific phenotypes including abnormal electrophysiological properties, Exp. Neurol., 2015, 263, 190–199 CrossRef CAS.
- S. M. Chambers, C. A. Fasano, E. P. Papapetrou, M. Tomishima, M. Sadelain and L. Studer, Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling, Nat. Biotechnol., 2009, 27(3), 275–280 CrossRef CAS.
- F. Bianchi, M. Malboubi, Y. Li, J. H. George, A. Jerusalem and F. Szele, et al., Rapid and efficient differentiation of functional motor neurons from human iPSC for neural injury modelling, Stem Cell Res., 2018, 32, 126–134 CrossRef CAS.
- M. J. Workman, R. G. Lim, J. Wu, A. Frank, L. Ornelas and L. Panther, et al., Large-scale differentiation of iPSC-derived motor neurons from ALS and control subjects, Neuron, 2023, 111(8), 1191–1204 e5 CrossRef CAS PubMed.
- L. Thiry, R. Hamel, S. Pluchino, T. Durcan and S. Stifani, Characterization of Human iPSC-derived Spinal Motor Neurons by Single-cell RNA Sequencing, Neuroscience, 2020, 450, 57–70 CrossRef CAS PubMed.
- M. J. Castellanos-Montiel, M. Chaineau, A. K. Franco-Flores, G. Haghi, D. Carrillo-Valenzuela and W. E. Reintsch, et al., An Optimized Workflow to Generate and Characterize iPSC-Derived Motor Neuron (MN) Spheroids, Cell, 2023, 12(4), 545 CrossRef CAS.
- L. Thiry, J. P. Clement, R. Haag, T. E. Kennedy and S. Stifani, Optimization of Long-Term Human iPSC-Derived Spinal Motor Neuron Culture Using a Dendritic Polyglycerol Amine-Based Substrate, ASN Neuro, 2022, 14, 17590914211073381 CrossRef CAS PubMed.
- K. Fujimori, M. Ishikawa, A. Otomo, N. Atsuta, R. Nakamura and T. Akiyama, et al., Modeling sporadic ALS in iPSC-derived motor neurons identifies a potential therapeutic agent, Nat. Med., 2018, 24(10), 1579–1589 CrossRef CAS PubMed.
- C. J. Alves, R. Dariolli, F. M. Jorge, M. R. Monteiro, J. R. Maximino and R. S. Martins, et al., Gene expression profiling for human iPS-derived motor neurons from sporadic ALS patients reveals a strong association between mitochondrial functions and neurodegeneration, Front. Cell. Neurosci., 2015, 9, 289 Search PubMed.
- N. Ichiyanagi, K. Fujimori, M. Yano, C. Ishihara-Fujisaki, T. Sone and T. Akiyama, et al., Establishment of In Vitro FUS-Associated Familial Amyotrophic Lateral Sclerosis Model Using Human Induced Pluripotent Stem Cells, Stem Cell Rep., 2016, 6(4), 496–510 CrossRef PubMed.
- H. Chen, K. Qian, Z. Du, J. Cao, A. Petersen and H. Liu, et al., Modeling ALS with iPSCs reveals that mutant SOD1 misregulates neurofilament balance in motor neurons, Cell Stem Cell, 2014, 14(6), 796–809 CrossRef PubMed.
- A. C. Devlin, K. Burr, S. Borooah, J. D. Foster, E. M. Cleary and I. Geti, et al., Human iPSC-derived motoneurons harbouring TARDBP or C9ORF72 ALS mutations are dysfunctional despite maintaining viability, Nat. Commun., 2015, 6, 5999 CrossRef.
- B. J. Wainger, E. Kiskinis, C. Mellin, O. Wiskow, S. S. Han and J. Sandoe, et al., Intrinsic membrane hyperexcitability of amyotrophic lateral sclerosis patient-derived motor neurons, Cell Rep., 2014, 7(1), 1–11 CrossRef.
- M. F. Burkhardt, F. J. Martinez, S. Wright, C. Ramos, D. Volfson and M. Mason, et al., A cellular model for sporadic ALS using patient-derived induced pluripotent stem cells, Mol. Cell. Neurosci., 2013, 56, 355–364 CrossRef PubMed.
- S. Morimoto, S. Takahashi, D. Ito, Y. Date, K. Okada and C. Kato, et al., Phase 1/2a clinical trial in ALS with ropinirole, a drug candidate identified by iPSC drug discovery, Cell Stem Cell, 2023, 30(6), 766–780 e9 CrossRef.
- K. Autar, X. Guo, J. W. Rumsey, C. J. Long, N. Akanda and M. Jackson, et al., A functional hiPSC-cortical neuron differentiation and maturation model and its application to neurological disorders, Stem Cell Rep., 2022, 17(1), 96–109 Search PubMed.
- S. Y. Cao, Y. Hu, C. Chen, F. Yuan, M. Xu and Q. Li, et al., Enhanced derivation of human pluripotent stem cell-derived cortical glutamatergic neurons by a small molecule, Sci. Rep., 2017, 7(1), 3282 Search PubMed.
- G. Wali, Y. Li, D. Abu-Bonsrah, D. Kirik, C. L. Parish and C. M. Sue, Generation of human-induced pluripotent-stem-cell-derived cortical neurons for high-throughput imaging of neurite morphology and neuron maturation, STAR Protoc., 2023, 4(2), 102325 CrossRef PubMed.
- T. Uchimura, J. Otomo, M. Sato and H. Sakurai, A human iPS cell myogenic differentiation system permitting high-throughput drug screening, Stem Cell Res., 2017, 25, 98–106 CrossRef.
- A. Tanaka, K. Woltjen, K. Miyake, A. Hotta, M. Ikeya and T. Yamamoto, et al., Correction: Efficient and Reproducible Myogenic Differentiation from Human iPS Cells: Prospects for Modeling Miyoshi Myopathy In Vitro, PLoS One, 2013, 8(12), e61540 CrossRef PubMed.
- S. Selvaraj, R. Mondragon-Gonzalez, B. Xu, A. Magli, H. Kim and J. Lainé, et al., Screening identifies small molecules that enhance the maturation of human pluripotent stem cell-derived myotubes, eLife, 2019, 8, e47970 CrossRef.
- C. Cheung and S. Sinha, Human embryonic stem cell-derived vascular smooth muscle cells in therapeutic neovascularisation, J. Mol. Cell. Cardiol., 2011, 51(5), 651–664 CrossRef PubMed.
- F. J. Steyn, R. Li, S. E. Kirk, T. W. Tefera, T. Y. Xie and T. J. Tracey, et al., Altered skeletal muscle glucose-fatty acid flux in amyotrophic lateral sclerosis, Brain Commun., 2020, 2(2), fcaa154 CrossRef PubMed.
- A. Badu-Mensah, X. Guo, C. W. McAleer, J. W. Rumsey and J. J. Hickman, Functional skeletal muscle model derived from SOD1-mutant ALS patient iPSCs recapitulates hallmarks of disease progression, Sci. Rep., 2020, 10(1), 14302 CrossRef PubMed.
- B. F. Vahsen, E. Gray, A. G. Thompson, O. Ansorge, D. C. Anthony and S. A. Cowley, et al., Non-neuronal cells in amyotrophic lateral sclerosis - from pathogenesis to biomarkers, Nat. Rev. Neurol., 2021, 17(6), 333–348 CrossRef PubMed.
- C. M. Castillo Bautista and J. Sterneckert, Progress and challenges in directing the differentiation of human iPSCs into spinal motor neurons, Front. Cell Dev. Biol., 2023, 10, 1089970 CrossRef PubMed.
- S. Sances, L. I. Bruijn, S. Chandran, K. Eggan, R. Ho and J. R. Klim, et al., Modeling ALS with motor neurons derived from human induced pluripotent stem cells, Nat. Neurosci., 2016, 19(4), 542–553 CrossRef PubMed.
- E. Giacomelli, B. F. Vahsen, E. L. Calder, Y. Xu, J. Scaber and E. Gray, et al., Human stem cell models of neurodegeneration: From basic science of amyotrophic lateral sclerosis to clinical translation, Cell Stem Cell, 2022, 29(1), 11–35 CrossRef PubMed.
- A. M. G. Ragagnin, S. Shadfar, M. Vidal, M. S. Jamali and J. D. Atkin, Motor Neuron Susceptibility in ALS/FTD, Front. Neurosci., 2019, 13, 532 CrossRef PubMed.
- J. Nijssen, L. H. Comley and E. Hedlund, Motor neuron vulnerability and resistance in amyotrophic lateral sclerosis, Acta Neuropathol., 2017, 133(6), 863–885 CrossRef.
- K. Kim, A. Doi, B. Wen, K. Ng, R. Zhao and P. Cahan, et al., Epigenetic memory in induced pluripotent stem cells, Nature, 2010, 467(7313), 285–290 CrossRef.
- L. de Boni, G. Gasparoni, C. Haubenreich, S. Tierling, I. Schmitt and M. Peitz, et al., DNA methylation alterations in iPSC- and hESC-derived neurons: potential implications for neurological disease modeling, Clin. Epigenet., 2018, 10, 13 CrossRef.
- R. Ho, S. Sances, G. Gowing, M. W. Amoroso, J. G. O'Rourke and A. Sahabian, et al., ALS disrupts spinal motor neuron maturation and aging pathways within gene co-expression networks, Nat. Neurosci., 2016, 19(9), 1256–1267 CrossRef PubMed.
- J. D. Miller, Y. M. Ganat, S. Kishinevsky, R. L. Bowman, B. Liu and E. Y. Tu, et al., Human iPSC-based modeling of late-onset disease via progerin-induced aging, Cell Stem Cell, 2013, 13(6), 691–705 CrossRef.
- E. Vera, N. Bosco and L. Studer, Generating Late-Onset Human iPSC-Based Disease Models by Inducing Neuronal Age-Related Phenotypes through Telomerase Manipulation, Cell Rep., 2016, 17(4), 1184–1192 CrossRef.
- G. Ciceri, A. Baggiolini, H. S. Cho, M. Kshirsagar, S. Benito-Kwiecinski and R. M. Walsh, et al., An epigenetic barrier sets the timing of human neuronal maturation, Nature, 2024, 626(8000), 881–890 CrossRef.
- J. Brettschneider, K. Del Tredici, J. B. Toledo, J. L. Robinson, D. J. Irwin and M. Grossman, et al., Stages of pTDP-43 pathology in amyotrophic lateral sclerosis, Ann. Neurol., 2013, 74(1), 20–38 CrossRef PubMed.
- R. B. Campenot, Local control of neurite development by nerve growth factor, Proc. Natl. Acad. Sci. U. S. A., 1977, 74(10), 4516–4519 CrossRef.
- W. X. Chen, J. G. Li, X. H. Wan, X. S. Zou, S. Y. Qi and Y. Q. Zhang, et al., Design of a microfluidic chip consisting of micropillars and its use for the enrichment of nasopharyngeal cancer cells, Oncol. Lett., 2019, 17(2), 1581–1588 Search PubMed.
- A. T. O'Neill, N. A. Monteiro-Riviere and G. M. Walker, Microfabricated curtains for controlled cell seeding in high throughput microfluidic systems, Lab Chip, 2009, 9(12), 1756–1762 RSC.
- P. Duc, M. Vignes, G. Hugon, A. Sebban, G. Carnac and E. Malyshev, et al., Human neuromuscular junction on micro-structured microfluidic devices implemented with a custom micro electrode array (MEA), Lab Chip, 2021, 21(21), 4223–4236 RSC.
- J. B. Nielsen, R. L. Hanson, H. M. Almughamsi, C. Pang, T. R. Fish and A. T. Woolley, Microfluidics: Innovations in Materials and Their Fabrication and Functionalization, Anal. Chem., 2020, 92(1), 150–168 CrossRef CAS.
- R. Gomez-Sjoberg, A. A. Leyrat, B. T. Houseman, K. Shokat and S. R. Quake, Biocompatibility and reduced drug absorption of sol-gel-treated poly(dimethyl siloxane) for microfluidic cell culture applications, Anal. Chem., 2010, 82(21), 8954–8960 CrossRef CAS PubMed.
- T. Ching, X. Nie, S.-Y. Chang, Y.-C. Toh and M. Hashimoto, Chapter 1 - Techniques and materials for the fabrication of microfluidic devices, in Principles of Human Organs-on-Chips, ed. M. Mozafari, Woodhead Publishing, 2023, pp. 1–36 Search PubMed.
- F. Nebuloni, Q. B. Do, P. R. Cook, E. J. Walsh and R. Wade-Martins, A fluid-walled microfluidic platform for human neuron microcircuits and directed axotomy, Lab Chip, 2024, 24(13), 3252–3264 RSC.
- T. Brinker, E. Stopa, J. Morrison and P. Klinge, A new look at cerebrospinal fluid circulation, Fluids Barriers CNS, 2014, 11, 10 CrossRef.
- K. C. Ng Kee Kwong, A. R. Mehta, M. Nedergaard and S. Chandran, Defining novel functions for cerebrospinal fluid in ALS pathophysiology, Acta Neuropathol. Commun., 2020, 8(1), 140 CrossRef CAS.
- P. Chennampally, A. Sayed-Zahid, P. Soundararajan, J. Sharp, G. A. Cox and S. D. Collins, et al., Author Correction: A microfluidic approach to rescue ALS motor neuron degeneration using rapamycin, Sci. Rep., 2021, 11(1), 19743 CrossRef CAS.
- J. Mandrioli, R. D'Amico, E. Zucchi, S. De Biasi, F. Banchelli and I. Martinelli, et al., Randomized, double-blind, placebo-controlled trial of rapamycin in amyotrophic lateral sclerosis, Nat. Commun., 2023, 14(1), 4970 CrossRef CAS.
- S. Halldorsson, E. Lucumi, R. Gomez-Sjoberg and R. M. T. Fleming, Advantages and challenges of microfluidic cell culture in polydimethylsiloxane devices, Biosens. Bioelectron., 2015, 63, 218–231 CrossRef CAS PubMed.
- C. Jensen and Y. Teng, Is It Time to Start Transitioning From 2D to 3D Cell Culture?, Front. Mol. Biosci., 2020, 7, 00033 CrossRef CAS.
- L. Rao, Y. Qian, A. Khodabukus, T. Ribar and N. Bursac, Engineering human pluripotent stem cells into a functional skeletal muscle tissue, Nat. Commun., 2018, 9(1), 126 CrossRef PubMed.
- A. Khodabukus, N. Prabhu, J. Wang and N. Bursac, In Vitro Tissue-Engineered Skeletal Muscle Models for Studying Muscle Physiology and Disease, Adv. Healthcare Mater., 2018, 7(15), e1701498 CrossRef.
- A. Shima, Y. Morimoto, H. L. Sweeney and S. Takeuchi, Three-dimensional contractile muscle tissue consisting of human skeletal myocyte cell line, Exp. Cell Res., 2018, 370(1), 168–173 CrossRef CAS PubMed.
- M. Afshar Bakooshli, E. S. Lippmann, B. Mulcahy, N. Iyer, C. T. Nguyen and K. Tung, et al., A 3D culture model of innervated human skeletal muscle enables studies of the adult neuromuscular junction, eLife, 2019, 8, e44530 CrossRef.
- R. Csapo, M. Gumpenberger and B. Wessner, Skeletal Muscle Extracellular Matrix - What Do We Know About Its Composition, Regulation, and Physiological Roles? A Narrative Review, Front. Physiol., 2020, 11, 253 CrossRef.
- A. Soles, A. Selimovic, K. Sbrocco, F. Ghannoum, K. Hamel and E. L. Moncada, et al., Extracellular Matrix Regulation in Physiology and in Brain Disease, Int. J. Mol. Sci., 2023, 24(8), 7049 CrossRef CAS.
- R. J. Bueno, C. Fernandez-Zapata, M. Salla, J. Campo Garcia, A. Alacam and O. Klein, et al., Generation of decellularized human brain tissue for investigating cell-matrix interactions: a proof-of-concept study, Front. Bioeng. Biotechnol., 2025, 13, 1578467 CrossRef PubMed.
- A. N. Cho, Y. Jin, Y. An, J. Kim, Y. S. Choi and J. S. Lee, et al., Microfluidic device with brain extracellular matrix promotes structural and functional maturation of human brain organoids, Nat. Commun., 2021, 12(1), 4730 CrossRef CAS.
- D. Lam, H. A. Enright, J. Cadena, S. K. G. Peters, A. P. Sales and J. J. Osburn, et al., Tissue-specific extracellular matrix accelerates the formation of neural networks and communities in a neuron-glia co-culture on a multi-electrode array, Sci. Rep., 2019, 9(1), 4159 CrossRef.
- T. Osaki, S. G. M. Uzel and R. D. Kamm, Microphysiological 3D model of amyotrophic lateral sclerosis (ALS) from human iPS-derived muscle cells and optogenetic motor neurons, Sci. Adv., 2018, 4(10), eaat5847 CrossRef CAS PubMed.
- K. Imamura, Y. Izumi, M. Nagai, K. Nishiyama, Y. Watanabe and R. Hanajima, et al., Safety and tolerability of bosutinib in patients with amyotrophic lateral sclerosis (iDReAM study): A multicentre, open-label, dose-escalation phase 1 trial, EClinicalMedicine, 2022, 53, 101707 CrossRef PubMed.
- O. L. Eichmuller and J. A. Knoblich, Human cerebral organoids - a new tool for clinical neurology research, Nat. Rev. Neurol., 2022, 18(11), 661–680 CrossRef PubMed.
- P. Korhonen, T. Malm and A. R. White, 3D human brain cell models: New frontiers in disease understanding and drug discovery for neurodegenerative diseases, Neurochem. Int., 2018, 120, 191–199 CrossRef CAS PubMed.
- A. E. Trevino, N. Sinnott-Armstrong, J. Andersen, S. J. Yoon, N. Huber and J. K. Pritchard, et al., Chromatin accessibility dynamics in a model of human forebrain development, Science, 2020, 367(6476), eaay1645 CrossRef CAS.
- C. A. Herring, R. K. Simmons, S. Freytag, D. Poppe, J. J. D. Moffet and J. Pflueger, et al., Human prefrontal cortex gene regulatory dynamics from gestation to adulthood at single-cell resolution, Cell, 2022, 185(23), 4428–4447.e28 CrossRef CAS PubMed.
- J. Temple, E. Velliou, M. Shehata and R. Levy, Current strategies with implementation of three-dimensional cell culture: the challenge of quantification, Interface Focus, 2022, 12(5), 20220019 CrossRef PubMed.
- A. M. Taylor, M. Blurton-Jones, S. W. Rhee, D. H. Cribbs, C. W. Cotman and N. L. Jeon, A microfluidic culture platform for CNS axonal injury, regeneration and transport, Nat. Methods, 2005, 2(8), 599–605 CrossRef CAS PubMed.
- K. A. Hosie, A. E. King, C. A. Blizzard, J. C. Vickers and T. C. Dickson, Chronic excitotoxin-induced axon degeneration in a compartmented neuronal culture model, ASN Neuro, 2012, 4(1), e00076 CrossRef PubMed.
- G. Robertson, T. J. Bushell and M. Zagnoni, Chemically induced synaptic activity between mixed primary hippocampal co-cultures in a microfluidic system, Integr. Biol., 2014, 6(6), 636–644 CrossRef CAS.
- Y. Ren, S.-H. Huang, S. Mosser, M. O. Heuschkel, A. Bertsch and P. C. Fraering, et al., A Simple and Reliable PDMS and SU-8 Irreversible Bonding Method and Its Application on a Microfluidic-MEA Device for Neuroscience Research, Micromachines, 2015, 6(12), 1923–1934 CrossRef.
- J. M. Faustino Martins, C. Fischer, A. Urzi, R. Vidal, S. Kunz and P. L. Ruffault, et al., Self-Organizing 3D Human Trunk Neuromuscular Organoids, Cell Stem Cell, 2020, 27(3), 498 CrossRef PubMed.
- X. Y. Zheng, H. L. Zhang, Q. Luo and J. Zhu, Kainic acid-induced neurodegenerative model: potentials and limitations, J. Biomed. Biotechnol., 2011, 2011, 457079 Search PubMed.
- D. I. Carrasco, E. K. Bichler, M. M. Rich, X. Wang, K. L. Seburn and M. J. Pinter, Motor terminal degeneration unaffected by activity changes in SOD1G93A mice; a possible role for glycolysis, Neurobiol. Dis., 2012, 48(1), 132–140 Search PubMed.
- J. L. Cheng, A. L. Cook, J. Talbot and S. Perry, How is Excitotoxicity Being Modelled in iPSC-Derived Neurons?, Neurotoxic. Res., 2024, 42(5), 43 Search PubMed.
- N. Tian, K. A. Hanson, A. J. Canty, J. C. Vickers and A. E. King, Microtubule-dependent processes precede pathological calcium influx in excitotoxin-induced axon degeneration, J. Neurochem., 2020, 152(5), 542–555 CrossRef PubMed.
- F. Ciarpella, R. G. Zamfir, A. Campanelli, E. Ren, G. Pedrotti and E. Bottani, et al., Murine cerebral organoids develop network of functional neurons and hippocampal brain region identity, iScience, 2021, 24(12), 103438 Search PubMed.
- J. Nijssen, J. Aguila, R. Hoogstraaten, N. Kee and E. Hedlund, Axon-Seq Decodes the Motor Axon Transcriptome and Its Modulation in Response to ALS, Stem Cell Rep., 2018, 11(6), 1565–1578 CrossRef PubMed.
- E. Ferrari, C. Palma, S. Vesentini, P. Occhetta and M. Rasponi, Integrating Biosensors in Organs-on-Chip Devices: A Perspective on Current Strategies to Monitor Microphysiological Systems, Biosensors, 2020, 10(9), 110 CrossRef.
- M. Gheorghiu, A short review on cell-based biosensing: challenges and breakthroughs in biomedical analysis, J. Biomed. Res., 2020, 35(4), 255–263 Search PubMed.
- R. De Michele, F. Carimi and W. B. Frommer, Mitochondrial biosensors, Int. J. Biochem. Cell Biol., 2014, 48, 39–44 CrossRef PubMed.
- S. G. Uzel, R. J. Platt, V. Subramanian, T. M. Pearl, C. J. Rowlands and V. Chan, et al., Microfluidic device for the formation of optically excitable, three-dimensional, compartmentalized motor units, Sci. Adv., 2016, 2(8), e1501429 CrossRef.
- D. Zhang, L. Zhou, Z. Nie, W. Zhang, F. Yu and R. Wang, ROS-responsive SERS probes for real-time monitoring of oxidative stress in traumatic brain injury, Chem. Commun., 2025, 61(67), 12510–12513 RSC.
- H. Braak, J. Brettschneider, A. C. Ludolph, V. M. Lee, J. Q. Trojanowski and K. Del Tredici, Amyotrophic lateral sclerosis--a model of corticofugal axonal spread, Nat. Rev. Neurol., 2013, 9(12), 708–714 CrossRef PubMed.
- E. Y. Liu, J. Russ, C. P. Cali, J. M. Phan, A. Amlie-Wolf and E. B. Lee, Loss of Nuclear TDP-43 Is Associated with Decondensation of LINE Retrotransposons, Cell Rep., 2019, 27(5), 1409–1421 e6 CrossRef PubMed.
- J. R. Highley, J. Kirby, J. A. Jansweijer, P. S. Webb, C. A. Hewamadduma and P. R. Heath, et al., Loss of nuclear TDP-43 in amyotrophic lateral sclerosis (ALS) causes altered expression of splicing machinery and widespread dysregulation of RNA splicing in motor neurones, Neuropathol. Appl. Neurobiol., 2014, 40(6), 670–685 CrossRef.
- M. S. Feiler, B. Strobel, A. Freischmidt, A. M. Helferich, J. Kappel and B. M. Brewer, TDP-43 is intercellularly transmitted across axon terminals, J. Cell Biol., 2015, 211(4), 897–911 Search PubMed.
- V. Cappello and M. Francolini, Neuromuscular Junction Dismantling in Amyotrophic Lateral Sclerosis, Int. J. Mol. Sci., 2017, 18(10), 2092 CrossRef PubMed.
- J. A. Umbach, K. L. Adams, C. B. Gundersen and B. G. Novitch, Functional neuromuscular junctions formed by embryonic stem cell-derived motor neurons, PLoS One, 2012, 7(5), e36049 CrossRef PubMed.
- K. A. Southam, A. E. King, C. A. Blizzard, G. H. McCormack and T. C. Dickson, Microfluidic primary culture model of the lower motor neuron-neuromuscular junction circuit, J. Neurosci. Methods, 2013, 218(2), 164–169 CrossRef PubMed.
- C. A. Blizzard, K. A. Southam, E. Dawkins, K. E. Lewis, A. E. King and J. A. Clark, et al., Identifying the primary site of pathogenesis in amyotrophic lateral sclerosis - vulnerability of lower motor neurons to proximal excitotoxicity, Dis. Models Mech., 2015, 8(3), 215–224 CrossRef PubMed.
- K. Stoklund Dittlau, E. N. Krasnow, L. Fumagalli, T. Vandoorne, P. Baatsen and A. Kerstens, et al., Human motor units in microfluidic devices are impaired by FUS mutations and improved by HDAC6 inhibition, Stem Cell Rep., 2021, 16(9), 2213–2227 CrossRef.
- A. Urzi, I. Lahmann, L. V. N. Nguyen, B. R. Rost, A. Garcia-Perez and N. Lelievre, et al., Efficient generation of a self-organizing neuromuscular junction model from human pluripotent stem cells, Nat. Commun., 2023, 14(1), 8043 CrossRef.
- J. Andersen, O. Revah, Y. Miura, N. Thom, N. D. Amin and K. W. Kelley, et al., Generation of Functional Human 3D Cortico-Motor Assembloids, Cell, 2020, 183(7), 1913–1929.e26 CrossRef PubMed.
- K. Imamura, Y. Izumi, A. Watanabe, K. Tsukita, K. Woltjen and T. Yamamoto, et al., The Src/c-Abl pathway is a potential therapeutic target in amyotrophic lateral sclerosis, Sci. Transl. Med., 2017, 9(391), eaaf3962 Search PubMed.
- B. J. Wainger, E. A. Macklin, S. Vucic, C. E. McIlduff, S. Paganoni and N. J. Maragakis, et al., Effect of Ezogabine on Cortical and Spinal Motor Neuron Excitability in Amyotrophic Lateral Sclerosis: A Randomized Clinical Trial, JAMA Neurol., 2021, 78(2), 186–196 Search PubMed.
- C. R. Bye, E. Qian, K. Lim, M. Daniszewski, F. C. Garton, B. C. Trần-Lê, H. H. Liang, T. Lin, J. G. Lock, D. E. Crombie, S. Morgan, Y. Hu, S. K. Barton, L. M. Palmer, E. Djouma, S. Kodikara, K.-A. Lê Cao, T. Dharmadasa, A. K. Henders, L. A. Ziser, M. C. Kiernan, K. Talbot, M. Needham, S. Fletcher, P. Talman, S. Mathers, N. R. Wray, A. W. Hewitt, A. Pebay and B. J. Turner, Large-scale drug screening in iPSC-derived motor neurons from sporadic ALS patients identifies a potential combinatorial therapy, Nat. Neurosci., 2025 DOI:10.1038/s41593-025-02118-7 , 41286450.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |