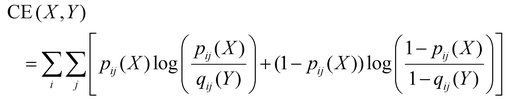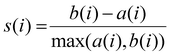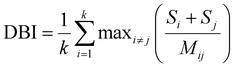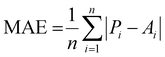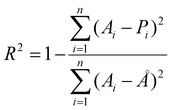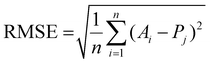Open Access Article
Open Access ArticleMachine learning framework to predict glass transition temperature in natural deep eutectic solvents: a step toward green functional materials
Durbek Usmanov ab,
Priyanka Yadavc,
Gerardo M. Casanola-Martin
ab,
Priyanka Yadavc,
Gerardo M. Casanola-Martin a,
Akshat S. Mallya
a,
Akshat S. Mallya c,
Seyedehelham Shirvanihosseinia,
Allison Hubel
c,
Seyedehelham Shirvanihosseinia,
Allison Hubel cd and
Bakhtiyor Rasulev
cd and
Bakhtiyor Rasulev *ab
*ab
aDepartment of Coatings and Polymeric Materials, North Dakota State University, Fargo, ND, USA. E-mail: bakhtiyor.rasulev@ndsu.edu
bMaterials and Nanotechnology (MNT) Program, North Dakota State University, Fargo, ND, USA
cDepartment of Mechanical Engineering, University of Minnesota, Minneapolis, MN, USA
dDepartment of Biomedical Engineering, University of Minnesota, Minneapolis, MN, USA
First published on 10th April 2026
Abstract
Natural Deep Eutectic Solvents (NADES) are a promising class of sustainable and environmentally-safe solvents with highly tunable physicochemical properties, including the glass transition temperature, which is critical for their functional performance, including ice control applications. Here, we present an interpretable machine learning (ML) framework to predict glass transition temperature from the molecular structure of NADES combination, integrating descriptor-based feature engineering, unsupervised clustering, and ensemble regression. Combination of components and their mixing ratios for forming NADES were utilized to generate specific multi-component descriptors to describe NADES for ML modeling. A set of multicomponent descriptors was calculated based on individual descriptors from chemically diverse components of NADES. As a result, a Random Forest (RF) model was developed to predict Tg values of NADES and the model achieved a very good performance with R2 values in a range of 0.87–0.93, for both training and test sets. The analysis of contributing factors by Shapley Additive exPlanations (SHAP) analysis identified key features highlighting contributions of 3D geometry, atomic mass distribution and electronic effects. Finally, our results demonstrate that ML approaches combined with mixture descriptors approach and interpretable modeling, enable accurate and chemically meaningful prediction of Tg, facilitating the rational design of NADES for applications in green chemistry and sustainable materials science.
Green foundation1. This work advances green chemistry by enabling data-driven design of natural deep eutectic solvents (NADES), biodegradable and low-toxicity alternatives to conventional organic solvents. We develop an interpretable machine-learning framework that predicts the glass transition temperature (Tg) directly from mixture-aware molecular descriptors, enabling virtual prescreening of formulations prior to synthesis.2. By reducing trial-and-error experimentation and unnecessary differential scanning calorimetry measurements, the approach lowers material consumption, solvent waste, and energy input associated with formulation development. This strategy shifts solvent discovery toward predictive, sustainability-driven design. 3. Importantly, the framework provides chemically interpretable guidance that links hydrogenbond network topology and molecular flexibility to thermal behavior, thereby supporting the rational selection of sustainable compositions. Future work will expand compositional diversity and integrate multi-objective optimization (e.g., viscosity, toxicity, water tolerance) to further strengthen environmentally responsible solvent design. |
1 Introduction
Climate change drives the global scientific community to search for novel materials that can replace conventional, often toxic substances with biodegradable alternatives without compromising essential physical properties. In this context, Natural Deep Eutectic Solvents (NADES) have emerged as a promising class of materials due to their biodegradability,2 low toxicity,3 and tunable physicochemical properties, such as high thermal stability,4 conductivity,5 adjustable viscosity,6 polarity,6 low volatility,7 water solubility,7 stability in different concentrations, and a high solubilization capacity for a wide range of pharmacologically active compounds.7 Despite their relatively short history of research, it is well-known that NADES demonstrate a unique ability to form stable hydrogen-bonded networks, making them promising sustainable alternatives for various applications, including as pesticides,8 absorption materials for gases,9 coatings,10 cosmetics,11 drug delivery systems,12 biosensors,13 solar cells,14 supercapacitors,15 battery electrolytes,16 anti-freezing or anti-icing agents,17,18 for cryopreservation19 and in green chemistry processes.20One of the crucial thermophysical properties that significantly influences the materials, particularly in coatings, encapsulation, and flexible electronics, is the glass transition temperature.21–23 Tg defines the temperature range at which a material transforms from a rigid, glassy state to a soft, rubbery, phase.24 This transition is essential for determining thermal stability, mechanical integrity, and usability of materials under varying environmental conditions. For NADES-based systems, controlling the Tg is vital in ensuring functionality in temperature-sensitive applications; however, systematic studies on their Tg behavior remain limited due to the complex, multicomponent nature of their hydrogen-bonded networks.21–23 In cryopreservation applications, higher cryoprotectant Tg values have been linked to lower critical cooling and warming rates for vitrification,25 reduced likelihood of damaging ice crystal formation,26 and the reduction of thermal stresses in vitrified samples, leading to lower chances of catastrophic mechanical failure.27 While Tg is commonly measured using experimental techniques such as differential scanning calorimetry (DSC), thermomechanical analysis (TMA), and dynamic mechanical analysis,28 computational modelling and machine learning are increasingly being adopted to predict Tg from molecular descriptors.29,30
These data-driven approaches offer valuable insights into the relationship between molecular structure and thermophysical behavior, accelerating the design of NADES and other next-generation materials for advanced technological applications. However, despite the growing interest in NADES, comprehensive studies on their glass transition behavior remain limited, primarily because of the complexity of their multicomponent hydrogen-bonded networks and the variability in their physicochemical properties.8,31 In recent years, accurately estimating the glass transition temperature of materials, particularly in confined geometries, has become increasingly important due to Tg's strong dependence on both composition and environmental factors.32 However, the experimental determination of Tg for newly developed material systems remains a significant challenge, as such measurements are often labor-intensive, time-consuming, and costly.23 Fortunately, the availability of experimental Tg datasets has opened the door to benchmarking computational approaches that offer a more efficient and scalable means of predicting Tg.
Beyond property prediction, machine learning is increasingly being used to optimize sustainable chemical processes, including biomass fractionation workflows.33,34 These advances underscore the broader potential of data-driven approaches in green chemistry, where predictive modeling can reduce experimental burden, accelerate optimization, and improve decision-making in complex multivariable systems. In this context, machine learning (ML)-based quantitative structure–property relationship (QSPR) models as efficient tools for solving complex challenges in chemistry, biology and materials science.35–44 These models directly link molecular structures to their physicochemical properties, offering a faster and more cost-effective alternative to traditional trial-and-error experimentation. Scientists have successfully applied QSPR models to a wide range of chemical systems, including the prediction of Tg, significantly advancing the rational design of new materials.30,45–51 Although many studies have predicted Tg values for various materials, researchers have yet to develop a dedicated dataset or model for Tg prediction in NADES. These advancements underscore the increasing effectiveness of ML-based approaches in capturing complex structure–property relationships within NADES systems. However, a review of the current literature reveals that most studies have predominantly focused on melting point,52,53 density,54,55 and viscosity of NADES,56,57 but only limited attention has been given to Tg. This gap highlights the urgent need for development of dedicated computational and predictive models for Tg, specifically designed to address the unique structural and physicochemical complexities of NADES.
Considering these limitations, in this study, we developed a ML-based QSPR model for predicting the glass transition temperature of NADES. We integrated an advanced combination of computational techniques to improve predictive accuracy and model interpretability. The Random Forest algorithm was selected as the primary modeling tool due to its robustness in capturing complex, nonlinear relationships across multivariate descriptor sets. The dataset, which comprises Tg values ranging from −122.87 to −52.54 °C, was analyzed using a combination of ML and dimensionality reduction methods. Specifically, Uniform Manifold Approximation and Projection (UMAP) was employed to explore the chemical space of NADES compounds, revealing meaningful structural clusters of NADES in the visualized chemical space. This unsupervised clustering guided the splitting of the dataset into training and test sets, thereby ensuring a structurally diverse and representative evaluation framework. To interpret the model predictions and assess the influence of individual molecular features on Tg, a SHAP approach was applied. This enabled a quantitative understanding of the most significant descriptors influencing the thermal behavior of NADES. This integrated approach not only enables accurate prediction of Tg but also provides valuable insights into the molecular determinants of glass transition behavior in eutectic solvent systems. Also, since today's materials science increasingly prioritizes environmentally friendly and sustainable alternatives, the developed model offers a powerful tool for the rational design of novel NADES formulations with tailored thermophysical properties for green and environmentally safe applications.
2 Methodology
2.1 Materials
L-Proline (Pro, ≥99%, Sigma Aldrich), betaine (Bet, ≥98%, Sigma Aldrich), guanidine hydrochloride (Gua, ≥99.5%, Thermo Scientific Chemicals), choline chloride (ChCl, 98%, Oakwood Chemical), sarcosine (Sar, 98%, Sigma Aldrich), L-lysine (Lys, ≥98%, Sigma Aldrich), trimethylamine N-oxide (TMAO, ≥98%, Thermo Scientific Chemicals), L-arginine (Arg, ≥98%, Sigma Aldrich), urea (U, ≥99.5%, Fluka Chemie GmbH), D-(+)-glucose (Glu, ≥99.5%, Sigma Aldrich), D-(−)-fructose (Fru, ≥99%, Sigma Aldrich), sucrose (Suc, puriss grade, Sigma Aldrich), D-(+)-trehalose dihydrate (Tre, ≥99%, Sigma Aldrich), D-sorbitol (Sor, ≥98%, MP Biomedicals), Xylitol (Xyl, ≥99%, Sigma Aldrich), ethylene glycol (EG, 99.8%, Sigma Aldrich), glycerol (Gly, 99.5%, Humco) and ultrapure-grade water was used from a Milli-Q filtration system to prepare the NADES samples.2.2 NADES preparation
NADES-forming components were added to capped glass vials (WHEATON) in the molar ratios specified in this work until a total mass of 15 g was achieved. The glass vials containing the mixtures were then placed in a water bath at 60 °C and stirred at 200 rpm for 24 h. After mixing, the samples were allowed to rest at room temperature for 72 h before performing further analyses. Polarized optical microscopy was performed for all NADES mixtures to confirm the absence of crystals.58 The NADES formulations included in the study were prepared using a mixture of sugars (glucose, sucrose, etc.), sugar alcohols (glycerol, sorbitol, etc.), amino acids (proline, lysine, etc.) and other metabolites like urea and trimethylamine N-oxide. Two, three and four-component mixtures were prepared in a wide range integer molar ratios. Water was added to the prepared NADES to create 20% w/w to 80% w/w solutions of diluted NADES. Details about the composition of the NADES formulations are included in Table S2.2.3 Thermophysical characterization
Differential scanning calorimetry was carried out using a TA Instruments Q1000 calorimeter. For each measurement, 15–20 mg of a NADES sample was placed into an aluminum pan (Tzero, TA Instruments) and sealed with a hermetic lid (Tzero, TA Instruments). The samples were first cooled from 20 °C to −150 °C at a constant rate of 10 °C min−1, followed by a 3 min isothermal equilibration at −150 °C to ensure thermal stability. Subsequently, the samples were heated from −150 °C back to 20 °C at 10 °C min−1. Thermophysical properties of the NADES samples were evaluated using TA Universal Analysis software (version 4.5A). The glass transition temperature1 was identified as the first local minimum observed in the plot of the derivative of heat flow with respect to temperature during the heating step.2.4 Computational data analysis and structural descriptors calculation
The structural characterization of NADES’ components began with the construction of individual molecular structures using ChemSketch software.59 The optimized components were subsequently processed by applying AlvaDesc code to generate a comprehensive array of molecular descriptors.60 For each NADES system, mixture-type descriptors were generated by combining the descriptors of individual components, multiplying on their respective molar ratios.61,62This approach produced an initial set of 5666 descriptors per each NADES system, spanning a broad spectrum of physicochemical, topological, geometrical, and electronic properties. These descriptors encompassed multiple-dimensional categories, including 0D, 1D, 2D, and 3D, and were drawn from various descriptor classes, such as constitutional indices, topological and connectivity metrics, 2D autocorrelations, geometrical parameters, atom-centered fragments, and quantum-chemical features.
To minimize redundancy and eliminate non-informative variables, descriptors exhibiting zero variance or near-constant values across the dataset were removed. Following this filtering process, a final set of 4316 descriptors was retained for further machine learning modeling. Importantly, this descriptor set was not used directly in its entirety for interpretation or final decision-making. Instead, feature relevance was further refined in a data-driven manner using Random Forest-based learning and SHAP analysis, which effectively reduced the dimensionality to a compact subset of influential descriptors governing Tg prediction. A detailed summary of the initial descriptor set is provided in the SI (Table S2).
These curated descriptors formed the input features for all subsequent predictive modeling, providing a high-dimensional and chemically meaningful representation of the NADES space for accurate prediction of the glass transition temperature. Besides, Fig. 1 illustrates the general workflow.
 | ||
| Fig. 1 Overview of the research workflow. Dataset preparation workflow (left), and machine learning workflow. | ||
2.5 Uniform manifold approximation and projection (UMAP)
UMAP is a nonlinear dimensionality reduction technique that constructs fuzzy topological representations of data by modeling the high-dimensional input space as a weighted graph. In this method, the probability of a connection between two data points in the high-dimensional manifold is expressed as:| pij = e−d(xi, xj)/σi |
| qij = (1 + a(yi − yj)2)−b |
Unlike other methods, UMAP minimizes a cross-entropy loss function defined as:
This formulation allows UMAP to preserve both local and global data structures, enabling it to capture meaningful relationships in complex high-dimensional datasets.
The two principal hyperparameters in UMAP are the number of neighbors and the minimum distances. The number of neighbors defines the size of the local neighborhood used for manifold approximation – lower values enhance the preservation of local structures, whereas higher values emphasize global relationships. The minimum distance controls how tightly data points are packed in the low-dimensional space: smaller values promote tight clustering, while larger values preserve more of the global data topology.
To assess the clustering structure in the UMAP – reduced space, k-means clustering was applied. The optimal number of clusters was determined using internal validation metrics, namely the Davies–Bouldin Index (DBI) and the Silhouette Score (SS).
The SS s(i) for each sample is defined as:
The DBI is defined as:
After selecting k = 6 as the optimal number of clusters, final cluster assignments were computed using k-means, and centroid coordinates were extracted from the UMAP embedding space. Prior to dimensionality reduction, all numerical descriptors were standardized using StandardScaler (zero mean, unit variance). UMAP was then applied using the umap-learn implementation with n_neighbors = 20, min_dist = 0.1, n_components = 2, metric = ‘euclidean’, and random_state = 42 to ensure reproducibility. UMAP projections and cluster validation were implemented using the UMAP-learn and Scikit-learn libraries in Python. These results confirm that the UMAP transformation not only preserves meaningful chemical relationships in the descriptor space but also facilitates robust unsupervised clustering of NADES formulations.
2.6 Machine learning model and evaluation metrics
In this study, in the best-performing Random Forest Regression63 model was employed to predict the Tg of the NADES systems. RF is a widely used ensemble learning method that builds a collection of decision trees and aggregates their outputs to produce robust predictions. The algorithm operates by constructing multiple regression trees from random subsets of the training data and features, and then averaging their outputs to mitigate variance and overfitting. This approach is convenient in handling high-dimensional descriptor spaces and complex nonlinear relationships.The RF model was implemented using the Scikit-learn library in Python. Hyperparameters – including the number of estimators (trees), maximum tree depth, and minimum samples per leaf – were optimized using grid search with 5-fold cross-validation to ensure generalization and minimize overfitting. The RF regression model was implemented in Python using the Scikit-learn library (RandomForestRegressor). Hyperparameter optimization was performed using grid search with 5-fold cross-validation (GridSearchCV, scoring = R2) and a fixed random seed (random_state = 42) to ensure reproducibility. The hyperparameter search space included the number of trees (n_estimators = 100–400), maximum tree depth (max_depth = 4–10), minimum number of samples required to split an internal node (min_samples_split = 2–10), and minimum number of samples per leaf (min_samples_leaf = 1–4). Cross-validation was conducted exclusively within the training set, following the UMAP-based structure-aware train–test splitting strategy. The held-out test set was not used during model selection and was reserved solely for final performance evaluation, thereby minimizing information leakage and overfitting.
To evaluate the performance of the model, the following metrics were calculated on the training, test, and external validation datasets:
Mean absolute error:64
Coefficient of determination (R2):
Root Mean Squared Error (RMSE):
These statistical indicators were selected to ensure a comprehensive evaluation of the model's accuracy, error distribution, and generalization capacity. A high R2, alongside low MAE and RMSE values, indicates the reliable predictive performance and robustness of the trained RF model.
2.7 Feature selection and model interpretability
To enhance model transparency and ensure the reliability of Tg predictions, we employed a dual strategy that integrates both data-driven feature selection and post hoc model interpretability analysis. To interpret feature contributions and quantify their influence on the model output, we applied SHAP, a model-agnostic interpretability method based on cooperative game theory. SHAP values were computed using the shap.TreeExplainer module (SHAP v0.44.1), which is optimized for ensemble tree models such as Random Forest.Prior to SHAP analysis, standard descriptor preprocessing steps were applied, including the removal of constant descriptors and highly correlated variables, to ensure numerical stability and reduce redundancy in the descriptor space. After initial descriptor preprocessing, SHAP-based feature importance analysis was used to identify the most informative variables. The final predictive models were constructed using the top 35 descriptors ranked by SHAP importance, thereby substantially reducing the dimensionality of the descriptor space relative to the number of samples. This dimensionality reduction step mitigates the risk of overfitting and improves model interpretability.
It should be noted that SHAP provides an additive decomposition of model predictions and does not explicitly reconstruct the full nonlinear functional form learned by the underlying machine learning model. Consequently, SHAP-based interpretations should not be viewed as mechanistic or causal descriptions of molecular interactions.
Nevertheless, SHAP remains a powerful interpretability tool for identifying dominant structure–property trends, directional effects, and threshold-like behaviours embedded within nonlinear models. In this study, SHAP analysis is used to reveal consistent, chemically plausible relationships between molecular descriptors and Tg, rather than to infer explicit nonlinear interaction mechanisms. The observed SHAP dependence patterns (e.g., saturation and sigmoidal responses) reflect the nonlinear decision boundaries learned by the model, providing physically meaningful insight into the factors governing vitrification in NADES.
SHAP analysis allowed for a detailed decomposition of Tg predictions into additive feature attributions, revealing both global feature importance and local instance-specific effects. Visualization tools, including beeswarm and summary plots, were used to identify the most influential descriptors. Among the top contributors were 3D-MoRSE and GETAWAY descriptors (e.g., Mor08p, Mor26p, R6m+) and topological autocorrelations (e.g., MATS4i), highlighting the importance of molecular geometry, polarizability, and electronic distribution in modulating the thermal behavior of NADES systems.
This framework not only improved the interpretability of our model but also provided mechanistic insights into structure–property relationships within the NADES chemical space, thereby supporting rational design and formulation strategies.
2.8 Visualization
All graphical analyses and visualizations were performed using Python (version 3.10.11) with the Matplotlib (version 3.7.1) and Seaborn (version 0.12.2) libraries. These tools were used to generate model performance plots (e.g., R2 comparisons, SHAP summary plots), feature importance rankings, UMAP projections, clustering diagrams, and correlation heatmaps. Custom plotting scripts ensure consistent visual styling, interpretability, and high-resolution rendering (≥300 dpi), making the figures suitable for publication in both print and digital formats. All visualization codes are fully reproducible and provided in the SI.3 Results
This work contributes to green chemistry by enabling data-driven pre-screening of NADES formulations, reducing experimental waste, solvent consumption, and energy-intensive trial-and-error synthesis. In this study, we used an in-house dataset comprising 263 NADES systems with experimentally measured Tg values to develop several machine learning methods for Tg prediction. This is a unique experimental dataset, collected by the group, since all the other public data available are scarce and made with different protocols. Therefore, the collection of 263 NADES with measured Tg under the same experimental protocol is a unique set for a proper ML study. To investigate Tg of NADES in detail, we systematically evaluated different algorithms using a curated dataset of NADES formed from various components’ combinations. The results presented in this work include model development steps, performance metrics, feature importance analysis based on SHAP analysis, and insights into molecular descriptors that most significantly influence Tg values.After careful data curation, the dataset of NADES was used to develop predictive models, which were trained and evaluated using a “mixture-based split” strategy, ensuring that structurally related compounds were not included in both the training and test sets simultaneously. This approach provides a more realistic assessment of the ML model's generalizability, especially for prospective applications involving prescreening of novel NADES formulations. The overall workflow of the study is shown in Fig. 1.
3.1 Experimental analysis
The glass transition temperature was determined from the DSC thermogram by analyzing the first derivative of heat flow with respect to temperature, where the peak of the first local maximum during the heating step was taken as Tg. Upon cooling, pure NADES (without dilution) samples exhibited vitrification without undergoing crystallization, thereby forming a metastable supercooled liquid state, eventually undergoing a glass transition into a non-equilibrium glassy phase. This was observed in their thermograms by noting the presence of only a glass transition without a melting peak, consistent with eutectic behavior.5,65,66 The Tg values of pure NADES were found to lie between −114.54 °C (Fruc![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) EG 1
EG 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 66) to −53.64 °C (Gly
66) to −53.64 °C (Gly![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Glu
Glu![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Sorb
Sorb![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Water 1
Water 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3), lower than those observed for the water-diluted samples. These findings agree with previous reports showing that NADES remain in the liquid state across a broad low-temperature range.66,67
3), lower than those observed for the water-diluted samples. These findings agree with previous reports showing that NADES remain in the liquid state across a broad low-temperature range.66,67
Water addition was found to disrupt the supramolecular organization of NADES, increase molecular mobility, and lower Tg.66,68 Tg decreased with increasing dilution up to 50% but remained nearly constant at higher dilutions.18 The reduction in Tg upon hydration reflected the well-known plasticizing role of water.5 Consistent with the Flory–Fox equation, Tg was observed to be directly proportional to molecular weight. NADES formulated with low-molecular-weight ethylene glycol reported the lowest Tg, whereas those with high-molecular-weight sorbitol exhibited higher Tg values.69
3.2 Dataset preparation for ML-QSPR modeling
A crucial first step before performing any ML modeling is a proper splitting of the dataset to avoid data leakage and achieve a balanced structural diversity distribution of the NADES dataset through unbiased train-test data splitting. For this, we applied the UMAP unsupervised classification approach70 to perform dimensionality reduction on the dataset and facilitate visualization. UMAP effectively projects the high-dimensional molecular features (descriptors) space into two dimensions while preserving both local and global topological structures.71,72 To quantitatively determine the optimal number of clusters, we evaluated the Silhouette Score (SS)73 and Davies–Bouldin index (DBI)74 across multiple cluster configurations (from 2 to 17 clusters). As shown in Fig. 2a, the SS reached its maximum (SS = 0.476) and the DBI its minimum (DBI = 0.695) at k = 6, indicating the optimal balance between cluster cohesion and separation. As shown in Fig. 2b, the UMAP revealed the presence of six well-defined clusters, each representing distinct regions of chemical space, thus confirming the molecular diversity within the dataset. These six clusters served as the foundation for splitting the dataset into training and test sets, ensuring that the model would be trained and evaluated on structurally diverse and representative subsets. Furthermore, the UMAP-based clustering approach provides insights into the underlying structural patterns of the NADES compounds, as seen in the distribution of the clusters. Each cluster exhibits distinct chemical characteristics, which may correlate with differences in its glass transition behavior. By using this structure-aware data partitioning strategy, we minimized potential data leakage and enhanced the predictive model's generalization ability.It is worth noting that water content can significantly influence the Tg value. To explore this, the importance of water concentration on the Tg within structurally similar NADES systems was investigated as well, and a comparative violin plot was generated (Fig. 3), highlighting various combinations of the same core components with differing water content. As can be noticed in the plot, the systems Bet![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Ure
Ure![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Wat, Bet
Wat, Bet![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Sor
Sor![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Wat, Bet
Wat, Bet![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Sor
Sor![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Ure
Ure![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Wat, and Bet
Wat, and Bet![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Lys
Lys![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) Wat exhibit minimal shifts and smooth transitions in Tg values as water concentration increased. Water concentration was explicitly reported for each formulation (Table S2) and treated as part of the mixture composition during descriptor generation for the machine learning model. The consistently narrow violin shapes across these systems indicate that the incorporation of water had only a moderate impact on their thermal behavior. This stability likely reflects robust intermolecular interactions among the primary components (e.g., betaine, sorbitol, urea, lysine), which are only marginally affected by hydration. Conversely, other NADES systems demonstrated broader or more irregular Tg distributions, with pronounced changes in response to variations in water. This suggests that their physicochemical properties are more sensitive to hydration, possibly due to weaker or more dynamic hydrogen bonding networks. These findings emphasize that specific NADES formulations preserve structural and thermal coherence even under changing moisture conditions, making them attractive candidates for applications requiring thermophysical stability. This analysis underscores the chemical and functional diversity within the NADES formulations, highlighting the need to employ advanced machine learning techniques to capture the complex, nonlinear relationships between molecular structure and physicochemical properties. The applied methodology enhances the interpretability and robustness of the predictive framework, enabling more accurate and generalizable predictions of the glass transition temperatures across a structurally diverse range of NADES systems.
Wat exhibit minimal shifts and smooth transitions in Tg values as water concentration increased. Water concentration was explicitly reported for each formulation (Table S2) and treated as part of the mixture composition during descriptor generation for the machine learning model. The consistently narrow violin shapes across these systems indicate that the incorporation of water had only a moderate impact on their thermal behavior. This stability likely reflects robust intermolecular interactions among the primary components (e.g., betaine, sorbitol, urea, lysine), which are only marginally affected by hydration. Conversely, other NADES systems demonstrated broader or more irregular Tg distributions, with pronounced changes in response to variations in water. This suggests that their physicochemical properties are more sensitive to hydration, possibly due to weaker or more dynamic hydrogen bonding networks. These findings emphasize that specific NADES formulations preserve structural and thermal coherence even under changing moisture conditions, making them attractive candidates for applications requiring thermophysical stability. This analysis underscores the chemical and functional diversity within the NADES formulations, highlighting the need to employ advanced machine learning techniques to capture the complex, nonlinear relationships between molecular structure and physicochemical properties. The applied methodology enhances the interpretability and robustness of the predictive framework, enabling more accurate and generalizable predictions of the glass transition temperatures across a structurally diverse range of NADES systems.
3.3 Development of ML-QSPR model
To assess confidently the Tg of NADES, the capabilities of several machine learning algorithms were analyzed, including RF,75 CatBoost,76 XGBoost,77 Gradient Boosting,78 LightGBM,79 Support Vector Regression (SVR),80 k-Nearest Neighbors (kNN),81 Gaussian Process,82 Artificial Neural Networks (ANN),83 and Lasso Regression.84 Each individual ML model was optimized through hyperparameter tuning, and the models’ performances were evaluated using the coefficient of determination (R2) for the training set, mean squared error,85 and R2 values for the test set (SI, Table S1). To establish a performance benchmark, we initially evaluated a set of baseline ML models. This included both linear and non-linear regression algorithms. Among these, the RF model demonstrated the most promising performance, achieving R2 of 0.87 for the test set and a low mean squared error (MSE = 0.0001), while maintaining a strong fit on the training data (R2 = 0.96), indicating good generalization (Table S1). In contrast, some models displayed tendencies toward overfitting; for instance, XGBoost and CatBoost achieved nearly perfect R2 values on the training set (0.99), their predictive performance declined on the test set to 0.75 and 0.89, respectively (Table S1), indicating limited generalization ability. Similarly, Gradient Boosting and LightGBM models exhibited high training R2 values but decreased test performance, suggesting that these models capture complex patterns in the training data, but they lack robustness when exposed to unseen data without further regularization. These results show that RF algorithm is highly effective in modeling the complex, nonlinear relationships between molecular descriptors and the Tg of NADES formulations. Its superior generalization performance underscores the robustness of ensemble tree-based approaches for capturing structure–property relationships in chemically diverse systems. Furthermore, the observed tendency of specific advanced boosting methods to overfit emphasizes the necessity of careful model selection and validation to ensure predictive reliability.To further enhance the predictive performance of the RF model, we conducted an extensive hyperparameter optimization using a grid search approach. Key parameters, including the number of trees [n_estimators], maximum tree depth [max_depth], minimum number of samples required to split an internal node [min_samples_split], and minimum number of samples needed for a leaf node [min_samples_leaf], were systematically varied across a defined range. GridSearchCV with 5-fold cross-validation was employed to evaluate each parameter combination, with R2 as the primary scoring metric. The optimal configuration identified through 5-fold cross-validation consisted of 180 trees [n_estimators = 180], a maximum depth of 6, min_samples_split = 2, and min_samples_leaf = 1. The hyperparameter-tuned RF model preserved high predictive accuracy while improving computational efficiency and model stability, ensuring that predictions of Tg remain both interpretable and generalizable across a chemically diverse NADES space. Although RF provides built-in feature importance measures such as mean decrease in Gini impurity86,87 or permutation importance,88 these approaches are limited by their global scope, bias toward high-cardinality or continuous variables, and instability in correlated descriptor spaces, which are common in cheminformatics. To overcome these shortcomings, we employed SHAP approach,89 a framework that ensures mathematical consistency, delivers both global rankings and local, sample-specific contributions, and distributes importance more equitably across correlated descriptors. By directly linking descriptor effects to predicted Tg values at multiple levels, SHAP offers a more reliable and chemically meaningful interpretation of structure–property relationships. SHAP was therefore used to complement RF feature importance, providing unbiased insights that enhance mechanistic interpretability and strengthen confidence in the model's predictive performance across the chemically diverse NADES space.
3.4 Model Interpretability via SHAP Analysis
An additional layer of analysis was conducted using SHAP approach to deepen our understanding of the internal decision-making process of the optimized model. This approach was implemented not to assess robustness, but rather to provide a better understanding of the relative importance of the molecular descriptors selected in the ML model and to explain how they influence the model's predictions. By decomposing each prediction into additive contributions from individual features, SHAP values provide a transparent and quantitative interpretation of how specific chemical descriptors influence the predicted glass transition temperature. Representative outcomes from this interpretability analysis are illustrated in Fig. 4.Before engaging in a more in-depth interpretation of SHAP-based feature attributions, it is essential to recognize that several factors inherently bound the interpretability of any ML model. These include the model's architectural constraints (e.g., ensemble tree-based structure in RF), the nature of the input representation (in this case, applied molecular mixture-descriptors), and the specific chemical space spanned by the training dataset. Therefore, the SHAP values discussed herein should be understood as explanations tied to the Tg predictions within the scope of the NADES systems represented in this study. Importantly, the limited contribution or apparent absence of specific physicochemical descriptors traditionally considered relevant for glass transition phenomena does not imply that these features lack significance across broader or alternative molecular contexts. Instead, their marginal influence in our model may reflect dataset-specific patterns or descriptor redundancy within the confined chemical space.
With this framework in mind, a SHAP beeswarm plot (Fig. 4a) was generated to visualize the relative impact of the top 20 selected molecular descriptors on the predicted glass transition temperature of NADES. Among these, molecular topological descriptors such as Mor08p, R6m+, MATS4i, and Mor26p emerged as the most influential features, each displaying a distinctive effect on model output. In this context, descriptors with higher positive SHAP values are associated with an increased predicted Tg value, whereas those with negative values tend to lower the predicted Tg value. The feature importance ranking, Fig. 4c, corroborates these findings, indicating that a small subset of descriptors accounts for the majority of the model's predictive ability. Further, the individualized SHAP analyses (Fig. 4b and d) provide a detailed understanding of how specific molecular descriptors contribute to Tg predictions in individual NADES systems. For instance, descriptor Mor08p is a 3D-MoRSE descriptor weighted by polarizability, exhibits a strong positive contribution to Tg, indicating that spatially extended and polarizable molecular structures tend to elevate glass transition temperatures. At the same time, R6m+ is a GETAWAY descriptor capturing mass distribution at topological lag 6, contributes negatively, suggesting that mass clustering at intermediate distances may reduce Tg. These trends are substantiated by SHAP dependence and force plots, which reveal both the directionality and magnitude of feature effects. A representative SHAP force plot, Fig. 4d, demonstrates the local interpretability of a single prediction: Mor08p contributed a significant positive shift (+0.572) toward the predicted Tg value, whereas R6m+ had a minor negative effect (−0.025). Together, these feature contributions reduced the prediction from the model's base value (2.26) to the final output (2.23). This visualization reinforces the utility of SHAP in uncovering feature-specific impacts within the Random Forest framework, offering mechanistic insights into the structural underpinnings of Tg behavior in NADES. Additionally, the presented figure illustrates a SHAP decision plot, which visualizes the contribution of individual molecular descriptors to the predicted values of the glass transition temperature expressed as Tg. The x-axis represents the model's output (predicted values), while the y-axis lists the most influential descriptors, ranked by their average SHAP importance. Each line on the plot corresponds to a single observation in the dataset, tracing how cumulative SHAP values (feature attributions) evolve from the model's base value to its final output for that instance. The color gradient encodes the normalized value of the respective descriptor: blue indicates lower values, whereas blue to pink denotes higher values.
Although the RF algorithm is inherently robust to multicollinearity and does not require strict linear independence among input features, where evaluated pairwise correlations remain essential for interpreting model's behavior. To this end, a Pearson correlation matrix was constructed for the top 15 descriptors ranked by SHAP importance, Fig. 4e. This visualization enables an assessment of feature redundancy and helps to identify potential duplication of encoded physicochemical information. For instance, descriptor pairs such as MATS4i-HATS2e and WHALES60_Rem-WHALES80_Rem exhibited strong cross-correlation (|r| > 0.85), indicating that they encode similar structural characteristics. In contrast, features like Mor08p and R6m+ displayed weak correlation (r ≈ –0.46), underscoring low similarity and high structural diversity among the most informative variables. Overall, the analysis confirms the presence of both partially redundant and chemically unique descriptors, ensuring broad and diverse coverage of the NADES chemical space. This reinforces the interpretability of the model and highlights the value of integrating SHAP-based feature importance with correlation-based filtering in molecular modeling workflows.
These results highlight the model's strong generalization ability and stable predictive accuracy across diverse datasets (Fig. 5). Moreover, the combination of rigorous regularization through optimized tree depth and node splitting, along with descriptor-level interpretability via SHAP, not only mitigates overfitting but also provides mechanistic insights, ensuring that complex nonlinear relationships within the descriptor space are captured in a transparent and chemically meaningful manner. To further validate the model's robustness and feature interpretability, we examined the distribution of the most influential descriptors across the training and test datasets. As shown in Fig. 4a and b, descriptors such as Mor08p, R6m+, MATS4i, and Mor26p exhibit consistent distributions between training and test sets, indicating good representativeness and minimal sampling bias. Specifically, Mor08p follows a near-normal distribution centered around 0.4 in both datasets, while R6m+ displays a positively skewed distribution, reflecting its sparsity in the chemical space. Similar trends are observed for MATS4i and Mor26p, where the descriptors’ values are well-aligned across the splits.
Although descriptors such as Mor08p and R6m+ do not explicitly encode hydrogen bonding, they capture structural features that indirectly influence it. Mor08p reflects the three-dimensional distribution of polarizability and is consistent with differences in molecular packing and supramolecular rigidity, whereas R6m+ describes medium-range mass distribution and can be associated with structural compactness and segmental mobility. Together, these effects are relevant to the stability of hydrogen-bond networks and the vitrification behavior of NADES.
Thus, the discussed visualizations together link structural diversity to thermal behavior, thereby enhancing the mechanistic interpretability of the model outputs. These distributional consistencies bolster the generalization ability of the optimized RF model and ensure that SHAP-derived feature contributions are not merely the result of sampling bias. Moreover, the consistency between feature importance and descriptor availability across datasets enhances the interpretability and chemical relevance of the model, supporting its use in virtual screening and rational NADES design.
3.5 Detailed descriptor interpretation
The glass transition behavior of NADES exhibits clear compositional dependence across chemically distinct formulation regimes defined by the nature of the constituent components and their relative ratios. Fig. 6 illustrates how the glass transition temperature of NADES varies across chemically distinct compositional regimes, as identified through UMAP-based clustering of the descriptor space and visualized using representative component systems. Rather than reflecting isolated descriptor effects, the observed trends primarily arise from differences in NADES composition, hydrogen-bond donor/acceptor balance, and water content, which collectively govern network rigidity and molecular mobility.Clusters associated with higher Tg values are predominantly composed of sugar-rich NADES systems, such as sucrose–glucose–fructose–water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 11) and glucose–fructose–water (1
11) and glucose–fructose–water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8). These systems feature dense, multidirectional hydrogen-bonding networks formed by polyhydroxylated carbohydrates, leading to enhanced structural rigidity and reduced segmental mobility. The presence of multiple hydroxyl groups per molecule promotes extensive intermolecular interactions, which stabilizes the amorphous phase and shifts Tg toward higher values. In these compositions, water acts as a plasticizer at low concentrations but becomes structurally integrated into the hydrogen-bonded network at higher ratios, contributing to network stabilization rather than disruption.
8). These systems feature dense, multidirectional hydrogen-bonding networks formed by polyhydroxylated carbohydrates, leading to enhanced structural rigidity and reduced segmental mobility. The presence of multiple hydroxyl groups per molecule promotes extensive intermolecular interactions, which stabilizes the amorphous phase and shifts Tg toward higher values. In these compositions, water acts as a plasticizer at low concentrations but becomes structurally integrated into the hydrogen-bonded network at higher ratios, contributing to network stabilization rather than disruption.
In contrast, clusters characterized by lower Tg values are enriched in amino acid- and choline-based NADES, such as proline–ethylene glycol (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3.3) and choline chloride–glycerol (1
3.3) and choline chloride–glycerol (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2). These systems exhibit more flexible hydrogen-bonding motifs, dominated by fewer donor–acceptor sites and increased conformational freedom of aliphatic chains. Ethylene glycol and glycerol introduce mobility through rotational flexibility, while zwitterionic or ionic components such as proline and choline chloride generate localized interactions that are less effective in producing rigid, three-dimensional networks. As a result, these formulations display lower resistance to molecular rearrangement and reduced glass transition temperatures.
2). These systems exhibit more flexible hydrogen-bonding motifs, dominated by fewer donor–acceptor sites and increased conformational freedom of aliphatic chains. Ethylene glycol and glycerol introduce mobility through rotational flexibility, while zwitterionic or ionic components such as proline and choline chloride generate localized interactions that are less effective in producing rigid, three-dimensional networks. As a result, these formulations display lower resistance to molecular rearrangement and reduced glass transition temperatures.
Intermediate clusters, including mixed systems such as glycerol–glucose–sorbitol–water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3), exhibit Tg values between the two extremes. These compositions combine rigid carbohydrate backbones with flexible polyol or amino-alcohol components, yielding partially constrained hydrogen-bonded networks. The coexistence of rigid and flexible molecular motifs produces heterogeneous interaction landscapes, reflected in broader Tg distributions and cluster overlap. This behavior highlights the non-additive nature of Tg in multicomponent eutectic systems, where collective interactions dominate over individual component properties.
3), exhibit Tg values between the two extremes. These compositions combine rigid carbohydrate backbones with flexible polyol or amino-alcohol components, yielding partially constrained hydrogen-bonded networks. The coexistence of rigid and flexible molecular motifs produces heterogeneous interaction landscapes, reflected in broader Tg distributions and cluster overlap. This behavior highlights the non-additive nature of Tg in multicomponent eutectic systems, where collective interactions dominate over individual component properties.
Importantly, the cluster-resolved analysis demonstrates that Tg in NADES is governed primarily by component chemistry and formulation strategy, rather than by isolated molecular descriptors. The machine-learning descriptors serve as an effective abstraction of these chemical features—capturing size, polarity, and interaction density—but the underlying physical origin of Tg variation remains rooted in hydrogen-bond topology, molecular flexibility, and water-mediated plasticization effects.
From a green chemistry perspective, these findings provide chemically intuitive design guidelines for tailoring Tg in NADES. Carbohydrate-rich, highly hydrogen-bonded systems are suitable for applications requiring elevated Tg and enhanced thermal stability, such as cryopreservation or solid-state encapsulation. Conversely, polyol- and amino acid-based NADES offer lower Tg and greater flexibility, making them attractive for low-temperature processing, coatings, and solvent applications. By linking machine-learning insights to concrete compositional motifs, this study advances the rational, sustainability-driven design of NADES with application-specific thermophysical properties.
3.6 Implications for process and formulation design
Accurate prediction of Tg enables the rational selection of NADES formulations for temperature-sensitive processes, such as cryopreservation, anti-icing, and sub-zero storage. By identifying compositions with Tg values safely below operating temperatures, the proposed ML framework allows prescreening of mechanically stable, amorphous systems without extensive experimental trial-and-error.In practical formulation design, Tg represents only one of several parameters governing the performance of NADES systems. Other properties, such as viscosity, toxicity, conductivity, cost, and solubilization capacity, may also strongly influence the suitability of a formulation for specific applications. Although the present study focuses on Tg prediction, the proposed descriptor-based machine learning framework is readily extendable to multi-property modelling. By combining Tg prediction with existing ML models for the viscosity, density, or surface tension of deep eutectic solvents, multi-objective screening strategies could be implemented to support rational NADES design. Such approaches would enable the identification of formulations that simultaneously satisfy multiple performance constraints, thereby accelerating the development of sustainable functional materials.
On the other hand, Tg must often be balanced against viscosity and water tolerance. Descriptor-level interpretation further indicates that molecular geometry, polarizability, and mass distribution simultaneously influence Tg and molecular mobility, providing guidance for navigating trade-offs between thermal stability, flow behavior, and hydration sensitivity. These insights support the development of multi-objective formulation strategies rather than single-property optimization.
From a process perspective, virtual Tg screening significantly reduces reliance on low-throughput DSC measurements by narrowing experimental validation to high-confidence candidates. As a result, the framework accelerates NADES formulation workflows while reducing material consumption and experimental cost.
If the required components are experimentally accessible, these formulations may be directly prioritized for preparation and validation. Otherwise, structurally related components, such as alternative sugars, polyols, amino acids, or quaternary ammonium derivatives, can be used to construct a virtual library of candidate systems across feasible molar ratios and water contents. For each candidate formulation, mixture descriptors are calculated using the same stoichiometry-weighted protocol applied in this study, and the trained Random Forest model is then used to predict Tg.
To ensure reliable interpretation, predicted candidates should also be evaluated within the applicability domain of the model, meaning that descriptor values remain within or close to the range represented in the training dataset. In this way, the model functions as a prescreening tool that reduces the number of experimental trials and accelerates the rational identification of NADES formulations with targeted low-temperature properties.
3.7 Limitations and transferability
Despite the strong predictive performance of the proposed machine-learning framework, several limitations should be acknowledged. First, the model is trained on a finite dataset of experimentally reported NADES systems and therefore reflects the chemical diversity, compositional ranges, and water contents represented in the available data. As a result, predictions for formulations that lie far outside the explored chemical space, such as systems involving unconventional hydrogen-bond donors or extreme stoichiometries, should be interpreted with caution. As noted in the SI data (Table S2), the Tg data used to train the ML models were based on single data points for each composition at each dilution. However, we have found that the Tg for NADES measured using the method described in this study have a low degree of variability.18,68 This experience gives us confidence in the reproducibility of the Tg measurements.Second, the descriptor-based representation provides an effective abstraction of component chemistry but does not explicitly encode specific intermolecular interactions or dynamic hydrogen-bond rearrangements. Consequently, the model captures statistically learned structure–property relationships rather than explicit mechanistic pathways governing glass transition behavior. While SHAP analysis improves interpretability, it does not establish causal links between individual descriptors and Tg.
Regarding transferability, the applicability of the model is expected to be highest for NADES composed of chemically related components and comparable formulation strategies, particularly polyol-, carbohydrate-, and amino acid-based systems. The applicability domain analysis further supports this view by identifying regions of descriptor space in which predictions are reliable. Extension of the model to fundamentally different eutectic systems or to non-natural DES formulations would require retraining or recalibration using representative experimental data.
Nevertheless, the overall modeling strategy, combining structure-aware data splitting, ensemble learning, and interpretability analysis, is transferable and can be readily adapted to other thermophysical properties or solvent classes. With the continued expansion of high-quality experimental datasets, the predictive scope and generalizability of such data-driven frameworks are expected to improve, supporting the rational and sustainable design of next-generation eutectic materials.
Although the present model was evaluated using a structure-aware held-out test set, prospective validation on newly prepared NADES formulations remains an important next step. Such validation would further establish the practical utility of the framework for formulation design and targeted screening.
4 Conclusions
In this work, we combined an in-house dataset with a data-driven framework to develop an ML model for predicting the Tg of NADES. A unique, internally consistent dataset of 263 NADES systems with Tg values ranging from −122.9 to −52.5 °C, measured under identical DSC protocols, was used to ensure reliable model training and validation. A stoichiometry-weighted combination of single-component descriptors generated mixture-specific molecular descriptors. UMAP-based clustering (k = 6) was used to guide structure-aware train-test splitting, preventing data leakage and ensuring evaluation on chemically distinct NADES families.Among the machine-learning algorithms tested, the RF model achieved the best balance between accuracy and generalization, with R2 = 0.93 (training) and R2 = 0.87 (test), whereas advanced boosting models showed higher overfitting. SHAP analysis identified a small subset of descriptors dominating Tg prediction, with the most influential being related to 3D molecular geometry, polarizability, atomic mass distribution, and electronic effects, directly linking Tg to molecular mobility and hydrogen-bond network rigidity.
Overall, this study demonstrates that Tg of NADES can be predicted with high accuracy using mixture-aware descriptors and interpretable machine learning. The proposed framework enables rapid prescreening and rational formulation of NADES with targeted thermal properties, reducing experimental effort and supporting scalable design of green solvents for cryopreservation, anti-icing, and materials applications.
Author contributions
D. U.: writing – original draft; D. U., G. C.-M., B. R.: software; D. U., G. C.-M., A. M., P. Y., S. S., A. H., B. R.: methodology; D. U., G. C.-M., A. H. B. R.: conceptualization; D. U., G. C.-M., A. M., P. Y., S. S., A. H., B. R.: writing – review and editing; A. H., B. R.: supervision; A. H., B. R.: resources; A. H., B. R.: funding acquisition.Conflicts of interest
Author A.H., P.Y., and A.S.M. are named inventors on U.S. Provisional Application No. 63/788,934 filed by the University of Minnisota relating to the technology described in this manuscript.Data availability
The authors declare that all data supporting the findings and those used for reproducing the figures in this paper are available within the paper and its supplementary information (SI). Supplementary information: experimental dataset and data used to develop the machine learning models (SI file S1 and file S2) and curated dataset. See DOI: https://doi.org/10.1039/d6gc01009a.Acknowledgements
A. H. and B. R. thank the U.S. DARPA for the support through the Award Number: HR001124C0315. A. H. receives partial support for this work from the NSF through the MRSEC (Award No. DMR-2011401) and the NNCI (Award No. ECCS-2025124) programs. Authors thank the Characterization Facility, University of Minnesota, where most of the experiments were carried out. Also, authors acknowledge the Center for Computationally Assisted Science and Technology (CCAST) at North Dakota State University, resources of which were used. This was made possible in part by National Science Foundation,90 MRI Award number 2019077 (Lead PI – B. Rasulev). Supercomputing support provided by the CCAST HPC System at NDSU is gratefully acknowledged.References
- A. Yetgin, Revolutionizing multi–omics analysis with artificial intelligence and data processing, Quant. Biol., 2025, 13(3), e70002 CrossRef PubMed.
- Y. T. Dai, J. van Spronsen, G. J. Witkamp, R. Verpoorte and Y. H. Choi, Natural deep eutectic solvents as new potential media for green technology, Anal. Chim. Acta, 2013, 766, 61–68 CrossRef CAS PubMed.
- A. Sharma and B.-S. Lee, Toxicity test profile for deep eutectic solvents: A detailed review and future prospects, Chemosphere, 2024, 350, 141097 CrossRef CAS PubMed.
- R. Alcalde, A. Gutiérrez, M. Atilhan and S. Aparicio, An experimental and theoretical investigation of the physicochemical properties on choline chloride - Lactic acid based natural deep eutectic solvent (NADES), J. Mol. Liq., 2019, 290, 110916 CrossRef CAS.
- R. Craveiro, I. Aroso, V. Flammia, T. Carvalho, M. T. Viciosa and M. Dionísio, et al., Properties and thermal behavior of natural deep eutectic solvents, J. Mol. Liq., 2016, 215, 534–540 CrossRef CAS.
- C. Spaggiari, L. Carbonell-Rozas, H. Zuilhof, G. Costantino and L. Righetti, Structural elucidation and long-term stability of synthesized NADES: A detailed physicochemical analysis, J. Mol. Liq., 2025, 424, 127105 CrossRef CAS.
- N. Aguilar, C. Benito, S. Martel-Martín, A. Gutiérrez, S. Rozas and P. A. Marcos, et al., Insights into Carvone: Fatty Acid Hydrophobic NADES for Alkane Solubilization, Energ Fuel, 2024, 38(24), 23633–23653 CrossRef CAS PubMed.
- A. Arunachalam, T. Oosterhoff, I. Breet, P. Dijkstra, R. A. M. Yunus and D. Parisi, et al., Harnessing the bio-adhesive power of natural deep eutectic solvents for trichome-inspired pest control, Commun. Mater., 2025, 6(1), 101 CrossRef.
- B. R. Wang, L. Lin, W. X. Zhang, C. F. Zhu, S. H. Ren and Y. C. Hou, et al., Specific heat capacities of deep eutectic solvents applicable for absorbing sulfur dioxide, Sep. Purif. Technol., 2025, 370, 133106 CrossRef CAS.
- J. Boiteux, M. Espino, S. Azcarate, M. F. Silva, F. J. V. Gomez and P. Pizzuolo, et al., NADES blend for bioactive coating design as a sustainable strategy for postharvest control, Food Chem., 2023, 406, 135054 CrossRef CAS PubMed.
- D. S. Freitas, D. Rocha, J. Santos, J. Noro, T. D. Tavares and M. O. Teixeira, et al., NADES-in-Oil Emulsions Enriched with Essential Oils for Cosmetic Application, Processes, 2025, 13(2), 374 CrossRef CAS.
- H. Li, K. N. Yang, Y. M. Yang, L. Q. Ding and X. X. Li, Natural deep eutectic solvents (NADES) in drug delivery systems: Characteristics, applications, and future perspectives, Int. J. Pharm., 2025, 675, 125509 CrossRef CAS PubMed.
- N. A. Aschemacher, C. M. Teglia, Á.S Siano, F. A. Gutierrez and H. C. Goicoechea, Development of an electrochemical sensor using carbon nanotubes and hydrophobic natural deep eutectic solvents for the detection of α-glucosidase activity in extracts of autochthonous medicinal plants, Talanta, 2024, 268(1), 125313 CrossRef CAS PubMed.
- C. L. Boldrini, N. Manfredi, F. M. Perna, V. Capriati and A. Abbotto, Eco-Friendly Sugar-Based Natural Deep Eutectic Solvents as Effective Electrolyte Solutions for Dye-Sensitized Solar Cells, ChemElectroChem, 2020, 7(7), 1707–1712 CrossRef CAS.
- D. Julião, M. Xavier and X. Mascarenhas, Deep eutectic solvents: viable sustainable electrolytes for supercapacitors, Mater. Today Energy, 2024, 42, 101432 CrossRef.
- C. Zhang, Y. Ding, L. Zhang, X. Wang, Y. Zhao and X. Zhang, et al., A Sustainable Redox-Flow Battery with an Aluminum-Based, Deep-Eutectic-Solvent Anolyte, Angew. Chem., Int. Ed., 2017, 56(26), 7454–7459 CrossRef CAS PubMed.
- X. Li, J.-Y. Li, M. F. Manzoor, Q.-Y. Lin, J.-l. Shen and L. Liao, et al., Natural deep eutectic solvent: A promising eco-friendly food bio-inspired antifreezing, Food Chem., 2024, 437, 137808 CrossRef CAS.
- A. S. Mallya, P. Yadav, S. Zakhia and A. Hubel, Low-temperature Characterization of Sugar Alcohol-based Type V Deep Eutectic Solvents for Anti-icing and Cryopreservation Applications, Sustain. Chem. Pharm., 2025, 48, 102256 CrossRef CAS PubMed.
- K. Hornberger, R. Li, A. R. C. Duarte and A. Hubel, Natural deep eutectic systems for nature-inspired cryopreservation of cells, AIChE J., 2021, 67(2), e17085 CrossRef CAS.
- M. T. Fu, H. M. Zhang, J. L. Bai, M. T. Cui, Z. Y. Liu and X. J. Kong, et al., Application of Deep Eutectic Solvents with Modern Extraction Techniques for the Recovery of Natural Products: A Review, ACS Food Sci. Technol., 2025, 5(2), 444–461 CrossRef CAS.
- V. Gupta, R. Thakur and A. B. Das, Effect of natural deep eutectic solvents on thermal stability, syneresis, and viscoelastic properties of high amylose starch, Int. J. Biol. Macromol., 2021, 187, 575–583 CrossRef CAS PubMed.
- Y. Tian, D.-W. Sun, L. Xu, T.-H. Fan, S.-T. Zhang and Z. Zhu, Bioinspired Cryoprotectants Enabled by Binary Natural Deep Eutectic Solvents for Sustainable and Green Cryopreservation, ACS Sustain. Chem. Eng., 2022, 10(23), 7677–7691 CrossRef CAS.
- R. Craveiro, V. I. B. Castro, M. T. Viciosa, M. Dionísio, R. L. Reis and A. R. C. Duarte, et al., Influence of natural deep eutectic systems in water thermal behavior and their applications in cryopreservation, J. Mol. Liq., 2021, 329, 115533 CrossRef CAS.
- W. Kauzmann, The Nature of the Glassy State and the Behavior of Liquids at Low Temperatures, Chem. Rev., 1948, 43(2), 219–256 CrossRef CAS.
- S. Bojic, A. Murray, B. L. Bentley, R. Spindler, P. Pawlik and J. L. Cordeiro, et al., Winter is coming: the future of cryopreservation, BMC Biol., 2021, 19(1), 56 CrossRef PubMed.
- M. Sansinena, M. V. Santos, G. Taminelli and N. Zaritky, Implications of storage and handling conditions on glass transition and potential devitrification of oocytes and embryos, Theriogenology, 2014, 82(3), 373–378 CrossRef CAS PubMed.
- S. Kavian, R. Sellers, G. A. Sanchez, C. Alvarez, G. Aguilar and M. J. Powell-Palm, Higher glass transition temperatures reduce thermal stress cracking in aqueous solutions relevant to cryopreservation, Sci. Rep., 2025, 15(1), 27903 CrossRef CAS PubMed.
- N. Rieke, J. Hancox, W. Li, F. Milletarì, H. R. Roth and S. Albarqouni, et al., The future of digital health with federated learning, NPJ Digit. Med., 2020, 3, 119 CrossRef PubMed.
- A. Karuth, A. Alesadi, W. Xia and B. Rasulev, Predicting glass transition of amorphous polymers by application of cheminformatics and molecular dynamics simulations, Polymer, 2021, 218, 123495 CrossRef CAS.
- G. M. Casanola-Martin, A. Karuth, H. Pham-The, H. González-Díaz, D. C. Webster and B. Rasulev, Machine learning analysis of a large set of homopolymers to predict glass transition temperatures, Commun. Chem., 2024, 7(1), 226 CrossRef CAS PubMed.
- L. B. Ayres, F. J. V. Gomez, M. F. Silva, J. R. Linton and C. D. Garcia, Predicting the formation of NADES using a transformer-based model, Sci. Rep., 2024, 14, 2715 CrossRef CAS PubMed.
- X. Xiao, D. Kong, X. Qiu, W. Zhang, F. Zhang and L. Liu, et al., Shape-Memory Polymers with Adjustable High Glass Transition Temperatures, Macromolecules, 2015, 48(11), 3582–3589 CrossRef CAS.
- M. Madadi, E. Kargaran, S. S. Hashemi, C. Sun, J. F. M. Denayer and K. Karimi, et al., Scalable Lignin Monomer Production Via Machine Learning-Guided Reductive Catalytic Fractionation of Lignocellulose, Adv. Sci., 2025, 12(42), e10496 CrossRef CAS PubMed.
- M. Madadi, E. Kargaran, S. Al Azad, M. Saleknezhad, E. Zhang and F. Sun, Machine learning-driven optimization of biphasic pretreatment conditions for enhanced lignocellulosic biomass fractionation, Energy, 2025, 326, 136241 CrossRef CAS.
- H. Wang, T. Fu, Y. Du, W. Gao, K. Huang and Z. Liu, et al., Scientific discovery in the age of artificial intelligence, Nature, 2023, 620(7972), 47–60 CrossRef CAS PubMed.
- C. Kaspar, B. J. Ravoo, W. G. van der Wiel, S. V. Wegner and W. H. P. Pernice, The rise of intelligent matter, Nature, 2021, 594(7863), 345–355 CrossRef CAS.
- T. Puzyn, B. Rasulev, A. Gajewicz, X. Hu, T. P. Dasari and A. Michalkova, et al., Using nano-QSAR to predict the cytotoxicity of metal oxide nanoparticles, Nat. Nanotechnol., 2011, 6(3), 175–178 CrossRef CAS PubMed.
- M. Pourmousa, S. Jain, E. Barnaeva, W. Jin, J. Hochuli and Z. Itkin, et al., AI-driven discovery of synergistic drug combinations against pancreatic cancer, Nat. Commun., 2025, 16(1), 4020 CrossRef CAS.
- E. N. Muratov, J. Bajorath, R. P. Sheridan, I. V. Tetko, D. Filimonov and V. Poroikov, et al., QSAR without borders, Chem. Soc. Rev., 2020, 49(11), 3525–3564 RSC.
- M. Popova, O. Isayev and A. Tropsha, Deep reinforcement learning for de novo drug design, Sci. Adv., 2018, 4(7), eaap7885 CrossRef CAS PubMed.
- D. Fourches, E. Muratov and A. Tropsha, Trust, but verify: on the importance of chemical structure curation in cheminformatics and QSAR modeling research, J. Chem. Inf. Model., 2010, 50(7), 1189–1204 CrossRef CAS PubMed.
- A. A. Toropov, B. F. Rasulev and J. Leszczynski, QSAR modeling of acute toxicity by balance of correlations, Bioorg. Med. Chem., 2008, 16(11), 5999–6008 CrossRef CAS PubMed.
- M. E. Erickson, M. Ngongang and B. Rasulev, A Refractive Index Study of a Diverse Set of Polymeric Materials by QSPR with Quantum-Chemical and Additive Descriptors, Molecules, 2020, 25(17), 3772 CrossRef CAS.
- N. Sizochenko, B. Rasulev, A. Gajewicz, E. Mokshyna, V. E. Kuz'min and J. Leszczynski, et al., Causal inference methods to assist in mechanistic interpretation of classification nano-SAR models, RSC Adv., 2015, 5(95), 77739–77745 RSC.
- H. Tran, R. Gurnani, C. Kim, G. Pilania, H.-K. Kwon and R. P. Lively, et al., Design of functional and sustainable polymers assisted by artificial intelligence, Nat. Rev. Mater., 2024, 9(12), 866–886 CrossRef CAS.
- J. M. Gregoire, L. Zhou and J. A. Haber, Combinatorial synthesis for AI-driven materials discovery, Nat. Synth., 2023, 2(6), 493–504 CrossRef CAS.
- X. Zhong, B. Gallagher, S. Liu, B. Kailkhura, A. Hiszpanski and T. Y.-J. Han, Explainable machine learning in materials science, NPJ Comput. Mater., 2022, 8(1), 204 CrossRef.
- L. Ahmed, B. Rasulev, S. Kar, P. Krupa, M. A. Mozolewska and J. Leszczynski, Inhibitors or toxins? Large library target-specific screening of fullerene-based nanoparticles for drug design purpose, Nanoscale, 2017, 9(29), 10263–10276 RSC.
- Y. Zhuravskyi, K. Iduoku, M. E. Erickson, A. Karuth, D. Usmanov and G. Casanola-Martin, et al., Quantitative Structure-Permittivity Relationship Study of a Series of Polymers, ACS Mater. Au, 2024, 4(2), 195–203 CrossRef CAS PubMed.
- A. Daghighi, G. M. Casanola-Martin, K. Iduoku, H. Kusic, H. González-Díaz and B. Rasulev, Multi-Endpoint Acute Toxicity Assessment of Organic Compounds Using Large-Scale Machine Learning Modeling, Environ. Sci. Technol., 2024, 58(23), 10116–10127 CrossRef CAS.
- A. Karuth, S. Szwiec, G. M. Casanola-Martin, A. Khanam, M. Safaripour and D. Boucher, et al., Integrated machine learning, computational, and experimental investigation of compatibility in oil-modified silicone elastomer coatings, Prog. Org. Coat., 2024, 193, 108526 CrossRef CAS.
- A. K. Lavrinenko, I. Y. Chernyshov and E. A. Pidko, Machine Learning Approach for the Prediction of Eutectic Temperatures for Metal-Free Deep Eutectic Solvents, ACS Sustain. Chem. Eng., 2023, 11(42), 15492–15502 CrossRef CAS.
- A. Khajeh, M. Shakourian-Fard and K. Parvaneh, Quantitative structure-property relationship for melting and freezing points of deep eutectic solvents, J. Mol. Liq., 2021, 321, 114744 CrossRef CAS.
- M. Abdollahzadeh, M. Khosravi, B. H. Kh. Masjidi, A. S. Behbahan, A. Bagherzadeh and A. Shahkar, et al., Estimating the density of deep eutectic solvents applying supervised machine learning techniques, Sci. Rep., 2022, 12(1), 4954 CrossRef CAS PubMed.
- L. B. Ayres, M. Bandara, C. D. McMillen, W. T. Pennington and C. D. Garcia, eutXG: A Machine-Learning Model to Understand and Predict the Melting Point of Novel X-Bonded Deep Eutectic Solvents, ACS Sustain. Chem. Eng., 2024, 12(30), 11260–11273 CrossRef CAS.
- V. Odegova, A. Lavrinenko, T. Rakhmanov, G. Sysuev, A. Dmitrenko and V. Vinogradov, DESignSolvents: an open platform for the search and prediction of the physicochemical properties of deep eutectic solvents, Green Chem., 2024, 26(7), 3958–3967 RSC.
- M. Mohan, K. D. Jetti, M. D. Smith, O. N. Demerdash, M. K. Kidder and J. C. Smith, Accurate Machine Learning for Predicting the Viscosities of Deep Eutectic Solvents, J. Chem. Theory Comput., 2024, 20(9), 3911–3926 CrossRef CAS PubMed.
- A. Joules, T. Burrows, I. P. Dosa and A. Hubel, Characterization of eutectic mixtures of sugars and sugar-alcohols for cryopreservation, J. Mol. Liq., 2023, 371, 120937 CrossRef CAS PubMed.
- A. D. Hunter, ACD/ChemSketch 1.0 (freeware); ACD/ChemSketch 2.0 and its tautomers, dictionary, and 3D plug-ins; ACD/HNMR 2.0; ACD/CNMR 2.0, ACS Publications, 1997 Search PubMed.
- A. Mauri, alvaDesc: A tool to calculate and analyze molecular descriptors and fingerprints, in Ecotoxicological QSARs, Springer, 2020, pp. 801–820 Search PubMed.
- R. A. Mahini, G. Casanola-Martin, S. A. Ludwig and B. Rasulev, MixtureMetrics: A comprehensive package to develop additive numerical features to describe complex materials for machine learning modeling, SoftwareX, 2024, 28, 101911 CrossRef.
- R. A. Mahini, G. Casanola-Martin, S. Szwiec, S. A. Ludwig and B. Rasulev, CombinatorixPy: Advancing mixture descriptors for computational chemistry, SoftwareX, 2025, 29, 102060 CrossRef.
- J. M. Stokes, K. Yang, K. Swanson, W Jin, A. Cubillos-Ruiz and N. M. Donghia, et al., A Deep Learning Approach to Antibiotic Discovery, Cell, 2020, 180(4), 688–702.e13 CrossRef CAS PubMed.
- Z. Tanoli, A. Fernández-Torras, U. O. Özcan, A. Kushnir, K. M. Nader and Y. Gadiya, et al., Computational drug repurposing: approaches, evaluation of in silico resources and case studies, Nat. Rev. Drug Discovery, 2025, 24(7), 521–542 CrossRef CAS PubMed.
- V. I. B. Castro, R. Craveiro, J. M. Silva, R. L. Reis, A. Paiva and A. R. C. Duarte, Natural deep eutectic systems as alternative nontoxic cryoprotective agents, Cryobiology, 2018, 83, 15–26 CrossRef CAS PubMed.
- Y. Dai, J. van Spronsen, G. J. Witkamp, R. Verpoorte and Y. H. Choi, Natural deep eutectic solvents as new potential media for green technology, Anal. Chim. Acta, 2013, 766, 61–68 CrossRef CAS PubMed.
- L. K. Savi, M. C. G. C. Dias, D. Carpine, N. Waszczynskyj, R. H. Ribani and C. W. I. Haminiuk, Natural deep eutectic solvents (NADES) based on citric acid and sucrose as a potential green technology: a comprehensive study of water inclusion and its effect on thermal, physical and rheological properties, Int. J. Food Sci. Technol., 2019, 54(3), 898–907 CrossRef CAS.
- A. S. Mallya, P. Yadav, S. Zakhia and A. Hubel, Computational Design of Natural Deep Eutectic Systems Using COSMO-RS for Ice Control Applications, ACS Sustain. Chem. Eng., 2025, 13(36), 14683–14692 CrossRef CAS PubMed.
- M. U. H. Joardder, M. H. Bosunia, M. M. Hasan, A. A. Ananno and A. Karim, Significance of Glass Transition Temperature of Food Material in Selecting Drying Condition: An In-Depth Analysis, Food Rev. Int., 2024, 40(3), 952–973 CrossRef CAS.
- L. McInnes, J. Healy, N. Saul and L. Großberger, UMAP: Uniform Manifold Approximation and Projection, J. Open Source Softw., 2018, 3(29), 861 CrossRef.
- M. Rugard, T. Jaylet, O. Taboureau, A. Tromelin and K. Audouze, Smell compounds classification using UMAP to increase knowledge of odors and molecular structures linkages, PLoS One, 2021, 16(5), e0252486 CrossRef CAS PubMed.
- N. Moshkov, T. Becker, K. Yang, P. Horvath, V. Dancik and B. K. Wagner, et al., Predicting compound activity from phenotypic profiles and chemical structures, Nat. Commun., 2023, 14(1), 1967 CrossRef CAS.
- P. J. Rousseeuw, Silhouettes: A graphical aid to the interpretation and validation of cluster analysis, J. Comput. Appl. Math., 1987, 20, 53–65 CrossRef.
- D. L. Davies and D. W. Bouldin, A Cluster Separation Measure, IEEE Trans. Pattern Anal. Mach. Intell., 1979, PAMI-1(2), 224–227 Search PubMed.
- L. Breiman, Random Forests, Mach. Learn., 2001, 45(1), 5–32 CrossRef.
- L. Prokhorenkova, G. Gusev, A. Vorobev, A. V. Dorogush and A. Gulin, CatBoost: unbiased boosting with categorical features. NIPS'18: Proceedings of the 32nd International Conference on Neural Information Processing Systems, 2018, pp. 6639–49.
- Xgboost: A scalable tree boosting system. Proceedings of the 22nd acm sigkdd international conference on knowledge discovery and data mining, ed. T. Chen and C. Guestrin, 2016.
- A. Natekin and A. Knoll, Gradient boosting machines, a tutorial, Front. Neurorobot., 2013, 7, 21 Search PubMed.
- G. Ke, Q. Meng, T. Finley, T. Wang, W. Chen and W. Ma, et al., LightGBM: A highly efficient gradient boosting decision tree, Adv. Neural Inf. Process. Syst., 2017, 30, 3146–3154 Search PubMed.
- A. J. Smola and B. Schölkopf, A tutorial on support vector regression, Stat. Comput., 2004, 14(3), 199–222 CrossRef.
- Z. Zhang, Introduction to machine learning: k-nearest neighbors, Ann. Transl. Med., 2016, 4(11), 218 CrossRef PubMed.
- C. K. I. Williams and C. E. Rasmussen, Gaussian processes for regression, Adv. Neural Inf. Process. Syst., 1995, 8, 514–520 Search PubMed.
- E. Grossi and M. Buscema, Introduction to artificial neural networks, Eur. J. Gastroenterol. Hepatol., 2007, 19(12), 1046–1054 CrossRef PubMed.
- R. Tibshirani, Regression shrinkage and selection via the lasso, J. R. Stat. Soc. Ser. B Stat. Methodol., 1996, 58(1), 267–288 CrossRef.
- G. Zengin, R. Ceylan, J. Katanic, A. Aktumsek, S. Matic and T. Boroja, et al., Exploring the therapeutic potential and phenolic composition of two Turkish ethnomedicinal plants – Ajuga orientalis L. and Arnebia densiflora (Nordm.) Ledeb, Ind. Crops Prod., 2018, 116, 240–248 CrossRef CAS.
- B. H. Menze, B. M. Kelm, R. Masuch, U. Himmelreich, P. Bachert and W. Petrich, et al., A comparison of random forest and its Gini importance with standard chemometric methods for the feature selection and classification of spectral data, BMC Bioinf., 2009, 10(1), 213 CrossRef PubMed.
- S. Nembrini, I. R. König and M. N. Wright, The revival of the Gini importance?, Bioinformatics, 2018, 34(21), 3711–3718 CrossRef CAS PubMed.
- A. Altmann, L. Toloşi, O. Sander and T. Lengauer, Permutation importance: a corrected feature importance measure, Bioinformatics, 2010, 26(10), 1340–1347 CrossRef CAS PubMed.
- S. M. Lundberg and S.-I. Lee, A unified approach to interpreting model predictions, Adv. Neural Inf. Process. Syst., 2017, 30, 4768–4777 Search PubMed.
- N. Li, A. Lewin, S. Ning, M. Waito, M. P. Zeller and A. Tinmouth, et al., Privacy-preserving federated data access and federated learning: Improved data sharing and AI model development in transfusion medicine, Transfusion, 2025, 65(1), 22–28 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |