 Open Access Article
Open Access ArticleAuLCA: augmented life cycle assessment for chemical data gaps
Maximilian G. Hoepfner†
 ab,
Dion Jakobs†
ab,
Dion Jakobs† a,
Lucas F. Santos
a,
Lucas F. Santos ab and
Gonzalo Guillén-Gosálbez
ab and
Gonzalo Guillén-Gosálbez *ab
*ab
aInstitute for Chemical and Bioengineering, Department of Chemistry and Applied Biosciences, ETH Zurich, Vladimir-Prelog-Weg 1, 8093 Zurich, Switzerland. E-mail: gonzalo.guillen.gosalbez@chem.ethz.ch
bNCCR Catalysis, Zürich CH-8093, Switzerland
First published on 14th April 2026
Abstract
Life cycle assessment (LCA) has become the prevalent tool to quantify the impact of chemical processes, yet data gaps remain a major obstacle towards its widespread adoption. Existing LCA databases cover a few thousand, mostly high-production-volume, chemicals; however, fine chemicals are often underrepresented. Here we introduce an augmented LCA (AuLCA) framework based on chemical reaction networks (CRN), mass-based impact propagation, and first principles-based energy estimations to predict the life cycle inventories and impacts of chemicals. By applying AuLCA to four case studies, we find good agreement with commercial data, with the accuracy level depending on the chemical reaction network's size and density. Overall, AuLCA is intended to support sustainable decision-making across chemical scales, particularly in early-stage decisions on chemical reaction pathways selection.
Green foundation1. We provide an algorithm to cover data gaps in life cycle assessment, the fundamental tool to quantify the environmental impact of chemical systems investigated in Green Chemistry. By combining reaction networks with mass-based allocation and first-principles energy consumption approximations, we estimate life cycle impacts to guide the discovery of greener synthesis routes, even when only scarce data are available.2. This work estimates the life cycle footprint of thousands of chemicals that are missing in commercial databases. Our approach simplifies the time-consuming LCA data collection phase, covering chemicals that previously lacked LCA data and were hard to model due to the multiple synthesis steps involved. 3. Beyond improving our LCA augmentation algorithm, i.e., by refining solvent and yield estimates, the crucial next step is to optimize synthesis pathways based on sustainability criteria and make the refined tool available to the Green Chemistry community so LCA becomes more widely adopted in the field. This will allow benchmarking chemical systems, including syntheses routes, to identify greener alternatives, providing measurable insights and trends to support sustainable development. |
Introduction
Since their inception in 1998, green chemistry principles have shaped chemistry towards a new paradigm where sustainability is not an afterthought but is a foundational goal.1,2 Today, chemists and engineers guided by these principles seek to design chemical processes and products with lower environmental footprint to ensure long-term ecological balance.3Despite the clarity of this overarching goal, the path towards a truly sustainable chemical industry remains uncertain. The current chemical industry represents 10% of the global greenhouse gas (GHG) emissions and is classified as a hard-to-abate sector, making it a critical target for sustainability-driven actions.4 Multiple green chemical technologies are being investigated, whose sustainability performance needs to be quantified using metrics to support experimental research, technology deployment, and policymaking.
The environmental impact of chemical routes was originally assessed via mass- and energy-based process level metrics such as the E-factor, Atom Economy, or Process Mass Intensity (PMI).5–7 Although these metrics address key environmental aspects such as resource utilization and waste prevention, they provide limited information on the full environmental footprint of complex molecules throughout their lifecycle.8,9 Complementing these metrics, Life Cycle Impact Assessment (LCIA) methods10 quantify the environmental, human health, and resources impacts of chemical systems across their full life cycle, encompassing the resource extraction, manufacturing, transportation, and end-of-life phases. LCA studies allow identifying environmentally detrimental processes,11–13 the occurrence of burden-shifting (collateral damage) across environmental categories,14–16 and the most critical parameters affecting environmental impacts.17,18
However, completing an LCA is a time- and resource-intensive task that requires detailed accounting of all material and energy flows along the life cycle of the reference product, most of which are hard to collect in practice. Consequently, LCAs are frequently only completed retrospectively for processes fully characterized and developed, thus reducing the opportunity for LCA results to influence early-stage chemical exploration.4,19 Although some early-stage LCAs have been conducted,11,20–22 performing full LCAs for hypothetical synthetic routes remains challenging due to lack of data. Moreover, even LCAs of already existing chemicals may face many data gaps,23 as discussed below, thus hampering the sustainable chemicals transition.
LCA databases, such as ecoinvent,24 only contain hundreds to thousands of chemicals, representing a small fraction of the over 279 million registered substances,25 which severely limits sustainability assessments. This is particularly true in fine chemicals (e.g., active compounds in pharma, pesticides, additives, etc.), whose synthesis typically involves multiple reaction steps entailing diverse reagents, solvents, and catalysts, which are seldom publicly disclosed.
Several predictive LCA (streamlined LCA) methodologies have been developed to cover LCA data gaps.26–29 They estimate cradle-to-gate or gate-to-gate LCA impacts using approximations, where the recent trend is to leverage machine learning algorithms to correlate basic features (e.g., molecular structure, thermodynamic properties, etc.) with environmental impacts.30–32 Commonly used machine learning methods include artificial neural networks,19,32–35 support vector machines and Gaussian process regressors (SVM/GPRs),33,36 and, more recently, transformers,29 amongst others.37,38 Additionally, optimization-based methods28,39 and similarity matrices40 have also been applied to the same problem.
Most streamlined LCA methodologies are based on regression approaches calibrated with a training set of chemical footprints. These methods are often based on the chemical's structure and properties while the underlying reaction pathway is not explicitly considered, although it plays a key role in the chemical's footprint. Ethylene, for example, could be synthesized from naphtha in the steam cracker, and also produced via dehydration of bioethanol, leading to completely different environmental footprints.
Moreover, such regression tools are trained for specific classes of chemicals and impact metrics, thus providing lower accuracies when extrapolating beyond the training set.
Here, we address these limitations by developing a novel Augmented Life Cycle Assessment (AuLCA) methodology that integrates chemical reaction networks, first-principles-based energy estimations, and mass-based propagation of LCA data. Using data from ecoinvent version 3.9.1,24 we show that AuLCA provides sensible predictions, more so when the reaction pathway is known.
Overall, AuLCA aims to facilitate the broad application of LCA to support better informed, transparent, and reliable sustainable decision-making across chemical scales.
Methods
AuLCA comprises several core components, which we describe next considering the phases of a standard LCA. First, the goal and scope of the LCA study are defined (Fig. 1A). Then, data collection and curation are performed (Fig. 1B). This involves constructing chemical reaction networks (CRNs) around a corpus of chemicals whose life cycle inventories (LCI) are known and which we here retrieve from a commercial database. These LCIs will be then propagated across the network. Next, the impacts of the missing chemicals will be predicted from the propagated LCIs of the molecules in the corpus and the gate-to-gate emissions linked to the associated reaction pathways (Fig. 1C). Finally, the predicted impacts are analysed. Here, a leave-one-out validation approach is applied to assess the performance of the algorithm (Fig. 1D) in four different case studies. | ||
| Fig. 1 Four-step framework of the AuLCA tool: (A) Goal and Scope definition, (B) data curation and network construction, (C) impact augmentation and (D) data analysis and validation of predictions. | ||
Step 1. Goal and scope of the analysis
The goal of the study is to compute the environmental footprint of 1 kg of chemical (functional unit) following a cradle-to-gate scope. The challenge here is that the LCI of this chemical is unknown and must be estimated before its impact can be computed.Step 2. Reaction network construction
We initiate our workflow to estimate the impact by defining a corpus of chemicals and collecting their associated LCIs, in our case retrieved from ecoinvent v3.9.1 database in accordance with ISO 14044/14040. To this end, similar filtering criteria as in Lucas et al.8 are applied using the brightway25 package41 in python v3.12 (Fig. 5 in the SI for more information). The LCIs are defined for the functional unit of 1 kg of chemical. Next, CRNs are built around the chemicals in the corpus by identifying the chemical neighbourhood around them (i.e., set of chemical reactions transforming the molecules in the corpus into other molecules). Due to the flexible input data strategy, both the corpus of chemicals as well as the CRNs can be updated to include the most recent data as well as user-specific tailored datasets. This structure ensures that common challenges, e.g., the increasing number of novel chemicals and new (and updated) characterization factors, could be easily addressed without changing the algorithm structure itself.Ideally, we would employ chemical databases to build CRNs, iteratively expanding the molecules in the corpus as many times as desired via chemical reactions retrieved from the chemical database. Alternatively, open databases (e.g., USPTO and the CJHIF chemical reaction dataset (CRD)42) might be used instead. Despite being free, the latter often contain fewer reactions, thus preventing the use of some molecules in the corpus (i.e., those not appearing in any reaction in the open CRN) and requiring some additional approximations in the calculations, as discussed later in the article.
The goal of the analysis is to estimate the LCIs for the nodes in the network with unknown footprint connected to the nodes in the corpus with known footprint.
Several data curation strategies might be required to prepare the CRNs before data augmentation (section 1 of the SI) to ensure they can be used for LCA augmentation, regardless of their source.
Step 3. Data augmentation
We first provide the general nomenclature followed in the method derivation. The CRN is represented as a directed bi-partite graph consisting of nodes (chemicals and reactions) and edges (reactant and product connections between reactions and chemicals). Reactions r belong to the set of all reactions R in the graph. The chemical nodes n belong to the set S := SKi ∪ SUi, which is given, at iteration i∈I of the algorithm, by the union of chemicals with known (in the corpus) or estimated (via data augmentation) LCA data (SKi), and the set of chemicals whose footprint we still seek to estimate (SUi). Hence, the intersection of SKi ∩ SUi is empty at any iteration i of the algorithm (SKi ∩ SUi = ∅), as the LCI of a chemical is either known or unknown, but not both at the same time. With proceeding iterations of the algorithm, chemicals from the set SUi are gradually added to the set SKi.Ultimately, all chemicals in SUi will belong to the set SKi after completing the calculations in the last iteration of the data augmentation step. At iteration zero, set SKi contains the chemicals whose LCI data have been retrieved from the commercial database (e.g., the corpus), denoted here as chemicals in the set SEI. Therefore, at iteration 0, SK0 = SEI applies.
Moreover, we define additional sets used in the derivation of the algorithm. Specifically, for all reactions r in the network (∀r∈R), we define the set of corresponding reactants linked to the incoming nodes n (SINr) based on their mass-based stoichiometric coefficient Vn,r∈![[Doublestruck R]](https://www.rsc.org/images/entities/char_e175.gif) 2 with r∈R and n∈S, as given below:
2 with r∈R and n∈S, as given below:
| SINr = {n∈S|νn,r < 0} ∀r∈R | (1) |
Conversely, for products (SOUTr) of a reaction r, we have:
| SOUTr = {n∈S|νn,r > 0} ∀r∈R | (2) |
Similarly, the set of reactions that produce (RPRODn) or consume (RCONSn) chemical node n, respectively, are defined as following:
| RPRODn = {r∈R|n∈SOUTr} ∀n∈S | (3) |
| RCONSn = {r∈R|n∈SINr} ∀n∈S | (4) |
In Fig. 2 an example of a reaction (a), with two reactants (1,2) and two products (3,4) is given. In iteration i = 0 of the algorithm, only 1 and 2 are modelled with LCIs retrieved from a commercial database, while in the next iteration i = 1, we shall compute the LCI for chemicals 3, 4, which will then join set SKi.
The data augmentation algorithm departs from the molecules in the commercial databases, e.g., the corpus, but in performing the data augmentation it may face data gaps (i.e., the footprint of some reactants might be missing when attempting to compute the footprint of the reaction products). This might happen because the chemical database contains reactions that depart from a molecule in the corpus but require additional reactants missing in the corpus. Hence, a strategy is required to rank the reactions based on data availability, prioritizing data augmentation in those reactions where more information is at hand. Therefore, to guide the LCI prediction across the CRN, we rank the reactions r and nodes n in the set SUi (those whose footprint is yet to be estimated) based on the herein introduced Availability Factor (AF).
The AF quantifies the amount of LCI data available in each reaction yielding each compound in SUi. In each iteration, the compounds and synthesis routes with the highest AF are prioritized for data augmentation.
Hence, parameter AFPi,n is first defined in every iteration i for every node n as follows:
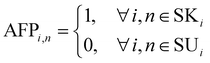 | (5) |
The set of nodes connected to reaction r, denoted by Nr, is defined as follows:
| Nr = SINr∪SOUTr | (6) |
The AF for node n, in reaction r, at iteration i, is calculated as:
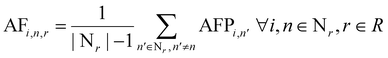 | (7) |
For each iteration i, we define the maximum AF over all reactions r involving nodes n as:
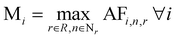 | (8) |
Finally, we can define RAFMaxi as the set of all reactions that actually reach the maximum AF in iteration i:
| RAFMaxi = {r∈R|n∈Nr, AFi,n,r = Mi} ∀i | (9) |
After identifying the order in which calculations will be performed, we next conduct the subsequent impact augmentation. Here, our goal is to determine the life cycle assessment inventories (and impacts LCAIk,n) of the missing chemicals. Impacts can be obtained from the LCI using characterisation factors CFj,k that map flow j, of all environmental flows J, to the impact category index k of all impact categories K.
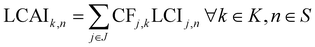 | (10) |
The life cycle inventory (LCIj,n) for every node n∈S in the reaction network is given by the LCIs embodied in the reactants in the reaction yielding the missing chemical, plus the LCI flows linked to the chemical transformation, including the reaction energy and the energy required in the downstream separations. In what follows, LCIRMj,n,r is the inventory given by the mass-based allocation of the LCIs of the corresponding reactants in node n for reaction r, LCIRj,n,r is the inventory linked to the reaction energy in reaction r for node n, while the LCI connected to the separation energy of reaction r is denoted by LCISj,r.
As different reactions might point to the same chemical (i.e., different alternative pathways might yield the same molecule), we shall compute the average of LCIs estimated across pathways as follows:
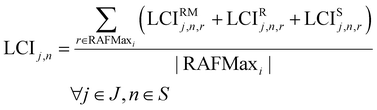 | (11) |
The reaction energy inventory LCIRj,n,r is calculated using the mass-based enthalpy of formation 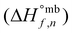 of all reactants and products of the reaction and normalizing by the stoichiometry of the node of interest. Moreover, once the energy requirements are computed, they are converted into the corresponding LCI using parameter LCIHEATj (i.e., inventory of the heating or cooling agent, retrieved from an environmental database).
of all reactants and products of the reaction and normalizing by the stoichiometry of the node of interest. Moreover, once the energy requirements are computed, they are converted into the corresponding LCI using parameter LCIHEATj (i.e., inventory of the heating or cooling agent, retrieved from an environmental database).
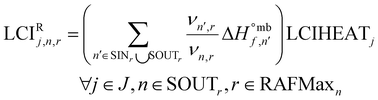 | (12) |
Similarly, the separation energy inventory LCISj,n,r is computed from the energy and solvent requirements for separations and the LCI of energy provision LCIHEATj, also retrieved from an environmental database.
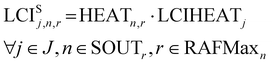 | (13) |
Here we estimate parameter HEATn,r, which quantifies the energy requirements for separations, following the heuristics in Gani et al.43 In essence, such heuristics provide suitable separation technologies for product n in reaction r. Once a suitable separation technology is identified, heuristics for energy and solvent requirements are applied. Herein, we focus on distillation, liquid–liquid extraction and recrystallization.
Finally, the inventory embodied in reactants LCIRMj,n,r is computed assuming a mass-based allocation method based on mass-based stoichiometry coefficients combined with the reactants’ life cycle inventories LCIj,n.
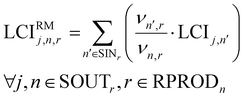 | (14) |
Hence, the overall algorithm (Algorithm 1) can be written in compact form as follows.
| Algorithm 1: augmented life cycle assessment (AuLCA) | ||
|---|---|---|
| 1 | i ← 0 | # Start at iteration 0 |
| 2 | SKi ← SEI | # Init. data |
| 3 | SUi ← Si /SKi | |
| 4 | Initialize LCAIk,n, LCIj,n for n∈SKi | # Corpus of LCA data |
| 5 | While SUi ≠ 0 | |
| 6 | r ← RAFMaxi | # Ranking |
| 7 | Compute LCAIk,n, LCIj,n | |
| 8 | SKi ← SKi ∪ n | |
| 9 | SUi ← SUi /n | |
| 10 | i ← i + 1 | # Next iteration |
| 11 | Return LCAIk,n | |
Fig. 3 illustrates how Algorithm 1 works within a simple reaction network, containing seven chemicals (1, 2, 3, 4, 5, 6, 7) and three reactions (a, b, c). In iteration i = 0, all chemicals in the corpus SEI = {1, 2, 4} are added to SKi. Meanwhile, SUi = {3, 5, 6, 7} ≠ ∅. Using the availability factor, n = 3 in reaction r = a, is selected as the next node, due to the highest availability factor being RAFMaxi,n = a. After computing LCAIn from LCIj,n, the algorithm then proceeds to the next iterations until the loop stops with SUi = 0. See more detailed examples in section 2 of the SI.
 | ||
| Fig. 3 Schematic of Algorithm 1, based on a simple reaction network with three reactions (a, b, c) and with seven chemicals. In total ten nodes, i.e., chemicals and reactions. | ||
Step 4. Data validation and analysis
Here the algorithm is validated against known data and insights are derived from the analysis of the LCA results generated, as discussed in the next section.Results
The performance of the algorithm is evaluated using data in ecoinvent (compare Fig. S5 in the SI). This database mostly contains high production volume chemicals, thereby affecting the accuracy of the results (see limitations and assumptions in the SI). An external validation using the IDEA database44 can also be found in the SI, which leads to similar results.Predictions for known synthesis routes
We first evaluate the algorithm performance by assuming that the reaction pathway is known, and only gate-to-gate emissions are unknown. To this end, we consider 13 exemplary organic chemicals in ecoinvent for the validation, reconstructing the associated reaction pathways based on the activity description and material flows in ecoinvent (Table 4 in the SI). For the prediction, we only consider one synthesis step at a time, namely the single-step transformation of a specific chemical in ecoinvent into one of the said 13 chemicals, all of which are also in ecoinvent. Subsequently, reaction networks containing only a single reaction were curated from the retrieved ecoinvent data. This approach serves as an internal benchmark, to ensure that the AuLCA algorithm accurately captures the environmental impacts when only gate-to-gate emissions need to be predicted for a known single-step synthesis path. Next, AuLCA was applied to each chemical focusing on predicting the IPCC 2021 global warming potential (GWP), which was then compared with the corresponding reference values in ecoinvent using the following metrics:45,46 root mean square error (RMSE), coefficient of determination (R2), mean absolute error (MAE), and the mean relative error (MRE).Fig. 4 shows that the predictions closely match the references (R2 = 0.97), with an RMSE of 0.74 kgCO2-eq per kg, an MAE of 0.52 kgCO2-eq per kg, and a mean relative error of 11%. Similar results are obtained when analysing the energy-related emissions rather than the total emissions (Table 4 of the SI). This analysis indicates that when reaction pathways are known predictions are accurate, suggesting that the mass allocation and gate-to-gate calculations work well in the cases analysed.
 | ||
| Fig. 4 Prediction performance of AuLCA based on manually selected chemicals from ecoinvent using the same reaction pathway. Values given in kgCO2eq per kg. | ||
Leave-one-out validation
We next evaluate the overall performance of AuLCA following a leave-one-out validation approach (LOOV). The LOOV was selected to use the available validation data to the maximum extent possible by ensuring that each compound in our dataset is utilized for validation while the model is trained on the remaining instances. Given the specific constraints of high-quality LCI benchmarks, LOOV provides a fair estimate of the model's generalization error and ensures a direct, comprehensive comparison against the employed LCA database. We define four different CRNs case studies, three based on open-source (OS) data of different sizes and one on Reaxys©‡ data47 (Table 1).| Networks | Open-source (OS) | Reaxys | ||
|---|---|---|---|---|
| Case study | CS I | CS II | CS III | CS IV |
| Reactions, r | 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
300![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
308![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500 500 |
| Chemicals, S | 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
140![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
360![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
310![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 000 |
| SK0 | 49 | 98 | 122 | 236 |
| SKLOOV | 12 | 34 | 41 | 110 |
We define the training set, e.g., the set SK0 corresponding to the corpus for each such case, as the intersection of all chemicals S in the graph (CRN) and the precomputed chemicals from ecoinvent v3.9.1 SEI (SK0 = S ∩ SEI). Since the set of all chemicals S within the CRN is different for each case study, the training set is different too. In addition, this set needs to be filtered, yielding a validation dataset SKLOOV that is used in the LOOV. Particularly, single atom compounds, PFAS, small complex molecules (e.g., SiCl4), inorganic chemicals, and heavy halogenated molecules were excluded from SK0 to build the validation set (SKLOOV⊆SK0)(full filtering criteria in section 3.2 of the SI). Again, due to differences among CRNs, the number of validation chemicals SKLOOV differs also across cases, affecting the comparisons.
During the LOOV, the LCI of one chemical in the validation set is removed from the training set SK0, and then predicted using AuLCA for each case, using the remaining known data for the other chemicals in the augmentation. Hence, the entire data augmentation procedure is repeated for as many times as chemicals in the validation set, leaving one of them out of the analysis at a time.
Table 2 summarizes the results. We find that larger networks often lead to better prediction performance, although recall that such performance is computed over training sets of different sizes. Specifically, in the OS case studies, moving from 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 to 100
000 to 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 and 300
000 and 300![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 reactions increased the size of the training set SK0 from 49 to 98 and 122, respectively. Note that the size of the training set SK0 does not increase in the same proportion as the network size does, e.g., for a tenfold increase (CS I vs. CS II), the size of the training set SK0 doubles only. This moderate growth of the training set SK0 can be explained by the fact that a high number of reactions in the OS networks do not contain any ecoinvent chemicals present in SEI. Consequently, expanding the network does not guarantee a commensurate expansion of the initial nodes in the set SK0. Regardless, performance tends to improve with larger training sets, e.g., larger corpus SK0, which improve data accuracy and enable a broader coverage of chemicals within the CRN, as discussed below. Furthermore, a higher degree of interconnection, especially within larger CRNs, allows for the identification of more alternative synthesis pathways, whereas the validation data typically rely on a single selected route. Since our approach averages values across all available pathways within the CRN, the prediction is strongly influenced by the diversity of the considered synthesis routes (i.e., number of routes in the network, where some will be more industrially relevant than other). In addition, sparse CRNs might be on the other hand unable to predict the right synthesis route due to lack of sufficient reaction connections within the network. Hence, there is a clear trade-off regarding network size, as larger networks will average values over a wider range of routes (being only some of them industrially relevant) and smaller ones might be unable to identify the most realistic pathway. This trade-off is discussed more in-depth later in the article.
000 reactions increased the size of the training set SK0 from 49 to 98 and 122, respectively. Note that the size of the training set SK0 does not increase in the same proportion as the network size does, e.g., for a tenfold increase (CS I vs. CS II), the size of the training set SK0 doubles only. This moderate growth of the training set SK0 can be explained by the fact that a high number of reactions in the OS networks do not contain any ecoinvent chemicals present in SEI. Consequently, expanding the network does not guarantee a commensurate expansion of the initial nodes in the set SK0. Regardless, performance tends to improve with larger training sets, e.g., larger corpus SK0, which improve data accuracy and enable a broader coverage of chemicals within the CRN, as discussed below. Furthermore, a higher degree of interconnection, especially within larger CRNs, allows for the identification of more alternative synthesis pathways, whereas the validation data typically rely on a single selected route. Since our approach averages values across all available pathways within the CRN, the prediction is strongly influenced by the diversity of the considered synthesis routes (i.e., number of routes in the network, where some will be more industrially relevant than other). In addition, sparse CRNs might be on the other hand unable to predict the right synthesis route due to lack of sufficient reaction connections within the network. Hence, there is a clear trade-off regarding network size, as larger networks will average values over a wider range of routes (being only some of them industrially relevant) and smaller ones might be unable to identify the most realistic pathway. This trade-off is discussed more in-depth later in the article.
| Networks | Open-source (OS) | Reaxys© | ||
|---|---|---|---|---|
| Case study | CS I | CS II | CS III | CS IV |
| RMSE [kgCO2eq per kg] | 3.80 | 3.65 | 2.79 | 2.97 |
| MAE [kgCO2eq per kg] | 2.53 | 2.52 | 1.98 | 2.27 |
| MRE [%] | 72.3 | 81.1 | 59.3 | 72.1 |
| R2 | 0.28 | 0.11 | 0.41 | −0.08 |
RMSE and MAE both improve when increasing the CRN and set SK0 size (Fig. 5). For example, comparing CS I and CS III, the RMSE is reduced from 3.80 kgCO2eq to 2.79 kgCO2eq, while the MAE decreases from 2.53 kgCO2eq to 1.98 kgCO2eq. The MRE follows a similar trend from CS I to CS III.
 | ||
| Fig. 5 Values given in kgCO2eq per kg. Correlations and residuals analysis of case studies. (A) case study I, (B) case study II, (C) case study III, (D) case study IV. Detailed overview of results can be found in Table 2. Grey area visualizes acceptable deviation from reference data with deviation of up to +100%/−50%. Sets of all employed chemicals used for LOOV can be found in the SI. | ||
The Reaxys©-based CRN strategy in CS IV shows comparable prediction accuracy, with an RMSE of 2.97 kgCO2eq, a MAE of 2.27 kgCO2eq, but a notably higher MRE of 72.1%. Note, however, that the training set SK0 and the validation set SKLOOV are nearly twice as large as in CS III, despite containing a similar number of reactions. This is because of the way we build the CRN using Reaxys© data, which allows including more ecoinvent chemicals in the corpus. The high MRE in CS IV arises because AuLCA tends to overestimate the GWP of low-GWP molecules, which are more frequent in the Reaxys©-based CRN due to the larger number of reactions involving such compounds.
While the prior performance metrics indicated good predictive performance, the R2 values across the case studies appear comparatively modest and inconsistent. These results can be attributed to the specific distribution of the validation data. Specifically, the low variance within the validation sets significantly penalizes the R2 calculation, resulting in low values even when absolute deviations are comparable small. This effect is particularly evident in CS IV, which includes multiple low-GWP compounds with a correspondingly low variance in the validation data. Consequently, these R2 values should be interpreted in the context of the data's narrow range rather than as a lack of model accuracy.
Fig. 5 (top) shows all the predicted values for the different cases. As seen, most of the chemicals, in all four case studies, are predicted with an error in the range of +100%/−50% (grey area in Fig. 6). However, the relative number of chemicals falling outside the grey area only slightly decreases as we move to larger networks (from 33.3% in CS I to 30.0% in CS IV). As in Table 2, the trend of the violin plots (Fig. 5, bottom) shows an increasing prediction accuracy from CS I to CS III. In CS IV, however, the violin plot reveals also a slight overestimation, i.e., distribution centered slightly above zero for the reasons previously discussed.
 | ||
| Fig. 6 Prediction error analysis for LOOV chemicals in CS III. Similar trend observed for case studies CS I, CS II and CS IV in the SI. | ||
To identify any biases in AuLCA, we analyse the prediction error in Fig. 6 (for CS III only, the other cases, which behave similarly, are provided in Fig. 8 of the SI), finding positive and negative prediction errors of similar magnitude. Hence, no strong systematic over- or underestimation is found across all predictions, suggesting the absence of any systematic bias. Further evaluation of the results shows that overestimation with higher errors is mainly associated with large scale, industrially synthesized, olefin-like chemicals, for which very well-established routes exist. Alternatively, their footprint is predicted by AuLCA using more complex synthesis pathways, thereby leading to larger GWPs. This mostly happens in CSI I–III, as the dataset contains only patents, some of which might have never been deployed at scale and may greatly differ from current industrial practice. In addition, this often leads to more synthesis steps and larger emissions.
Moreover, in CS IV, the standard routes to produce chemicals are averaged together with others, less common (and complex) ones, thereby leading as well to overpredictions. Conversely, underestimation often occurs in some molecules (e.g., acetylene) requiring reactants (i.e., oxygen contributes to 90% of its GWP) missing in the network and, therefore, assumed through proxies (see eqn (S13) in the SI) that tend to underestimate their true value. This emphasizes the importance of the AF-ranking to produce more accurate estimates, as discussed next.
Effect of the AF on the prediction
We next investigate the effect of the AF on the predictions made (Fig. 7). For this, we counted the number of unknown chemicals (SUi) within the CRNs in the OS case studies. To this end, we defined an AF threshold, which limits the augmentation to those chemicals generated in reactions with a minimum level of information available (i.e., minimum amount of LCI data available).Without setting an AF threshold, all chemicals can be computed. Introducing a modest threshold of 0.25 (i.e., at least 25% of reactants and products in each reaction must be known) reduces coverage: CS II and CS III are slightly affected, with 79% and 82% of chemicals still computable, whereas CS I drops to 68%. This effect is even more pronounced for a threshold of 0.5. Higher AF thresholds, e.g., >0.75, leave only a negligible fraction of chemicals computable for all cases.
This result indicates the poor degree of interconnections within smaller CRNs, such as in CS I. Here, most chemicals are computed using a limited number of reactions strongly affected by an AF threshold, while this does not happen to the same extent in CS II and CS III, which contain more reactions. Regardless of the case, it is obvious that approximations (eqn (1) in the SI) are required for making predictions in all networks, and less so in larger networks due to more interconnections. Notably, Reaxys© provides more alternative routes and many links between chemicals, thus being virtually insensitive to the AF threshold.
Fig. 8 provides an illustrative network example for the chemical morpholine in CS II and CS III. Imposing larger AFs leads to more convoluted networks because the algorithm tries to avoid proxies. This requires expanding further the synthesis routes to fully connect the ecoinvent chemicals with the target molecule without relying on so many approximations. Certainly, only two reactions remain the same in both cases, while the number of ecoinvent chemicals (stars) used in the predictions is similar in the two cases. The selected pathways for CS III are, hence, longer, involving more reactions and chemicals to bypass proxies (triangles).
 | ||
| Fig. 8 Visualization of sub-networks for CS II (green, left) and CS III (blue, right), to identify the computational graph for the case study chemical morpholine. | ||
With a larger repertoire of reactions, missing compounds can be inferred from preceding steps via impact propagation, thereby avoiding the use of less accurate proxies. In smaller CRNs, such as the 100k OS example (green, left side), fewer reactions are available, so missing chemicals cannot be inferred and must be assumed through proxies: CS III requires only two assumptions to compute morpholine's GWP, and CS II six. This observed behavior underlines the advantage of larger networks to reduce the number of necessary proxies. However, they also lead to more complex routes that might also differ from more direct ones, particularly when analyzing well-established bulk chemicals involving fewer synthesis steps like those in ecoinvent. Numerical examples indicate that avoiding proxies is more critical than minimizing pathway length, although the effect can be chemical–specific. Proxies fail to distinguish high- and low-impact compounds because they assign the average impact of the reactants. Consequently, it is preferable to predict missing compounds via longer, information-preserving routes, provided the added uncertainty remains low, as these better capture compound-specific impacts.
Gate-to-gate impact contributions
Fig. 9 shows the impact of reaction energy, separation, and mass-based allocation in case study CS III (similar patterns found for the remaining cases are shown in Fig. S9 of the SI). As seen, consistent with previous work,8 the prediction is dominated by the impact embodied in reactants (gate-to-gate impacts based on mass-based allocation in the range of 4.88–7.25 kgCO2-eq, CI 95%). This is because reactants already include gate-to-gate impacts from preceding steps, so applying mass-based allocation along a pathway propagates these upstream, energy-related impacts. Moreover, the distribution of the mass-based allocation impact is heavily influenced by the selection of the CRN.Energy requirements in separations clearly exceed those in the reaction. This is because of the often mild conditions at the reaction step, leading to small energy needs in contrast to the energy requirements of energy-intensive separation processes, particularly distillation. Furthermore, reaction energy impacts related to cooling duties for exothermic reactions were neglected and so were energy losses. Moreover, energy requirements in separations are more spread due to the different processes assumed for the individual reactions in a pathway.
Applicability to alternative LCIA methods
Due to the fact that the GWP is currently (arguably) the most critical metric for quantifying the environmental impact, the AuLCA algorithm was validated exclusively against the IPCC GWP 202148 methodology. Regardless, the AuLCA algorithm is fundamentally designed to generate LCIs compatible with any assessment framework and is inherently capable of integrating other alternative life cycle impact assessment (LCIA) methods, such as ReCiPe 201649 or EF 3.1.50Conclusions and outlook
Here we introduced a first principles-based method for estimating LCA data for fine chemicals. Our data augmentation algorithm AuLCA showed promising performance for covering data gaps, especially when considering the same pathway for reference values. The algorithm can easily perform hundreds of thousands of LCAs, demonstrating its scalability to large data sets.However, our approach is inherently limited by the quality and diversity of the input chemicals in the corpus and the CRNs employed in the LCA data propagation. In particular, the distribution and variance of chemicals in the corpus could be improved by including more fine chemicals. In addition, chemical diversity and network connectivity in the CRNs could be enhanced to maximize overall coverage and, thus, the predictive power.
A key next step is the integration of multiple reaction databases, potentially combining foundational CRNs, such as in CS IV, with more specialized CRNs, such as in CS I to CS III. This integration will expand the algorithm's capabilities across a broader chemical space and allow it to operate with higher AF thresholds. Moreover, gate-to-gate estimates will have to be refined, including the footprint of solvents and catalysts while considering more accurate yields. Based on the observed predictive performance, the AuLCA algorithm could provide a robust platform for generating preliminary LCA estimates. It could be leveraged to guide sustainable decision-making, with an emphasis on the selection of synthesis routes, particularly in the early stages of chemical process development, where data is often limited. Furthermore, the next version of the AuLCA framework could be provided in the form of a toolbox for LCA practitioners and chemists to generate environmental footprint estimates for a plethora of chemicals. Due to the open data structure, AuLCA will be able to support user–defined data to be tailored to the specific setups, including the integration of multiple LCIA methods, e.g., ReCiPe 2016.
Overall, AuLCA could support early-stage design decisions in a plethora of applications, with strong emphasis on guiding the sustainable scale-up of active compounds production in the pharma industry. Specifically, automating the evaluation of alternative routes, currently performed manually, would enable chemists and process engineers to identify and prioritize the most sustainable synthesis options more efficiently, helping to advance the goals of Green Chemistry in both research and industrial practice.
Author contributions
Authors M. G. H. and D. J. contributed equally. M. G. H.: conceptualization, data curation, formal analysis, investigation, methodology, software, validation, visualization, writing – original draft, writing – review and editing; D. J.: conceptualization, data curation, formal analysis, investigation, methodology, software, validation, visualization, writing – original draft, writing – review and editing; L. F. S.: conceptualization, investigation, methodology, supervision, writing – review and editing; G. G-G.: conceptualisation, validation, writing – original draft, writing – review and editing, supervision, project administration, funding acquisition.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI) and additional data is available through Zenodo with the following https://doi.org/10.5281/zenodo.17962518. Code could be made partly available upon reasonable request. Supplementary information is available. See DOI: https://doi.org/10.1039/d5gc06892d.Acknowledgements
This publication was created as part of NCCR Catalysis (grant number 225147), from the Swiss National Science Foundation. The authors would like to also acknowledge the financial support from the Swiss National Science Foundation (Project LEARN-D, grant number 214877). Part of the data used in this work was retrieved from Reaxys.com via API. Elsevier holds all rights to this data.References
- P. T. Anastas and J. C. Warner, Green Chemistry: Theory and Practice, Oxford University Press, 2000 Search PubMed.
- E. S. Beach, Z. Cui and P. T. Anastas, Energy Environ. Sci., 2009, 2, 1038 RSC.
- ACS Symposium Series, American Chemical Society, Washington, DC, 2000, pp. 1–6 Search PubMed.
- A. González-Garay, N. Mac Dowell and N. Shah, Discover Chem. Eng., 2021, 1, 2 CrossRef.
- R. A. Sheldon, ACS Sustainable Chem. Eng., 2018, 6, 32–48 CrossRef CAS.
- F. Roschangar, R. A. Sheldon and C. H. Senanayake, Green Chem., 2015, 17, 752–768 RSC.
- R. A. Sheldon, Green Chem., 2017, 19, 18–43 RSC.
- E. Lucas, A. J. Martín, S. Mitchell, A. Nabera, L. F. Santos, J. Pérez-Ramírez and G. Guillén-Gosálbez, Green Chem., 2024, 26, 9300–9309 RSC.
- S. Eichwald, H. Ostovari, H. Minten, J. Meyer-Waßewitz, D. Förtsch and N. Von Der Assen, Green Chem., 2025, 27, 10153–10168 RSC.
- ISO 14044, International Organization for Standardization, 2006.
- H. A. Van Kalkeren, A. L. Blom, F. P. J. T. Rutjes and M. A. J. Huijbregts, Green Chem., 2013, 15, 1255 RSC.
- D. Kralisch, D. Ott and D. Gericke, Green Chem., 2015, 17, 123–145 RSC.
- O. G. Griffiths, R. E. Owen, J. P. O'Byrne, D. Mattia, M. D. Jones and M. C. McManus, RSC Adv., 2013, 3, 12244 RSC.
- V. Tulus, J. Pérez-Ramírez and G. Guillén-Gosálbez. 10.1039/D1GC02623B.
- M. Pillich, J. Schilling, L. Bosetti and A. Bardow, Green Chem., 2024, 26, 10893–10906 RSC.
- I. Ioannou, J. Javaloyes-Antón, J. A. Caballero and G. Guillén-Gosálbez, ACS Sustainable Chem. Eng., 2023, 11, 1949–1961 CrossRef CAS PubMed.
- H. Minten, B. D. Vandegehuchte, B. Jaumard, R. Meys, C. Reinert and A. Bardow, Green Chem., 2024, 26, 8728–8743 RSC.
- D. Faust Akl, D. Poier, S. C. D'Angelo, T. P. Araújo, V. Tulus, O. V. Safonova, S. Mitchell, R. Marti, G. Guillén-Gosálbez and J. Pérez-Ramírez, Green Chem., 2022, 24, 6879–6888 RSC.
- G. Wernet, S. Papadokonstantakis, S. Hellweg and K. Hungerbühler, Green Chem., 2009, 11, 1826 RSC.
- R. K. Helling and D. A. Russell, Green Chem., 2009, 11, 380 RSC.
- S. Righi, A. Morfino, P. Galletti, C. Samorì, A. Tugnoli and C. Stramigioli, Green Chem., 2011, 13, 367–375 RSC.
- D. Ott, S. Borukhova and V. Hessel, Green Chem., 2016, 18, 1096–1116 RSC.
- D. Cespi, Green Chem., 2025, 27, 12107–12114 RSC.
- G. Wernet, C. Bauer, B. Steubing, J. Reinhard, E. Moreno-Ruiz and B. Weidema, Int. J. Life Cycle Assess., 2016, 21, 1218–1230 CrossRef.
- Chemical Abstract Services, CAS Registry, https://www.cas.org/cas-data/cas-registry, (accessed May 21, 2025).
- R. G. Hunt, T. K. Boguski, K. Weitz and A. Sharma, in The International Journal of Life Cycle Assessment, Springer Science and Business Media LLC, 1998, vol. 3 Search PubMed.
- Design for Innovative Value Towards a Sustainable Society, Springer Netherlands, Dordrecht, 2012, pp. 609–614 Search PubMed.
- R. Calvo-Serrano and G. Guillén-Gosálbez, ACS Sustainable Chem. Eng., 2018, 6, 7109–7118 CrossRef CAS.
- D. Zhang, Z. Wang, C. Oberschelp, E. Bradford and S. Hellweg, ACS Sustainable Chem. Eng., 2024, 12, 2700–2708 CrossRef CAS PubMed.
- A. Ghoroghi, Y. Rezgui, I. Petri and T. Beach, Int. J. Life Cycle Assess., 2022, 27, 433–456 CrossRef.
- B. Neupane, F. Belkadi, M. Formentini, E. Rozière, B. Hilloulin, S. F. Abdolmaleki and M. Mensah, Sustainable Prod. Consumption, 2025, 56, 37–53 CrossRef.
- P. Karka, S. Papadokonstantakis and A. Kokossis, in Computer Aided Chemical Engineering, Elsevier, 2019, vol. 46, pp. 97–102 Search PubMed.
- S. Sharafi, A. Kazemi and Z. Amiri, J. Cleaner Prod., 2023, 408, 137242 CrossRef CAS.
- X. Zhu, C.-H. Ho and X. Wang, ACS Sustainable Chem. Eng., 2020, 8, 11141–11151 CrossRef.
- R. Song, A. A. Keller and S. Suh, Environ. Sci. Technol., 2017, 51, 10777–10785 CrossRef CAS PubMed.
- J. Kleinekorte, J. Kleppich, L. Fleitmann, V. Beckert, L. Blodau and A. Bardow, ACS Sustainable Chem. Eng., 2023, 11, 9303–9319 CrossRef CAS.
- B. Zhao, C. Shuai, P. Hou, S. Qu and M. Xu, Environ. Sci. Technol., 2021, 55, 8439–8446 CrossRef CAS PubMed.
- R. Calvo-Serrano, M. González-Miquel, S. Papadokonstantakis and G. Guillén-Gosálbez, Comput. Chem. Eng., 2018, 108, 179–193 CrossRef CAS.
- A. König, K. Ulonska, A. Mitsos and J. Viell, Energy Fuels, 2019, 33, 1659–1672 CrossRef.
- P. Hou, J. Cai, S. Qu and M. Xu, Environ. Sci. Technol., 2018, 52, 5259–5267 CrossRef CAS PubMed.
- C. Mutel, J. Open Source Software, 2017, 2, 236 CrossRef.
- Reaction SMILES CRD 1.37M dataset, R. van der Lingen, 2025 Search PubMed.
- C. A. Jaksland, R. Gani and K. M. Lien, Chem. Eng. Sci., 1995, 50, 511–530 CrossRef CAS.
- IDEA Ver.3.3.3 Regionalized type JPN+GLO (2024 September 02), National Institute of Advanced Industrial Science and Technology (AIST), Research Institute of Science for Safety and Sustainability, Research Laboratory for IDEA Search PubMed.
- D. S. Young, Handbook of regression methods, Chapman & Hall, CRC Press, imprint of Taylor & Francis Group, Boca Raton, FL London New York, 2017 Search PubMed.
- P. Dumre, S. Bhattarai and H. K. Shashikala, in 2024 4th International Conference on Technological Advancements in Computational Sciences (ICTACS), IEEE, Tashkent, Uzbekistan, 2024, pp. 1856–1861 Search PubMed.
- Elsevier, Reaxys, 2025. https://www.reaxys.com/, accessed March 2025.
- V. Masson-Delmotte, P. Zhai, A. Pirani, S. L. Connors, C. Péan, S. Berger, N. Caud, Y. Chen, L. Goldfarb, M.I. Gomis, M. Huang, K. Leitzell, E. Lonnoy and J. B. R. Matthews, T. K. Maycock, T. Waterfield, O. Yelekçi, R. Yu, and B. Zhou (eds.), Climate change 2021: the physical science basis: summary for policymakers: working group I contribution to the sixth Assessment report of the Intergovernmental Panel on Climate Change, IPCC, Geneva, Switzerland, 2021 Search PubMed.
- M. A. J. Huijbregts, Z. J. N. Steinmann, P. M. F. Elshout, G. Stam, F. Verones, M. Vieira, M. Zijp, A. Hollander and R. Van Zelm, Int. J. Life Cycle Assess., 2017, 22, 138–147 CrossRef.
- Suggestions for updating the organisation environmental footprint (OEF) method, ed. L. Zampori and R. Pant, and European Commission, Publications Office, Luxembourg, 2019 Search PubMed.
Footnotes |
| † The authors M. G. H and D. J. contributed equally. |
| ‡ Copyright © 2022 Elsevier Limited except certain content provided by third parties. Reaxys is a trademark of Elsevier Limited. |
| This journal is © The Royal Society of Chemistry 2026 |



