 Open Access Article
Open Access ArticleStructure–function mapping of host defense peptide mimics reveals strategies for antibacterial activity and eradication of mature bacterial biofilms
Xueyang
Yin†
 a,
Yufang
Bi†
a,
Xiehe
Wang†
a,
Yufang
Bi†
a,
Xiehe
Wang†
 a,
Jing
Ren
b,
Jinrong
Yao
a,
Jing
Ren
b,
Jinrong
Yao
 *a,
Xin
Chen
*a,
Xin
Chen
 acd,
Shengjie
Ling
acd,
Shengjie
Ling
 *ac and
Zhengzhong
Shao
*ac and
Zhengzhong
Shao
 *ad
*ad
aState Key Laboratory of Molecular Engineering of Polymers, Research Center of AI for Polymer Science, Department of Macromolecular Science, Fudan University, Shanghai 200433, China. E-mail: yaoyaojr@fudan.edu.cn; lingshengjie@fudan.edu.cn; zzshao@fudan.edu.cn
bSchool of Physical Science and Technology, ShanghaiTech University, Shanghai 201210, China
cShanghai Stomatological Hospital & School of Stomatology, Fudan University, Shanghai 200433, China
dLaboratory of Advanced Materials, Fudan University, Shanghai 200433, China
First published on 9th March 2026
Abstract
Bacterial biofilms shield embedded cells behind extracellular polymeric substance (EPS) barriers, frustrate antibiotic penetration, and protect persisters, demanding materials that show broad antimicrobial activity coupled with efficient biofilm clearance. We establish a design framework for host defense peptide (HDP) mimics using poly(α-amino acid) copolymers with a cationic lysine backbone (degree of polymerization ≈20) and tunable hydrophobic modules—valine (Val), norvaline (Nva), phenylalanine (Phe), and N-ε-carbobenzoxy-lysine (CBL). Random ring-opening copolymerization of α-amino acid N-carboxyanhydride (NCA) (0–60% hydrophobic feed) yields amphiphilic libraries whose composition programs the secondary structure: Val biases β-sheet formation, high Nva favors α-helices, Phe promotes ordered states via π–π stacking, and CBL side-chain amides build stabilizing hydrogen-bond networks. Across MIC/MBC testing, S. aureus and P. aeruginosa exhibit distinct requirements: the cationic charge density is the primary determinant for penetrating EPS and clearing mature Gram-positive biofilms, whereas moderate hydrophobicity—notably Nva and CBL—optimizes activity against Gram-negative pathogens by balancing the solubility, membrane affinity, and EPS interactions. Self-assembly refines performance further; nanoscale aggregates with elevated surface potential (e.g., Lys60CBL40) deepen penetration and accelerate persister eradication. Critically, the antibacterial potency against planktonic cells does not predict the antibiofilm efficacy, revealing mechanistic decoupling that necessitates independent tuning. These results deliver a quantitative structure–function map and actionable rules for engineering amphiphilic polymers that recapitulate the functional hallmarks of HDPs, enabling simultaneous planktonic killing and robust clearance of mature biofilms and offering promising candidates for multidrug-resistant, biofilm-associated infections.
Green foundation1. This study establishes a sustainable polymerization strategy for host defense peptide mimics via propylene-oxide-assisted N-carboxyanhydride (NCA) chemistry, achieving green synthesis of biodegradable poly(α-amino acid) copolymers with tunable amphiphilicity and potent antibacterial/biofilm-clearing functions.2. The process eliminates inert gas protection and anhydrous conditions, replaces hazardous acid scavengers with benign propylene oxide, and markedly reduces solvent use, energy input, and chemical waste; efficacy: ≥80% S. aureus biofilm clearance at 125–175 μg mL−1, 50–80% P. aeruginosa biomass reduction at 200 μg mL−1, MIC 3.13 μg mL−1; nanoscale (+17.3 mV) aggregates deepen EPS penetration. 3. The method enables scalable, benign-by-design production of antimicrobial polymers derived from renewable amino acids. Future research will further establish it as a safer, low-waste alternative to antibiotic-based infection control while advancing sustainable healthcare and green manufacturing. |
1. Introduction
Bacterial biofilms represent a major challenge for antimicrobial therapy.1 Compared with planktonic cells, biofilm-embedded bacteria are shielded by an extracellular polymeric substance (EPS) matrix that impedes drug penetration and protects dormant persisters, rendering conventional antibiotics largely ineffective.2 Biofilms are common in chronic wounds, implants, and device-associated infections, where antibiotic overuse further accelerates resistance development.3 These challenges underscore the urgent need for new antimicrobial strategies with broad-spectrum activity, high penetration, and low resistance potential.Host defense peptide (HDP) mimics have emerged as promising materials for combating bacterial infections and biofilm-associated diseases.4,5 However, most current efforts focus on molecular activity5 rather than on the efficiency and sustainability of their synthesis. In recent years, the green chemistry community has emphasized quantifiable metrics—such as the E-factor,6 process mass intensity (PMI),7 energy consumption, and life-cycle considerations8—as essential criteria for evaluating chemical processes. These quantitative indicators provide a unifying framework for assessing the environmental improvement and resource efficiency. Integrating these concepts into the development of bioactive polymers enables a direct link between molecular design and sustainable manufacturing.9
In parallel, it has become increasingly important to evaluate the environmental impact of antimicrobial strategies themselves. Conventional antibiotics often persist in aquatic environments and contribute to the propagation of antimicrobial resistance genes,10 whereas host defense peptides and their synthetic analogues are typically biodegradable and exhibit minimal long-term persistence.11 From a green chemistry standpoint, HDP-based systems therefore offer the potential to reduce environmental loading and resistance risk while maintaining the therapeutic efficacy.12
In this context, the present study explores how structural variations in cationic copolypeptides influence both biological efficacy and process efficiency. By combining compositional tuning with analysis of material and energy use, we establish composition–structure–function relationships that are relevant not only to antibacterial performance but also to process greenness. This dual perspective bridges molecular functionality with sustainable synthesis, in line with the current direction of green chemistry research. Furthermore, by comparing the sustainability profile of peptide-based antimicrobials with conventional antibiotics on a functional basis (i.e., achieving ≥90% biofilm clearance), this study also highlights how biomimetic peptide design can contribute to the broader sustainability goals of green chemistry through lower persistence, reduced dosage, and improved life-cycle performance.
2. Materials and methods
2.1 Chemicals and reagents
All reagents were used as received without further purification unless otherwise stated. The main chemicals included N-ε-tert-butyloxycarbonyl-L-lysine (Boc-L-Lys, 97%), L-norvaline (L-Nva, 99%), tetrahydrofuran (THF, 99.9%, anhydrous), methyloxirane (≥99.5%), triphosgene (99%), anhydrous magnesium sulfate (AR), ethyl acetate (distilled grade), n-hexane (AR), L-valine N-carboxyanhydride (L-Val NCA, 98%), L-phenylalanine N-carboxyanhydride (L-Phe NCA, 98%), N-ε-carbobenzoxy-lysine NCA (L-CBL NCA, 98%), lithium bis(trimethylsilyl)amide (LiHMDS, 97%), and trifluoroacetic acid (TFA, ≥99.5%). All solvents were supplied by commercial vendors and used directly in synthesis and polymerization procedures. General handling and analytical procedures followed established protocols.132.2 Synthesis and characterization of NCA monomers
The successful synthesis of Boc-L-Lys NCA was confirmed by 1H and 13C NMR (Fig. S2 and S3). In the 1H NMR spectrum (400 MHz, CDCl3), the characteristic NCA ring proton appears at δ 6.88 (s, 1H); the α-CH of lysine at δ 4.33 (dd, J = 6.8, 4.7 Hz, 1H); a singlet at δ 3.14 (2H) corresponds to the ε-CH2; multiplets at δ 2.07–1.94 and 1.91–1.77 (each 1H) and 1.63–1.45 (4H) arise from the aliphatic methylenes; and the Boc tert-butyl group resonates at δ 1.45 (s, 9H). The 13C NMR spectrum (100 MHz, CDCl3) further corroborates the structure, showing carbonyl carbons at δ 169.9 (NCA) and 152.1 (carbamate), the Boc quaternary carbon near δ 77.3, the α-carbon at δ 57.5, and aliphatic/tert-butyl carbons at δ 29.2, 28.4, and 21.1. These assignments are consistent with previous reports for Boc-protected lysine NCAs.14
The successful synthesis of L-Nva NCA was confirmed by 1H NMR (Fig. S5). In the 1H NMR spectrum (400 MHz, CDCl3), the characteristic NCA ring proton appears at δ 6.88 (s, 1H); the α-CH resonates at δ 4.34 (dd, J = 6.9, 5.3 Hz, 1H); signals for the aliphatic methylenes are observed at δ 1.91 (ddd, J = 15.9, 9.7, 5.4 Hz, 1H), 1.86–1.71 (m, 1H), and 1.60–1.36 (m, 2H); and the terminal CH3 appears as a triplet at δ 0.99 (t, J = 7.3 Hz, 3H).
2.3 Synthesis of random copolymers
2.4 Evaluation of process efficiency and greenness
Quantitative indicators commonly adopted in green chemistry were employed to evaluate the process efficiency, including the E-factor (ratio of waste to product), PMI, and energy consumption per kilogram of product. PMI (process mass intensity) was defined as: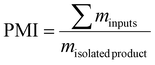 | (1) |
Unless otherwise noted, PMI refers to materials PMI (excludes solvents and water); the complete PMI (cPMI) including solvents/water is provided in the SI together with solvent-recovery assumptions.15–17 These metrics provide a direct measure of material and energy utilization efficiency during polymer synthesis.
For comparison, a conventional α-pinene/HCl-based NCA synthesis was analyzed as a representative benchmark. The present moisture-tolerant, catalyst-free route proceeds at ambient temperature within 3 h, in contrast to 60 °C for 12 h under nitrogen protection required by the traditional process. This modification leads to an approximately twofold reduction in energy demand and a decrease of E-factor and PMI values from 51.2 to 22.5 and from 11 to 5, respectively (Table 1). Such quantitative evaluation allows comparison of the synthesis efficiency on an environmental basis consistent with current green chemistry practices.18–23
| Parameter | Conventional α-pinene/HCl route | Moisture-tolerant epoxide-neutralized route (this work) | Improvement (%) |
|---|---|---|---|
| Conventions: PMI excludes solvents/water unless noted; cPMI (gross & net; 90% recovery for THF/EtOAc) is provided in the SI S1. | |||
| Reaction temperature/time | 60 °C × 12 h (under N2) | 25 °C × 3 h (ambient, sealed vial) | ≈80 ↓ thermal input |
| Catalyst/acid neutralizer | α-Pinene (10 eq.) + triphosgene | Propylene oxide (10 eq.) + triphosgene | — (eliminates hazardous acid gas) |
| Inert-gas requirement | Continuous N2/Ar purge | Not required (sealed system) | 100 ↓ |
| Yield (%) | ∼60% | 74% | +20 ↑ |
| Main by-products | HCl + chlorinated residues | 1-Chloro-2-propanol/2-chloro-1-propanol (acid-catalyzed ring-opening products of PO); removable during work-up | Non-corrosive/low toxicity |
| Energy consumption (kWh kg−1 product) | 10 | 5 | −50 ↓ |
| E-factor (waste/product) | 51.2 | 22.5 | −56 ↓ |
| Process mass intensity (PMI) | 11 | 5 | −55 ↓ |
| Overall assessment | High energy input and corrosive by-products | Low energy input, benign by-products, higher yield | — |
2.5 Antibacterial activity evaluation
The antibacterial activity of the cationic polypeptide polymers was evaluated by determining the minimum inhibitory concentration (MIC), minimum bactericidal concentration (MBC), and the killing efficiency against persister cells. MIC tests were conducted following a standard protocol described previously.24S. aureus (ATCC 6538) and P. aeruginosa (ATCC 9027) (obtained from the Shanghai Collection of Microorganisms) were inoculated into 10 mL LB medium and incubated at 37 °C with shaking for 12 h. The bacterial cultures were harvested by centrifugation, resuspended, and diluted with Mueller–Hinton (MH) broth to the desired concentration.In a sterile 96-well plate, serial two-fold dilutions of the polymer solution were prepared in MH medium, and 50 μL of each dilution was mixed with an equal volume of bacterial suspension (2 × 105 CFU per mL). Wells containing only bacterial suspension and only MH broth served as the control and blank, respectively. After static incubation at 37 °C for 9 h, the optical density at 600 nm (OD600) was measured using a multimode microplate reader (Cytation 3, BioTek). Bacterial viability was calculated using the following eqn (2).
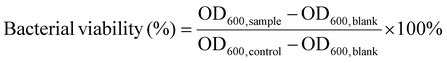 | (2) |
For MBC determination, 3.5 μL of the bacterial suspension from each MIC well was spotted onto LB agar plates and incubated at 37 °C for 12 h; the lowest polymer concentration yielding no visible colonies was recorded as the MBC. Persister cell killing assays were performed using S. aureus (ATCC 6538) grown to the logarithmic phase in LB medium. The cells were collected, resuspended in MH broth to 108 CFU per mL, and treated with ciprofloxacin (CIP, 10 × MIC) for 18 h under static conditions to induce persister formation. The persister cells were divided into two groups: one treated with CIP (10 × MIC) and the other with the poly(α-amino acid) solution (4 × MIC). At 6 h intervals, aliquots were withdrawn, plated on LB agar, and incubated overnight at 37 °C for colony counting. The number of surviving colonies was used to quantify the bactericidal efficiency of the polymers against antibiotic-tolerant persister cells.
2.6 Antibiofilm activity evaluation
The antibiofilm performance of the cationic polypeptide polymers was evaluated by assessing their ability to inhibit biofilm formation and disrupt the preformed biofilms of S. aureus (ATCC 6538) and P. aeruginosa (ATCC 9027). For inhibition assays, bacterial cells were harvested by centrifugation and resuspended in MH broth supplemented with 1% glucose to a final concentration of 2 × 106 CFU per mL. Polymer solutions were serially two-fold diluted in MH medium containing 1% glucose in tissue-culture-treated 96-well plates, with a final volume of 50 μL per well. Subsequently, 50 μL of the bacterial suspension was added to each well and mixed thoroughly, followed by static incubation at 37 °C for 24 h. Wells containing only MH broth (100 μL) served as negative controls, while those containing 50 μL MH and 50 μL bacterial suspension served as positive controls.For disruption assays, mature S. aureus biofilms were established by inoculating 100 μL of a 105 CFU per mL bacterial suspension (prepared in MH medium) into tissue-culture-treated 96-well plates and incubating statically at 37 °C for 24 h. The supernatant was then discarded, and the wells were gently rinsed with PBS to remove planktonic cells. Diluted polymer solutions (200 μL) were carefully added to each well containing preformed biofilms, followed by incubation at 37 °C for 24 h. The viability of biofilm-embedded bacteria was quantified using an MTT assay.
For P. aeruginosa biofilms, bacterial cells were harvested, resuspended in SOB medium, and adjusted to 105 CFU per mL. A 100 μL aliquot of this suspension was transferred into tissue-culture-treated 96-well plates and incubated statically at 37 °C for 36 h to allow biofilm maturation. The supernatant was removed, and the wells were washed once with PBS to remove planktonic bacteria. Polymer solutions were serially diluted in MH medium from an initial concentration of 200 μg mL−1, and 200 μL of each dilution was added to the biofilm-containing wells. After static incubation at 37 °C for 9 h, MTT staining was performed to determine bacterial metabolic activity.
For MTT staining, the treated biofilms were rinsed gently with PBS, and 100 μL of MTT working solution (0.5 mg mL−1 in PBS) was added to each well. The plates were incubated at 37 °C for 4 h in the dark. After removing the supernatant, 100 μL of DMSO was added to dissolve the formazan crystals, and the absorbance at 570 nm was measured using a multimode microplate reader. Bacterial viability was calculated using eqn (3):
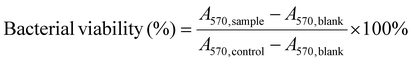 | (3) |
Confocal laser scanning microscopy (CLSM, Nikon C2+) was employed to visualize the biofilm morphology and cell viability after polymer treatment. Prior to imaging, biofilms were stained with 25 μL SYTO 9 (3 μM) and 25 μL propidium iodide (PI, 15 μM) for 5 min. SYTO 9 was excited at ∼500 nm with emission at ∼530 nm and PI was excited at ∼493 nm with emission at ∼636 nm.
For surface morphology observation, biofilms on cell-culture coverslips were treated with the respective polymer solutions and examined using field-emission scanning electron microscopy (SEM, Gemini SEM 500, Zeiss). Following treatment, the biofilms were rinsed gently with PBS and fixed overnight at 4 °C with 4% glutaraldehyde. After fixation, samples were dehydrated through a graded ethanol series (30, 50, 70, 80, 90, 95, and 100% v/v), dried, and subjected to SEM analysis to evaluate the extent of biofilm disruption.
2.7 Cytotoxicity
NIH/3T3 cells were evaluated by CCK-8 viability analysis after 24 h exposure to polymers at 50, 100, 125, 150, 175, and 200 µg mL−1; cells were seeded at 8 × 103 cells per well in DMEM + 10% FBS and the absorbance was read at 450 nm. The full protocol is provided in the SI.2.8 Characterization
Molecular weights were determined by gel permeation chromatography (GPC) in DMF containing 0.01 M LiBr at 50 °C (flow rate 1.0 mL min−1) using PolarGel-L (and, when required, PolarGel-M) columns. 1H NMR spectra of polymers were acquired in D2O on a Bruker AVANCE III HD spectrometer to verify composition and deprotection. Circular dichroism (CD) spectra were recorded on a Chirascan spectropolarimeter (Applied Photophysics) at 25 °C using polymer solutions (200 µg mL−1 in PBS, pH 7.4) in 1 mm quartz cuvettes over 195–280 nm (1 nm interval, 50 nm min−1 scan rate, 2 nm bandwidth); the mean residue ellipticity was calculated as [θ] (deg cm2 dmol−1) = mdeg/(l × c), where mdeg is the CD signal, l is the path length (mm), and c is the molar concentration (mM). Critical aggregation concentrations (CACs) were determined with 8-anilino-1-naphthalenesulfonic acid (ANS, 10 µM) as a fluorescent probe by preparing polymers at 0.03–400 µg mL−1 in PBS (pH = 7.4) or PB (pH = 5.5), incubating for 1 h at room temperature in the dark, and recording emission spectra from 420–670 nm (λex = 360 nm); CAC values were obtained from the intersection of two linear fits of fluorescence intensity versus log[polymer].3. Results and discussion
3.1 Design strategy for constructing host defense peptide mimics
Although HDP mimics display broad structural diversity, they generally share two hallmarks: cationic charge, which promotes electrostatic adsorption to anionic bacterial membranes, and amphiphilicity, which enables insertion into lipid bilayers and disruption of membrane integrity.25 Guided by these features, we established a polymer library with varied cationic and hydrophobic modules to explore structure–function relationships.26Poly(α-amino acids) were selected as the cationic framework due to their resemblance to natural HDPs and tunable side chains. Primary amines were introduced as charge carriers, as they facilitate initial membrane binding and have been reported to surpass tertiary and quaternary amines in membrane perturbation.27
Hydrophobicity was tuned by four representative monomers. Valine offered moderate hydrophobicity to balance solubility and insertion; norvaline increased side-chain flexibility and thus backbone conformational freedom; phenylalanine, with an aromatic ring, enhanced lipophilicity and rigidity, favoring stable membrane insertion; and CBL introduced amide groups, enabling hydrogen bonding with membrane phosphates and enhancing activity.28
Finally, the chain length was set at ∼20 residues, informed by previous reports that polymers shorter than this lack insertion capability, whereas longer chains suffer from reduced solubility, altered charge density, and diminished activity.29,30 This design ensured a balance among conformational adaptability, solubility, and antibacterial performance, providing a robust platform for subsequent studies.
The process operates under mild conditions and affords Boc-L-Lys NCA as white crystals (yield = 74%). Relative to the α-pinene/HCl route, the reaction temperature and duration are markedly reduced, resulting in roughly half the energy consumption and approximately 40% lower overall waste generation. These outcomes illustrate that modifications at the process level can directly enhance the material and energy efficiency while maintaining the product yield. Our comparisons cover conventional chlorinated protocols and moisture-tolerant scavenger strategies reported in the literature,25,26 with side-by-side conditions summarised in Table 1.
3.2 Synthesis of HDP mimics via NCA ring-opening polymerization
To systematically probe how the hydrophobic content affects antibacterial and antibiofilm performance, we synthesized a series of copolymers with the general formula P(Lysx%-co-Yy%)n (n = 20), where Y represents Val, Nva, Phe, or CBL. For clarity, they are denoted as LysxValy, LysxNvay, LysxPhey, and LysxCBLy, with x and y indicating the molar ratios of Lys and the hydrophobic unit.Cationic Boc-L-Lys NCA and four hydrophobic NCAs (L-Val, L-Nva, L-Phe, L-CBL) were synthesized and copolymerized through random NCA ring-opening, yielding amphiphilic polymers containing both cationic (Lys) and hydrophobic segments. To avoid solubility loss from excessive hydrophobicity, feed ratios were limited to 0–60%, producing representative libraries for each class (Fig. 1).31
1H NMR confirmed that incorporated monomer ratios closely matched feed compositions, validating reproducibility, although Val-rich copolymers (≥50%) showed slightly lower incorporation due to reduced solubility in THF during polymerization. Gel permeation chromatography (GPC) revealed number-average molecular weights of 3–5 kDa (≈20 residues) and dispersities of 1.3–1.7. LysxPhey and LysxCBLy showed narrower distributions (Đ ≈ 1.3), while Val- and Nva-rich copolymers were broader (Đ ≈ 1.4–1.7), likely reflecting β-sheet tendencies of Val or higher chain flexibility of Nva. Val incorporation >40% also led to solubility loss, limiting accurate GPC analysis, whereas Nva-containing copolymers retained partial solubility (Table 2). From a process standpoint, maintaining solution polymerisation under ambient THF conditions defines an operational window that minimises solvent load and enables solvent recovery, consistent with Green Chemistry practice.33
| Series | GPC | NMR | S. aureus 6538 | P. aeruginosa 9027 | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| M n | Đ | DP | (x%![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) y%) y%) |
MIC | MBC | MIC | MBC | SMIC80 | ||
| LysxValy | Lys90Val10 | 5000 | 1.50 | 23 | 91![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 9 9 |
6.25 | 6.25 | 25 | 50 | 100 |
| Lys80Val20 | 4200 | 1.73 | 21 | 83![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 17 17 |
6.25 | 6.25 | 25 | 50 | 100 | |
| Lys70Val30 | 4200 | 1.75 | 22 | 73![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 27 27 |
6.25 | 12.5 | 25 | 50 | 50 | |
| Lys60Val40 | 4700 | 1.74 | 27 | 64![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 36 36 |
6.25 | 12.5 | 12.5 | 25 | 50 | |
| Lys50Val50 | — | — | — | 56![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 44 44 |
12.5 | 200 | 25 | 100 | 100 | |
| Lys40Val60 | — | — | — | 48![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 52 52 |
25 | 50 | 25 | 200 | 200 | |
| LysxNvay | Lys90Nva10 | 5500 | 1.41 | 25 | 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 10 |
6.25 | 6.25 | 50 | 50 | 100 |
| Lys80Nva20 | 5000 | 1.41 | 25 | 81![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 19 19 |
6.25 | 6.25 | 25 | 50 | 50 | |
| Lys70Nva30 | 4100 | 1.55 | 22 | 70![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30 |
6.25 | 12.5 | 12.5 | 25 | 50 | |
| Lys60Nva40 | 3700 | 1.60 | 21 | 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40 40 |
12.5 | 25 | 3.13 | 25 | 25 | |
| Lys50Nva50 | 3000 | 1.73 | 18 | 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 50 |
25 | 100 | 3.13 | 12.5 | 25 | |
| Lys40Nva60 | 2800 | 1.76 | 19 | 43![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 57 57 |
25 | 50 | 6.25 | 25 | 50 | |
| LysxPhey | Lys90Phe10 | 4900 | 1.32 | 22 | 90![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 10 |
6.25 | 6.25 | 50 | 100 | 100 |
| Lys80Phe20 | 4900 | 1.30 | 23 | 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20 |
6.25 | 6.25 | 25 | 50 | 100 | |
| Lys70Phe30 | 4800 | 1.32 | 24 | 70![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 30 |
6.25 | 6.25 | 12.5 | 50 | 50 | |
| Lys60Phe40 | 4500 | 1.32 | 23 | 62![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 38 38 |
6.25 | 12.5 | 6.25 | 12.5 | 50 | |
| Lys50Phe50 | 4300 | 1.31 | 23 | 52![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 48 48 |
12.5 | 12.5 | 6.25 | 25 | 50 | |
| Lys40Phe60 | 4400 | 1.33 | 25 | 38![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 62 62 |
25 | 50 | 12.5 | 50 | 100 | |
| LysxCBLy | Lys90CBL10 | 5100 | 1.29 | 22 | 92![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 8 8 |
6.25 | 6.25 | 50 | 100 | 100 |
| Lys80CBL20 | 5100 | 1.30 | 21 | 80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 20 |
6.25 | 6.25 | 25 | 25 | 50 | |
| Lys70CBL30 | 5100 | 1.31 | 21 | 68![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 32 32 |
6.25 | 12.5 | 12.5 | 12.5 | 25 | |
| Lys60CBL40 | 5200 | 1.27 | 21 | 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40 40 |
12.5 | 12.5 | 12.5 | 12.5 | 50 | |
| Lys50CBL50 | 5300 | 1.27 | 21 | 51![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 49 49 |
12.5 | 25 | 12.5 | 25 | 50 | |
| Lys40CBL60 | 5500 | 1.28 | 21 | 48![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 52 52 |
25 | 50 | 25 | 50 | 100 | |
Because NCA copolymerization is statistical, short-range sequences are heterogeneous, with alternating, blocky, or mixed motifs. Importantly, this heterogeneity did not compromise the antibacterial or antibiofilm performance; rather, it recapitulated the structural diversity of natural HDPs and enriched the library for subsequent structure–activity studies.
3.3 Conformational analysis of HDP mimics as a function of composition
The secondary structures of HDP mimics in aqueous environments were investigated using far-UV circular dichroism (CD) spectroscopy in PBS, which better simulates physiological conditions. Spectra between 190–250 nm capture amide transitions and reveal conformational states: random coils show a positive band at ∼216 nm and a negative band near 197 nm; α-helices exhibit minima at 208 and 222 nm and β-sheets display a negative band near 216 nm and a positive signal around 195 nm.In the LysxValy series (Fig. 2a), 10–40% Val yielded spectra characteristic of random coils, consistent with electrostatic repulsion among Lys residues. At 50% Val, mixed α-helix/β-sheet features emerged, while 60% Val favored β-sheet structures with a strong 217 nm minimum. This progression agrees with the known β-sheet bias of Val, driven by steric restrictions from isopropyl side chains.34–36 LysxNvay (Fig. 2b) displayed similar trends at 10–50% Nva, but at 60% formed stable α-helices, indicated by [θ]222/[θ]208 ratio >1, suggesting interhelical stabilization.37
 | ||
| Fig. 2 CD spectra of (a) LysxValy, (b) LysxNvay, (c) LysxPhey and (d) LysxCBLy in PBS buffer (pH 7.4) at 25 °C. | ||
For LysxPhey (Fig. 2c), increasing the Phe content weakened the 197 nm negative band, which became positive at 60%, reflecting π–π stacking that promotes ordered conformations. LysxCBLy polymers (Fig. 2d) also transitioned toward α-helical or β-sheet structures with rising CBL content, with mixed structures already evident at 40%. This effect likely stems from amide-mediated hydrogen-bonding networks, in line with reports that self-assembling antimicrobial peptides stabilize secondary structures through hydrogen bonding and aromatic interactions.38
At 10–40% hydrophobic content, all four series adopted random coil conformations, again attributable to Lys–Lys electrostatic repulsion. Raising the pH, which partially neutralizes cationic groups and mimics anionic bacterial surfaces, induced helix formation in Lys90CBL10. This environmental responsiveness demonstrates that conformational adaptability is an intrinsic feature of HDP mimics and may underlie their capacity to tune antibacterial activity.
3.4 Structure–function relationships governing the antibacterial activity of HDP mimics against planktonic bacteria
We evaluated the minimum inhibitory concentrations (MICs) and minimum bactericidal concentrations (MBCs) of the polymers against S. aureus and P. aeruginosa (Table 2). Against S. aureus, all four series maintained strong activity at 10–40% hydrophobic content (MIC <12.5 μg mL−1, MBC <25 μg mL−1). LysxValy and LysxPhey were the most potent (MIC = 6.25 μg mL−1), comparable to natural peptides such as lactoferricin B and superior to many reported AMPs.39–43At ≥50% hydrophobicity, trends diverged. LysxValy and LysxNvay showed reduced bactericidal activity, with MBCs rising to 100–200 μg mL−1, likely due to lowered charge density and polymer self-association. In contrast, LysxPhey and LysxCBLy retained robust activity, as aromatic stacking or amide hydrogen bonding stabilized interactions when the cationic density decreased.
Against P. aeruginosa, LysxValy displayed a bell-shaped activity curve, peaking at intermediate Val levels, whereas LysxNvay maintained low MICs across a broader range (e.g., 3.13 μg mL−1 at Lys50Nva50). LysxPhey and LysxCBLy were the most effective at 30–50% hydrophobic content. Notably, large CBL-rich aggregates retained activity comparable to single-chain analogs, indicating that the balance of cationic and hydrophobic groups, rather than the assembly state, governs efficacy.
CD analysis linked these outcomes to the secondary structure. Ordered conformations at high Val or Nva content correlated with diminished bactericidal efficiency, as functional groups became sequestered within folded or aggregated states. This is consistent with previous reports where enforced helicity reduced AMP activity.40,44,45
Comparisons across species revealed that LysxValy and LysxPhey were more effective against Gram-positive bacteria, while LysxNvay and LysxCBLy favored Gram-negative pathogens. This dichotomy reflects cell envelope differences: Gram-positive bacteria rely primarily on electrostatic interactions with teichoic acids (Fig. S10a), whereas Gram-negative species require additional hydrophobicity to penetrate their lipopolysaccharide-rich outer membrane (Fig. S10b). Despite the random sequence distribution of these copolymers, the overall amphiphilicity proved to be sufficient to reproduce the key functional features of natural HDPs.46
In parallel with MICs, NIH/3T3 viability assays show that cytotoxicity varies with the hydrophobic pendant group at equal charge density, consistent with the amphiphilic-balance window observed for antibacterial/antibiofilm activity. This structure dependence motivates prioritizing pendant-group chemistry when modeling and synthesizing next-generation mimics (Fig. S11).28
3.5 Non-bactericidal mechanisms and structural determinants underlying biofilm inhibition
In addition to bactericidal activity, HDP mimics inhibited biofilm formation through non-lethal pathways, including interference with bacterial adhesion, quorum sensing, and EPS secretion. Under biofilm-promoting conditions, all four series suppressed P. aeruginosa biofilms in a manner consistent with their antibacterial potency, with LysxNvay showing the strongest inhibition up to 50% Nva before slightly declining (Table 2).For S. aureus, however, inhibition patterns diverged from antibacterial trends. LysxValy copolymers effectively killed planktonic cells at 10–40% Val but required ∼4 × MIC to block biofilm formation, underscoring the tolerance conferred by rapid adhesion and EPS deposition. Notably, Lys50Val50, despite weak bactericidal activity (MBC 200 μg mL−1), suppressed >80% of biofilm biomass at 50 μg mL−1, while Lys40Val60, though more bactericidal, showed poor antibiofilm performance (Fig. 3a). These discrepancies highlight mechanistic distinctions between cell killing and biofilm inhibition, consistent with natural peptides such as LL-37, which blocks biofilm development at sub-MIC levels through adhesion and signaling interference.47
Mechanistic insights suggest that cationic polymers can act at sub-lethal doses by charge neutralization or “bridging effects” that cluster bacteria and reduce surface colonization,48,49 or by disrupting metabolism and quorum sensing.50 For example, Lys50Val50 assemblies likely adhere to bacterial surfaces, altering hydrophobicity and preventing adhesion.51
LysxNvay polymers also displayed strong antibiofilm effects, with Lys50Nva50 fully suppressing S. aureus biofilms at 50 μg mL−1, below its MBC (Fig. 3b). Similarly, LysxPhey required higher multiples of MBC at low Phe content but became more efficient at 40–50%, consistent with π–π interactions stabilizing ordered conformations (Fig. 3c). LysxCBLy showed potent inhibition at 10–40% CBL but diminished efficacy at higher loadings, likely due to the formation of large aggregates that limited cell-surface interactions (Fig. 3d).
Altogether, these results show that antibiofilm activity is not solely dictated by bactericidal potency. Instead, it arises from synergistic contributions of cationic density, moderate hydrophobicity, and specific non-cytotoxic mechanisms, highlighting design opportunities for polymers that target biofilm formation independently of bacterial killing.
3.6 Structural regulation and functional performance in clearing mature biofilms
We next assessed the ability of selected HDP mimics to eradicate established biofilms. Against S. aureus, polymers with 10–40% hydrophobic content were the most effective. CLSM imaging with SYTO 9/PI staining revealed that Lys90Val10 and Lys80Val20 induced extensive bacterial death throughout biofilms, whereas Lys70Val30 and Lys60Val40 showed residual viability in deeper layers. SEM confirmed pronounced structural damage, including ruptured membranes and collapsed surfaces (Fig. 4a and b). Quantitative assays showed that Lys90Val10 cleared 80% of biofilms at 150 μg mL−1, while Lys80Val20 required 175 μg mL−1; Lys70Val30 removed only ∼60% at 200 μg mL−1, and Lys40Val60 was largely inactive (Fig. 4c). | ||
| Fig. 4 (a) Three-dimensional confocal laser scanning microscopy (3D CLSM) images of mature S. aureus biofilms after 24 h treatment with 200 μg mL−1 LysxValy. (b) SEM images of biofilms treated with LysxValy. (c) Viability of biofilms incubated with LysxValy†, LysxNvay, LysxPhey, or LysxCBLy (*p < 0.05, **p < 0.01, *p < 0.001 vs. control). † Data adapted from ref. 52, with permission from the Royal Society of Chemistry. Inset: MTT-stained biofilm after polymer treatment. | ||
In the LysxNvay series, the clearance capacity declined sharply as the cationic content decreased. Lys90Nva10 achieved ∼80% removal at 150 μg mL−1, but polymers with >20% Nva were ineffective. LysxPhey followed a similar trend: Lys90Phe10 and Lys80Phe20 reduced ∼80% at 150–175 μg mL−1, whereas higher Phe contents lost efficacy. In contrast, LysxCBLy polymers maintained strong performance, with 10–20% CBL clearing >80% of biomass at 125–175 μg mL−1, while Lys70CBL30 retained ∼60% clearance at 200 μg mL−1 (Fig. 4c). Collectively, these results underscore that the cationic density dominates EPS penetration and biofilm clearance.
To disentangle charge effects from hydrophobic chemistry, we compared Lys90CBL10, Lys90BA10, Lys90Leu10, and Lys100. Despite the identical charge ratios, Lys90CBL10 achieved near-complete clearance (95%) at 200 μg mL−1, outperforming Lys90BA10 (70%) (Fig. 5a). The enhanced activity of CBL arises from amide-mediated hydrogen bonding with EPS and membranes.16,53 Persister assays confirmed that the amphiphilic balance accelerates clearance: Lys90CBL10, Lys90Val10, and Lys90BA10 eradicated persisters within 18 h, while Lys100 required 24 h (Fig. 5b). Consistent with these trends, the biofilm viability decreased in a dose-dependent manner, with LysxCBLy and LysxValy showing the steepest reductions, and LysxPhey intermediate and LysxNvay the least (Fig. 5c). Thus, additional cationic charge alone is insufficient; moderate hydrophobicity enhances penetration and killing depth.
For P. aeruginosa, polymers with 20–50% hydrophobicity were the most effective. LysxNvay and LysxCBLy disrupted the biofilm structure, increased red fluorescence (Fig. 6a and b), and reduced biomass by 50–80% at 200 μg mL−1 (Fig. 6c–h).
In contrast, LysxValy and LysxPhey showed limited activity except at specific compositions (Fig. 7a–c). Notably, Lys60CBL40 was effective against both S. aureus and P. aeruginosa, likely due to its higher surface potential (+17.3 mV) compared with Lys50Nva50 (+6.9 mV), enabling stronger EPS penetration and adsorption.
Integrating these findings, we constructed a structure–function map (Fig. 7d). High cationic content was decisive for Gram-positive activity, while moderate hydrophobicity enhanced the efficacy against Gram-negative biofilms. Polymers relying solely on charge lacked broad-spectrum antibiofilm potency. Rational amphiphilic designs, balancing charge, hydrophobicity, and assembly properties, therefore represent the optimal strategy for developing HDP mimics capable of eradicating mature biofilms.
3.7 Discussion
With antibiotic resistance intensifying and conventional therapies failing against biofilm-associated infections, this study provides a systematic framework linking the polymer structure to antibacterial and antibiofilm functions. By comparing poly(α-amino acid) copolymers with varied hydrophilic–hydrophobic compositions, we identified how the cationic density, hydrophobicity, and assembly behaviour cooperatively shape biological outcomes.Our results demonstrate that the cationic charge density is the primary driver of EPS penetration and biofilm clearance. Polymers with high Lys content, such as Lys90Val10 and Lys90CBL10, readily permeated and disrupted mature S. aureus biofilms, while reducing charge below ∼60% led to a sharp loss of efficacy. This underscores the central role of electrostatics in overcoming the biofilm barrier.
Hydrophobicity further modulated activity, especially against P. aeruginosa. Polymers incorporating Nva or CBL performed best at intermediate hydrophobic ratios (20–40%), where solubility, membrane affinity, and EPS interactions were optimally balanced. CBL's amide side chains provided additional hydrogen bonding, conferring advantages over Val, Phe, or ester-containing BA.
Self-assembly also proved critical. Lys60CBL40 formed nanoscale aggregates with higher surface potential (+17.3 mV), enabling deep EPS penetration and persister eradication, whereas Lys50Nva50, despite similar composition, lacked comparable activity due to its lower charge density and weaker assemblies. Persister assays confirmed that amphiphilicity, not charge density alone, governs clearance kinetics: Lys90CBL10, Lys90Val10, and Lys90BA10 eradicated persisters faster than fully cationic Lys100.
Importantly, antibacterial potency against planktonic bacteria and antibiofilm efficacy were not linearly correlated. While electrostatic adsorption dominates activity against Gram-positive bacteria, moderate hydrophobicity is essential for breaching Gram-negative outer membranes and their biofilms. This distinction highlights that separate mechanisms underlie these two activities, requiring differentiated design strategies.
In addition to improvements in process greenness,54 the comparative sustainability of peptide-based antimicrobials relative to conventional antibiotics merits attention. Antibiotic production and environmental fate analyses have shown that active pharmaceutical ingredient synthesis accounts for nearly 40–45% of the total life-cycle environmental impact,55 and persistent residues contribute to antimicrobial resistance dissemination. In contrast, antimicrobial peptides and their mimics are enzymatically degradable and exhibit effective antibiofilm activity at 10–100 μg mL−1.56,57 When compared on a functional unit basis (≥90% mature biofilm clearance), the lower mass and energy demands per antibacterial effect indicate that HDP-based systems inherently align with green chemistry principles by reducing the persistence, resistance potential, and material intensity.32
Overall, our study clarifies the cooperative roles of cationic density, moderate hydrophobicity, and self-assembly in defining the dual functionality of HDP mimics (Fig. 8). These insights provide design principles for amphiphilic polymers capable of both killing planktonic bacteria and clearing mature biofilms, offering a rational path toward next-generation biomimetic therapeutics for multidrug-resistant and chronic biofilm-associated infections.
4. Conclusion
This study elucidates composition–structure–function relationships in cationic copolypeptides for biofilm eradication and demonstrates that the underlying synthetic route achieves high efficiency with reduced material and energy demands. The moisture-tolerant, catalyst-free NCA process yields 74% conversion—an improvement over conventional peptide-synthesis routes—while lowering the E-factor and PMI by about 50% and halving the energy requirement. These quantitative reductions provide clear evidence of an environmentally improved process consistent with the current principles of green chemistry.Beyond process optimization, host defense peptide mimics represent an environmentally responsible antimicrobial platform. Compared with small-molecule antibiotics—which often show persistence in water environments and contribute to antimicrobial resistance—biodegradable peptides offer comparable efficacy with lower functional-unit mass and minimal long-term environmental footprint. This work therefore underscores how the rational design and sustainable synthesis of HDP-based materials can jointly advance the dual objectives of biomedical performance and green chemistry sustainability.
Conflicts of interest
The authors declare no conflict of interest.Data availability
The data supporting the findings of this study are available within the article and its supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5gc05877e.Acknowledgements
This work was supported by the National Key R&D Program of China (2025YFA0923503), the National Natural Science Foundation of China (52322305 and 52473098), and the starting grant of Fudan University.References
- J. W. Costerton, P. S. Stewart and E. P. Greenberg, Science, 1999, 284, 1318–1322 CrossRef CAS PubMed
.
- H.-C. Flemming and J. Wingender, Nat. Rev. Microbiol., 2010, 8, 623–633 CrossRef CAS PubMed
.
- C. R. Arciola, D. Campoccia and L. Montanaro, Nat. Rev. Microbiol., 2018, 16, 397–409 CrossRef CAS PubMed
.
- W. Zhang, S. Deng, M. Zhou, J. Zou, J. Xie, X. Xiao, L. Yuan, Z. Ji, S. Chen, R. Cui, Z. Luo, G. Xia and R. Liu, Biomater. Sci., 2022, 10, 4515–4524 RSC
.
- Z. Si, W. Zheng, D. Prananty, J. Li, C. H. Koh, E.-T. Kang, K. Pethe and M. B. Chan-Park, Chem. Sci., 2022, 13, 345–364 RSC
.
- R. A. Sheldon, Green Chem., 2017, 19, 18–43 RSC
.
- J. Li, E. M. Simmons and M. D. Eastgate, Green Chem., 2017, 19, 127–139 RSC
.
- E. Lucas, A. J. Martín, S. Mitchell, A. Nabera, L. F. Santos, J. Pérez-Ramírez and G. Guillén-Gosálbez, Green Chem., 2024, 26, 9300–9309 RSC
.
- M. Di Consiglio, E. Sturabotti, B. Brugnoli, A. Piozzi, L. M. Migneco and I. Francolini, Polym. Chem., 2023, 14, 432–442 RSC
.
- D. G. J. Larsson and C.-F. Flach, Nat. Rev. Microbiol., 2022, 20, 257–269 CrossRef CAS PubMed
.
- C. A. Davis and E. M.-L. Janssen, Environ. Int., 2020, 134, 105271 CrossRef CAS PubMed
.
- P. Ma, Y. Wu, W. Jiang, N. Shao, M. Zhou, Y. Chen, J. Xie, Z. Qiao and R. Liu, Biomater. Sci., 2022, 10, 4193–4207 RSC
.
- Y.-M. Wu, K. Chen, J. Wang, W. Dai, H. Yu, X. Xie, M. Chen and R.-H. Liu, Nat. Protoc., 2025, 20, 709–726 CrossRef CAS PubMed
.
- Z.-Y. Tian, Z. Zhang, S. Wang and H. Lu, Nat. Commun., 2021, 12, 5810 CrossRef CAS PubMed
.
- D. Reinhardt, F. Ilgen, D. Kralisch, B. König and G. Kreisel, Green Chem., 2008, 10, 1170–1181 RSC
.
- H. C. Erythropel,
et al.
, Green Chem., 2018, 20, 1929–1961 RSC
.
- F. Roschangar, R. A. Sheldon and C. H. Senanayake, Green Chem., 2017, 19, 281–285 RSC
.
- J. L. Osorio-Tejada, F. Ferlin, L. Vaccaro and V. Hessel, Green Chem., 2023, 25, 9760–9778 RSC
.
- T. Yamaki,
et al.
, Green Chem., 2024, 26, 3758–3766 RSC
.
- A. Bardow, J. Pérez-Ramírez, S. Sala and L. Vaccaro, Green Chem., 2024, 26, 11016–11018 RSC
.
- V. G. Zuin, I. Eilks, M. Elschami and K. Kümmerer, Green Chem., 2021, 23, 1594–1608 RSC
.
- P. M. Nowak, Green Chem., 2023, 25, 4625–4640 RSC
.
- C. Jiménez-González, C. S. Ponder, Q. B. Broxterman and J. B. Manley, Org. Process Res. Dev., 2011, 15, 912–917 CrossRef
.
- D. Zhang, Y. Qian, S. Zhang, P. Ma, Q. Zhang, N. Shao, F. Qi, J. Xie, C. Dai, R. Zhou, Z. Qiao, W. Zhang, S. Chen and R. Liu, Sci. China Mater., 2019, 62, 604–610 CrossRef
.
- H. R. Kricheldorf, Angew. Chem., Int. Ed., 2006, 45, 5752–5784 CrossRef CAS PubMed
.
- C. A. Stewart and C. A. VanderWerf, J. Am. Chem. Soc., 1954, 76, 1259–1264 CrossRef CAS
.
- E. F. Palermo, D.-K. Lee, A. Ramamoorthy and K. Kuroda, J. Phys. Chem. B, 2011, 115, 366–375 CrossRef CAS PubMed
.
- D. S. S. M. Uppu, M. M. Konai, U. Baul, P. Singh, T. K. Siersma, S. Samaddar, S. Vemparala, L. W. Hamoen, C. Narayana and J. Haldar, Chem. Sci., 2016, 7, 4613–4623 RSC
.
- G. N. Tew, R. W. Scott, M. L. Klein and W. F. DeGrado, Acc. Chem. Res., 2010, 43, 30–39 CrossRef CAS PubMed
.
- H. Etayash and R. E. W. Hancock, Pharmaceutics, 2021, 13, 1820 CrossRef CAS PubMed
.
- G. J. M. Habraken, K. H. R. M. Wilsens, C. E. Koning and A. Heise, Polym. Chem., 2011, 2, 1322–1330 RSC
.
- H.-K. Lin, C.-H. Dai, T. F. Jamison and K. F. Jensen, Angew. Chem., Int. Ed., 2017, 56, 8870–8873 CrossRef CAS PubMed
.
- V. Tulus, J. Pérez-Ramírez and G. Guillén-Gosálbez, Green Chem., 2021, 23, 9881–9893 RSC
.
- T. Yang, T. Xue, J. Mao, Y. Chen, H. Tian, A. Bartolome, H. Xia, X. Yao, C. V. Kumar, J. Cheng and Y. Lin, J. Am. Chem. Soc., 2024, 146, 5823–5833 CrossRef CAS PubMed
.
- N. J.-A. Chan, D. Gu, S. Tan, Q. Fu, T. G. Pattison, A. J. O'Connor and G. G. Qiao, Nat. Commun., 2020, 11, 1630 CrossRef PubMed
.
- S. Yamashita and T. Yamashita, Proc. Natl. Acad. Sci. U. S. A., 1975, 72, 941–942 CrossRef CAS PubMed
.
- S. Y. Lau, A. K. Taneja and R. S. Hodges, J. Biol. Chem., 1984, 259, 13253–13261 CrossRef CAS PubMed
.
- H. Liu, Z. Song, Y. Zhang, B. Wu, D. Chen, Z. Zhou, H. Zhang, S. Li, X. Feng, J. Huang and H. Wang, Nat. Mater., 2025, 24, 1295–1306 CrossRef CAS PubMed
.
- J. Turner, Y. Cho, N.-N. Dinh, A. J. Waring and R. I. Lehrer, Antimicrob. Agents Chemother., 1998, 42, 2206–2214 CrossRef CAS PubMed
.
- J. Harder, J. Bartels, E. Christophers and J.-M. Schröder, J. Biol. Chem., 2001, 276, 5707–5713 CrossRef CAS PubMed
.
- H. Raghuraman and A. Chattopadhyay, Biosci. Rep., 2007, 27, 189–223 CrossRef CAS PubMed
.
- Ø. Samuelsen, H. H. Haukland, H. Jenssen, M. Krämer, K. Sandvik, H. Ulvatne and L. H. Vorland, FEBS Lett., 2005, 579, 3421–3426 CrossRef PubMed
.
- Y. Ge, D. L. MacDonald, K. J. Holroyd, C. Thornsberry, H. Wexler and M. Zasloff, Antimicrob. Agents Chemother., 1999, 43, 782–788 CrossRef CAS PubMed
.
- T. L. Raguse, E. A. Porter, B. Weisblum and S. H. Gellman, J. Am. Chem. Soc., 2002, 124, 12774–12785 CrossRef CAS PubMed
.
- A. M. Carmona-Ribeiro and L. D. de Melo Carrasco, Int. J. Mol. Sci., 2013, 14, 9906–9946 CrossRef PubMed
.
- M. A. Gelman, B. Weisblum, D. M. Lynn and S. H. Gellman, Org. Lett., 2004, 6, 557–560 CrossRef CAS PubMed
.
- J. Overhage, A. Campisano, M. Bains, E. C. W. Torfs, B. H. A. Rehm and R. E. W. Hancock, Infect. Immun., 2008, 76, 4176–4182 CrossRef CAS PubMed
.
- L. L. Foster, S.-i. Yusa and K. Kuroda, Antibiotics, 2019, 8, 61 CrossRef CAS PubMed
.
- L. R. Martinez, M. R. Mihu, G. Han, S. Frases, R. J. B. Cordero, A. Casadevall, A. J. Friedman, J. M. Friedman and J. D. Nosanchuk, Biomaterials, 2010, 31, 669–679 CrossRef CAS PubMed
.
- H. Lu, C. Tu, T. Zhou, W. Zhang, Y. Zhan, J. Ding, X. Wu, Z. Yang, W. Cao, L. Deng, C. Gao and F. Xu, Chem. Eng. J., 2022, 436, 135130 CrossRef CAS
.
- Y. Yang, Y. Qian, M. Zhang, S. Hao, H. Wang, Y. Fan, R. Liu, D. Xu and F. Wang, J. Mater. Sci. Technol., 2023, 133, 77–88 CrossRef CAS
.
- Y. Bi, X. Chen, F. Luo, X. Wang, X. Chen, J. Yao and Z. Shao, Biomater. Sci., 2024, 12, 5337–5348 RSC
.
- D. S. S. M. Uppu, S. Samaddar, J. Hoque, M. M. Konai, P. Krishnamoorthy, B. R. Shome and J. Haldar, Biomacromolecules, 2016, 17, 3094–3102 CrossRef CAS PubMed
.
- O. Al Musaimi, B. G. de la Torre and F. Albericio, Green Chem., 2020, 22, 996–1018 RSC
.
- K. Yang, B. Lv, H. Shen, G. Jing and Z. Zhou, Int. J. Life Cycle Assess., 2021, 26, 64–75 CrossRef CAS
.
- G. Batoni, G. Maisetta and S. Esin, Biochim. Biophys. Acta, 2016, 1858, 1044–1060 CrossRef CAS PubMed
.
- E. Galdiero, L. Lombardi, A. Falanga, G. Libralato, M. Guida and R. Carotenuto, Pharmaceutics, 2019, 11, 322 CrossRef CAS PubMed
.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |






