Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Lessons learned on how to reengineer enzymes for improved performance in ionic liquids: insights from BSLA saturation mutagenesis library
Shuaiqi
Meng
ab,
Yu
Ji
*ab,
Maxine
Yew
 c,
Leilei
Zhu
c,
Leilei
Zhu
 *c and
Ulrich
Schwaneberg
*c and
Ulrich
Schwaneberg
 *ab
*ab
aState Key Laboratory of Green Biomanufacturing, National Energy R&D Center for Biorefinery, Beijing Key Lab of Bioprocess, College of Life Science and Technology, Beijing Advanced Innovation Center for Soft Matter Science and Engineering, Beijing University of Chemical Technology, Beijing 100029, China. E-mail: yuji@buct.edu.cn; u.schwaneberg@biotec.rwth-aachen.de
bInstitute of Biotechnology, RWTH Aachen University, Worringerweg 3, 52074 Aachen, Germany
cState Key Laboratory of Engineering Biology for Low-Carbon Manufacturing, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, 32 West 7th Avenue, Tianjin 300308, PR China. E-mail: zhu_ll@tib.cas.cn
First published on 23rd February 2026
Abstract
Ionic liquids (ILs) are attractive reaction media in biocatalysis due to their excellent substrate solubilization properties and their promotion of mild and often environmentally friendly reaction conditions. However, enzyme activity is typically reduced at IL concentrations above 10%. In recent decades, continuous efforts in enzyme engineering have aimed to improve enzyme resistance to ILs, yet achieving robust enzymes remains challenging. This review summarizes research efforts over the past decades aimed at improving IL resistance of enzymes, spanning mechanistic insights and engineering strategies. Analyses of enzyme–IL interactions revealed that the primary effect of ILs is the stripping of water molecules from the enzyme surface. Subsequently, a comprehensive site-saturation mutagenesis (SSM) library of Bacillus subtilis lipase A (BSLA), covering all 181 positions, provided a systematic basis for understanding IL tolerance. Screening this library in the presence of four [BMIM]-based ILs ([BMIM]Cl, [BMIM]Br, [BMIM]I, and [BMIM][TfO]) revealed a general engineering principle: the hydration shell of enzymes is a key determinant of IL resistance. Finally, strategies to identify functional positions associated with improved IL resistance and to efficiently recombine beneficial substitutions are discussed. These engineering approaches minimize experimental effort while maximizing enzyme performance in ILs, providing a powerful and broadly applicable framework for the future design of IL-tolerant enzymes.
Green foundation1. Ionic liquids are promising green solvents for enzyme catalysis, offering low volatility, tunable properties, and potential to replace harmful organic solvents; however, limited enzyme resistance has restricted their wider use.2. This review elaborates on the interactions between enzymes and ionic liquids, and discusses state-of-the-art strategies to enhance enzyme tolerance, providing pathways to unlock their sustainable potential in biocatalysis and expand industrial relevance. 3. Future advances could focus on designing even more robust enzymes, integrating ionic liquids with renewable feedstocks, and scaling applications to maximize their role in sustainable manufacturing. |
1. Introduction
Enzymatic biocatalysis is widely used in synthesis of valuable compounds such as natural products, pharmaceuticals, and biofuels.1–4 Enzymes are generally designed by nature to efficiently perform reactions in aqueous environment. However, many valuable compounds and natural polymers have poorly solubility in water, which restricts substrate loading, reduces volumetric productivity, and ultimately constrains overall biocatalytic performance.5–9 Ionic liquids (ILs) offer a promising alternative reaction medium capable of overcoming productivity and performance limitations for insoluble compounds in enzyme-catalyzed reactions thereby enhancing catalytic efficiency and process productivity.10 For example, Lozano et al. employed lipase Novozym 435 for the synthesis of panthenyl monoacyl esters in [C12mim][BF4], achieving up to 90% conversion and 100% selectivity.11 In another example, Zhao et al. synthesized a novel ionic liquid, [CH3OCH2CH2-Im-t-BuOH][Tf2N], which enabled ε-caprolactone polymerization with conversions of up to 74%, compared with only 37% under solvent-free conditions.12 Driven by the advantage of ILs, research interest in IL-based biocatalysis has grown rapidly. The applications of ILs have been reported across a wide range of enzyme, including lipases,13–17 cellulases,18,19 PETases,20 and xylanases,21 and laccases.22–24ILs possess unique properties such as strong solvating power, negligible vapor pressure, and high thermal stability, which make them especially attractive for biocatalytic processes.25 In simple terms, ILs are molten salts with melting points typically below 100 °C. They are generally composed of organic cations (e.g. imidazolium, pyridinium, alkylated ammonium) paired with organic or inorganic anions (e.g. chloride, nitrate, tetrafluoroborate, hexafluorophosphate, methylsulfate), as illustrated in Fig. 1.26,27 ILs are often referred to as “designer solvents” as the cationic and anionic components can be independently varied.28 This tunability allows the fine adjustment of key solvent properties, such as polarity, viscosity, miscibility, and hydrophobicity or hydrophilicity. Moreover, ILs can be mixed with other solvents to form homogeneous aqueous or multiphasic systems, expanding their versatility in application. In such systems, ionic composition can be tailored to manipulate solvent–solute interactions by altering physicochemical properties.29 For instance, ILs can be designed to be immiscible with low-polarity or non-polar organic solvents, thereby enabling selective extraction and product separation.30
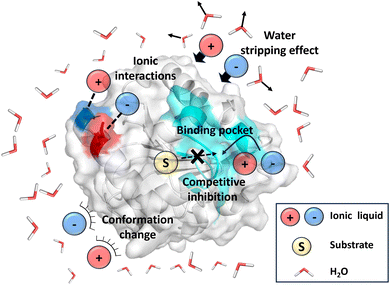
The main limitation to the broad range of IL applications is the limited availability of enzymes capable of withstanding high IL concentrations while maintaining adequate process stability.31–33 The selection of the cation–anion pair plays a critical role in determining the stability and compatibility of IL–enzyme systems. Certain classes of ILs are known to be toxic to whole cells and proteins, often causing irreversible deactivation of enzymatic function. For instance, CALB retains only 3% of its catalytic activity after incubation in [BMIM][NO3], primarily due to strong hydrogen bonding between the anions and the enzyme, which disrupts its secondary structure and leads to inactivation.34 The general effects of ILs on enzyme activity and stability include (Fig. 2):
(a) Ionic interactions (e.g., salt bridges), particularly on the enzyme surface, which can destabilize the protein's secondary structure;35,36
(b) Water-stripping effects, where ILs remove essential hydration layers from the enzyme surface, reducing flexibility and altering conformation of enzyme;37,38
(c) Occupation of the enzyme's active site by IL molecules, which can act as reversible inhibitors by blocking substrate binding;39
(d) Destabilization of the folded state of proteins, potentially leading to unfolding, aggregation, and loss of catalytic activity.40,41
Both the anion and the cation part of IL can significantly influence the activity of enzyme depending on their interactions with water and the specific enzyme involved.42 For example, in imidazolium-based ILs used for cellulose dissolution, the anion is generally considered as the dominant factor responsible for cellulase deactivation.43 In contrast, for enzymes such as Bacillus subtilis lipase A (BSLA) and CALB, imidazolium cations with varying alkyl chain lengths have more pronounced effect on enzyme structure and stability.37,41,44,45
The influence of individual ions on protein stability can be briefly understood through the Hofmeister series, which classifies ions as either kosmo tropes (structure stabilizers) or chaotropes (structure disruptors) based on their ability to order or disorder water molecules.46 Based on the Hofmeister series, an overall trend in anion-induced destabilization is observed in the following order: C4H4O62− > SO42−>HPO42− > C3H5O(CO2)33− > CH3CO2− > HCO3− > CrO42− > Cl− > NO3− > ClO3−.47 The corresponding sequence for cations is: Li+ > K+ ≈ Na+ > NH4+ > Mg2+.47 However, the Hofmeister series does not always reliably predict enzyme behavior in ILs. For instance, while halide anions such as Cl− and Br− are generally considered denaturing agents, CALB has shown 6-fold improvement in activity in the presence of 1-decyl-3-methylimidazolium chloride.48 The effects of ILs on proteins also influenced by various factors such as protein structure, IL concentration, temperature, and pH.49,50
Overall, the influence of ILs on enzymes is complex. Several successful strategies have been reported to enhance enzyme resistance to ILs through enzyme engineering51–53 and chemical modification.15,54,55 An example of chemical modifications is the modification of CALB structure with betaine ionic liquids of different chain lengths resulting in improved enzyme activity, thermal stability and DMSO tolerance.15 The flexibility of CALB was enhanced by the IL modifier which results in increased number of water molecules surrounding the enzymatic active sites. Among the enzyme engineering reports several cases employed site-saturation mutagenesis libraries (SSM).13,56 A representative example is the BSLA SSM library, which encompasses 3620 BSLA variants and represents the complete natural sequence diversity generated by single amino acid substitutions at all BSLA positions.13 The BSLA-SSM libraries were screened in four ionic liquids: [BMIM][Cl], [BMIM][Br], [BMIM][I], and [BMIM][TfO]. The generated datasets offer a comprehensive study for quantifying and analyzing enzyme resistance to ILs, providing the first insight into how lipases can be rationally redesigned to function effectively in ILs. This dataset provides a robust model system for elucidating general design principles governing enzyme resistance in IL environments.
The application of ILs as solvents for enzymatic reactions has been extensively summarized in previous reviews, which provide comprehensive overviews of their physicochemical properties, effects on catalysis, and potential process advantages.57–61 As these aspects have been well covered elsewhere, they are not the focus of the present review. Instead, this review concentrates on enzyme engineering strategies aimed at evolving enzymes with enhanced tolerance toward ILs. Using large-scale mutational fitness landscape analysis of BSLA as an entry point, we comprehensively discuss enzyme behavior in IL environments, elucidate molecular mechanisms of enzyme–IL interactions, and summarize engineering and rational recombination strategies. Based on these systematic analyses, generalizable design principles for the development of IL-tolerant enzymes are developed, providing a framework to guide future biocatalyst design in non-conventional reaction media.
2. Enzyme behavior in ILs
Gaining insights into enzyme behavior in ionic liquids will be illustrated on the example of BSLA. Firstly, will explore the change of BSLA's properties in four different ILs ([BMIM]Cl, [BMIM]Br, [BMIM]I, and [BMIM][TfO]). This will involve a detailed analysis of how ILs influence the enzyme's hydrogen bond network, salt bridge network, flexibility, and hydration shell. Secondly, the specific roles of cations and anions are discussed, with emphasis on how these interactions affect overall enzyme performance in IL environments.2.1 Structural changes of enzymes in ILs
Enzymes properties are significantly affected by interactions between anions, cations, cosolvent water with proteins.62 Enzymes are by natural evolution designed to operate efficiently within cell environments and in aqueous solutions. Exposure to ionic liquids (ILs) often results in reduced enzyme activity.13,60 In the case of BSLA, extensive MD-simulations were conducted using Gromacs to investigate its behavior in IL environments. Each production simulation included 100 ns runs performed in triplicate to thoroughly examine the enzyme's behavior in ILs (Fig. 3).63–65 Overall, the structure of BSLA remains preserved in ILs ([BMIM]Cl, [BMIM]Br, [BMIM]I, and [BMIM][TfO]), which is reflected in the RMSD values.64 In binary mixtures of IL and water (18.3% v/v [BMIM][Cl], 13.2% v/v [BMIM][Br], 10.0% v/v [BMIM][I], and 15.0% v/v [BMIM][TfO] in water), BSLA conformation has alterations when exposed to ionic liquids in contrast to the aqueous phase. For instance, BSLA structure shows a partial swelling in all four ILs, as evidenced by an augmented radius of gyration and increased solvent accessibility surface areas.64 Furthermore, a reduction in the number of H-bonds and a decrease in number of salt bridges are observed in the presence of ILs.64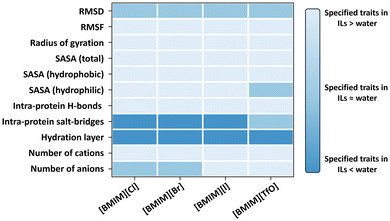 | ||
| Fig. 3 Geometrical properties and protein solvation phenomenon investigation for BSLA WT in 4 ILs ([BMIM]Cl, [BMIM]Br, [BMIM]I, and [BMIM][TfO]). Color representation highlights higher (light blue), unchanged (blue), and lower (dark blue) enzyme performances in ILs relative to that in an aqueous environment.63–65 This figure adapted from ref. 63–65 with permission from Royal Society of Chemistry, copyright © 2019,63 and American Chemical Society. Copyright © 2019.64 | ||
2.2 Molecular interactions between enzymes and ILs
Ionic interactions between ILs and proteins dictate enzymatic performance in ILs. The impact of four ionic liquids ([BMIM]Cl, [BMIM]Br, [BMIM]I, and [BMIM][TfO]) on BSLA was investigated.64,66 BSLA was simulated in these four ILs using Gromacs with the GROMOS96 54a7 force field. Each production simulation included 100 ns runs performed in triplicate, within a 10 Å simulation box. The spatial distribution function reveals the ions distribution around BSLA (see Fig. 4). [BMIM] + has more predominant interactions with BSLA than halogen anions (Cl−, Br−, and I−).64 In particular, approximately 30–40 BMIM+ cations interacted within the first solvation shell of the BSLA surface, whereas the number of binding molecules was less than 10 Cl−, Br−, and I− anions, respectively.64 In these instances, BMIM+ cation interacts more favorably with the BSLA surface in the case of halogenated-BMIM, potentially playing a dominant role in the reduction of catalytic activity of BSLA in ILs. Interestingly, the TfO− anion interacts more frequently with BSLA compared to the halogen anions.64 Due to higher electrostatic and counterion effects, TfO− anions and BMIM+ cations interact with similar regions of the BSLA surface and display similar interact patterns. Overall, the binding intensity of anions follows a trend of TfO− > I− > Br− > Cl− (Fig. 4).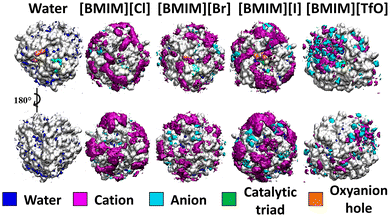 | ||
| Fig. 4 The BSLA properties in ILs. Spatial distribution function (SDF) for the solvent distribution of the BSLA surface in water and ILs simulations.64 Color code: gray, enzyme surface; orange, oxyanion hole; green, catalytic triad; blue, water; purple, BMIM+, and cyan, anions. This figure reproduced from ref. 64 with permission from Copyright © 2019, American Chemical Society. | ||
The radial distribution function elucidates the orientation of ions and thus unveils the binding mode between ions and BSLA. For BMIM+ cation, the initial solvation shells appeared within approximately 5 Å.64 BMIM+ cation primarily interacts with BSLA using its hydrophobic tail, indicating that the interaction between BMIM+ and BSLA occurs through hydrophobic interactions. Additionally, π–π interactions between BMIM+ cation and Y139 and W42 of BSLA are also observed.64 Modifying the protein surface potential has demonstrated to have effects on BMIM+ cation binding.66 These observations suggested that multiple factors contribute to the interaction between ions and BSLA. As for the anions, all halogen ions exhibit the first solvation shell at approximately 2.25 Å, while TfO− has its initial solvation shell at 5 Å. The hydrophobic C-terminal of TfO− exhibits a preferential interaction with BSLA.64
Ionic liquids could interact more strongly with BSLA surfaces than water.67 Notably, the distribution of water on enzyme surface is reduced in ILs compared to pure water environment.64,66 The dominant surface interactions of ionic species and the displacement of structurally essential water molecules from the BSLA surface collectively resulted in significant attenuation of enzymatic activity. Compared with halogen anion, the BMIM+ cations displayed a strong affinity for the enzyme surface, especially towards the catalytic triad.63,64 Meanwhile, the binding of both anions and cations of [BMIM][TfO] on BSLA surface contributed to the stripping off of essential water molecules.64
3. Engineering strategies for improving enzyme resistance ILs
Reports to improve ILs resistance comprise directed evolution approaches,68 rational approaches,69 and combined approaches.70 The following sections introduce and discuss these approaches, with particular emphasis on the BSLA model system. This chapter is divided into four main parts: part 3.1 describes the previous directed evolution campaign on enzymes for improved IL resistance; part 3.2 discusses the rational design strategy and the necessity of the BSLA-SSM library, which reflects the complete natural diversity of BSLA at each position; part 3.3 discusses the statistical analysis of the BSLA-SSM library, which identified the hydration shell as a key factor strongly correlating with enzyme IL resistance; parts 3.4 and 3.5 elaborate on two approaches for improving enzyme performance in ILs: surface charge engineering and introducing hydrophobic amino acids at buried sites. Finally, part 3.6 discusses the CompassR strategy for efficiently recombining the beneficial substitutions.3.1 Directed evolution studies
Directed evolution relies on random and iterative mutagenesis to introduce diversity. High throughput screening is then employed to identify variants with the desired characteristics. This approach allows the engineering of enzymes without prior knowledge of their structure and mechanism, and its effectiveness has been demonstrated in many cases. For example, a directed evolution campaign screening 1500 BSLA clones identified the M1 variant (M134N/N138S/L140S), which exhibited nearly double the specific activity (16.9 vs. 9.4 U mg−1) and improved resistance (233% vs. 111%) at 9 vol% 1-butyl-3-methylimidazolium trifluoromethanesulfonate compared its WT.71 Notably, all substitutions in the beneficial M1 variant are located on the enzyme surface near the substrate-binding cleft.71 These substitutions are proposed to enlarge and increase the polarity of the substrate-binding environment, thereby facilitating improved accommodation of the substrate p-nitrophenyl butyrate (pNPB).71 In addition, the introduction of polar residues is likely to strengthen interactions with surrounding water molecules, potentially contributing to an enhanced local hydration layer.In another case, laccase from T. versicolor was evolved through two rounds of error-prone PCR (epPCR), screening a total of 2800 clones. This effort yielded the variant M3 (F265S/A318 V) with a 4.5-fold improved activity compared to the WT in the presence of 15% (v/v) [EMIM] [EtSO4].68 The two substitutions are distant from each other but both are located on the enzyme surface.68 Relative folding free-energy calculations (ΔΔGfold) indicated that A318 V contributes to enzyme stabilization, whereas F265S is slightly destabilizing, suggesting that their combined effect may be synergistic rather than additive.68
However, despite directed evolution has been proven to be effective, its drawbacks are equally notable, including the extensive screening load and its inability to investigate the entire protein sequence, rather only a limited number of protein variants can be examined at a time. It is therefore challenging to rely on directed evolution alone to provide a comprehensive understanding of how to enhance enzyme resistance to ILs, as a prerequisite for protein engineering based on rational design.
3.2 BSLA-SSM library empowers semi-rational design
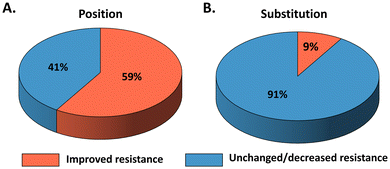
To improve enzyme resistance in ILs, extensive efforts have been devoted to developing engineering strategies. Surface engineering represent one of the most widely applied approaches. The principle is to modulate enzyme–IL interactions in order to mitigate the detrimental effects of ILs on protein structure and function.16 For example, Zhou et al. reported a BSLA variant containing eight surface substitutions (A20E/K44E/Y49E/R57E/G111D/A132D/M134E/G158E), which exhibited up to 17.2-fold improvement in resistance in [BMIM][Cl] compared with its WT.16 Notably, this variant also displayed significantly enhanced thermostability, with its melting temperature (Tm) increasing from 51 °C to 97 °C.16 In another case, Nordwald et al. employed two-dimensional heteronuclear single quantum coherence NMR to assess IL-induced perturbations in BSLA by analyzing chemical shift changes. The identified beneficial variant (G158E/K44E/R57E/Y49E) showed more than a seven-fold increase in resistance in [BMIM][Cl] relative to the WT.69 In general, improved IL resistance is often associated with enhanced protein stability and reduced unfavourable interactions with ionic liquids. While these studies have provided valuable mechanistic insights and demonstrated that surface engineering can enhance enzyme performance in ILs, most of them rely on a limited number of substitutions or case-specific designs, which restrict their generality. To move beyond isolated examples and enable a systematic, quantitative evaluation of enzyme–IL interactions, the BSLA-SSM library was developed as a model system that allows comprehensive assessment of enzyme fitness across multiple IL environments.BSLA has a minimal α/β-hydrolase fold, making it an ideal model enzyme for studying IL resistance.72 To gather a comprehensive understanding of enzyme IL resistance and elucidate the fitness landscape of the BSLA library, Schwaneberg et al. generated a BSLA-site saturation mutagenesis (SSM) library comprising 3620 variants.13 This library offered the 20 naturally occurring amino acids of BSLA at all 181 positions through SSM. The evaluation of resistance of four imidazolium-based ILs ([BMIM]Cl, [BMIM]Br, [BMIM]I, and [BMIM][TfO]) revealed that 6%–13% (13% for [BMIM][Cl], 6% for [BMIM][Br], 7% for [BMIM][I], and 8% for [BMIM][TfO]) of all substitutions show increased ILs resistance than BSLA WT (Fig. 5A and Table 1). These beneficial substitutions were distributed across 50%–69% (69% for [BMIM][Cl], 50% for [BMIM][Br], 52% for [BMIM][I], and 57% for [BMIM][TfO]) of all BSLA positions (Fig. 5B).13,73 The BSLA WT consists of 43% non-polar, 34% polar, 9% negatively charged, and 14% positively charged amino acids. Generally, there is a preference for substitutions by chemically different amino acids (such as aromatic to polar/aliphatic/charged amino acids) over chemically similar ones.13 Therefore, using mutagenesis methods, such as SDM and SeSaM, to introduce chemically different substitutions could be considered to enhance the enzyme IL resistance.74 Among the 20, naturally charged amino acids are more favored as beneficial substitutions (Table 1).13
| Ionic liquids | Amino acid type % (variant number) | ||||
|---|---|---|---|---|---|
| Positively charged | Negatively charged | Non-polar | Polar | Total | |
| a The entire BSLA-SSM library contains 3620 variants. b All percentage values are normalized to BSLA amino acid composition, as BSLA amino acid composition is not evenly distributed (43% non-polar, 34% polar, 9% negatively charged, and 14% positively charged amino acid). The following amino acid classification was used: nonpolar: G, A, V, L, I, M, F, W, P. Polar: S, T, C, Y, N, Q. Negatively charged: D, E. Positively charged: K, R, H. | |||||
| [BMIM][Cl] | 29.1% (85) | 31.4% (59) | 20.4% (183) | 19.1% (135) | 100% (462) |
| [BMIM][Br] | 28.5% (37) | 32.3% (27) | 21.8% (87) | 17.4% (55) | 100% (206) |
| [BMIM][I] | 28.6% (49) | 34.5% (38) | 20.7% (109) | 16.1% (67) | 100% (263) |
| [BMIM][TfO] | 24.9% (46) | 34.5% (41) | 19.0% (108) | 21.6% (97) | 100% (292) |
On average, the ratio of beneficial substitutions of BSLA was 8.5%, and for positions, it was 57%. The high ratio makes it easy to obtain key positions and promising variants in simple calculational design. Analyses that largely focus on limited variants may lead to conclusions that deviate from the actual mechanism, resembling a “blind person touching an elephant”. Harrar et al. compared 22 previously described structure-based approaches that aimed at increasing enzyme IL resistance.73 Surprisingly, most of the approaches performed worse than random mutagenesis, and only 2 methods (leveraging experimental information on thermostability and targeting the structural weak spots of enzymes) showed improved prediction accuracy.73 This comprehensive study showed that global design principles of enzymes cannot be found from the analysis on limited variants. It highlights the importance of considering sufficient prior information from large and diverse datasets to enhance enzyme IL resistance.
3.3 Enzyme hydration shell determines the enzyme IL resistance
As discussed in Section 2, the conformations of BSLA are altered upon exposure to ILs. This change can be attributed to IL ions displacing water molecules from the first hydration shell of the enzyme. Such dehydration weakens surface hydrogen-bond networks and salt bridges and, in some cases, perturbs local secondary structural elements, ultimately contributing to the reduction in enzyme activity.5,64 Therefore, enhancing the hydration shell represents a key engineering strategy for improving enzyme resistance to ILs, which was supported by previous experimental and computational studies. For example, Wallraf et al. employed KnowVolution for engineering laccases LCC2 from Trametes versicolor.70 By introducing relatively hydrophilic substitutions A310D/A312P/A318R, the variant OM3 exhibited a 4.5-fold improvement in activity in the presence of 15% (v/v) [EMIM][EtSO4].70 Zhao et al. conducted molecular dynamics (MD) simulations to examine the behavior of 60 BSLA substitutions in 15% (v/v) [BMIM][TfO], comprising 50 beneficial substitutions and 10 non-beneficial substitutions.63 The simulation results demonstrated that, in water, all substitutions exhibited similar patterns of surface water molecules. However, in [BMIM][TfO], 72% of beneficial substitutions showed an enhanced hydration layer, while 70% of non-beneficial substitutions had a damaged hydration layer.63The importance of the hydration layer in driving improved enzyme IL resistance was further explored by Cui et al. through comprehensive MD simulations on 25 BSLA substitutions at 19 positions from the BSLA-SSM library.75 They evaluated 45 molecular observables, including geometrical properties, solvation phenomena, and BSLA-solvent interaction energies, to assess the dynamic and structural behavior of BSLA substitutions. Surprisingly, most observables exhibited unpredictable behavior in both beneficial and non-beneficial variants.75 Certain observables, such as [BMIM]+-residue contact frequency and the distance between Ile12-N and Met78-N, showed significance in a specific IL but not in another IL. Among all observables, the only general factor related to IL resistance was the enzyme hydration layer. Thus, tailoring enzymes with high IL resistance based on enhancing the hydration layer was feasible. Cui et al. further generated beneficial variants using the stepwise recombination method (Fig. 6).75 These variants showed an evidently positive correlation with the number of water molecules around substituted positions (up to R2 = 0.96 in [BMIM][TfO] and R2 = 0.85 in [BMIM]Cl, Fig. 6B). Importantly, the mutagenesis effects were not limited to the mutagenesis site; the locally improved hydration level could “pass” to the entire protein, thereby enhancing the hydration shell of the entire protein (Fig. 6C).75 In summary, the determinant factor influencing the difference in IL resistance is the hydration shell surrounding the enzyme. Appropriate engineering strategies could be developed based on enhancing the hydration shell to improve enzyme IL resistance.
 | ||
| Fig. 6 Solvation behaviors of BSLA variants in 18.3% (v/v) [BMIM]Cl and 15.0% (v/v) [BMIM][TfO].75 (A) Visualization of the targeted mutation positions in BSLA. (B) Influence of the hydration shell on the IL resistance of the BSLA variants relative to the WT. (C) Spatial distribution of water, BMIM+, and Cl−/TfO− occupancy on the surface of BSLA variants. This Figure reproduced from ref. 75 with permission from Copyright © 2022, American Chemical Society. | ||
3.4 Surface charge engineering
Analysis of the BSLA-SSM library revealed the beneficial substitutions present in both surface-exposed and buried regions, with differences in substitution patterns in these regions.13 More than 70% of beneficial substitutions were identified in exposed position (Fig. 7). For surface-exposed positions, an effective strategy is to modify the distribution of surface charges.13 By introducing or rearranging charged residues, it is possible to modulate electrostatic interactions between the enzyme and IL components. Such modifications can reduce unfavorable ion accumulation at the protein surface, promote water retention in the first hydration shell, and thereby help maintain native hydrogen-bond networks and structural integrity.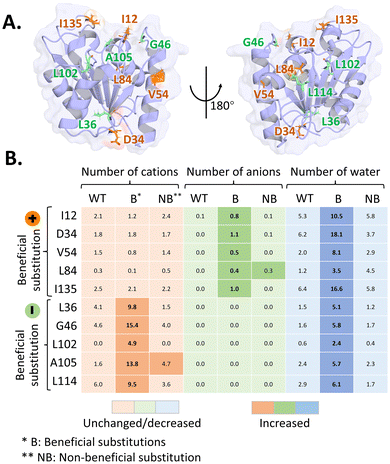 | ||
| Fig. 7 Effect of surface charged substitutions on BSLA variants.66 (A) Visualization of the targeted mutation positions in BSLA. At the targeted positions, beneficial substitutions introducing positively charged residues are marked in orange, whereas beneficial substitutions introducing negatively charged residues are marked in green. (B) Heatmaps showing the number of BMIM+ cations, Cl− anions, and water molecules surrounding the substituted sites in BSLA wild type and variants. In the top five variants, beneficial substitutions introduce positively charged residues, whereas in the bottom five variants, beneficial substitutions introduce negatively charged residues. This Figure adapted from ref. 66 with permission from Copyright © 2022, American Chemical Society. | ||
The effects of positively and negatively charged substitutions on enzyme–IL interactions are not equivalent and can lead to distinct solvation behaviors. Charged residues tend to preferentially interact with oppositely charged IL ions, thereby modulating the local ionic environment around the protein surface. Consequently, the choice of introduced residues can, in principle, be guided by the dominant ionic species present in a given IL system. In the case of BSLA in [BMIM]-based ILs, the bulky [BMIM]+ cation exerts a stronger perturbing effect on the enzyme surface than smaller anions such as Cl−, Br−, or I−.64 Therefore, introducing positively charged residues is more effective in repelling [BMIM]+ cations and promoting the retention of water molecules at the enzyme surface.66
Pramanik and colleagues systematically investigated the effect of charged substitutions on BSLA IL resistance.66 In terms of the ability to form salt bridges, their exploration revealed that negatively charged substitutions might be more prone to form additional salt bridges compared to positively charged substitutions.66 Meanwhile, the charged nature of substitution could influence the repulsion or attraction towards ILs (Fig. 7). Compared with the BSLA WT, positively charged substitutions resulted in the repulsion of BMIM+ cations and the attraction of anions. Conversely, negatively charged substitutions repulsed anions and attracted cations. MD simulation demonstrated that positive substitutions showed strong repulsion to BMIM+ cations and weak attraction to anions, while the negative substitutions had obvious attraction to BMIM+ cations.66 Therefore, positively charged substitutions allowed for the retention of more water molecules at the BSLA surface than the negative ones. Analysis of the BSLA-SSM library also showed that most of the positions favorable for resistance improvement are located in the BMIM + binding region, thus weakening the interaction between BMIM and the enzyme.13,63,66,75
3.5 Introducing hydrophobic amino acids at buried sites
Less than 30% of beneficial substitutions occur at buried sites in the protein.13 At these buried positions, polar and hydrophobic substitutions resulted in more improved variants than charged substitutions.13,63 This is likely because buried positions are predominantly composed of hydrophobic amino acids, making it difficult to introduce stabilizing charged residues into the protein interior. Moreover, internal residues have limited direct influence on modulating enzyme–IL interactions or maintaining the surface hydration shell.In this context, beneficial substitutions at buried positions are more likely to enhance intrinsic protein stability rather than directly affect interactions with ILs. Given that the hydrophobic network exists within the protein, a potential strategy is to introduce hydrophobic amino acids at buried sites, thereby enhancing the hydrophobic interactions near the substitutions and therefore stabilizing the protein.63
3.6 CompassR – beneficially recombine the beneficial substitutions
In earlier discussions, the challenge of identifying beneficial substitutions has been resolved through analyzing the BSLA-SSM model library. However, new challenges emerged regarding how to effectively recombine these beneficial substitutions. Simply recombining more than two or three beneficial substitutions may not necessarily result in further improved enzyme variants.76 In some cases, such recombination could even lead to damage to protein functions.77 It is estimated that at least 80% of substitutions with deleterious effects can be attributed to severe destabilization of the protein fold.78To tackle this challenge, Schwaneberg et al. developed the CompassR strategy, which aims to guide the recombination of beneficial substitutions.79,80 The principle of CompassR is to utilize the relative free energy of folding (ΔΔGfold) to predict the effects of substitutions in recombinant variants (Fig. 8). This strategy could efficiently enhance enzyme properties by recombining beneficial substitutions that contribute to intrinsic stability, resulting in enzymes with improved resistance in ILs. An application of the CompassR strategy is the generation of BSLA variant M1a (F17S/V54K/D64N/D91E/G155N), which exhibited up to a 6.7-fold higher resistance against 40% (v/v) [BMIM]Cl, 5.6-fold in 80% (v/v) [BMIM]Br, 5.0-fold in 30% (v/v) [BMIM][TfO], and 2.7-fold in 10% (v/v) [BMIM]I.56 The power of the CompassR strategy suggests the possibility of recombining more than five beneficial substitutions, paving the way for the design of enzymes with much-improved IL resistance in the future.
 | ||
| Fig. 8 Enzyme engineering pipeline for engineering high IL-resistant lipases. The parent lipase was diversified and screened/designed, yielding beneficial substitutions with improved hydration layer. These substitutions were then recombined using the CompassR strategy to preserve the enzyme's inherent stability, resulting in stable recombinants with enhanced ILs resistance. Part of this figure was reproduced from ref. 81 with permission from Copyright © 2020, American Chemical Society. | ||
When sufficient computational resources are available, MD simulations could also be employed to predict beneficial recombinants.82 For example, these simulations could monitor changes in hydration layers, thus predicting ILs resistance of variants. While CompassR is based on thermodynamic stability analysis, molecular dynamics focuses on kinetic analysis. By combining the strengths of both methods, more effective reorganization strategies can be developed. Furthermore, these data can feed into data-driven approaches such as machine learning to reduce experimental workload and further improve IL resistance of variants.
4. Conclusion
The key lesson in improving resistance to ILs is the necessity of uncovering the molecular insights by full diversity generation of enzymes. After studying BSLA-SSM library, a general engineering principle is that the hydration layer surrounding the enzyme determines the enzyme resistance to ILs. By enhancing this hydration shell, e.g., through surface charge engineering, has proven to be an effective strategy for improving the enzyme IL resistance. Furthermore, recombined with CompassR strategy (based on relative free energy of folding) offers a promising approach to guide the recombination of beneficial substitutions. Therefore, the integration of enhancing hydration shell and advanced recombination strategy enables the engineering of enzymes that function effectively in highly concentrated or even pure ILs in the future.Conflicts of interest
There are no conflicts to declare.Data availability
This article is a review and does not report new primary data. All data discussed are available in the cited literature.Acknowledgements
Prof. Yu Ji acknowledges support from the Fundamental Research Funds for the Central Universities (buctrc202526, JD2528). This work was supported by the National Key Research and Development Program of China (2025YFF0521500). This study was also supported by the Tianjin Synthetic Biotechnology Innovation Capacity Improvement Project (TSBICIP-IJCP-003) and the International Partnership Program of the Chinese Academy of Sciences (Grant No. 306GJHZ2025003BS).References
- J.-M. Choi, S.-S. Han and H.-S. Kim, Biotechnol. Adv., 2015, 33, 1443–1454 CrossRef CAS PubMed.
- S. Galanie, D. Entwistle and J. Lalonde, Nat. Prod. Rep., 2020, 37, 1122–1143 RSC.
- S. Meng, Y. Ji, L. Zhu, G. V. Dhoke, M. D. Davari and U. Schwaneberg, Biotechnol. Adv., 2022, 61, 108051 CrossRef CAS PubMed.
- S. Meng, Z. Li, Y. Du, X. Cheng, B. Nian, S. Fu, X. Song and H. Cui, Angew. Chem., Int. Ed., 2026, e25086, DOI:10.1002/anie.202525086.
- P. Xu, S. Liang, M. H. Zong and W. Y. Lou, Biotechnol. Adv., 2021, 51, 107702 CrossRef CAS PubMed.
- S. Meng, J. Guo, Z. Li, K. Nie, H. Xu, T. Tan and L. Liu, Enzyme Microb. Technol., 2019, 131, 109417 CrossRef CAS PubMed.
- U. Markel, D. F. Sauer, J. Schiffels, J. Okuda and U. Schwaneberg, Angew. Chem., Int. Ed., 2019, 58, 4454–4464 CrossRef CAS PubMed.
- A. V. Shivange, J. Marienhagen, H. Mundhada, A. Schenk and U. Schwaneberg, Curr. Opin. Chem. Biol., 2009, 13, 19–25 CrossRef CAS PubMed.
- C. Zeymer and D. Hilvert, Annu. Rev. Biochem., 2018, 87, 131–157 CrossRef CAS PubMed.
- T. Itoh, Chem. Rev., 2017, 117, 10567–10607 CrossRef CAS PubMed.
- P. Lozano, E. Alvarez, S. Nieto, R. Villa, J. M. Bernal and A. Donaire, Green Chem., 2019, 21, 3353–3361 RSC.
- H. Zhao and C. Toe, Process Biochem., 2020, 98, 59–64 CrossRef CAS.
- V. J. Frauenkron-Machedjou, A. Fulton, L. Zhu, C. Anker, M. Bocola, K. E. Jaeger and U. Schwaneberg, ChemBioChem, 2015, 16, 937–945 CrossRef CAS PubMed.
- Z. Lu, M. Chen, T. Jin, B. Nian and Y. Hu, Int. J. Biol. Macromol., 2023, 248, 125894 CrossRef CAS PubMed.
- Y. Xue, X.-G. Zhang, Z.-P. Lu, C. Xu, H.-J. Xu and Y. Hu, Front. Bioeng. Biotechnol., 2022, 10, 850890 CrossRef PubMed.
- Y. Zhou, B. Pérez, W. Hao, J. Lv, R. Gao and Z. Guo, Biochem. Eng. J., 2019, 148, 195–204 CrossRef CAS.
- W. Cheng and B. Nian, Molecules, 2023, 28, 5848 CrossRef CAS PubMed.
- M. Maleki, S. Ariaeenejad and G. H. Salekdeh, Bioresour. Technol., 2022, 345, 126536 CrossRef CAS PubMed.
- S. Summers, C. Kraft, S. Alamdari, J. Pfaendtner and J. L. Kaar, ACS Sustainable Chem. Eng., 2020, 8, 11299–11307 CrossRef CAS.
- Z. Zara, D. Mishra, S. K. Pandey, E. Csefalvay, F. Fadaei, B. Minofar and D. Řeha, Molecules, 2022, 27, 119 CrossRef CAS PubMed.
- H. Hebal, N. Boucherba, B. Binay and O. Turunen, Biocatal. Biotransform., 2021, 39, 242–259 CrossRef CAS.
- S. Haj Kacem, S. Galai, A. Pérez de los Ríos, F. J. Hernández Fernández and I. Smaali, J. Chem. Technol. Biotechnol., 2017, 93, 174–183 CrossRef.
- T. Itoh and Y. Takagi, ACS Sustainable Chem. Eng., 2021, 9, 1443–1458 CrossRef CAS.
- J. Jia, P. Xue, L. Ma, K. Shi and R. Li, Biochem. Eng. J., 2022, 187, 108607 CrossRef CAS.
- A. T. Silva, C. Teixeira, E. F. Marques, C. Prudencio, P. Gomes and R. Ferraz, ChemMedChem, 2021, 16, 2604–2611 CrossRef CAS PubMed.
- L. C. Henderson and D. J. Eyckens, Front. Chem., 2019, 7, 263 CrossRef PubMed.
- R. Hayes, G. G. Warr and R. Atkin, Chem. Rev., 2015, 115, 6357–6426 CrossRef CAS PubMed.
- I. Newington, J. M. Perez-Arlandis and T. Welton, Org. Lett., 2007, 9, 5247–5250 CrossRef CAS PubMed.
- R. A. Sheldon, R. M. Lau, M. J. Sorgedrager, F. van Rantwijk and K. R. Seddon, Green Chem., 2002, 4, 147–151 RSC.
- N. Dib, C. M. O. Lepori, N. M. Correa, J. J. Silber, R. D. Falcone and L. Garcia-Rio, Polymers, 2021, 13, 1378 CrossRef CAS PubMed.
- N. J. Turner, Nat. Chem. Biol., 2009, 5, 567–573 Search PubMed.
- V. I. Pârvulescu and C. Hardacre, Chem. Rev., 2007, 107, 2615–2665 CrossRef PubMed.
- B. Singh, G. Bulusu and A. Mitra, J. Phys. Chem. B, 2015, 119, 392–409 CrossRef CAS PubMed.
- H. T. Imam, V. Krasňan, M. Rebroš and A. C. Marr, Molecules, 2021, 26, 4791 CrossRef CAS PubMed.
- H. Cui, L. Eltoukhy, L. Zhang, U. Markel, K.-E. Jaeger, M. D. Davari and U. Schwaneberg, Angew. Chem., Int. Ed., 2021, 60, 11448–11456 CrossRef CAS PubMed.
- L. Bui-Le, C. J. Clarke, A. Bröhl, A. P. S. Brogan, J. A. J. Arpino, K. M. Polizzi and J. P. Hallett, Commun. Chem., 2020, 3, 55 CrossRef CAS PubMed.
- S. Pramanik, G. V. Dhoke, K.-E. Jaeger, U. Schwaneberg and M. D. Davari, ACS Sustainable Chem. Eng., 2019, 7, 11293–11302 Search PubMed.
- M. A. Mohammad Latif, N. M. Micaêlo and M. B. Abdul Rahman, RSC Adv., 2014, 4, 48202–48211 Search PubMed.
- Z. Li, D. Roccatano, M. Lorenz, R. Martinez and U. Schwaneberg, J. Biotechnol., 2014, 169, 87–94 CrossRef CAS PubMed.
- H. Weingärtner, C. Cabrele and C. Herrmann, Phys. Chem. Chem. Phys., 2012, 14, 415–426 RSC.
- K. G. Sprenger, J. G. Plaks, J. L. Kaar and J. Pfaendtner, Phys. Chem. Chem. Phys., 2017, 19, 17426–17433 RSC.
- Y. Wang, H. Wang, L. Chen, W. Wang, Z. Yang, Z. Xue and T. Mu, Green Chem., 2023, 25, 4685–4695 RSC.
- M. B. Turner, S. K. Spear, J. G. Huddleston, J. D. Holbrey and R. D. Rogers, Green Chem., 2003, 5, 443–447 Search PubMed.
- E. M. Nordwald, J. G. Plaks, J. R. Snell, M. C. Sousa and J. L. Kaar, ChemBioChem, 2015, 16, 2456–2459 CrossRef CAS PubMed.
- H. S. Kim, D. Eom, Y.-M. Koo and Y. G. Yingling, Phys. Chem. Chem. Phys., 2016, 18, 22062–22069 Search PubMed.
- A. Kumar and P. Venkatesu, Int. J. Biol. Macromol., 2014, 63, 244–253 CrossRef CAS PubMed.
- K. P. Gregory, G. R. Elliott, H. Robertson, A. Kumar, E. J. Wanless, G. B. Webber, V. S. J. Craig, G. G. Andersson and A. J. Page, Phys. Chem. Chem. Phys., 2022, 24, 12682–12718 RSC.
- S. P. Ventura, L. D. Santos, J. A. Saraiva and J. A. Coutinho, Green Chem., 2012, 14, 1620–1625 RSC.
- H. Zhao, J. Chem. Technol. Biotechnol., 2016, 91, 25–50 CrossRef CAS PubMed.
- N. V. Veríssimo, F. A. Vicente, R. C. de Oliveira, B. Likozar, R. P. d. S. Oliveira and J. F. B. Pereira, Biotechnol. Adv., 2022, 61, 108055 CrossRef PubMed.
- V. J. Frauenkron-Machedjou, A. Fulton, L. L. Zhu, C. Anker, M. Bocola, K. E. Jaeger and U. Schwaneberg, ChemBioChem, 2015, 16, 937–945 CrossRef CAS PubMed.
- J. N. Pedersen, Y. Zhou, Z. Guo and B. Pérez, Biotechnol. Bioeng., 2019, 7, 1795–1812 CrossRef PubMed.
- A. Schindl, M. L. Hagen, S. Muzammal, H. A. Gunasekera and A. K. Croft, Front. Chem., 2019, 7, 347 CrossRef CAS PubMed.
- A. P. S. Brogan, L. Bui-Le and J. P. Hallett, Nat. Chem., 2018, 10, 859–865 CrossRef CAS PubMed.
- G. Ma, Z. Zhang, M. Chen, Y. Zhang, B. Nian and Y. Hu, J. Sci. Food Agric., 2024, 104, 2669–2678 CrossRef CAS PubMed.
- H. Cui, S. Pramanik, K.-E. Jaeger, M. D. Davari and U. Schwaneberg, Green Chem., 2021, 23, 3474–3486 RSC.
- M. Naushad, Z. A. Alothman, A. B. Khan and M. Ali, Int. J. Biol. Macromol., 2012, 51, 555–560 CrossRef CAS PubMed.
- T. Itoh, Chem. Rec., 2023, 23, e202200275 CrossRef PubMed.
- K. C. Badgujar, V. C. Badgujar and B. M. Bhanage, Curr. Opin. Green Sustainable Chem., 2022, 36, 100621 Search PubMed.
- M. Moniruzzaman, K. Nakashima, N. Kamiya and M. Goto, Biochem. Eng. J., 2010, 48, 295–314 CrossRef CAS.
- J. Dupont, B. C. Leal, P. Lozano, A. L. Monteiro, P. Migowski and J. D. Scholten, Chem. Rev., 2024, 124, 5227–5420 CrossRef CAS PubMed.
- N. Prabhu and K. Sharp, Chem. Rev., 2006, 106, 1616–1623 CrossRef CAS PubMed.
- J. Zhao, V. J. Frauenkron-Machedjou, A. Fulton, L. Zhu, M. D. Davari, K. E. Jaeger, U. Schwaneberg and M. Bocola, Phys. Chem. Chem. Phys., 2018, 20, 9600–9609 RSC.
- S. Pramanik, G. V. Dhoke, K.-E. Jaeger, U. Schwaneberg and M. D. Davari, ACS Sustainable Chem. Eng., 2019, 7, 11293–11302 CrossRef CAS.
- T. El Harrar, B. Frieg, M. D. Davari, K. E. Jaeger, U. Schwaneberg and H. Gohlke, Comput. Struct. Biotechnol. J., 2021, 19, 4248–4264 CrossRef CAS PubMed.
- S. Pramanik, H. Cui, G. V. Dhoke, C. B. Yildiz, M. Vedder, K.-E. Jaeger, U. Schwaneberg and M. D. Davari, ACS Sustainable Chem. Eng., 2022, 10, 2689–2698 CrossRef CAS.
- M. Chaudhary, S. Pal, K. Kumari, I. Bahadur, G. Gambhir and P. Singh, Adv. Appl. Ionic Liq., 2023, 417–441 Search PubMed.
- H. Liu, L. Zhu, M. Bocola, N. Chen, A. C. Spiess and U. Schwaneberg, Green Chem., 2013, 15, 1348–1355 RSC.
- E. M. Nordwald, G. S. Armstrong and J. L. Kaar, ACS Catal., 2014, 4, 4057–4064 CrossRef CAS.
- A.-M. Wallraf, H. Liu, L. Zhu, G. Khalfallah, C. Simons, H. Alibiglou, M. D. Davari and U. Schwaneberg, Green Chem., 2018, 20, 2801–2812 RSC.
- J. Zhao, N. Jia, K. E. Jaeger, M. Bocola and U. Schwaneberg, Biotechnol. Bioeng., 2015, 112, 1997–2004 CrossRef CAS PubMed.
- G. van Pouderoyen, T. Eggert, K. E. Jaeger and B. W. Dijkstra, J. Mol. Biol., 2001, 309, 215–226 CrossRef CAS PubMed.
- T. El Harrar, M. D. Davari, K. E. Jaeger, U. Schwaneberg and H. Gohlke, Comput. Struct. Biotechnol. J., 2022, 20, 399–409 CrossRef CAS PubMed.
- J. Pottkämper, P. Barthen, N. Ilmberger, U. Schwaneberg, A. Schenk, M. Schulte, N. Ignatiev and W. R. Streit, Green Chem., 2009, 11, 957–965 RSC.
- H. Cui, L. Zhang, C. B. Yildiz, L. Eltoukhy, L. Cheng, K.-E. Jaeger, U. Schwaneberg and M. D. Davari, ACS Sustainable Chem. Eng., 2022, 10, 15104–15114 CrossRef CAS.
- C. G. Acevedo-Rocha, A. Li, L. D'Amore, S. Hoebenreich, J. Sanchis, P. Lubrano, M. P. Ferla, M. Garcia-Borras, S. Osuna and M. T. Reetz, Nat. Commun., 2021, 12, 1621 Search PubMed.
- M. Wittmund, F. Cadet and M. D. Davari, ACS Catal., 2022, 12, 14243–14263 CrossRef CAS.
- N. Tokuriki and D. S. Tawfik, Curr. Opin. Struct. Biol., 2009, 19, 596–604 CrossRef CAS PubMed.
- H. Cui, H. Cao, H. Cai, K. E. Jaeger, M. D. Davari and U. Schwaneberg, Chem. – Eur. J., 2020, 26, 643–649 CrossRef CAS PubMed.
- H. Cui, K. E. Jaeger, M. D. Davari and U. Schwaneberg, Chem. – Eur. J., 2021, 27, 2789–2797 Search PubMed.
- H. Cui, L. Zhang, L. Eltoukhy, Q. Jiang, S. K. Korkunç, K.-E. Jaeger, U. Schwaneberg and M. D. Davari, ACS Catal., 2020, 10, 14847–14856 CrossRef CAS.
- X. Wang, A. Li, X. Li and H. Cui, Chem. – Eur. J., 2024, 30, e202303889 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |