Open Access Article
Open Access ArticleAssessing the diversity and functional profile of the “microbial proteome” in fermented foods
Laura
Winkler†
,
Ayesha
Awan†
 *,
Nicole M.
Rideout
and
Manuel
Kleiner
*,
Nicole M.
Rideout
and
Manuel
Kleiner
 *
*
Department of Plant and Microbial Biology, North Carolina State University, Raleigh, NC, USA. E-mail: manuel_kleiner@ncsu.edu; aawan2@ncsu.edu
First published on 26th March 2026
Abstract
Fermented foods are staples in diets worldwide and are known for their health benefits. Microorganisms are the key to fermented food production as they convert raw substrates into digestible, nutritious, and health-promoting products. While microbes are essential for fermented food production, their contribution to the dietary protein profile of the final food product in terms of microbial biomass is largely unknown. We analyzed proteins from 17 fermented foods using metaproteomics to identify and quantify microbial and food-derived proteins. We found that microbial proteins contribute up to 11% of the total protein content in fermented foods and comprise as much as 60% of the total number of identified proteins. These microbial proteins included many for central functions in microbial cells, such as glycolysis enzymes, translation machinery, and chaperones, as well as proteins for specialized functions that are important for the ecological niches in food fermentation, such as carbohydrate degrading enzymes and proteases. Some of these microbial proteins, such as proteases, could impact gut physiology. These findings highlight the substantial contribution of microbial proteins to the nutritional and functional profile of fermented foods, which may have important implications for interactions with the gut microbiota and health outcomes.
Introduction
Fermented foods have been an integral part of the human diet for more than 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 years. They are consumed globally due to their enhanced shelf life, improved sensory properties, and health benefits. They include foods and beverages that are made through microbial conversion of the raw food substrate.1–4 While originally fermentation processes were mainly used as a food preservation method and to improve the taste and texture of certain foods, fermented products account for up to 40% of global food consumption today.5,6 This includes different kinds of bread, meat, cheese, alcoholic beverages, yogurt, sauerkraut, kimchi, and miso. For instance, the average annual consumption of plain bread in Turkey is 104 kg per person, miso in Japan is 7 kg per person, and yogurt in the Netherlands is 25 L per person.5,7
000 years. They are consumed globally due to their enhanced shelf life, improved sensory properties, and health benefits. They include foods and beverages that are made through microbial conversion of the raw food substrate.1–4 While originally fermentation processes were mainly used as a food preservation method and to improve the taste and texture of certain foods, fermented products account for up to 40% of global food consumption today.5,6 This includes different kinds of bread, meat, cheese, alcoholic beverages, yogurt, sauerkraut, kimchi, and miso. For instance, the average annual consumption of plain bread in Turkey is 104 kg per person, miso in Japan is 7 kg per person, and yogurt in the Netherlands is 25 L per person.5,7
Beyond preservation and taste enhancement, fermentation improves the nutritional and functional profile of foods. It can enhance protein digestibility and micronutrient bioavailability in plant-based foods, particularly legumes, cereals, and pseudocereals.8 Proteins can also be hydrolyzed into bioactive peptides with antioxidant, antimicrobial, anti-diabetic, and anti-cancer properties.9,10 Fermented foods are also recognized as sources of probiotics and bioactive compounds, such as lactic acid and B vitamins, with potential benefits for gut health and immune modulation.1,4,11 While clinical studies link the consumption of fermented foods like yogurt and kimchi to reduced risks of cardiovascular disease, type 2 diabetes, and overall mortality, evidence supporting the effectiveness of most fermented foods for gastrointestinal health and disease is currently quite limited.12–14
One key aspect to consider when uncovering the mechanisms underlying the beneficial health effects of fermented foods is the microbial biomass, and in particular microbial protein, consumed as part of the fermented foods. When consuming fermented foods, it is often assumed that the dietary protein is mainly from the raw substrate, and microbial proteins are not given much consideration. The extent to which proteins in the raw food substrate are converted into microbial proteins, and the identities of those microbial proteins, are unknown. Microbial biomass generated during fermentation may significantly contribute to the total protein content ingested during fermented food consumption. The contribution of microbes to fermented foods is rarely quantified or characterized, and their nutritional and functional roles are underexplored.
While various -omics approaches have been applied to study the microbiome of fermented foods, metaproteomics, the large-scale study of proteins from microbial communities, remains underutilized.15–17 Liquid chromatography–tandem mass spectrometry (LC-MS/MS) has only recently gained traction in food proteomics,18 highlighting a knowledge gap about the protein composition of fermented foods, particularly the role of microbial proteins.
To determine the contribution of microbial proteins to fermented foods and estimate how much of the input substrate is converted to microbial biomass, we performed metaproteomic analyses on 17 fermented and 3 non-fermented food sources, including dairy products, sourdough bread, fermented cabbage, and cacao. These fermented foods encompass a variety of dairy and plant-based foods and were selected to represent different food substrates, microbial compositions, and fermentation processes used in commonly consumed and easily accessible fermented foods worldwide, allowing us to explore how substrate and microbial ecology influence the proteomic and functional profiles of fermented foods. The non-fermented foods analyzed in the study included milk, wheat, and tofu (soy) and represent the unfermented baseline of the corresponding fermented products. Comparing these non-fermented food substrates to their fermented counterparts allows for characterization of changes in the original food substrate's proteome as a result of microbial fermentation. We found that microbial proteins comprised a large portion of the total number of identifiable proteins in fermented foods. These microbial proteins make a substantial contribution to the total dietary protein content of these fermented foods and may possess distinct functional properties relevant to host health.
Methods
Sample acquisition
We selected fermented foods based on products commonly mentioned in the literature.19,20 The foods were purchased from different suppliers or donated by people who make specific fermented foods in their homes. We also purchased three different non-fermented food sources, which serve as negative controls (NC). The selected foods are described in Table 1 and SI Table S1. We analyzed three technical replicates of each of the fermented and non-fermented foods listed in Table 1 for this study.| Food source | Brand or homemade |
|---|---|
| Dairy yogurt | Siggi's |
| Dairy sour cream | Daisy |
| Cottage cheese | Good culture |
| Plant-based yogurt | So delicious |
| Plant-based sour cream | Forager project |
| Miso paste | Miso master® |
| Tempeh | Lightlife |
| Plain yeast bread | The essential baking company |
| Sourdough bread purchased | EuroClassic |
| Sourdough starter | Homemade |
| Sourdough bread | Homemade |
| Fermented cabbage | Homemade |
| French brie cheese | Boar's head |
| Soy sauce | Kikkoman |
| Cacao | NavitasOrganics |
| Kefir day 1 fermented | Homemade |
| Kefir day 2 fermented | Homemade |
| Tofu (NC) | House foods |
| Wheat flour (NC) | White lilly |
| Cow milk (NC) | Simple truth organic |
DNA extraction
We extracted DNA from each food sample using the DNeasy® PowerSoil® Pro kit (Qiagen GmbH, Germany) following the manufacturer's instructions with small changes. 250 mg of each sample was loaded into the provided PowerBead tubes and 800 μL of CD1 solution was added. Cell lysis and homogenization was done using the Bead Ruptor Elite (Omni International) at 6 m s−1, 00:45 min run, 00:30 min dwell, 3 cycles. For cow milk and tofu, this step was repeated 2 times, and for cacao, fermented cabbage, and wheat flour, this step was repeated 3 times. The samples were centrifuged at 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 1 min, and 200 µL of solution CD2 was added to the supernatant and mixed. Samples were centrifuged again, and 600 µL of solution CD3 was mixed with the supernatant. 650 µL of lysate was added to an MB spin column, the column was centrifuged at 15
000g for 1 min, and 200 µL of solution CD2 was added to the supernatant and mixed. Samples were centrifuged again, and 600 µL of solution CD3 was mixed with the supernatant. 650 µL of lysate was added to an MB spin column, the column was centrifuged at 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g, and the flow-through was discarded. This step was repeated until all the lysate had been applied to the column. The column was placed into a new collection tube, and 500 µL of solution EA was added, followed by centrifugation. The flow-through was discarded, 500 µL of C5 solution was added, and the column was centrifuged again. After that, the column was placed into a new collection tube and centrifuged for 2 min. Then, the column was placed into a 1.5 mL elution tube, and 100 µL of solution C6 was added to the center of the filter membrane. After centrifugation for 1 min, the eluted DNA was ready for downstream processing. The DNA quantity and quality were measured using the DeNovix® DS-11 FX + Spectrophotometer/Fluorometer (DeNovix Inc.). Extracted DNA was stored at −20 °C.
000g, and the flow-through was discarded. This step was repeated until all the lysate had been applied to the column. The column was placed into a new collection tube, and 500 µL of solution EA was added, followed by centrifugation. The flow-through was discarded, 500 µL of C5 solution was added, and the column was centrifuged again. After that, the column was placed into a new collection tube and centrifuged for 2 min. Then, the column was placed into a 1.5 mL elution tube, and 100 µL of solution C6 was added to the center of the filter membrane. After centrifugation for 1 min, the eluted DNA was ready for downstream processing. The DNA quantity and quality were measured using the DeNovix® DS-11 FX + Spectrophotometer/Fluorometer (DeNovix Inc.). Extracted DNA was stored at −20 °C.
Library preparation and sequencing methods
Sequencing was done by CosmosID (CosmosID Inc.). DNA libraries were prepared using the Watchmaker DNA Library Prep Kit (7K0019-1K) and compatible Twist Universal Adapter System. Genomic DNA was fragmented using a mastermix of Watchmaker Frag/AT Buffer and Frag/AT Enzyme Mix. Twist Universal Adapters (10×) and Twist HT Unique Dual Indexes (2×) were added to each sample, followed by 7 cycles of PCR to construct the DNA libraries. The final DNA libraries were purified using AMPure magnetic beads (Beckman Coulter) and eluted in nuclease-free water. Following elution, the libraries were quantified using the Qubit™ fluorometer dsDNA HS Assay Kit. Libraries were then circularized using the Element Adept library compatibility workflow and sequenced on the Element AVITI platform using the AVITI 2 × 150 Cloudbreak sequencing kit.Post sequencing data analysis
We analyzed the quality of the sequencing reads using FastQC version 0.12.1,21 and determined the base quality scores, GC content, and adapter contamination. We then merged forward and reverse reads using the reformat.sh function in BBTools version 39.01 using the default overlap and mismatch parameters.22 Adapters and low-quality sequences were trimmed, and contaminant phiX174 sequences were removed using Trim Galore version 0.6.10.23 We used phyloflash version 3.4.2 to map reads to 16S rRNA gene sequences and determine which taxa were present in the samples.24Sample preparation and protein extraction
We lyophilized all liquid food samples, including cow milk, soy sauce, and kefir, to concentrate them. We weighed 150 mg of wet sample (such as yogurt) and 100 mg of dry sample (such as bread) into 2 mL Lysing Matrix E (MP Biomedicals™) tubes and added 1 mL of SDS-lysis buffer (5% w/v) in 50 mM TEAB, pH 7.5. We lysed samples using a Bead Ruptor Elite (Omni International) at the following settings: 6.45 m s−1, 00:45 min run, 01:00 min dwell, 5 cycles. Subsequently, the samples were heated for 10 min at 95 °C and then centrifuged for 5 min at 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g, and the supernatant was retained.
000g, and the supernatant was retained.
To extract proteins and prepare peptides, we followed the Suspension Trapping (S-Trap) sample preparation method for bottom-up proteomics analysis.25 We mixed 6.4 µL of 500 mM DTT with the entire lysate and incubated for 10 min at 95 °C. Then, we added 12.7 µL of 500 mM IAA solution and incubated in a centrifuge tube shaker at room temperature (RT) and 600 rpm for 1 min. The lysates were then further incubated in the dark for 30 min. After the incubation, 16 µL of 12%-phosphoric acid solution and 6 times the sample volume of binding/wash buffer (100 mM TEAB in 90% MetOH) was added.
For the next steps, the flow-through was discarded after every centrifugation step. 1200 µL of sample was loaded on the S-TRAP-mini-80 column (PROTIFI™) and centrifuged at 4000g for 30 s. The column was washed with 400 µL of wash/binding buffer and centrifuged at 4000g for 30 s. This step was repeated 3 times in total. Without adding more buffer, the column was centrifuged one more time at 4000g for 30 s. The column was then placed in a new 2 mL sample tube, and proteins were digested by adding 0.8 µg MS-grade trypsin (Thermo Scientific Pierce™, Rockford, IL, USA), solubilized in 40 µL digestion/elution buffer 1 (50 mM TEAB in HPLC water) to the column, and incubating it overnight at 37 °C in a wet chamber. The next day, peptides were eluted off the column by first adding 80 µL of elution buffer 1 to the column, incubating for 10 min at 37 °C, and centrifuging at 4000g for 1 min. Then 80 µL of elution buffer 2 (0.2% formic acid in HPLC water) was added and centrifuged at 4000g for 1 min. Finally, 80 µL of elution buffer 3 (50% acetonitrile and 0.2% formic acid in HPLC water) was added and centrifuged at 4000g for 1 min. Acetonitrile from eluted peptides was removed using a vacuum centrifuge, and the peptides were resuspended in ∼30 µL elution buffer 2. Peptide concentrations were determined using the Micro BCA™ Protein Assay Kit (Thermo Scientific) according to the manufacturer's instructions.
LC-MS/MS analysis
We analyzed peptides from food samples using 1D-LC-MS/MS as previously described.26,27 We loaded 1 μg of peptides onto a 5 mm, 300 μm ID C18 556 Acclaim® PepMap100 pre-column (Thermo Fisher Scientific) with loading solvent A (2% acetonitrile, 0.05% TFA) using an UltiMate™ 3000 RSLCnano Liquid Chromatograph (Thermo 558 Fisher Scientific). An EASY-Spray analytical column heated to 60 °C (PepMap RSLC C18, 2 μm material, 75 cm × 75 μm, Thermo Fisher Scientific) was used to separate the peptides. A 140 min gradient at a flow rate of 300 nl min−1 was used for peptide separation of which the first 102 minutes of the gradient went from 95% eluent A (0.1% formic acid) to 31% eluent B (0.1% formic acid, 80% acetonitrile), then 18 min from 31 to 50% B, and 20 min at 99% B. One to two 100% acetonitrile wash runs were performed between each sample to minimize carryover. The eluted peptides were ionized using an Easy-Spray source and analyzed in an Exploris 480 hybrid quadrupole-Orbitrap mass spectrometer (Thermo Fisher Scientific) with the following parameters: m/z 445.12003 lock mass, normalized collision energy 24, 25 s dynamic exclusion, and exclusion of ions of +1 charge state. Full MS scans were acquired for 380 to 1600 m/z at a resolution of 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 and a maximum IT time of 200 ms. Data-dependent MS2 was performed for the 15 most abundant ions at a resolution of 15 000 and maximum IT of 100 ms.
000 and a maximum IT time of 200 ms. Data-dependent MS2 was performed for the 15 most abundant ions at a resolution of 15 000 and maximum IT of 100 ms.
Database construction and protein identification
For each of the fermented foods, we constructed non-redundant protein sequence databases following the previously described principles.28 These databases comprised annotated protein sequences from the Proteomes section of UniProt corresponding to the bacterial and eukaryotic ingredients present in each food source. The database for each food source is unique, and the species to include in the database were selected based on the current literature, the ingredients list of the food, and our taxonomic analysis of the metagenomic data. The database composition for each food source is detailed in SI Table S2. In order to remove redundant sequences, the protein sequences of each strain/species were clustered using cd-hit (Version 4.7) with an identity threshold of 95%.29We searched the MS2 spectra against the respective food-source-specific database using the PEAKS® X + software and quantified proteins using area under the curve30–32 with the following settings: precursor correction: min charge = 1, max charge = 10; precursor mass error tolerance = 10 ppm using monoisotopic mass; fragment ion error tolerance = 0.05 Da; enzyme = trypsin, semispecific; maximum missed cleavages = 2; maximum allowed variable PTMs per peptide = 3, including carbamidomethylation, oxidation (M) and deamidation (NQ). Advanced options: DeNovo Sequencing, FDR decoy-fusion, identification of unspecified PTMs with PEAKS PTM, and identification of more mutations with PEAKS SPIDER. For the downstream analysis, protein tables were filtered for an FDR of 5%, proteins −10 lg![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P ≥ 15, and ≥2 unique peptides without significant peptides, and a DeNovo only score of 50% was used. The raw data are provided in SI Table S4.
P ≥ 15, and ≥2 unique peptides without significant peptides, and a DeNovo only score of 50% was used. The raw data are provided in SI Table S4.
Data processing, statistical analysis, and data visualization
We used total sum scaling to (1) normalize the relative abundances of proteins within each sample33 and to (2) normalize at the organism level to estimate the relative abundances of proteins within each organism in a sample. We calculated the abundance of each microbial species using its proteinaceous biomass.34 We did this by summing up the peptide spectra of all proteins with at least 2 protein unique peptides assigned to each given species. GraphPad Prism (Version 10.2.3), Microsoft Excel, and ggplot235 in R (version 2024.12.1)36 were used for data processing and visualization. The pheatmap package in R was used to make the heatmap.37Results
Microbial proteins comprise a substantial portion of the total protein content in fermented foods
We used metaproteomics to analyze the protein content of 17 different fermented foods and 3 non-fermented foods (Fig. 1). The total number of identified proteins in each food source ranged from 680 total proteins in dairy sour cream to beyond 3000 total proteins in homemade sourdough bread (Fig. 1A). The number of identified microbial proteins ranged from 119 proteins in Miso to over 1000 different proteins in Brie cheese. The relative abundance of all microbial proteins out of the total detected proteins ranged from 0.6% in cacao to more than 11% in plain yeast bread (Fig. 1B). | ||
| Fig. 1 Microbial proteins contribute a large amount of dietary protein to fermented foods. A. Contribution of the different identified microbial proteins to the overall number of identified proteins (mean and standard deviation are shown; 3 technical replicates were analyzed). The percentage above each food source indicates the proportion of identified microbial proteins relative to the total number of proteins detected. B. Relative abundance of microbial and non-microbial (food substrate) protein in different fermented food sources (n = 3). Cow milk, wheat flour, and tofu served as unfermented negative controls. Peptides were identified using the PEAKS software, allowing for semi-tryptic peptides. Relative proteinaceous biomass was determined as described in Kleiner et al. (2017) for each sample, and the results were averaged by food.34 | ||
The number of microbial proteins identified in a fermented food depends on at least two factors. First, the overall abundance of microbial protein in the food, because detection of proteins by metaproteomics only works if their relative abundance in the sample is above the detection limit. If the fermenting microorganisms did not grow to an abundance where they make up a relevant amount of the protein in the food, their proteins are hard to detect. Hence, the measured relative abundance of detected proteins is directly related to the abundance of a microbial species in the food. Second, the number of different microbial taxa present in the food, because each taxon contributes additional differentiable proteins to the food. This means that if the fermenting microbial community has more microbial taxa, more microbial proteins may be detected. Some fermented foods, such as soy sauce and cocoa, had little to no microbial proteins detected, likely due to a low abundance of microorganisms and thus microbial proteins in these food types.
The proportion and diversity of microbial proteins were much higher than the food substrate proteins in 5 out of the 16 fermented foods analyzed. In Brie cheese, for example, out of the 1573 different proteins present, 1023 of these (65%) were microbial proteins. This pattern was observed in almost all dairy products, especially in dairy yogurt, dairy sour cream, and dairy cottage cheese, with the exception of kefir, in which microbial proteins only made up 23–27% of the total number of proteins. In plant-based dairy alternative fermented foods, the number of microbial proteins ranged from 16% of all proteins in plant-based sour cream to 43% in plant-based yogurt. In the bread samples, microbial proteins contributed 17–19% to the overall protein number. The proportion of microbial proteins by number was lower in miso (9%) and fermented cabbage (2%). In contrast, 32–57% of the overall number of proteins in tempeh and soy sauce were of microbial origin.
Looking at total protein biomass regardless of protein identity, the percentage of each sample that could be attributed to microbial protein biomass varied between different fermented foods, with 0.5–2.5% in dairy products, 0.8–2% in plant-based dairy alternatives, 0.5–1.2% in soy based products, 5% in cabbage, and 5–11% in bread products. These results show that microbial proteins contribute a large proportion of the protein, by both number of different proteins and abundance relative to non-microbial proteins, to the overall protein content of fermented foods.
Fermented foods harbor distinct, food-specific microbial communities
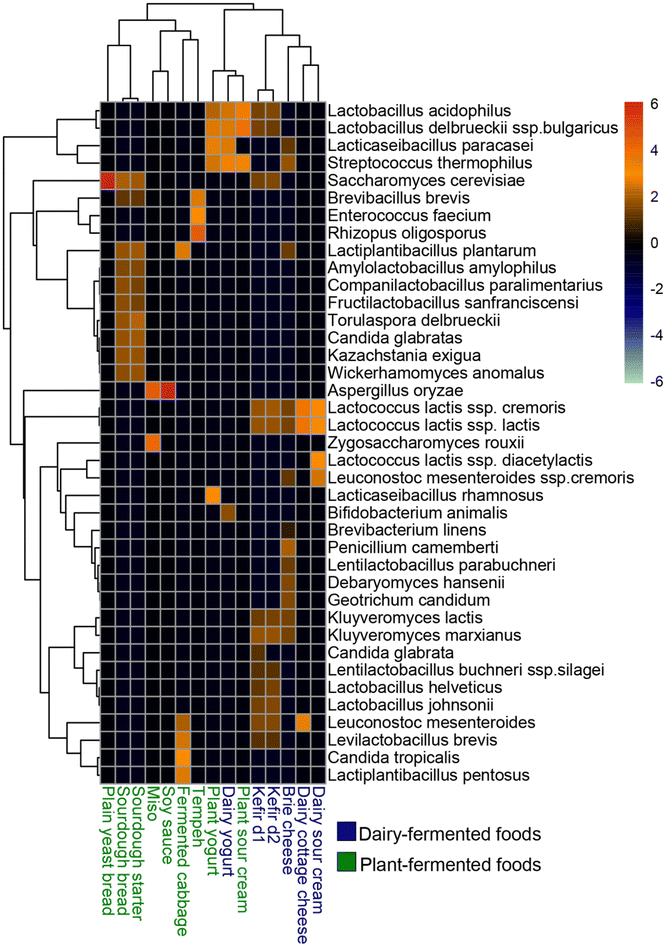
As expected, microbial community composition differed strongly between dairy and plant-based fermented foods, as well as between different dairy products such as yogurt and cheese (Fig. 2). Among all the fermented foods analyzed, brie cheese and kefir had the highest number of microbial species, while plain yeast bread and soy sauce had the lowest. While both dairy and plant yogurts contained Streptococcus thermophilus, Lacticaseibacillus paracasei, Lactobacillus delbrueckii ssp. bulgaricus and Lactobacillus acidophilus, only dairy yogurt contained Bifidobacterium animalis, and only plant yogurt contained Lacticaseibacillus rhamnosus. Dairy and plant sour creams had distinct microbial compositions, and dairy sour cream clustered with dairy cottage cheese, while plant sour cream clustered with plant and dairy yogurts. Dairy sour cream contained Lactococcus lactis strains and Leuconostoc mesenteroides, while plant sour cream contained L. acidophilus, L. delbrueckii, and S. thermophilus.Although both dairy-based fermented foods and their plant alternatives mainly contained Bacillota species, the specific taxa differed. Brie cheese and kefir had the greatest diversity of microbes compared to other dairy fermented foods and contained both bacterial and fungal phyla. Bacterial taxa in brie cheese included the Bacillota species L. lactis ssp. lactis, L. lactis ssp. cremoris, and S. thermophilus, as well as Actinobacteria species Brevibacterium linens. Fungal taxa in brie cheese included Ascomycota species Kluyveromyces lactis, Debaryomyces hansenii, Penicillium camemberti, and Geotrichum candidum.
Kefir, a fermented milk product, contained the fungal species Candida glabrata, Kluyveromyces marxianus, K. lactis, and Saccharomyces cerevisiae as well as the bacterial species L. acidophilus, Lactobacillus johnsonii, Lactobacillus helveticus, Lentilactobacillus buchneri, L. mesenteroides, L. delbrueckii, and L. lactis. Kefir fermented for 2 days, compared to 1 day of fermentation, had a higher abundance of L. johnsonii and L. helveticus and a lower abundance of C. glabrata. Legume (soy) based fermented foods like tempeh, soy, and miso were dominated by fungal species, including Rhizopus oligosporus in tempeh, Zygosaccharomyces rouxii in miso, and Aspergillus oryzae in both miso and soy sauce. Tempeh also contained the bacterial species Enterococcus faecium and Brevibacillus brevis.
Fermented cereal-based foods, including plain yeast bread and sourdough bread, both contained S. cerevisiae. However, sourdough contained a greater diversity of both Ascomycota fungal species, including Wickerhamomyces anomalus, C. glabrata, Torulaspora delbrueckii, Kazachstania exigua, and Bacillota bacterial species, including Lactiplantibacillus plantarum, Amylolactobacillus amylophilus, Companilactobacillus paralimentarius, Fructilactobacillus sanfranciscensis, and B. brevis. Fermented cabbage or sauerkraut, another plant-fermented food, also contained fungal and bacterial species, including Candida tropicalis, Lactiplantibacillus pentosus, L. plantarum, and Levilactobacillus brevis.
The overall microbial community composition was substrate-specific, with both dairy and plant-fermented foods dominated by Bacillota and Ascomycota species. L. delbrueckii ssp. bulgaricus, L. lactis, and L. acidophilus were specific to mainly dairy fermented products, and Lactiplantibacillus species were associated with plant substrates containing complex carbohydrates. Fungal species were mainly associated with legume and cereal-based fermented foods.
Distinct profiles of microbial and non-microbial proteins across various fermented foods
We quantified the relative abundances of microbial and food substrate derived non-microbial proteins across 14 dairy, plant, grain, and legume based fermented foods, as well as their unfermented food substrates (Fig. 3). In most of the fermented food samples, the diversity of microbial proteins was greater than that of non-microbial food substrate dietary proteins. Comparison with the unfermented food counterparts revealed that microbial fermentation significantly alters the protein composition of these foods.In unfermented dairy milk alpha-S2-casein, alpha-S1-casein, and alpha-lactalbumin were the most abundant non-microbial proteins. In fermented dairy products such as yogurt and cottage cheese, alpha-S1-casein, alpha-S2-casein, kappa casein, and alpha lactalbumin were the most abundant cow-derived proteins, while beta-casein and beta lactalbumin were higher in abundance in kefir and sour cream. Brie cheese had a high abundance of alpha-S2-casein and kappa casein.
Microbial proteins in dairy fermented foods varied by product type. Yogurt had a high abundance of chaperone proteins, glyceraldehyde-3-phosphate dehydrogenase (GAPDH), and enolase. Cottage cheese and sour cream had a high abundance of elongation factor Tu, DNA-binding protein HU, enolases and tagatose-1,6-diphosphate aldolase. Kefir had a high abundance of GAPDH, elongation factor Tu, and enolase. Brie cheese had a high abundance of mainly fungal proteins, including peptide hydrolase and dipeptidyl-peptidase V.
Plant-based alternatives to dairy fermented foods had distinct profiles of non-microbial food substrate proteins compared to their dairy counterparts, including high abundance of cocosin 1, vicillin-like antimicrobial peptides, glutelin type A1, and cupin domain containing protein. However, the microbial proteins, including GAPDH, enolase, DNA-binding proteins, chaperones, and elongation factor Tu were the most abundant microbial proteins in plant-based and dairy yogurts and sour creams.
Unfermented wheat had a high abundance of seed storage proteins (e.g., 7.14% abundance of plant lipid transfer/seed storage protein), glutenin (10% summed abundance of all subunits), and a variety of amylase and trypsin inhibitors (>20% summed relative abundance), whereas its fermented counterparts, such as yeast bread and sourdough, maintained a high abundance of seed storage proteins (14% abundance of plant lipid transfer/seed storage protein) but had a lower abundance of amylase and trypsin inhibitors (∼14% summed relative abundance). Grain-based fermented foods, plain yeast bread, and sourdough had different microbial protein profiles reflective of differences in their microbial community and diversity (Fig. 2). While both breads had abundant yeast-derived proteins with high abundance of GAPDH and enolase, sourdough had a much higher diversity of low-abundance bacterial proteins.
Similarly, unfermented soy-based tofu had a high abundance of seed storage proteins, including glycinin and beta-conglycinin, while fermented soy based products, including tempeh and miso, had reduced abundance of these seed storage proteins and increased abundances of protease inhibitors, including the kunitz trypsin inhibitor and the BBI serine inhibitor (SI Table S4). In terms of microbial proteins, soy-based fermented food products had a high abundance of microbial carbohydrate degradation enzymes and proteases. Tempeh had a high abundance of microbial phosphoenolpyruvate carboxykinase (PEPCK), elongation factor 1-A, ATP synthase, and malate dehydrogenase. Miso had a high abundance of microbial glucan-1,4-alpha-glucosidase, alpha amylase, aspergillopepsin-1, and neutral protease 2.
Overall, these results indicate significant shifts in protein composition between fermented foods and their unfermented counterparts. Unfermented food substrate proteins, such as caseins, glutelins, vicilins, and glycinin, are reduced in relative abundance as they are degraded by microbes and used to produce new microbial proteins.
Discussion
While it is known that fermented foods positively impact health outcomes, the underlying factors driving these effects are not well characterized.38–40 Thus, it is crucial to characterize the distinct microbial and non-microbial components of fermented foods to better understand how they might interact with the host and the gut microbiota upon consumption. While many different fermented foods and beverages from all over the world have been investigated using different -omics approaches, including metagenomics, metatranscriptomics, and metabolomics, the metaproteomic approach remains underutilized despite the fact that proteins are key nutritional and functional components of these foods.41–43In this study, we used metaproteomics to study fermented foods on the protein level, infer specific microbial and non-microbial proteins, and characterize their functional profiles. Our metaproteomics results show that microbial proteins comprise a substantial proportion of total protein content in fermented foods. This shows that microorganisms not only contribute to the fermentation process itself but also to the overall nutritional profile of fermented food through the conversion of substrate protein into microbial protein. These results offer avenues for investigation beyond the known probiotic effects of fermented foods, as the microbial proteins consumed as part of these foods may have direct nutritional and immune-modulatory effects. The large microbial protein diversity present in these foods also translates to a broader functional potential, including carbohydrate-degrading enzymes, proteases, bioactive peptides and other compounds that can influence the host's digestion and interact with the gut microbiota in the colon.44,45
As expected, the most abundant microbial proteins found in fermented foods were associated with central metabolism and core physiological functions; these proteins included glycolysis proteins, translation factors, chaperones, carbohydrate degrading enzymes, and proteases.41,42 In dairy fermented foods, which are dominated by bacteria, microbial proteins involved in protein biosynthesis, carbohydrate metabolism, and energy production were most abundant. In contrast, in plant-based fermented foods that were mostly fermented by fungi, such as miso, soy sauce, and tempeh, carbohydrate-degrading enzymes (CAZymes) and proteases were more abundant.46
The presence of microbial enzymes such as CAZymes and proteases is noteworthy as it points to substrate-specific adaptations that can impact gut health. CAZymes are crucial for the breakdown of carbohydrates and can support the breakdown of dietary fiber, increasing the accessibility of fermentable carbohydrates to the gut microbiota.47–49 This indicates that fungi found in fermented foods, such as A. oryzae (soy sauce and miso) and R. oligosporus (tempeh), may play a role in carbohydrate accessibility to other microbes in the gut, potentially promoting short-chain fatty acid (SCFA) production, gut barrier integrity, and anti-inflammatory effects. Microbial proteases, found in brie cheese, soy sauce, miso, and tempeh, can act on substrates such as soy proteins to generate bioactive peptides that can affect health, including impacting angiotensin–converting enzyme (ACE) activity, nutrient processing and absorption, and modulating immune and gut barrier responses.50–52 Hence, microbial proteins in fermented foods can contribute functional molecules that, beyond basic nutrition, can interact with the gut microbiota and impact host physiology. While our results did not focus on profiling the functional activity of these health-relevant microbial proteins, future studies should investigate their activities and how they might interact with the host and other gut microbes.
In addition to increasing microbial protein biomass in fermented foods, microbial fermentation also altered the nutritional landscape of foods by modifying the abundance of functional and health-relevant food substrate proteins through metabolic conversion. These changes include degrading and altering the abundances of anti-nutritional protease inhibitors, such as alpha-amylase/trypsin inhibitors in wheat, Kunitz trypsin and BBI inhibitors in soy, as well as of potential food allergens and immunogenic proteins, including glutelin, vicillin, and casein proteins.8,53,54 These findings have important implications in the context of previous studies showing that the detection and in vitro IgE reactivity of known food allergens is altered post microbial fermentation and highlight the potential for engineering the fermentation process to enhance microbial proteolytic activity against anti-nutritional factors or allergenic food proteins.53–55
Furthermore, the observation that different food types are dominated by different microbial taxa suggests that fermented foods carry a substrate-specific core functional microbiome. The identified taxa in the dairy products and plant-based products in this study aligned with findings of previous studies that used DNA-based approaches to profile microbes. Dairy fermented foods such as brie, kefir, and yogurt contained bacterial species including Lactococcus lactis, Streptococcus thermophilus, Lacticaseibacillus paracasei, and Lactobacillus delbrueckii, often alongside fungi like Kluyveromyces lactis and Candida glabrata, while plant-based fermented foods such as tempeh, miso, soy sauce, sourdough, and sauerkraut were enriched in Lactiplantibacillus species adapted to complex carbohydrate substrates and fungi like Rhizopus oligosporus, Aspergillus oryzae, and Zygosaccharomyces rouxii.56,57
The sourdough microbiome has been extensively studied, revealing that sourdough contains a large diversity of microorganisms, especially yeasts.58–60 We included the strains identified in those previous studies in the protein database for this study, and our results based on proteinaceous biomass differed slightly. We found that S. cerevisiae was the dominant fungus, followed by C. glabrata, K. exigua, T. delbrueckii, and W. anomalus, in line with previous studies.61–63 However, the most abundant lactobacillus bacteria in our sourdough were L. plantarum (9.7%), followed by F. sanfranciscensis (1.7%), while other studies have reported the reverse trend in the abundance patterns of these two bacteria using a 16S rRNA gene and ITS amplicon sequencing approach.58 This difference between our results and prior studies is likely due to the fact that metaproteomics measures the proteinaceous mass contribution of a species to a community, while DNA sequencing-based approaches provide insights into gene copy numbers contributed by species. This suggests that using proteinaceous biomass instead of DNA-based measures may be more representative of the quantitative profile of the microbial community in fermented foods, as well as its contribution to the nutritional and functional profile of the food.34
A limitation of this study is that fermented foods, especially the homemade kind, can lack a standardized manufacturing process, which can introduce variability based on ingredients, microbial inocula, and fermentation conditions used. This inherent variability, in addition to the fact that we only used one product or homemade batch per fermented food type, limits our ability to make generalizing statements about the microbial community composition in specific fermented foods. However, we do think that the trends we found when comparing microbial community composition across different fermented foods provide qualitative insights into overall differences in microbial community composition between fermented food types. Future studies could investigate variability in the microbial proteome of specific fermented food types using multiple products or homemade batches of the same fermented food type.
Overall, this study underscores the importance of considering microbial proteins as a key component of the nutritional and functional value of fermented foods. While the direct effects of some fungal and bacterial proteins and enzymes on human health remain speculative due to digestive degradation, their influence on the immunogenicity or functionality of other proteins warrants further investigation beyond the scope of this study. Our use of metaproteomics has laid the groundwork for such future research into the potential health-promoting effects of microbial proteins in fermented food products.
Conflicts of interest
The authors declare no competing interests.Author contributions
Laura Winkler: experimental design, data collection, data analysis, writing, and editing the manuscript; Ayesha Awan: conceptualization of the study, experimental design, data collection, data analysis, writing, and editing the manuscript; Nicole Rideout: experimental design, data collection, data analysis, editing; Manuel Kleiner: funding, conceptualization of the study, experimental design, data processing, writing, and editing the manuscript.Data availability
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD070912.64 The metagenomic raw reads were submitted to NCBI SRA under the bioproject identifier PRJNA1365831.Supplementary information (SI): Table S1: Details of each of the commercially obtained fermented foods, including source, label specified food substrate and microbes; Table S2: Table listing each of the microbial and non-microbial species used to construct the protein database that was used in the proteomics analysis; Table S3: The relative abundance of the proteomic biomass of each microbial species in the fermented foods analyzed; Table S4: The raw protein abundances identified in each of the three replicates of the fermented foods analyzed in the proteomic analysis. See DOI: https://doi.org/10.1039/d5fo05039a.
Acknowledgements
We would like to thank Jessie Maier for providing the homemade sourdough starter and bread, Sadije Jakupi for providing the fermented cabbage, and Sandra Kelberlau for providing the kefir. We would also like to thank Dr Heather Maughan for editing the manuscript and for her insightful suggestions. All LC-MS/MS measurements were made in the Molecular Education, Technology, and Research Innovation Center (METRIC) at North Carolina State University. This work was supported by the National Institutes of Health under Awards R35GM138362 and R01DK118024, and the European Union under Award Number 2023 1 DE01 KA131 HED 000120145.References
- M. L. Marco, D. Heeney, S. Binda, C. J. Cifelli, P. D. Cotter and B. Foligné, et al., Health benefits of fermented foods: microbiota and beyond, Curr. Opin. Biotechnol., 2017, 44, 94–102 CrossRef CAS PubMed.
- H. Sahm, G. Antranikian, K. P. Stahmann and R. Takors, Industrielle Mikrobiologie, Springer Berlin Heidelberg Imprint Springer Spektrum, Berlin, Heidelberg, 1st edn, 2013 Search PubMed.
- J. P. Tamang, P. D. Cotter, A. Endo, N. S. Han, R. Kort and S. Q. Liu, et al., Fermented foods in a global age: East meets West, Compr. Rev. Food Sci. Food Saf., 2020, 19(1), 184–217 CrossRef PubMed.
- Industrial microbiology, ed. D. B. Wilson, H. Sahm, K. P. Stahmann and M. Koffas, Wiley-VCH, Weinheim, 2020, p. 397 Search PubMed.
- L. Cuamatzin-García, P. Rodríguez-Rugarcía, E. G. El-Kassis, G. Galicia, M. D. L. Meza-Jiménez and B.-L. MaDR, et al., Traditional Fermented Foods and Beverages from around the World and Their Health Benefits, Microorganisms, 2022, 10(6), 1151 CrossRef PubMed.
- E. Dimidi, S. Cox, M. Rossi and K. Whelan, Fermented Foods: Definitions and Characteristics, Impact on the Gut Microbiota and Effects on Gastrointestinal Health and Disease, Nutrients, 2019, 11(8), 1806 CrossRef CAS PubMed.
- S. Chilton, J. Burton and G. Reid, Inclusion of Fermented Foods in Food Guides around the World, Nutrients, 2015, 7(1), 390–404 CrossRef PubMed.
- A. Kårlund, C. Gómez-Gallego, J. Korhonen, O. M. Palo-oja, H. El-Nezami and M. Kolehmainen, Harnessing Microbes for Sustainable Development: Food Fermentation as a Tool for Improving the Nutritional Quality of Alternative Protein Sources, Nutrients, 2020, 12(4), 1020 CrossRef PubMed.
- Q. Guo, P. Chen and X. Chen, Bioactive peptides derived from fermented foods: Preparation and biological activities, J. Funct. Foods, 2023, 101, 105422 CrossRef CAS.
- L. Peres Fabbri, A. Cavallero, F. Vidotto and M. Gabriele, Bioactive Peptides from Fermented Foods: Production Approaches, Sources, and Potential Health Benefits, Foods, 2024, 13(21), 3369 CrossRef CAS PubMed.
- Biochemical, physiological, and molecular aspects of human nutrition, ed. M. H. Stipanuk and M. A. Caudill, Elsevier, St. Louis, Missouri, 4th edn, 2019, p. 959 Search PubMed.
- M. Chen, Q. Sun, E. Giovannucci, D. Mozaffarian, J. E. Manson and W. C. Willett, et al., Dairy consumption and risk of type 2 diabetes: 3 cohorts of US adults and an updated meta-analysis, BMC Med., 2014, 12(1), 215 CrossRef PubMed.
- S. S. Soedamah-Muthu, G. Masset, L. Verberne, J. M. Geleijnse and E. J. Brunner, Consumption of dairy products and associations with incident diabetes, CHD and mortality in the Whitehall II study, Br. J. Nutr., 2013, 109(4), 718–726 CrossRef CAS.
- L. C. Tapsell, Fermented dairy food and CVD risk, Br. J. Nutr., 2015, 113(S2), S131–S135 CrossRef CAS PubMed.
- O. Nikoloudaki, F. Aheto, R. Di Cagno and M. Gobbetti, Synthetic microbial communities: A gateway to understanding resistance, resilience, and functionality in spontaneously fermented food microbiomes, Food Res. Int., 2024, 192, 114780 CrossRef CAS PubMed.
- M. Kleiner, Metaproteomics: Much More than Measuring Gene Expression in Microbial Communities, mSystems, 2019, 4(3), e00115–e00119 CrossRef CAS.
- T. Van Den Bossche, J. Armengaud, D. Benndorf, J. A. Blakeley-Ruiz, M. Brauer and K. Cheng, et al., The microbiologist's guide to metaproteomics, iMeta, 2025, 4(3), e70031 CrossRef CAS PubMed.
- C. Aydoğan, Critical review of new advances in food and plant proteomics analyses by nano-LC/MS towards advanced foodomics, TrAC, Trends Anal. Chem., 2024, 176, 117759 CrossRef.
- Handbook of food and beverage fermentation technology, ed. Y. H. Hui, Marcel Dekker, New York, 2004, p. 148. (Food science and technology) Search PubMed.
- Handbook of Animal-Based Fermented Food and Beverage Technology, ed. Y. H. Hui and E. Ö. Evranuz, CRC Press, Boca Raton, 2nd edn, 2012 Search PubMed.
- S. Andrews, FastQC: A Quality Control Tool for High Throughput Sequence Data.
- B. Bushnell, BBTools software package, 2014 Search PubMed.
- F. Krueger, F. James, P. Ewels, E. Afyounian, M. Weinstein and B. Schuster-Boeckler, et al., FelixKrueger/TrimGalore: v0.6.10 - add default decompression path, Zenodo, 2023 [cited 2026 Feb 3]. Available from: https://zenodo.org/record/5127898 Search PubMed.
- H. R. Gruber-Vodicka, B. K. B. Seah and E. Pruesse, phyloFlash: Rapid Small-Subunit rRNA Profiling and Targeted Assembly from Metagenomes, mSystems, 2020, 5(5), e00920–20 CrossRef CAS PubMed.
- A. Zougman, P. J. Selby and R. E. Banks, Suspension trapping (STrap) sample preparation method for bottom–up proteomics analysis, Proteomics, 2014, 14(9), 1006–1000 CrossRef CAS PubMed.
- A. Awan, A. Bartlett, J. A. Blakeley-Ruiz, T. Richie, C. M. Theriot and M. Kleiner, Dietary protein from different sources escapes host digestion and is differentially modified by gut microbiota, Food Funct., 2025, 16(18), 7154–7168, 10.1039/D5FO01132A.
- A. Mordant and M. Kleiner, Evaluation of Sample Preservation and Storage Methods for Metaproteomics Analysis of Intestinal Microbiomes, Microbiol. Spectr., 2021, 9(3), e01877–e01821, DOI:10.1128/Spectrum.01877-21.
- J. A. Blakeley-Ruiz and M. Kleiner, Considerations for constructing a protein sequence database for metaproteomics, Comput. Struct. Biotechnol. J., 2022, 20, 937–952 CrossRef CAS PubMed.
- W. Li and A. Godzik, Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences, Bioinformatics, 2006, 22(13), 1658–1659 CrossRef CAS PubMed.
- PEAKS® Studio X+, Bioinformatics solutions Inc., 2019.
- R. Vitorino, S. Guedes, F. Trindade, I. Correia, G. Moura and P. Carvalho, et al., De novo sequencing of proteins by mass spectrometry, Expert Rev. Proteomics, 2020, 17(7–8), 595–607 CrossRef CAS PubMed.
- J. Zhang, L. Xin, B. Shan, W. Chen, M. Xie and D. Yuen, et al., PEAKS DB: De Novo Sequencing Assisted Database Search for Sensitive and Accurate Peptide Identification, Mol. Cell. Proteomics, 2012, 11(4), M111.010587 CrossRef PubMed.
- M. A. Dillies, A. Rau, J. Aubert, C. Hennequet-Antier, M. Jeanmougin and N. Servant, et al., A comprehensive evaluation of normalization methods for Illumina high-throughput RNA sequencing data analysis, Briefings Bioinf., 2013, 14(6), 671–683 CrossRef CAS PubMed.
- M. Kleiner, E. Thorson, C. E. Sharp, X. Dong, D. Liu and C. Li, et al., Assessing species biomass contributions in microbial communities via metaproteomics, Nat. Commun., 2017, 8(1), 1558 CrossRef PubMed.
- H. Wickham, ggplot2: elegant graphics for data analysis, Springer international publishing, Cham, 2nd edn, 2016, p. 1. (Use R!) Search PubMed.
- R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria, 2023; Available from: https://www.R-project.org/ Search PubMed.
- R. Kolde, Pheatmap: pretty heatmapsR package version 201912726 Search PubMed.
- A. Mukherjee, S. Breselge, E. Dimidi, M. L. Marco and P. D. Cotter, Fermented foods and gastrointestinal health: underlying mechanisms, Nat. Rev. Gastroenterol. Hepatol., 2024, 21(4), 248–266 CrossRef CAS PubMed.
- V. Valentino, R. Magliulo, D. Farsi, P. D. Cotter, O. O'Sullivan and D. Ercolini, et al., Fermented foods, their microbiome and its potential in boosting human health, Microb. Biotechnol., 2024, 17(2), e14428 CrossRef CAS PubMed.
- I. Park and M. Mannaa, Fermented Foods as Functional Systems: Microbial Communities and Metabolites Influencing Gut Health and Systemic Outcomes, Foods, 2025, 14(13), 2292 CrossRef CAS PubMed.
- C. Aydoğan, Critical review of new advances in food and plant proteomics analyses by nano-LC/MS towards advanced foodomics, TrAC, Trends Anal. Chem., 2024, 117759 CrossRef.
- E. S. Okeke, R. E. Ita, E. J. Egong, L. E. Udofia, C. L. Mgbechidinma and O. D. Akan, Metaproteomics insights into fermented fish and vegetable products and associated microbes, Food Chem. Mol. Sci., 2021, 3, 100045 CrossRef CAS PubMed.
- L. Yang, W. Fan and Y. Xu, Metaproteomics insights into traditional fermented foods and beverages, Compr. Rev. Food Sci. Food Saf., 2020, 19(5), 2506–2529 CrossRef PubMed.
- C. Y. E. Tachie, J. O. Onuh and A. N. A. Aryee, Nutritional and potential health benefits of fermented food proteins, J. Sci. Food Agric., 2024, 104(3), 1223–1233 CrossRef CAS PubMed.
- J. P. Tamang, D. H. Shin, S. J. Jung and S. W. Chae, Functional Properties of Microorganisms in Fermented Foods, Front. Microbiol., 2016, 7, 578 Search PubMed . Available from:: https://journal.frontiersin.org/Article/10.3389/fmicb.2016.00578/abstract.
- P. V. Gavande, A. Goyal and C. M. G. A. Fontes, Carbohydrates and Carbohydrate-Active enZymes (CAZyme): An overview, in Glycoside Hydrolases, Elsevier, 2023, pp. 1–23. [cited 2025 May 12]. Available from: https://linkinghub.elsevier.com/retrieve/pii/B9780323918053000125 Search PubMed.
- S. C. Di Rienzi and R. A. Britton, Adaptation of the Gut Microbiota to Modern Dietary Sugars and Sweeteners, Adv. Nutr., 2020, 11(3), 616–629 CrossRef PubMed.
- A. E. Kaoutari, F. Armougom, J. I. Gordon, D. Raoult and B. Henrissat, The abundance and variety of carbohydrate-active enzymes in the human gut microbiota, Nat. Rev. Microbiol., 2013, 11(7), 497–504 CrossRef PubMed.
- C. A. Lozupone, J. I. Stombaugh, J. I. Gordon, J. K. Jansson and R. Knight, Diversity, stability and resilience of the human gut microbiota, Nature, 2012, 489(7415), 220–230 CrossRef CAS PubMed.
- J. Khan, W. N. I. W. M. Zain and M. N. Islam, Involvement of ACE2 in the intestinal transport of amino acids: Possible health and nutritional consequences in altered expression, Bangladesh J. Med. Sci., 2023, 22(4), 729–733 CrossRef.
- M. Mirzaei, S. Mirdamadi, M. R. Ehsani and M. Aminlari, Production of antioxidant and ACE-inhibitory peptides from Kluyveromyces marxianus protein hydrolysates: Purification and molecular docking, J. Food Drug Anal., 2018, 26(2), 696–705 CrossRef CAS PubMed.
- X. L. Zhu, K. Watanabe, K. Shiraishi, T. Ueki, Y. Noda and T. Matsui, et al., Identification of ACE-inhibitory peptides in salt-free soy sauce that are transportable across caco-2 cell monolayers, Peptides, 2008, 29(3), 338–344 CrossRef CAS PubMed.
- K. E. El Mecherfi, S. D. Todorov, M. A. Cavalcanti de Albuquerque, S. Denery-Papini, R. Lupi and T. Haertlé, , et al., Allergenicity of Fermented Foods: Emphasis on Seeds Protein-Based Products, Foods, 2020, 9(6), 792 CrossRef CAS PubMed.
- D. Günal-Köroğlu, G. Karabulut, G. Ozkan, H. Yılmaz, B. Gültekin-Subaşı and E. Capanoglu, Allergenicity of Alternative Proteins: Reduction Mechanisms and Processing Strategies, J. Agric. Food Chem., 2025, 73(13), 7522–7546 CrossRef PubMed.
- X. Huang, D. Schuppan, L. E. Rojas Tovar, V. F. Zevallos, J. Loponen and M. Gänzle, Sourdough Fermentation Degrades Wheat Alpha-Amylase/Trypsin Inhibitor (ATI) and Reduces Pro-Inflammatory Activity, Foods, 2020, 9(7), 943 CrossRef CAS PubMed.
- D. A. Anagnostopoulos, E. A. Karamani, S. Letsiou, A. Tsiartsafis, N. Chatzimpyrou and I. S. Boziaris, et al., Bacterial and fungal community composition in dairy products and plant-based dairy alternatives using high throughput sequencing, Food Res. Int., 2025, 221, 117552 CrossRef CAS PubMed.
- M. Palmnäs-Bédard, A. de Santa Izabel, J. Dicksved and R. Landberg, Characterization of the Bacterial Composition of 47 Fermented Foods in Sweden, Foods, 2023, 12(20), 3827 CrossRef PubMed.
- E. A. Landis, A. M. Oliverio, E. A. McKenney, L. M. Nichols, N. Kfoury and M. Biango-Daniels, et al., The diversity and function of sourdough starter microbiomes, eLife, 2021, 10, e61644 CrossRef CAS PubMed.
- A. T. Reese, A. A. Madden, M. Joossens, G. Lacaze and R. R. Dunn, Influences of Ingredients and Bakers on the Bacteria and Fungi in Sourdough Starters and Bread, mSphere, 2020, 5(1), e00950–e00919 CrossRef CAS PubMed.
- L. De Vuyst, A. Comasio and S. V. Kerrebroeck, Sourdough production: fermentation strategies, microbial ecology, and use of non-flour ingredients, Crit. Rev. Food Sci. Nutr., 2023, 63(15), 2447–2479 CrossRef CAS PubMed.
- G. Huys, H. M. Daniel and L. De Vuyst, Taxonomy and Biodiversity of Sourdough Yeasts and Lactic Acid Bacteria, in Handbook on Sourdough Biotechnology, ed. M. Gobbetti and M. Gänzle, Springer US, New York, NY, 2013, pp. 105–154. Available from: https://link.springer.com/10.1007/978-1-4614-5425-0_5 Search PubMed.
- C. Urien, J. Legrand, P. Montalent, S. Casaregola and D. Sicard, Fungal Species Diversity in French Bread Sourdoughs Made of Organic Wheat Flour, Front. Microbiol., 2019, 10, 201 CrossRef PubMed.
- B. García-Béjar, P. Fernández-Pacheco, J. Carreño-Domínguez, A. Briones and M. Arévalo-Villena, Identification and biotechnological characterisation of yeast microbiota involved in spontaneous fermented wholegrain sourdoughs, J. Sci. Food Agric., 2023, 103(15), 7683–7693 CrossRef PubMed.
- Y. Perez-Riverol, J. Bai, C. Bandla, D. García-Seisdedos, S. Hewapathirana and S. Kamatchinathan, et al., The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences, Nucleic Acids Res., 2022, 50(D1), D543–D552 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |