Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
A fructo-oligosaccharide and sea buckthorn complex ameliorates glucose and lipid metabolism in type 2 diabetic (T2DM) rats via metabolic and anti-inflammatory pathways
Ling
Chen†
 ab,
Hualin
Liu†
ab,
Hualin
Liu†
 ab,
Yanwu
Chen
ab,
Qianya
Xiao
ab,
Zijin
Huang
ab,
Ziying
Li
ab,
Xiaofei
Zhang
ab,
Xueying
Fan
ab,
Meichen
Xiao
ab,
Xiaomin
Li
*abc and
Yuan Yao
Chen
ab,
Yanwu
Chen
ab,
Qianya
Xiao
ab,
Zijin
Huang
ab,
Ziying
Li
ab,
Xiaofei
Zhang
ab,
Xueying
Fan
ab,
Meichen
Xiao
ab,
Xiaomin
Li
*abc and
Yuan Yao
Chen
 *ab
*ab
aResearch & Development Division, Perfect (Guangdong) Co., Ltd, Zhongshan, China. E-mail: yfglblxm@126.com; yuanyaochen0401@163.com
bResearch & Development Division, Perfect Life & Health Institute, Zhongshan, China
cSchool of Traditional Chinese Materia Medica, Shenyang Pharmaceutical University, Shenyang, Liaoning 110016, China
First published on 24th April 2026
Abstract
This study investigated the effects and underlying mechanisms of a fructo-oligosaccharide (FOS) and sea buckthorn complex (FS) on blood glucose and lipid metabolism in type 2 diabetic mellitus (T2DM) rats. The T2DM model was induced by a high-fat diet (HFD) combined with alloxan administration. Prior to rat experiments, the synergistic hypoglycemic and hypolipidemic effects of FOS and sea buckthorn have been verified by zebrafish experiments. FS intervention significantly reduced fasting blood glucose, triglycerides (TG), total cholesterol (TC), and low-density lipoprotein cholesterol (LDL-C) in rats, although high-density lipoprotein cholesterol (HDL-C) showed a non-significant increasing trend. 16S rDNA sequencing revealed that FS significantly reshaped gut microbiota; it enriched beneficial genera (e.g., Akkermansia, Bifidobacterium) and suppressed inflammation-associated taxa (e.g., Escherichia–Shigella). Transcriptomic analysis indicated that FS reversed aberrant expression of inflammation-related genes (e.g., Retnlg, Retnlb, Bmp2 in the colon) and metabolism-related genes (e.g., Gpam, Cpt1a in the liver), activating pathways including fat digestion and absorption and fatty acid metabolism, while inhibiting immune-inflammatory pathways (e.g., the “TNF signaling pathway”, “Th17 cell differentiation”). Serum bile acid (BA) profiling showed FS modulated levels of primary and secondary BAs (e.g. CA, α-MCA, HDCA, GHDCA), restoring metabolic homeostasis. Pearson correlation analyses demonstrated robust associations among key microbiota, BAs, blood glucose/lipid indices, and farnesoid X receptor (FXR/Nr1h4) expression, supporting a regulatory network involving the gut–liver axis. This study elucidates the multi-target mechanisms by which FS ameliorates glucolipid metabolic disorders in T2DM, via gut microbiota remodeling, attenuation of intestinal/hepatic inflammation, and BA metabolism crosstalk, providing a theoretical basis for prebiotic-based precision nutrition.
1. Introduction
Type 2 diabetes mellitus (T2DM) is a chronic metabolic disorder primarily characterized by impaired insulin secretion and/or insulin resistance and has become a major global public health challenge.1,2 Current pharmacological interventions (e.g., metformin and GLP-1 receptor agonists) partially regulate blood glucose but often fail to control lipid disorders and cause adverse effects.3 Moreover, long-term use of these medications is associated with adverse effects including gastrointestinal disturbances and vitamin B12 deficiency.4 These limitations underscore the urgent need for novel therapeutic strategies. Beyond insulin dysfunction, accumulating evidence highlights gut microbiota dysbiosis and gut–liver axis disruption as key pathogenic factors:5 T2DM patients exhibit reduced beneficial short-chain fatty acid (SCFA)-producing bacteria (Akkermansia muciniphila, Faecalibacterium prausnitzii) and increased endotoxin-producing taxa (Escherichia coli),6–8 which impair intestinal barrier integrity and trigger systemic inflammation.Prebiotics exhibit unique advantages in the management of T2DM by regulating the gut microbiota.9 Among them, dietary administration of fructo-oligosaccharides (FOS) can significantly ameliorate multiple metabolic abnormalities in T2DM mice induced by a high-fat diet (HFD), including insulin resistance, systemic inflammation, oxidative stress, dyslipidemia, and hepatic steatosis.10 A meta-analysis involving 607 adult T2DM participants further indicated that FOS supplementation is positively associated with reduced fasting insulin levels and increased high-density lipoprotein cholesterol (HDL-C).11 The potential mechanisms underlying these benefits include promoting the proliferation of beneficial bacteria such as Bifidobacterium,10,12 facilitating gut bacterial fermentation to produce SCFAs and regulating bile acids (BAs), both of which interact with host metabolism,13,14 as well as inhibiting the growth of pathogenic bacteria, reducing the production of lipopolysaccharide (LPS), and restoring intestinal barrier integrity to alleviate inflammation.15,16 Sea buckthorn (Hippophae rhamnoides L.) is a traditional Chinese herbal medicine rich in bioactive components (polyphenols, fatty acids, vitamins, and phytosterols)17 and it also exerts potential beneficial effects on metabolic disorders.18–20In vitro and in vivo experiments (including cell lines, animal models, and human patients) have shown that sea buckthorn can lower blood lipid, blood pressure, and blood glucose levels and regulate key metabolites;21 its antioxidant compounds further alleviate intestinal oxidative damage,22 creating a favorable microenvironment for the gut microbiota. 16S rDNA sequencing analysis further confirms that sea buckthorn treatment significantly reduces body weight and blood glucose levels in diabetic mice, while restoring the normal abundance of Bifidobacterium, Lactobacillus and Bacillus.23
However, most current studies focus on single-component interventions. Despite the comprehensive metabolic benefits of FOS, their effect on hepatic lipid metabolism remains limited when used alone. For sea buckthorn, although it regulates the gut microbiota and improves metabolic indicators, there is still insufficient evidence to confirm its role in “gut microbiota–BAs” crosstalk. Therefore, whether the combination of FOS and sea buckthorn exerts a synergistic effect on T2DM remains an open question. To address this gap, we first performed a preliminary zebrafish experiment to verify the synergistic hypoglycemic and hypolipidemic effects of FOS and sea buckthorn, which provided a basis for subsequent rat experiments (see SI file1.docx for details). Based on this, we hypothesize that the FOS–sea buckthorn complex (FS) exerts synergistic ameliorative effects on T2DM via the “gut microbiota–BAs–host” axis. Using multi-omics technologies (16S rDNA sequencing, transcriptomics, serum BA profiling), this study aims to: (1) elucidate FS-induced changes in gut microbiota composition and their associations with BA profiles; (2) decipher FS's regulatory effects on colon inflammatory genes and liver metabolic genes and key pathways; (3) identify predictive biomarkers for FS efficacy. This work will lay a theoretical foundation for prebiotic-based precision nutrition strategies targeting T2DM.
2. Materials and methods
2.1 Animals, T2DM model and FS complex
Male specific pathogen-free (SPF) Sprague-Dawley rats (Rattus norvegicus, weight 130–170 g) were purchased from Zhuhai Bestest Biotechnology Co., Ltd (Zhuhai, Guangdong, China) and housed in an SPF-grade environment. The rats were housed in alternating light and dark cycles at a temperature of 22 °C ± 2 °C, humidity of 60% ± 10%, and differential pressure of ≥10 Pa. T2DM was induced by feeding the rats a high-fat diet combined with low-dose alloxan administration, resulting in insulin resistance and T2DM development. The FS complex applied in this study was mainly composed of FOS and sea buckthorn powder, supplied by Perfect (Guangdong) Co., Ltd (Zhongshan, Guangdong, China). To clarify the chemical basis of the FS complex, quantitative analysis of its key components was conducted using high-performance liquid chromatography (HPLC, Agilent 1260 Infinity III RID). The purity of FOS was determined to be 94.9%, with its major polymeric components including kestose (50.1%), nystose (40.3%), and fructofuranosylnystose (4.5%). Fresh sea buckthorn fruits were characterized by their rich bioactive constituents, where the total flavonoid content (expressed as rutin equivalent) was 0.61% (w/w) (UV-Vis Spectrophotometry). The primary flavonoid components were identified as narcissin, rutin, isorhamnetin-3-O-glucoside-7-O-rhamnoside, isorhamnetin-3-O-glucoside, isoquercetin, and kaempferol, with contents of 21.48 mg per 100 g, 7.37 mg per 100 g, 4.98 mg per 100 g, 3.12 mg per 100 g, 1.31 mg per 100 g, and 0.42 mg per 100 g, respectively (HPLC). The total content of organic acids accounted for 8.38% (w/w), among which malic acid, succinic acid, tartaric acid, adipic acid, and citric acid were the main components, with the corresponding contents of 4.6%, 2.0%, 0.88%, 0.7%, and 0.2% (w/w), respectively (HPLC). Detailed chromatographic profiles and component quantification data for FOS and fresh sea buckthorn fruits are provided in the SI (Fig. S1 and S2 and SI file2.docx).2.2 Experimental design
Following a 5-day acclimatization period, the rats were randomly divided into two initial groups: the control group (Con, n = 8) and the high-fat group (n = 16). The Con group was fed a standard maintenance diet, while the high-fat group received a HFD. Both groups had free access to food and water throughout the acclimatization and initial dietary intervention phases. After 7 days of dietary intervention, rats displaying obesity resistance (20% with slight weight gain) were excluded based on weight gain criteria. The remaining high-fat group rats were further divided into two groups: the model (Mod) group and the FS group, each group consisted of six rats (Fig. 1A). During the subsequent intervention period the Con group continued on a normal chow diet and received a daily gavage of sterile water at a volume of 1 mL per 100 g body weight (BW); the Mod group remained on the HFD and received the same daily gavage of sterile water (1 mL per 100 g BW) as the Con group. The FS group was maintained on the HFD and received a daily gavage of the FS complex at a dose of 2.06 g per kg BW (gavage volume standardized to 1 mL per 100 g BW). On Day 56 of the experiment, all rats (Con, Mod, and FS groups) were subjected to a 24 hour fast (with free access to water). On Day 57, the Mod group and FS group were injected with alloxan at a dose of 105 mg per kg BW. Thereafter, the respective dietary regimens and daily gavage treatments were continued until the animals were sacrificed. All animal procedures were performed in accordance with the ARRIVE guidelines for the reporting of in vivo experiments and approved by the Institutional Animal Care and Use Committee (WM-GD-DW001) of the Perfect (GuangDong) Co., Ltd (Zhongshan, China).2.3 Oral glucose tolerance test
The oral glucose tolerance test (OGTT) was performed on Day 63 of the experiment. All rats first underwent a 12-hour fast (with no access to food or water), after which their fasting blood glucose (FBG) level (0-hour blood glucose value) was measured. Prior to glucose administration, each group received their respective scheduled gavage treatment, with the entire procedure completed within 20 minutes. Immediately, all rats were administered an oral glucose load at a dose of 2.5 g per kg BW. Blood glucose levels were subsequently measured at 0.5 and 2 h post-glucose administration. To evaluate the oral glucose tolerance of the rats, the area under the blood glucose curve (AUC) from 0 to 2 hours (integrating the 0-, 0.5-, and 2-hour blood glucose values) was calculated.2.4 Sample collection
Fecal samples were collected from all rats on Day 62. Naturally excreted fecal samples were collected under sterile conditions: sterile EP tubes were gently positioned near the rats’ anus to capture fresh excreta. On Day 64, rats were anesthetized prior to sample collection. For blood collection, retro-orbital blood sampling was performed to obtain 2–5 mL of whole blood, which was immediately centrifuged to separate serum. Both fecal samples and serum aliquots were stored at −80 °C until subsequent analysis. For tissue sampling, liver and colon tissues were harvested immediately after blood collection. These tissues were first snap-frozen in liquid nitrogen to preserve RNA integrity and then transferred to −80 °C storage. For pathological analysis of liver tissue, liver tissue specimens were heat-fixed at 60 °C for 1 h, followed by hematoxylin and eosin (H&E) staining as previously described.24 Additionally, serum levels of four blood lipid parameters, triglycerides (TG), total cholesterol (TC), high-density lipoprotein cholesterol (HDL-C), and low-density lipoprotein cholesterol (LDL-C), were quantified for each group.2.5 Fecal 16S rDNA amplicon sequencing and microbiome analysis
Microbial genomic DNA was extracted from fecal samples using the MagPure Stool DNA Kit (Guangzhou Magen Biotechnology Co., Ltd, Guangzhou, China) and quantified with a NanoDrop 2000c spectrophotometer (Thermo Fisher Scientific, Waltham, MA, USA) following the manufacturer's instructions. The V3–V4 hypervariable region of bacterial 16S rDNA was amplified with universal primers (forward: 5′-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG-3′, reverse: 5′-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC-3′) under the following thermal cycling conditions: initial denaturation at 94 °C for 5 min; 25 cycles of denaturation at 94 °C for 30 s, annealing at 58 °C for 30 s, and extension at 72 °C for 30 s; final extension at 72 °C for 5 min. Amplified PCR products were purified using VAHTS DNA Clean Beads (NanJing Vazyme BioTech Co., Ltd, Nanjing, China), followed by index PCR and library construction. Library quality was assessed using a Qubit 3.0 fluorometer with the Qubit™ 1× dsDNA HS Assay Kit. Paired-end sequencing (PE250) was performed on an Illumina MiSeq platform (Illumina, San Diego, CA, USA). Raw sequencing data were demultiplexed based on unique barcodes and quality-filtered with FastQC (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and Trimmomatic.25 Clean reads were imported into QIIME2 for amplicon sequence variants (ASVs).26 DADA2 plugin was used for denoising, chimera removal, and deduplication to generate amplicon sequence variants (ASVs) and the feature table.27 Rarefaction analysis was performed in QIIME2 on the feature table (subsequently exported to R software, v4.1.2). After normalization, each sample retained 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 756 reads for downstream analysis. Microbial community analysis, including α-diversity, β-diversity, and taxonomic classification, was performed using R. The gut microbiota signature of each group was established by Linear discriminant analysis Effect Size (LEfSe).28
756 reads for downstream analysis. Microbial community analysis, including α-diversity, β-diversity, and taxonomic classification, was performed using R. The gut microbiota signature of each group was established by Linear discriminant analysis Effect Size (LEfSe).28
Mediation analysis was conducted to evaluate whether the effect of the intervention group (Mod = 0 vs. FS = 1) on metabolic outcomes (TG, TC, HDL-C, LDL-C, OGTT) was mediated by key gut microbial biomarker taxa (performed in R). For each mediator–outcome pair, a simple mediation model was fitted using ordinary least squares regression, estimating the following paths: a: the effect of the group on the mediator (X → M), b: the effect of the mediator on the outcome adjusted for group (M → Y|X), c′: the direct effect of the group on the outcome (X → Y|M), c: the total effect of the group on the outcome (X → Y). The indirect effect was computed as the product of coefficients ab. To account for the small sample size, a non-parametric bootstrap procedure with 2000 resamples was used to estimate 95% confidence intervals (CIs) for the indirect effect using the percentile method. A mediation effect was considered statistically significant if its 95% CI did not include zero. We applied the Benjamini–Hochberg procedure to control the FDR. Given the exploratory nature of the analysis and limited statistical power, primary findings are reported for mediation effects meeting an FDR threshold of <0.2, consistent with practices in exploratory omics research.
2.6 RNA sequencing and transcriptome analysis
Total RNA was extracted from liver and intestine tissues using TRIzol reagent (Invitrogen, Carlsbad, CA, USA), following the protocol described previously.29 RNA sequencing was performed by Gene Denovo Corporation (https://www.genedenovo.com/about/), and clean data were generated. Subsequently, clean data were quality-filtered by FastQC. High-quality sequences were processed with Salmon for quantification of the transcript abundance.30 The resulting transcript abundance matrix was analyzed with DESeq2 (https://github.com/thelovelab/DESeq2) to identify the differential expressed genes (DEGs). Genes with an adjusted P-value (p-adjust) < 0.05 and absolute log2 fold change (|log2FC|) > 1.5 were defined as DEGs. These DEGs were further subjected to Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis and Gene Set Enrichment Analysis (GSEA) using the R package clusterProfiler.31 To control type I error in multiple tests, the false discovery rate (FDR) was calculated with the Benjamini–Hochberg procedure (implemented in the R function p-adjust).2.7 Serum bile acid analysis
BA standards were accurately weighed to prepare a concentrated stock solution (1000 μg mL−1) in methanol. The concentrated stock solution was diluted with 30% methanol to generate a series of ten standard curve concentrations. All standard stock solutions and working solutions were stored at −20 °C. Serum samples were mixed with 600 μL of prechilled (−20 °C) methanol in centrifuge tubes, followed by centrifugation at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 10 min at 4 °C. The supernatant (400 μL) was evaporated to dryness using a vacuum concentrator, and the resulting residue was reconstituted with 100 μL of 30% methanol. The reconstituted solution was filtered through a 0.22 μm membrane, and the filtrate was transferred to sample vials for liquid chromatography-mass spectrometry (LC-MS) analysis. BA quantification was performed using an EXion LC liquid chromatograph coupled with an AB6500 Plus mass spectrometer (AB SCIEX, Framingham, MA, USA). The targeted metabolic analysis of BAs was outsourced to PANOMIX Biotech Co., Ltd (Suzhou, China).
000g for 10 min at 4 °C. The supernatant (400 μL) was evaporated to dryness using a vacuum concentrator, and the resulting residue was reconstituted with 100 μL of 30% methanol. The reconstituted solution was filtered through a 0.22 μm membrane, and the filtrate was transferred to sample vials for liquid chromatography-mass spectrometry (LC-MS) analysis. BA quantification was performed using an EXion LC liquid chromatograph coupled with an AB6500 Plus mass spectrometer (AB SCIEX, Framingham, MA, USA). The targeted metabolic analysis of BAs was outsourced to PANOMIX Biotech Co., Ltd (Suzhou, China).
2.8 Statistical analysis
All statistical analyses were performed using R software. For microbiological data evaluation, multiple R packages, including psych,32 tidyverse,33 ggplot2,34 and igraph,35 were used for data analysis and visualization. p-Values were calculated using the Adonis test based on 999 permutations. Group differences were assessed using the Mann–Whitney U test, with p-values adjusted (p-adjust) for multiple comparisons using the Benjamini–Hochberg procedure. A p-value of less than 0.05 was considered statistically significant. All figures were generated using R software.3. Results
3.1 FS ameliorates blood glucose and lipid profiles in T2DM rats
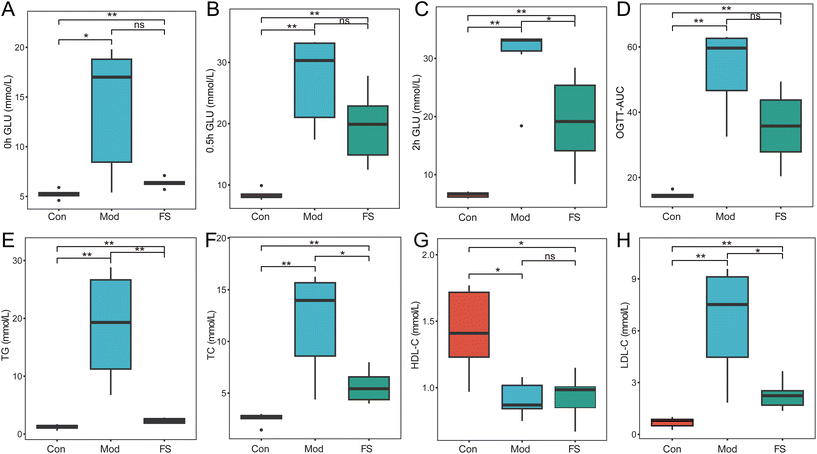
Following a 10–12 h fast, an OGTT was conducted on day 63 to assess glucose tolerance in rats. As depicted in Fig. 2, compared with the Con group, the Mod group exhibited significantly elevated fasting blood glucose (0 h GLU, Fig. 2A, P < 0.05), blood glucose at 0.5 h post-gavage (0.5 h GLU, Fig. 2B, P < 0.01), and blood glucose at 2 h post-gavage (2 h GLU, Fig. 2C, P < 0.01). In contrast to the Mod group, the FS group showed a significant reduction in blood glucose at 2 h post-gavage (Fig. 2C, P < 0.05). Fasting and 0.5 h post-gavage glucose levels were also lower in the FS group, though differences were not statistically significant (Fig. 2A and B). Similarly, OGTT AUC in the FS group trended lower relative to the Mod group, with no significant difference (Fig. 2D). These results indicate that FS improves glucose regulation in T2DM rats. For lipid profiles, the Mod group had significantly higher levels of triglycerides (TG, Fig. 2E, P < 0.01), total cholesterol (TC, Fig. 2F, P < 0.01), and low-density lipoprotein cholesterol (LDL-C, Fig. 2H, P < 0.01) compared with the Con group. Supplementation with FS markedly reduced these elevated lipid indices (Fig. 2E and F, P < 0.01; Fig. 2H, P < 0.05). Additionally, high-density lipoprotein cholesterol (HDL-C) levels were significantly decreased in the Mod group compared to the Con group (Fig. 2G, P < 0.05), yet no significant difference was observed between the Mod and FS groups (Fig. 2G). These findings demonstrate that FS ameliorates glucose and lipid homeostasis in T2DM rats.3.2 FS modulates the community structure of gut bicrobiota in T2DM rats
16S rDNA amplicon sequencing was used to characterize fecal microbial communities in rats. Compared with the Con group, the Mod group showed a significant decrease in α-diversity. In contrast to the Mod group, the FS group exhibited a further reduction in α-diversity (Fig. 3A and B, P < 0.05; the Wilcoxon test). β-Diversity analysis (PCoA and Adonis tests) revealed distinct clustering between the Con and Mod groups (Fig. 3C, R2 = 0.525, P = 0.003), as well as between the Mod and FS groups (Fig. 3C, R2 = 0.492, P = 0.002). Subsequent analyses of the fecal microbiota composition showed that the gut microbiota in all groups was predominately composed of Firmicutes, Bacteroidota, and Verrucomicrobiota at the phylum level (Fig. S3) and primarily consisted of Bacteroides, Akkermansia, Muribaculaceae uncultured bacterium, Parabacteroides, and Blautia (top 20 genera, Fig. 3D) at the genus level.LEfSe analysis was performed to identify group-specific microbial biomarkers. A total of 12 biomarkers were identified in the Mod group, including Desulfovibrionaceae uncultured, Escherichia Shigella, Parasutterella, Candidatus_Stoquefichus, Hungatella, Faecalitalea, Mucispirillum, Phocea, Clostridioides, Butyricimonas, Holdemania, and Eisenbergiella (Fig. 3E, LDA > 3.2, P < 0.05). In the FS group, taxa such as Akkermansia, Parabacteroides, Blautia, Ruminococcus torques group, Lachnoclostridium, Flavonifractor, Bacteroidales uncultured, Bilophila, Bifidobacterium, Ruminococcus gauvreauii group, and Coriobacteriaceae-UCG-002 were significantly more abundant (Fig. 3E, LDA > 3.2, P < 0.05). Mediation analysis revealed significant indirect effects of specific gut microbial taxa in mediating the metabolic benefits of FS intervention (Fig. S4). Specifically, Phocae significantly mediated the FS-induced reductions in TC, LDL-C, and OGTT outcomes, with indirect effects of −7.43, −4.92, and −21.44, respectively (Fig. S4A–C). Additionally, Desulfovibrionaceae_uncultured mediated the decrease in OGTT with an indirect effect of −32.07 (Fig. S4D). These results demonstrate that FS regulates the composition and structure of gut microbiota in T2DM rats.
3.3 FS regulates the transcription of genes associated with colonic inflammatory processes in T2DM rats
Gut microbiota and their metabolites influence systemic health through extensive connections between the intestine and other tissues.36 To explore the mechanism of FS action, transcriptome analysis was performed on colonic tissues. Principal component analysis (PCA) showed distinct clustering among the three groups, indicating divergent gene expression profiles across Con, Mod and FS groups (Fig. 4A). When comparing Mod vs. Con, 384 DEGs were identified: 160 genes were downregulated and 224 genes were upregulated in Mod (Fig. 4B). Elevated FDRs were observed in the Mod group compared to the Con group for both upregulated (e.g., Retnlg, Retnlb, Bmp2) and downregulated genes (e.g., Ckm, Klf15, Slc15a2) (SI file3.xlsx). In the FS vs. Mod comparison, 219 DEGs were identified: 135 genes were downregulated and 84 genes were upregulated in FS (Fig. 4C). Relative to the Mod, FS reversed approximately one-third of the DEGs observed in Mod vs. Con (SI file3.xlsx).GSEA based on KEGG pathways showed that, compared with Mod, FS activated metabolism-related pathways, such as “Fat digestion and absorption”, “Starch and sucrose metabolism” and “Carbohydrate digestion and absorption” (Fig. 4D, left panel). Conversely, immune-related pathways, including “Intestinal immune network for IgA production”, “Natural killer cell mediated cytotoxicity” and “TNF signaling pathway” were suppressed in FS vs. Mod, whereas these pathways were activated in the Mod vs. Con (Fig. 4D, right panel). To further elucidate the molecular mechanisms underlying the beneficial effects of FS intervention, we performed enrichment analysis of DEGs in key signaling pathways. Notably, within the “IL-17 signaling pathway”, the majority of genes that were significantly upregulated in the model group (Mod vs. Con) were effectively restored following FS treatment (Fig. S5). These results suggest that FS enhances the metabolic capacity and alleviates intestinal inflammation in T2DM rats.
3.4 FS regulates the transcription of genes related to hepatic metabolism and inflammation in T2DM rats
H&E staining of liver sections revealed increased lipid droplets and lymphocyte infiltration in the Mod group, whereas FS treatment reduced hepatic lipid accumulation and lymphocyte infiltration (Fig. S6). To elucidate the underlying mechanisms, transcriptome analysis was performed on liver tissues. PCA demonstrated clear distinction between Mod and the other two groups (Fig. 5A). A total of 1044 DEGs were identified between Con and Mod: 409 genes were downregulated and 635 genes were upregulated in Mod (Fig. 5B). Compared with Con, the Mod group showed higher FDR values for upregulated genes (e.g., Gpam, Lifr, Srebf1) and downregulated genes (e.g., Slc25a47, Ugp2, Cpt1a) (SI file4.xlsx). In the FS vs. Mod comparison, 1120 DEGs were identified: 684 genes were downregulated and 436 genes were upregulated in FS (Fig. 5C). Relative to Mod, FS reversed approximately half of the DEGs observed in Mod vs. Con (SI file4.xlsx).GSEA-KEGG enrichment analysis showed that, compared with the Mod, FS activated metabolism-related pathways including “pyruvate metabolism”, “fatty acid metabolism” and “lipoic acid metabolism” (Fig. 5D, left panel). Conversely, immune-related pathways including “intestinal immune network for IgA production”, “natural killer cell mediated cytotoxicity” and “Th17 cell differentiation” were suppressed in FS vs. Mod, whereas these pathways were activated in Mod vs. Con (Fig. 5D, right panel). Further enrichment analysis of DEGs across key metabolic pathways revealed that the FS effectively reversed the transcriptional suppression induced by a HFD. Specifically, in the mod group (Mod vs. Con), numerous genes involved in “fatty acid degradation” (Fig. S7), “pyruvate metabolism” (Fig. S8), and the “Citrate cycle (TCA cycle)” (Fig. S9) were significantly downregulated. Remarkably, FS intervention largely restored the expression of these genes (FS vs. Mod). Overall, KEGG enrichment analysis of DEGs indicated that “metabolic pathway” was the most significantly enriched signaling pathway across groups (Fig. S10). These findings demonstrate that FS enhances hepatic metabolic capacity and attenuates liver inflammation in T2DM rats.
3.5 FS modulates serum bile acid levels in T2DM rats
Serum BA concentrations were quantified across all groups. Partial least squares-discriminant analysis (PLS-DA) showed a marked separation in serum BA profiles between Con and Mod (Fig. 6A). Similarly, a significant distinction in serum BA composition was observed between Mod and FS (Fig. 6B). Subsequent analyses of major differential serum BAs revealed that for primary BAs, levels of cholic acid (CA, Fig. 6C), chenodeoxycholic acid (CDCA, Fig. 6D), and α-MCA (Fig. 6F) were reduced in Mod vs. Con but showed partial recovery in FS (though not all differences reached significance). Taurochenodeoxycholic acid (TCDCA) increases significantly in Mod and FS compared with Con (Fig. 6E). Beta-muricholic acid (β-MCA) showed a significant increase in both FS vs. Mod and FS vs. Con (Fig. 6G). For secondary BAs, HDCA and GHDCA were markedly downregulated in Mod vs. Con (P < 0.01) and remained low in FS (Fig. 6H and I). Compared to the Con group, the Mod group showed a significant reduction in ursodeoxycholic acid (UCA), which was restored by FS treatment (Fig. 6J). While lithocholic acid (LCA) did not differ significantly across groups (Fig. 6K), allo-cholic acid (ACA) was significantly decreased in Mod vs. Con and displayed a recovering, albeit non-significant, trend in the FS group (Fig. 6L). These results indicate that FS modulates serum BA profiles, including both primary and secondary BAs, in T2DM rats.3.6 Correlations between key microbiota, bile acids, and blood glucose/lipid parameters
Correlation analyses were performed to assess associations among key differential gut microbiota, BAs, Nr1h4 expression (Fig. S11), and blood glucose/lipid parameters (P < 0.05, |r| > 0.6), with results used to construct the network in Fig. 7 (see SI file5.xlsx for details). Specifically, for microbiota–glucose/lipid correlations, genera such as Bilophila, Coriobacteriaceae-UCG-002, Bifidobacterium, Parabacteroides, Ruminococcus gauvreauii group, and Bacteroidales uncultured were negatively correlated with TG, whereas Desulfovibrionaceae uncultured, Phocea, Mucispirillum, Holdemania, Hungatella, and Escherichia-Shigella showed positive correlations with TG. Consistent with the aforementioned analyses, the beneficial effects of the FS intervention on TC, LDL-C, and OGTT were partially mediated through its modulation of Phocae and Desulfovibrionaceae_uncultured (Fig. S4). Besides, Parasutterella was positive related to OGTT, TC, and LDL-C. Holdemania was positive related to OGTT, TC, TG and LDL-C. Regarding BAs and glucose/lipid correlations, HDCA and GHDCA were negatively correlated with TG, and moreover, TG, TC, LDL-C and OGTT results exhibited consistent positive intercorrelations with parallel association patterns with BAs and microbiota. Notably, Bifidobacterium (a representative probiotic, further identified as Bifidobacterium pseudolongum) exhibited significant negative correlations with TC, TG and LDL-C, as well as with the microbial taxa Desulfovibrionaceae uncultured, Phocea and Hungatella; Conversely, it showed positive associations with the BAs HDCA, GHDCA and glycodeoxycholic acid (GDCA) and with the bacterial genera Coriobacteriaceae-UCG-002, Parabacteroides and Bilophila. Nr1h4, a receptor closely involved in BA metabolism, exhibited expression levels in both the liver and colon that were significantly positively correlated with GDCA. Additionally, hepatic Nr1h4 expression showed a strong positive correlation with HDCA and GHDCA, while colonic Nr1h4 expression was significantly negatively correlated with TC and LDL-C. Collectively, these correlations suggest that FS intervention modulates the gut microbiota composition and metabolite levels, thereby influencing blood glucose and lipid homeostasis.Guided by established BA metabolism pathways,37–39 we further analyzed the molecular and microbial alterations associated with FS intervention (Fig. S12). Notably, FS treatment significantly upregulated the hepatic expression of key BA-modifying enzymes, including cytochrome P450 family 27 subfamily A member 1 (CYP27A1) and UGTs (UDP-glucuronosyltransferases) (Fig. S13). Concurrently, in the intestinal microbiota, FS markedly increased the relative abundance of Bifidobacterium—a genus known to harbor bile salt hydrolase (BSH) activity, and Blautia—a bacterial group implicated in 7α-dehydroxylation of BAs. These coordinated changes in both host enzymatic machinery and gut microbial composition suggest that CYP27A1, UGTs, Bifidobacterium, and Blautia may collectively play pivotal roles in mediating the lipid-lowering effects of FS through modulation of BA metabolism.
4. Discussion
In this study, multi-omics techniques were used to systematically elucidate the intervention mechanisms of FS in T2DM rats. The results showed that FS could ameliorate glycolipid metabolic disorders in multiple dimensions by remodeling the gut microbiota structure, regulating host metabolic and anti-inflammatory pathways, and restoring BA homeostasis.4.1 FS ameliorates glucose and lipid metabolic homeostasis in diabetic rats
Dysregulation of blood glucose and lipids represents a hallmark pathological feature of T2DM and its associated cardiovascular complications.40 In this study, FS significantly reduced fasting blood glucose and 2 hour postprandial glucose levels during an OGTT in T2DM rats. Notably, while FS induced a significant decline in 2 h glucose, the OGTT AUC only showed a near-significant decreasing trend (P = 0.0649); a parallel trend was also observed for 0.5 h glucose (P = 0.0921). This discrepancy stems from the characteristics of the indices: OGTT AUC integrates glucose dynamics across 0 h, 0.5 h, and 2 h, such that intra-group variability at each time point (e.g., the larger fluctuation of 0.5 h glucose, Fig. 2B) accumulates to amplify the overall variability of AUC, reducing its statistical power. In contrast, 2 h glucose (a single time point) exhibited more concentrated metabolic responses in this cohort (narrower error bar in Fig. 2C, i.e., smaller intra-group variability), enabling the detection of inter-group differences even with a limited sample size (n = 6 per group). These near-significant trends still suggest that FS may benefit overall glucose tolerance, a potential effect masked by cumulative variability of integrated indices and a small sample size. Future studies with expanded cohorts can verify FS's role in improving global glucose tolerance. FS also downregulated serum levels of TG, TC, and LDL-C, indicating its potential therapeutic value in managing diabetes-related metabolic dysregulation.41–43 These beneficial effects appear to be mediated by distinct yet complementary mechanisms. FOS enhance insulin sensitivity via gut microbiota-derived SCFAs, while sea buckthorn constituents modulate hepatic lipid metabolism by suppressing lipogenesis and promoting fatty acid β-oxidation.44,45 These findings are consistent with reports on other prebiotic compounds, suggesting shared pathways involving the regulation of hepatic lipid metabolism.46,47 Notably, the lack of significant effects on HDL-C levels may reflect intervention-specific pharmacokinetic properties or may require longer treatment durations to elicit detectable changes.4.2 Functional significance of FS in remodeling the gut microbiota structure of T2DM rats
Gut microbiota dysbiosis is a key pathogenic factor in T2DM, characterized by a reduction in beneficial bacteria and enrichment of pro-inflammatory bacteria.48–50 This study found that FS could selectively enrich beneficial bacteria such as Akkermansia, Bifidobacterium, Blautia and Parabacteroides, while inhibiting pro-inflammatory bacteria including Escherichia and Desulfovibrionaceae. This is highly consistent with the microbiota-regulating properties of FOS and sea buckthorn: FOS serve as a preferential carbon source for Bifidobacterium, which exhibits stronger FOS utilization capacity than other intestinal bacteria due to unique metabolic advantages (e.g., higher β-fructosidase activity and more efficient transport systems), thereby promoting Bifidobacterium proliferation through nutritional competition.51,52 Meanwhile, sea buckthorn polysaccharides can repair the intestinal barrier and provide a mucous layer colonization environment for Akkermansia.53,54 These beneficial microbial populations have been well-documented to exert anti-diabetic effects through multiple mechanisms: (1) enhancement of glucose and lipid homeostasis via the production of SCFAs, (2) improvement of insulin sensitivity, (3) strengthening of intestinal barrier function, and (4) reduction of systemic inflammation associated with metabolic dyslipidemia.55–60 In contrast, pathogenic microbes are known to exacerbate systemic inflammation through LPS-mediated mechanisms, thereby promoting insulin resistance in T2DM.61 These microbiota-derived inflammatory factors activate innate immune pathways, which in turn disrupt glucose metabolism.62–64 In addition, several biomarkers in the Model group have been reported to be detrimental to health. A study including 1544 participants revealed that Parasutterella was positively associated with BMI and type 2 diabetes, whereby the link to L-cysteine might be relevant in type 2 diabetes development and the link to the fatty acid biosynthesis pathway for body weight gain in obesity development.65Holdemania, a member of the Erysipelotrichaceae family, is implicated in glucose and lipid metabolic disorder and may contribute to the pathogenesis of obesity and T2DM.66–68Phocea is a potential biomarker influencing the progression and complications of obesity-induced type 2 diabetes and is also associated with energy metabolism and insulin sensitivity.69,70 Our finding suggests that Phocae and Desulfovibrionaceae_uncultured may serve as key microbial intermediaries linking FS-induced gut microbiota remodeling to systemic lipid and glucose homeostasis. Furthermore, the negative correlation between Bifidobacterium and TC, TG, LDL-C (Fig. 7), as well as the positive correlation between Escherichia-Shigella and TG, further confirms a causal relationship between the microbiota regulation by FS and the improvement of glucose and lipid metabolism. Collectively, these findings identify targeted microbiota modulation as a potential therapeutic strategy to break the vicious cycle between chronic inflammation and metabolic dysfunction in T2DM.71–73The diabetic condition was associated with a marked reduction in gut microbial α-diversity, consistent with previous reports linking microbial dysbiosis to metabolic dysfunction.74 Of note is that the α-diversity in the FS group was further decreased compared with that in the Mod group (Fig. 3A and B). This phenomenon does not indicate deterioration of the microbiota; instead, it may represent a manifestation of “functional remodeling”—FS selectively enriches metabolically dominant bacteria (e.g., Bifidobacterium) and competitively inhibits the growth of other non-dominant bacteria, thereby forming a “beneficial bacteria-dominated” microbiota structure. The long-term ecological consequences of this phenomenon require systematic evaluation through well-controlled longitudinal studies.
4.3 FS modulates metabolic and inflammatory pathways in the colon and liver of T2DM rats
Transcriptomic analysis revealed that FS can synchronously reverse abnormal gene expression in the colon and liver of diabetic rats, forming a “intestinal anti-inflammation-hepatic metabolism” synergistic effect. In colonic tissue, FS significantly downregulated the overexpression of inflammation-related genes (Retnlg, Retnlb, Bmp2) induced by T2DM (SI file3.xlsx). Under pathological conditions, the overexpression of resistin-like molecules such as those encoded by Retnlb can aggravate intestinal inflammation and disrupt intestinal barrier integrity by activating the nuclear factor kappa-B (NF-κB) pathway. In contrast, downregulation of this gene can reduce the entry of LPS into the bloodstream, thereby alleviating systemic insulin resistance.75,76 This study further verified that the abnormally upregulated NF-κB pathway in the Mod group was significantly downregulated after FS intervention (Fig. 4D), which was consistent with the aforementioned mechanism. Meanwhile, FS significantly upregulated the expression of metabolism-related genes (Ckm, Klf15, Slc15a2) in the colon (SI file3.xlsx). Among these, Klf15 can enhance the body's ability to regulate postprandial glucose absorption by promoting the expression of glucose transporter 2 (GLUT2);77Ckm is involved in the regulation of energy metabolism by modulating the creatine phosphate shuttle system,78 and preabsorptive upper small intestinal protein-sensing mechanisms mediated by Slc15a2 (PepT1) exert beneficial effects on whole-body glucose homeostasis.79 The “IL-17 signaling pathway” has been consistently implicated in the pathogenesis of chronic inflammatory conditions, including metabolic syndrome.80 Our results demonstrate that FS intervention effectively restores the dysregulated expression of key genes within this pathway, thereby supporting the hypothesis that targeted modulation of the “IL-17 signaling pathway” can attenuate intestinal inflammation and ameliorate associated metabolic disturbances. This finding aligns with previous evidence showing that aberrant activation of the IL-17 axis contributes to insulin resistance, adipose tissue inflammation, and systemic metabolic dysfunction.80,81 Collectively, these findings indicate that FS intervention alleviates colonic inflammatory responses in T2DM rats which may represent a mechanistic link between gut immune homeostasis and improved glucose-lipid metabolism.The liver plays a crucial role in maintaining systemic glucose and lipid homeostasis by regulating glycogen synthesis, glycogen breakdown, as well as lipid oxidation and storage.82,83 This study found that FS could significantly reverse hepatic metabolic gene dysregulation induced by T2DM, specifically manifested as follows: FS targeted and regulated the expression of key genes involved in hepatic lipid metabolism, restoring their levels to the physiological range observed in the Con group (SI file4.xlsx). On the one hand, FS downregulated key lipogenic genes, including Gpam, Lifr, and Srebf1: inhibition of Gpam improves glucose tolerance and lipid metabolism by reducing diacylglycerol accumulation and enhancing insulin signaling.84 Wang et al. confirmed that downregulation of Lifr expression in cells significantly inhibits adipogenic differentiation of preadipocytes, reducing the number of adipocytes by approximately two-thirds.85Srebf1 integrates the regulation of glucose and lipid metabolism by directly activating genes involved in fatty acid and TC biosynthesis.86 On the other hand, FS upregulated the expression of Cpt1a, Slc27a5 and Ugp2: Cpt1a encodes carnitine palmitoyltransferase 1A (CPT1A), a key rate-limiting enzyme in fatty acid β-oxidation; Slc27a5 can synergize with Cpt2 to regulate lipid metabolism by enhancing the catabolism and transport of fatty acids and BAs.87,88 Clinical sample studies have shown that the expression level of UGP2 in patients with non-alcoholic fatty liver disease (NAFLD) is negatively correlated with the expression of lipid synthesis-related genes, suggesting that this gene may be involved in inhibiting lipid synthesis.89
Further transcriptomic pathway enrichment analysis showed that FS exerts “bidirectional synergistic” regulation on the liver (Fig. 5D): on the one hand, FS activates core metabolic pathways, including “pyruvate metabolism, “fatty acid metabolism”, and “fat digestion and absorption” pathways. Similarly, enrichment analysis of DEGs revealed significant upregulation of key metabolic pathways, including “fatty acid degradation” (Fig. S7), “pyruvate metabolism” (Fig. S8), and the “TCA cycle” (Fig. S9). Another study indicated that hydrolyzed guar gum altered fatty acid degradation ameliorates HFD-induced obesity.90 Under aerobic conditions, glucose is generally assumed to be burned fully by tissues via the TCA cycle to carbon dioxide in mammalian tissues.91 Activation of the “pyruvate metabolism” pathway promotes the conversion of glucose to energy, thus the restoration of pyruvate metabolism and TCA cycle activity by FS suggests improved glucose utilization and oxidative phosphorylation, which are critical for glucose metabolic regulation and help reduce lipid accumulation in the liver.92,93 On the other hand, FS inhibits pro-inflammatory signaling networks, including the intestinal IgA immune networks, natural killer (NK) cell-mediated cytotoxicity pathway, and T helper 17 (Th17) cell differentiation pathway. The inhibitory effect of FS on Th17 cell differentiation and NK cell cytotoxicity suggests that it possesses significant anti-inflammatory activity and this effect may be achieved through immunomodulatory mechanisms.94 In addition, pathological analysis of liver tissue further confirmed that FS intervention significantly reduces hepatic lipid deposition and lymphocyte infiltration, directly verifying its hepatoprotective and anti-inflammatory effects.95,96
4.4 FS restores bile acid homeostasis in T2DM rats
BAs are not only key cofactors for the digestion and absorption of dietary fats, but also serve as important signaling molecules involved in the regulation of glucose and lipid metabolism homeostasis.97 This study elucidates the multidimensional regulatory effects of FS intervention on the BA metabolic pathway (Fig. S12). BAs are synthesized in hepatocytes via cytochrome P450-mediated oxidation of cholesterol through two distinct biosynthetic pathways: the classical and alternative pathways.37 In both pathways, FS significantly upregulates the expression of CYP27A1, thereby enhancing the biosynthesis of CA and CDCA. With respect to BA conjugation and excretion, FS upregulates the hepatic expression of UGTs, promoting the formation of glucurono- and sulfo-conjugated BAs. This modification increases BA hydrophilicity and biliary excretion efficiency, effectively alleviating intrahepatic BA accumulation and associated lipid deposition. In the intestine, BAs are further metabolized by microbial enzymes derived from the gut microbiota.37–39 Notably, FS intervention markedly enriches the abundance of Bifidobacterium, which encodes bacterial BSH that catalyzes the deconjugation of glycine- or taurine-conjugated BAs to generate free CA and CDCA. Additionally, the observed elevation in HDCA levels is likely mediated by microbial 7α-dehydroxylation activity conferred by Blautia. Collectively, FS exerts coordinated regulation at key nodes of BA homeostasis, including CYP27A1-mediated hepatic biosynthesis and microbiota-dependent secondary BA generation, thus reshaping the global BA profile.In the present study, FS was found to significantly regulate the serum BA profile: it not only increased the levels of significantly decreased primary BAs (CDCA, α-MCA) and secondary BA (UCA) in the Mod group, restoring them to the levels of the Con group, but also significantly elevated the levels of HDCA and GHDCA in the Mod group, though these two secondary BAs in the FS group were still significantly lower than those in the Con group (Fig. 6), a distinct regulatory pattern that may be attributed to the specific modulation of the gut microbiota–BA axis by FS.
Combined with the observation that CDCA, the synthetic precursor of HDCA, has returned to normal levels in the FS group (Fig. 6D), it suggests that the decrease in HDCA is not due to insufficient substrate availability but rather the inhibition of the microbial transformation process. The conversion of primary BAs into HDCA specifically requires 7α-dehydroxylase activity, an enzymatic function encoded by specific gut bacteria, including members of Clostridium, Eubacterium, and Blautia.37–39 In our study, among these known 7α-dehydroxylating taxa, only Blautia exhibited significant upregulation following FS intervention (Fig. S12); given the absence of notable changes in other canonical 7α-dehydroxylating genera, the overall capacity for secondary BA transformation—particularly toward HDCA—appears to be limited under the current experimental conditions. Thus, despite FS intervention promoting the enrichment of Blautia, the functional output of 7α-dehydroxylation may still remain constrained due to the lack of a broader consortium of dehydroxylating microbes, which ultimately leads to the decrease in HDCA levels. The homeostasis of the BA pool is regulated by a complex network consisting of host synthesis and microbial metabolism. The significant increase in α/β-muricholic acid levels in the FS group (Fig. 6F and G) indicates a shift in host BA synthesis toward the alternative pathway, which in turn alters the composition of the primary BA pool and affects the substrate supply for secondary BA synthesis. The decrease in HDCA levels may be a manifestation of the overall remodeling of the BA profile rather than a single metabolic defect.98 The restoration of normal levels of other secondary BAs such as UCA and LCA in this study further confirms that FS intervention exerts a selective effect on BA metabolic pathways, and this selectivity is particularly prominent in the HDCA synthetic pathway. The failure of HDCA to fully return to normal levels may be associated with the intervention duration and the persistent effect of a HFD. Although 8-week intervention was sufficient to reverse some metabolic indicators, the severe dysbiosis and metabolic disorders induced by a long-term high-fat diet combined with chemical induction may require a longer period to fully restore the function of specific microbiota related to HDCA synthesis. Meanwhile, the continuous intake of a high-fat diet during the intervention period in the FS group may have maintained an intestinal microenvironment unfavorable for the proliferation of HDCA-producing microbiota, thereby offsetting the effect of prebiotics to a certain extent.
Beyond the distinct regulation of HDCA, the restorative effects of FS on other BAs are consistent with previous reports on FOS intervention, which enhances beneficial metabolites like MCA through gut microbiota-mediated 6α-hydroxylation of BAs.99–101 Existing studies have shown that in diabetic patients, CDCA can promote the secretion of glucagon-like peptide-1 (GLP-1) by activating the G protein-coupled BA receptor 1 (GPBAR1).102 Meanwhile, CDCA also possesses the functions of regulating cholesterol metabolism, protecting liver function, and maintaining intestinal barrier integrity.103 Studies on rodents have demonstrated that the taurine-conjugated form of UCA (T-UCA) can improve blood glucose levels, and this effect may be achieved by regulating endoplasmic reticulum (ER) stress in metabolic tissues and/or pancreatic β-cells.104,105 ER stress induces lipogenesis via sterol regulatory element-binding protein SREBP-1c, resulting in hepatic lipid accumulation.106 Additionally, HDCA was reported to significantly increase the abundances of probiotic bacteria such as Parabacteroides distasonis, which can enhance lipid catabolism through modulating the gut–liver axis, thereby alleviating non-alcoholic fatty liver disease.107
Notably, TCDCA was significantly elevated in both Mod and FS groups compared to Con (Fig. 6D). Previous studies have confirmed that a HFD can induce an increase in serum TCDCA concentration in rodents, with the core mechanism as follows: HFD-induced gut microbiota dysregulation impairs BSH activity, inhibiting TCDCA deconjugation and degradation; accumulated TCDCA then accumulates in the upper small intestine (USI) and ileum, crosses the blood–brain barrier to activate farnesoid X receptor (FXR) in the dorsal vagal complex (DVC), which not only blocks insulin signaling in DVC neurons to exacerbate systemic insulin resistance,108 but also disrupts local nutrient-sensing glucoregulatory pathways, consistent with another finding that HFD rats lose the glucose tolerance-improving effect of intestinal nutrient infusion, while transplantation of healthy USI microbiota restores this function by reducing TCDCA and inhibiting FXR.109 However, no reduced BSH activity or significant changes in FXR-encoding gene (Nr1h4) expression were observed in the Mod group, indicating a distinct mechanism for HFD-induced TCDCA elevation from previous studies. We hypothesize that this is likely due to HFD-increased TC—a primary BA precursor—which enhances substrate availability, promotes the synthesis of conjugated BAs like TCDCA, and ultimately leads to its accumulation. The failure to reduce TCDCA levels in the FS group may be attributed to the insufficient increase in the abundance of BSH-producing gut microbiota within the current intervention period. Our study revealed that among BSH-producing intestinal bacteria, only the abundance of Bifidobacterium was significantly increased (Fig. S12). However, the elevated abundance of a single bacterial genus is insufficient to synthesize an adequate amount of BSH, which hinders the efficient conversion and metabolism of TCDCA, ultimately leading to its accumulation in the body. In summary, the elevation of TCDCA levels in both groups may be the result of the combined effect of enhanced hepatic BA synthesis and insufficient gut microbial degradation capacity.
In conclusion, FS can improve the systemic glucose and lipid metabolism homeostasis in rats with T2DM by regulating the homeostasis of the BA pool.
4.5 Multi-omics analyses elucidate the potential mechanism of FS-mediated improvement of glucose and lipid metabolism in T2DM rats
The regulatory network of “gut microbiota–BAs–glycose/lipids–Nr1h4” constructed via Pearson correlation analysis (Fig. 7) provides direct evidence for the multi-target mechanism of action of FS. Previous studies have confirmed that gut microbiota can alter the size and composition of the total BA pool in the enterohepatic system, which is consistent with the results of this study.110 Specifically, the abundance of the key probiotic Bifidobacterium showed a significant negative correlation with TC and TG (|r| > 0.6), and this association may be mediated by enhancing the conversion of cholesterol to HDCA and GHDCA.62Parabacteroides exhibited a similar correlation pattern to Bifidobacterium, while Akkermansia showed no significant correlation with blood glucose or blood lipid parameters. Furthermore, the abundance of Hungatella was strongly negatively correlated with colonic Nr1h4 expression (|r| > 0.8), suggesting that this genus may promote intestinal cholesterol absorption, which is consistent with previous reports linking it to increased cardiovascular risk.111 The inhibitory effect of FS on Hungatella (Fig. 3E) further improves its “anti-inflammatory–metabolic” synergistic regulatory mechanism.After FS intervention, the expression of hepatic Nr1h4 was positively correlated with CDCA, HDCA and GHDCA (|r | >0.6), confirming the core role of FXR in regulating hepatic lipid synthesis and excretion. In contrast, colonic Nr1h4 was negatively correlated with TC and LDL-C (|r| > 0.6), suggesting that intestinal FXR signaling may be involved in inhibiting cholesterol absorption. Meanwhile, the positive correlation between colonic Nr1h4 expression and CDCA indicates that intestinal FXR can inhibit hepatic BA synthesis through the fibroblast growth factor 15 (FGF15) pathway and promote the release of glucagon-like peptide-1 (GLP-1) by activating the G protein-coupled receptor (GPBAR1), ultimately synergistically regulating glycolipid metabolism.112,113
We further explored the molecular and microbial changes associated with FS intervention (Fig. S5). Notably, FS treatment significantly upregulated the hepatic expression of CYP27A1 and UGTs and increased the abundance of Bifidobacterium and Blautia. These coordinated changes in both host enzymatic machinery and gut microbial composition suggest that CYP27A1, UGTs, Bifidobacterium, and Blautia may collectively play pivotal roles in mediating the lipid-lowering effects of FS through modulation of BA metabolism.
4.6 The absorption, distribution, metabolism, and excretion properties of FOS and sea buckthorn
To further clarify the pharmacodynamic basis of FS-mediated metabolic improvements, it is necessary to elaborate on the absorption, distribution, metabolism, and excretion (ADME) properties of its core components (FOS and sea buckthorn constituents) and distinguish their direct systemic effects from indirect regulation via gut microbiota.As a water-soluble dietary fiber, FOS resist hydrolysis by digestive enzymes in the saliva, stomach and small intestine and are barely absorbed in the upper gastrointestinal tract, thus reaching the large intestine in an intact form. In the colon, FOS are selectively fermented by commensal beneficial bacteria (e.g., Bifidobacterium and Lactobacillus) to generate SCFAs including acetate, propionate and butyrate.114 Owing to their poor bioavailability in the upper gastrointestinal tract, FOS lack direct systemic biological activity and their favorable effects on glycolipid metabolism are instead mediated indirectly via gut microbiota modulation, a mechanism that involves promoting probiotic proliferation through nutritional competition and subsequent enhancement of SCFA production.114 Further evidence for this indirect regulatory mode comes from antibiotic intervention assays and [14C]-FOS tracer experiments, which demonstrated complete abrogation of FOS-associated metabolic protective effects following gut microbiota depletion.115,116 In contrast, sea buckthorn contains diverse bioactive components (e.g., flavonoids, polysaccharides, triterpenic acids) with distinct ADME characteristics, leading to a combination of indirect and direct regulatory effects. Sea buckthorn polysaccharide (SP) can promote the production of SCFAs by remodeling the structure of gut microbiota and then regulate the expression of proteins related to hepatic lipid metabolism (e.g., p-AMPKα, PPARα, FAS, etc.) through the gut microbiota–SCFAs–liver axis, thereby exerting lipid-lowering effects.22,117 Rutin in total flavones of sea buckthorn can be microbially converted into absorbable quercetin aglycone in the ileum and colon;118 isorhamnetin-3-O-glucoside can be metabolized by bacteria into its aglycone, isorhamnetin.119 However, isorhamnetin, kaempferol, and quercetin can be absorbed via oral administration and enter the systemic circulation, possessing the potential to exert direct systemic effects;120 triterpenic acids target the liver, activate the AMPK/Nrf2/NF-κB signaling pathway, and suppress the expression of fatty acid synthase (FAS), directly alleviating hepatic lipid deposition and oxidative stress.121
In summary, FS may maintain glycolipid metabolic homeostasis through a dual regulatory mode: on the one hand, by improving the gut microbiota structure and restoring BA profiles to activate the FXR-mediated multi-tissue metabolic regulatory network;122 on the other hand, by leveraging the distinct ADME characteristics of its core components (indirect regulation by FOS and combined direct–indirect effects by sea buckthorn constituents), which together synergistically alleviate metabolic disorders in T2DM rats.
5. Conclusion
Administration of FS is associated with improved glucose and lipid metabolism in T2DM rats, accompanied by multi-target regulatory effects including gut microbiota remodeling, attenuation of hepatic and intestinal inflammation, and modulation of BA and fatty acid metabolism. Furthermore, mediation analysis revealed that the beneficial effects of FS intervention on TC, LDL-C, and oral OGTT outcomes were partially mediated through the modulation of Phocae and Desulfovibrionaceae_uncultured (Fig. S4), which may serve as key microbial intermediaries linking FS-induced gut microbiota remodeling to systemic lipid and glucose homeostasis. These findings suggest that the metabolic benefits of FS are likely associated with the reshaping of the gut microbiota, although further experiments such as fecal microbiota transplantation or antibiotic intervention are still needed to substantiate the definitive causal and mechanistic relationships.6. Research limitations and future perspectives
Only male Sprague-Dawley (SD) rats were used and gender differences were not considered. Male-specific factors, particularly testosterone levels, may act as confounding variables that affect the study results. Testosterone, a key male sex hormone, has been confirmed to regulate glucolipid metabolism by modulating skeletal muscle glucose transport and lipid accumulation. For instance, in T2DM rats, testosterone and its metabolite 5α-dihydrotestosterone can activate the translocation of glucose transporter 4 (GLUT-4) in skeletal muscle, thereby improving glucose utilization.123 Testosterone deficiency is associated with increased ectopic lipid accumulation in the liver, which exacerbates dysregulation of glucolipid metabolism.124 Meanwhile, testosterone also interacts with gut microbiota: in male patients with T2DM;125 testosterone deficiency is correlated with altered gut microbiota composition, such as increased abundance of Blautia and Lachnospirales and the phylum Firmicutes in the gut is positively associated with serum testosterone levels in elderly men.126 Since our study focused on the effect of FS on glucolipid metabolism and gut microbiota in T2DM rats, the high testosterone level in male models may have synergistically or antagonistically affected the experimental effects of FS. This confounding effect may limit the generalizability of our findings to female models, as females have distinct sex hormone profiles (e.g., dominant estrogen) and corresponding differences in gut microbiota composition.127 Therefore, the effect of FS in female models should be verified in subsequent research and studies with gonadectomized models or hormone supplementation experiments are recommended to further clarify the interaction between sex hormones, FS, and the gut microbiota–glucolipid metabolism axis.Author contributions
Ling Chen contributed to conceptualisation, data curation, formal analysis, methodology and writing; Hualin Liu: contributed to conceptualisation, data curation, methodology, software and writing; Yanwu Chen contributed to data curation and methodology; Qianya Xiao contributed to data curation and methodology; Zijin Huang contributed to data curation and methodology; Ziying Li contributed to data curation and methodology; Xiaofei Zhang contributed to data curation and methodology; Xueying Fan contributed to data curation and methodology; Meichen Xiao contributed to data curation and methodology; Xiaomin Li contributed to funding acquisition, conceptualisation, project administration and supervision; Yuan Yao Chen contributed to funding acquisition, conceptualisation, project administration, supervision and writing.Conflicts of interest
There are no conflicts to declare.Data availability
All data generated in this study have been deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA) under the BioProject accession number PRJNA1356967. All relevant data are publicly available through this accession.The supplementary information accompanying this article comprises 13 supplementary figures (Fig. S1–Fig. S13) and five supplementary files. These include data on the synergistic effects of fructo-oligosaccharides and sea buckthorn on glucose, triglyceride, and total cholesterol levels in zebrafish larvae (Supplementary File 1), supplementary tables (Table S1 and Table S2, Supplementary File 2), transcription profiles of colonic-associated genes (Supplementary File 3), transcription profiles of liver-related genes (Supplementary File 4), and correlation analyses between key microbiota, bile acids, and blood glucose and lipid parameters (Supplementary File 5). See DOI: https://doi.org/10.1039/d5fo04996b.
Acknowledgements
This work was funded by grants from the Perfect (Guangdong) Co., Ltd and Perfect Life & Health Institute Co., Ltd, Zhongshan, China.References
- GBD 2021 Diabetes Collaborators, Global, Regional, and National Burden of Diabetes from 1990 to 2021, with Projections of Prevalence to 2050: A Systematic Analysis for the Global Burden of Disease Study 2021, Lancet, 2023, 402(10397), 203–234, DOI:10.1016/S0140-6736(23)01301-6.
- D. J. Magliano and E. J. Boyko, IDF Diabetes Atlas 10th edition scientific committee, in IDF Diabetes Atlas, International Diabetes Federation, Brussels, 2021 Search PubMed.
- M. J. Davies, V. R. Aroda, B. S. Collins, R. A. Gabbay, J. Green, N. M. Maruthur, S. E. Rosas, S. Del Prato, C. Mathieu, G. Mingrone, P. Rossing, T. Tankova, A. Tsapas and J. B. Buse, Management of Hyperglycemia in Type 2 Diabetes, 2022. A Consensus Report by the American Diabetes Association (ADA) and the European Association for the Study of Diabetes (EASD), Diabetes Care, 2022, 45(11), 2753–2786, DOI:10.2337/dci22-0034.
- Metformin-induced vitamin B12 deficiency can cause or worsen distal symmetrical, autonomic and cardiac neuropathy in patients with diabetes – PubMed, https://pubmed.ncbi.nlm.nih.gov/35491956/, accessed 2025-08-19.
- L. Liu, J. Zhang, Y. Cheng, M. Zhu, Z. Xiao, G. Ruan and Y. Wei, Gut Microbiota: A New Target for T2DM Prevention and Treatment, Front. Endocrinol., 2022, 13, 958218, DOI:10.3389/fendo.2022.958218.
- J. Qin, Y. Li, Z. Cai, S. Li, J. Zhu, F. Zhang, S. Liang, W. Zhang, Y. Guan, D. Shen, Y. Peng, D. Zhang, Z. Jie, W. Wu, Y. Qin, W. Xue, J. Li, L. Han, D. Lu, P. Wu, Y. Dai, X. Sun, Z. Li, A. Tang, S. Zhong, X. Li, W. Chen, R. Xu, M. Wang, Q. Feng, M. Gong, J. Yu, Y. Zhang, M. Zhang, T. Hansen, G. Sanchez, J. Raes, G. Falony, S. Okuda, M. Almeida, E. LeChatelier, P. Renault, N. Pons, J.-M. Batto, Z. Zhang, H. Chen, R. Yang, W. Zheng, S. Li, H. Yang, J. Wang, S. D. Ehrlich, R. Nielsen, O. Pedersen, K. Kristiansen and J. Wang, A Metagenome-Wide Association Study of Gut Microbiota in Type 2 Diabetes, Nature, 2012, 490(7418), 55–60, DOI:10.1038/nature11450.
- K. H. Allin, V. Tremaroli, R. Caesar, B. A. H. Jensen, M. T. F. Damgaard, M. I. Bahl, T. R. Licht, T. H. Hansen, T. Nielsen, T. M. Dantoft, A. Linneberg, T. Jørgensen, H. Vestergaard, K. Kristiansen, P. W. Franks, IMI-DIRECT consortium, T. Hansen, F. Bäckhed and O. Pedersen, Aberrant Intestinal Microbiota in Individuals with Prediabetes, Diabetologia, 2018, 61(4), 810–820, DOI:10.1007/s00125-018-4550-1.
- H. Wu, V. Tremaroli, C. Schmidt, A. Lundqvist, L. M. Olsson, M. Krämer, A. Gummesson, R. Perkins, G. Bergström and F. Bäckhed, The Gut Microbiota in Prediabetes and Diabetes: A Population-Based Cross-Sectional Study, Cell Metab., 2020, 32(3), 379–390, DOI:10.1016/j.cmet.2020.06.011.
- D. Davani-Davari, M. Negahdaripour, I. Karimzadeh, M. Seifan, M. Mohkam, S. J. Masoumi, A. Berenjian and Y. Ghasemi, Prebiotics: Definition, Types, Sources, Mechanisms, and Clinical Applications, Foods, 2019, 8(3), 92, DOI:10.3390/foods8030092.
- X. Ding, Y. Chen, L. Du, J. Li, X. Meng, H. Lv, B. Tong, G. Niu, T. Jian and J. Chen, Benefits of Inulin and Fructo-Oligosaccharides on High Fat Diet-Induced Type 2 Diabetes Mellitus by Regulating the Gut Microbiota in Mice, J. Nutr. Biochem., 2025, 141, 109908, DOI:10.1016/j.jnutbio.2025.109908.
- F. Liu, M. Prabhakar, J. Ju, H. Long and H.-W. Zhou, Effect of Inulin-Type Fructans on Blood Lipid Profile and Glucose Level: A Systematic Review and Meta-Analysis of Randomized Controlled Trials, Eur. J. Clin. Nutr., 2017, 71(1), 9–20, DOI:10.1038/ejcn.2016.156.
- L. Zhao, F. Zhang, X. Ding, G. Wu, Y. Y. Lam, X. Wang, H. Fu, X. Xue, C. Lu, J. Ma, L. Yu, C. Xu, Z. Ren, Y. Xu, S. Xu, H. Shen, X. Zhu, Y. Shi, Q. Shen, W. Dong, R. Liu, Y. Ling, Y. Zeng, X. Wang, Q. Zhang, J. Wang, L. Wang, Y. Wu, B. Zeng, H. Wei, M. Zhang, Y. Peng and C. Zhang, Gut Bacteria Selectively Promoted by Dietary Fibers Alleviate Type 2 Diabetes, Science, 2018, 359(6380), 1151–1156, DOI:10.1126/science.aao5774.
- W. Wang, Z. Fan, Q. Yan, T. Pan, J. Luo, Y. Wei, B. Li, Z. Fang and W. Lu, Gut Microbiota Determines the Fate of Dietary Fiber-Targeted Interventions in Host Health, Gut Microbes, 2024, 16(1), 2416915, DOI:10.1080/19490976.2024.2416915.
- C. Le Bourgot, E. Apper, S. Blat and F. Respondek, Fructo-Oligosaccharides and Glucose Homeostasis: A Systematic Review and Meta-Analysis in Animal Models, Nutr. Metab., 2018, 15, 9, DOI:10.1186/s12986-018-0245-3.
- Q. Ma, R. Du, P. Long, K. Sun, Y. Wang, Y. Yang, X. Shen and L. Gao, The Protective Effects of Burdock Fructooligosaccharide on Preterm Labor Through Its Anti-Inflammatory Action, Int. J. Mol. Sci., 2025, 26(6), 2659, DOI:10.3390/ijms26062659.
- N. Pengrattanachot, L. Thongnak and A. Lungkaphin, The Impact of Prebiotic Fructooligosaccharides on Gut Dysbiosis and Inflammation in Obesity and Diabetes Related Kidney Disease, Food Funct., 2022, 13(11), 5925–5945, 10.1039/d1fo04428a.
- Y. Chen, Y. Cai, K. Wang and Y. Wang, Bioactive Compounds in Sea Buckthorn and Their Efficacy in Preventing and Treating Metabolic Syndrome, Foods, 2023, 12(10), 1985, DOI:10.3390/foods12101985.
- J. Wang, W. Zhang, D. Zhu, X. Zhu, X. Pang and W. Qu, Hypolipidaemic and Hypoglycaemic Effects of Total Flavonoids from Seed Residues of Hippophae Rhamnoides L. in Mice Fed a High-Fat Diet, J. Sci. Food Agric., 2011, 91(8), 1446–1451, DOI:10.1002/jsfa.4331.
- C. Song, J. Du and H. Ge, Research of Hippophae rhamnoides fruits on serum lipids and liver protection effects in high-fat-diet rats, Weisheng Yanjiu, 2015, 44(4), 628–631 Search PubMed.
- Y. Bai, J. Li, X. Wu, M. Zhang, Y. Zhang, P. Chen, J. Ma, S. Zhang, H. Zhang, X. Li and Z. Yang, Mult-Omics Analysis Reveals the Lipid-Lowering Effects of Sea Buckthorn and Milk Thistle Solid Beverage in Hyperlipidemic Rats, Phytomedicine, 2025, 144, 156920, DOI:10.1016/j.phymed.2025.156920.
- Y. Chen, Y. Cai, K. Wang and Y. Wang, Bioactive Compounds in Sea Buckthorn and Their Efficacy in Preventing and Treating Metabolic Syndrome, Foods, 2023, 12(10), 1985, DOI:10.3390/foods12101985.
- Q. Ouyang, X. Li, Y. Liang and R. Liu, Sea Buckthorn Polysaccharide Ameliorates Colitis, Nutrients, 2024, 16(9), 1280, DOI:10.3390/nu16091280.
- H. Yuan, F. Shi, L. Meng and W. Wang, Effect of Sea Buckthorn Protein on the Intestinal Microbial Community in Streptozotocin-Induced Diabetic Mice, Int. J. Biol. Macromol., 2018, 107(Pt A), 1168–1174, DOI:10.1016/j.ijbiomac.2017.09.090.
- J. Li, L. Zhang, J. Xin, L. Jiang, J. Li, T. Zhang, L. Jin, J. Li, P. Zhou, S. Hao, H. Cao and L. Li, Immediate Intraportal Transplantation of Human Bone Marrow Mesenchymal Stem Cells Prevents Death from Fulminant Hepatic Failure in Pigs, Hepatology, 2012, 56(3), 1044–1052, DOI:10.1002/hep.25722.
- A. M. Bolger, M. Lohse and B. Usadel, Trimmomatic: A Flexible Trimmer for Illumina Sequence Data, Bioinformatics, 2014, 30(15), 2114–2120, DOI:10.1093/bioinformatics/btu170.
- E. Bolyen, J. R. Rideout, M. R. Dillon, N. A. Bokulich, C. C. Abnet, G. A. Al-Ghalith, H. Alexander, E. J. Alm, M. Arumugam, F. Asnicar, Y. Bai, J. E. Bisanz, K. Bittinger, A. Brejnrod, C. J. Brislawn, C. T. Brown, B. J. Callahan, A. M. Caraballo-Rodríguez, J. Chase, E. K. Cope, R. Da Silva, C. Diener, P. C. Dorrestein, G. M. Douglas, D. M. Durall, C. Duvallet, C. F. Edwardson, M. Ernst, M. Estaki, J. Fouquier, J. M. Gauglitz, S. M. Gibbons, D. L. Gibson, A. Gonzalez, K. Gorlick, J. Guo, B. Hillmann, S. Holmes, H. Holste, C. Huttenhower, G. A. Huttley, S. Janssen, A. K. Jarmusch, L. Jiang, B. D. Kaehler, K. B. Kang, C. R. Keefe, P. Keim, S. T. Kelley, D. Knights, I. Koester, T. Kosciolek, J. Kreps, M. G. I. Langille, J. Lee, R. Ley, Y.-X. Liu, E. Loftfield, C. Lozupone, M. Maher, C. Marotz, B. D. Martin, D. McDonald, L. J. McIver, A. V. Melnik, J. L. Metcalf, S. C. Morgan, J. T. Morton, A. T. Naimey, J. A. Navas-Molina, L. F. Nothias, S. B. Orchanian, T. Pearson, S. L. Peoples, D. Petras, M. L. Preuss, E. Pruesse, L. B. Rasmussen, A. Rivers, M. S. Robeson, P. Rosenthal, N. Segata, M. Shaffer, A. Shiffer, R. Sinha, S. J. Song, J. R. Spear, A. D. Swafford, L. R. Thompson, P. J. Torres, P. Trinh, A. Tripathi, P. J. Turnbaugh, S. Ul-Hasan, J. J. J. van der Hooft, F. Vargas, Y. Vázquez-Baeza, E. Vogtmann, M. von Hippel, W. Walters, Y. Wan, M. Wang, J. Warren, K. C. Weber, C. H. D. Williamson, A. D. Willis, Z. Z. Xu, J. R. Zaneveld, Y. Zhang, Q. Zhu, R. Knight and J. G. Caporaso, Reproducible, Interactive, Scalable and Extensible Microbiome Data Science Using QIIME 2, Nat. Biotechnol., 2019, 37(8), 852–857, DOI:10.1038/s41587-019-0209-9.
- B. J. Callahan, P. J. McMurdie, M. J. Rosen, A. W. Han, A. J. A. Johnson and S. P. Holmes, DADA2: High-Resolution Sample Inference from Illumina Amplicon Data, Nat. Methods, 2016, 13(7), 581–583, DOI:10.1038/nmeth.3869.
- N. Segata, J. Izard, L. Waldron, D. Gevers, L. Miropolsky, W. S. Garrett and C. Huttenhower, Metagenomic Biomarker Discovery and Explanation, Genome Biol., 2011, 12(6), R60, DOI:10.1186/gb-2011-12-6-r60.
- H. M. Hassan, Q. Cai, X. Liang, J. Xin, K. Ren, J. Jiang, D. Shi, Y. Lu, T. Li, Y. Shang, L. He, X. Chen, S. Sun, P. Li, B. Guo, J. Chen, H. Yang, W. Hu, X. Chen and J. Li, Transcriptomics Reveals Immune-Metabolism Disorder in Acute-on-Chronic Liver Failure in Rats, Life Sci. Alliance, 2021, 5(3), e202101189, DOI:10.26508/lsa.202101189.
- R. Patro, G. Duggal, M. I. Love, R. A. Irizarry and C. Kingsford, Salmon Provides Fast and Bias-Aware Quantification of Transcript Expression, Nat. Methods, 2017, 14(4), 417–419, DOI:10.1038/nmeth.4197.
- T. Wu, E. Hu, S. Xu, M. Chen, P. Guo, Z. Dai, T. Feng, L. Zhou, W. Tang, L. Zhan, X. Fu, S. Liu, X. Bo and G. Yu, clusterProfiler 4.0: A Universal Enrichment Tool for Interpreting Omics Data, Innovation, 2021, 2(3), 100141, DOI:10.1016/j.xinn.2021.100141.
- psych: Procedures for Personality and Psychological Research - Northwestern Scholars, https://www.scholars.northwestern.edu/en/publications/psych-procedures-for-personality-and-psychological-research (accessed 2025-08-21).
- H. Wickham, M. Averick, J. Bryan, W. Chang, L. D. McGowan, R. François, G. Grolemund, A. Hayes, L. Henry, J. Hester, M. Kuhn, T. L. Pedersen, E. Miller, S. M. Bache, K. Müller, J. Ooms, D. Robinson, D. P. Seidel, V. Spinu, K. Takahashi, D. Vaughan, C. Wilke, K. Woo and H. Yutani, Welcome to the Tidyverse, J. Open Source Softw., 2019, 4(43), 1686, DOI:10.21105/joss.01686.
- H. Wickham, ggplot2: Elegant Graphics for Data Analysis, Springer-Verlag New York, DOI:10.1007/978-3-319-24277-4.
- G. Csárdi, T. Nepusz, V. Traag, S. Horvát, F. Zanini, D. Noom, K. Müller, M. Salmon and M. Antonov, C. Z. I. details, igraph author, Igraph: Network Analysis and Visualization, 2025, https://cran.r-project.org/web/packages/igraph/index.html (accessed 2025-08-21) Search PubMed.
- X. Su, Y. Gao and R. Yang, Gut Microbiota-Derived Tryptophan Metabolites Maintain Gut and Systemic Homeostasis, Cells, 2022, 11(15), 2296, DOI:10.3390/cells11152296.
- W. Jia, G. Xie and W. Jia, Bile Acid-Microbiota Crosstalk in Gastrointestinal Inflammation and Carcinogenesis, Nat. Rev. Gastroenterol. Hepatol., 2018, 15(2), 111–128, DOI:10.1038/nrgastro.2017.119.
- W. Jia, Y. Li, K. C. P. Cheung and X. Zheng, Bile Acid Signaling in the Regulation of Whole Body Metabolic and Immunological Homeostasis, Sci. China:Life Sci., 2024, 67(5), 865–878, DOI:10.1007/s11427-023-2353-0.
- J. Chen, Y. Xiao, D. Li, S. Zhang, Y. Wu, Q. Zhang and W. Bai, New Insights into the Mechanisms of High-Fat Diet Mediated Gut Microbiota in Chronic Diseases, iMeta, 2023, 2(1), e69, DOI:10.1002/imt2.69.
- American Diabetes Association, 6. Glycemic Targets: Standards of Medical Care in Diabetes-2021, Diabetes Care, 2021, 44(Suppl 1), S73–S84, DOI:10.2337/dc21-S006.
- V. K. Ridaura, J. J. Faith, F. E. Rey, J. Cheng, A. E. Duncan, A. L. Kau, N. W. Griffin, V. Lombard, B. Henrissat, J. R. Bain, M. J. Muehlbauer, O. Ilkayeva, C. F. Semenkovich, K. Funai, D. K. Hayashi, B. J. Lyle, M. C. Martini, L. K. Ursell, J. C. Clemente, W. Van Treuren, W. A. Walters, R. Knight, C. B. Newgard, A. C. Heath and J. I. Gordon, Gut Microbiota from Twins Discordant for Obesity Modulate Metabolism in Mice, Science, 2013, 341(6150), 1241214, DOI:10.1126/science.1241214.
- E. E. Canfora, J. W. Jocken and E. E. Blaak, Short-Chain Fatty Acids in Control of Body Weight and Insulin Sensitivity, Nat. Rev. Endocrinol., 2015, 11(10), 577–591, DOI:10.1038/nrendo.2015.128.
- T. Yang, Y. Liu, L. Li, Y. Zheng, Y. Wang, J. Su, R. Yang, M. Luo and C. Yu, Correlation between the triglyceride-to-high-density lipoprotein cholesterol ratio and other unconventional lipid parameters with the risk of prediabetes and Type 2 diabetes in patients with coronary heart disease: a RCSCD-TCM study in China, Cardiovasc. Diabetol., 2022, 21, 93, DOI:10.1186/s12933-022-01531-7.
- B. Schnabl and D. A. Brenner, Interactions between the Intestinal Microbiome and Liver Diseases, Gastroenterology, 2014, 146(6), 1513–1524, DOI:10.1053/j.gastro.2014.01.020.
- P. D. Cani, R. Bibiloni, C. Knauf, A. Waget, A. M. Neyrinck, N. M. Delzenne and R. Burcelin, Changes in Gut Microbiota Control Metabolic Endotoxemia-Induced Inflammation in High-Fat Diet-Induced Obesity and Diabetes in Mice, Diabetes, 2008, 57(6), 1470–1481, DOI:10.2337/db07-1403.
- A. Wahlström, S. I. Sayin, H.-U. Marschall and F. Bäckhed, Intestinal Crosstalk between Bile Acids and Microbiota and Its Impact on Host Metabolism, Cell Metab., 2016, 24(1), 41–50, DOI:10.1016/j.cmet.2016.05.005.
- F. Li, C. Jiang, K. W. Krausz, Y. Li, I. Albert, H. Hao, K. M. Fabre, J. B. Mitchell, A. D. Patterson and F. J. Gonzalez, Microbiome Remodelling Leads to Inhibition of Intestinal Farnesoid X Receptor Signalling and Decreased Obesity, Nat. Commun., 2013, 4, 2384, DOI:10.1038/ncomms3384.
- J. Qin, Y. Li, Z. Cai, S. Li, J. Zhu, F. Zhang, S. Liang, W. Zhang, Y. Guan, D. Shen, Y. Peng, D. Zhang, Z. Jie, W. Wu, Y. Qin, W. Xue, J. Li, L. Han, D. Lu, P. Wu, Y. Dai, X. Sun, Z. Li, A. Tang, S. Zhong, X. Li, W. Chen, R. Xu, M. Wang, Q. Feng, M. Gong, J. Yu, Y. Zhang, M. Zhang, T. Hansen, G. Sanchez, J. Raes, G. Falony, S. Okuda, M. Almeida, E. LeChatelier, P. Renault, N. Pons, J.-M. Batto, Z. Zhang, H. Chen, R. Yang, W. Zheng, S. Li, H. Yang, J. Wang, S. D. Ehrlich, R. Nielsen, O. Pedersen, K. Kristiansen and J. Wang, A Metagenome-Wide Association Study of Gut Microbiota in Type 2 Diabetes, Nature, 2012, 490(7418), 55–60, DOI:10.1038/nature11450.
- S. A. Joyce, J. MacSharry, P. G. Casey, M. Kinsella, E. F. Murphy, F. Shanahan, C. Hill and C. G. M. Gahan, Regulation of Host Weight Gain and Lipid Metabolism by Bacterial Bile Acid Modification in the Gut, Proc. Natl. Acad. Sci. U. S. A., 2014, 111(20), 7421–7426, DOI:10.1073/pnas.1323599111.
- P. D. Cani, A. Everard and T. Duparc, Gut Microbiota, Enteroendocrine Functions and Metabolism, Curr. Opin. Pharmacol., 2013, 13(6), 935–940, DOI:10.1016/j.coph.2013.09.008.
- J. Gu, B. Mao, S. Cui, X. Tang, Z. Liu, J. Zhao and H. Zhang, Bifidobacteria Exhibited Stronger Ability to Utilize Fructooligosaccharides, Compared with Other Bacteria in the Mouse Intestine, J. Sci. Food Agric., 2022, 102(6), 2413–2423, DOI:10.1002/jsfa.11580.
- J. Gu, S. Cui, X. Tang, Z. Liu, J. Zhao, H. Zhang, B. Mao and W. Chen, Effects of Fructooligosaccharides (FOS) on the Composition of Cecal and Fecal Microbiota and the Quantitative Detection of FOS-Metabolizing Bacteria Using Species-Specific Primers, J. Sci. Food Agric., 2022, 102(12), 5301–5311, DOI:10.1002/jsfa.11884.
- S. Y. Geerlings, I. Kostopoulos, W. M. De Vos and C. Belzer, Akkermansia Muciniphila in the Human Gastrointestinal Tract: When, Where, and How?, Microorganisms, 2018, 6(3), 75, DOI:10.3390/microorganisms6030075.
- Z. Song, Z. Qiao, J. Liu, L. Han, X. Chen and Y. Wang, Sea Buckthorn Berries Alleviate Ulcerative Colitis via Regulating Gut Faecalibaculum Rodentium-Mediated Butyrate Biosynthesis, Phytomedicine, 2025, 139, 156490, DOI:10.1016/j.phymed.2025.156490.
- X. Liu, B. Mao, J. Gu, J. Wu, S. Cui, G. Wang, J. Zhao, H. Zhang and W. Chen, Blautia—a new functional genus with potential probiotic properties?, Gut Microbes, 2021, 13(1) DOI:10.1080/19490976.2021.1875796.
- X. Tong, J. Xu, F. Lian, X. Yu, Y. Zhao, L. Xu, M. Zhang, X. Zhao, J. Shen, S. Wu, X. Pang, J. Tian, C. Zhang, Q. Zhou, L. Wang, B. Pang, F. Chen, Z. Peng, J. Wang, Z. Zhen, C. Fang, M. Li, L. Chen and L. Zhao, Structural Alteration of Gut Microbiota during the Amelioration of Human Type 2 Diabetes with Hyperlipidemia by Metformin and a Traditional Chinese Herbal Formula: A Multicenter, Randomized, Open Label Clinical Trial, mBio, 2018, 9(3), e02392–17, DOI:10.1128/mBio.02392-17.
- C. Depommier, A. Everard, C. Druart, H. Plovier, M. Van Hul, S. Vieira-Silva, G. Falony, J. Raes, D. Maiter, N. M. Delzenne, M. de Barsy, A. Loumaye, M. P. Hermans, J.-P. Thissen, W. M. de Vos and P. D. Cani, Supplementation with Akkermansia Muciniphila in Overweight and Obese Human Volunteers: A Proof-of-Concept Exploratory Study, Nat. Med., 2019, 25(7), 1096–1103, DOI:10.1038/s41591-019-0495-2.
- H. Plovier, A. Everard, C. Druart, C. Depommier, M. Van Hul, L. Geurts, J. Chilloux, N. Ottman, T. Duparc, L. Lichtenstein, A. Myridakis, N. M. Delzenne, J. Klievink, A. Bhattacharjee, K. C. H. van der Ark, S. Aalvink, L. O. Martinez, M.-E. Dumas, D. Maiter, A. Loumaye, M. P. Hermans, J.-P. Thissen, C. Belzer, W. M. de Vos and P. D. Cani, A Purified Membrane Protein from Akkermansia Muciniphila or the Pasteurized Bacterium Improves Metabolism in Obese and Diabetic Mice, Nat. Med., 2017, 23(1), 107–113, DOI:10.1038/nm.4236.
- L. Zhao, F. Zhang, X. Ding, G. Wu, Y. Y. Lam, X. Wang, H. Fu, X. Xue, C. Lu, J. Ma, L. Yu, C. Xu, Z. Ren, Y. Xu, S. Xu, H. Shen, X. Zhu, Y. Shi, Q. Shen, W. Dong, R. Liu, Y. Ling, Y. Zeng, X. Wang, Q. Zhang, J. Wang, L. Wang, Y. Wu, B. Zeng, H. Wei, M. Zhang, Y. Peng and C. Zhang, Gut Bacteria Selectively Promoted by Dietary Fibers Alleviate Type 2 Diabetes, Science, 2018, 359(6380), 1151–1156, DOI:10.1126/science.aao5774.
- P. D. Cani, A. M. Neyrinck, F. Fava, C. Knauf, R. G. Burcelin, K. M. Tuohy, G. R. Gibson and N. M. Delzenne, Selective Increases of Bifidobacteria in Gut Microflora Improve High-Fat-Diet-Induced Diabetes in Mice through a Mechanism Associated with Endotoxaemia, Diabetologia, 2007, 50(11), 2374–2383, DOI:10.1007/s00125-007-0791-0.
- P. D. Cani, J. Amar, M. A. Iglesias, M. Poggi, C. Knauf, D. Bastelica, A. M. Neyrinck, F. Fava, K. M. Tuohy, C. Chabo, A. Waget, E. Delmée, B. Cousin, T. Sulpice, B. Chamontin, J. Ferrières, J.-F. Tanti, G. R. Gibson, L. Casteilla, N. M. Delzenne, M. C. Alessi and R. Burcelin, Metabolic Endotoxemia Initiates Obesity and Insulin Resistance, Diabetes, 2007, 56(7), 1761–1772, DOI:10.2337/db06-1491.
- R. Caesar, C. S. Reigstad, H. K. Bäckhed, C. Reinhardt, M. Ketonen, G.Ö Lundén, P. D. Cani and F. Bäckhed, Gut-Derived Lipopolysaccharide Augments Adipose Macrophage Accumulation but Is Not Essential for Impaired Glucose or Insulin Tolerance in Mice, Gut, 2012, 61(12), 1701–1707, DOI:10.1136/gutjnl-2011-301689.
- F. Bäckhed, H. Ding, T. Wang, L. V. Hooper, G. Y. Koh, A. Nagy, C. F. Semenkovich and J. I. Gordon, The Gut Microbiota as an Environmental Factor That Regulates Fat Storage, Proc. Natl. Acad. Sci. U. S. A., 2004, 101(44), 15718–15723, DOI:10.1073/pnas.0407076101.
- H. Tilg and A. R. Moschen, Microbiota and Diabetes: An Evolving Relationship, Gut, 2014, 63(9), 1513–1521, DOI:10.1136/gutjnl-2014-306928.
- L. Henneke, K. Schlicht, N. A. Andreani, T. Hollstein, T. Demetrowitsch, C. Knappe, K. Hartmann, J. Jensen-Kroll, N. Rohmann, D. Pohlschneider, C. Geisler, D. M. Schulte, U. Settgast, K. Türk, J. Zimmermann, C. Kaleta, J. F. Baines, J. Shearer, S. Shah, G. Shen-Tu, K. Schwarz, A. Franke, S. Schreiber and M. Laudes, A Dietary Carbohydrate - Gut Parasutterella - Human Fatty Acid Biosynthesis Metabolic Axis in Obesity and Type 2 Diabetes, Gut Microbes, 2022, 14(1), 2057778, DOI:10.1080/19490976.2022.2057778.
- T. Zhao, L. Zhan, W. Zhou, W. Chen, J. Luo, L. Zhang, Z. Weng, C. Zhao and S. Liu, The Effects of Erchen Decoction on Gut Microbiota and Lipid Metabolism Disorders in Zucker Diabetic Fatty Rats, Front. Pharmacol., 2021, 12, 647529, DOI:10.3389/fphar.2021.647529.
- Z. A. Barandouzi, A. R. Starkweather, W. A. Henderson, A. Gyamfi and X. S. Cong, Altered Composition of Gut Microbiota in Depression: A Systematic Review, Front. Psychiatry, 2020, 11, 541, DOI:10.3389/fpsyt.2020.00541.
- J.-H. Jang, M.-J. Yeom, S. Ahn, J.-Y. Oh, S. Ji, T.-H. Kim and H.-J. Park, Acupuncture Inhibits Neuroinflammation and Gut Microbial Dysbiosis in a Mouse Model of Parkinson's Disease, Brain, Behav., Immun., 2020, 89, 641–655, DOI:10.1016/j.bbi.2020.08.015.
- J. M. Moreno-Navarrete, M. Serino, V. Blasco-Baque, V. Azalbert, R. H. Barton, M. Cardellini, J. Latorre, F. Ortega, M. Sabater-Masdeu, R. Burcelin, M.-E. Dumas, W. Ricart, M. Federici and J. M. Fernández-Real, Gut Microbiota Interacts with Markers of Adipose Tissue Browning, Insulin Action and Plasma Acetate in Morbid Obesity, Mol. Nutr. Food Res., 2018, 62(3) DOI:10.1002/mnfr.201700721.
- Y. Wang, M. Ouyang, X. Gao, S. Wang, C. Fu, J. Zeng and X. He, Phocea, Pseudoflavonifractor and Lactobacillus Intestinalis: Three Potential Biomarkers of Gut Microbiota That Affect Progression and Complications of Obesity-Induced Type 2 Diabetes Mellitus, Diabetes, Metab. Syndr. Obes.: Targets Ther., 2020, 13, 835–850, DOI:10.2147/DMSO.S240728.
- P. J. Turnbaugh, R. E. Ley, M. A. Mahowald, V. Magrini, E. R. Mardis and J. I. Gordon, An Obesity-Associated Gut Microbiome with Increased Capacity for Energy Harvest, Nature, 2006, 444(7122), 1027–1031, DOI:10.1038/nature05414.
- A. Everard, C. Belzer, L. Geurts, J. P. Ouwerkerk, C. Druart, L. B. Bindels, Y. Guiot, M. Derrien, G. G. Muccioli, N. M. Delzenne, W. M. de Vos and P. D. Cani, Cross-Talk between Akkermansia Muciniphila and Intestinal Epithelium Controls Diet-Induced Obesity, Proc. Natl. Acad. Sci. U. S. A., 2013, 110(22), 9066–9071, DOI:10.1073/pnas.1219451110.
- L. Zhao, F. Zhang, X. Ding, G. Wu, Y. Y. Lam, X. Wang, H. Fu, X. Xue, C. Lu, J. Ma, L. Yu, C. Xu, Z. Ren, Y. Xu, S. Xu, H. Shen, X. Zhu, Y. Shi, Q. Shen, W. Dong, R. Liu, Y. Ling, Y. Zeng, X. Wang, Q. Zhang, J. Wang, L. Wang, Y. Wu, B. Zeng, H. Wei, M. Zhang, Y. Peng and C. Zhang, Gut Bacteria Selectively Promoted by Dietary Fibers Alleviate Type 2 Diabetes, Science, 2018, 359(6380), 1151–1156, DOI:10.1126/science.aao5774.
- S. I. Sayin, A. Wahlström, J. Felin, S. Jäntti, H.-U. Marschall, K. Bamberg, B. Angelin, T. Hyötyläinen, M. Orešič and F. Bäckhed, Gut Microbiota Regulates Bile Acid Metabolism by Reducing the Levels of Tauro-Beta-Muricholic Acid, a Naturally Occurring FXR Antagonist, Cell Metab., 2013, 17(2), 225–235, DOI:10.1016/j.cmet.2013.01.003.
- A. Kushiyama, H. Sakoda, N. Oue, M. Okubo, Y. Nakatsu, H. Ono, T. Fukushima, H. Kamata, F. Nishimura, T. Kikuchi, M. Fujishiro, K. Nishiyama, H. Aburatani, S. Kushiyama, M. Iizuka, N. Taki, J. Encinas, K. Sentani, N. Ogonuki, A. Ogura, S. Kawazu, W. Yasui, Y. Higashi, H. Kurihara, H. Katagiri and T. Asano, Resistin-Like Molecule β Is Abundantly Expressed in Foam Cells and Is Involved in Atherosclerosis Development, Arterioscler. Thromb. Vasc. Biol., 2013, 33(8), 1986–1993, DOI:10.1161/ATVBAHA.113.301546.
- Y. Shi, N. Zhu, Y. Qiu, J. Tan, F. Wang, L. Qin and A. Dai, Resistin-like molecules: a marker, mediator and therapeutic target for multiple diseases, Cell Commun. Signaling, 2023, 21(18) DOI:10.1186/s12964-022-01032-w.
- S. Gray, M. W. Feinberg, S. Hull, C. T. Kuo, M. Watanabe, S. S. Banerjee, A. DePina, R. Haspel and M. K. Jain, The Krüppel-like Factor KLF15 Regulates the Insulin-Sensitive Glucose Transporter GLUT4*, J. Biol. Chem., 2002, 277(37), 34322–34328, DOI:10.1074/jbc.M201304200.
- Y. Su, Three-Dimensional Network of Creatine Metabolism: From Intracellular Energy Shuttle to Systemic Metabolic Regulatory Switch, Mol. Metab., 2025, 100, 102228, DOI:10.1016/j.molmet.2025.102228.
- H. J. Dranse, T. M. Z. Waise, S. C. Hamr, P. V. Bauer, M. A. Abraham, B. A. Rasmussen and T. K. T. Lam, Physiological and therapeutic regulation of glucose homeostasis by upper small intestinal PepT1-mediated protein sensing, Nat. Commun., 2018, 9, 1118, DOI:10.1038/s41467-018-03490-8.
- R. Pappu, V. Ramirez-Carrozzi and A. Sambandam, The Interleukin-17 Cytokine Family: Critical Players in Host Defence and Inflammatory Diseases, Immunology, 2011, 134(1), 8–16, DOI:10.1111/j.1365-2567.2011.03465.x.
- M. Sumarac-Dumanovic, D. Stevanovic, A. Ljubic, J. Jorga, M. Simic, D. Stamenkovic-Pejkovic, V. Starcevic, V. Trajkovic and D. Micic, Increased Activity of Interleukin-23/Interleukin-17 Proinflammatory Axis in Obese Women, Int. J. Obes., 2009, 33(1), 151–156, DOI:10.1038/ijo.2008.216.
- M. C. Petersen, D. F. Vatner and G. I. Shulman, Regulation of Hepatic Glucose Metabolism in Health and Disease, Nat. Rev. Endocrinol., 2017, 13(10), 572–587, DOI:10.1038/nrendo.2017.80.
- C. Postic, R. Dentin and J. Girard, Role of the Liver in the Control of Carbohydrate and Lipid Homeostasis, Diabetes Metab., 2004, 30(5), 398–408, DOI:10.1016/s1262-3636(07)70133-7.
- C. A. Nagle, J. An, M. Shiota, T. P. Torres, G. W. Cline, Z.-X. Liu, S. Wang, R. L. Catlin, G. I. Shulman, C. B. Newgard and R. A. Coleman, Hepatic Overexpression of Glycerol-Sn-3-Phosphate Acyltransferase 1 in Rats Causes Insulin Resistance, J. Biol. Chem., 2007, 282(20), 14807–14815, DOI:10.1074/jbc.M611550200.
- G. Wang, G. Li, A. Song, Y. Zhao, J. Yu, Y. Wang, W. Dai, M. Salas, H. Qin, L. Medrano, J. Dow, A. Li, B. Armstrong, P. T. Fueger, H. Yu, Y. Zhu, M. Shao, X. Wu, L. Jiang, J. Campisi, X. Yang and Q. A. Wang, Distinct Adipose Progenitor Cells Emerging with Age Drive Active Adipogenesis, Science, 2025, 388(6745), eadj0430, DOI:10.1126/science.adj0430.
- J. D. Horton, J. L. Goldstein and M. S. Brown, SREBPs: Activators of the Complete Program of Cholesterol and Fatty Acid Synthesis in the Liver, J. Clin. Invest., 2002, 109(9), 1125–1131, DOI:10.1172/JCI15593.
- Q. Gao, G. Zhang, Y. Zheng, Y. Yang, C. Chen, J. Xia, L. Liang, C. Lei, Y. Hu, X. Cai, W. Zhang, H. Tang, Y. Chen, A. Huang, K. Wang and N. Tang, SLC27A5 deficiency activates NRF2/TXNRD1 pathway by increased lipid peroxidation in HCC, Cell Death Differ., 2020, 27, 1086–1104, DOI:10.1038/s41418-019-0399-1.
- Y. Guo, Z. Zhang, Z. Wen, X. Kang, D. Wang, L. Zhang, M. Cheng, G. Yuan and H. Ren, Mitochondrial SIRT2-Mediated CPT2 Deacetylation Prevents Diabetic Cardiomyopathy by Impeding Cardiac Fatty Acid Oxidation, Int. J. Biol. Sci., 2025, 21(2), 725–744, DOI:10.7150/ijbs.102834.
- J. Chen, Y. Zhou, Z. Liu, Y. Lu, Y. Jiang, K. Cao, N. Zhou, D. Wang, C. Zhang, N. Zhou, K. Shi, L. Zhang, L. Zhou, Z. Wang, H. Zhang, K. Tang, J. Ma, J. Lv and B. Huang, Hepatic Glycogenesis Antagonizes Lipogenesis by Blocking S1P via UDPG, Science, 2024, 383(6684), eadi3332, DOI:10.1126/science.adi3332.
- X. Fu, Z. Liu, R. Li, J. Yin, H. Sun, C. Zhu, Q. Kong, H. Mou and S. Nie, Amelioration of Hydrolyzed Guar Gum on High-Fat Diet-Induced Obesity: Integrated Hepatic Transcriptome and Metabolome, Carbohydr. Polym., 2022, 297, 120051, DOI:10.1016/j.carbpol.2022.120051.
- S. Hui, J. M. Ghergurovich, R. J. Morscher, C. Jang, X. Teng, W. Lu, L. A. Esparza, T. Reya, N. L. Zhan, J. Y. Guo, E. White and J. D. Rabinowitz, Glucose Feeds the TCA Cycle via Circulating Lactate, Nature, 2017, 551(7678), 115–118, DOI:10.1038/nature24057.
- N. K. H. Yiew and B. N. Finck, The Mitochondrial Pyruvate Carrier at the Crossroads of Intermediary Metabolism, Am. J. Physiol. Endocrinol. Metab., 2022, 323(1), E33–E52, DOI:10.1152/ajpendo.00074.2022.
- N. E. Sunny, S. Satapati, X. Fu, T. He, R. Mehdibeigi, C. Spring-Robinson, J. Duarte, M. J. Potthoff, J. D. Browning and S. C. Burgess, Progressive Adaptation of Hepatic Ketogenesis in Mice Fed a High-Fat Diet, Am. J. Physiol. Endocrinol. Metab., 2010, 298(6), E1226–E1235, DOI:10.1152/ajpendo.00033.2010.
- E. Vivier, D. H. Raulet, A. Moretta, M. A. Caligiuri, L. Zitvogel, L. L. Lanier, W. M. Yokoyama and S. Ugolini, Innate or Adaptive Immunity? The Example of Natural Killer Cells, Science, 2011, 331(6013), 44–49, DOI:10.1126/science.1198687.
- F. H. Karlsson, V. Tremaroli, I. Nookaew, G. Bergström, C. J. Behre, B. Fagerberg, J. Nielsen and F. Bäckhed, Gut Metagenome in European Women with Normal, Impaired and Diabetic Glucose Control, Nature, 2013, 498(7452), 99–103, DOI:10.1038/nature12198.
- H. K. Pedersen, V. Gudmundsdottir, H. B. Nielsen, T. Hyotylainen, T. Nielsen, B. A. H. Jensen, K. Forslund, F. Hildebrand, E. Prifti, G. Falony, E. Le Chatelier, F. Levenez, J. Doré, I. Mattila, D. R. Plichta, P. Pöhö, L. I. Hellgren, M. Arumugam, S. Sunagawa, S. Vieira-Silva, T. Jørgensen, J. B. Holm, K. Trošt, MetaHIT Consortium, K. Kristiansen, S. Brix, J. Raes, J. Wang, T. Hansen, P. Bork, S. Brunak, M. Oresic, S. D. Ehrlich and O. Pedersen, Human Gut Microbes Impact Host Serum Metabolome and Insulin Sensitivity, Nature, 2016, 535(7612), 376–381, DOI:10.1038/nature18646.
- M. Trauner, T. Claudel, P. Fickert, T. Moustafa and M. Wagner, Bile Acids as Regulators of Hepatic Lipid and Glucose Metabolism, Dig. Dis., 2010, 28(1), 220–224, DOI:10.1159/000282091.
- J. Y. L. Chiang, Bile Acid Metabolism and Signaling, Compr. Physiol., 2013, 3(3), 1191–1212, DOI:10.1002/cphy.c120023.
- C. Jiang, C. Xie, F. Li, L. Zhang, R. G. Nichols, K. W. Krausz, J. Cai, Y. Qi, Z.-Z. Fang, S. Takahashi, N. Tanaka, D. Desai, S. G. Amin, I. Albert, A. D. Patterson and F. J. Gonzalez, Intestinal Farnesoid X Receptor Signaling Promotes Nonalcoholic Fatty Liver Disease, J. Clin. Invest., 2015, 125(1), 386–402, DOI:10.1172/JCI76738.
- T. Le Roy, M. Llopis, P. Lepage, A. Bruneau, S. Rabot, C. Bevilacqua, P. Martin, C. Philippe, F. Walker, A. Bado, G. Perlemuter, A.-M. Cassard-Doulcier and P. Gérard, Intestinal Microbiota Determines Development of Non-Alcoholic Fatty Liver Disease in Mice, Gut, 2013, 62(12), 1787–1794, DOI:10.1136/gutjnl-2012-303816.
- A. Vrieze, E. Van Nood, F. Holleman, J. Salojärvi, R. S. Kootte, J. F. W. M. Bartelsman, G. M. Dallinga-Thie, M. T. Ackermans, M. J. Serlie, R. Oozeer, M. Derrien, A. Druesne, J. E. T. Van Hylckama Vlieg, V. W. Bloks, A. K. Groen, H. G. H. J. Heilig, E. G. Zoetendal, E. S. Stroes, W. M. de Vos, J. B. L. Hoekstra and M. Nieuwdorp, Transfer of Intestinal Microbiota from Lean Donors Increases Insulin Sensitivity in Individuals with Metabolic Syndrome, Gastroenterology, 2012, 143(4), 913–916, DOI:10.1053/j.gastro.2012.06.031.
- S. Fiorucci and E. Distrutti, Chenodeoxycholic Acid: An Update on Its Therapeutic Applications, Handb. Exp. Pharmacol., 2019, 256, 265–282, DOI:10.1007/164_2019_226.
- X. Li, X. Yao, X. Zhang, X. Dong, S. Chi, B. Tan, S. Zhang and S. Xie, Effects of Dietary Chenodeoxycholic Acid Supplementation in a Low Fishmeal Diet on Growth Performance, Lipid Metabolism, Autophagy and Intestinal Health of Pacific White Shrimp, Litopenaeus Vannamei, Fish Shellfish Immunol., 2022, 127, 1088–1099, DOI:10.1016/j.fsi.2022.07.045.
- U. Özcan, E. Yilmaz, L. Özcan, M. Furuhashi, E. Vaillancourt, R. O. Smith, C. Z. Görgün and G. S. Hotamisligil, Chemical Chaperones Reduce ER Stress and Restore Glucose Homeostasis in a Mouse Model of Type 2 Diabetes, Science, 2006, 313(5790), 1137–1140, DOI:10.1126/science.1128294.
- Y. Y. Lee, S. H. Hong, Y. J. Lee, S. S. Chung, H. S. Jung, S. G. Park and K. S. Park, Tauroursodeoxycholate (TUDCA), Chemical Chaperone, Enhances Function of Islets by Reducing ER Stress, Biochem. Biophys. Res. Commun., 2010, 397(4), 735–739, DOI:10.1016/j.bbrc.2010.06.022.
- A. H. Lee, E. F. Scapa, D. E. Cohen and L. H. Glimcher, Regulation of Hepatic Lipogenesis by the Transcription Factor XBP1, Science, 2008, 320, 1492–1496, DOI:10.1126/science.1158042.
- J. Kuang, J. Wang, Y. Li, M. Li, M. Zhao, K. Ge, D. Zheng, K. C. P. Cheung, B. Liao, S. Wang, T. Chen, Y. Zhang, C. Wang, G. Ji, P. Chen, H. Zhou, C. Xie, A. Zhao, W. Jia, X. Zheng and W. Jia, Hyodeoxycholic Acid Alleviates Non-Alcoholic Fatty Liver Disease through Modulating the Gut-Liver Axis, Cell Metab., 2023, 35(10), 1752–1766, DOI:10.1016/j.cmet.2023.07.011.
- S.-Y. Zhang, R. J. W. Li, Y.-M. Lim, B. Batchuluun, H. Liu, T. M. Z. Waise and T. K. T. Lam, FXR in the Dorsal Vagal Complex Is Sufficient and Necessary for Upper Small Intestinal Microbiome-Mediated Changes of TCDCA to Alter Insulin Action in Rats, Gut, 2021, 70(9), 1675–1683, DOI:10.1136/gutjnl-2020-321757.
- T. M. Z. Waise, Y.-M. Lim, Z. Danaei, S.-Y. Zhang and T. K. T. Lam, Small Intestinal Taurochenodeoxycholic Acid-FXR Axis Alters Local Nutrient-Sensing Glucoregulatory Pathways in Rats, Mol. Metab., 2021, 44, 101132, DOI:10.1016/j.molmet.2020.101132.
- S. I. Sayin, A. Wahlström, J. Felin, S. Jäntti, H.-U. Marschall, K. Bamberg, B. Angelin, T. Hyötyläinen, M. Orešič and F. Bäckhed, Gut Microbiota Regulates Bile Acid Metabolism by Reducing the Levels of Tauro-beta-muricholic Acid, a Naturally Occurring FXR Antagonist, Cell Metab., 2013, 17, 225–235, DOI:10.1016/j.cmet.2013.01.003.
- A. Genoni, C. T. Christophersen, J. Lo, M. Coghlan, M. C. Boyce, A. R. Bird, P. Lyons-Wall and A. Devine, Long-Term Paleolithic Diet Is Associated with Lower Resistant Starch Intake, Different Gut Microbiota Composition and Increased Serum TMAO Concentrations, Eur. J. Nutr., 2020, 59(5), 1845–1858, DOI:10.1007/s00394-019-02036-y.
- S. Fiorucci and E. Distrutti, Chenodeoxycholic Acid: An Update on Its Therapeutic Applications, Handb. Exp. Pharmacol., 2019, 256, 265–282, DOI:10.1007/164_2019_226.
- S. Fiorucci and E. Distrutti, Chenodeoxycholic Acid: An Update on Its Therapeutic Applications, in Bile Acids and Their Receptors, ed. S. Fiorucci and E. Distrutti, Springer International Publishing, Cham, 2019, pp. 265–282, DOI:10.1007/164_2019_226.
- G. R. Gibson, K. P. Scott, R. A. Rastall, K. M. Tuohy, A. Hotchkiss, A. Dubert-Ferrandon, M. Gareau, E. F. Murphy, D. Saulnier, G. Loh, S. Macfarlane, N. Delzenne, Y. Ringel, G. Kozianowski, R. Dickmann, I. Lenoir-Wijnkook, C. Walker and R. Buddington, Dietary Prebiotics: Current Status and New Definition, Food Sci. Technol. Bull.: Funct. Foods, 2010, 7, 1–19 Search PubMed.
- J. Miyamoto, Y. Ando, A. Nishida, M. Yamano, S. Suzuki, H. Takada and I. Kimura, Fructooligosaccharides Intake during Pregnancy Improves Metabolic Phenotype of Offspring in High Fat Diet-Induced Obese Mice, Mol. Nutr. Food Res., 2024, 68(9), e2300758, DOI:10.1002/mnfr.202300758.
- T. Oku and S. Nakamura, Fructooligosaccharide: Metabolism through Gut Microbiota and Prebiotic Effect, Food Nutr. J., 2017, 2, 128, DOI:10.29011/2575-7091.100028.
- Y. Lan, Q. Sun, Z. Ma, J. Peng, M. Zhang, C. Wang, X. Zhang, X. Yan, L. Chang, X. Hou, R. Qiao, A. Mulati, Y. Zhou, Q. Zhang, Z. Liu and X. Liu, Seabuckthorn Polysaccharide Ameliorates High-Fat Diet-Induced Obesity by Gut Microbiota-SCFAs-Liver Axis, Food Funct., 2022, 13(5), 2925–2937, 10.1039/d1fo03147c.
- D. Ulluwishewa, C. A. Montoya, L. Mace, E. A. Rettedal, K. Fraser, W. C. McNabb, P. J. Moughan and N. C. Roy, Biotransformation of Rutin in In Vitro Porcine Ileal and Colonic Fermentation Models, J. Agric. Food Chem., 2023, 71(33), 12487–12496, DOI:10.1021/acs.jafc.3c00980.
- L. Du, M. Zhao, J. Xu, D. Qian, S. Jiang, E. Shang, J. Guo and J. Duan, Analysis of the Metabolites of Isorhamnetin 3-O-Glucoside Produced by Human Intestinal Flora in Vitro by Applying Ultraperformance Liquid Chromatography/Quadrupole Time-of-Flight Mass Spectrometry, J. Agric. Food Chem., 2014, 62, 2489–2495, DOI:10.1021/jf405261a.
- G. Li, X. Zeng, Y. Xie, Z. Cai, J. C. Moore, X. Yuan, Z. Cheng and G. Ji, Pharmacokinetic Properties of Isorhamnetin, Kaempferol and Quercetin after Oral Gavage of Total Flavones of Hippophae Rhamnoides L. in Rats Using a UPLC-MS Method, Fitoterapia, 2012, 83(1), 182–191, DOI:10.1016/j.fitote.2011.10.012.
- L. Ren, Y. Xie, R. Wang, Y. Zuo, S. Zhang, F. Tie, Q. Dong, H. Wang and N. Hu, Sea Buckthorn Triterpenic Acids Alleviates Metabolic Dysfunction-Associated Steatotic Liver Disease via Modulation of Liver Biochemistry, Lipid Metabolism and AMPK/Nrf2/NF-κB Signaling Pathway, Food Biosci., 2025, 68, 106611, DOI:10.1016/j.fbio.2025.106611.
- W. Deng, W. Fan, T. Tang, H. Wan, S. Zhao, Y. Tan, K. A. Oware, J. Tan, J. Li and S. Qu, Farnesoid X Receptor Deficiency Induces Hepatic Lipid and Glucose Metabolism Disorder via Regulation of Pyruvate Dehydrogenase Kinase 4, Oxid. Med. Cell. Longevity, 2022, 2022, 3589525, DOI:10.1155/2022/3589525.
- N. Horii, K. Sato, N. Mesaki and M. Iemitsu, Increased Muscular 5α-Dihydrotestosterone in Response to Resistance Training Relates to Skeletal Muscle Mass and Glucose Metabolism in Type 2 Diabetic Rats, PLoS One, 2016, 11(11), e0165689, DOI:10.1371/journal.pone.0165689.
- 1667-P: Effects of Low Androgenic Hormone Levels on Glycemic Metabolism and Ectopic Lipid Accumulation | Diabetes | American Diabetes Association, https://diabetesjournals.org/diabetes/article/74/Supplement_1/1667-P/158391/1667-P-Effects-of-Low-Androgenic-Hormone-Levels-on (accessed 2026-02-10).
- S. Liu, R. Cao, L. Liu, Y. Lv, X. Qi, Z. Yuan, X. Fan, C. Yu and Q. Guan, Correlation Between Gut Microbiota and Testosterone in Male Patients With Type 2 Diabetes Mellitus, Front. Endocrinol., 2022, 13, 836485, DOI:10.3389/fendo.2022.836485.
- M. Matsushita, K. Fujita, D. Motooka, K. Hatano, J. Hata, M. Nishimoto, E. Banno, K. Takezawa, S. Fukuhara, H. Kiuchi, Y. Pan, T. Takao, A. Tsujimura, S. Yachida, S. Nakamura, W. Obara, H. Uemura and N. Nonomura, Firmicutes in Gut Microbiota Correlate with Blood Testosterone Levels in Elderly Men, World J. Mens Health, 2022, 40(3), 517–525, DOI:10.5534/wjmh.210190.
- Z. Ma and W. Li, How and Why Men and Women Differ in Their Microbiomes: Medical Ecology and Network Analyses of the Microgenderome, Adv. Sci., 2019, 6, 1970140, DOI:10.1002/advs.201902054.
Footnote |
| † These authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |