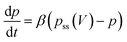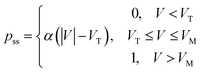Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Inducing nonlinear conductance and emergent memristance in open pores using blockers
Raman Dhiman a,
Priyanshu R. Guptab,
Nicholas X. Armendarezab,
Lyssa Lashusc,
Kyler Grogana,
Malika Raob,
Aida Ficab,
Ronald J. Voglera,
Dominic Bujanosa,
Harekrushna Beheraab,
Stephen A. Sarles
a,
Priyanshu R. Guptab,
Nicholas X. Armendarezab,
Lyssa Lashusc,
Kyler Grogana,
Malika Raob,
Aida Ficab,
Ronald J. Voglera,
Dominic Bujanosa,
Harekrushna Beheraab,
Stephen A. Sarles d and
Manish Kumar*ab
d and
Manish Kumar*ab
aJohn J. McKetta Jr. Department of Chemical Engineering, The University of Texas at Austin, 200 E. Dean Keeton Street, Austin, TX 78712, USA. E-mail: manish.kumar@utexas.edu; Tel: +1 619-917-7392
bFariborz Maseeh Department of Civil, Architectural and Environmental Engineering, The University of Texas at Austin, 301 E. Dean Keeton Street, Austin, TX 78712, USA
cAiiso Yufeng Li Family Department of Chemical and Nano Engineering, University of California, San Diego, CA, USA
dDepartment of Mechanical, Aerospace and Biomedical Engineering, University of Tennessee, Knoxville, Tennessee 37916, USA
First published on 27th April 2026
Abstract
Memristance in biomembrane systems is an emerging area of research relevant to biocomputing. Work has so far focused on voltage-driven insertion of pore-forming peptides in droplet interface bilayer systems, design of asymmetric pores, and lipid membrane poration. However, only a few instances of always-open, pore-forming channels have been used to design and study the emergence of memristive behaviors in biomimetic membranes. In this work, we investigate the Outer Membrane Protein F (OmpF) as a model nanopore. Due to its nanoscale pore size (∼1 nm diameter), OmpF can be transiently or partially blocked by a variety of molecules, including antibiotics, polyamines, and polypeptides. We reconstituted OmpF into droplet interface bilayers and examined electrical signatures of different blocking agents using cyclic voltammetry. We show that binding of some of these agents to pore-lining residues induces strongly nonlinear current–voltage (I–V) responses, which, in some instances, give rise to memristive behavior. We find that the interaction of ampicillin and spermine with OmpF at the tested conditions was insufficient to produce nonlinearity or memristance. However, interaction between arginine polypeptides and OmpF leads to length and charge-dependent nonlinear conductance with emergent memristance in the case of arginine nonapeptide. Molecular dynamics simulations showed that arginine pentapeptides (Arg-5) block the pore longitudinally as a stretched chain whereas bulkier and highly charged arginine nonapeptides (Arg-9) block the pore transversally by compaction of the peptide chain. Although both blocking conformations were sufficient to exhibit nonlinear dynamics, only the transversal blocking conformation led to emergent memristance, likely due to longer-lived interactions between binding sites and the Arg-9.
Introduction
In recent years, with the advancement of artificial intelligence, demand for energy required for computation has increased exponentially, and global power demand for computation is expected to surpass global power production by 2030.1 Traditional computing devices follow the Von Neumann architecture, where the processing unit and memory unit are two separate entities.2 Data needs to be continuously transferred from one unit to another.2 This data transfer, which can cost up to 1000 times that of actual computation costs, is referred to as the Von Neumann bottleneck.3 The inefficiency caused by this bottleneck motivates research into devices where memory and processing units can be co-located.Memristive devices or memristors4,5 are two-terminal devices in which memory and processing units are co-located.6 The conductance of these devices depends on their history of voltage and current.5
Leon Chua and co-workers showed that potassium and sodium ion channels in the Hodgkin–Huxley axon could be described as memristors.7 This work, along with others, has inspired work on biological memristors as a building block for brain-inspired computing hardware, including ion channel-doped biomembrane-based memristors.8–17
Alamethicin-doped membranes have been extensively studied in droplet interface bilayer (DIB) systems.8,12,13,15 Alamethicin peptides form transmembrane ion channels following voltage-driven insertion into lipid membranes. In alamethicin-doped membranes, memristance emerges from the time difference between insertion and exit of the pore-forming peptides from lipid membranes as well as the time delay in area and thickness changes of the membrane due to voltage, which influences the total number of open channels that conduct ions.8 There are limited examples in the literature of non-voltage-gated protein channels that act as memristors. In one study, Koner et al. showed that a gramicidin-doped biomembrane serves as a memristor where memristance emerges from the dynamics of the reconfiguration of gramicidin.9 Paulo et al. demonstrated how hydrophobically gated nanopores can act as memristors.16 Very recently, Mayer et al. reported on open beta-barrel proteins (aerolysin, alpha-HL, and MspA) exhibiting memristance because of gated ion transport governed by lumen charge.17
OmpF is one of the major diffusion pores in the outer membrane of E. coli.18 OmpF is a homotrimer where each monomer is made up of 16-stranded antiparallel beta-barrels.19 The L3 loop joins β-strands 5 and 6 of the barrel and forms the constriction zone in each pore.19 Literature reports suggest that OmpF itself exhibits gating behavior above certain voltage thresholds, but this behavior is highly dependent on experimental conditions20 with varying voltage threshold values reported in the literature.20,21
OmpF is also extensively studied for its antibiotic binding and transport properties as it is one of the pathways for antibiotic entry into the cell.18,22,23 There are conflicting reports on the antibiotic ampicillin blocking the OmpF pore.18,24,25 Earlier studies indicated ampicillin binding and blocking of OmpF pores,18,25 but it was later shown that ampicillin shows negligible blocking of the OmpF pore. Rather, it was the degraded ampicillin product (penicilloic acid) that is actually responsible for OmpF blocking.24
There are also studies to understand the interaction between OmpF pores and polyamines, where spermine (a type of polyamine) significantly reduces the activity of OmpF pores.26,27 Spermine reduces the activity of OmpF not by plugging the pore, but rather by stabilizing the closed conformation of the pore.26
Lamichhane et al. studied the binding of arginine polypeptide chains with OmpF28 and reported no translocation. However, another study using secondary blocker molecules attached to the periplasmic side of OmpF established that tri-arginine indeed translocates through OmpF whereas hepta-arginine is bulky enough to preclude translocation through OmpF pores.29 We hypothesized that memristance could be induced in an open pore, such as OmpF, using voltage-responsive blocker molecules, inspired by voltage-gated ion channels, in which the probability of channel opening and closing produces memristance.7 This functionality is produced by blocker molecules that act as a pseudo-gate and the probability of blocking and unblocking of the molecule (dictated by its residence time on specific residues) will produce emergent memristance. In this work, we exploited the blocking of OmpF using various blocker molecules and attempted to induce memristance in an OmpF-doped DIB biomembrane system. We also conducted molecular dynamics simulations to understand the binding of blocker molecules with OmpF. To the best of our knowledge, this work is the first demonstration of the use of ion channel extrinsic blocking to induce memristance.
Methods
Materials
1,2-Diphytanoyl-sn-glycero-3-phosphocholine (DPhPC) lipids (25 mg ml−1 in chloroform) were obtained from Avanti Research. Ampicillin sodium salt and spermine were purchased from Sigma-Aldrich, arginine peptides were custom synthesized by GenScript Biotech.Preparation of vesicle solution
25 mg ml−1 of DPhPC stock in chloroform was added to a round-bottom flask along with 2–3 ml of additional chloroform. A rotary evaporator was used to evaporate chloroform under reduced pressure and to prepare thin lipid films. The flask was then placed in a vacuum oven at room temperature for ≥2 hours to ensure complete removal of chloroform under reduced pressure. The residual lipid film was rehydrated using the appropriate volume of buffer (100 mM potassium chloride (KCl), 10 mM 3-(N-morpholino)propanesulfonic acid (MOPS), pH 7.0) and extruded (11 passes) using Avanti’s mini extruder to form an aqueous vesicle solution at a target lipid concentration of 2 mg ml−1. A 0.2 µm PC membrane was used for extrusion.Purification of OmpF
OmpF was expressed and purified according to procedures reported before.30Blockers tested
The blockers tested included ampicillin (net charge at pH 7(n): −0.75),24 spermine (n = +4), tri-arginine peptide (Arg-3) (n = +3), penta-arginine peptide (Arg-5) (n = +5), hepta-arginine peptide (Arg-7) (n = +7), and nona-arginine peptide (Arg-9) (n = +9).Preparation of electrodes
Rounded Ag/AgCl electrodes are needed to securely interact with the droplets within the droplet interface bilayer (DIB) system. The tips of the 0.127 mm diameter silver wire (Ag) electrodes were rounded by melting. Then, silver/silver chloride (Ag/AgCl) electrodes were prepared by dipping the ends of Ag electrodes into bleach for at least 20 minutes. The tips of these electrodes were then dipped in melted agarose to coat their ends and make them hydrophilic. The electrodes were connected to a patch clamp amplifier (HEKA, EPC 10). The electrode tips were then submerged in a hexadecane oil bath. The positions of the electrodes were manipulated by using micromanipulators.Formation of droplet interface bilayers
400 nL of aqueous vesicle solution was pipetted onto each of the electrode tips in oil. After allowing 10–15 minutes for monolayer formation around the droplets, we used micromanipulators to bring the droplets into contact to form bilayers. Once the bilayer formed, we waited 3 minutes for it to stabilize before investigating its voltage-responsive properties.Insertion of OmpF in droplet interface bilayer
Vesicles were premixed with OmpF (at a lipid to protein mass ratio (LPR) of 2000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) and blocker molecules (different concentrations) before dispensing the droplet vesicles onto the electrodes. For inserting OmpF into DIB membranes, a droplet of prepared mixture of vesicles with OmpF and blocker molecules was added to the source electrode and a droplet of vesicles without any OmpF or blocker molecules was added to the grounded electrode (with the exception of the ampicillin case, where a droplet of vesicles with OmpF was added to the source electrode and a droplet of vesicles with blocker molecules was added to the ground electrode). Droplet interface bilayers were formed as described above. Then three DC voltage pulses of −100 mV each were applied for 120 s (with a wait time of 5 min in between).
1) and blocker molecules (different concentrations) before dispensing the droplet vesicles onto the electrodes. For inserting OmpF into DIB membranes, a droplet of prepared mixture of vesicles with OmpF and blocker molecules was added to the source electrode and a droplet of vesicles without any OmpF or blocker molecules was added to the grounded electrode (with the exception of the ampicillin case, where a droplet of vesicles with OmpF was added to the source electrode and a droplet of vesicles with blocker molecules was added to the ground electrode). Droplet interface bilayers were formed as described above. Then three DC voltage pulses of −100 mV each were applied for 120 s (with a wait time of 5 min in between).
For experiments of inducing memristance in the OmpF pore, pre-insertion of OmpF in the bilayer before memristance testing is desirable. To achieve this, the relative net charges of the protein and blocker molecules were utilized. OmpF is negatively charged and was added to the droplet attached to the source electrode, whereas blocker molecules were added to the source side or ground droplet, depending on whether they were positively charged or negatively charged, respectively (Table 1 and Fig. 1). We surmised that applying a negative potential at the source electrode would attract negatively charged molecules to the ground electrode, whereas positively charged molecules would move in the opposite direction. This procedure ensured that the initial few sweeps could be used to insert OmpF in bilayers, whereas the blocker molecule would move away from the bilayer, hence essentially separating the insertion of OmpF dynamics from the blocking dynamics of the blocker molecules. A similar approach to driving molecules towards bilayers has been used before in the literature29
| Protein/blocker molecules | Net charge at pH 7 | Side in which protein/blocker molecules are added | Direction of force experienced during initial sweeps for OmpF insertion |
|---|---|---|---|
| OmpF | Negative | Source | Towards bilayer |
| Ampicillin | Negative | Ground | Away from bilayer |
| Spermine | Positive | Source | Away from bilayer |
| Arg-3 | Positive | Source | Away from bilayer |
| Arg-5 | Positive | Source | Away from bilayer |
| Arg-7 | Positive | Source | Away from bilayer |
| Arg-9 | Positive | Source | Away from bilayer |
Electrophysiology
A HEKA EPC 10 amplifier was used to conduct electrophysiological studies, including current–voltage (I–V) sweeps. For the memristance tests in particular, zero-mean continuous triangular voltage sweeps were applied on the droplet interface bilayer system with an amplitude of ±150 mV and varying sweep rates. A 1 kHz Bessel filter was used during the measurement. A 100 Hz display filter was applied for post-processing of the data.Molecular dynamics simulations
The initial coordinates for the OmpF porin (PDB ID: 2OMF) embedded in a 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC) bilayer were generated using the CHARMM-GUI Membrane Builder.31 Three distinct systems were constructed: (i) OmpF in POPC (control), (ii) OmpF/POPC with an Arg5 peptide (Arg-5), and (iii) OmpF/POPC with an Arg9 peptide (Arg-9). Peptides were placed in the extracellular vestibule, and overlapping water molecules were removed using a 2 Å distance cutoff. The systems were solvated with TIP3P water molecules and neutralized with NaCl to mimic experimental ionic strength conditions.Simulations were performed using NAMD 3.0 with the CHARMM36m force field. Additional CHARMM-based equilibration was performed, plus 10 ns of equilibration before introducing the peptides. Long-range electrostatic interactions were computed using the Particle Mesh Ewald (PME) method, and van der Waals interactions were treated with a force-based switching function between 10 and 12 Å.
A timestep of 2 fs was used, and a constant temperature of 303.15 K was maintained using Langevin dynamics. Pressure was regulated at 1.01325 bar using the Nosé–Hoover Langevin piston method. A constant external electric field of −0.5 kcal (mol Å e)−1 was applied along the z-axis, corresponding to a transmembrane potential of approximately 2.5 V.32 While this potential exceeds experimental voltages, it is a standard approach in porin simulations to accelerate sampling and overcome the free energy barriers of ion/peptide transport within computationally accessible timescales.32–35
For each of the three systems (control, Arg5, and Arg9), three independent simulation replicas were executed for 15 ns each, starting from different initial velocity distributions to ensure statistical robustness.
Results and discussion
A bilayer without any protein acts as a capacitor and allows very minimal ion transport, as illustrated in Fig. 2A, and the current–voltage (I–V) curve shows an almost flat horizontal line showing minimal transport and large capacitance. We conducted preliminary screening experiments with six distinct blocker molecules (representing three classes: antibiotics, polyamines, and polypeptides) to study OmpF blocking and their ability to induce nonlinear behaviors and memristance in OmpF-based DPhPC lipid membranes using the DIB method (Fig. 1). These blockers included ampicillin, spermine, tri-arginine peptides (Arg-3), penta-arginine peptides (Arg-5), hepta-arginine peptides (Arg-7), and nona-arginine peptides (Arg-9). As seen in Fig. 2, I–V sweeps for preliminary evaluation of memristive behavior showed no signs of emergent memristance or distinctive nonlinear behaviors for the OmpF–membrane system in the presence of ampicillin (0.13 µM) or spermine (0.3 mM) (blocker concentrations cover the range of blocker to pore molar ratio of ∼5 to 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000). Nonlinear I–V behaviors and emergent memristance were observed with arginine peptides of increasing length (Fig. 3 & 4).
000). Nonlinear I–V behaviors and emergent memristance were observed with arginine peptides of increasing length (Fig. 3 & 4).
 | ||
| Fig. 2 I–V sweeps for zero mean triangular voltage input show no emergent nonlinear behavior or emergent memristance for ampicillin or spermine molecules driven through OmpF pores. (A) The I–V sweep for the DIB with no OmpF and no blocker molecules showing minimal transport. (B) The I–V sweep for the DIB with OmpF pore, but no blocker molecules, produces a straight line. (C) The I–V sweep for the DIB containing the OmpF pore, along with ampicillin molecules. Here, we also observe almost linear I–V curves representing no/negligible memristance as evidenced by a lack of a pinched hysteresis loop. (D) The I–V sweep for the system containing the OmpF pore with spermine molecules. Here, we do observe a slightly different curve, but no memristive behavior. The sweep rate applied to generate A was 10 mV s−1, and for the other three plots, 20 mV s−1. The area between the curves is because of the large capacitance of the bilayer, as explained elsewhere36 (typical values of capacitance for these experiments were 400–800 pF, except for the spermine case, where that value was 50–60 pF). | ||
Emergence of nonlinear I–V curves and memristance by OmpF blocking by Arg-9
OmpFs inserted into the bilayer allow conductive transport of ions, resulting in a linear I–V profile (in this case with minimal contribution of capacitance) (Fig. 3A–C). Arginine is a positively charged amino acid and can interact with negatively charged residues on the L3 loop within the OmpF pore (Fig. S1). The interaction of Arg-9 with OmpF (Fig. 3D) induced nonlinearity and emergent memristance in the system. When the voltage was increased from 0 to 150 mV, initially the current also increased linearly, indicating nearly constant conductance (Fig. 3E). But above a certain threshold voltage, the conductance decreased with increasing applied voltage. This trend indicates a tendency for increased blocking with increasing voltage, presumably because of the blocker molecules being driven to and then interacting strongly with particular charged residues within the pore. In the reverse cycle from 150 mV to 0 mV, the conductance increased (indicated by the increase in slope of the I–V curve) till it reached the value reached in the 0 mV to 150 mV direction. This reversible change in conductance indicates voltage-dependent blockage of the OmpF pore. It also indicates that a persistent potential is required to keep the pore blocked, and reducing the potential initiates the unblocking process. The I–V curve of the protein pore blocked with Arg-9 molecules at a concentration of 0.003 mM showed a hysteresis loop (Fig. 3F) and a symmetric, pinched I–V hysteresis loop, meeting one of the main criteria for classifying systems as possessing emergent memristance. To confirm whether the hysteresis is memristive rather than a distortion caused by capacitive currents, we varied the sweep rate of the triangle voltage waveform to test for hysteresis at various frequencies. Fig. S5 shows the results that capacitive I–V characteristics are only observable at very high frequencies (>100 mV s−1). When we adjusted for these by subtracting the capacitive current associated with a DIB device, we observed hysteresis effects at higher frequencies. We hypothesize that Arg-9, because of its higher charge and larger size, binds in a manner that the blocking and unblocking kinetics become sufficiently differentiated, leading to memristance.A similar pattern is observed during the negative voltage cycle (0 to −150 mV and back). It is worth noting that we did not add any blocker to the other side of the membrane (i.e., the opposite droplet). Further investigation is needed to understand the reasoning behind this observed phenomena.
Emergence of nonlinear dynamics by Arg-3, Arg-5, and Arg-7
OmpF exhibited nonlinear I–V behaviors in the presence of Arg-3, Arg-5, and Arg-7 at certain concentrations, although they did not result in significant I–V hysteresis (Fig. 4). Arg-3 at a concentration of 0.05 mM showed some transient blocking, but this blocking was not sufficient to significantly alter the linearity of the I–V curve (Fig. 4A). However, at 1 mM, Arg-3 induced a nonlinear response in the OmpF I–V trace at positive voltages, but no memristance (Fig. 4B). Arg-5 and Arg-7 also induced nonlinear I–V behavior, although they also did not exhibit memristance (Fig. 4C–F). We observed minimal blocking in the case of Arg-3 and Arg-5 during the negative voltage cycle (0 to −150 mV and back), whereas we observed blocking during the negative voltage cycle for Arg-7 (similar to Arg-9 as discussed above). According to literature, Arg-3 can translocate through the pore, whereas Arg-7 does not.29 Our MD simulation results also show that Arg-5 could translocate across the pore, whereas Arg-9 did not under the conditions simulated. Literature results combined with our MD simulations suggest that although Arg-3 and Arg-5 can translocate through the pore, the concentration on the other side of the pore does not reach a level during I–V sweeps to result in nonlinear IV relationships at negative voltages. Similar to the case of Arg-9, the reason for blocking observed during the negative cycle for Arg-7 is unclear and further investigations are necessary to provide a convincing explanation for this observation.Theoretical model
We developed a probabilistic model inspired by the Hodgkin–Huxley model37 to describe ion channel dynamics underlying OmpF blocking and the resulting IV behavior. In this model, current can be simply modeled by single channel conductance, the number of channels, and their open probability:| I = GuANa(1 − p)V |
where β is the rate constant of pore blocking, and pss is the steady state probability of pore blockage at a particular voltage. pss is a static function of the transmembrane voltage, which we define as
where α is the proportionality constant representing the dependency of pore blocking probability on voltage. VT is the threshold voltage after which pore blocking starts. VM is the voltage at which probability of pore blockage becomes 1. The blocking of pores is responsible for the nonlinearity of the curve (hence it is dependent on α), whereas the memristance is dependent on how quickly the probability reaches its steady state and hence is dependent on β.
Plots in Fig. 5 are results from the model with assumed values of α and β. A value of single channel conductance, Gu = 0.28 nS,38 was approximated from literature (shown in SI) and used, and the number of inserted channels per bilayer (A × Na) was assumed as 100, as the number of pores inserted varied mostly in the range of 20–200 for our experiments, as shown in Fig. S2 and S4. In Fig. 5A, the curves resulting from a low value of α indicate that the probability of blocking at steady state is lower compared to the higher value of α, resulting in curves seen in Fig. 5B and C. A higher value of β indicates that dp/dt, the rate of change of probability of blocking, is high, and thus the probability of blocking reaches its steady state relatively rapidly. Therefore, no memristance is observed with the higher value of β compared to the lower value of β, where we do observe memristance (indicated by the pinched hysteresis loop).
Molecular dynamics simulation
Molecular dynamics simulations of a single OmpF pore reveal a pronounced reduction in ionic flux across the pores blocked with arginine peptide variants relative to the control. MD inherently samples short timescales, limiting the number of ion conduction events that can be observed (a conductance of 300 pS corresponds to ∼2 crossings per 10 ns window). To obtain better statistics within the accessible simulation timeline, we applied a transmembrane voltage equivalent to 2.8 V. The use of unphysiologically high voltages in our MD simulations is consistent with prior studies.32,33,39–42 We lightly restrained lipid phosphorus atoms and Cα atoms of the equilibrated protein, a stabilization strategy that preserves lipid bilayer, and the protein’s secondary structure while allowing side-chain mobility.43,44 Under the simulation conditions, the system without any peptides sustains a steady ionic current of 880 ± 21 pA, whereas the OmpF pore in the presence Arg-5 and Arg-9 peptides exhibited substantial current suppression, with mean currents of 240 ± 15 pA and 208 ± 15 pA, respectively (Fig. 6). Notably, during the initial ∼2 ns, the currents observed in both Arg-5 and Arg-9 trajectories closely follow the same trend as the control simulation. However, this transient conductive phase is quickly followed by a rapid transition to a predominantly blocked state, marked by extended periods of reduced flux punctuated by rare ion-translocation events.Visualizations of the trajectories showed that while both peptides, Arg-5 and Arg-9, reside within the beta-barrel, they exhibit distinct conformational behaviors within the protein pore. Arg-9 exhibits a compact, globular structure that stably occupies the central pore axis, generating a large steric and electrostatic barrier to entering the constriction created by loop L3. In contrast for Arg-5, the peptide unfolds and a subset of arginine residues (Arg1, Arg2, and Arg3) bind in the pore lumen, and constrict the pore along the pore length rather than just at the entrance (Fig. 7).
These distinctions are further highlighted by plotting the radius of gyration (Rg) and end-to-end distance (Ree) distributions (Fig. 8). For Arg-9, the trajectory shows a clear transition toward a collapsed, globular state, where the Ree drops significantly over time (from ∼18 Å to <10 Å) while Rg remains tightly clustered around 6.8 Å. This quantitative shift mirrors the visual transition into a stable, globular plug. Conversely, the Arg-5 plots reveals a very gradual increase in the end-to-end distance, indicating the “unfolding” behavior observed in the visualizations. The peptide remains physically small enough to fit within the barrel but stretches its backbone to allow side chains of Arg1–3 to protrude into the lumen. Together, these metrics confirm that Arg-9 blocks the pore through a single, dense steric bulk, whereas Arg-5 achieves occlusion through a more elongated, multi-point constriction along the pore axis.
The functional consequence of these structural differences is most evident under high applied voltage (equivalent to 3.5 V), where a complete translocation event was observed for Arg-5 (Fig. 9). In this trajectory, the longitudinal orientation of the Arg-5 peptide allowed it to thread through the constriction zone and eventually escape into the bulk solvent on the trans side. This exit resulted in a full recovery of the ionic current to levels indistinguishable from the control (pore-only) simulations. Arg-9, on the other hand, remained securely lodged within the barrel throughout the same simulation timescale. These results indicate that while Arg-5 can effectively constrict the pore at lower potentials, its ‘unfolded’ binding mode is more susceptible to voltage-driven translocation than the compact ‘plug’ formed by Arg-9.
Overall, MD simulations reveal a difference in the configuration of Arg-5 vs. Arg-9 when blocking the OmpF pore. Both molecules clearly demonstrated a drop in conductance when they block the pore. However, as shown in the Rg vs. Ree plot (Fig. 8) Arg-5 polypeptide adopts an elongated, unfolded conformation that occupies the pore longitudinally, whereas Arg-9 collapses into a compact globular conformation, occluding the pore transversely. Fig. 7 illustrates this difference in conformation taken by Arg-5 vs. Arg-9 when blocking the pore. These results suggest that both Arg-5 and Arg-9 can block the OmpF pore and we believe that this blocking leads to nonlinearity in the I–V curves. However, the binding mode for smaller peptides like Arg-5 is strikingly different from the binding mode of Arg-9. Transversal blocking of the pore by Arg-9 potentially introduces a lag in unbinding kinetics and results in emergent memristance. On the other hand, longitudinal blocking of the pore by smaller peptides like Arg-5 does not create any lag in the unbinding process, and hence we did not observe emergent memristance. Further work is needed to explain how this difference in binding mode may result in emergent memristance.
Conclusions
Three different classes of blocker molecules were investigated for their potential to induce nonlinearity and pinched hysteresis (a signature of emergent memristance) in OmpF-doped biomembranes. Whereas ampicillin and spermine were unable to produce nonlinearity and memristance in the system, bulkier and highly charged arginine polypeptides (Arg-3, Arg-5, Arg-7, and Arg-9) were able to induce nonlinearity in the OmpF doped biomembrane. Some emergent memristance was also observed in the case of Arg-9. MD simulations indicate that both Arg-5 and Arg-9 can block the OmpF pore and reduce the conductance. However, simulations also indicated a difference in blocking conformation for Arg-5 and Arg-9. Where Arg-5 unfolded and blocked the OmpF pore longitudinally, Arg-9 collapsed into a globular structure and blocked the pore transversally. Both blocking conformations have resulted in nonlinear dynamics, but emergent memristance in Arg-9 is likely the result of transversal and likely long-lived blocking of the pore, whereas longitudinal blocking in the smaller peptide (Arg-5) was unable to lead to such emergent memristance. Further investigation is required to understand how this difference in binding conformation translates to emergent memristance. A theoretical probabilistic model was also built and used to show the transformation of linear dynamics to nonlinear dynamics by varying the α (proportionality constant) parameter and the transformation of nonlinear dynamics to memristance by varying the β (rate constant of pore blocking) parameter. This study shows that open pores can exhibit memristance dynamics in the presence of blocker molecules.Looking forward, the results of this work contribute to the wider application of artificial membrane systems mimicking biological neurons, which rely on ion channel dynamics, both voltage gating and molecule-binding mediated, to produce signalling. The device presented in this work utilizes a blocking mechanism that is among a growing library of literature examples of ion channel-based dynamics that seek to emulate neuronal and synaptic signalling. Furthermore, like other biologically derived computing systems, our device operates in the nanoampere and millivolt range, resulting is significantly lower power consumption than silicon-based devices. The field of biomimetic nanofluidics for neuromorphic applications seeks to leverage the low-power signalling of ion channel-based devices to achieve the computational efficiency of the brain.
Author contributions
R. D. and M. K. conceptualized the project. R. D., L. L., K. G. and M. R. performed the experiments. P. R. G. performed the molecular dynamics simulations. N. X. A. developed the theoretical model. R. J. V., K. G. and D. B. purified the protein. A. F., H. B. and S. S. helped with experimental design and manuscript editing. R. D., P. R. G. and M. K. performed the analysis and wrote the manuscript with the help of other co-authors.Conflicts of interest
There are no conflicts to declare.Data availability
All the data used in this manuscript have been uploaded to Texas Data Repository and can be accessed using the following link: https://doi.org/10.18738/T8/MTA7H4.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d6fd00008h.
Acknowledgements
This work was supported by the WoodNext Foundation. AI assistance was used for brainstorming ideas and used to generate initial codes for plotting the data using MATLAB and Python.Notes and references
- C. D. Danesh, C. M. Shaffer, D. Nathan, R. Shenoy, A. Tudor, M. Tadayon, Y. Lin and Y. Chen, Synaptic Resistors for Concurrent Inference and Learning with High Energy Efficiency, Adv. Mater., 2019, 31(18), 1808032 CrossRef PubMed.
- G. S. Syed, M. Le Gallo and A. Sebastian, Non von Neumann computing concepts, Elsevier, 2024, pp. 11–35 Search PubMed.
- D. Ivanov, A. Chezhegov, M. Kiselev, A. Grunin and D. Larionov, Neuromorphic artificial intelligence systems, Front. Neurosci., 2022, 16 DOI:10.3389/fnins.2022.959626.
- L. O. Chua and K. Sung Mo, Memristive devices and systems, Proc. IEEE, 1976, 64(2), 209–223 Search PubMed.
- L. Chua, Memristor-The missing circuit element, IEEE Trans. Circ. Theor., 1971, 18(5), 507–519 Search PubMed.
- G. Zhou, Z. Wang, B. Sun, F. Zhou, L. Sun, H. Zhao, X. Hu, X. Peng, J. Yan and H. Wang, et al., Volatile and Nonvolatile Memristive Devices for Neuromorphic Computing, Adv. Electron. Mater., 2022, 8(7), 2101127 CrossRef CAS.
- L. Chua, V. Sbitnev and H. Kim, Hodgkin–Huxley Axon Is Made Of Memristors, Int. J. Bifurcat. Chaos, 2012, 22(03), 1230011 CrossRef.
- J. S. Najem, G. J. Taylor, R. J. Weiss, M. S. Hasan, G. Rose, C. D. Schuman, A. Belianinov, C. P. Collier and S. A. Sarles, Memristive Ion Channel-Doped Biomembranes as Synaptic Mimics, ACS Nano, 2018, 12(5), 4702–4711 CrossRef CAS PubMed.
- S. Koner, J. S. Najem, M. S. Hasan and S. A. Sarles, Memristive plasticity in artificial electrical synapses via geometrically reconfigurable, gramicidin-doped biomembranes, Nanoscale, 2019, 11(40), 18640–18652 RSC.
- J. J. Maraj, J. S. Najem, J. D. Ringley, R. J. Weiss, G. S. Rose and S. A. Sarles, Short-Term Facilitation-Then-Depression Enables Adaptive Processing of Sensory Inputs by Ion Channels in Biomolecular Synapses, ACS Appl. Electron. Mater., 2021, 3(10), 4448–4458 CrossRef CAS.
- J. J. Maraj, K. P. T. Haughn, D. J. Inman and S. A. Sarles, Sensory Adaptation in Biomolecular Memristors Improves Reservoir Computing Performance, Adv. Intell. Syst., 2023, 5(8), 2300049 CrossRef.
- N. X. Armendarez, A. S. Mohamed, A. Dhungel, M. R. Hossain, M. S. Hasan and J. S. Najem, Brain-Inspired Reservoir Computing Using Memristors with Tunable Dynamics and Short-Term Plasticity, ACS Appl. Mater. Interfaces, 2024, 16(5), 6176–6188 CrossRef CAS PubMed.
- A. S. Mohamed, N. X. Armendarez, M. S. Hasan and J. S. Najem, Memimpedance-Based Biomolecular Device for Adaptive Physical Reservoir Computing, Adv. Intell. Syst., 2025, 7(11), 2500281 CrossRef.
- Z. Li, S. K. Myers, J. Xiao, Y. Li, N. Noy, A. Leuski and A. Noy, Neuromorphic ionic computing in droplet interface synapses, Sci. Adv., 2025, 11(30), eadv6603 CrossRef CAS PubMed.
- S. A. Sarles, J. S. Najem and A. S. Mohamed, Voltage-responsive biomimetic membranes and ion channels for neuromorphic computing, npj Unconv. Comput., 2025, 2(1), 26 CrossRef.
- G. Paulo, K. Sun, G. Di Muccio, A. Gubbiotti, B. Morozzo Della Rocca, J. Geng, G. Maglia, M. Chinappi and A. Giacomello, Hydrophobically gated memristive nanopores for neuromorphic applications, Nat. Commun., 2023, 14, 8390 CrossRef CAS PubMed.
- S. F. Mayer, M. F. Mitsioni, P. Robin, L. VanDen Heuvel, N. Ronceray, M. J. Marcaida, L. A. Abriata, L. F. Krapp, J. S. Anton and S. Soussou, et al., Lumen charge governs gated ion transport in β-barrel nanopores, Nat. Nanotechnol., 2026, 21, 116–124 CrossRef CAS PubMed.
- B. K. Ziervogel and B. Roux, The Binding of Antibiotics in OmpF Porin, Structure, 2013, 21(1), 76–87 CrossRef CAS PubMed.
- S. W. Cowan, T. Schirmer, G. Rummel, M. Steiert, R. Ghosh, R. A. Pauptit, J. N. Jansonius and J. P. Rosenbusch, Crystal structures explain functional properties of two E. coli porins, Nature, 1992, 358(6389), 727–733 CrossRef CAS PubMed.
- K. M. Robertson and D. P. Tieleman, Molecular basis of voltage gating of OmpF porin, Biochem. Cell Biol., 2002, 80(5), 517–523 CrossRef CAS PubMed.
- D. Jeanteur, T. Schirmer, D. Fourel, V. Simonet, G. Rummel, C. Widmer, J. P. Rosenbusch, F. Pattus and J.-M. Pagès, Structural and Functional Alterations of a Colicin-Resistant Mutant of OmpF Porin from Escherichia coli, Proc. Natl. Acad. Sci. U. S. A., 1994, 91(22), 10675–10679 CrossRef CAS PubMed.
- A. H. Delcour, Outer membrane permeability and antibiotic resistance, Biochim. Biophys. Acta, Proteins Proteomics, 2009, 1794(5), 808–816 CrossRef CAS PubMed.
- E. Hajjar, A. Bessonov, A. Molitor, A. Kumar, K. R. Mahendran, M. Winterhalter, J.-M. Pagès, P. Ruggerone and M. Ceccarelli, Toward Screening for Antibiotics with Enhanced Permeation Properties through Bacterial Porins, Biochemistry, 2010, 49(32), 6928–6935 CrossRef CAS PubMed.
- I. Ghai, H. Bajaj, J. Arun Bafna, H. A. El Damrany Hussein, M. Winterhalter and R. Wagner, Ampicillin permeation across OmpF, the major outer-membrane channel in Escherichia coli, J. Biol. Chem., 2018, 293(18), 7030–7037 CrossRef CAS PubMed.
- E. M. Nestorovich, C. Danelon, M. Winterhalter and S. M. Bezrukov, Designed to penetrate: Time-resolved interaction of single antibiotic molecules with bacterial pores, Proc. Natl. Acad. Sci. U. S. A., 2002, 99(15), 9789–9794 CrossRef CAS PubMed.
- R. Iyer and A. H. Delcour, Complex Inhibition of OmpF and OmpC Bacterial Porins by Polyamines, J. Biol. Chem., 1997, 272(30), 18595–18601 CrossRef CAS PubMed.
- R. Iyer, Z. Wu, P. M. Woster and A. H. Delcour, Molecular basis for the polyamine-OmpF porin interactions: inhibitor and mutant studies, J. Mol. Biol., 2000, 297(4), 933–945 CrossRef CAS PubMed.
- U. Lamichhane, T. Islam, S. Prasad, H. Weingart, K. R. Mahendran and M. Winterhalter, Peptide translocation through the mesoscopic channel: binding kinetics at the single molecule level, Eur. Biophys. J., 2013, 42(5), 363–369 CrossRef CAS PubMed.
- J. Wang, J. A. Bafna, S. P. Bhamidimarri and M. Winterhalter, Small-Molecule Permeation across Membrane Channels: Chemical Modification to Quantify Transport across OmpF, Angew. Chem., Int. Ed., 2019, 58(14), 4737–4741 CrossRef PubMed.
- R. J. Vogler, K. E. Kimball, D. D. Bujanos, H. Behera, B. J. Pedretti, K. Grogan, N. A. Lynd, B. D. Freeman and M. Kumar, Rapid heat-driven formation of 2D nanosheets from membrane proteins and block copolymers, J. Membr. Sci., 2025, 720, 123732 CrossRef CAS.
- S. Jo, T. Kim, V. G. Iyer and W. Im, CHARMM-GUI: A web-based graphical user interface for CHARMM, J. Comput. Chem., 2008, 29(11), 1859–1865 CrossRef CAS PubMed.
- J. Gumbart, F. Khalili-Araghi, M. Sotomayor and B. Roux, Constant electric field simulations of the membrane potential illustrated with simple systems, Biochim. Biophys. Acta, Biomembr., 2012, 1818(2), 294–302 CrossRef CAS PubMed.
- B. Roux, The Membrane Potential and its Representation by a Constant Electric Field in Computer Simulations, Biophys. J., 2008, 95(9), 4205–4216 CrossRef CAS PubMed.
- C. Kutzner, H. Grubmüller, B. L. de Groot and U. Zachariae, Computational Electrophysiology: The Molecular Dynamics of Ion Channel Permeation and Selectivity in Atomistic Detail, Biophys. J., 2011, 101(4), 809–817 CrossRef CAS PubMed.
- F. Sofos, T. E. Karakasidis and D. Spetsiotis, Molecular dynamics simulations of ion separation in nano-channel water flows using an electric field, Mol. Simul., 2019, 45(17), 1395–1402 CrossRef CAS.
- B. Sun, Y. Chen, M. Xiao, G. Zhou, S. Ranjan, W. Hou, X. Zhu, Y. Zhao, S. A. T. Redfern and Y. N. Zhou, A Unified Capacitive-Coupled Memristive Model for the Nonpinched Current–Voltage Hysteresis Loop, Nano Lett., 2019, 19(9), 6461–6465 CrossRef CAS PubMed.
- A. L. Hodgkin and A. F. Huxley, A quantitative description of membrane current and its application to conduction and excitation in nerve, J. Physiol., 1952, 117(4), 500–544 CrossRef CAS PubMed.
- S. A. Ionescu, S. Lee, N. G. Housden, R. Kaminska, C. Kleanthous and H. Bayley, Orientation of the OmpF Porin in Planar Lipid Bilayers, ChemBioChem, 2017, 18(6), 554–562 CrossRef CAS PubMed.
- A. Aksimentiev and K. Schulten, Imaging α-Hemolysin with Molecular Dynamics: Ionic Conductance, Osmotic Permeability, and the Electrostatic Potential Map, Biophys. J., 2005, 88(6), 3745–3761 CrossRef CAS PubMed.
- Y. Li, R. Sun, H. Liu and H. Gong, Molecular dynamics study of ion transport through an open model of voltage-gated sodium channel, Biochim. Biophys. Acta, Biomembr., 2017, 1859(5), 879–887 CrossRef CAS PubMed.
- F. Khalili-Araghi, E. Tajkhorshid and K. Schulten, Dynamics of K+ Ion Conduction through Kv1.2, Biophys. J., 2006, 91(6), L72–L74 CrossRef CAS PubMed.
- C. Maffeo, S. Bhattacharya, J. Yoo, D. Wells and A. Aksimentiev, Modeling and Simulation of Ion Channels, Chem. Rev., 2012, 112(12), 6250–6284 CrossRef CAS PubMed.
- M. B. Ulmschneider, C. Bagnéris, E. C. McCusker, P. G. Decaen, M. Delling, D. E. Clapham, J. P. Ulmschneider and B. A. Wallace, Molecular dynamics of ion transport through the open conformation of a bacterial voltage-gated sodium channel, Proc. Natl. Acad. Sci. U. S. A., 2013, 110(16), 6364–6369 CrossRef CAS PubMed.
- A. Aksimentiev, Deciphering ionic current signatures of DNA transport through a nanopore, Nanoscale, 2010, 2(4), 468 RSC.
| This journal is © The Royal Society of Chemistry 2026 |